Abstract
RAS oncoproteins are molecular switches associated with critical signaling pathways that regulate cell proliferation and differentiation. Mutations in the RAS family, mainly in the KRAS isoform, are responsible for some of the deadliest cancers, which has made this protein a major target in biomedical research. Here we demonstrate that a designed bis-histidine peptide derived from the αH helix of the cofactor SOS1 binds to KRAS with high affinity upon coordination to Pd(II). NMR spectroscopy and MD studies demonstrate that Pd(II) has a nucleating effect that facilitates the access to the bioactive α-helical conformation. The binding can be suppressed by an external metal chelator and recovered again by the addition of more Pd(II), making this system the first switchable KRAS binder, and demonstrates that folding-upon-binding mechanisms can operate in metal-nucleated peptides. In vitro experiments show that the metallopeptide can efficiently internalize into living cells and inhibit the MAPK kinase cascade.
Similar content being viewed by others
Introduction
RAS proteins are oncogenic supervillains mutated in almost half of all cancers1. Many of these mutations drive some of the most lethal cancer types, including about 95% of pancreatic, 45% of colorectal cancers, and 33% of lung adenocarcinomas2. More specifically, KRAS, the most frequently mutated isoform, is responsible for most RAS cancers and generates highly aggressive and life-threatening phenotypes3. These features have made KRAS a key target in the search for new antitumor agents for over three decades4,5. Physiologically, RAS proteins are binary molecular switches that oscillate between an inactive, GDP-bound, state and an active RAS-GTP complex, aided by specific guanine nucleotide exchange factor proteins (GEFs), such as SOS1, which stimulate the GDP/GTP exchange and activation, and GTPase-activating proteins (GAPs) that catalyze the hydrolysis of GTP and inactivation in the RAS-GDP state. Certain mutations, such as KRASG12C or KRASG12V, compromise the GTPase activity of RAS proteins, locking them in their active state, thereby inducing the aberrant activation of several signaling pathways that ultimately lead to uncontrolled cell growth and proliferation, invasiveness, angiogenesis, and metastasis6,7. Activation of RAS-GDP by SOS1 is induced by the insertion of the αH helix of the exchange factor, which opens up the GTP binding pocket and reduces the affinity for the trinucleotide8. It has been shown, that conformationally restricted peptidomimetics of the SOS1 αH helix featuring helix-capping modifications9,10, hydrocarbon staples11, and SOS1 protein mimics12, bind KRAS with high affinity, and competitively inhibit RAS activation by SOS1.
In contrast to these constitutively active inhibitors, locked into stable α-helical conformations by permanent linkages, stimuli-responsive peptides that change their affinity in response to an external input could offer new opportunities in basic research, as well as for spatiotemporal controlled therapeutic applications. Thus, going beyond the classic metal-stabilized α-helices13,14,15,16, and considering our previous studies in the field of DNA binding17,18,19,20, we hypothesized that metal chelation could be applied for the development of stimuli-responsive KRAS peptide inhibitors21,22. Indeed, here we show that the peptide αH-His2, engineered to include two i,i + 4 His chelating residues in a short fragment of the SOS1 αH helix, binds KRAS only after addition of a Pd(II) source. We also show that such coordination is reversible, and that the metallopeptide αH-His2[Pd] is capable of inhibiting KRAS-activated pathways in live cells. Moreover, we also found that complete preorganization of the peptide α-helix is not required for efficient binding and inhibition, and that αH-His2[Pd] operates through a coupled folding-upon-binding mechanism, typical of Intrinsically Disordered Proteins.
Results
Design and synthesis of the metal-responsive KRAS peptide inhibitor
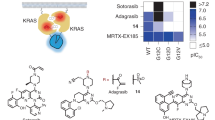
Based on the reported examples of SOS1 αH peptidomimetics11, as well as on the structural details of the SOS1 αH inserted between the switch regions of KRAS (PDB codes 1NVW and 1BKD)8,23, we chose the fragment comprising SOS1 residues Phe929 to Asn944, which spans the full αH helix for our experiments. Inspection of the hypothetical αH/RAS complex revealed that positions Tyr933 and Ile937 would be located in the solvent-exposed face of such αH helix fragment, and distanced so that their side chains, when replaced by His residues, would be in register and appropriately oriented to chelate a Pd(II) center (Fig. 1). In addition to the Tyr933His and Ile937His mutations, the αH sequence in our final design included two additional Arg residues in its N-terminus, for increased solubility and cellular uptake, and the sequence was capped with 6-carboxytetramethylrhodamine (TMR) for quantification and fluorescence monitoring (Fig. 1, αH-His2). The peptides were synthesized following standard Fmoc solid-phase peptide synthesis methods on an automated microwave synthesizer24,25.
a Detail of the αH helix of SOS1 complexed with RAS (PDB code 1BKD) indicating the residues Tyr933 and Ile937 that are replaced by the metal chelating His residues; the structure of SOS1 is shown in white. b Sequence of the natural αH helix of SOS1 and the mutant peptide αH-His2 used in this study, highlighting the two engineered His residues. c Conformational control by Pd(II) chelation. d Wenxiang diagram of the αH-His279,80, indicating the residues aligned in the outer face of the αH helix, including the two histidine residues in consecutive turns of the α-helix.
Addition of Pd(II) increases the α-helical content of the peptide αH-His2
Mixing the peptide αH-His2 with 1 equiv. of cis-PdCl2(en) gave the expected metallopeptide, αH-His2[Pd] from now on, as confirmed by NMR by following the chemical shift changes observed at the His ring protons, and by MS (see below and Fig. S2). Circular dichroism studies showed that metal coordination moderately increases the α-helical content of the peptide. Thus, a solution of αH-His2 displayed the typical CD signature of a random coil peptide and a low-intensity band at 222 nm (≈−2200 deg cm2 dmol−1); addition of one equivalent of cis-PdCl2(en) induced a modest increase in the negative intensity of this band (≈−5 700 deg cm2 dmol−1) that is consistent with a relatively small α-helical content of about 22% (Fig. 2, left; Fig. S3)26. Earlier experiments in our laboratory show that square planar cis-Pd(II) complexes display the ideal coordination properties to promote the effective folding of the helical conformation while other geometry (trans-[Pd(PPh3)2Cl2]), or other metal ions, such as Cu(II), Co(II), Ni(II), Fe(II), do not stabilize the active conformation22. Interestingly, the addition of a small excess of the Pd(II) chelator diethyldithiocarbamate (DEDTC), gave rise to a marked reduction in the helicity, supporting the key role of the metal in stabilizing the α-helical conformation. Subsequent addition of cis-PdCl2(en) led to a recovery of the helicity, thus confirming that the system can be easily switched between the metallo-stapled and the metal-free forms (Fig. 2, right).
a Circular dichroism of a 20 µM solution of αH-His2 (dashed line) and the spectrum 30 min after addition of 1 eq. of cis-PdCl2(en); b CD signal at 222 nm for: A 20 µM solution of αH-His2; B same solution after addition of Pd(II) and formation of the αH-His2[Pd] metallopeptide; C solution in B after addition of 5 eq. of DEDTC; D solution in C after addition of 20 eq. of cis-PdCl2(en). The experiments were carried out in 10 mM phosphate buffer pH 7.5, 100 mM of NaCl and 10% TFE, 25 °C. Mean residue molar ellipticity ([Θ]MRE) was calculated considering an 18-mer.
NMR spectroscopy demonstrates that the αH-His2 core sequence partially folds into an α-helical conformation upon metal chelation
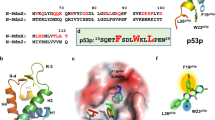
We characterized in detail the conformational effects of the Pd(II) coordination by NMR spectroscopy. All peptide residues were unambiguously assigned combining spin system identification in the TOCSY experiment and sequential assignment via NOEs (SI, pages S7-S11). In agreement with the CD studies, we found that the free peptide αH-His2 is mostly unstructured in solution, with a certain propensity to populate an α-helical conformation in the central part of the sequence, as deduced from the pattern of NH-NH NOEs observed for residues 929-940. The addition of cis-PdCl2(en) (up to 3 equiv.) led to a significant folding of the peptide chain, as deduced from several long-range NOEs between the side chains of Phe929, Ile932, and Leu934. These NOEs were absent in the free peptide and suggest that in addition to locking the α-helical conformation in the vicinity of the coordinating residues His933 and His937—from residues Gly931 to Leu938—metal chelation also promotes the formation of a N-terminal helical conformation from residues Arg928 to Phe930 (Fig. 3, Methods).
a Experimental NMR conformation in solution (lowest energy) in yellow; NOEs are indicated as dotted lines superimposed with a representative conformation of the MD simulation (in blue). b, c Aromatic-side chain regions of the NOESY spectra. b native peptide. c in the presence of cis-PdCl2(en). Residues participating in NOEs are labeled. In the left panel only intra-residue NOEs are detected whereas in the right panel, both intra and inter residue NOEs are observed, indicating the presence of a helical conformation upon Pd binding. PDB structures available as Supplementary Data 1.
Molecular modeling provides further insight into the conformational dynamics and metal coordination of the peptide αH-His2
To understand the conformational effect of the metal chelation in the α-helical conformation, we ran a series of 2 µs Gaussian accelerated molecular dynamics (GaMD) simulations of the peptide αH-His2 in its free form and coordinated with Pd(II). For the latter, the starting model was generated by performing protein-ligand docking of the Pd(II) metal center to the peptide using approaches updated for metal ligands27,28,29,30. The simulations show in all cases a largely disordered N-terminal region (from residues Arg927 to Ile932). In the case of the metal-free peptide, followed by an unstable C-terminal α-helix that undergoes multiple folding/unfolding cycles until, at the end of the simulation, the conformational sampling leads to a long-lasting ensemble of unfolded structures (Supplementary Information, Figs. S4a and S5a). This indicates that the peptide has some intrinsic helical propensity, albeit insufficient to reach a stable fold. This result is supported by the experimental evidence that points to a largely disordered state in this metal-free form. In contrast, the simulation of the metallopeptide αH-His2[Pd] with Nτ coordination in both His led to a more stable fold with substantial α-helical propensity throughout the 2 µs simulation. The core region from residues His933 to Glu941, where the coordination occurs, rarely deviates from a canonical α-helix and when it does, it adopts other organized secondary structures (e.g., 310 helix). No transition to a random coil is observed (Figs. S4b and S5b). A comparison between the MD most sampled conformation of the αH-His2[Pd] and the lowest energy NMR conformer in solution shows an excellent agreement (Fig. 3). An additional control MD run with an alternative coordination mode (Nτ-His933/Nπ-His937) showed a marginally stable helical fold with frequent deviations of the ideal α-helix geometry and even completely unfolded structures. Such poor structuring effect is clearly shown by the representative structure of the most populated sub-ensemble, which shows a kink in the α-helix at the coordination site (Figs. S4c and S5c). Altogether, the computational study provides an excellent match with the spectroscopic data, confirming the enhancement of α-helical conformation through dual coordination of the His residues through their Nτ atoms.
The metallopeptide αH-His2[Pd] binds KRASwt
The above studies demonstrate that coordination to the Pd(II) modulates the conformation of the peptide αH-His2 in solution, nucleating the formation of an incipient α-helix. We anticipated that this metal-induced folding might facilitate the interaction of the peptide with KRAS. Indeed, mixing a 15 nM solution of the unstructured αH-His2 in absence of Pd(II) with increasing concentrations of wild-type KRASwt did not produce any increase in the fluorescence anisotropy of the TMR fluorophore at 559 nm, indicating that the apopeptide, αH-His2, has low binding affinity for KRASwt. In contrast, when the titration was repeated with a preformed palladium/peptide complex, αH-His2[Pd], we observed a marked increase in the anisotropy, which is consistent with binding to KRASwt and formation of a larger complex with slower rotational diffusion. The increase in anisotropy could be fitted to a simple 1:1 binding model with an approximate dissociation constant of 240 nM (Fig. 4a). Similar titration profiles were obtained when αH-His2[Pd] was incubated with the oncogenic mutants KRASG12C and KRASG12V, which displayed also tight binding with dissociation constants of 345 nM and 294 nM, respectively (Fig. S6). Importantly, the addition of DEDTC to the solution of the complex αH-His2[Pd]/KRASwt18,31,32, resulted in a marked decrease of the TMR anisotropy, indicating the disassembly of the metallopeptide αH-His2[Pd], and disengagement of αH-His2 from KRASwt. Furthermore, the addition of fresh cis-PdCl2(en) to that solution restored the original anisotropy, indicating that the released peptide αH-His2 was still able to chelate Pd(II) metal center and to bind once again as αH-His2[Pd] to KRASwt (Fig. 4b), thereby confirming the reversibility of the system.
a Normalized emission anisotropy at 559 nm of a 15 nM solution of αH-His2[Pd] with increasing concentrations of KRASwt. The best fit to a 1:1 binding model is also shown43,44; b anisotropy values for: A, a saturated αH-His2[Pd]/KRAS complex; B, same solution after addition of 50 eq. of DEDTC; C, solution in B after addition of 50 eq. of cis-Cl2Pd(en). Data are mean ± SEM for experiments performed in technical triplicate and are representative of at least three biological replicates performed with independent preparations of recombinant KRASwt. All experiments carried out at 25 °C. Data were subjected to a One-Way ANOVA statistical analysis, ***p < 0.001.
Taken together, these data confirm that the palladium clip is working as a facilitator of the helix folding rather than as a traditional staple in which the covalent link fixes the bioactive α-helical conformation33. In other words, the coordination to the Pd(II) appears to provide just enough thermodynamic stabilization for the peptide to display folding-upon-binding properties such as those typically observed for Intrinsically Disordered Proteins34,35. A similar effect has been described for short trigger sequences that mediate the folding of coiled-coil peptides and allow the peptides to surpass a critical threshold of stability36,37. These results are also consistent with the region of the metal clip being highly structured, as shown by NMR and MD studies, despite the low total helical content of the metallopeptide as measured by CD.
The metallopeptide αH-His2[Pd] inhibits KRASwt GTP exchange
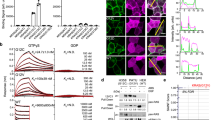
Having confirmed the high affinity of the complex αH-His2[Pd] for KRASwt, we studied its effect in the nucleotide release process associated with biological inactivation of this protein38. Therefore, we first measured the intrinsic nucleotide release rate by monitoring the fluorescence emission of the complex of KRASwt with the GTP analog mantGTP, 2′-/3′-O-(N-methylanthraniloyl) guanosine-5′-triphosphate, which displays low fluorescence in solution, but becomes strongly emissive when associated with KRASwt21,22. As expected, the KRASwt/mantGPT complex was very stable, and fluorescent mantGTP was released very slowly, with a half-life of about 7 min (Fig. 5). The half-life of this complex was somewhat decreased by the presence of the free peptide αH-His2 (Fig. 5 right, t1/2 ≈ 5 min), but the addition of the preformed metallopeptide αH-His2[Pd] drastically increased the rate of nucleotide release, so that the measured half-time of the KRASwt/mantGPT complex was reduced to about 3 min under the same conditions (Fig. 5, right).
Preformed αH-His2[Pd] increases the displacement of mantGTP bound to KRASwt. a normalized emission at 430 nm. Black triangles: intrinsic nucleotide displacement curve of a 1 μM solution of the preformed KRASwt/mantGTP complex; white circles, same solution in the presence of 10 μM αH-His2; black circles: same KRASwt/mantGTP solution in the presence of 10 μM αH-His2[Pd]. b half-life of the kinetic traces shown on the left. A intrinsic KRASwt/mantGTP complex; B with αH-His2; C with αH-His2[Pd]. Data were subjected to a One-Way ANOVA statistical analysis, ***p < 0.001.
The metallopeptide αH-His2[Pd] inhibits the ERK1/2 signaling pathway in cells
The coordinative basis of our stapled peptide raised the question of whether it could be used in the complex cellular environment. Thus, to assess its capacity to antagonize the KRAS activation in living mammalian cells, we first monitored its cellular uptake by fluorescence microscopy. Incubation of lung carcinoma A549 cells with αH-His2 revealed a rather poor internalization (Fig. 6a), but when this peptide was previously incubated with cis-PdCl2(en), the resulting metallopeptide, αH-His2[Pd], showed an intense intracellular fluorescence signal (Fig. 6b). Therefore, the palladium clip facilitates the internalization39. We then analyzed the activation of the MAPK RAF-MEK-ERK cascade, a well-documented KRAS effector pathway implicated in cell proliferation, survival, and differentiation (Fig. 6c)40. As expected, incubation of serum-starved A549 cells41 with either the apopeptide αH-His2 or with cis-PdCl2(en) failed to induce changes in the levels of ERK phosphorylation, as measured by blotting the cell lysates with pERK specific antibodies. In contrast, the addition of the preformed complex αH-His2[Pd] to the cell culture led to a significant decrease in the levels of phosphorylated ERK (Fig. 6d), thus demonstrating that the palladapeptide is capable of inhibiting ERK phosphorylation pathways in a dose-responsive manner.
a, b Effect of Pd(II) coordination in cellular internalization (A549 cells). a Incubation with αH-His2 (10 µM, 30 min at 37 °C); b incubation with the preformed αH-His2[Pd] complex, prepared by mixing an equimolar mixture of αH-His2 and cis-PdCl2(en) in water for 10 min (10 μM, 30 min at 37 °C). The cells were washed with PBS before being observed under the microscope. Incubations were made in Dulbecco´s modified Eagle medium (DMEM) completed with 5% of fetal bovine serum (FBS-DMEM). λexc = 550 nm, λem = 590-650 nm; c Scheme of the MAPK pathway and the proposed inhibition of the SOS1-mediated activation of KRAS by αH-His2[Pd] that should lead to reduced phosphorylation of ERK; d Addition of cis-PdCl2(en) to αH-His2 led to a palladium peptide complex that inhibits the RAF-MEK-ERK mitogen-activated protein kinase (MAPK) pathway downstream of KRAS in living cells, as deduced from the decrease in the amount of phosphorylated pERK. Serum-starved A549 cells were incubated either with αH-His2, the preformed metallopeptide αH-His2[Pd], or with cis-PdCl2(en) at the indicated doses for 4 h. Cellular lysates were subjected to Western blot analysis by using antibodies to phospho- and total ERK1/2. Densitometry measurement of the Western blot bands. The ratio between phospho-ERK1/2 and total ERK1/2 are indicated as fold change with respect to activation conditions (without FBS, lane 2). Activation percentage is shown above each column. Original WB and micrographies available as Supplementary Data 2.
Discussion
Peptidomimetics, particularly α-helix mimics, are promising platforms for the modulation of “undruggable” protein targets42. To date, the α-helical peptide inhibitors reported in the literature typically rely on the covalent linkages43,44,45, including carbon staples46,47,48, hydrogen bond surrogates49,50,51, lactam bridges52,53, or cysteine crosslinks52,53, among others54,55. These covalent modifications afford peptides that are constitutively active, but having at hand stimuli-responsive peptidomimetics, which can be (de)activated at will, would open interesting opportunities both in basic research and biomedicine. In this context, we envisioned the application of metal coordination to provide increased conformational stability and stimuli-responsive properties to a short α-helix peptidomimetic.
As model system, we chose the SOS1 αH helix/KRAS complex because of the intrinsic relevance of the KRAS GTPase in cancer as well as because there are clear precedents of short peptides derived from the αH helix that selectively bind to KRAS21,22. Furthermore, the available structural data of the SOS1/KRAS complex allowed us to rationally engineer the parent sequence to introduce the required residues for Pd(II) coordination in the exposed face of the αH helix in its complex with KRAS54,55,56. Thus, inspection of the reported structures of the SOS1/KRAS complexes suggested the replacement of Tyr933 and Ile937 with two His residues to create the metal-binding site33; these mutations, together with two additional Arg residues at the N-terminus ensured the solubility of the resulting peptide (RRFFGIHLTNHLKTEEGN). No further optimization of the sequence to increase its intrinsic α helical propensity was attempted. Indeed, the software AGADIR, accessible via http://agadir.crg.es/, predicts that the sequence of the αH-His2 peptide at 25 °C and pH 7 is only 12% helical56,57,58.
Having at hand the desired peptide, αH-His2, we used a combination of circular dichroism, NMR spectroscopy, and computational methods to study its coordination properties and the effect of Pd(II) binding in its folding. Both CD and NMR indicated that the free peptide αH-His2 was largely unstructured in solution and also showed a very modest stabilization of the α helical conformation in the presence of the metal ion. Thus, addition of Pd(II) results in a small increase in the negative molar ellipticity signal at 222 nm from ≈−2 200 to −5 700 deg cm2 dmol−1, and in the appearance of several long-range NOEs between the side chains of Phe929, Ile932, and Leu934. Molecular Dynamics studies confirm the Nτ coordination of both His residues, and the partial folding of the peptide from residues Arg928 to Phe930. These effects were reversible, and addition of a Pd(II) chelator reduced the helical content of the peptide.
Despite the modest effect of Pd(II) chelation in the overall helical content of the αH-His2 peptide, Pd(II) coordination led to a dramatic change in the affinity for KRAS, so that while no significant binding was observed by fluorescence anisotropy when the free peptide αH-His2 was incubated with KRAS, titrations with the metallopeptide αH-His2[Pd] resulted in clear binding isotherms with tight dissociation constants of 345 nM and 294 nM for KRASG12C and KRASG12V, respectively. This effect is consistent with an induced folding-upon-binding mechanism facilitated by the nucleation of the bioactive α helical conformation when the Pd(II) coordinates the two His residues. Therefore, although the coordination of the metal ion does not provide enough energy to fold the metallopeptide αH-His2[Pd] into an α-helix by itself, it is enough to do so in the presence of KRAS and the additional interactions with the protein that stabilize the bound helix. This behavior reproduces that of Intrinsically Disordered Proteins or regions of proteins, which are typically involved in protein-protein interactions regulating signaling pathways, that rely on their transition from a disordered to a folded state upon recognition of their physiological partners58. This mechanism allows high specificity, ultra-sensitivity, and switching behavior59, which might also be desirable properties in designed peptide inhibitors. As expected from the initial coordination tests, addition of a Pd(II) chelator disassembles the metallopeptide αH-His2[Pd] and takes apart the αH-His2[Pd]/KRAS complex. Importantly, the formation of the αH-His2[Pd]/KRAS complex has functional consequences, so that the inhibition of the GTPase activity of KRAS can be modulated in vitro by the presence of Pd(II).
Finally, the preformed metallopeptide αH-His2[Pd] was internalized by A549 cells, as demonstrated by fluorescence microscopy, and the peptide inhibited the MAPK RAF-MEK-ERK cascade in a dose-responsive manner, as demonstrated by the measured decrease in the levels of phosphorylated ERK. To our knowledge, this is the first demonstration of a designed metallopeptide that can modulate a signaling pathway in living cells.
In this proof-of-concept work we challenge the current paradigm for the design of α-helix peptidomimetics, which seeks to maximize the α-helical content to avoid the entropic penalty associated with the folding of the peptide into its bioactive helical conformation. We demonstrate that such drastic preorganization is not required and that marginally stable helices can act as effective inhibitors through a folding-upon-binding mechanism, just like the natural Intrinsically Disordered Proteins that exploit this process to modulate protein activity. Moreover, we also report, for the first time, a strategy to obtain a switchable α-helical peptidomimetics that modify their folding and binding affinity to their targets in response to an external stimulus, namely, the presence of a Pd(II) metal ion. We applied this strategy for the design of a short peptide derived from the αH helix of the cofactor SOS1 that binds the oncogenic protein KRAS with high affinity upon coordination to Pd(II). NMR and MD studies provide a detailed atomistic picture of the nucleating effect of the metal center and support a folding-upon binding mechanism typical of Intrinsically Disordered Proteins60. In consonance with the metal-induced activation, the metal-free peptide, αH-His2, is biologically inactive in cellular assays, but the metallopeptide αH-His2[Pd] induced a significant inhibitory effect in KRAS-promoted pathways. Our approach uncovers new possibilities for the design of dynamic α-helix peptidomimetics that exploit the folding-upon-binding mechanism and can be extended to other biological targets.
Methods
Peptide synthesis
The peptide αH-His2 was synthesized on a Liberty Blue Lite automatic microwave assisted peptide synthesizer from CEM Corporation, following the manufacturer’s recommended procedures. The synthesis was on a 0.1 mmol scale using a 0.5 mmol/g load H-Rink amide ChemMatrix resin. Amino acids were coupled in 5-fold excess using DIC (N,N'-Diisopropylcarbodiimide) as activator, Oxime as base, and DMF as solvent. Couplings were conducted for 4 min at 90 °C. Deprotection of the temporal Fmoc protecting group was performed with 20% piperidine in DMF for 1 min at 75 °C. TMR was manually coupled to the N-terminus of the sequence by incubation of the resin-bound peptide with 3 eq. of 5-carboxytetramethylrhodamine (0.15 mmol, 64.5 mg), 3 eq. of HATU, and 5 eq. of DIEA 0.2 M in DMF for 60 min. Cleavage/deprotection was done by treating the resin-bound peptide for 2 h with 900 μL TFA, 50 μL CH2Cl2, 25 μL H2O and 25 μL TIS (1 mL of cocktail/40 mg resin). The resin was filtered, and the filtrate was added onto ice-cold diethyl ether. After ~20 min, the precipitate was centrifuged and washed again with of ice-cold ether. The solid residue was dried under argon and redissolved in water/CH3CN (1:1) for HPLC purification on a semipreparative Agilent 1100 series LC using a Phenomenex Luna–C18 (250 × 10 mm) reverse-phase column and a linear gradient (5 to 75% B over 40 min, 4 mL/min.; A: H2O 0.1% TFA, B: CH3CN 0.1% TFA). Collected fractions with pure products were freeze-dried to afford the desired peptide. The peptide was analyzed by analytical UHPLC-MS with an Agilent 1200 series LC/MS using an SB C18 (1.8 μm, 2.1 × 50 mm) analytical column from Phenomenex with a linear gradient from 5 to 95% of solvent B for 20 min at a flow rate of 0.35 mL/min (A: water with 0.1% TFA, B: acetonitrile with 0.1% TFA). Compounds were detected by UV absorption at 222, 270, and 330 nm. Electrospray Ionization Mass Spectrometry (ESI/MS) was performed with an Agilent 6120 Quadrupole LC/MS model in positive scan mode using direct injection of the purified peptide solution into the MS detector.
Circular Dichroism
Circular Dichroism was measured on a Jasco-715 coupled with a thermostat Nestlab RTE-111 using a 2 mm Hellma cuvette at 25 °C and the following settings: Acquisition range, 300-195 nm; band width, 2.0 nm; resolution, 0.2 nm; accumulation, 5 scans; sensitivity, 10 mdeg; response time, 0.25 s; speed, 100 nm/min. The spectra are the average of 5 scans and were processed using the “smooth” macro in the program KaleidaGraph (v 3.5 by Synergy Software).
Fluorescence Anisotropy and switch experiments
Measurements were made with a Jobin-Yvon Fluoromax-3, (DataMax 2.20), coupled to a Wavelength Electronics LFI − 3751 temperature controller, and using a 1 mL Hellma micro cuvette. Settings: integration time, 2.0 s; excitation slit width, 5.0 nm; emission slit width, 20.0 nm; excitation wavelength, 559 nm; emission wavelength, 585 nm. Aliquots of a stock solution of protein were added onto 15 nM solutions of αH-His2 or αH-His2[Pd] in Tris-HCl buffer 20 mM, pH 7.5, 100 mM NaCl, and the anisotropy was recorded after each addition. Experimental data correspond to the mean of three independent titration experiments and are representative of at least three biological replicates performed with independent preparations of recombinant proteins. All experiments carried out at 25 °C. Before use the corresponding purified protein, the glycerol was removed using a Zeba Spin Desalting Columns (Thermo fisher) that had been equilibrated with anisotropy buffer (Tris-HCl buffer 20 mM, pH 7.5, 100 mM NaCl). For the switch experiment, after completing the titration at saturating concentrations of the protein, 50 eq. of DEDTC (relative to the metallopeptide) was added to the solution and the anisotropy was recorded. After that, 50 eq. of cis-PdCl2(en) was added (relative to the metallopeptide) and the anisotropy was recorded again.
High-resolution NMR
NMR experiments were recorded on a Bruker Avance III 600-MHz spectrometer (IRB Barcelona) equipped with a quadruple (1H, 13C, 15N, 31P) resonance cryogenic probe head and a z-pulse field gradient unit at 298 K using a 1 mM solution of αH-His2 peptide—in the presence or absence of cis-PdCl2(en)—1D proton spectra were recorded with a sweep width of 12000 Hz and 32 k data points. A total of 16 scans were accumulated with an acquisition time of 2.05 s. A Watergate w5 composite pulse was used to suppress the water signal. 1H 2D-TOCSY and NOESY experiments were acquired in 90% H2O/10% D2O and used to assign the spin systems corresponding to the peptide resonances61,62,63. For the 2D-NOESY experiments, mixing times of 300, 150, and 80 ms were acquired to minimize the impact of spin-diffusion in the assignments. Spin-locking fields of 8 kHz and 50 ms mixing time was used for the 2D-TOCSY experiments. All 2D spectral widths were 8000 Hz. The data size was 512 points in F1, indirect dimension, and 2048 points in F2, direct dimension. For each F1 value, 48 transients were accumulated in the NOESY and 32 in the TOCSY experiments respectively. Data were processed with a combination of exponential and shifted sine-bell window functions for each dimension followed by automated baseline- and phase correction using TopSpin 3.5 (© Bruker 2020). The 512 × 2k data matrices were zero-filled to 2k × 2k (NOESY and TOCSY). Structure Calculation: To prove that the primary structure (composition and connectivity) corresponds to the theoretical sequence, we have followed the sequence assignment strategy63. We identified the characteristic spin system of every residue in the sequence, using the 2D-TOCSY experiment, and every spin system was connected to the following one via NOEs observed from the side chain of a given residue (i) to the amide proton of the following residue (i + 1) as well as from the amide proton of (i) to the amide of (i + 1). The full spin analysis as well as the assignment of the NOEs were carried out manually using CARA software64,65. Distance restraints derived from the NOESY experiments were used for the NMR-based model building of the peptide in solution using unambiguously assigned peaks exclusively and the program CNS 1.2 (Crystallography and NMR system)66. The protocol consisted of an implicit water simulated-annealing of 120 structures using 8,000 cooling steps followed by an explicit water refinement of the calculated structures using all experimental restraints during 1200 steps. The Pd coordination was not explicitly included in the calculation. To display the metal coordination, we manually optimized the His rotamers to facilitate the coordination and the Pd was added to the final model displayed in Fig. 2.
Computational methods
Ligand-protein docking exploration with GOLD30, and GaudiMM28, was first performed to discern the possible coordination modes of the Pd(II) complex with the αH-His2 peptide. Then, the αH-His2[Pd] cores were parametrized with the Amber tool MCPB.py67, based on the bonded model approach. QM calculations were performed with the DFT formalism as implemented in Gaussian 09, combined with the hybrid functional B3LYP68. The basis set employed was 6-31 g(d,p) for C, H and N atoms, while the SDD basis set with f-polarization function was used for the metal ion, with a pseudo-potential for the core electrons69. All these calculations were combined with the Grimme’s D3 correction for dispersion70. The complexes were embedded in a solvent-polarizable dielectric continuum model (SMD, water)71. Once these QM calculations were obtained, the parameters were created for those atoms coordinating the Pd(II) ion through Seminario’s approach72. The point charges were computed based on the RESP protocol73. Once the parameters were obtained, the systems were set up with AMBER18 tleap to build the topology and the coordinates file. The force field used for the protein atoms was the ff14SB74, improved force field recommended for the simulation of peptides. For the remaining atoms, the GAFF force field was applied; while the parameters obtained from the MCPB.py calculation were used for the Pd(II) ion. Explicit water TIP3P solvent was also considered in the complexes75, introducing neutrality by the addition of three and one chloride ions for the metallic and free peptide, respectively. The systems were embedded into a cubic box containing between 2700 and 3000 water molecules, 10 Å from the protein to the edge of the box. The topology and the coordinates files achieved for each complex were then used for a first 10 ns classical Molecular Dynamics (MD) simulation. The coordinates of the last structure were used for the Gaussian accelerated Molecular Dynamics (GaMD) simulations76, which allows an extensive exploration of the conformational space. The AMBER ff14SB force field in an NVT ensemble was also used with SHAKE algorithm. A boost on both dihedral a total potential energy was applied, producing GaMDs of at least 2 μs for each system. To prove the correct exploration of the conformational space, PCA analysis were applied to the trajectories. Despite GaMD allows extensive conformational exploration, simulations were started from different putative folded geometries of the peptide to assure unbiased predictions to evaluate the progression of the system from the two extreme situations (data not shown).
Protein Expression and Purification
The constructs pDNA2.0 6H-TEV-KRAS, pDNA2.0 6H-TEV-KRAS G12V and pDNA2.0 6H-FLAG-TEV-KRAS G12C were purchased from Addgene. BL21 cells (Invitrogen) were transform with the constructs and grown in Luria broth (LB) to OD 600 0.7 and induced with 250 mM isopropyl β-D-1-thiogalactopyranoside (IPTG) for 16 h at 16 °C. Cells were pelleted and resuspended in lysis buffer (20 mM sodium phosphate, pH 8.0, 500 mM NaCl, 10 mM imidazole, 1 mM 2-mercaptoethanol (BME), 5% (vol/vol) glycerol). After centrifugation, the pellet was frozen until use. Then, protease inhibitor (EDTA free) and lysozyme (1 mg/mL) were added, and the pellet was sonicated for 6 min. (35 sec. on /10 sec. off). Protein was purified over an IMAC (immobilized metal affinity chromatography) following HisPur Ni-NTA protocols (Thermo Fisher). Protein was concentrated in a 10-kDa Amicon Ultra-15 (Millipore) aliquoted with 15% glycerol, and then flash-frozen and stored at −20 °C.
Nucleotide Release assays
Loading of fluorescent-labeled GDP (mantGTP) to KRASwt was conducted following previous reports8,77. Purified KRASwt was buffer-exchanged in Zeba Spin Desalting Columns (Thermo Fisher) in loading buffer (20 mM Tris-HCl [pH 7.5], 50 mM NaCl, 4 mM EDTA and 1 mM DTT). The resulted eluted KRASwt, after measuring their concentration, was incubated with 10-fold molar excess of mantGTP (Abcam) for 1.5 h at 20 °C in the dark. Reactions were supplemented with 10 mM MgCl2 and incubated for 1 h on ice. Free nucleotide was removed using a Zeba Spin Desalting Columns (Thermo Fisher) that had been equilibrated with the reaction buffer (20 mM Tris-HCl, pH 7.5, 50 mM NaCl, 1 mM MgCl2 and 1 mM DTT). The effect of peptides on the intrinsic rate of nucleotide release was monitored using the decrease in fluorescence with time as mantGTP dissociates from KRAS in a 100 μL reaction mixture (96-well plate) of 1 μM KRASwt/mantGPT complex and 10 μM of αH-His2 or αH-His2[Pd]. Fluorescence was excited at 370 nm and emission was monitored at 430 nm during 30 min, using a Tecan Infinite M Plex plate reader. The data were processed using the program GraphPad Prism.
Cell internalization and inhibition of ERK1/2
All steps were performed on a sterile clean bench Telstar AV-100 at rt. Solutions stored in a fridge were warmed beforehand in a water bath (37 °C). Unless otherwise specified, all incubations were performed in DMEM containing 5% of fetal bovine serum (FBS-DMEM). Cell Culture: A549 cell line was cultured in DMEM (Dulbecco´s modified Eagle´s medium), 5 mM glutamine, penicillin (100 units/mL) and streptomycin (100 units/mL), all from Invitrogen. Proliferating cultures were maintained in a 5% CO2 humidified incubator at 37 °C. For all the experiments, cells were seeded in the corresponding well at the indicated concentration 2 days before treatment. A549 cells were seeded on glass-bottom plates 48 h (150.000 cell/ml) before treatment. Culture medium was removed and DMEM containing 5% fetal bovine serum (FBS-DMEM) and αH-His2 or αH-His2[Pd] (10 µM) were added. The metallopeptide was made just before the addition to cells by pre-incubating with metal complexes (1:1) in water for 10 min. After 30 min of incubation with cells, these were washed twice with PBS and replaced with fresh FBS-DMEM to observe under the microscope with adequate filters. Digital pictures of the different samples were taken under identical conditions of gain and exposure. A549 cells were seeded on glass-bottom plates (150.000 cell/ml) 48 h before treatment. Culture medium was removed, the cells was serum-starved and αH-His2, αH-His2[Pd] or cis-PdCl2(en) were added at different concentrations and incubated for 4 h. After two washes with PBS, the cells were lysed in Laemmli buffer containing Tris pH 6,8 1 M, Glycerol, SDS, Bromophenol Blue, and β-mercaptoethanol, and heated at 95 °C for 10 min. The samples were separated by SDS-12.5% polyacrylamide gel and transferred to nitrocellulose membrane, to probed by Western blotting. Levels of total ERK2 and phosphorylated ERK were detected with anti-ERK1/2 and phospho-ERK1/2) antibodies, respectively and revealed with LuminataTM Classico Western HRP substrate following manufacter´s protocol. The visualization was performed with ChemiDoc MP imaging system by Biorad, and the processing of the images and densitometry quantification was carried out with ImageLab program.
Fluorescence microscopy
All images were obtained with an Andor Zyla mounted on a Nikon TiE. LED excitation wavelength 550 nm. Filter cube TRITC-B-000 (Semrock): BP 543/22 nm, LP 593/40 nm, and DM 562 nm. Images were further processed with Image J.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The PDB structures of the NMR and Molecular Dynamics studies are available as Supplementary Data 1; the original WB and microscope images are available as Supplementary Data 2. All other datasets generated and analyzed during the current study are available from the corresponding authors on reasonable request.
References
Sanchez-Vega, F. et al. Oncogenic signaling pathways in the cancer genome Atlas. Cell 173, 321–337.e10 (2018).
Forbes, S. A. et al. COSMIC: exploring the world’s knowledge of somatic mutations in human cancer. Nucleic Acids Res. 43, D805–D811 (2015).
Hobbs, G. A., Der, C. J. & Rossman, K. L. RAS isoforms and mutations in cancer at a glance. J. Cell Sci. 129, 1287–1292 (2016).
Ledford, H. Cancer: the Ras renaissance. Nature 520, 278–280 (2015).
Spiegel, J., Cromm, P. M., Zimmermann, G., Grossmann, T. N. & Waldmann, H. Small-molecule modulation of Ras signaling. Nat. Chem. Biol. 10, 613–622 (2014).
Wittinghofer, A. & Waldmann, H. Ras—a molecular switch involved in tumor formation. Angew. Chem. Int. Ed. 39, 4192–4214 (2000).
Rajalingam, K., Schreck, R., Rapp, U. R. & Albert, S. Ras oncogenes and their downstream targets. Biochim. Biophys. Acta 1773, 1177–1195 (2007).
Margarit, S. M. et al. Structural evidence for feedback activation by Ras.GTP of the Ras-specific nucleotide exchange factor SOS. Cell 112, 685–695 (2003).
Patgiri, A., Yadav, K. K., Arora, P. S. & Bar-Sagi, D. An orthosteric inhibitor of the Ras-Sos interaction. Nat. Chem. Biol. 7, 585–587 (2011).
Yoo, D. Y., Hauser, A. D., Joy, S. T., Bar-Sagi, D. & Arora, P. S. Covalent targeting of Ras G12C by rationally designed peptidomimetics. ACS Chem. Biol. 15, 1604–1612 (2020).
Leshchiner, E. S. et al. Direct inhibition of oncogenic KRAS by hydrocarbon-stapled SOS1 helices. Proc. Natl Acad. Sci. USA 112, 1761–1766 (2015).
Hong, S. H. et al. A Sos proteomimetic as a pan-Ras inhibitor. Proc. Natl. Acad. Sci. USA 118, e2101027118 (2021).
Ghadiri, M. R. & Fernholz, A. K. Peptide architecture. Design of stable α-helical metallopeptides via a novel exchange-inert ruthenium(III) complex. J. Am. Chem. Soc. 112, 9633–9635 (1990).
Ghadiri, M. R. & Choi, C. Secondary structure nucleation in peptides. Transition metal ion stabilized α-helices. J. Am. Chem. Soc. 112, 1630–1632 (1990).
Kelso, M. J., Hoang, H. N., Appleton, T. G. & Fairlie, D. P. The first solution stucture of a single α-helical turn. A pentapeptide α-helix stabilized by a metal clip. J. Am. Chem. Soc. 122, 10488–10489 (2000).
Ma, M. T., Hoang, H. N., Scully, C. C. G., Appleton, T. G. & Fairlie, D. P. Metal clips that induce unstructured pentapeptides to be α-helical in water. J. Am. Chem. Soc. 131, 4505–4512 (2009).
Rodríguez, J., Mosquera, J., Vázquez, M. E. & Mascareñas, J. L. Nickel-promoted recognition of long DNA sites by designed peptide derivatives. Chemistry 22, 13474–13477 (2016).
Rodríguez, J., Mosquera, J., García-Fandiño, R., Vázquez, M. E. & Mascareñas, J. L. A designed DNA binding motif that recognizes extended sites and spans two adjacent major grooves. Chem. Sci. 7, 3298–3303 (2016).
Learte-Aymamí, S., Rodríguez, J., Vázquez, M. E. & Mascareñas, J. L. Assembly of a ternary metallopeptide complex at specific DNA sites mediated by an AT-hook adaptor. Chemistry 26, 8875–8878 (2020).
Gómez-González, J. et al. Dynamic stereoselection of peptide helicates and their selective labeling of DNA replication foci in cells*. Angew. Chem. Int. Ed. Engl. 60, 8859–8866 (2021).
Learte-Aymamí, S., Vidal, C., Gutiérrez-González, A. & Mascareñas, J. L. Intracellular reactions promoted by Bis(histidine) miniproteins stapled using palladium(II) complexes. Angew. Chem. Int. Ed. Engl. 59, 9149–9154 (2020).
Learte-Aymamí, S., Curado, N., Rodríguez, J., Vázquez, M. E. & Mascareñas, J. L. Metal-dependent DNA recognition and cell internalization of designed, basic peptides. J. Am. Chem. Soc. 139, 16188–16193 (2017).
Boriack-Sjodin, P. A., Margarit, S. M., Bar-Sagi, D. & Kuriyan, J. The structural basis of the activation of Ras by Sos. Nature 394, 337–343 (1998).
Coin, I., Beyermann, M. & Bienert, M. Solid-phase peptide synthesis: from standard procedures to the synthesis of difficult sequences. Nat. Protoc. 2, 3247–3256 (2007).
Vanier, G. S. Microwave-assisted solid-phase peptide synthesis based on the Fmoc protecting group strategy (CEM). Methods Mol. Biol. 1047, 235–249 (2013).
Shepherd, N. E., Hoang, H. N., Abbenante, G. & Fairlie, D. P. Single turn peptide α helices with exceptional stability in water. J. Am. Chem. Soc. 127, 2974–2983 (2005).
Jones, G., Willett, P., Glen, R. C., Leach, A. R. & Taylor, R. Development and validation of a genetic algorithm for flexible docking. J. Mol. Biol. 267, 727–748 (1997).
Rodríguez-Guerra Pedregal, J., Sciortino, G., Guasp, J., Municoy, M. & Maréchal, J.-D. GaudiMM: A modular multi-objective platform for molecular modeling. J. Comput. Chem. 38, 2118–2126 (2017).
Di Meo, T. et al. Functionalized artificial bidomain proteins based on an α-solenoid protein repeat scaffold: a new class of artificial diels–alderases. ACS Omega 4, 4437–4447 (2019).
Sciortino, G., Garribba, E. & Maréchal, J.-D. Validation and applications of protein-ligand docking approaches improved for metalloligands with multiple vacant sites. Inorg. Chem. 58, 294–306 (2019).
Kuzmic, P. Program DYNAFIT for the analysis of enzyme kinetic data: application to HIV proteinase. Anal. Biochem. 237, 260–273 (1996).
Kuzmič, P. In Methods in Enzymology Vol. 467 (ed. Abelson, J.) 247–280 (Academic Press, 2009).
Toto, A. et al. Templated folding of intrinsically disordered proteins. J. Biol. Chem. 295, 6586–6593 (2020).
Kammerer, R. A. et al. An autonomous folding unit mediates the assembly of two-stranded coiled coils. Proc. Natl Acad. Sci. USA 95, 13419–13424 (1998).
Lee, D. L., Lavigne, P. & Hodges, R. S. Are trigger sequences essential in the folding of two-stranded alpha-helical coiled-coils? J. Mol. Biol. 306, 539–553 (2001).
Rojas, R. J., Kimple, R. J., Rossman, K. L., Siderovski, D. P. & Sondek, J. Established and emerging fluorescence-based assays for G-protein function: Ras-superfamily GTPases. Comb. Chem. High. Throughput Screen. 6, 409–418 (2003).
Jeganathan, S., Müller, M. P., Ali, I. & Goody, R. S. Assays for nucleotide competitive reversible and irreversible inhibitors of Ras GTPases. Biochemistry 57, 4690–4699 (2018).
Hiratsuka, T. New ribose-modified fluorescent analogs of adenine and guanine nucleotides available as substrates for various enzymes. Biochim. Biophys. Acta 742, 496–508 (1983).
Roberts, P. J. & Der, C. J. Targeting the Raf-MEK-ERK mitogen-activated protein kinase cascade for the treatment of cancer. Oncogene 26, 3291–3310 (2007).
Jung, Y. D., Nakano, K., Liu, W., Gallick, G. E. & Ellis, L. M. Extracellular signal-regulated kinase activation is required for up-regulation of vascular endothelial growth factor by serum starvation in human colon carcinoma cells. Cancer Res. 59, 4804–4807 (1999).
Miller, M. The importance of being flexible: the case of basic region leucine zipper transcriptional regulators. Curr. Protein Pept. Sci. 10, 244–269 (2009).
de Araujo, A. D. et al. Comparative α-helicity of cyclic pentapeptides in water. Angew. Chem. Int. Ed. Engl. 53, 6965–6969 (2014).
Bernal, F., Tyler, A. F., Korsmeyer, S. J., Walensky, L. D. & Verdine, G. L. Reactivation of the p53 tumor suppressor pathway by a stapled p53 peptide. J. Am. Chem. Soc. 129, 2456–2457 (2007).
Cromm, P. M., Spiegel, J. & Grossmann, T. N. Hydrocarbon stapled peptides as modulators of biological function. ACS Chem. Biol. 10, 1362–1375 (2015).
Walensky, L. D. & Bird, G. H. Hydrocarbon-stapled peptides: principles, practice, and progress. J. Med. Chem. 57, 6275–6288 (2014).
Chapman, R. N., Dimartino, G. & Arora, P. S. A highly stable short alpha-helix constrained by a main-chain hydrogen-bond surrogate. J. Am. Chem. Soc. 126, 12252–12253 (2004).
Patgiri, A., Jochim, A. L. & Arora, P. S. A hydrogen bond surrogate approach for stabilization of short peptide sequences in α-helical conformation. Acc. Chem. Res. 41, 1289–1300 (2008).
Patgiri, A., Menzenski, M. Z., Mahon, A. B. & Arora, P. S. Solid-phase synthesis of short α-helices stabilized by the hydrogen bond surrogate approach. Nat. Protoc. 5, 1857–1865 (2010).
Harrison, R. S. et al. Downsizing human, bacterial, and viral proteins to short water-stable alpha helices that maintain biological potency. Proc. Natl Acad. Sci. USA 107, 11686–11691 (2010).
Hoang, H. N. et al. A novel long-range n to π* interaction secures the smallest known α-helix in water. Angew. Chem. Int. Ed. Engl. 58, 18873–18877 (2019).
Shepherd, N. E., Abbenante, G. & Fairlie, D. P. Consecutive cyclic pentapeptide modules form short α-helices that are very stable to water and denaturants. Angew. Chem. Int. Ed. 43, 2687–2690 (2004).
Fairlie, D. P. & Dantas de Araujo, A. Review stapling peptides using cysteine crosslinking. Biopolymers 106, 843–852 (2016).
Jackson, D. Y., King, D. S., Chmielewski, J., Singh, S. & Schultz, P. G. General approach to the synthesis of short α-helical peptides. J. Am. Chem. Soc. 113, 9391–9392 (1991).
Horne, W. S. & Grossmann, T. N. Proteomimetics as protein-inspired scaffolds with defined tertiary folding patterns. Nat. Chem. 12, 331–337 (2020).
Rezaei Araghi, R. & Keating, A. E. Designing helical peptide inhibitors of protein-protein interactions. Curr. Opin. Struct. Biol. 39, 27–38 (2016).
Muñoz, V. & Serrano, L. Elucidating the folding problem of helical peptides using empirical parameters. II. Helix macrodipole effects and rational modification of the helical content of natural peptides. J. Mol. Biol. 245, 275–296 (1995).
Muñoz, V. & Serrano, L. Elucidating the folding problem of helical peptides using empirical parameters. III. Temperature and pH dependence. J. Mol. Biol. 245, 297–308 (1995).
Muñoz, V. & Serrano, L. Elucidating the folding problem of helical peptides using empirical parameters. Nat. Struct. Biol. 1, 399–409 (1994).
Teilum, K., Olsen, J. G. & Kragelund, B. B. On the specificity of protein–protein interactions in the context of disorder. Biochem. J. 478, 2035–2050 (2021).
Tsomaia, N. Peptide therapeutics: targeting the undruggable space. Eur. J. Med. Chem. 94, 459–470 (2015).
Bax, A. & Davis, D. G. Practical aspects of two-dimensional transverse NOE spectroscopy. J. Magn. Reson. 63, 207–213 (1985).
Macura, S. & Ernst, R. R. Elucidation of cross relaxation in liquids by two-dimensional N.M.R. spectroscopy. Mol. Phys. 41, 95–117 (1980).
Wuthrich, K. NMR of Proteins and Nucleic Acids. (John Wiley & Sons, 1986).
Ramirez-Espain, X., Ruiz, L., Martin-Malpartida, P., Oschkinat, H. & Macias, M. J. Structural characterization of a new binding motif and a novel binding mode in group 2 WW domains. J. Mol. Biol. 373, 1255–1268 (2007).
Martín-Gago, P. et al. A tetradecapeptide somatostatin dicarba-analog: synthesis, structural impact and biological activity. Bioorg. Med. Chem. Lett. 24, 103–107 (2014).
Brunger, A. T. Version 1.2 of the crystallography and NMR system. Nat. Protoc. 2, 2728–2733 (2007).
Li, P. & Merz, K. M. Jr MCPB.py: A python based metal center parameter builder. J. Chem. Inf. Model. 56, 599–604 (2016).
Becke, A. D. Density‐functional thermochemistry. III. The role of exact exchange. J. Chem. Phys. 98, 5648–5652 (1993).
Bergner, A., Dolg, M., Küchle, W., Stoll, H. & Preuß, H. Ab initio energy-adjusted pseudopotentials for elements of groups 13–17. Mol. Phys. 80, 1431–1441 (1993).
Grimme, S., Antony, J., Ehrlich, S. & Krieg, H. A consistent and accurate ab initio parametrization of density functional dispersion correction (DFT-D) for the 94 elements H-Pu. J. Chem. Phys. 132, 154104 (2010).
Marenich, A. V., Cramer, C. J. & Truhlar, D. G. Universal solvation model based on solute electron density and on a continuum model of the solvent defined by the bulk dielectric constant and atomic surface tensions. J. Phys. Chem. B 113, 6378–6396 (2009).
Seminario, J. M. Calculation of intramolecular force fields from second-derivative tensors. Int. J. Quantum Chem. 60, 1271–1277 (1996).
Bayly, C. I., Cieplak, P., Cornell, W. & Kollman, P. A. A well-behaved electrostatic potential based method using charge restraints for deriving atomic charges: the RESP model. J. Phys. Chem. 97, 10269–10280 (1993).
Maier, J. A. et al. ff14SB: improving the accuracy of protein side chain and backbone parameters from ff99SB. J. Chem. Theory Comput. 11, 3696–3713 (2015).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Miao, Y. et al. Improved reweighting of accelerated molecular dynamics simulations for free energy calculation. J. Chem. Theory Comput. 10, 2677–2689 (2014).
McCarthy, M. J. et al. Discovery of high-affinity noncovalent allosteric KRAS inhibitors that disrupt effector binding. ACS Omega 4, 2921–2930 (2019).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Chou, K. C., Zhang, C. T. & Maggiora, G. M. Disposition of amphiphilic helices in heteropolar environments. Proteins 28, 99–108 (1997).
Chou, K.-C., Lin, W.-Z. & Xiao, X. Wenxiang: a web-server for drawing wenxiang diagrams. Nat. Sci. (Irvine) 03, 862–865 (2011).
Acknowledgements
Financial support from the Spanish grants RTI2018-099877-B-I00, CTQ2015-70698-R, Orfeo-cinqa network CTQ2016-81797-REDC, Red the Péptidos en Biomedicina y Nanociencia, RED2018-102417-T; Centro Singular de Investigación de Galicia accreditation 2019-2022, ED431G 2019/03 and the European Union (European Regional Development Fund - ERDF) and AGAUR (SGR-50); the European Research Council (Advanced Grant No. 340055), are gratefully acknowledged. S. Learte-Aymamí thanks the MINECO for her Ph.D. fellowship. G. Sciortino thanks the Spanish MICINN Juan de la Cierva program, FJC2019-039135-I. We also acknowledge institutional funding from MINECO through the Centers of Excellence Severo Ochoa Award given to IRB Barcelona, as well as from the CERCA Program of the Generalitat de Catalunya. M.J.M. is an ICREA Programme Investigator. Molecular graphics performed with UCSF Chimera, developed by the Resource for Biocomputing, Visualization, and Informatics at the UCSF, with support from NIH P41-GM10331178. We also want to thank Dr. Huy Hoang and Prof. David P. Fairlie, from The University of Queensland, Brisbane, for sharing the NMR structure of the peptide Ac-[KAAAD]-NH2 used in our preliminary modeling. Finally, we also thank R. Menaya-Vargas for her technical assistance with cell experiments.
Author information
Authors and Affiliations
Contributions
Conceptualization, M.E.V. and J.L.M; methodology, J.R.C., J.-D.M., M.J.M., J.L.M., and M.E.V.; Validation, S.L.A., P.M.M., L.R.M., G.S., J.R.C., J.-D.M., M.J.M., J.L.M., and M.E.V.; formal analysis, S.L.A., P.M.M., L.R.M., G.S., J.R.C., J.-D.M, and M.J.M.; investigation, S.L.A., P.M.M., M.J.M., L.R.M. and G.S.; resources, M.E.V., J.L.M., M.J.M. and J.-D.M.; data curation, S.L.A., P.M.M., L.R.M., G.S. and J.R.C.; writing—original draft, M.E.V., J.L.M., M.J.M. and J.-D.M.; writing—review and editing, S.L.A., P.M.M., L.R.M., G.S., J.R.C., J.-D.M., M.J.M., J.L.M., and M.E.V.; visualization, J.R.C., J.-D.M, M.J.M., J.L.M., and M.E.V.; supervision, J.R.C., J.-D.M., M.J.M., J.L.M., and M.E.V.; project administration, M.E.V.; funding acquisition, M.E.V., J.L.M., M.J.M. and J.-D.M. Author Contributions are detailed as Supplementary Note 1 in the Supplementary Information file.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Chemistry thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Learte-Aymamí, S., Martin-Malpartida, P., Roldán-Martín, L. et al. Controlling oncogenic KRAS signaling pathways with a Palladium-responsive peptide. Commun Chem 5, 75 (2022). https://doi.org/10.1038/s42004-022-00691-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42004-022-00691-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.