Abstract
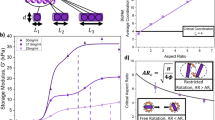
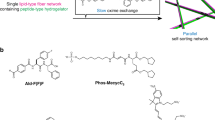
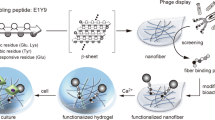
Glycosylation alters protein form and function by establishing intermolecular forces that mediate specific interactions while preventing non-specific aggregation. Self-assembled peptide nanofibers modified with carbohydrates are increasingly used as biomaterials to mimic glycosylated protein function, yet the influence of carbohydrate conjugates on nanofiber structure remains poorly defined. Here we show that a dense carbohydrate surface layer can facilitate hierarchical organization of peptide nanofibers into anisotropic networks. Glycosylated peptide nanofibers remain dispersed in dilute conditions, whereas non-glycosylated nanofibers tend to aggregate. In crowded conditions, some glycosylated nanofibers laterally associate and align. This behavior depends on carbohydrate chemistry, particularly hydroxyls, suggesting involvement of short-range attractive forces. Macroscopic gels fabricated from densely glycosylated peptide nanofibers are resistant to non-specific interactions with proteins, mammalian cells, and bacteria, yet selectively bind lectins, analogous to natural low-fouling mucosal barriers. Collectively, these observations demonstrate that glycosylation can inform structure in addition to endowing function to peptide-based supramolecular biomaterials.
Similar content being viewed by others
Introduction
Sequential self-assembly of proteins over multiple length scales underlies formation of countless structural and functional biomaterials throughout living systems. For example, the cytoskeleton is a dynamic network of anisotropic protein filaments that rearranges to give cells their shape, facilitate their movement, direct intracellular traffic, and regulate signal transduction1. Likewise, organization of proteins and proteoglycans into the extracellular matrices of multicellular organisms guides all aspects of cell physiology by engaging cell surface receptors, non-covalently regulating the transport of extracellular signals, and providing physical cues2. Inspired by these observations, synthetic supramolecular biomaterials assembled from peptides and peptide analogs are becoming increasingly more common3. However, encoding hierarchical self-assembly into these systems remains a challenge, despite established sequence-structure guidelines that have informed design of hundreds of peptide-based molecules that can form fibers with nano-scale features (i.e., “nanofibers”)4.
Achieving hierarchical order requires self-assembly pathways that minimize erroneous kinetically trapped structures5. In natural fibrillar protein networks, this is facilitated by fine balances of attractive and repulsive forces that establish checkpoints in the free energy landscape. For example, collagen fibril assembly is activated by enzymatic removal of globular N- and C-terminal propeptides from pro-collagen triple helices6, while microtubule assembly is regulated by GTP/GDP binding to β-tubulin7. In contrast, synthetic peptides and their analogs tend to assemble into nanofibers that subsequently aggregate into random entangled networks because non-specific inter-fiber attractive forces outweigh repulsive forces8. Manipulating system temperature or pH can promote hierarchical order among synthetic nanofibers by balancing intermolecular forces9,10,11. Likewise, shear flow, magnetic field, and ultrasound can be applied to induce peptide nanofiber alignment12,13,14,15. Alternatively, amino acid sequences can be tailored to promote nanofiber lateral association16,17, although these examples have been limited to electrostatic and aromatic–aromatic interactions involving the few natural charged or hydrophobic amino acids, respectively. We envision that moving beyond the limited range of molecular motifs provided by amino acids will provide new opportunities to encode sequential self-assembly within systems of peptide-based nanofibers, leading to supramolecular biomaterials demonstrating hierarchical order over multiple length scales.
Posttranslational modification of proteins with carbohydrates (i.e., “glycosylation”) can stabilize folded conformations, prevent aggregation, and establish selective molecular recognition by way of attractive and repulsive forces that promote specific intermolecular interactions while limiting non-specific aggregation18,19. Notably, glycosylation facilitates hierarchical assembly of various natural protein-based structures, including acetylcholine receptors, flagella, and mucinous liquid crystals20,21,22. Peptide-based nanofibers modified with carbohydrates are finding increasing use as biomaterials for various applications, including growth factor delivery, lectin recognition, cell binding, and tissue regeneration23,24,25,26,27,28,29,30. Yet, the influence of glycosylation on synthetic peptide self-assembly has received only limited attention focused primarily on hydrogel formation, nanofiber morphology, and nanofiber solubility31,32.
Here we demonstrate that a dense carbohydrate surface layer facilitates hierarchical organization of β-sheet peptide nanofibers into anisotropic networks that are aligned over multiple length scales. Further, these anisotropic networks resist non-specific cell, bacteria, and protein interactions, yet selectively recognize lectins, due to the collective activity of carbohydrates assembled into a highly ordered supramolecular architecture.
Results
Glycosylated peptide nanofiber alignment in the gel state
QQKFQFQFEQQ (“Q11”) is a synthetic peptide that self-assembles into β-sheet nanofibers in aqueous media, which entangle into self-supporting gels at high (~mM) concentrations33. We observed that a variant of Q11 terminated with asparagine-linked N-acetylglucosamine (“GQ11”, Fig. 1a) formed birefringent self-supporting gels at mM concentrations (Fig. 1b–d), which consisted of nanofibers that were aligned over several microns (Fig. 1e, f and Supplementary Fig. 1). GQ11 gels produced a highly anisotropic two-dimensional (2D) small-angle X-ray scattering (SAXS) pattern (Fig. 1g). Employing an established fitting algorithm34, the nematic order parameter, S, was found to be 0.83, indicating a high degree of fiber alignment across the 1 mm beam diameter. The characteristic fiber–fiber correlation length within GQ11 gels estimated from the reciprocal of the half-width at half maximum, Γ, was ~5 nm (Fig. 1h). This suggested that GQ11 β-sheet nanofibers aligned via lateral association rather than entwining, where the latter is the more common morphology for hierarchically ordered β-sheet peptide nanofibers35,36,37.
Glycosylation facilitates organization of β-sheet peptide nanofibers into anisotropic networks. a Chemical structure of GQ11 and b self-assembly of GQ11 into gels of aligned β-sheet nanofibers. GQ11 forms c self-supporting hydrogels that were d birefringent when viewed between cross-polarizers. e, f GQ11 hydrogels consisted of directionally oriented nanofibers when viewed using field-emission scanning electron microscopy (scale bar = 1 μm in e and 0.5 μm in f). g Small-angle X-ray scattering two-dimensional diffraction pattern and h radial intensity profile suggested anisotropic nanofiber orientation within GQ11 gels
Glycosylated peptide nanofiber alignment in crowded systems
To gain mechanistic understanding of GQ11 alignment, we compared nanofiber morphology at sub-gelling (~μM) concentrations under dilute and crowded conditions. Non-adsorbing macromolecular crowders establish depletion forces that increase effective concentration38, which were used here to mimic the environment of GQ11 nanofibers in the concentrated gel state. GQ11 nanofibers under dilute conditions were dispersed and randomly oriented, with no obvious aggregation or lateral association (Fig. 2a, b and Supplementary Fig. 2a). In contrast, GQ11 nanofibers formed aligned bundles in the presence of Ficoll and poly(ethylene glycol) (PEG) (Fig. 2c, d and Supplementary Fig. 2b–e). GQ11 nanofibers did not laterally associate or align in the presence of sucrose, the carbohydrate monomer unit of Ficoll (Supplementary Fig. 2f), suggesting that the observed morphological changes were due to depletion forces induced by the presence of a macromolecule in solution. Although kinetics of nanofiber lateral association and alignment were not investigated explicitly, all transmission electron microscopy (TEM) samples were prepared within 5 min of introducing the crowding agent, suggesting that formation of inter-fiber contacts was rapid. The drying required to visualize nanofibers with TEM could induce structural artifacts, such as lateral association and alignment, which may not be observed in solution. To characterize the morphology of GQ11 nanofibers in the presence of a crowder in native conditions, we visualized samples with cryo-electron microscopy (cryo-EM) (Fig. 2e and Supplementary Fig. 3). Similar to samples visualized with TEM, GQ11 nanofibers were laterally associated and highly aligned when viewed with cryo-EM in the presence of Ficoll, demonstrating that the alignment observed in samples viewed using TEM was not a drying artifact. Interestingly, though, aligned GQ11 nanofibers viewed with cryo-EM in the presence of Ficoll appeared twisted, as indicated by periodically alternating light and dark regions (Fig. 2e), whereas aligned GQ11 nanofibers viewed after drying with TEM appeared straighter (Fig. 2c, d). This suggested that drying may have an effect on the morphology of individual nanofibers.
GQ11 nanofibers laterally associate and align in crowded conditions. GQ11 nanofibers were dispersed when viewed using transmission electron microscopy in a water and b phosphate-buffered saline (PBS). GQ11 nanofibers were aligned in the presence of Ficoll when viewed using c, d transmission electron microscopy and e cryo-electron microscopy. f GQ11 nanofibers selectively bound wheat germ agglutinin (WGA), a GlcNAc-binding lectin, but not Concanavalin A (ConA) or bovine serum albumin (BSA), suggesting carbohydrates were on the nanofiber surface and accessible to the surrounding aqueous environment. Data presented as mean ± standard deviation (n = 3). Inset: Co-localization of GQ11 nanofibers and WGA-coated gold beads (scale bar = 50 nm). GQ11 nanofibers were also aligned in the presence of g BSA, h, i ConA, and j trifluoroethanol (TFE) when viewed using TEM (scale bar = 100 nm)
To determine whether GlcNAc groups were accessible on the nanofiber surface, we characterized GQ11 binding to lectin and non-lectin proteins. GQ11 nanofibers bound wheat germ agglutinin (WGA), a lectin that recognizes GlcNAc (Fig. 2f). In contrast, GQ11 did not bind bovine serum albumin (BSA) or Concanavalin A (ConA), which lack GlcNAc-binding properties (Fig. 2f). Together these results indicated that GQ11–WGA interactions were specifically mediated by glycans that were displayed on the nanofiber surface and accessible in the surrounding aqueous environment. Notably, although BSA and ConA did not bind to GQ11 nanofibers, the presence of the proteins in solution induced nanofiber lateral association and alignment similar to Ficoll and PEG (Fig. 2g–i and Supplementary Fig. 4a, b). This suggested a general role for excluded volume effects in GQ11 nanofiber lateral association that were independent of the type of macromolecular crowder that was present in solution. From these observations, we inferred that non-specific aggregation of GQ11 nanofibers was prevented by favorable interactions between surface glycans and surrounding water, while conditions that maximized water entropy at the expense of solute entropy promoted nanofiber lateral association into anisotropic networks. We further probed the role of water by`introducing trifluoroethanol (TFE), which has weaker hydrogen bonding capacity than water39. GQ11 nanofibers aligned in 25% TFE (Fig. 2j and Supplementary Fig. 4c), similar to their behavior in the presence of the macromolecular crowders, Ficoll, PEG, BSA, and ConA. Collectively, these observations suggested that decorating self-assembled peptide nanofibers with a moiety that interacts strongly with water, such as a carbohydrate, can impart a checkpoint in the free energy landscape that facilitates hierarchical assembly by establishing repulsive forces that minimize kinetically trapped aggregate formation.
Role of hydroxyls on GQ11 nanofiber association and alignment
Carbohydrates interact strongly with water via their multiple hydroxyl moieties. We studied the role of GlcNAc hydroxyl moieties in GQ11 hierarchical self-assembly by characterizing the morphology of nanofibers formed from a Q11 variant terminated with peracetylated GlcNAc (“AcGQ11”). Self-supporting AcGQ11 gels were weakly birefringent (Fig. 3a, b) and consisted of randomly organized fibers (Fig. 3c and Supplementary Fig. 5). AcGQ11 produced a 2D SAXS scatter plot characteristic of a nearly isotropic sample (Fig. 3d), and SAXS intensity decay in the low q range was consistent with the predicted profile for a collection of randomly oriented rigid rods (Fig. 3e). Likewise, SAXS spectra angular profiles for GQ11 and AcGQ11 were significantly different (Fig. 3f) and represent a substantial decrease in the degree of alignment as evident by the reduction in the spectral order parameter from S = 0.83 to S = 0.53, respectively. At sub-gelling conditions, AcGQ11 nanofibers were dispersed in phosphate-buffered saline (PBS), similar to GQ11, but did not align in the presence of Ficoll or TFE (Fig. 3g and Supplementary Fig. 6). Taken together, these data demonstrated an important role for hydroxyl moieties in GQ11 nanofiber lateral association and alignment.
Hierarchical self-assembly of glycosylated peptides depends on hydroxyl moieties. AcGQ11 forms a self-supporting hydrogels that b were weakly birefringent when viewed under cross-polarizers. c AcGQ11 hydrogels consist of randomly arranged nanofibers when viewed using field-emission scanning electron microscope (scale bar = 1 μm). SAXS d 2D scatter plot, e radial, and f angular intensity profiles suggest random distribution of AcGQ11 nanofibers relative to GQ11 nanofibers. g AcGQ11 nanofibers were dispersed in phosphate-buffered saline (PBS) and did not align in the presence of Ficoll. h TQ11 and i SQ11 nanofibers showed a tendency to aggregate in PBS and did not align in the presence of Ficoll. j LacNAcQ11 and k LacDiNAcQ11 nanofibers were dispersed in PBS; however, only LacNAcQ11 nanofibers aligned in the presence of Ficoll (scale bar = 100 nm in g–k)
To determine whether a single hydroxyl group appended onto Q11 was sufficient to facilitate alignment, we characterized the morphology of nanofibers formed from Q11 variants terminated with threonine and serine (i.e., “TQ11” and “SQ11”, respectively). TQ11 and SQ11 nanofibers demonstrated a tendency to aggregate in PBS and did not align in crowded environments (Fig. 3h, i and Supplementary Fig. 7). These observations suggested that either the number of hydroxyl groups or their arrangement on GlcNAc were important for GQ11 hierarchical assembly. We further interrogated the influence of hydroxyls on nanofiber alignment by synthesizing Q11 variants terminated with the disaccharides Gal-β1,4-GlcNAc (“LacNAc”, 6 OH groups) or GalNAc-β1,4-GlcNAc (“LacDiNAc”, 5 OH groups) (Supplementary Fig. 8a, b). LacNAcQ11 and LacDiNAcQ11 nanofibers were dispersed in PBS (Fig. 3j, k, top and Supplementary Fig. 8c, e), similar to GQ11. In crowded environments, LacNAcQ11 nanofibers formed aligned bundles, whereas LacDiNAcQ11 did not (Fig. 3j, k, bottom and Supplementary Fig. 8d, f). Collectively, these data suggested that nanofiber lateral association depended on carbohydrate chemistry rather than hydroxyl number alone. For LacDiNAcQ11, the GalNAc acetamido may disrupt inter-fiber interactions that are otherwise favorable for LacNAcQ11 having a hydroxyl. Similar dependence on carbohydrate chemistry for encoding hierarchical order is observed in many natural systems, such as multicellular association in sponge and melanoma cells40,41, mucins gels42, glycosynapse formation43, and myelin sheath compaction44.
Influence of glycan density on GQ11 nanofiber alignment
The observed hydroxyl dependence of GQ11 hierarchical assembly suggested that nanofiber lateral association and alignment may be mediated by hydrogen bonding. Carbohydrate hydrogen bond lengths are typically 1–2 nm45, which is comparable to the distance between N-termini of anti-parallel β-strands in peptide nanofibers46. To probe relationships between inter-carbohydrate distance and nanofiber alignment, we first co-assembled GQ11 peptide with a non-glycosylated control peptide, NQ11, at different molar ratios (χ) (Supplementary Fig. 9). Nanofibers with low carbohydrate density (χGQ11 = 0–0.25) were prone to aggregation in PBS, whereas those with higher carbohydrate density (χGQ11 ≥ 0.5 GQ11) remained dispersed (Fig. 4a, left). Nanofibers with a high carbohydrate density aligned in crowded environments, while nanofibers with a low carbohydrate density aggregated (Fig. 4a, right).
Spatial aspects of carbohydrate presentation govern nanofiber lateral association and alignment. a Nanofibers of GQ11 and NQ11 co-assembled at different molar ratios transitioned from a tendency to non-specifically aggregate toward alignment with increasing GQ11 mole fraction. b GQ11G nanofibers were dispersed in phosphate-buffered saline (PBS) and aligned in the presence of Ficoll. GQ11 nanofibers with c a short, 2 amino acid linker (2aaGQ11) tended to aggregate in PBS and did not align in the presence of Ficoll, whereas GQ11 nanofibers with d a longer, 8 amino acid linker (8aaGQ11) were dispersed in PBS and aligned in the presence of Ficoll, similar to GQ11 having a 4 amino acid linker. Scale bar = 100 nm in a–d
We also characterized the morphology of glycosylated nanofibers assembled from a pair of charge-complementary Q11 variants (i.e., QQKFKFKFKQQ (“CATCH+”) and QQEFEFEFEQQ (“CATCH−”)). CATCH peptides co-assemble into β-sheets when mixed at an equimolar ratio47. Nanofibers assembled from a variant of CATCH+ terminated with Asn-linked GlcNAc and non-glycosylated CATCH− were dispersed in neutral aqueous buffer, similar to GQ11/NQ11 nanofibers where χGQ11 = 0.5. However, glycosylated CATCH nanofibers did not laterally associate or align in the presence of Ficoll (Supplementary Fig. 10a, b), in contrast to GQ11/NQ11 nanofibers where χGQ11 = 0.5. These observations suggested that lateral association and alignment of glycosylated peptide nanofibers depends on the peptide itself. This may be due to differences in nanofiber morphology. Compared to GQ11 nanofibers, CATCH nanofibers are often much more twisted and tortuous when viewed with TEM (Supplementary Fig. 10c).
Based on these observations, we further characterized relationships between physical aspects of GlcNAc display and hierarchical assembly of GQ11 nanofibers in crowded conditions using TEM. First, we decreased inter-carbohydrate distance along the nanofiber by synthesizing a Q11 variant modified with asparagine(GlcNAc) at both the N- and C-termini (“GQ11G”). GQ11G nanofibers were dispersed in PBS and aligned in the presence of Ficoll (Fig. 4b and Supplementary Fig. 11a–c), similar to GQ11. Finally, we varied the distance between the nanofiber and carbohydrate by synthesizing GQ11 variants with amino acid linkers separating asparagine(GlcNAc) from Q11 that were either half or twice the length of the parent GQ11 molecule (i.e., “2aaGQ11” or “8aaGQ11”, respectively). 2aaGQ11 nanofibers aggregated in PBS and did not align in crowded environments (Fig. 4c and Supplementary Fig. 11d, e), whereas 8aaGQ11 nanofibers were dispersed in PBS and aligned under crowded conditions (Fig. 4d and Supplementary Fig. 11f, g).
Molecular interactions mediating GQ11 nanofiber alignment
To characterize the molecular interactions involved in the lateral association and alignment of GQ11 nanofibers, we first analyzed 100% GQ11 and 100% NQ11 (i.e., 0% GQ11) nanofibers using circular dichroism (CD) and Fourier-transform infrared spectroscopy (FTIR). In solution, GQ11 and NQ11 nanofibers adopted similar structures that were rich in β-sheets, as indicated by their comparable CD and FTIR spectra (Fig. 5a, b), which suggested that glycosylation did not induce significant changes in the structure of Q11 nanofibers. In the gel state, however, FTIR spectra of GQ11 had a new peak at ~1668 cm−1 that was absent in FTIR spectra of NQ11 (Fig. 5c and Supplementary Fig. 12). A similar peak is seen in samples of α-chitin and has been assigned to stretching of GlcNAc carbonyl groups of one chitin polymer that are hydrogen bonded to amine groups on a neighboring chitin polymer48. These observations suggested the formation of hydrogen bonds involving GlcNAc groups on GQ11 nanofibers upon transition from the solution to the gel state.
Characterization of molecular interactions mediating association and alignment of GQ11 nanofibers. GQ11 and NQ11 nanofibers adopted similar secondary structures under dilute conditions when analyzed with a Fourier-transform infrared spectroscopy and b circular dichroism. c In the gel state, GQ11 had a peak at 1668 cm−1 attributed to the GlcNAc C=O group that was absent in NQ11 gels, whereas peaks at 1620 cm−1 related to the peptide backbone were unchanged. d GQ11 nanofibers demonstrated a strong increase in ellipticity at 203 nm as a function of Ficoll concentration, whereas e NQ11 ellipticity at 203 nm remained unchanged in the presence of Ficoll. When compared to GQ11 nanofibers alone (f), GQ11 nanofibers also demonstrated an increase in ellipticity at 203 nm in the presence of soluble GlcNAc (g). In contrast, when compared to NQ11 alone (h), ellipticity of NQ11 was unchanged in the presence of soluble GlcNAc (i). In g and i, “Obs.” denotes the observed spectrum for a sample of nanofiber+GlcNAc, whereas “Exp.” denotes the expected spectrum based on the sum of the nanofiber and GlcNAc spectra shown under “Individual”. Note that in both f and h, the same representative spectrum is used for the GlcNAc sample
Next, we analyzed GQ11 and NQ11 in dilute and crowded conditions using CD. Ellipticity at 203 nm significantly increased in CD spectra of GQ11 with increasing Ficoll concentration (Fig. 5d), whereas NQ11 spectra remained unchanged (Fig. 5e). This increase in ellipticity for GQ11 nanofibers in crowded conditions was suggestive of interactions involving carbonyl groups in either the peptide backbone or GlcNAc. Taken together with the FTIR spectra of GQ11 gels, we inferred that this change in ellipticity was due to intermolecular interactions involving the GlcNAc carbonyl groups. However, this interpretation was further complicated by reports that anisotropic samples can distort CD spectra due contributions from linear birefringence and linear dichroism49, as seen previously with aligned amyloid protofilaments50. Thus these solution-phase spectroscopic changes may be attributable to the lateral association and alignment of GQ11 nanofibers observed in crowded conditions using TEM. To account for this possibility, we analyzed samples containing either GQ11 or NQ11 nanofibers with or without soluble GlcNAc using CD. We observed increases in ellipticity at 203 nm for samples of GQ11 plus soluble GlcNAc (Fig. 5f, g and Supplementary Fig. 13a), whereas ellipticity was unchanged for samples of NQ11 plus soluble GlcNAc (Fig. 5h, i and Supplementary Fig. 13b). N,N’,N triacetylchitotriose, a GlcNAc oligomer, did not induce lateral association or alignment of GQ11 nanofibers (Supplementary Fig. 14), suggesting that the changes in ellipticity observed for GQ11 in crowded conditions were due to interactions involving carbonyl groups, not linear birefringence or linear dichroism. Further, the observation that GlcNAc interacted with GQ11 but not NQ11, which only differ in having GlcNAc or not, strongly suggests that nanofiber lateral association involves carbohydrate–carbohydrate interactions (CCI). CCI are weak and stabilized by multivalent avidity effects, as demonstrated previously with glycopolymers, glycomicelles, and glyconanoparticles characterized with analytical techniques such as Langmuir monolayers, modified enzyme-linked immunosorbent assay, surface plasmon resonance, quartz crystal microbalance, and calorimetry51,52,53,54,55,56,57. Here the smaller change in ellipticity observed for GQ11 nanofibers mixed with soluble GlcNAc (Fig. 5g) vs. nanofibers in crowded conditions (Fig. 5d) could be due in part to weaker affinity between GlcNAc immobilized in a multivalent configuration, such as on a peptide nanofiber, and monovalent GlcNAc in solution. However, further studies are needed to quantitatively measure the affinity of intermolecular interactions between carbohydrates on β-sheet peptide nanofibers.
Non-fouling properties of GQ11 hydrogels
Within living systems, carbohydrates assembled into multivalent configurations demonstrate emergent functional properties that are not shared by their monovalent counterparts. For example, dense carbohydrate presentation at biological interfaces, such as the glycocalyx and secreted mucinous layers, confers resistance to non-specific biological interactions while encoding selective biomolecule recognition58. Resistance is often attributed to a combination of carbohydrate hydrophilicity and steric repulsion, while recognition is enabled by carbohydrate clustering and multivalency. Here we characterized protein interactions with GQ11 and NQ11 gels by analyzing the release of encapsulated BSA and WGA into bulk aqueous media. Nearly all encapsulated BSA was released from GQ11 gels, whereas a significant fraction of encapsulated WGA was retained (Fig. 6a, b). In contrast, NQ11 gels released nearly all encapsulated BSA and WGA (Fig. 6a, b). Together, these data demonstrated that glycosylation endowed peptide nanofiber gels with selective recognition of a GlcNAc-binding lectin, where high binding affinity likely resulted from assembling carbohydrates into a multivalent configuration.
GQ11 gels selectively recognized a GlcNAc-binding protein but otherwise resisted non-specific biological interactions. a Bovine serum albumin rapidly released from NQ11 and GQ11 hydrogels, whereas b wheat germ agglutinin was selectively retained by GQ11 hydrogels. Dashed lines in a and b represent mass of protein encapsulated within the gel. GQ11 hydrogels were more resistant to non-specific RAW264.7 macrophage adhesion than NQ11 hydrogels in the presence of serum as determined c quantitatively via metabolic activity and d, e qualitatively via light microscopy. f–h GQ11 hydrogels were also more resistant to non-specific NIH3T3 fibroblast adhesion in the presence of serum than NQ11 hydrogels. i Glass surfaces coated with GQ11 hydrogels resisted non-specific NIH3T3 fibroblast adhesion. j Region of GQ11 hydrogel represented by white box in i. In contrast, k regions of glass coated with NQ11 hydrogels were fouled by NIH3T3 fibroblasts. l Region of NQ11 hydrogel represented by white box in k. m GQ11 hydrogels were also more resistant to E. coli adhesion than NQ11 hydrogels. In i–l, nanofibers were stained green with Thioflavin T (ThT) and cells were stained red with CellTracker®. Scale bar = 100 μm in d, e, g–i, and k. Data presented as mean ± standard deviation in a–c, f, and m. **p < 0.005, and ***p < 0.0005 using Student’s t test (n = 3)
We also characterized GQ11 gel resistance to non-specific mammalian cell adhesion. Both macrophages and fibroblasts attached to the surface of NQ11 gels in the presence of serum, whereas significantly fewer cells attached to GQ11 gels (Fig. 6c–h and Supplementary Fig. 15). Regions of glass slides coated with GQ11 had few adherent fibroblasts, whereas regions coated with NQ11 were fouled (Fig. 6i–l and Supplementary Fig. 16). Collectively, these observations suggested that NQ11 gels were susceptible to non-specific adsorption of serum proteins that mediate cell adhesion, while GQ11 gels were more resistant to non-specific protein adsorption. This was likely due to steric repulsion and hydrophilicity endowed by GlcNAc monosaccharides organized into a dense layer on the surface of the nanofiber network. Further, these results suggested that GlcNAc-binding receptors expressed by macrophages or fibroblasts, such as Endo180 or CD20659,60, do not interact with GlcNAc groups on GQ11 gels in a manner that enables cell adhesion.
To characterize the biodegradability of GQ11 and NQ11 hydrogels, we treated each with trypsin, a protease that was expected to cleave both peptides at the amide linkage C-terminal to the Lys residue of Q11 (QQK*FQFQFEQQ). Mass spectra for each gel before and after trypsin treatment demonstrated that both peptides were digested (Supplementary Fig. 17), indicating that both NQ11 and GQ11 hydrogels are biodegradable.
Finally, we evaluated GQ11 gel resistance to bacterial adhesion. Compared to control surfaces, GQ11 gels reduced the number of viable adherent Escherichia coli to <10%, whereas NQ11 gels only reduced bacteria colonization to ~40% (Fig. 6m). Interestingly, AcGQ11 gels demonstrated comparable resistance to E. coli adhesion as GQ11 gels, whereas their resistance to fibroblast and macrophage adhesion was weaker (Supplementary Fig. 18). These observations suggested that both glycosylation and nanofiber alignment may contribute to GQ11 resistance to mammalian cell adhesion, while only glycosylation is necessary to prevent E. coli adhesion. Previous reports have demonstrated that chitin nanocrystals embedded within electrospun polymer membranes can confer non-fouling properties through a combination of increased hydrophilicity that prevents cell adhesion and the antimicrobial properties of deacetylated glucosamine groups (i.e., chitosan) produced via acid hydrolysis during chitin nanocrystal extraction61,62. Here we propose that resistance to E. coli adhesion on GQ11 gels primarily results from the increased hydrophilicity established by nanofiber glycosylation because our materials are devoid of deacetylated glucosamine units having antimicrobial properties.
Discussion
Here we show that glycosylation facilitates hierarchical assembly of β-sheet peptide nanofibers into anisotropic networks that are ordered over multiple length scales. Lateral association and alignment of glycosylated peptide nanofibers is dependent on both carbohydrate chemistry and density. The data in Figs. 4 and 5 support a role for short-range hydrogen bonding between GQ11 nanofibers as a mediator of lateral association. The low energy and reversibility of hydrogen bonding affords a key advantage over longer range and omnidirectional interactions in hierarchical self-assembly. Namely, weak interactions that can be stabilized by near-neighbor cooperativity may extend inter-fiber contact time while also permitting nanofiber rotation and sliding, which together have been theorized to facilitate fiber alignment63. Importantly, the multiple co-localized hydrogen bonds afforded by a carbohydrate stabilized inter-fiber interactions to an extent that was not possible with mono-hydroxylated threonine and serine, yet were not so robust as to kinetically trap nanofibers as misaligned aggregates. Although beyond the scope of this report, elucidating specific inter-fiber interactions involved in GQ11 lateral association will likely open up new avenues to rationally design glycosylated peptides that hierarchically assemble into precise supramolecular architectures.
Our data demonstrate that anisotropic GQ11 networks resist non-specific cell, bacteria, and protein interactions, yet selectively recognize lectins, analogous to natural densely glycosylated materials. Compared to many previously reported glycoconjugates designed for low-fouling biomaterial applications, such as polysaccharides, peptoids, and alkanethiols64,65,66, GQ11 is unique because it can physically crosslink into self-supporting gels. Given the important role of macrophages, fibroblasts, and bacteria in establishing the foreign body reaction to implants67, these observations suggest that GQ11 gels may find use as either low-fouling biomaterials or surface coatings. This potential is further supported by the reported low immunogenicity of Q1168, which suggests that GQ11 will demonstrate favorable immunological compatibility.
From our observations, we postulate that GQ11 hierarchical assembly proceeds through a sequential mechanism. First, GQ11 peptides self-assemble into nanofibers that resist non-specific aggregation due to water–carbohydrate interactions established by a dense surface glycan layer (Fig. 7a). Then, excluded volume effects drive nanofiber collapse into bundles, where relatively weak inter-fiber interactions mediate lateral association while facilitating nanofiber rotation and sliding to correct misalignments, ultimately resulting in system convergence on the most thermodynamically favorable state of aligned nanofibers (Fig. 7b). Nanofiber twisting due to amino acid chirality would lead to radial display of carbohydrates at a regular interval that provides inter-fiber contact points in all directions (Fig. 7c). Hierarchical assembly via lateral association rather than entwining would suggest that GQ11 nanofiber bundles can grow radially with no geometric constraint, because the entropic penalty resulting from untwisting of subsequent layers in entwined structures would be avoided35,36,37. Carbohydrates displayed on the outermost nanofibers of a network interact with water to endow resistance to non-specific biological interactions, yet also encode selective biomolecule recognition (Fig. 7d). Together, these observations demonstrate that glycosylation can alter the structure and function of supramolecular biomaterials by establishing attractive and repulsive forces that dictate intermolecular interactions, analogous to the influence of glycosylation on protein form and function19. We envision that modifying self-assembling peptides with the diverse carbohydrate chemistries found throughout nature will lead to supramolecular biomaterials with a broad range of new emergent properties that are advantageous for various biomedical and biotechnological applications.
Proposed role of glycosylation in hierarchical self-assembly and emergent function of GQ11 nanofibers. Glycans on GQ11 nanofibers interact with water (a), forming a hydration layer that minimizes nanofiber non-specific aggregation under dilute conditions (b). Excluded volume effects in crowded environments drive nanofiber bundling (b), while short-range inter-fiber interactions facilitate lateral association and long-range alignment. c Carbohydrates are radially projected at regular intervals due nanofiber twisting established by amino acid chirality, which provides inter-fiber contact points in all directions to facilitate lateral association without geometric constraint. d Glycans on the outermost layer of nanofibers interact favorably with water to prevent non-specific biological interactions, yet can specifically recognize a carbohydrate-binding protein
Methods
Peptide synthesis and purification
Amino acids and amide resin were purchased from Novabiochem. 2-(7-Aza-1H-benzotriazole-1-yl)-1,1,3,3-tetramethyluronium hexafluorophosphate (HATU), 1-hydroxy-7-azabenzotriazole (HOAt), N,N-diisopropylethylamine (DIEA), dimethylformamide, trifluoroacetic acid (TFA), diethyl ether, and methanol were purchased from Fisher Scientific. Piperidine, 1,8-Diazabicyclo[5.4.0] undec-7-ene, triisopropylsilane (TIS), and sodium methoxide were purchased from Sigma-Aldrich. Peptides GQ11 (N(GlcNAc)SGSGQQKFQFQFEQQ), GQ11G (N(GlcNAc)SGSGQQKFQFQFEQQGSGSN(GlcNAc)), 2aaGQ11 (N(GlcNAc)SGQQKFQFQFEQQ), 8aaGQ11 (N(GlcNAc)SGSGSGSGQQKFQFQFEQQ), NQ11 (NSGSGQQKFQFQFEQQ), AcGQ11 (N((Ac)3-GlcNAc)SGSGQQKFQFQFEQQ), TQ11 (TSGSGQQKFQFQFEQQ), and SQ11 (SSGSGQQKFQFQFEQQ) were synthesized following standard Fmoc solid phase peptide synthesis protocols with DIEA/HOAt/HATU activation, according to previously reported methods27. Peptides were cleaved with TFA/TIS/water (95:2.5:2.5) cocktail. Peptides were precipitated with diethyl ether, dried, resuspended in distilled water, and freeze dried. GQ11, GQ11G, 2aaGQ11, and 8aaGQ11 were further treated with sodium methoxide and precipitated in methanol for deacetylation. All peptides were purified using an Ultimate 3000 HPLC equipped with a C18 column. Peptide molecular weight was assessed using matrix-assisted laser desorption/ionization–time of flight (MALDI-TOF) in a Bruker Microflex LRF system and α-cyano-4-hydroxycinnamic acid as the matrix (Supplementary Fig. 19). Peptide purity was >95% for all studies (Supplementary Fig. 20).
Nanofiber and gel preparation
To prepare nanofibers, lyophilized peptide powders were dissolved in ultrapure water at 5 mM, diluted to working concentration in 1× PBS (137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 1.8 mM KH2PO4), and incubated overnight at room temperature. To prepare gels, peptides were dissolved in ultrapure water at 5 mM, diluted to 4.5 mM using 10× PBS, and incubated overnight at room temperature. Gels were either formed by placing peptide solution into the bottom of a 2 mL scintillation vial or by placing between two glass slides rendered hydrophobic with Sigmacote (Sigma-Aldrich), which were separated with 3 mm spacers.
Birefringence
All gels were cast in 2 mL scintillation vials, placed in front of a linearly polarized light source, and imaged using a Nikkon digital camera equipped with cross-polarizers.
Field-emission scanning electron microscopy
For visualization of gels using FE-SEM, gels were dehydrated in a series of ethanol/water solutions progressing from 30% to 100% ethanol. Dehydrated gels were critical point dried for 1 h in a Bal-tec critical point dryer. Samples were sputter coated with 8 nm gold using a Denton Desk V Sputter System and imaged using a Hitachi SU5000 Schottky Field-Emission Variable Pressure SEM. Sample preparation and image collection were done on equipment maintained by the University of Florida Interdisciplinary Center for Biotechnology Research.
Transmission electron microscopy
For visualization of nanofibers with TEM, nanofiber solutions in PBS were prepared as described above. Nanofibers were diluted to 250 μM in 1× PBS or 1× PBS with macromolecular crowder. Crowders included: bovine serum albumin (10 μM, Fisher Scientific); ConA (10 μM, Fisher Scientific); PM70 Ficoll (100 mg/mL, GE Healthcare); PEG ([MW = 2000] 40% (w/v), Alfa Aesar); D-sucrose (100 mg/ml, Fisher Scientific); N,N’,N-triacetylchitotriose (200 μM, Sigma Aldrich); or TFE (Acros Organics). Within 5 min, solutions of nanofibers ± macromolecular crowder were adsorbed onto Formvar/carbon grids (FCF400-CU-UB, Electron Microscopy Sciences) by placing liquid sample on top of the grid for 1 min. Grids were dried by tilting onto a Kimwipe (Kimberly-Clark) and negatively stained with a 2% aqueous solution of uranyl acetate. For WGA-gold labeling of nanofibers, samples adsorbed onto grids were placed upside-down on top of a 20 μL drop of gold-conjugated WGA (EY Laboratories, Inc., GP-2101-15, 10 mg/mL) for 5 min prior to negative staining with 2% uranyl acetate. Samples were washed 3× with 1× PBS prior to negative staining to remove any loosely adsorbed background WGA-gold. Samples were imaged using a Hitachi 7000 TEM housed in the University of Florida Interdisciplinary Center for Biotechnology Research.
Cryo-electron microscopy
All sample preparation for cryo-EM was performed by the University of Florida Interdisciplinary Center for Biotechnology Research EM Core Facility. Three-microliter aliquots of 1× PBS containing GQ11 (250 μM) and Ficoll (100 mg/mL) were applied to C-flat holey carbon grids (Protochips, Inc.) and vitrified using a Vitrobot™ Mark IV (FEI Co.) operated at 4 °C and with ~90% humidity in the control chamber. The vitrified sample was stored under liquid nitrogen and transferred into a Gatan cryo-holder (Model 626/70) for imaging. The sample was examined using a 4k × 4k CCD camera (Gatan, Inc.) on a Tecnai G2 F20-TWIN Transmission Electron Microscope (FEI Co.) operated at a voltage of 200 kV using low-dose conditions (~20 e/Å2). Images were recorded with a defocus of approximately 3 µm to improve contrast.
Small angle X-ray scattering
To quantify the average fiber orientation over macroscopic scales within self-assembled peptide nanofiber hydrogels, we performed SAXS measurements, sampling volumes of approximately 1 mm3. Hydrogels were prepared in aqueous buffer, pipetted into thin-walled amorphous quartz capillaries (10 μm wall thickness), and flame sealed. SAXS data were collected with a 2D wire detector (Bruker Nanostar) for approximately 18 h and analyzed using a combination of Fit2D software and custom written analysis code. The scattering patterns lay on a coordinate system of wave-vectors in reciprocal space, given by
where λ = 1.54 Å is the Cu Kα X-ray wavelength and 2θ is the scattering angle. Sample anisotropy can be quantified by measuring the nematic order parameter from these data, given by34,
Here ϕ is the azimuthal angle in reciprocal space and f(ϕ) is the orientational distribution function, which is determined by fitting a Gaussian line-shape to the relative intensity at a fixed q as a function of the azimuthal angle69. We average over a narrow q-range (0.13 < q < 0.5) to determine f(ϕ) for each sample.
Fourier transform infrared spectroscopy
Peptide secondary structure was characterized using attenuated total reflectance FTIR. A 3 μL nanofiber solution in water or PBS (4.5 mM) was deposited onto a diamond-coated ZnSe crystal and dried at room temperature. Spectra were collected with a PerkinElmer Spectrum 100 spectrometer equipped with a KBr beam splitter. The data presented corresponds to the average of 4 scans at a resolution of 4 cm−1.
Circular dichroism
Peptide secondary structure was characterized under dilute and crowded conditions using CD on an Aviv Model 430 spectrometer or an Applied Photophysics Chiroscan V100 spectrophotometer. For analysis of peptide secondary structure and characterization of peptides in crowded conditions, nanofibers were prepared by dissolving lyophilized peptide powders in ultrapure water at 5 mM and diluting to 0.15 mM in 1× CD potassium phosphate buffer (10 mM Na2HPO4, 1.8 mM KH2PO4, 2.7 mM KCl, 137 mM KF, pH 7.4). Here sodium chloride was substituted with potassium fluoride due to strong absorbance by Cl− ions in the ultraviolet range. For analysis of nanofiber binding to GlcNAc, nanofibers were prepared by dissolving lyophilized peptides in water at 10 mM and then diluting ten-fold in 1× CD potassium phosphate buffer. Samples were then diluted to 0.25 mM in 1× CD potassium phosphate buffer containing 0.6 mM GlcNAc. Spectra reported correspond to the average of 3 runs after baseline subtraction and with dynode values <500 V.
Protein binding to nanofibers
Nanofibers (1 mM) in PBS were incubated for 1 h at room temperature with 10 μM of WGA (Sigma-Aldrich L9640), ConA (MP Biomedicals 15071001), or BSA (MP Biomedicals). Nanofibers were sedimented by centrifugation at 11,300 × g for 5 min, and supernatant was analyzed for unbound protein by measuring tryptophan fluorescence (excitation 280 nm, emission 345 nm) using a SpectraMax M3 spectrophotometer (Molecular Devices), similar to previously reported methods27.
Protein release from hydrogels
PBS 70 μL solution containing 4.5 mM peptide plus 10 μM BSA or WGA was added to the bottom of 96-well plates and allowed to gel overnight. Protein release was initiated by adding 200 μL of fresh 1× PBS on top of the gels. For all groups at each time point, 100 μL of supernatant was transferred into a black 96-well, glass-bottom plate, and protein concentration was measured via tryptophan fluorescence (excitation λ = 280 nm, emission λ = 345 nm) using a SpectraMax M3 spectrophotometer (Molecular Devices). After fluorescence measurements, supernatants were transferred back to the corresponding well containing gels until the following time point. Fluorescence measurements were performed 0.5, 1, 2, 4, 8, and 24 h.
Expression and purification of β-1,4-galactosyltransferase 1
Human placental β-1,4-galactosyltransferase 1 Y284L (GalNAcT) mutant was previously reported30. Sequence can be found in Supplementary Data 1 and 2. GalNAcT gene (Genscript) was inserted into pET(+)-21d vector between NcoI and XhoI sites, transformed into TOP 10 E. coli, and selected on lysogeny broth (LB) agar plates containing 100 μg/mL ampicillin. Positive clones were used to inoculate 5 mL of LB and culture was grown overnight at 37 °C on an orbital shaker (225 rpm). Genes were isolated from culture using a Plasmid Mini-prep Kit (Qiagen) following the manufacturer’s instructions and sequenced at the University of Florida Interdisciplinary Center for Biotechnology Research. Genes containing the correct sequence were transformed into Origami B (DE3) E. coli for expression and then plated on LB agar plates containing 100 μg/mL ampicillin and 50 μg/mL of kanamycin A for selection. Positive clones were picked and used to inoculate 5 mL of 2×TY media containing antibiotics and allowed to grow overnight at 37 °C on an orbital shaker at 225 rpm. Cultures were expanded in 1 L of 2×TY media containing antibiotics and incubated at 37 °C on an orbital shaker until they reached an optical density of 0.6 at 600 nm. To induce protein expression, 0.5 mM isopropyl β-d-1-thiogalactopyranoside (IPTG, Fisher Scientific) was added and cultures were incubated for 18 h at 18 °C shaking at 225 rpm. Bacteria were collected by centrifugation at 16,300 × g for 10 min and washed 3 times with PBS. Bacteria were lysed by incubating with 10× BugBuster (EMD Millipore) diluted to working concentration with HEPES buffer (20 mM) containing lysozyme (Fisher) and deoxyribonuclease I (Worthington) for 15 min at room temperature. Lysed bacteria were cleared by centrifugation at 42,800 × g for 15 min at 4 °C. Supernatant containing protein was loaded into columns containing HisPurTM cobalt resin (Fisher), washed with 15 column volumes of HEPES buffer (50 mM), and eluted with an imidazole gradient. Imidazole was removed from the pure protein fractions by dialyzing against 2 L of HEPES buffer containing 10 mM of MnCl2. Protein molecular weight was confirmed with sodium dodecyl sulfate-polyacrylamide gel electrophoresis gel stained with Coomassie Blue.
Enzymatic carbohydrate conversion
LacNAcQ11 and LacDiNAcQ11 were prepared following previously established methods27,30. For LacNAcQ11, per 1 nmol of GQ11, 1.5 μg of β-1,4-galactosyltransferase from bovine milk (β-1,4-GalT, Sigma-Aldrich G5507) and 5 nmol of uridine diphosphate galactose (UDP-Gal, Carbosynth MU06699) were added in 20 mM HEPES buffer with 10 mM MnCl2 (pH 7.5). For LacDiNAcQ11, per 1 nmol of GQ11, 2 μg of β-1,4-galactosyltransferase 1 Y285L mutant (β-1,4GalT Y285L), and 10 nmol of uridine diphosphate n-acetylgalactosamine (UDP-GalNAc, Carbosynth MU04515) were added in 50 mM HEPES buffer containing 10 mM MnCl2 (pH 7.5). Enzymatic conversions were carried out at 37 °C for 18 h. Enzymatic conversions were confirmed with MALDI-TOF (Supplementary Fig. 8a, b). To remove enzymes and unreacted species, nanofibers were washed 3 times by centrifuging at 11,300 × g for 5 min, removing supernatant, and resuspending in PBS.
Mammalian cell adhesion to gels and gel-coated surfaces
RAW264.7 macrophages and NIH 3T3 fibroblasts were from ATCC. For mammalian cell studies, RAW264.7 macrophages and NIH 3T3 fibroblasts were grown in Dulbecco’s Modified Eagle Medium (Corning) supplemented with 10% fetal bovine serum (Hyclone) and 1% penicillin–streptomycin (Gibco) at 37 °C and 5% CO2. Cells (100 μL, 5 × 105 cells/mL) were seeded on top of GQ11 and NQ11 gels (55 μL) formed on the bottom of a 96-well plate, as described above. Cells were incubated for 24 h at 37 °C and 5% CO2. Loose cells were removed by washing gels 5 times with 1× PBS. Cells were imaged in bright field with a Zeiss Axio Observer Microscope. Cell metabolic activity was measured using CellTiter-Blue® Cell Viability Assay (Promega) according to the manufacturer’s instructions. To study fibroblast adhesion to gel-coated surfaces, glass microscope slides were cleaned with ethanol and coated with a thin layer of 5 mM GQ11 or NQ11 solution. Slides were maintained at 37 °C for 1 h before placing them in tissue culture plates. Fibroblasts were fluorescently labeled with CellTracker™ Red CMTPX Dye (ThermoFisher) before seeding onto gel-coated or bare glass regions at a density of 5 × 105 cells/mL. Fibroblasts were incubated for 8 h at 37 °C and 5% CO2. Slides were immersed three times in PBS to remove loose cells and imaged using a Zeiss Axio Observer Microscope equipped with a red excitation/emission filter (546/608 nm).
Bacterial cell adhesion to hydrogels
One ShotTM TOP10 Chemically Competent E. coli containing ampicillin-resistant pET(+)-21d vector were grown in LB medium at 37 °C on an orbital shaker (225 rpm) until they reached an optical density of 0.1 (λ = 600 nm). Bacteria in LB media (100 μL) were added onto the top of GQ11 and NQ11 gels (55 μL) cast on the bottom of 96-well plates, as described above. Samples were incubated at room temperature for 4 h, after which samples were washed 5 times with 1× PBS. Bound remaining bacteria were quantified with BacTiter-GloTM (Promega) according to the manufacturer’s instructions.
Trypsin digest
Five microliters of 5 mM NQ11 or GQ11 gel was digested with 1 µL of 0.25% Trypsin at 37 °C for 24 h. Samples were mixed 1:1 with α-cyano-4-hydroxycinnamic acid (Sigma) (10 mg/mL) in 70% acetonitrile and 30% water (both containing 0.1% TFA). Two microliters of sample was spotted and dried onto a MALDI-TOF plate and scanned. Samples were analyzed using MALDI-TOF in a Bruker Daltonics Autoflex maX with a smart beam II UV laser.
Statistical analysis
Protein, bacteria, and cell experiments were conducted in triplicate. Statistical differences between groups were analyzed using an unpaired t test with a confidence interval of 95% or analysis of variance (ANOVA) with Tukey’s post hoc in the GraphPad Prism software.
T test comparing GQ11 and NQ11 only
All experimental and control groups were n = 3 for macrophage, fibroblast, and bacterial cell adhesion to bulk gels, and the data were reported as average ± standard deviation. Statistical differences between groups were analyzed using unpaired Student’s t test. In macrophage adhesion studies, p = 0.0048 and F value is 7.771. In fibroblast adhesion studies, p = 0.0002 and F value is 180. In bacteria adhesion studies, p = 0.0005 and F value is 3.073.
One-way ANOVA to include AcGQ11
All experimental and control groups were n = 3 for macrophage, fibroblast, and bacterial cell adhesion to bulk gels, and the data were reported as average ± standard deviation. Statistical differences between groups were analyzed using ANOVA with Tukey’s post hoc in the GraphPad Prism software. In macrophage adhesion studies, p = 0.002 for GQ11 vs. NQ11, p = 0.0027 for GQ11 vs. AcGQ11, and no significant (n.s.) difference between NQ11 vs. AcGQ11. F value is 24.17. In fibroblast adhesion studies, ****p < 0.0001 for GQ11 vs. NQ11, p = 0.0017 for GQ11 vs. AcQ11 and p = 0.003 for NQ11 vs. AcGQ11. F value is 114.7. In bacteria adhesion studies, ****p < 0.0001 for GQ11 vs. NQ11 and NQ11 vs. AcGQ11 and p = 0.0013 for GQ11 vs. AcGQ11. F value is 127.
Data availability
All materials and raw data are available upon request from the corresponding author.
References
Vignaud, T., Blanchoin, L. & Thery, M. Directed cytoskeleton self-organization. Trends Cell Biol. 22, 671–682 (2012).
Muiznieks, L. D. & Keeley, F. W. Molecular assembly and mechanical properties of the extracellular matrix: a fibrous protein perspective. Biochim. Biophys. Acta 1832, 866–875 (2013).
Webber, M. J., Appel, E. A., Meijer, E. W. & Langer, R. Supramolecular biomaterials. Nat. Mater. 15, 13–26 (2016).
Du, X., Zhou, J., Shi, J. & Xu, B. Supramolecular hydrogelators and hydrogels: from soft matter to molecular biomaterials. Chem. Rev. 115, 13165–13307 (2015).
Whitelam, S. Hierarchical assembly may be a way to make large information-rich structures. Soft Matter 11, 8225–8235 (2015).
Mouw, J. K., Ou, G. & Weaver, V. M. Extracellular matrix assembly: a multiscale deconstruction. Nat. Rev. Mol. Cell Biol. 15, 771–785 (2014).
Conde, C. & Caceres, A. Microtubule assembly, organization and dynamics in axons and dendrites. Nat. Rev. Neurosci. 10, 319–332 (2009).
Tantakitti, F. et al. Energy landscapes and functions of supramolecular systems. Nat. Mater. 15, 469–476 (2016).
Zhang, S. et al. A self-assembly pathway to aligned monodomain gels. Nat. Mater. 9, 594–601 (2010).
Chen, Y. R., Gan, H. X. & Tong, Y. W. pH-controlled hierarchical self-assembly of peptide amphiphile. Macromolecules 48, 2647–2653 (2015).
Huang, Z. et al. Responsive nematic gels from the self-assembly of aqueous nanofibres. Nat. Commun. 2, 459 (2011).
Lowik, D. W. P. M. et al. A highly ordered material from magnetically aligned peptide amphiphile nanofiber assemblies. Adv. Mater. 19, 1191–1195 (2007).
Pappas, C. G. et al. Alignment of nanostructured tripeptide gels by directional ultrasonication. Chem. Commun. (Camb.) 51, 8465–8468 (2015).
Wall, B. D. et al. Aligned macroscopic domains of optoelectronic nanostructures prepared via shear-flow assembly of peptide hydrogels. Adv. Mater. 23, 5009–5014 (2011).
Wallace, M., Cardoso, A. Z., Frith, W. J., Iggo, J. A. & Adams, D. J. Magnetically aligned supramolecular hydrogels. Chem. Eur. J. 20, 16484–16487 (2014).
Hu, Y. et al. Electrostatic-driven lamination and untwisting of beta-sheet assemblies. ACS Nano 10, 880–888 (2016).
Zhou, J., Du, X., Gao, Y., Shi, J. & Xu, B. Aromatic-aromatic interactions enhance interfiber contacts for enzymatic formation of a spontaneously aligned supramolecular hydrogel. J. Am. Chem. Soc. 136, 2970–2973 (2014).
Eichler, J. & Koomey, M. Sweet new roles for protein glycosylation in prokaryotes. Trends Microbiol. 25, 662–672 (2017).
Lis, H. & Sharon, N. Protein glycosylation. Structural and functional aspects. Eur. J. Biochem. 218, 1–27 (1993).
Gehle, V. M., Walcott, E. C., Nishizaki, T. & Sumikawa, K. N-glycosylation at the conserved sites ensures the expression of properly folded functional ACh receptors. Mol. Brain Res. 45, 219–229 (1997).
Parker, J. L., Day-Williams, M. J., Tomas, J. M., Stafford, G. P. & Shaw, J. G. Identification of a putative glycosyltransferase responsible for the transfer of pseudaminic acid onto the polar flagellin of Aeromonas caviae Sch3N. Microbiologyopen 1, 149–160 (2012).
Verdugo, P. Supramolecular dynamics of mucus. Cold Spring Harb. Perspect. Med. 2, 1–14 (2012).
Caliskan, O. S., Sardan Ekiz, M., Tekinay, A. B. & Guler, M. O. Spatial organization of functional groups on bioactive supramolecular glycopeptide nanofibers for differentiation of mesenchymal stem cells (MSCs) to brown adipogenesis. Bioconjug. Chem. 28, 740–750 (2017).
Gunay, G. et al. Antigenic GM3 lactone mimetic molecule integrated mannosylated glycopeptide nanofibers for the activation and maturation of dendritic cells. ACS Appl Mater. Interfaces 9, 16035–16042 (2017).
Lee, S. S. et al. Sulfated glycopeptide nanostructures for multipotent protein activation. Nat. Nanotechnol. 12, 821–829 (2017).
Liu, J. et al. Peptide glycosylation generates supramolecular assemblies from glycopeptides as biomimetic scaffolds for cell adhesion and proliferation. ACS Appl Mater. Interfaces 8, 6917–6924 (2016).
Restuccia, A., Tian, Y. F., Collier, J. H. & Hudalla, G. A. Self-assembled glycopeptide nanofibers as modulators of galectin-1 bioactivity. Cell Mol. Bioeng. 8, 471–487 (2015).
Ustun Yaylaci, S. et al. Supramolecular GAG-like self-assembled glycopeptide nanofibers induce chondrogenesis and cartilage regeneration. Biomacromolecules 17, 679–689 (2016).
Restuccia, A. & Hudalla, G. A. Tuning carbohydrate density enhances protein binding and inhibition by glycosylated beta-sheet peptide nanofibers. Biomater. Sci. 6, 2327–2335 (2018).
Restuccia, A. et al. Evaluation of self-assembled glycopeptide nanofibers modified with N,N’-diacetyllactosamine for selective galectin-3 recognition and inhibition. ACS Biomater. Sci. Eng. 4, 3451–3459 (2018).
Roytman, R. et al. Exploring the self-assembly of glycopeptides using a diphenylalanine scaffold. Org. Biomol. Chem. 9, 5755–5761 (2011).
Tsuzuki, T., Kabumoto, M., Arakawa, H. & Ikeda, M. The effect of carbohydrate structures on the hydrogelation ability and morphology of self-assembled structures of peptide-carbohydrate conjugates in water. Org. Biomol. Chem. 15, 4595–4600 (2017).
Collier, J. H. & Messersmith, P. B. Enzymatic modification of self-assembled peptide structures with tissue transglutaminase. Bioconjug Chem. 14, 748–755 (2003).
Gennes, P. G. D. & Prost, J. The Physics of Liquid Crystals 2nd edn (Clarendon Press; Oxford University Press, Oxford, 1993).
Hall, D. M., Bruss, I. R., Barone, J. R. & Grason, G. M. Morphology selection via geometric frustration in chiral filament bundles. Nat. Mater. 15, 727–732 (2016).
Aggeli, A. et al. Hierarchical self-assembly of chiral rod-like molecules as a model for peptide beta-sheet tapes, ribbons, fibrils, and fibers. Proc. Natl. Acad. Sci. USA 98, 11857–11862 (2001).
Weisel, J. W., Nagaswami, C. & Makowski, L. Twisting of fibrin fibers limits their radial growth. Proc. Natl. Acad. Sci. USA 84, 8991–8995 (1987).
Zhou, H. X., Rivas, G. & Minton, A. P. Macromolecular crowding and confinement: biochemical, biophysical, and potential physiological consequences. Annu. Rev. Biophys. 37, 375–397 (2008).
Roccatano, D., Colombo, G., Fioroni, M. & Mark, A. E. Mechanism by which 2,2,2-trifluoroethanol/water mixtures stabilize secondary-structure formation in peptides: a molecular dynamics study. Proc. Natl Acad. Sci. USA 99, 12179–12184 (2002).
Bucior, I., Scheuring, S., Engel, A. & Burger, M. M. Carbohydrate-carbohydrate interaction provides adhesion force and specificity for cellular recognition. J. Cell Biol. 165, 529–537 (2004).
Eggens, I. et al. Specific interaction between Lex and Lex determinants - a possible basis for cell recognition in preimplantation embryos and in embryonal carcinoma cells. J. Biol. Chem. 264, 9476–9484 (1989).
Haugstad, K. E. et al. Enhanced self-association of mucins possessing the T and Tn carbohydrate cancer antigens at the single-molecule level. Biomacromolecules 13, 1400–1409 (2012).
Hakomori, S. Carbohydrate-to-carbohydrate interaction, through glycosynapse, as a basis of cell recognition and membrane organization. Glycoconj. J. 21, 125–137 (2004).
Boggs, J. M. et al. Participation of galactosylceramide and sulfatide in glycosynapses between oligodendrocyte or myelin membranes. FEBS Lett. 584, 1771–1778 (2010).
Jeffrey, G. A. & Lewis, L. Cooperative aspects of hydrogen-bonding in carbohydrates. Carbohyd. Res. 60, 179–182 (1978).
Makin, O. S. & Serpell, L. C. Structures for amyloid fibrils. FEBS J. 272, 5950–5961 (2005).
Seroski, D. T. et al. Co-assembly tags based on charge complementarity (CATCH) for installing functional protein ligands into supramolecular biomaterials. Cell. Mol. Bioeng. 9, 335–350 (2016).
Focher, B., Naggi, A., Torri, G., Cosani, A. & Terbojevich, M. Structural differences between chitin polymorphs and their precipitates from solutions - evidence from Cp-Mas C-13-Nmr, Ft-Ir and Ft-Raman spectroscopy. Carbohyd. Polym. 17, 97–102 (1992).
Davidsson, A., Norden, B. & Seth, S. Measurement of oriented circular-dichroism. Chem. Phys. Lett. 70, 313–316 (1980).
Adachi, R. et al. Flow-induced alignment of amyloid protofilaments revealed by linear dichroism. J. Biol. Chem. 282, 8978–8983 (2007).
Seah, N., Santacroce, P. V. & Basu, A. Probing the lactose center dot GM3 carbohydrate-carbohydrate interaction with glycodendrimers. Org. Lett. 11, 559–562 (2009).
Matsuura, K. et al. Surface plasmon resonance study of carbohydrate-carbohydrate interaction between various gangliosides and Gg3-carrying polystyrene. Biomacromolecules 5, 937–941 (2004).
Yoon, S. J. et al. Interaction of N-linked glycans, having multivalent GlcNAc termini, with GM3 ganglioside. Glycoconj. J. 23, 639–649 (2006).
Yoon, S. J. et al. Interaction of N-linked glycans, having multivalent GlcNAc termini, with GM3 ganglioside. Glycoconj. J. 24, 181–181 (2007).
Bavireddi, H., Bharate, P. & Kikkeri, R. Probing carbohydrate-carbohydrate interactions by photoswitchable supramolecular glycoclusters. Chem. Commun. (Camb.) 49, 3988–3990 (2013).
Matsuura, K., Tsuchida, A., Okahata, Y., Akaike, T. & Kobayashi, K. A quartz-crystal microbalance study of adsorption behaviors of artificial glycoconjugate polymers onto chemically modified gold surfaces and their interactions with lectins. Bull. Chem. Soc. Jpn. 71, 2973–2977 (1998).
de la Fuente, J. M. & Penades, S. Understanding carbohydrate-carbohydrate interactions by means of glyconanotechnology. Glycoconj. J. 21, 149–163 (2004).
Reitsma, S., Slaaf, D. W., Vink, H., van Zandvoort, M. A. & oude Egbrink, M. G. The endothelial glycocalyx: composition, functions, and visualization. Pflug. Arch. 454, 345–359 (2007).
Sheikh, H., Yarwood, H., Ashworth, A. & Isacke, C. M. Endo180, an endocytic recycling glycoprotein related to the macrophage mannose receptor is expressed on fibroblasts, endothelial cells and macrophages and functions as a lectin receptor. J. Cell Sci. 113, 1021–1032 (2000).
Taylor, M. E., Bezouska, K. & Drickamer, K. Contribution to ligand binding by multiple carbohydrate-recognition domains in the macrophage mannose receptor. J. Biol. Chem. 267, 1719–1726 (1992).
Jalvo, B., Mathew, A. P. & Rosal, R. Coaxial poly(lactic acid) electrospun composite membranes incorporating cellulose and chitin nanocrystals. J. Membr. Sci. 544, 261–271 (2017).
Goetz, L. A., Jalvo, B., Rosal, R. & Mathew, A. P. Superhydrophilic anti-fouling electrospun cellulose acetate membranes coated with chitin nanocrystals for water filtration. J. Membr. Sci. 510, 238–248 (2016).
Yu, X. P. & Carlsson, A. E. Kinetics of filament bundling with attractive interactions. Biophys. J. 87, 3679–3689 (2004).
Ham, H. O., Park, S. H., Kurutz, J. W., Szleifer, I. G. & Messersmith, P. B. Antifouling glycocalyx-mimetic peptoids. J. Am. Chem. Soc. 135, 13015–13022 (2013).
Holland, N. B., Qiu, Y., Ruegsegger, M. & Marchant, R. E. Biomimetic engineering of non-adhesive glycocalyx-like surfaces using oligosaccharide surfactant polymers. Nature 392, 799–801 (1998).
Hederos, M., Konradsson, P. & Liedberg, B. Synthesis and self-assembly of galactose-terminated alkanethiols and their ability to resist proteins. Langmuir 21, 2971–2980 (2005).
Anderson, J. M., Rodriguez, A. & Chang, D. T. Foreign body reaction to biomaterials. Semin. Immunol. 20, 86–100 (2008).
Rudra, J. S., Tian, Y. F., Jung, J. P. & Collier, J. H. A self-assembling peptide acting as an immune adjuvant. Proc. Natl Acad. Sci. USA 107, 622–627 (2010).
Oldenbourg, R., Wen, X., Meyer, R. B. & Caspar, D. L. D. Orientational distribution function in nematic tobacco-mosaic-virus liquid-crystals measured by X-ray diffraction. Phys. Rev. Lett. 61, 1851–1854 (1988).
Acknowledgements
This research was supported by NSF Career (DMR-1455201) to G.A.H. and NSF Career (DMR-1352043) to T.E.A. J.K was partially supported by an NIH T32 basic microbiology and infectious diseases training grant (5T32AI007110-34). The authors would like to acknowledge Dr. Mavis Agbandje-McKenna and Dr. Robert McKenna for providing access to the CD spectrometer in the UF Center for Structural Biology and to Dr. Michael Harris for access to the CD spectrometer in the UF Department of Chemistry. Cryo-EM samples were prepared and imaged by Rodolfo Alvarado at the University of Florida ICBR.
Author information
Authors and Affiliations
Contributions
G.A.H. conceived the project, designed experiments, analyzed data, and contributed to the writing and editing of the manuscript. A.R. conducted most of the experiments, analyzed data, prepared figures, and contributed to writing and editing of the manuscript. D.T.S. conducted experiments, analyzed data, and prepared figures. K.K. contributed to FE-SEM experiments. C.S.O. conducted SAXS experiments and together with T.E.A. analyzed SAXS data. J.J.K. contributed to CD experiments. K.R.K. contributed to data analysis. S.A.F. contributed to peptide synthesis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Restuccia, A., Seroski, D.T., Kelley, K.L. et al. Hierarchical self-assembly and emergent function of densely glycosylated peptide nanofibers. Commun Chem 2, 53 (2019). https://doi.org/10.1038/s42004-019-0154-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42004-019-0154-z
This article is cited by
-
Ultrasensitive electrochemical sensor based on synergistic effect of Ag@MXene and antifouling cyclic multifunctional peptide for PD-L1 detection in serum
Microchimica Acta (2024)
-
Multi-responsive chitosan-based hydrogels for controlled release of vincristine
Communications Chemistry (2023)
-
Charge guides pathway selection in β-sheet fibrillizing peptide co-assembly
Communications Chemistry (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.