Abstract
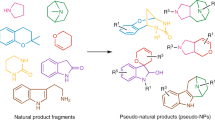
Natural products offer unexplored molecular frameworks for the development of chemical leads and innovative drugs. However, the structural complexity of natural products compared with synthetic drug-like molecules often limits the scaffold hopping potential of natural-product-inspired molecular design. Here we introduce a holistic molecular representation incorporating pharmacophore and shape patterns, which facilitates scaffold hopping from natural products to isofunctional synthetic compounds. This computational approach captures simultaneously the partial charge, atom distributions and molecular shape. In a prospective application, we use four natural cannabinoids as queries in a chemical database search for novel synthetic modulators of human cannabinoid receptors. Of the synthetic compounds selected by the new method, 35% are experimentally confirmed as active. These cannabinoid receptor modulators are structurally less complex than their respective natural product templates. The results of this study validate this holistic molecular representation for hit and lead finding in drug discovery.
Similar content being viewed by others
Introduction
Natural products have inspired numerous pharmacologically active lead compounds that have entered clinical trials1,2,3,4,5,6,7,8,9,10. Natural products possess desirable molecular frameworks as starting points for small molecule drug discovery11 as they contain larger fractions of sp3-hybridized bridgehead atoms, chiral centers and diverse pharmacophores5,9,12. However, the majority of natural products in the Dictionary of Natural Products13 (DNP) do not have immediate synthetic counterparts5. This is partly due to a lack of dedicated research tools and methods to harvest the full potential of natural products for drug discovery, especially for designing ligands when scarce or no target structural information is available. In such a situation, molecular descriptor analysis can support early drug discovery by enabling ligand-based scaffold hopping for hit and lead finding14,15,16.
Molecular descriptors are numerical representations computationally derived from the underlying molecular structure17. Molecular descriptors have been mainly used for reductionist representations that capture certain individual molecular features, such as fragments18 or atom/bond properties19. However, the structural differences between natural and synthetic compounds limit the scaffold hopping potential of these single-feature representations20.
To this end, we have developed a novel molecular representation to transfer relevant structural and pharmacophore information encoded in natural products to synthetically accessible compounds through similarity-based approaches. In contrast to the conventional single-feature descriptors, these molecular descriptors are holistic, capturing many molecular properties, such as geometric interatomic distances, molecular shape, and the partial charge distribution. From this representation, the new Weighted Holistic Atom Localization and Entity Shape (WHALES) descriptors are obtained.
For proof-of-concept, we employ WHALES to prospectively screen a large library of commercially available compounds, using four phytocannabinoids as natural product queries. Based on this computational analysis, seven out of the twenty compounds identified modulate human cannabinoid receptors (CB1, CB2) with low-micromolar potencies, agonistic and antagonistic activity, and different subtype selectivity. Five out of the seven active scaffolds are novel compared to the known cannabinoid receptor ligands from ChEMBL(v23)21 and SureChEMBL22. These results demonstrate that WHALES descriptors capture functionally relevant molecular features and enable scaffold hopping from natural products to bioactive synthetic mimetics.
Results
WHALES descriptors
We designed the WHALES descriptors to encode information on geometric interatomic distances, molecular shape, and atomic properties in a holistic way. The respective molecular feature distributions are computed from locally centered atom distances, drawing inspiration from a recently proposed data analysis method23. For each atom position in a three-dimensional (3D) molecular conformation, the spatial distribution of the surrounding molecular atoms is captured by a weighted atom-centered covariance matrix, which is used to normalize the interatomic distances to account for local feature distributions. The obtained interatomic distances are proportional to the remoteness of each atom from the center of local atomic distributions, measured in variance units. Additionally, to account for potential ligand-receptor interaction patterns (“pharmacophore” features), the contribution of each atom to the atom-centered covariance matrix is weighted by atomistic partial charges, as explained below.
Step 1 Atom-centered covariance matrix calculation
Let X be the matrix of the atom coordinates, containing as many rows as there are non-hydrogen atoms (n) and three columns corresponding to the 3D coordinates of each non-hydrogen atom. The distribution of atoms and their partial charges around any j-th atom is captured using an atom-centered weighted covariance matrix (Sw(j)),
where (xi − xj) are the differences between the 3D coordinates of the j-th atomic center and those of any i-th atom, while |δi| is the absolute value of the partial charge of the i-th atom. The atom-centered covariance is computed for any non-hydrogen atom of the molecule. The weighted covariance matrix is influenced by the density and partial charges of atoms surrounding j. In particular, Sw(j) can be thought of as an ellipsoid centered on j, whose principal axes are oriented in the three orthogonal directions of maximum atom-centered variance; the greater the variance, the longer the corresponding axis of the ellipsoid. This weighted covariance ellipsoid is influenced by (Supplementary Figure 1): (i) the distribution of the atoms surrounding j, since the ellipsoid axes are oriented in the directions of maximal molecular extension; and (ii) the distribution of the atomic properties, which causes a rotation of the atom-centered covariance ellipsoid toward the locations of high absolute partial charge (|δi|) densities.
Step 2 Atom-centered Mahalanobis distance calculation
From Sw(j), the atom-centered Mahalanobis (ACM) distance from the center j to any i-th atom is calculated as follows:
All of the pairwise normalized interatomic distances calculated according to Eq. 2 are collected in the ACM matrix (Fig. 1c): Each i-th row of the matrix represents how the i-th atom is “globally perceived” by other atoms, while each j-th column contains the distances from atom j to all the other atoms, where j itself is the center of the molecular feature space. Thus, a column represents how an atom “globally perceives” all the remaining atoms. Atoms located in the directions of high variance will have a smaller relative distance from the atomic center than atoms located in low-variance regions, e.g., atoms residing off the main molecular axis. Due to the normalization procedure based on Sw(j), the ACM distance is dimensionless and asymmetric.
Overview of the WHALES concept. a Starting from a given molecular graph, 3D energy minimization and partial charge calculation are performed. In this work, MMFF9425 energy-minimized conformation and Gasteiger-Marsili partial charges24 were utilized. b The coordinates and the computed partial charges are used to calculate the atom-centered weighted covariance (Eq. 1). A schematic representation of the centered covariance (blue ellipsoid) is shown for atom C2. The ellipsoid axes represent the directions and magnitude of maximal atom-centered covariance. c The atom-centered covariances are utilized to calculate the ACM distance matrix (Eq. 2). From the ACM, the remoteness (Eq. 3) and isolation degree (Eq. 4) of the j-th atom are calculated as the j-th row average (Avg.) and the j-th column minimum (Min.), respectively. Descriptor values of negatively charged atoms are assigned a negative sign (Eq. 6, not shown in Fig. 1). d WHALES descriptors are calculated as the deciles, the minimum and the maximum of isolation degree (Isol.), remoteness (Rem.) and isolation-remoteness ratio (IR), to obtain a molecular size-independent representation
Step 3 Calculation of atomic indices
From the ACM matrix, three indices are calculated for each atom (Fig. 1c):
-
(1)
Remoteness (Rem), which is the ACM matrix row-average, calculated as follows:
$${\rm Rem}(j) = \frac{{\mathop {\sum}\nolimits_{i = 1}^n {{\rm ACM}(j,i)} }}{{n - 1}},$$(3)where n is the number of non-hydrogen atoms. Remoteness is high for atoms with large ACM distances from many atomic centers (global information);
-
(2)
Isolation degree (Isol), which is the ACM matrix column minimum (excluding the atomic center):
$${\rm Isol}(j) = {\mathrm{min}}_i\left( {{\rm ACM}(i,j)} \right)\quad i\,\≠ \,j$$(4)The isolation degree represents the distance of the j-th object from its nearest atom neighbor. The isolation degree is high for atoms located in “peripheral” regions of the molecule, i.e., atoms are surrounded by a few close atoms (local information);
-
(3)
Isolation-Remoteness ratio, calculated as:
The Isolation-Remoteness ratio (IR) simultaneously accounts for the local and global information of each atom, assuming high values for atoms residing off the main molecular axis (i.e., high-isolation degree) and a small relative distance from most of the atomic centers (i.e., low remoteness).
The remoteness, isolation degree values and their ratio calculated for negatively charged atoms are assigned a negative sign, as follows:
This procedure allows to distinguish positively and negatively charged atoms having the same values of isolation degree and remoteness.
Step 4 WHALES descriptors calculation
Because the number of calculated atomic indices depends on the number of non-hydrogen atoms of the molecule, a binning procedure is applied to obtain a fixed-length representation, enabling the straightforward comparison of molecules with different numbers of atoms. In particular, the WHALES descriptors are calculated as deciles plus minimum and maximum of (i) atomic isolation degrees, (ii) remoteness values, and (iii) isolation/remoteness ratios. Thus, each molecule is characterized by the same number of descriptors (i.e., 11 values for each atomic index, for a total of 33 descriptors), regardless of the number of atoms considered (Fig. 1d). WHALES descriptors are invariant to any roto-translation of molecular coordinates and robust to small conformational changes (Supplementary Figure 2).
For this present proof-of-concept study, Gasteiger-Marsili partial charges24 and MMFF9425 energy-minimized structures were used for WHALES calculations. However, the WHALES descriptors can be computed using any type of energy-minimized structures and partial charge scheme as input, e.g., quantum-chemistry derived partial charges26.
Scaffold hopping from natural products
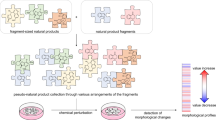
To assess the potential of WHALES for scaffold hopping from natural products, it was compared to extended-connectivity fingerprints18 (ECFPs), which represent the molecule as a set of fragments that are radially grown from each non-hydrogen atom. ECFPs are a benchmark in virtual screening campaigns27 due to their widespread availability in numerous software tools, ease of calculation and intuitiveness to chemists. WHALES and ECFPs were compared to detect differences in their representation of natural products compared to synthetic compounds. To this end, we compared 210,119 entries from the DNP with a set of 3,383,942 commercially available synthetic compounds. Each DNP natural product was used as a query to rank the remaining DNP and commercial compounds on a similarity-basis, using WHALES (Euclidean distance) or ECFP (Jaccard-Tanimoto distance) descriptors. WHALES led to a statistically higher (p < 0.001, Wilcoxon signed-rank test28) number of natural compound synthetic neighbors than ECFPs on average (Fig. 2a). Among the 200 nearest natural product neighbors, an average of 26% of the synthetic compounds were concentrated in the top-20 positions for WHALES, compared to 9% for ECFPs (Fig. 2b). This difference reflects the largely different chemical space representations obtained with the two descriptors (Fig. 2c). WHALES descriptors suggest synthetic compound “bridging regions” that connect clusters of natural products. In contrast, the fragment-based perception of ECFPs leads to a clear separation between synthetic compounds and natural products. This comparative study indicates that WHALES may be better suited for scaffold hopping between natural products and synthetic compounds than ECFPs. Thus, we applied WHALES to a prospective virtual screening on the Cannabinoid Receptor (CB) using natural cannabinoids as queries. Retrospective analysis of WHALES on CB actives annotated in ChEMBL showed that WHALES descriptors have a higher scaffold hopping potential on this target when compared to ECFPs (Fig. 3).
Similarity search using WHALES and ECFPs with natural products as queries. A total of 210,119 NPs were utilized as queries on 3,383,942 commercially available compounds (WHALES = Euclidean distance on Gaussian-normalized values; ECFPs = Jaccard-Tanimoto index). a Percentage of commercially available synthetic neighbors of each DNP natural product according to the selected molecular description (i.e., ECFPs and WHALES). Given portions of the list (i.e., 10, 20, 50, 100, and 200 neighbors) are displayed. Boxplots show median, mean (dot), 1st and 3rd quartiles (solid line), 95th percentile (whisker), and 99th percentile (squares). The average number of neighbors of each NP retrieved from WHALES (p < 0.001, Wilcoxon signed-rank test28) and the median number, up to 50 neighbors (p < 0.001, Kruskal–Wallis H-test29), are significantly larger than those retrieved from ECFPs. b The relative distribution of synthetic neighbors of NPs in the first 200 positions. Several portions of the similarity ranks are considered, as indicated by colors (1–10, 10–20, 20–50, 10–100, and 100–200 neighbors of NP); the larger the bar for a given portion of the list, the larger the average number of synthetic neighbors of NPs in that portion. c Network analysis of a randomly compiled set of 15,000 natural products (green) and 15,000 synthetic compounds (red); lines represent similarity relationships between the compounds (circles), which are colored according to their type (natural or synthetic compounds, respectively in green and red). Left: minimum spanning tree obtained with ECFPs; right: minimum spanning tree obtained with WHALES
Retrospective analysis of ECFP and WHALES scaffold hopping abilities on known cannabinoid receptor actives. Experimental activity values on CB1 and CB2 were retrieved from ChEMBL (v23). Active compounds (EC50, IC50, Ki or Kd ≤10 µM) were used as queries to rank the remaining compounds on a similarity basis (ECFP: Jaccard-Tanimoto index; WHALES: Euclidean distance). For each rank, the relative scaffold diversity of actives was computed as the number of unique scaffolds32 present in the actives of the top 1% list over the total number of actives found in the top 1% list. Boxplots show median (black line), mean (white dot), 1st and 3rd quartiles (lines), 5th and 95th percentiles (whiskers); gray dots represent the raw values
Prospective virtual screening
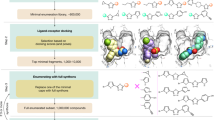
For prospective screening, we selected four of the most abundant active constituents of the cannabis plant (Cannabis sativa) as queries29, namely (Fig. 4): (1) (-)-trans-∆9-tetrahydrocannabinol (THC), (2) (-)-cannabidiol (CBD), (3) (-)-cannabinol (CBO), and (4) (-)-trans-∆9-tetrahydrocannabivarin (THCV). 1 and 3 act as agonists or partial agonists on CB1 and CB2 in vitro. Compound 4 shows dose-dependent agonism on CB2 and CB1 in vivo, respectively, while the mechanism of action of 2 is still under debate29,30,31. Each phytocannabinoid was used in turn to perform a similarity-based virtual screening on the commercial library, with the Euclidean distance calculated on WHALES descriptors as a ranking criterion. The compounds were sorted according to the sum of their reciprocal ranks obtained with each query. The 20 top-ranked synthetic compounds were selected and, without any additional exclusion criteria applied, tested in vitro for their modulatory activity on human CB1 and CB2 receptors.
Natural product queries (1–4) and novel CB modulators (5–11). In vitro activities are reported in Table 1
The WHALES-based virtual screening protocol led to the identification of seven active compounds (35% of the selected compounds), with activity values (EC/IC50 and KB) in the low micro- or nanomolar range and different selectivity profiles (Table 1). Scaffold analysis of the core rings and atomic frameworks32 of the synthetic hits revealed that five out of the seven actives not only differ in their structure from the natural product queries, but they also possess a novel scaffold that is not contained in any of the CB actives (EC/IC50 or Ki/D < 50 μM, 6188 compounds) annotated in ChEMBL21 or in the patent literature (SureCHEMBL)22 (Fig. 5). This result demonstrates that the WHALES method is suitable for retrieving isofunctional synthetic mimetics of bioactive natural products.
Scaffold analysis of known CB ligands from ChEMBL. The most frequently occurring atomic frameworks (Murcko scaffolds)32 in all actives on CB1 and CB2 annotated in ChEMBL23 (EC50, IC50, Ki, KD < 50 μM; 6188 compounds). Only the scaffold of 8 and 9 was present in the CB actives annotated in ChEMBL
Among the novel actives, one non-selective agonist (5, CB1: EC50 = 3.1 ± 0.5 μM; CB2: EC50 = 1.8 ± 0.6 μM) and three selective CB1 agonists (6, EC50 = 4.3 ± 0.7 μM; 7, EC50 > 30 μM; 9, EC50 = 1.0 ± 0.2 μM) were identified. These hits inherited the prevalent agonistic activity from the utilized natural cannabinoids with different selectivity profiles. Computational ligand docking (Fig. 6) suggests that 5 and 6 might act through similar binding poses and interaction patterns to their most similar natural-product templates according to WHALES (THCV [4] and THC [1], respectively). The non-selective antagonist 8 (CB1: IC50 = 10.1 ± 0.7 μM, KB = 8.8 μM; CB2: IC50 = 27.0 ± 0.8 μM, KB = 1.8 μM) and two selective CB1 antagonists (10, IC50 = 3.2 ± 0.5 μM, KB = 0.9 μM; 11, IC50 = 1.3 ± 0.2 μM, KB = 0.2 μM) were also identified.
Predicted binding poses of non-selective agonist 5 and CB1-selective agonist 6 in CB1/CB2 active sites. CB1: PDB-ID = 5XRA; CB2: homology model. The hits were compared with their most similar NP according to WHALES. Docking was performed with MOE on MMFF49x energy-minimized structures, which were ranked and refined by London dG and Alpha HB scores36. Key interactions are shown with dashed lines. a Active compound 6 (light blue) in comparison with THCV (4, green) in the active site of CB1; b active compound 5 (orange) in comparison with THCV (4, green) in the in the active site of CB1; c active compound 5 (orange) in comparison with THC (1, green) in the modeled active site of CB2
The similarity of the predicted binding poses of 5 and 6 to their natural product templates highlights that WHALES descriptors did indeed capture the pharmacophore of phytocannabinoids in terms of shape and partial charge distributions. At the same time, the presence of active hits with antagonistic activity and/or presumably novel receptor pocket interaction patterns demonstrate that the WHALES representation is sufficiently flexible to allow for the discovery of novel ligand-binding motifs. This is due to the “fuzziness” of the WHALES descriptors, which represent molecules by how their pharmacophore properties are distributed in 3D space without any explicit fragment, ring system or atom type information. Considering commercial building block availability, retrosynthetic analysis suggests that the bioactive hits can be prepared in three or fewer steps and are thus more easily synthetically accessible than the natural product queries (Supplementary Figure 3).
The screening library ranks obtained by considering only molecular shape (i.e., WHALES without any charge-based weighting, Eq. 1) or only charge (i.e., deciles of Gasteiger-Marsili partial charges) have a low correlation with those obtained by WHALES (Kendall rank correlation coefficient τ < |0.08|). None of the active hits were scored in the top 1000 of the screening compounds with the shape-only and charge-only descriptions. These results confirm the holistic character of WHALES descriptors, which grasp “emergent” structural features of NPs that cannot be captured by describing single aspects separately.
To assess the ability of WHALES to identify novel actives compared with existing tools, we compared the prospective virtual screening results with six common molecular descriptors (ECFP18, FeatMorgan11, RDKit33, MACCS 16634, AtomPair35 fingerprints, CATS15) and four pharmacophore screening protocols (MOE pharmacophore search36, LigandScout37 ligand-based pharmacophore search, ShAEP38, UFSRAT39). The virtual screening protocol was performed starting from the natural product queries (1–4) on the commercial screening library, using the benchmark methods and the same ranking protocol as in the productive WHALES run (Supplementary Note 1). None of the novel active hits discovered by WHALES were scored in the top 100 lists obtained with any of these alternative methods (Supplementary Table 2). This outcome clearly supports the use of WHALES in medicinal chemistry workflows for the discovery of novel active scaffolds.
Discussion
The results of this study demonstrate the suitability of this holistic virtual screening method for scaffold hopping from natural products to isofunctional synthetic compounds. The WHALES-based molecular representation bridges the gap between natural product and synthetic compound chemical spaces and leads to “bridging regions” of synthetic compounds that connect clusters of natural products. With 35% of the top-ranked compounds exhibiting low-micromolar in vitro activities, WHALES is at least competitive with other screening protocols. Importantly, WHALES proved suitable for retrieving novel active compounds and scaffolds that were not found by other methods for similarity searching. The cannabinoid receptor modulators obtained are structurally less complex than the natural product templates. These results clearly highlight the effectiveness of this novel holistic approach to harvest the potential of natural products by obtaining synthetically accessible, natural product-inspired bioactive compounds and to explore uncharted chemical space regions.
Methods
Compound preparation pre-processing and descriptor calculation
Compound structures were de-salted and protonated (considering a pH = 7) prior to descriptors calculation. Molecular geometry was optimized using the MMFF9425 force field with 1000 iterations and 10 starting conformers for each compound; the minimum energy conformation was used for subsequent descriptor calculation. Gasteiger-Marsili24 partial charges were computed using the RDKit module33.
Preliminary analysis
Extended-connectivity fingerprints18 (ECFPs) were calculated using Dragon 740 (size = 1024 bit; 2 bits per pattern, length = 0–4 bonds). Prim’s41 minimum spanning trees were generated on 15,000 DNP and 15,000 commercial non-overlapping compounds, which were selected randomly. Molecular scaffolds were defined according to Bemis-Murcko32 molecular frameworks using the RDKit module33.
Commercial library
The library was assembled from commercially available synthetic compounds from four providers: Asinex (http://www.asinex.com/libraries-html/) (Elite, Fragments, Gold and Platinum collections), ChemBridge screening compound collection (http://www.chembridge.com), Enamine advanced and HTS collections (http://www.enamine.net), and Specs screening compounds (https://www.specs.net).
Prospective screening
Phytocannabinoid structures were retrieved from the scientific literature. Structure optimization and descriptor calculation were performed as explained above. Each query was used to perform virtual screening based on Euclidean distance on Gaussian-normalized WHALES values. The virtual screening results of each commercial library compound were merged and sorted according to the sum of their reciprocal ranks on each query. The top-20 screening compounds were purchased from ChemBridge, Enamine, and Specs.
In vitro biological characterization
Screening compounds were purchased and assayed in vitro for agonism and antagonism on cannabinoid receptors CB1 and CB2 in functional test systems. For agonistic characterization, CHO cells over-expressing the respective human GPCR were incubated with varying concentrations of each compound for 20 min and cAMP response was quantified by homogenous time-resolved FRET (HTRF). For antagonistic characterization, varying concentrations of the test compounds in competition with a fixed agonist concentration were used. CP55940 (CB1 agonist, EC50 = 0.035 nM), WIN55212-2 (CB2 agonist, EC50 = 0.21 nM), AM281 (CB1 antagonist, IC50 = 10 nM) and AM630 (CB2 antagonist, IC50 = 0.9 µM) served as reference compounds. For each test compound concentration, a relative cAMP response compared to the respective reference compound was recorded. All experiments were independently repeated at least twice, and results were reported as the mean ± standard error. EC/IC50 values were calculated from dose-response curves using a four-parameter nonlinear regression (Supplementary Figure 4). These assays were performed by Cerep (Celle-L’Evescault, France; www.eurofinsdiscoveryservices.com; assay reference numbers 1744, 1745, 1746, 1747) on a fee-for-service basis.
Docking and homology modeling
The crystal structure of human CB1 in complex with agonist AM11542 (PDB-ID: 5XRA)42 was prepared for docking in MOE (v2016.0802)36. Energy minimization was performed using the Amber10:EHT force field. For each ligand, 60 poses were generated, their energy was minimized using MMFF94x force field within a rigid receptor, and they were ranked by London dG score36. The ten top-scoring poses were refined, re-scored using Alpha HB scoring, and visually inspected. Re-docking of the crystallized ligand led to a small RMSD value (0.39 Å). A homology model of CB2 (UniProt ID: P34972) was obtained with MODELLER43, using the prepared CB1 structure as the template. The initial template and target alignment was obtained by Muscle44 and then manually adjusted (Supplementary Figure 5). The ligand was retained to consider induced fit effects.
Data and code availability
The authors declare that the data supporting the findings of this study are available within the paper and its supplementary information. Python code implementing WHALES descriptors is deposited as an open source repository on GitHub (https://github.com/grisoniFr/whales_descriptors.git).
Change history
20 September 2018
The original PDF and HTML versions of this Article contained errors in Eqs. (4) and (6). In Eq. (4), the not equal symbol displayed incorrectly as #, and in Eq. (6), the greater than symbol was displayed in place of the less than symbol. This has been corrected in both the PDF and HTML versions of the Article.
References
Molinari, G. Natural products in drug discovery: present status and perspectives. in Pharmaceutical Biotechnology 13–27 (Springer, New York 2009).
Koehn, F. E. & Carter, G. T. The evolving role of natural products in drug discovery. Nat. Rev. Drug Discov. 4, 206–220 (2005).
Patridge, E., Gareiss, P., Kinch, M. S. & Hoyer, D. An analysis of FDA-approved drugs: natural products and their derivatives. Drug Discov. Today 21, 204–207 (2015).
Harvey, A. L. Natural products in drug discovery. Drug Discov. Today 13, 894–901 (2008).
Lee, M. L. & Schneider, G. Scaffold architecture and pharmacophoric properties of natural products and trade drugs: application in the design of natural product-based combinatorial libraries. J. Comb. Chem. 3, 284–289 (2001).
Brown, D. G., Lister, T. & May-Dracka, T. L. New natural products as new leads for antibacterial drug discovery. Bioorg. Med. Chem. Lett. 24, 413–418 (2014).
van Hattum, H. & Waldmann, H. Biology-oriented synthesis: harnessing the power of evolution. J. Am. Chem. Soc. 136, 11853–11859 (2014).
Rodrigues, T., Reker, D., Schneider, P. & Schneider, G. Counting on natural products for drug design. Nat. Chem. 8, 531–541 (2016).
Harvey, A. L., Edrada-Ebel, R. & Quinn, R. J. The re-emergence of natural products for drug discovery in the genomics era. Nat. Rev. Drug Discov. 14, 111–129 (2015).
Grabowski, K., Baringhaus, K.-H. & Schneider, G. Scaffold diversity of natural products: inspiration for combinatorial library design. Nat. Prod. Rep. 25, 892–904 (2008).
Morrison, K. C. & Hergenrother, P. J. Natural products as starting points for the synthesis of complex and diverse compounds. Nat. Prod. Rep. 31, 6–14 (2014).
Henkel, T., Brunne, R. M., Müller, H. & Reichel, F. Statistical investigation into the structural complementarity of natural products and synthetic compounds. Angew. Chem. Int. Ed. 38, 643–647 (1999).
Dictionary of Natural products (DNP) database, v20.1. CRC Press, Taylor & Francis, http://dnp.chemnetbase.com. (2011).
Martin, E. J. & Critchlow, R. E. Beyond mere diversity: tailoring combinatorial libraries for drug discovery. J. Comb. Chem. 1, 32–45 (1999).
Reutlinger, M. et al. Chemically advanced template search (CATS) for scaffold-hopping and prospective target prediction for ‘orphan’molecules. Mol. Inf. 32, 133–138 (2013).
Grisoni, F. et al. Matrix-based molecular descriptors for prospective virtual compound screening. Mol. Inf. 36, 1600091 (2017).
Todeschini, R. & Consonni, V. Molecular Descriptors for Chemoinformatics. (Wiley VCH, Weinheim, 2009).
Rogers, D. & Hahn, M. Extended-connectivity fingerprints. J. Chem. Inf. Model. 50, 742–754 (2010).
Bade, R., Chan, H.-F. & Reynisson, J. Characteristics of known drug space. Natural products, their derivatives and synthetic drugs. Eur. J. Med. Chem. 45, 5646–5652 (2010).
Rodrigues, T., Reker, D., Kunze, J., Schneider, P. & Schneider, G. Revealing the macromolecular targets of fragment-like natural products. Angew. Chem. Int. Ed. 54, 10516–10520 (2015).
Gaulton, A. et al. ChEMBL: a large-scale bioactivity database for drug discovery. Nucleic Acids Res. 40, D1100–D1107 (2011).
Papadatos, G. et al. SureChEMBL: a large-scale, chemically annotated patent document database. Nucleic Acids Res. 44, D1220–D1228 (2015).
Todeschini, R., Ballabio, D., Consonni, V., Sahigara, F. & Filzmoser, P. Locally centred Mahalanobis distance: a new distance measure with salient features towards outlier detection. Anal. Chim. Acta 787, 1–9 (2013).
Gasteiger, J. & Marsili, M. Iterative partial equalization of orbital electronegativity — a rapid access to atomic charges. Tetrahedron 36, 3219–3228 (1980).
Halgren, T. A. Merck molecular force field. I. Basis, form, scope, parameterization, and performance of MMFF94. J. Comput. Chem. 17, 490–519 (1996).
Finkelmann, A. R., Göller, A. H. & Schneider, G. Robust molecular representations for modelling and design derived from atomic partial charges. Chem. Commun. 52, 681–684 (2016).
Vogt, M., Stumpfe, D., Geppert, H. & Bajorath, J. Scaffold hopping using two-dimensional fingerprints: true potential, black magic, or a hopeless endeavor? Guidelines for virtual screening. J. Med. Chem. 53, 5707–5715 (2010).
Wilcoxon, F. Individual comparisons by ranking methods. Biom. Bull. 1, 80–83 (1945).
Pertwee, R. The diverse CB1 and CB2 receptor pharmacology of three plant cannabinoids: Δ9-tetrahydrocannabinol, cannabidiol and Δ9-tetrahydrocannabivarin. Br. J. Pharmacol. 153, 199–215 (2008).
Turner, S. E., Williams, C. M., Iversen, L. & Whalley, B. J. Molecular pharmacology of phytocannabinoids. in Phytocannabinoids: Unraveling the Complex Chemistry and Pharmacology of Cannabis sativa eds. (Kinghorn, A. D., Falk, H., Gibbons, S. & Kobayashi, J.) 61–101 (Springer International Publishing, 2017).
Martínez-Pinilla, E. et al. Binding and signaling studies disclose a potential allosteric site for cannabidiol in cannabinoid CB2 receptors. Front. Pharmacol. 8, 244 (2017).
Bemis, G. W. & Murcko, M. A. The properties of known drugs. 1. Molecular frameworks. J. Med. Chem. 39, 2887–2893 (1996).
RDKit: Open-source cheminformatics; http://www.rdkit.org (2017).
MACCS-II, MDL Information Systems Inc, San Leandro, CA, USA (1987).
Carhart, R. E., Smith, D. H. & Venkataraghavan, R. Atom pairs as molecular features in structure-activity studies: definition and applications. J. Chem. Inf. Comput. Sci. 25, 64–73 (1985).
Chemical Computing Group ULC. Molecular Operating Environment (MOE), 2013.08. Montreal, QC, Canada, H3A 2R7. (2017).
Wolber, G. & Langer, T. LigandScout: 3-D pharmacophores derived from protein-bound ligands and their use as virtual screening filters. J. Chem. Inf. Model. 45, 160–169 (2005).
Vainio, M. J., Puranen, J. S. & Johnson, M. S. ShaEP: molecular overlay based on shape and electrostatic potential. J. Chem. Inf. Model. 49, 492–502 (2009).
Shave, S. et al. UFSRAT: ultra-fast shape recognition with atom types–the discovery of novel bioactive small molecular scaffolds for FKBP12 and 11βHSD1. PLoS ONE 10, e0116570 (2015).
Kode srl. Dragon version 7.0.6, 2016, https://chm.kode-solutions.net (2016).
Prim, R. C. Shortest connection networks and some generalizations. Bell Labs Tech. J. 36, 1389–1401 (1957).
Hua, T. et al. Crystal structures of agonist-bound human cannabinoid receptor CB1. Nature 547, 468–471 (2017).
Fiser, A. & Šali, A. Modeller: generation and refinement of homology-based protein structure models. Methods Enzymol. 374, 461–491 (2003).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Acknowledgements
The authors thank Petra Schneider and Cyrill Brunner for technical support. This research was financially supported by the Swiss National Science Foundation (grant no. IZSEZ0_177477 to G.S.). D.M. was supported by an ETH Zurich Postdoctoral Fellowship (grant no. 16-2 FEL-07).
Author information
Authors and Affiliations
Contributions
F.G. and G.S. designed the study. F.G. developed WHALES with the support of V.C., R.T., and G.S. and performed the analysis. F.G., G.S., J.A.H., and D.M. analyzed and discussed the results. D.M. curated the experimental results. F.G. performed the docking study with the support of S.G.T.; S.G.T. developed the homology model. F.G. and G.S. wrote the manuscript. All authors contributed to manuscript revision and approved the final version.
Corresponding author
Ethics declarations
Competing interests
G.S. declares a potential financial conflict of interest in his role as life science industry consultant and cofounder of inSili.com GmbH, Zurich. All the remaining authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Grisoni, F., Merk, D., Consonni, V. et al. Scaffold hopping from natural products to synthetic mimetics by holistic molecular similarity. Commun Chem 1, 44 (2018). https://doi.org/10.1038/s42004-018-0043-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42004-018-0043-x
This article is cited by
-
Artificial intelligence for natural product drug discovery
Nature Reviews Drug Discovery (2023)
-
Structural derivatization strategies of natural phenols by semi-synthesis and total-synthesis
Natural Products and Bioprospecting (2022)
-
Identifying new topoisomerase II poison scaffolds by combining publicly available toxicity data and 2D/3D-based virtual screening
Journal of Cheminformatics (2019)
-
Automated de novo molecular design by hybrid machine intelligence and rule-driven chemical synthesis
Nature Machine Intelligence (2019)
-
Scaffold-Hopping from Synthetic Drugs by Holistic Molecular Representation
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.