Abstract
Light-driven proton pumps, such as proteorhodopsin, have been proposed as an energy source in the field of synthetic biology. Energy is required to power biochemical reactions within artificially created reaction compartments like proto- or nanocells, which are typically based on either lipid or polymer membranes. The insertion of membrane proteins into these membranes is delicate and quantitative studies comparing these two systems are needed. Here we present a detailed analysis of the formation of proteoliposomes and proteopolymersomes and the requirements for a successful reconstitution of the membrane protein proteorhodopsin. To this end, we apply design of experiments to provide a mathematical framework for the reconstitution process. Mathematical optimization identifies suitable reconstitution conditions for lipid and polymer membranes and the obtained data fits well to the predictions. Altogether, our approach provides experimental and modeling evidence for different reconstitution mechanisms depending on the membrane type which resulted in a surprisingly similar performance.
Similar content being viewed by others
Introduction
Methodologies from biology and engineering are combined in the bottom-up approach in synthetic biology1,2, aiming at building a biological system with a desired functionality from the bottom by using dedicated building blocks. An example is the design and construction of artificial proto-cells, cell-like objects exhibiting fundamental functionalities2. Despite the low complexity of proto-cells, their design and implementation requires in-depth knowledge about cellular machineries and their assembly. This knowledge could ultimately lead to the development of synthetic systems not found in nature, which can be utilized for example in industrial production of biotechnological goods or pharmaceuticals2,3. An important biomedical application of such synthetic systems is mimicking fundamental metabolic processes3,4,5. Within cells, reactions often occur inside specialized compartments, where membrane proteins mediate the transport of substrates and products6. The membrane protein forms a pore and allows passive diffusion up to a specific size. Larger and more complex molecules or gated processes, which only allow passage upon a certain criterion (e.g., voltage or molecular recognition), usually require a source of energy.
Building artificial proto-cells requires reconstitution of membrane proteins into the compartment’s membrane to facilitate exchange of metabolites. Different approaches and techniques have evolved to reconstitute them into lipid membranes. The research resulted in an applicable framework for detergent mediated reconstitutions into liposomes which is still used in variations today7,8. For reconstitutions into polymer membranes, which structurally resemble lipid membranes but are composed of amphiphilic polymers no such framework exists making the reconstitution of functional proteins challenging. Only brief guidelines have been proposed and the potentially very different interactions between polymers, detergents and proteins compared to lipid-based systems have not been studied in detail9, underlining the need for a detailed comparative study and formulation of a comprehensive framework10. Two highly important requirements need to be fulfilled to apply this concept in engineering: reproducibility and predictability. So far, the proposed approaches fail to satisfy at least one of these requirements. Design of experiments (DoE) is a method, which has emerged in the 1930s11. The underlying idea is to devise an experimental plan that samples a given parameter space optimally and thus keeps the number of experimental runs low and uses them efficiently. In contrast to the common one-factor-at-a-time method (OFAT or OVAT), which keeps factors constant and varies only one, DoE can identify interactions and requires fewer runs with the same precision in estimating the factors’ effects. Subsequently, the whole parameter space is interpolated via linear model regression. The derived model can in turn be optimized to find experimental conditions that yield a predicted response.
Within the past two decades, block copolymers have emerged as a synthetic alternative to natural phospholipids as membrane building blocks3,12,13. These polymers assemble into similar structures as phospholipids, but they can have a much higher molecular weight, longer chainlength and therefore be more robust3,14. The amphiphilic block copolymers are composed of a hydrophobic and one (AB-type, diblock) or two (ABA-type, triblock) hydrophilic blocks. Hence, similar to lipids block copolymers can self-assemble to form spherical particles, worm-like structures, hollow vesicles, and planar membrane15,16. In contrast to lipids, polymers can be adjusted to meet specific needs: the membrane thickness, rigidity, and permeability can be controlled by tuning the block length and the hydrophilic to hydrophobic block ratio3,15. Beside poly(butadiene)-block-poly(ethylene oxide) (PB-PEO) diblock copolymers, poly(2-methyloxazoline)-block-poly-(dimethysiloxane) (PMOXA-PDMS) diblock or poly(2-methyloxazoline)-block-poly-(dimethysiloxane)-block-poly(2-methyloxazoline) (PMOXA-PDMS-PMOXA) triblock copolymers are commonly used for self-assembly involving proteins or other biomolecules. Their low glass transition temperature and resulting flexibility, as well as lateral diffusion properties make them good candidates for the reconstitution of membrane proteins3. The combination of polymer membranes (based on e.g. PDMS-PMOXA, PB-PEO, or other polymer blocks3) and the efficiency and selectivity of biological components such as enzymes and membrane proteins combines the “best of both worlds” and can be exploited towards building synthetic nanoscale devices3. Such molecular factories can be envisioned performing enzymatic production or degradation of specific compounds (antibiotics, etc.) or take over a desired functionality17. Examples described in the past decade were facilitated by the progress of structural biology and the derived methods for membrane protein reconstitution18,19,20,21,22,23. Yet, in case of polymer systems progress was slower due to their lower prominence and commercial availability. Thus far, mainly robust membrane proteins have been used for reconstitution in polymer membranes and the goal to combine biological with synthetic parts from chemistry has only partly been achieved3.
More sophisticated systems require the presence of an energy source such as the generation and upkeep of proton gradients in addition to the reconstitution in the synthetic membrane. Bacteriorhodopsin (BR) and proteorhodopsin (PR) are well-known for their ability to form proton gradients upon illumination24,25,26. BR has been extensively studied over the years, from solving its crystal structure to using it as a light-triggered conductor22. PR has a similar structure to BR but is more accessible to genetic engineering and can be easily expressed in Escherichia coli. Examples from possible modifications are the adjustment of its absorption wavelength, integration of a chemical on/off-switch and the attachment of a hydrophilic protein to guide the orientation during insertion into the membrane, which is crucial for its functionality27,28,29,30.
In this work, we employ DoE to optimize reconstitution of a proteorhodopsin-green fluorescent protein fusion protein (PR-GFP) into liposomes and polymersomes. GFP guides the orientation during the insertion process into preformed vesicles due to its hydrophilic nature, which impedes passage through the hydrophobic part of the membrane29. Additionally, GFP’s fluorescence enables detection of the protein in the resulting assemblies. 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC) and PMOXA17-PDMS65-PMOXA17 (ABA) are used as phospholipid or block copolymer building blocks3,31, whereby DOPC serves as a benchmark and allows a direct comparison of the two systems. The proteovesicle formation under varying pH values, detergent (n-octyl-β-d-glucopyranoside, OG) and PR-GFP concentrations is investigated in a first step to identify factor combinations that yield uniform vesicles containing PR-GFP. Subsequently, PR-GFP’s function is assessed. Based on the data obtained, the reconstitution is optimized for DOPC liposomes and ABA polymersomes to yield fully functional proteovesicles.
Results
Workflow and overview
Within this project, we used a definitive screening design (DSD) proposed recently by Jones and Nachtsheim32,33, which focuses on efficiency and reduces the number of experimental trials. In contrast to the classical sequential approach (screening, effect estimation, and optimization), it is possible to apply a one-step screening and optimization to the process of interest (Fig. 1). Additionally, the factors’ significance was estimated via stepwise regression and only significant factors were kept in the model equations33. The factors lipid/polymer-to-protein ratio (LPR or PPR, w/w), detergent concentration (OG) and pH value were determined to be critical for a functional reconstitution of PR-GFP7,8. The buffer composition and especially the salt concentration can have a strong impact on the vesicle formation and stability of the protein in solution, too high or too low concentrations would lead to agglomeration and loss-of-protein during its reconstitution34,35. Here, 150 mM KCl is used throughout the experiments which was determined to be suitable for PR-GFP17,29. The size of the formed vesicles was determined via dynamic light scattering (DLS) and fluorescent correlation spectroscopy (FCS) and was used as one response. As FCS only detects objects associated with PR-GFP, we could potentially detect different vesicle populations. Evidence for a good correlation of sizes obtained by DLS/FCS and micrographs haven been presented for liposomes36,37 and PDMS-PMOXA polymersomes38,39,40,41, even though direct methods like (cryo-)TEM are considered more precise. The polydispersity index (PdI), obtained from cumulants analysis, was utilized as a measure for homogeneity. In order to measure PR-GFP’s proton pumping capability, we encapsulated the pH-responsive fluorescent dye pyranine and calculated the pH change over time during illumination via the change of fluorescence intensity23.
Schematic visualization of the workflow using design of experiments to achieve a functional reconstitution. The assembly of phospholipids or triblock copolymers together with membrane proteins is an induced self-assembly process and the resulting structure depends on the starting conditions. A priori knowledge about the factors (e.g., buffer conditions, protein concentration, and membrane concentration) is usually lacking and optimal results cannot be achieved. Having two different membrane building blocks further increases complexity. (I) By using DoE and defining influential factors along with characteristic responses one can devise an experimental plan to investigate the systems behavior in resource-efficient way. (II) Here, the effect of the factors pH value, detergent concentration and membrane-to-protein ratio on the proteovesicles’ characteristics size, homogeneity, and functionality was investigated. (III) The results were used to fit a model for detailed analysis of the process and subsequent optimization, which allowed to find optimal assembly conditions to yield the desired functional proteoliposomes and proteopolymersomes
The screening was split in two steps and the corresponding experimental plans for each run can be found in Supplementary Tables 1-3. Assuming that the formation of a proton gradient should work best with PR-GFP reconstituted in large unilamellar vesicles, we investigated the vesicle formation via film rehydration in the presence of PR-GFP for DOPC and ABA7,42. These results defined a parameter region, which fulfilled the predetermined criteria for size and homogeneity. In a second experimental run, this region of interest was investigated in more detail and the proton pumping activity was measured. Combining both datasets for the regression, a second-degree polynomial model in the form of
with y being a response, x a factor and β a coefficient, was derived for each response. The first sum represents all linear terms, the second all quadratic terms and the last sum the interaction terms. The multi-response optimization was carried out with the help of corresponding desirability functions43. We carried out an optimization toward maximum size, homogeneity, and proton pumping activity.
Formation of DOPC and ABA proteovesicles
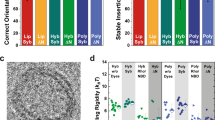
In the first round of experiments, the OG concentration was varied between 0.5 and 2% and a clear tendency toward bigger sizes at detergent concentrations above 1% is observed in case of the proteoliposomes (Fig. 2a). The observed sizes are well below the pore size of the last extrusion step (200 nm). This effect is pronounced at pH 6 and 8, as well as low LPR values. However, at pH 6 and LPR 80/0.05 mg/mL PR-GFP, the measured sizes diverge clearly (Fig. 2a). The objects detected by FCS are in the range of 45 nm whereas DLS detects objects around 110 nm. A similar difference is seen at pH 6, LPR 135/0.03 mg/mL PR-GFP, as well as at pH 7, LPR 25/0.16 mg/mL PR-GFP and 80/0.05 mg/mL PR-GFP, albeit not as pronounced as at lower pH values. This indicates the presence of two populations and a tendency of PR-GFP to partition into smaller sized structures. Imaging the sample via transmission electron microscopy (TEM) confirmed the presence of two populations (Supplementary Figure 5). Estimating the median number of PR-GFP per proteoliposome via FCS provides further evidence (Supplementary Figure 21 and Supplementary Tables 14 and 15). At pH 6, LPR 80 and 0.5% OG, only a median value of 18 PRG-FP molecules (first quartile (Q1): 2/third quartile (Q3): 30) is detected whereas at pH 7, LPR 80 and 1.25% OG 32 PR-GFP molecules (Q1: 20/Q3: 54) are detected.
Interaction plots of the proteovesicle formation. The interaction plots show the behavior of formed (a) DOPC proteoliposomes and b ABA proteopolymersomes toward changing reconstitution conditions. The data are grouped by pH value, different LPR or PPR values are indicated by the data point’s symbol and the used OG concentration are presented on the x-axis. The dotted lines show a potential trend-line and serve as a guide to the eye only. Error bars represent the standard error of the mean, n = 3. DLS, FCS and exemplary TEM data can be found in Supplementary Figures 1, 3, 5 and 6
The proteopolymersomes’ formation at various conditions is different from the liposomes (Fig. 2b). First of all, the observed sizes are smaller, ranging from 30 to 100 nm. Second, we observed that the sizes determined by FCS and DLS disagree stronger compared to the liposomes (Fig. 2b). At pH 6, 1.25% OG and PPR 135/0.03 mg/mL PR-GFP, FCS reports sizes of 51 ± 26 nm on average whereas DLS estimates 90 ± 11 nm. Similar to the observations made with the proteoliposomes, the results indicate two vesicle populations and the partitioning of PR-GFP into the smaller population. TEM shows an increase of small spherical objects as seen in Supplementary Figure 6. It should be noted that the preparation method used in this work always lead to the formation of small micelle-like objects. However, in contrast to the proteoliposomes, the phenomenon is pronounced at medium to high detergent concentration (1.25% and 2% OG) and nearly disappears at pH 8, as can be seen at pH 8, PPR 25 and 1.25% OG. The number of PR-GFP molecules increases to 31 (Q1: 21/Q3: 59), which is a threefold increase compared to the 11 PR-GFP molecules (Q1: 2/Q3: 46) detected at pH 6, 1.25% OG and PPR 135/0.03 mg/mL PR-GFP (Supplementary Figure 22).
The PdI value was used as an indicator for the homogeneity of the vesicle populations. At low LPR values, acidic and basic conditions, the proteoliposomes are more homogeneous (Fig. 2a). At neutral pH no clear trend can be observed. Contrary to the liposomes, the pH has a strong influence on the PdI of the proteopolymersomes (Fig. 2b). At pH 6, values range from 0.17 to 0.33 whereas at pH 8, it only slightly deviates around 0.17.
The measurements from the second round of experiments (Supplementary Table 2) fit within the trends observed in the first round experiments (compare Fig. 2 and Supplementary Figures 8 and 9). The data for the size and the PdI were combined with the first set of experiments to increase the accuracy of the model (Supplementary Tables 4-7). Moreover, we used the models derived from the first set of experiments to predict the outcome of the second one and to validate our approach. The size of PRGFP-containing liposomes increases with higher amounts of OG at pH 6 and pH 8, whereas at pH 7 the size remains stable (Supplementary Figure 8). A low LPR benefits the formation of larger proteoliposomes.
The trend that increasing pH values are beneficial for the formation of larger proteopolymersomes is further supported by additional data, including new pH values fitting into the new experimental boundaries (Supplementary Figure 9). The largest proteopolymersomes containing PRGFP are obtained at a pH range from 7.25 to 8. Similarly, the new data points of the PdI measurement fit as anticipated (Supplementary Figure 8 and 9). A low LPR is beneficial for a low PdI when reconstituting PR-GFP into DOPC lipsomes. Across all measured pH values, the lowest PdI was obtained when a LPR of 25/0.16 mg/mL PR-GFP was used. Contrary to the proteoliposomes, the pH value has a bigger influence on the PdI of the proteopolymersomes, with the lowest values being present at pH >7 (Supplementary Figure 9). The upcoming analysis of the model fitted to the data will offer some explanations for the observed phenomena.
Model analysis
A second-degree polynomial model (Equation 1) was fitted to the two datasets via forward stepwise regression, allowing a deeper analysis of the behavior of the two membrane systems (Fig. 3). From a modeling perspective it should be noted that the variance is higher in case of the polymersomes and cannot be explained well by the model (compare Supplementary Tables 8, 9 and 11, 12). The formation of proteopolymersomes under the tested conditions results in a higher variety of sizes that cannot be explained purely by batch-to-batch variations as the designated blocking variables in the model are mostly insignificant (blocking variables represent any non-controllable environmental condition which can have an influence on the experiment). Our approach to select a sub-region of the parameter space which yields well-formed vesicles (illustrated in Fig. 3, see also Table 1) allowed us to verify predictions based on the models obtained in the first DSD. Comparing the overall means of the responses to the predictions (Supplementary Figure 18) results in no statistical significant difference. Looking at the two membrane systems it can be concluded that the formation of proteoliposomes and proteopolymersomes is different (Table 2). Only one combination, namely pH 8, LPR 135/0.03 mg/mL and 0.5% OG, resulted in nearly identical results for lipid and polymer assemblies (Supplementary Figures 1 and 3). The only significant factor shared among all models for both membrane types is the LPR or PPR (Supplementary Tables 8–13). This is rather surprising as previous studies suggested that the detergent concentration would be the most influential factor during the protein reconstitution7,8. Indeed, the size of the LPR/PPR coefficient is comparable to or higher than OG’s (Supplementary Tables 8 and 11).
Contour plots. a The contour plot visualizes the proteolipsomes size, modeled from the first round of experiments depending on the LPR and OG concentration, whereas the pH value is fixed at 7. The color gradient indicates different sizes and shows a trend towards a region within the design space which yields homogeneous, large proteoliposomes (highlighted by the red box). b This region of interest applies to all three responses as shown in the stacked contour plots for both membrane types. Red boxes encompass the subregions of the design space which was subsequently screened for proteovesicles with functioning PR-GFP. Contour plots visualizing the responses’ behavior for the other factor combinations can be found in the Supplementary Figures 12–15
PR-GFP concentration (Supplementary Tables 1–3) plays an important role in all three responses, FCS, DLS, and PdI, for both membrane types and appears in linear and interaction terms, as described above and in Supplementary Figure 11. Additionally, the detergent concentration appears to have the highest influence on the proteoliposome formation; whereas, the pH value alone does not. It is only significant for the description of the FCS data. The pH value naturally has an effect on proteins, which is reflected by highly significant interaction term pH*LPR (Supplementary Table 9). In case of the proteopolymersomes, the pH value has considerable more influence on the homogeneity of the vesicle population. This trend is verified further in the second DSD (Supplementary Table 12, Supplementary Figures 2 and 4) and supports the assumption that polymer membranes are different from lipid membranes when used for protein reconstitution (see also Table 2). It should be assumed that the pH affects the polymer self-assembly or the pH value affects PR-GFP, which then in turn interacts with the polymer assembly. It was stated in literature that detergents do not interact in the same way with polymersomes as with liposomes9. The detergent micelles coexist with the vesicles up to a threshold limit and a further increase leads to dissolution of the polymer membranes9. This observation is supported by the data and results from this study and the role of the pH should move into the focus of research. The effect of the detergent concentration on the reconstitution of PR-GFP is interesting. As described in literature, the formation of proteoliposomes and successful reconstitution of membrane proteins into them is highly dependent on the type and amount of used detergent8. This is again reflected by the obtained model which includes LPR*OG interaction terms for the FCS’ description. This mirrors the requirement of the correct amount of detergent necessary to destabilize the liposome membrane and allows transfer of the protein during detergent removal into it. However, this term is completely absent in case of the proteopolymersomes’ models. Generally, OG terms are only significant in two (FCS and DLS) of three cases (Supplementary Table 12).
Proton pumping
The formation of a proton gradient should depend on the number of pumps present, their orientation and their structural integrity44. Thus, the highest proton-pumping activity is expected to be found within the derived region yielding homogeneous, large proteovesicles. The measurement was based on the encapsulation of the pH-sensitive molecular probe pyranine. Its change of fluorescence intensity was used to calculate the internal change of pH (Supplementary Figures 7 and 24)23. The starting pH value of the samples was always 7.2, as described in Methods section.
The highest decrease of 0.11 pH units was observed in proteoliposomes formed at pH 7 using 0.75% of OG and a LPR of 25/0.16 mg/mL PR-GFP (Fig. 4 and Supplementary Figure 10). However, also the variance was highest using these conditions. Proteoliposomes formed at pH 6 resulted in more reproducible gradients of 0.08–0.09 pH units. Overall, the highest gradients formed were observed using a low LPR of 25/0.16 mg/mL PR-GFP and non-basic pH. The gradients formed within the proteopolymersomes were lower on average (Fig. 4 and Supplementary Figure 10). A decrease of 0.04 pH units at pH 6.5, PPR 97.5/0.04 mg/mL PR-GFP and 0.5% OG was the lowest observed; whereas, the combination of pH 8, PPR 125/0.03 mg/mL PR-GFP and 0.5% OG resulted in a decrease of 0.09 pH units.
The change of the internal pH value of the proteovesicles over time with and without illumination. PR-GFP’s proton pumping capability is triggered during the illumination at 530 ± 10 nm and consequently the internal pH value changes. Shown are the samples with the highest and the lowest observed pH gradient for both membrane types (mean, n = 3)
In DOPC liposomes, the amplitude of the pH gradient is influenced by all three parameters during reconstitution (Supplementary Table 10). The lower the LPR (i.e., the more pumps are present), or the lower the detergent concentration, the higher the activity (Fig. 5a). Furthermore, a near-neutral pH value in the range of 6.5–6.8 during reconstitution is beneficial for PR-GFP’s activity.(Fig. 5a). Looking at the contour of LPR and OG one can see a clear gradient towards the low factor settings (LPR 25–40, 0.75–1.2% OG, see Supplementary Figures 15-17). A likely explanation is that in a low detergent regime, the vesicular structure remains largely intact, allowing a primarily oriented insertion of numerous proteins into the membrane7.
The modeled response surfaces of the pH gradient depending on the two most influential factors are shown. DOPC proteoliposomes are shown in panel a and ABA proteopolymersomes in b. The third factor is fixed at its central value (a: LPR = 50, b: OG = 1.12 %). The preference of acidic pH values and low detergent concentrations for DOPC proteoliposomes becomes clearly visible (a). Contrary, the formed pH gradient becomes larger at basic pH values in case of the ABA proteopolymersomes, as well as higher PPR values (b)
In terms of pH, the opposite behavior is observed in polymersomes (Fig. 5b). A pH of 8 is suited best for alarge gradient amplitude. Similar to the DOPC liposomes, the OG concentration is suitable in the lower range. However, the PPR moves toward a lower number of proteins available for insertion (Fig. 5b). Comparing the number of PR-GFP molecules detected by FCS after reconstitution with the starting LPR/PPR (Supplementary Figure 23) it is clearly visible that for proteoliposomes a decreasing LPR leads to more PR-GFP per vesicle. However, this seems not to be true for polymersomes, where the number of PR-GFP per vesicle remains largely constant and even an opposite trend can be observed. The similar amplitude of the proteopolymersomes’ pH gradient (Fig. 4) with fewer pumps can be explained by a reduced back diffusion3,44,45. The permeability of protons through the PMOXA-PDMS-PMOXA membranes is lower and thus fewer pumps are necessary to achieve the same gradient38,46. Contrary to earlier reports47, the thicker polymersome membrane does not seem to inhibit PR-GFP’s functionality and lower gradient amplitudes are more likely to be attributed to lower amounts of PR-GFP present in the membrane. This underlines again the different mechanisms for lipid and polymer membranes resulting in a functional reconstitution, mainly the role of the detergent concentration and the pH (see Table 2).
Optimization
For the optimization, the gradient formation, the proteovesicle size and PdI were used as a target to reach large and uniform vesicles. The model equations 1–8 in the Supplementary Note 1 were used for the optimization process, which was carried by the use of desirability functions43. The optimization process is explained exemplarily for the proton gradient formation in the Supplementary Discussion and Supplementary Figure 19.
The optimal conditions and their anticipated responses are summarized in Table 3.
The optimization follows the observed trends discussed before: a slight acidic pH value of 6 in combination with a low LPR and medium-to-low amount of OG leads to the formation of highly homogeneous proteoliposomes which build-up a proton gradient upon illumination. These results are very similar to the conditions determined experimentally earlier28,29. ABA triblock polymers follow a different route. The pH should be in the basic regime around 8 and the PPR at 112/0.04 mg/mL PR-GFP. A detergent concentration of 0.82% (w/v) is found to be optimal, which is higher compared to the DOPC system and surprising taking the observed negative effects of OG on the vesicle formation into account (Supplementary Figure 2).
Verification
As a last step, the built up framework was put to a test. We used the derived optimal conditions to carry out the reconstitutions into DOPC and ABA membranes (Table 3). Additionally, control reconstitutions were carried out under the same conditions but without PRGFP in order to confirm the measured response.
In both cases, proteoliposomes and proteopolymersomes, the measured pH gradient was much higher than expected from the predictions (Fig. 6). In case of the proteoliposomes, 0.10 pH units were expected whereas the measurement resulted in 0.18 pH units (131 PR-GFP molecules (Q1: 110/Q3: 157)). Similarly, PRGFP’s performance was higher in ABA membranes than expected, 0.12 pH units compared to the predicted 0.08 pH units (8 PR-GFP molecules (Q1: 7/Q3: 10)). It should be noted that the measured results are within the prediction interval, ranging from 0.04 to 0.19 pH units in case of the proteoliposomes and 0.03 till 0.17 pH units for the proteopolymersomes, even though the offset to the prediction is large. Vesicles not carrying PR-GFP have also shown a reaction toward illumination which was, however, either small (0.06 pH units, DOPC) or showing the opposite behavior (0.02 pH units, ABA). It should further be noted that looking at the kinetics in Supplementary Figure 20 the dynamic of the control is different compared to the actual samples. Similar behavior observed in literature is likely attributed to the fluctuations in the fluorescence signal due to residual pyranine on the exterior vesicle membrane48,49.
Comparison of the actual measurements, their controls and predictions for all four responses. Error bars represent the standard error of the mean, n = 3. In case of the predicted values, the bars represent the prediction interval containing 95% of the outcomes. The underlying data are shown via dots which are jittered slightly to avoid overlapping. Liposomes and polymersomes prepared under the same conditions as the samples but without PR-GFP were used as controls
In terms of size, the difference between prediction, control, and actual measurement are present; whereas, the proteoliposomes are smaller than expected, but within the prediction interval. However, the controls fall out of that range. Similar behavior is observed in case of the ABA proteopolymersomes, however their control FCS results are in good agreement with the predictions. For both membranes, the controls’ PdI is much higher than expected and higher compared to PRGFP-containing vesicles.
Discussion
Although membrane protein reconstitutions have been carried out for decades, examples for their application for the design of synthetic devices are rare and usually of model-like simplicity3. The inherent complexity of this approach provides a demanding challenge. With our study, we provide a possible framework to this field, showing an example of a thoroughly designed approach. Design of experiments has proven to be an excellent scaffold, which can be used as a guide to optimize relevant factors impacting the reconstitution conditions, which are crucial to the formation of a functional system.
The easy accessibility of DoE allows detailed analysis and verified assumptions from literature and revealed new insights. Even though the models and results obtained via DoE are only valid for their specific case, the high coherence of this work supports the claim that the underlying method can be applied to further systems with different membrane protein/detergent/membrane combinations. More specifically, the created framework can be expanded with other membrane proteins, having different structures compared to the purely α-helical proteorhodopsin. As more complex membrane proteins usually require the use of milder detergents such as n-decyl-β-d-Maltopyranoside (DM) or n-dodecyl-β-d-Maltopyranoside (DDM), other factors could be easily integrated into the experimental design and data assessment process.
Our results support the assumption that PMOXA-PDMS-PMOXA block copolymer membranes require very different conditions for the reconstitution of PR-GFP and potentially other alpha-helical membrane proteins. Even low OG concentrations can be disadvantageous for the vesicle integrity and the pH value during formation and reconstitution has a larger influence than expected. Only the lipid/polymer-to-protein ratio is a shared factor among the two membrane types but less PR-GFP seems to be incorporated into polymersomes. However, our data indicate a similar pH gradient amplitude in the proteopolymersomes. A future study could investigate this phenomenon. It will be also interesting to apply our methodology to lipid/polymer hybrid systems and their use as platform for membrane proteins as the combined properties in terms of stability and biocompatibility would be beneficial50,51,52,53. Most importantly, the potential optimization has been shown to yield functional proteovesicles composed of the lipid DOPC or an ABA block copolymer. The predictions were tested out for both systems and verified that DoE is an excellent approach to fulfill the above stated requirements of reproducibility and predictability. Thus, our framework allows access to two highly important characteristics of engineering: reproducibility and predictability. Our example shows the application of molecular engineering from protein design up to a mathematical model in order to achieve a functional product with the desired properties. Applying this methodology to further polymer (different block compositions, diblock copolymers) and lipid systems, as well as to other membrane proteins and detergent types, will potentially result in the realization of a library containing the necessary information about the interplay of these buildings blocks and the identification of general mechanisms and important factors governing the assembly of proto-cells.
Methods
Materials
All chemicals were provided by Sigma-Aldrich (Switzerland) if not stated otherwise. Experiments were carried out in triplicates and their standard deviation used to estimate the error if not stated otherwise. Lipids were supplied by Avanti Polar Lipids (USA) and were supplied as a powder, solubilized in chloroform and stored at −20 °C. The used detergents were supplied by Affymetrix (USA).
Cultivation of Escherichia coli
The cultivation of E. coli and expression and purification of PR-GFP was done essentially as described before28,29. E. coli carrying the PR-GFP containing pLEMO plasmid29 (kindly provided by Prof. Daniel Müller, ETH Zürich) was grown in LB-Miller medium at 30 °C and 180 rpm. An overnight culture was grown in the presence of 100 µg/mL ampicillin. The sterile medium was inoculated with 1% (v/v) of the overnight culture and 100 µg/mL ampicillin and 34 µg/mL chloramphenicol was added. The optical density (λ = 600 nm) was measured during growth and the expression triggered at a density of 0.8–1 via the addition of 0.1 mM β-d-1-thiogalactopyranoside (IPTG) and 5 μM all-trans-retinal. Subsequently, cells were incubated for additional 3 h, collected by centrifugation (3–18 K, Sigma) at 4000×g for 20 min and the supernatant removed. As a last step, the pellets were collected and suspended in 20 mM Tris-HCl, 100 mM NaCl, pH 7.4 and stored at −20 °C until preparation of the membrane.
Membrane preparation
The frozen cells were thawed and subsequently lysed by using a French press (EmulsiFlex, Avestin), operated at 1500 bar. The resulting lysate was first centrifuged (3–18 K, Sigma) at 4000×g for 20 min in order to remove cell debris and then at 150,000×g for 1 h (Optima XE-90, Beckman Coulter) to isolate the membrane. The pellet was homogenized, washed with 20 mM Tris-HCl, 100 mM NaCl, 10% (w/v) glycerol, pH 7.4 two times, aliquoted to 1 mL and stored at −80 °C until further use.
Purification
The isolation of PR out of its native membrane was carried out by using His-tag chromatography. The crude membrane fraction was solubilized in 7 mL buffer (20 mM Tris-HCl, pH 7.4, 300 mM NaCl, 10% (w/v) Glycerol) and 2.5% Cymal-5. The solution was protected from light and placed on an orbital shaker at room temperature overnight. On the next day, 1 mL Ni-NTA resin (Quiagen) was washed three times with the solubilization buffer and added to another 7 mL buffer with 30 mM imidazole resulting in a total volume of 15 mL. Subsequently, the suspension was placed in a an orbital shaker for 3 h to ensure complete binding to the resin. The suspension was transferred into spin-columns (Promega) and the flow-through was collected. 20 mM Tris-HCl, pH 7.4, 300 mM NaCl, 30 mM imidazole, 10% (m/v) glycerol containing 0.4% Cymal-5 were used to wash the resin and the flow-through was collected again. In order to remove residual washing buffer, the spin-column was centrifuged with 200×g for 1 min. The end of the column containing the resin was cutoff and the column sealed with parafilm. Volume of 450 μL of 20 mM KPi buffer, pH 7.4, 150 mM NaCl, 10% (w/v) glycerol, 400 mM imidazole and 0.4% Cymal-5 were added to elute the protein. It was incubated for 1 h on an orbital shaker at room temperature protected from light. The tube was placed in a 2 mL reaction tube and centrifuged in a bench-top centrifuge for 1 min at maximum speed. A spin-filter (30 kDa cutoff) was used with 20 mM KPi buffer, pH 7.4, 150 mM NaCl, 10% (w/v) glycerol and 0.4% Cymal-5 to increase the protein concentration and remove the residual imidazole. Volume of 1 μL of 1 mM TCEP was added after purification and the concentration determined by measuring the absorbance at 280 nm (ε = 96,510 L mol-1 cm-1). Typical concentrations were around 10–20 mg/mL. The purified PR-GFP was stored at 4 °C and used within 3 days.
Vesicle preparation and reconstitution of PR-GFP
The vesicles were formed using a variation of the film rehydration method28,54. A certain volume of the DOPC (in chloroform) or PMOXA17-PDMS65-PMOXA17 (in ethanol) solution was transferred into 1.25 mL flat bottom flasks and dried under high vacuum for 90 min. The resulting film was rehydrated with an aqueous buffer which always contained 20 mM potassium phosphate buffer, 150 mM potassium chloride and OG. The pH values and OG concentrations were adjusted accordingly to the experimental plan (Supplementary Tables 1-3). The final volume was 500 µL and the membrane concentration 4 mg/mL. If the vesicles were used for proton translocation measurements, 100 µM of pyranine was added. The solutions were stirred overnight at room temperature and protected from light. Subsequently, the vesicle preparation were homogenized by extrusion (11× times) through a polycarbonate membrane (200 nm, Nucleopore, Whatman). Now the necessary volume of PR-GFP was added according to the desired LPR/PPR (Supplementary Tables 1-3) and the protein-vesicle suspension stirred for 30–60 min. The dilution during the addition of PR-GFP to the reconstitution buffer lowers the concentration of CYMAL-5 by factor of ~20 below the cmc (0.12%) and thus, the impact of residual Cymal-5 was considered negligible55. Afterwards the samples were transferred into dialysis tubes (15 kDa cutoff, Visking) and dialyzed against 20 mM KPi and 150 mM KCl for 48 h. The pH was the same as the sample pH value. Furthermore, ~100 mg of SM-2 biobeads (Bio-Rad) were added to ensure a constant dialysis gradient56. After the dialysis was complete, the samples were again extruded with a 200 nm membrane to ensure a homogeneous solution, remove any formed aggregates and purified by eluting them through a G-25 MiniTrap size exclusion chromatography column (GE Healthcare), equilibrated with 20 mM potassium phosphate buffer, 150 mM KCl, pH 7.2. The final volume was 1 mL and the membrane concentration 2 mg/mL.
Dynamic light scattering
The samples were measured in a Zetaziser Nano (Malvern) at 25 °C. A HeNe laser (λ = 633 nm) was uses as a light source. The samples were not diluted and allowed to equilibrate for 120 s.
Fluorescence correlation spectroscopy
The FCS measurements were performed as already described40. Briefly, an inverted microscope (Axiovert 200 M, Zeiss), equipped with a laser scanning microscopy module LSM 510 (Zeiss) and a ConfoCor2 (Zeiss) module was used. A 488 nm HeNe laser was focused into the 5 μL sample using a 488 nm dichroic mirror and a 40× water immersion objective. The emission beam was guided through a 70 μm pinhole and detected. The autocorrelation curve was fitted by using the equation
with τ D being the diffusion time, equivalent to the decay time of the autocorrelation curve. The equation
with D being the diffusion coefficient was used to calibrate ω, the radius of the confocal volume, by using the known fluorescent dye orgeon green 48857. Finally, the Stokes-Einstein equation
with T being the temperature (298 K), k B the Boltzmann constant and η the viscosity was used to calculate the hydrodynamic radius r. Measurements were performed in 10 s intervals and 30 repetitions. The number of PR-GFP molecules was determined by estimating the molecular brightness of a single PR-GFP molecule in 20 mM potassium phosphate buffer, 150 mM KCl, pH 7.2, and 0.4% Cymal-5 which resulted in 2.4 ± 0.19 counts per molecule (cpm). Dividing the vesicles’ molecular brightness by PRGFP’s one results in an estimate of the number of PR-GFP molecules per vesicle.
Proton pumping assay
To detect PR-GFP’s ability to transport protons across a membrane when put under illumination we followed the well-established pyranine assay23,58. The measurements were carried out in a fluorescence spectrometer (LS55, Perkin Elmer), illuminating the sample with an 100 W xenon lamp (Intralux 4100, Volpi), utilizing a fiber guide to place the beam directly over the sample. The wavelength was adjusted to 530 ± 10 nm by using a band-pass filter (Thorlabs). The samples were measured undiluted and in the dark for 30 min in order to equilibrate them. Afterwards the measurement was carried out under illumination, whereby the illumination was cycled between 50 s on and 10 s off. The fluorescence measurement was done during the off cycle to avoid interference. After the illumination measurement, the sample was measured for another 30 min in the dark to observe the re-equilibration of the fluorescence signal. The temperature was controlled at 20 ± 1 °C. The measurement data from the first 30 min in the dark was used for a linear fit whose slope was used as a correction factor for the measurement in order to remove potential artifacts from pH drift. The fluorescence intensity data were normalized by using \({\Delta F}_{{\mathrm{460}}}/F_{{\mathrm{460}}}\). In order to calculate the gradients’ amplitude in pH units, we used a calibration curve (Supplementary Figure 24).
Experimental design and data assessment
We followed the design proposed by Jones and Nachtsheim called definitive screening design which allows a one-step screening and optimization process34,35. All experimental designs were created using the DoE module of the software JMP (SAS). The factors were chosen to be the pH value, the lipid-to-protein or polymer-to-protein ratio (LPR or PPR) and the OG concentration in % (w/v). Their high, middle and low settings can be found in Supplementary Tables 1 - 3. All designs were repeated three times in total to assess the variance. The data assessment were done by using the software R (Version 3.4). The model equations were derived by using a stepwise forward regression variant which enforces effect heredity, thus higher order effects are only included together with their corresponding linear effect59. All possible models were fitted and the one with the highest adjusted R2 was chosen. Blocking variables were used to take the possible batch-to-batch variation into account. The equations in Supplementary Note 1 were used for the optimization.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Bujara, M. & Panke, S. Engineering in complex systems. Curr. Opin. Biotechnol. 21, 586–591 (2010).
Hammer, D. A. & Kamat, N. P. Towards an artificial cell. FEBS Lett. 586, 2882–2890 (2012).
Palivan, C. G. et al. Bioinspired polymer vesicles and membranes for biological and medical applications. Chem. Soc. Rev. 45, 377–411 (2016).
Langowska, K., Palivan, C. G. & Meier, W. Polymer nanoreactors shown to produce and release antibiotics locally. Chem. Commun. 49, 128–130 (2013).
Lomora, M., Gunkel-Grabole, G., Mantri, S. & Palivan, C. G. Bio-catalytic nanocompartments for in situ production of glucose-6-phosphate. Chem. Commun. 53, 10148–10151 (2017).
Marguet, M., Bonduelle, C. & Lecommandoux, S. Multicompartmentalized polymeric systems: towards biomimetic cellular structure and function. Chem. Soc. Rev. 42, 512–529 (2013).
Rigaud, J.-L. & Lévy, D. Reconstitution of membrane proteins into liposomes. Methods Enzymol. 372, 65–86 (2003).
Seddon, A. M., Curnow, P. & Booth, P. J. Membrane proteins, lipids and detergents: not just a soap opera. Biochim. Biophys. Acta 1666, 105–117 (2004).
Pata, V., Ahmed, F., Discher, D. E. & Dan, N. Membrane solubilization by detergent: resistance conferred by thickness. Langmuir 20, 3888–3893 (2004).
Grzelakowski, M., Cherenet, M. F., Shen, Y. & Kumar, M. A framework for accurate evaluation of the promise of aquaporin based biomimetic membranes. J. Membr. Sci. 479, 223–231 (2015).
Weissman, S. A. & Anderson, N. G. Design of Experiments (DoE) and process optimization. a review of recent publications. Org. Process Res. Dev. 19, 1605–1633 (2015).
Nardin, C., Hirt, T., Leukel, J. & Meier, W. Polymerized ABA triblock copolymer vesicles. Langmuir 16, 1035–1041 (2000).
Nardin, C., Widmer, J., Winterhalter, M. & Meier, W. Amphiphilic block copolymer nanocontainers as bioreactors. Eur. Phys. J. E Soft Matter Biol. Phys. 4, 403–410 (2001).
Discher, D. E. Polymer vesicles. Science 297, 967–973 (2002).
Le Meins, J.-F. & Sandre, O. & Lecommandoux, S. Recent trends in the tuning of polymersomes’ membrane properties. Eur. Phys. J. E Soft Matter 34, 14 (2011).
Renggli, K. et al. Selective and responsive nanoreactors. Adv. Funct. Mater. 21, 1241–1259 (2011).
Hirschi, S. et al. Engineering and assembly of protein modules into functional molecular systems. Chim. Int. J. Chem. 70, 398–401 (2016).
Althoff, T., Davies, K. M., Schulze, S., Joos, F. & Kühlbrandt, W. GRecon: a method for the lipid reconstitution of membrane proteins. Angew. Chem. 124, 8468–8472 (2012).
Jackman, J., Knoll, W. & Cho, N.-J. Biotechnology applications of tethered lipid bilayer membranes. Materials 5, 2637–2657 (2012).
Krupinski, J. & Hammes, G. G. Steady-state ATP synthesis by bacteriorhodopsin and chloroplast coupling factor co-reconstituted into asolectin vesicles. Proc. Natl Acad. Sci. USA 83, 4233–4237 (1986).
Reimhult, E. & Kumar, K. Membrane biosensor platforms using nano- and microporous supports. Trends Biotechnol. 26, 82–89 (2008).
Tunuguntla, R. H. et al. Bioelectronic light-gated transistors with biologically tunable performance. Adv. Mater. 27, 831–836 (2015).
Verchère, A., Broutin, I. & Picard, M. Photo-induced proton gradients for the in vitro investigation of bacterial efflux pumps. Sci. Rep. 2, 306 (2012).
Bamann, C. Bamberg, E. Wachtveitl, J. & Glaubitz, C. Proteorhodopsin. Biochim. Biophys. Acta 1837, 614–625 (2014).
Rigaud, J.-L., Pitard, B. & Levy, D. Reconstitution of membrane proteins into liposomes: application to energy-transducing membrane proteins. Biochim. Biophys. Acta 1231, 223–246 (1995).
Grote, M. & O’Malley, M. A. Enlightening the life sciences: the history of halobacterial and microbial rhodopsin research. FEMS Microbiol. Rev. 35, 1082–1099 (2011).
Beja, O., Spudich, E. N., Spudich, J. L., Leclerc, M. & DeLong, E. F. Proteorhodopsin phototrophy in the ocean. Nature 411, 786–789 (2001).
Harder, D. et al. Engineering a chemical switch into the light-driven proton pump proteorhodopsin by cysteine mutagenesis and thiol modification. Angew. Chem. 128, 8992–8995 (2016).
Ritzmann, N. et al. Fusion domains guide the oriented insertion of light-driven proton pumps into liposomes. Biophys. J. 113, 1181–1186 (2017).
Wang, W.-W., Sineshchekov, O. A., Spudich, E. N. & Spudich, J. L. Spectroscopic and photochemical characterization of a deep ocean proteorhodopsin. J. Biol. Chem. 278, 33985–33991 (2003).
Lörcher, S. & Meier, W. Cosolvent fractionation of PMOXA- b -PDMS- b -PMOXA: Bulk separation of triblocks from multiblocks. Eur. Polym. J. 88, 575–585 (2017).
Jones, B. & Nachtsheim, C. J. A class of three-level designs for definitive screening in the presence of second-order effects. J. Qual. Technol. 43, 1–15 (2011).
Jones, B. & Nachtsheim, C. J. Definitive screening designs with added two-level categorical factors. J. Qual. Technol. 45, 121–129 (2013).
Peiró-Salvador, T., Ces, O., Templer, R. H. & Seddon, A. M. Buffers may adversely affect model lipid membranes: a cautionary tale. Biochemistry 48, 11149–11151 (2009).
Brasnett, C., Longstaff, G., Compton, L. & Seddon, A. Effects of cations on the behaviour of lipid cubic phases. Sci. Rep. 7, 8229 (2017).
Isaksson, S. et al. Protein-containing lipid bilayers intercalated with size-matched mesoporous silica thin films. Nano Lett. 17, 476–485 (2017).
Amstad, E. et al. Triggered release from liposomes through magnetic actuation of iron oxide nanoparticle containing membranes. Nano Lett. 11, 1664–1670 (2011).
Lomora, M. et al. Polymersomes with engineered ion selective permeability as stimuli-responsive nanocompartments with preserved architecture. Biomaterials 53, 406–414 (2015).
Ibrahimova, V. et al. Photosensitizer localization in amphiphilic block copolymers controls photodynamic therapy efficacy. Nanoscale 9, 11180–11186 (2017).
Einfalt, T. et al. Stimuli-triggered activity of nanoreactors by biomimetic engineering polymer membranes. Nano Lett. 15, 7596–7603 (2015).
Itel, F. et al. Molecular organization and dynamics in polymersome membranes: a Lateral Diffusion Study. Macromolecules 47, 7588–7596 (2014).
Choi, H.-J., Lee, H. & Montemagno, C. D. Toward hybrid proteo-polymeric vesicles generating a photoinduced proton gradient for biofuel cells. Nanotechnology 16, 1589–1597 (2005).
Derringer, G. Simultaneous optimization of several response variables. J. Qual. Technol. 12, 214–219 (1980).
Seigneuret, M. & Rigaud, J. L. Analysis of passive and light-driven ion movements in large bacteriorhodopsin liposomes reconstituted by reverse-phase evaporation. 2. Influence of passive permeability and back-pressure effects upon light-induced proton uptake. Biochemistry 25, 6723–6730 (1986).
Seigneuret, M. & Rigaud, J. L. Analysis of passive and light-driven ion movements in large bacteriorhodopsin liposomes reconstituted by reverse-phase evaporation. 1. Factors governing the passive proton permeability of the membrane. Biochemistry 25, 6716–6722 (1986).
Craciun, I., Denes, A. S., Gunkel-Grabole, G., Belluati, A. & Palivan, C. G. Surfaces decorated with polymeric nanocompartments for ph reporting. Helv. Chim. Acta 101, e1700290 (2018).
Kuang, L. et al. “Frozen” block copolymer nanomembranes with light-driven proton pumping performance. ACS Nano 8, 537–545 (2014).
Choi, H.-J. & Montemagno, C. D. Artificial organelle: ATP synthesis from cellular mimetic polymersomes. Nano Lett. 5, 2538–2542 (2005).
Choi, H.-J., Germain, J. & Montemagno, C. D. Effects of different reconstitution procedures on membrane protein activities in proteopolymersomes. Nanotechnology 17, 1825–1830 (2006).
Beales, P. A., Khan, S., Muench, S. P. & Jeuken, L. J. C. Durable vesicles for reconstitution of membrane proteins in biotechnology. Biochem. Soc. Trans. 45, 15–26 (2017).
Seneviratne, R. et al. A reconstitution method for integral membrane proteins in hybrid lipid-polymer vesicles for enhanced functional durability. Methods (2018). In the press https://doi.org/10.1016/j.ymeth.2018.01.021
Le Meins, J.-F., Schatz, C., Lecommandoux, S. & Sandre, O. Hybrid polymer/lipid vesicles: state of the art and future perspectives. Mater. Today 16, 397–402 (2013).
Khan, S., Li, M., Muench, S. P., Jeuken, L. J. C. & Beales, P. A. Durable proteo-hybrid vesicles for the extended functional lifetime of membrane proteins in bionanotechnology. Chem. Commun. 52, 11020–11023 (2016).
Kumar, M., Grzelakowski, M., Zilles, J., Clark, M. & Meier, W. Highly permeable polymeric membranes based on the incorporation of the functional water channel protein Aquaporin Z. Proc. Natl Acad. Sci. USA 104, 20719–20724 (2007).
Tunuguntla, R. et al. Lipid bilayer composition can influence the orientation of proteorhodopsin in artificial membranes. Biophys. J. 105, 1388–1396 (2013).
Philippot, J., Mutaftschiev, S. & Liautard, J. P. A very mild method allowing the encapsulation of very high amounts of macromolecules into very large (1000 nm) unilamellar liposomes. Biochim. Biophys. Acta 734, 137–143 (1983).
Müller, C. B. et al. Precise measurement of diffusion by multi-color dual-focus fluorescence correlation spectroscopy. Europhys. Lett. 83, 46001 (2008).
Seigneuret, M. & Rigaud, J.-L. Use of the fluorescent pH probe pyranine to detect heterogeneous directions of proton movement in bacteriorhodopsin reconstituted large liposomes. FEBS Lett. 188, 101–106 (1985).
Lawson, J. Design and analysis of experiments with R. (CRC Press, Taylor & Francis Group, 2015).
Acknowledgements
The authors would like to thank Martina and Hans-Peter Garni for their help with developing and testing the fluorescence measurement setup. Gabriele Persy is thanked for measuring the TEM images and Sven Kasper for the polymer synthesis. The research leading to these results has received funding from the University of Basel, the Swiss Nanoscience Institute, the Swiss National Science Foundation and the NCCR Molecular Systems Engineering. The authors are grateful for this support.
Author information
Authors and Affiliations
Contributions
R.G. designed the studies, developed the analysis and mathematical modeling. R.G., J.T., N.R., C.A., and A.S. designed, performed, and analyzed the experiments. G.G.G., D.F., D.M., and W.M. provided scientific guidance. All authors contributed to writing the manuscript and approved its final form.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Goers, R., Thoma, J., Ritzmann, N. et al. Optimized reconstitution of membrane proteins into synthetic membranes. Commun Chem 1, 35 (2018). https://doi.org/10.1038/s42004-018-0037-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42004-018-0037-8
This article is cited by
-
Proton gradients from light-harvesting E. coli control DNA assemblies for synthetic cells
Nature Communications (2021)
-
En route to dynamic life processes by SNARE-mediated fusion of polymer and hybrid membranes
Nature Communications (2021)
-
The chemistry of cross-linked polymeric vesicles and their functionalization towards biocatalytic nanoreactors
Colloid and Polymer Science (2021)
-
Artificial intelligence application for rapid fabrication of size-tunable PLGA microparticles in microfluidics
Scientific Reports (2020)
-
Reconstitution of FoF1-ATPase-based biomimetic systems
Nature Reviews Chemistry (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.