Abstract
Until recently the total synthesis of insulin, with its characteristic heterodimeric structure crosslinked by two interchain and one intrachain disulfide (SS) bridge, remained largely an unsolved challenge. By optimizing the synthesis and directed disulfide crosslinking of the two chains, and by applying biomimetic monocomponent proinsulin approaches, efficient insulin syntheses have been realized. Here we report the optimization and characterisation of an alternative strategy, oxidative native chain assembly. In this method unprotected A- and B-chains assemble oxidatively under thermodynamic control to afford bovine pancreatic insulin in 39% yield. Folding is found to proceed predominantly via structured 1SS* and 2SS* intermediates with a common interchain CysA20‒CysB19 disulfide. These results suggest that native chain assembly, long considered inefficient, may represent a reasonable strategy to access insulin variants. This is supported by the synthesis of human insulin and human type-II relaxin in yields of up to 49 and 47%, respectively, although the application to human insulin ValA16 variant is unsuccessful.
Similar content being viewed by others
Introduction
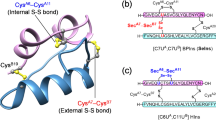
The endocrine hormone insulin regulates energy storage and glucose metabolism, and, thus, in diabetic patients where this system malfunctions, glucose levels have to be controlled in a life saving manner through subcutaneous injection of insulin. This globular miniprotein hormone (5.8 kDa) consists of two peptide chains, the A-chain (Ins-A, 21 amino acid residues) with its intramolecular disulfide bridge (CysA6–CysA11) and the B-chain (Ins-B, 30 amino acid residues), which are crosslinked by two interchain disulfide (SS) bridges (i.e., CysA7–CysB7 and CysA20–CysB19)1. In nature, insulin is produced as proinsulin, a single polypeptide chain of 86 amino acid residues in which the C-terminus of the B-chain is linked to the N-terminus of the A-chain by a 35-residue connecting C-peptide. In vivo, the proinsulin folds efficiently in the endoplasmic reticulum of the pancreatic β-cells, and is then converted to the mature two-chain insulin molecule by proteolytic processing of the prohormone2,3. Based on this knowledge a biomimetic approach was developed for rDNA industrial production of human insulins for therapeutic use which relies on the folding and enzymatic processing of recombinant proinsulin4,5. In vitro, the overall yield of folding and enzymatic processing of proinsulin into insulin can reach up to 70%6.
Early attempts in the 1960s of oxidative assembly of natural7 and of synthetic A- and B-chains8,9,10 into insulin led to disappointingly low yields of about 1–5%, in agreement with statistical analyses which predicted even significantly lower yields of folded insulin (i.e., <0.05%)11. However, yields of correctly folded insulin from dithiothreitol (DTT)-induced scrambled insulin could be sensibly improved under optimized conditions such as by the use of S-sulfonated A- and B-chains12 and in presence of protein disulfide isomerase (PDI)13,14 or its fragment with a redox active site15 to 30–60% and 25–30%, respectively. Moreover, studies on crosslinked insulin A- and B-chains clearly revealed that even short non-peptidic bridges designed from the 3D structure of insulin16 are sufficient to allow the correct oxidative folding of such single-component insulin precursors into the native disulfide topology at yields similar to those of proinsulin17,18,19. All these results strongly suggested that the A- and B-chains of insulin contain sufficient structural information for correct oxidative folding if the entropic penalty of associating the two chains into a globular structure is somehow reduced or even prevented.
The general improvements in the SPPS synthesis of larger peptides by the use of isoacyl amide bonds20,21,22,23,24,25,26 and optimization of the orthogonal protection of multiple cysteine residues27,28,29 led more recently to improved total syntheses of insulin by directed assembly of the two chains6,30,31,32,33. However, it was the approach via the single-component insulin precursors that allowed the greatest advances in efficient new total synthesis of insulin and other members of the insulin superfamily34,35,36,37,38.
Recently we demonstrated that the selenocysteine (Sec) strategy, i.e., introduction of two selenocysteine residues instead of cysteine residues to facilitate selective formation of a native disulfide topology,28,39,40 represents an alternative useful approach for an efficient and facile synthesis of active insulin analogs41. Indeed just by mixing the reduced Ins-A and Ins-B of bovine pancreatic insulin (BPIns), each containing one Sec residue in place of Cys7, at pH 10 and 4 °C, selenoinsulin, a long-lasting insulin analog having a SecA7–SecB7 diselenide (SeSe) bridge, was obtained in a yield of up to 27%. The selenoinsulin retained the same three-dimensional structure, hence the same biological activity as the wild-type in the cascade phosphorylation reactions in vitro41.
The success of selenoinsulin preparation suggested the possibility that active insulin may easily be obtained by chain assembly of reduced Ins-A and Ins-B without using an explicit interchain tether nor orthogonal protections on the Cys residues if selective formation of a native disulfide bridge between Ins-A and Ins-B can be induced. If such a classical protocol, namely native chain assembly (NCA), could be developed, the protocol would potentially be applicable to two-chain assembly of various members of the insulin superfamily. Indeed, it was previously reported that similar approaches are efficient for human type-II relaxin (HRlx-2)42,43. However, such a simple approach did not yield native fold for human insulin ValA16 variant44 and human relaxin-345.
In this study, we report NCA conditions for the synthesis of wild-type BPIns from A- and B-chains without applying modifications or protections on the peptide chains. To access this goal, the two-chain oxidative folding pathways of BPIns are first determined and then optimized. Based on the obtained NCA conditions, human insulin (HIns) and human type-II relaxin (HRlx-2) are both produced in a yield of nearly 50%. On the other hand, a possible limitation of NCA is evidenced by the unsuccessful application to the two-chain assembly of human insulin ValA16 variant that is known as a 'nonfoldable' insulin44 unless a single-component insulin precursor is applied35.
Results
Oxidative folding of BPIns with NCA
To find an effective NCA protocol, we tried to identify the major two-chain oxidative folding pathways of BPIns. When reduced Ins-A (RA) and Ins-B (RB) of BPIns, which had been prepared by reduction of native BPIns (N) with an excess amount of DTT46, were mixed in a pH 10.0 buffer solution containing ethylenediaminetetraacetic acid (EDTA) (1 mM) and urea (0.9 M) (Fig. 1a), RA was gradually converted into partially oxidized 1SSA and then fully oxidized 2SSA, which had one and two intramolecular SS linkages, respectively (eq. 1).
HPLC chromatograms obtained from the double-chain oxidative folding of BPIns. a Mixing of RA (227 μM) and RB (222 μM). b Mixing of 1SSA (225 μM) and RB (233 μM) c Mixing of 2SSA (201 μM) and RB (211 μM). d Mixing of RA (232 μM) and 1SSB (186 μM). The oxidative folding was performed at pH 10.0 and 0 °C in the presence of 0.9 M urea. Before HPLC analyses, aliquots of the folding solution were pre-treated with aqueous 2-aminoethyl methanethiosulfonate (AEMTS)62 to quench the reaction by blocking the free SH groups into -SSCH2CH2NH3+. Peaks A–E and F–H represent components of 1SSA and 2SSA, respectively. See Table 1 for the individual structure of these species and Supplementary Figs. 1 and 2 for their HPLC chromatograms
Similarly but much more slowly, RB was oxidized to 1SSB as well (eq. 2).
The intramolecular SS formation behaviors of Ins-A and Ins-B are consistent with our previous kinetic analysis of the SS formation46. Relative populations of RA, 1SSA, and 2SSA for Ins-A, those of RB and 1SSB for Ins-B, and the yield of N are plotted in Fig. 2a as a function of the reaction time. It can be seen that N is slowly generated after accumulation of 1SSA in the solution, suggesting that 1SSA couples with RB to form a productive heterodimeric transient species (TS), which would be oxidized to N with oxidants present in the solution, such as oxygen, 2SSA, and 1SSB (eq. 3).
Time courses of relative populations of disulfide-interediates and the yield of N. The reaction conditions for panels a, b, c, and d were the same as panels a, b, c, and d, respectively, of Fig. 1
Indeed, the populations of 1SSA and RB tended to decrease with generation of N. However, the total amount of N generated was much smaller than the decreased amounts of 1SSA and RB, indicating that alternative oxidation of these species to 2SSA and 1SSB took place slowly but significantly in the solution. As a result, the yield of N, which reached up to 9% after 100 h, did not significantly increase in a longer reaction time, during which 1SSA and RB were no longer available at high concentrations.
To support the chain assembly scheme of eq. 3, oxidized Ins-A (1SSA or 2SSA) and oxidized Ins-B (1SSB) were prepared in situ by treatment of RA and RB, respectively, with appropriate amounts of 3,4-dihydroxy-1-selenolane Se-oxide (DHSox), which is a strong oxidant for peptides to form disulfide bonds rapidly, selectively, and quantitatively47. The resulting species were subsequently mixed with the reduced counter-chain (i.e., RB or RA). When one equivalent of DHSox was reacted with RA, 1SSA species were generated mainly along with unreacted RA and fully oxidized 2SSA (Fig. 1b, 0 h). To the resulting mixture was added a RB solution. The populations of 1SSA and RB only slightly changed in course of time, while generation of N was observed after 1 h (Fig. 2b). The faster formation and higher yield of N than those in the chain assembly of RA and RB (Fig. 2a) are consistent with the coupling scheme of eq. 3.
On the other hand, when two equivalents of DHSox were reacted with RA, only 2SSA species were observed (Fig. 1c, 0 h). Subsequent mixing of generated 2SSA with RB resulted in rapid redistribution of the SS species within 0.5 h probably through an intermolecular SH/SS exchange reaction (Figs. 1c and 2c). In this case, generation of N was much faster than that seen in Figs. 1a and 1b, and the yield of N was increased up to 17%, suggesting the presence of an alternative pathway to N (eq. 4) in addition to eq. 3.
When 1SSB was mixed with RA, a similar pre-folding event, i.e., redistribution of RA and 1SSB into 1SSA and RB, respectively, was observed (Figs. 1d and 2d). Thus, 1SSA and RB were obtained as major components after 10 h, leading to accelerated formation of N. The final yield of N increased up to 15%. It should be noted that swap species, which are metastable misfolded 3SS intermediates with a non-native interchain SS linkage (e.g., CysA6–CysB7 or CysA11–CysB7) and are kinetically trapped on the folding pathways,48,49 were also generated as minor products as seen in Fig. 1.
Characterization of key SS* intermediates in NCA
The disulfides present in the 1SSA and 2SSA species could be unambiguously determined by comparing the retention times on HPLC chromatograms with those of authentic samples prepared by the solid-phase peptide synthesis (SPPS) (for their HPLC chromatograms see Supplementary Figs. 1 and 2). Relative populations of the 1SSA and 2SSA components estimated at different temperatures are summarized in Table 1. The results clearly show that the species of peak B, which has a native SS linkage, is the thermodynamically most stable among the six 1SSA species and also that its stability gradually increases by lowering the temperature (Table 1, right column). Similarly, the species of peak G, which has the native SS linkage, is the most populated among the three 2SSA species. It is, therefore, most likely that [CysA6‒CysA11]A (peak B) and [CysA6‒CysA11, CysA7‒CysA20]A (peak G) would preferentially react with RB to produce TS and TS′, respectively. This is consistent with the previous reports that suggested preferential formation of CysA6‒CysA11 linkage in Ins-A before interchain disulfide bond formation between Ins-A and Ins-B chains50,51.
To identify key SS species (SS*) after the chain assembly, we carried out reductive unfolding of native BPIns. When N was reacted with an excess amount of DTT (Fig. 3a), small but distinct peaks (2SS* and 1SS*) were transiently observed in addition to RA and RB. The transient species could be assigned by MS analysis (Supplementary Fig. 3) and chymotrypsin digestion analysis (Supplementary Fig. 4 and Supplementary Table 1) as metastable chain-coupled species having two native SS linkages of CysA6‒CysA11 and CysA20‒CysB19 (for 2SS*) and one native SS linkage of CysA20‒CysB19 (for 1SS*). 2SS* and 1SS* should be major ingredients of the key transient species (TS and/or TS′) proposed in the two-chain assembly of Ins-A and Ins-B. Interestingly, the species corresponding to 2SS* was observed in the NCA solution (Fig. 1). However, 1SS* was not observed during NCA probably due to overlapping with a large peak of RB.
Reductive unfolding of BPIns and CD spectrum of 2SS*. a HPLC chromatograms obtained from a reductive unfolding experiment. BPIns (0.95 mg mL−1) was reacted with 0.5 mM DTT in 25 mM sodium bicarbonate buffer (pH 10.0) at 0 °C. b Comparison of the CD spectrum of 2SS* (solid line) with that of BPIns (dashed line). The spectra were measured at the concentration of 23.5 μM at 4 °C and pH 7.0 using a quartz cell with a light path length of 1 mm
Major NCA folding pathways of BPIns
Taking the aforementioned results together, we derived an oxidative folding pathway of BPIns from the two chains as shown in Fig. 4. The 1SSA and 2SSA intermediates should contain all possible components with one or two disulfides, but the major components having a native CysA6‒CysA11 SS linkage. When minor components of 1SSA, such as peaks A and D of Table 1, reacts with RB through intermolecular SH/SS exchange, 1SS* having a CysA20‒CysB19 SS bridge would be formed and then oxidized to 2SS* through formation of the CysA6‒CysA11 SS linkage. In another pathway the CysA20‒CysB19 SS linkage (i.e., 1SS*) might be formed from RA and RB by direct oxidation as observed in the folding studies of proinsulin50. However such interchain disulfide formation should be much slower than the intrachain disulfide formation.
On the other hand, when the major component of 1SSA (peak B of Table 1) reacts with both RB and an oxidant, 2SS* would be generated directly through formation of a CysA20‒CysB19 SS bridge keeping the native CysA6‒CysA11 linkage. The resultant 2SS* would be converted to N through formation of the third native SS linkage, rather than be broken to 2SSA and RB through intramolecular SH/SS exchange, because 2SS* would have thermodynamic stability to some extent due to significant presence of α-helices as seen in Fig. 3b. Another effective pathway to N must exist from 2SSA. The SH/SS exchange reaction between the major component (peak G) and RB would produce TS′ having two native SS linkages. Thus, 2SS* must be an important ingredient of TS′.
According to the revealed NCA folding pathways of BPIns (Fig. 4), it is obvious that the CysA20‒CysB19 SS bridge is more easily formed between Ins-A and Ins-B than the other SS bridge, i.e., CysA7‒CysB7. This is consistent with the previous report52 and the fact that CysA7‒CysB7 is solvent-exposed53 and easily broken as seen in the reductive unfolding of BPIns (Fig. 3a). Previously, existence of two possible precursors with intrachain and interchain disulfide bonds, i.e., [CysA6‒CysA11, CysA7‒CysB7]AB and [CysA6‒CysA11, CysA20‒CysB19]AB, have been suggested in folding studies of both proinsulin and two-chain insulin50,51,54. The present work clearly showed that [CysA6‒CysA11, CysA20‒CysB19]AB is a major precursor of N during the oxidative folding at pH 10.0 and 0 °C.
In the two-chain folding of selenoinsulin, which has an interchain SeSe bridge in place of the CysA7‒CysB7 SS bridge of BPIns, it was proposed that the interchain SecA7‒SecB7 SeSe bridge is formed earlier than the CysA20‒CysB19 SS bridge due to larger thermodynamic stability of a SeSe bond than that of a SS bond41. Thus, in addition to the major pathways shown in Fig. 4, there would be minor double-chain folding pathways that proceed through formation of the CysA7‒CysB7 SS bridge before formation of the CysA20‒CysB19 SS bridge.
Optimization of the NCA conditions
According to the major NCA folding pathways of BPIns (Fig. 4), the folding yield could be improved by enhancing the probability of the formation of 2SS*. For this purpose, we explored optimal NCA folding conditions for BPIns by applying various pHs and temperatures in the presence of various additives. The results are summarized in Table 2 and Supplementary Table 2. A crucial factor must be the pH of the buffer solution because it can regulate the reactivity of SH groups and the conformation of Ins-A and Ins-B46. Indeed, when the two-chain assembly between 1SSA and RB was carried out at 4 °C at different pH values, the yield of N became maximal (13%) at pH 10.0 (Table 2, entries 1–5). This is probably due to the higher reactivity of the SH groups under more basic conditions as well as the robust structure of Ins-B formed at around pH 10,46. which can impede the undesired intrachain SS formation. Thus, Ins-B would preferentially form a SS bridge with Ins-A. At pH 11.0, however, the yield of N was only 3% due to preferential formation of a SS linkage within Ins-B.
Based on the populations of the SS components (Table 1), it is expected that 2SS*, which is a key heterodimeric intermediate, would be generated effectively by lowering the temperature, because the SS intermediates that have a native CysA6–CysA11 SS linkage in the 1SSA and 2SSA intermediate ensembles are more populated at lower temperatures. In addition, the low temperature would also stabilize 2SS* to prevent the reversible conversion to 1SSA or 2SSA. To carry out the chain assembly reaction below 0 °C, ethylene glycol (EG) (20%) was added to the buffer solution as an anti-freezing agent,55 which can simultaneously stabilize protein structures56,57.
When the temperature was decreased to ‒10 °C at pH 10.0, the yield of N increased up to 28% expectedly (Table 2, entries 6 and 7). Under this condition, generation of a large amount of 2SS* was indeed observed (Supplementary Fig. 5). In order to enhance the efficiency of NCA, sugar-based additives, such as xylitol (Xy) and inositol (Ino), which are known as stronger stabilizers of secondary and/or tertiary protein structure than EG,56,57 were subsequently tested instead of EG. The yield of N was not significantly enhanced with these additives, but the highest yield (30%) was achieved with Ino (0.25 M) (Table 2, entry 8). It should be noted that in the presence of these additives the concentration of urea can be reduced to 0.4 M. The lower concentration of urea must assist formation of N by stabilizing 2SS* relative to that generated in the presence of urea at a higher concentration. To further enhance the efficiency, we changed the NCA condition from 1SSA + RB to 2SSA + RB. Under this condition (entry 9), the folding yield was highest (34%) beyond that obtained from chain assembly folding of selenoinsulin41. When [CysA6‒CysA11, CysA7‒CysA20]A (peak G) was reacted with RB, the folding yield was not improved against our expectation (entry 10), and significant amount of swap species were generated. Although the yields achieved by NCA of the A- and B-chains in a 1:1 ratio is significantly higher than the previously reported yields (1–5%) for oxidative assembly of the two natural or synthetic chains under optimized conditions,8,9,10 an enormously long time (~10 days) was required until the reaction was completed. In order to solve this problem, NCA of RA and RB was then attempted in the presence of PDI, which is the most representative SS-forming and SS-reshuffling enzyme in an oxidative folding58. The GSSG and GSH were employed as redox reagents to promote the catalytic reaction of PDI (entry 11). As the result, the NCA at –10 °C and pH 10.0 was completed in 3 days (72 h) affording N in a yield of 39% (Supplementary Fig. 6). It should be noted that the efficiency of NCA was significantly decreased without PDI under the same NCA conditions in terms of both the velocity and yield (24%). The results obtained under other NCA conditions are summarized in Supplementary Table 2.
Applications of the NCA protocol to insulin families
To demonstrate the versatility as well as limitation of the developed NCA protocol, we applied it to the preparation of two additional hormone molecules and the variant of the insulin superfamily. The first synthetic target was human insulin (HIns), which differs from BPIns by ThrA8, IleA10, and ThrB30 instead of AlaA8, ValA10, and AlaB30. To avoid aggregation of the poorly soluble A-chain (HIns-A), it was synthesized with an isoacyl segment between ThrA8 and SerA9 as reported by Liu26 in a yield of 12.6%. On the other hand, the reduced B-chain of HIns (HIns-B) was prepared in a yield of 15.8% without applying the O-acyl isopeptide method. Before undertaking the NCA of HIns, it was confirmed that isoacyl HIns-A was immediately (<3 min) converted into the wild-type of HIns-A under the NCA conditions, i.e., in 25 mM sodium bicarbonate buffer solution at pH 10.0 and –10 °C (Supplementary Fig. 7). When the NCA of HIns-A and HIns-B chains was performed under the same conditions of entries 6 and 9 in Table 2, native HIns was obtained in yields of 18% and 12%, respectively, which are lower than the yields of BPIns. More strikingly, when the optimized condition with PDI, i.e., entry 11 in Table 2, was employed, native HIns was obtained in yields of 40% and 49% based on B-chain, by mixing reduced A-chain and B-chain in 1:1 and 2:1 ratios, respectively, in the presence of GSSG/GSH and PDI (Supplementary Fig. 8). The results of NCA for HIns are summarized in Supplementary Table 3. The yield of 49% achieved by NCA significantly exceeds the yields of the classical two-chain assembly method (1‒5%),8,9,10 but it is still below the yield of ~70% achieved by using a short ester linkage between the A- and B-chains34. or in vitro conversion of proinsulin6.
The second target protein was human type-II relaxin (HRlx-2), the amino acid sequence of which has 25% homology to that of HIns (Supplementary Fig. 9). HRlx-2 is known as an important pregnancy hormone helping relaxation of the uterus during pregnancy and delivery59. Recently, an additional function of HRlx-2 was suggested in treatment of acute heart failure probably due to the existence of the relaxin receptors in a peripheral nerve system60. Chemical synthesis of this insulin-like small protein has been object of intensive synthetic studies as HRlx-2 is difficult to isolate from mammalian tissues61. It was previously reported that native HRlx-2 can be obtained in a yield of about 30% by mixing the reduced A- and B-chains at pH 10.8 and 4 °C through a chain combination pathway similar to Fig. 442. More efficient preparation of HRlx-2 (48% yield) was achieved by application of the Met(O)B25 B-chain with increased solubility in water43. To apply the NCA method to HRlx-2, the reduced native A-chain (RRlx-A) was prepared by following the previous report46. Similarly, the reduced native B-chain (RRlx-B) was synthesized by SPPS and purified by RP-HPLC. Then the NCA protocol was applied for preparation of native HRlx-2 (NRlx). A solution of 1SSRlx-A, which was prepared by reaction of one equivalent of DHSox with RRlx-A at pH 10.0, was mixed with RRlx-B at 15 °C. The products were analyzed by RP-HPLC after quenching the reaction with AEMTS (Supplementary Fig. 9). The correctly folded HRlx-2 was indeed obtained almost exclusively with a yield of 47% in 26 h. The yield was comparable to the value reported in the literature43 despite the easier protocol.
The third target was human insulin ValA16 variant, which is known as a nonfoldable insulin analog by the oxidative two-chain assembly,44 although a DesDi approach has recently succeeded in the synthesis of the correctly folded ValA16 variant via a single-component insulin precursor35. It was, therefore, not surprising that the native fold of the ValA16 variant was not attained under NCA conditions. Indeed, formation of only oxidized species of A-chain and oligomers of B-chain was observed in the HPLC chromatograms, indicating a failure of coupling between the A- and B-chains (Supplementary Fig. 10). The result suggests that the NCA protocol is applicable only to foldable insulin and the analogs.
Discussion
We have characterized the major two-chain oxidative folding pathways of BPIns at pH 10 as shown in Fig. 4. When the reduced A-chain (RA) and B-chain (RB) were mixed in a buffer solution, the native CysA6‒CysA11 SS bond formed most abundantly in A-chain, and then A- and B-chains would couple together to form metastable heterodimeric intermediates, 1SS* and 2SS* both containing the native disulfide bridge CysA20‒CysB19. These key folding intermediates, which could even be observed upon reduction of native BPIns (N) by excess DTT, are then converted to N through formation of the other SS bridge.
On the basis of the two-chain folding pathways of BPIns revealed in this study, the NCA conditions were optimized. As a result, the yield of BPIns could be increased up to 34% at pH 10.0 and –10 °C in the presence of EG. Moreover, by utilizing PDI as a catalyst, the reaction time was significantly shortened and the yield was further increased to 39%. When a similar NCA protocol was applied to human insulin (HIns) and human type-II relaxin (HRlx-2), native HIns was obtained in a yield of 49% based on the B-chain when the B-chain was mixed with two equivalents of the A-chain, and HRlx-2 was obtained in a yield of 47%. However, limitation of NCA was also evidenced by the unsuccessful application to the two-chain folding of human insulin ValA16 variant, which is known as nonfoldable insulin44.
In conclusion, the NCA protocol reinvestigated herein may offer simple and promising approach to the total chemical synthesis of various insulins without using any genetic recombinant methodologies. The major two-chain folding pathways of BPIns (Fig. 4) were essentially the same as those of the proinsulin, although the folding yield (up to 39%) was lower than that of the proinsulin (~70%). This confirms that the C-peptide in proinsulin does not play an important role in the achievement of the proper folding but rather decreases the entropic penalty of the crosslinking reaction between A- and B-chains. Indeed, NCA just relies on the thermodynamic stability of the native fold, which properly guides component A- and B-chains to assemble into the native disulfide framework. If such a chain coupling is prohibited by any reasons, as evidenced in the case of the ValA16 variant, NCA would not produce a native insulin fold. Although the chain assembly reaction proceeds slowly, the simple experimental procedure, i.e., just mixing the A- and B-chains with no modification on the functional groups, would be a merit of the NCA protocol. Moreover, this protocol may expand accsess to foldable insulin analogs, which cannot be obtained by the rDNA technology, because all the processes, including SPPS and NCA, are based on chemical reactions, which would enable insertion of unnatural amino acids into the primary sequence.
Methods
Materials
BPIns was purchased from Sigma Aldrich, Japan, and used without purification. 2-Aminoethyl methanethiosulfonate (AEMTS)62 and 3,4-dihydroxy-1-selenolane Se-oxide (DHSox)63 were synthesized according to the literature methods. Plasmid used for expression of PDI was constructed as described previously64. PDI was overexpressed in Escherichia coli. Cells were harvested and homogenized. Recombinant protein was purified by a combination of several types of chromatography. Purity of PDI was confirmed to be over 95% by SDS-PAGE and protein concentration was determined by BCA method65. All other general reagents were commercially available and used without further purification.
General two-chain folding of BPIns with NCA
Solutions of the reduced A-chain (RA) and B-chain (RB), which were obtained by the reduction of native BPIns with DTT,46 were prepared by dissolving RA (0.4 mg) or RB (1.0 mg) in a sodium bicarbonate buffer solution (25 mM at pH 10.0, 120 μL) containing urea (1 or 2 M, respectively) and EDTA (1 mM). The concentrations of RA and RB were determined by UV absorbance at 275 nm using the molar extinction coefficients (3100 and 2690 M−1 cm−1, respectively46). Then, the solutions were diluted with the same buffer solution to adjust the concentrations to be 710–750 μM. To the RA solution (70 μL), DHSox (0, 1, or 2 eq) dissolved in the same buffer solution (5 μL) was added. Similarly, DHSox (0, or 1 eq) dissolved in the same buffer solution (10 μL) was added to the RB solution (80 μL). The prepared solutions were chilled at 4 °C for 15–30 min. The folding was initiated by vigorously mixing the RA, 1SSA, or 2SSA solution (75 μL) with the RB or 1SSB solution (75 μL) for 15 s, and the mixture was immediately diluted with a sodium bicarbonate buffer solution (25 mM at pH 10.0, 100 μL) containing EDTA (1 mM) and a various concentration of sugar-based additives at various temperatures (–10–4 °C). In the case of NCA using glutathione and PDI (i.e., the condition for entry 11 in Table 2), a sodium bicarbonate buffer solution (25 mM at pH 10.0, 100 μL) containing EDTA (1 mM), EG (25%), GSSG (0.5 mM), GSH (2.5 mM), and PDI (10 μM) was added to the mixture solution (150 μL) of RA and RB. The temperature was controlled within ± 0.3 °C by using a dry thermostatic chamber. After a certain period of time, the aliquot (20 μL) was transferred into an aqueous solution of 2-aminoethyl methanethiosulfonate (AEMTS) (8 mg mL−1, 200 μL) as the thiol-blocking reagent, chilled at 4 °C in a micro-centrifuge tube. After 15 min, the sample solution was diluted with an aqueous TFA solution (0.1%, 800 μL) and stored at −30 °C. The sample solution (1 mL) was analyzed by using the HPLC system equipped with a sample solution loop (1 mL) and a Tosoh TSKgel ODS-100V 4.6 × 150 reverse-phase (RP) column, which was equilibrated with a 80:20 (v/v) mixture of TFA (0.1%) in water (eluent A) and TFA (0.1%) in acetonitrile (eluent B) at a flow rate of 1.0 ml min−1. A solvent gradient (i.e., the ratio of eluent B linearly increased from 20 to 36% in 0–15 min, from 36% to 39% in 15–20 min, from 39 to 40% in 20–23 min) was applied. The folding intermediates produced were detected by the absorbance at 280 nm. The NCA yield based on A- or B-chain was estimated from the peak area on the RP-HPLC chromatograms.
Reductive unfolding of BPIns
Reductive unfolding was initiated by mixing the solutions of BPIns (1 mg mL−1, 475 μL) and DTT (10 mM, 25 μL) in sodium bicarbonate buffer at pH 10.0 and 0 °C. After a certain period of time, an aliquot (60 μL) was transferred into an aqueous AEMTS solution (8 mg mL−1, 200 μL) chilled at 0 °C in a micro-centrifuge tube. After 15 min, the mixture was diluted with an aqueous TFA solution (0.1%, 800 μL) and stored at −30 °C. The sample solutions were analyzed by RP-HPLC. The HPLC conditions were the same as those described above.
Enzymatic digestion of 2SS* and 1SS*
Solutions of BPIns, 2SS*, and 1SS*, which were collected through an analytical RP-HPLC column (Tosoh TSKgel ODS-100V reverse-phase 150 × 4.6 column), were lyophilized. The obtained white powder was dissolved in an aqueous TFA solution (0.1%, 7 μL) and subsequently diluted with an appropriate volume of a Tris-HCl buffer solution (100 mM at pH 8.0) so that the final concentration of the peptide is approximately 1 mg mL−1. To the peptide solution (100 μL) was added α-chymotrypsin (Sigma-Aldrich Japan) solution (1.5 mg mL−1, 1 μL) in 1 mM HCl, and the reaction was progressed at 23 °C for 120 min. The reaction was quenched by addition of an aqueous TFA solution (5%, 10 μL). The sample solution (20 μL) containing the digested peptide fragments was analyzed on a Jeol JMS-T100LP spectrometer connected to an Agilent 1200 series HPLC system equipped with a 20 μL sample solution loop and a Tosoh TSKgel ODS-100V 4.6 × 150 RP-column. The column was first equilibrated with 0.1% formic acid in water (eluent A) at a flow rate of 0.8 mL/min. After injection of the sample solution, a solvent gradient was applied by linearly increasing a ratio of 0.1% formic acid in acetonitrile (eluent B) from 0 to 15% in 0–30 min, from 15 to 30% in 30–50 min, from 30 to 40% in 50–60 min, and from 40 to 80% in 60–63 min.
SPPS of BPIns [CysA6‒CysA20]A
Fmoc-Asn(Trt)-CLEAR acid resin (360 mg, 0.1 mmol) was swelled with NMP for 1 h at room temperature. After removing the NMP, the peptide chain was elongated by using an automated microwave peptide synthesizer (Liberty Blue, CEM corporation). The obtained resin was washed with MeOH (×3) and dried in vacuo to yield H-Gly-Ile-Val-Glu(OBut)-Gln(Trt)-Cys(Trt)-Cys(MPM)-Ala-Ser(But)-Val-Cys(MPM)-Ser(But)-Leu-Tyr(But)-Gln(Trt)-Leu-Glu(OBut)-Asn(Trt)-Tyr(But)-Cys(Trt)-Asn(Trt)-CLEAR acid resin (520 mg). A portion of the resin (103 mg, 21 μmol) was treated with TFA cocktail (TFA: triisopropylsilane:H2O = 92.5:2.5:2.5, 2.5 mL) for 2 h at 4 °C. After the removal of TFA by N2 stream, the crude peptide was precipitated with Et2O, washed with Et2O (×3), and dried in vacuo. The obtained crude peptide, H-Gly-Ile-Val-Glu-Gln-Cys-Cys(MPM)-Ala-Ser-Val-Cys(MPM)-Ser-Leu-Tyr-Gln-Leu-Glu-Asn-Tyr-Cys-Asn-OH, was dissolved in sodium bicarbonate buffer (200 mM at pH 9.0) solution containing 6 M guanidine hydrochloride (Gdn HCl) and 20% (v/v) DMSO and then shaken for 48 h at room temperature. The resulting solution was purified by RP-HPLC to give H-Gly-Ile-Val-Glu-Gln-Cys-Cys(MPM)-Ala-Ser-Val-Cys(MPM)-Ser-Leu-Tyr-Gln-Leu-Glu-Asn-Tyr-Cys-Asn-OH. The MPM-protected [CysA6‒CysA20] was dissolved in TFA cocktail (TFA:2,2′-dipyridyldisulfide:thioanisole = 92.5:5:2.5, 2.5 mL) and incubated for 1 h at room temperature to give H-Gly-Ile-Val-Glu-Gln-Cys-Cys(Pys)-Ala-Ser-Val-Cys(Pys)-Ser-Leu-Tyr-Gln-Leu-Glu-Asn-Tyr-Cys-Asn-OH, where Pys represents a 2-pyridylsulfanyl group. After the removal of TFA by N2 stream, the crude peptide was precipitated with Et2O, washed with Et2O (×3), and dried in vacuo. The precipitate was dissolved in H2O-acetonitrile (1:1 v/v) containing 6 M Gdn HCl and 0.1% TFA, and the solution was treated with tris(2-carboxyethyl)phosphine (TCEP) hydrochloride (12.5 mg, 42 μmol) for 1 h at room temperature. The resulting sample solution was purified by RP-HPLC to give H-Gly-Ile-Val-Glu-Gln-Cys-Cys-Ala-Ser-Val-Cys-Ser-Leu-Tyr-Gln-Leu-Glu-Asn-Tyr-Cys-Asn-OH. Yield: 476 nmol, 2.3% from the starting resin. ESI-MS, found: 1169.5, calcd for [M + 2 H]2+: 1169.8. Amino acid analysis (AAA): Asp2.05Ser1.69Glu3.67Gly1.22Ala1.07Val1.82Ile0.63Leu2Tyr1.99.
SPPS of the other 1SSA species
Similar procedures were applied for SPPS of the 1SSA species of BPIns other than [CysA6‒CysA20]A. [CysA7‒CysA20]A: Yield, 3.2%; ESI-MS, found: 1169.5, calcd for [M + 2 H]2+: 1169.8; Amino acid analysis, Asp2.05Ser1.59Glu3.84Gly1.21Ala1.04Val1.78Ile0.70Leu2Tyr1.95. [CysA6‒CysA11]A: Yield, 0.3%; ESI-MS, found: 1169.5, calcd for [M + 2 H]2+: 1169.8; Amino acid analysis, Asp2.21Ser1.99Glu4.44Gly1.71Ala1.11Val1.99Ile0.70Leu2Tyr1.91. [CysA7‒CysA11]A: Yield, 0.7%; ESI-MS, found: 1169.6, calcd for [M + 2 H]2+: 1169.8; Amino acid analysis, Asp2.06Ser1.71Glu3.86Gly1.28Ala1.04Val1.64Ile0.64Leu2Tyr1.96. [CysA11‒CysA20]A: Yield, 3.0%; ESI-MS, found: 1169.5, calcd for [M + 2 H]2+: 1169.8; Amino acid analysis, Asp2.03Ser2.07Glu3.72Gly1.61Ala1.13Val1.68Ile0.62Leu2Tyr1.91. [CysA6‒CysA7]A: Yield, 1.9%; ESI-MS, found: 1169.5, calcd for [M + 2 H]2+: 1169.8; Amino acid analysis, Asp2.10Ser1.80Glu4.23Gly1.44Ala1.08Val1.72Ile0.65Leu2Tyr1.97.
Identification of the SS intermediates
The AEMTS-blocked folding intermediates fractionated by HPLC were collected and lyophilized. The resulting white powder of each intermediate was dissolved in an aqueous TFA solution (0.1%). The number of SS linkages involved in the intermediate was determined by measurement of the molecular weight with ESI(+)-MS spectrometry. The SS positions of 1SSA (peaks A‒E in Fig. 1b and Supplementary Fig. 1) were determined by comparing the HPLC retention times for peaks A‒E with those for the reference samples obtained by SPPS (see above). The reference sample solutions were prepared as follows. The synthesized sample (50 nmol), e.g., [CysA7‒CysA11]A, was dissolved in aqueous acetonitrile (50%, 300 μL) containing TFA (0.1%), and the portion (20 μL) was added to the AEMTS solution (8 mg mL−1) in water (200 μL). After dilution with aqueous TFA (0.1%, 800 μL), the sample solution was injected onto the HPLC system under the same analytical conditions as those applied in the HPLC analysis of peaks A‒E.
Similarly, the SS positions of 2SSA were determined by comparing the HPLC retention times for peaks F‒H (Fig. 1c and Supplementary Fig. 1) with those for the reference samples, i.e., [CysA7‒CysA11, CysA6‒CysA20]A, [CysA6‒CysA11, CysA7‒CysA20]A, and [CysA6‒CysA7, CysA11‒CysA20]A, which were obtained by oxidation of the synthesized 1SSA samples with DHSox (4 eq) in a sodium bicarbonate buffer solution (25 mM at pH 10.0) containing 1 M urea.
SPPS of O-acyl iso-human insulin A-chain and its ValA16 mutant
Fmoc-Asn(Trt)-CLEAR acid resin (0.40 mmol g−1, 500 mg, 0.20 mmol) was swelled with NMP for 30 min at room temperature. After washing with NMP, the peptide chain was elongated by the Fmoc-SPPS. Fmoc group was removed by treating with 20% piperidine/NMP for 20 min. The amino acids (0.80 mmol each) were activated by mixing with 1 M DCC/NMP (1.2 mL) and 1 M HOBt/NMP (1.2 mL) at room temperature for 30 min, and the coupling reaction was carried out at 50 °C for 1 h. At Thr8-Ser9 position, Boc-Ser[Fmoc-Thr(But)]-OH (280 mg, 0.48 mmol) was used as an O-acyl isopeptide component. After elongation, H-Gly-Ile-Val-Glu(OBut)-Gln(Trt)-Cys(Trt)-Cys(Trt)-Thr(But)-OCH2CH(NHBoc)CO-Ile-Cys(Trt)-Ser(But)-Leu-Tyr(But)-Gln(Trt)-Leu-Glu(OBut)-Asn(Trt)-Tyr(But)-Cys(Trt)-Asn(Trt)-resin (1.15 g) was obtained. A part of the resin (330 mg, 57.2 µmol) was treated with a TFA cocktail (TFA/H2O/phenol/thioanisole/triisopropylsilane, 82.5/5/5/5/2.5, 5 mL) at room temperature for 2 h. TFA was removed under nitrogen stream and the peptide was precipitated with diethyl ether. After washing twice with ether, the precipitate was dried under vacuum. The crude peptide was purified by RP-HPLC to give H-Gly-Ile-Val-Glu-Gln-Cys-Cys-Thr-OCH2CH(NH2)CO-Ile-Cys-Ser-Leu-Tyr-Gln-Leu-Glu-Asn-Tyr-Cys-Asn-OH (7.19 µmol, 12.6% yield). MALDI-TOF mass, found: m/z 2383.9, calcd: 2384.7 for (M + H)+. Amino acid analysis: Asp2.46Thr0.85Ser1.96Glu4.30Gly1Val0.71Ile1.34Leu2.47Tyr2.42.
For the synthesis of L16V mutant of A chain, starting from Fmoc-Asn(Trt)-CLEAR acid resin (0.40 mmol/g, 250 mg, 0.10 mmol), the peptide chain was elongated essentially according to the method described above, except that Fmoc-Val-OH was used at Leu16 position. After the chain assembly, H-Gly-Ile-Val-Glu(OBut)-Gln(Trt)-Cys(Trt)-Cys(Trt)-Thr(But)-OCH2CH(NHBoc)CO-Ile-Cys(Trt)-Ser(But)-Leu-Tyr(But)-Gln(Trt)-Val-Glu(OBut)-Asn(Trt)-Tyr(But)-Cys(Trt)-Asn(Trt)-resin (457 mg) was obtained. A part of the resin (115 mg, 25.2 µmol) was treated with a TFA cocktail (1.5 mL) at room temperature for 2 h. TFA was removed under nitrogen stream and the peptide was precipitated with diethyl ether. After washing twice with ether, the precipitate was dried under vacuum. The crude peptide was purified by RP-HPLC to give H-Gly-Ile-Val-Glu-Gln-Cys-Cys-Thr-OCH2CH(NH2)CO-Ile-Cys-Ser-Leu-Tyr-Gln-Val-Glu-Asn-Tyr-Cys-Asn-OH (1.88 µmol, 7.5% yield). MALDI-TOF mass, found: m/z 2368.8, calcd: 2369.0 for (M + H)+. Amino acid analysis: Asp2.03Thr0.85Ser1.59Glu4.23Gly1Val1.85Ile1.41Leu1.01Tyr1.98.
SPPS of human insulin B-chain
Starting from Fmoc-Thr(But)-Alko resin (0.56 mmol g−1, 357 mg, 0.20 mmol), peptide chain was elongated by the method as for human insulin A chain. After the chain assembly, H-Phe-Val-Asn(Trt)-Gln(Trt)-His(Trt)-Leu-Cys(Trt)-Gly-Ser(But)-His(Trt)-Leu-Val-Glu(OBut)-Ala-Leu-Tyr(But)-Leu-Val-Cys(Trt)-Gly-Glu(OBut)-Arg(Pbf)-Gly-Phe-Phe-Tyr(But)-Thr(But)-Pro-Lys(Boc)-Thr(But)-resin (1.25 g) was obtained. A part of the resin (101 mg, 16.2 µmol) was treated with a TFA cocktail (TFA/H2O/phenol/thioanisole/triisopropylsilane, 82.5/5/5/5/2.5, 1.5 mL) at room temperature for 2 h. TFA was removed under nitrogen stream and the peptide was precipitated with diethyl ether. After washing twice with ether, the precipitate was dried under vacuum. The crude peptide was purified by RP-HPLC to give H-Phe-Val-Asn-Gln-His-Leu-Cys-Gly-Ser-His-Leu-Val-Glu-Ala-Leu-Tyr-Leu-Val-Cys-Gly-Glu-Arg-Gly-Phe-Phe-Tyr-Thr-Pro-Lys-Thr-OH (2.55 µmol, 15.8% yield). MALDI-TOF mass, found: m/z 3431.0, calcd: 3430.9 for (M + H)+. Amino acid analysis: Asp0.94Thr1.85Ser0.79Glu2.89Pro1.29Gly3Ala1.01Val2.78Leu3.82Tyr2.42Phe2.97Lys1.04His1.86Arg1.01.
Two-chain folding of HIns with NCA
The same NCA protocol as that had been applied for BPIns was applied to the two-chain assembly of native HIns-A (50 nmol) (or its L16V mutant (50 nmol)) and HIns-B (50 nmol) synthesized above. The AEMTS-quenched sample solutions were analyzed by RP-HPLC with slight modification of the solvent-gradient condition. The ratio of eluent B was 20 to 30% in 0–5 min, from 30 to 35% in 5–13.5 min, from 35 to 0% in 13.5–14 min, and kept constant in 14–20 min at a flow rate of 0.8 mL/min.
Two-chain folding of HRlx-2 with NCA
The NCA synthesis of HRlx-2 was performed as follows. RRlx-A (0.2 mg) and RRlx-B (0.1 mg), which were obtained by SPPS synthesis,46,66 were dissolved in a sodium bicarbonate buffer solution (25 mM at pH 10.0, 15 or 30 μL, respectively) containing urea (3 or 6 M, respectively) and EDTA (1 mM). The solutions of RRlx-A and RRlx-B were diluted with the same buffer solution (75 μL or 30 μL, respectively) without urea. The concentrations of RRlx-A and RRlx-B were determined by UV absorbance at 275 nm using the molar extinction coefficients (1650 and 8300 M−1 cm−1, respectively46,66). Then, the solutions were diluted with the same buffer solution to adjust the concentrations to be 500–720 μM. To the RRlx-A solution (35 μL), DHSox (1 eq) dissolved in the same buffer solution (5 μL) was added so as to obtain 1SSRlx-A as a major product in the solution, and the solution was chilled at 4 °C for 5 min. The folding was initiated by vigorously mixing the 1SSRlx-A solution (20 μL) with the RRlx-B solution (20 μL) for 15 s, and the mixture was immediately diluted with a sodium bicarbonate buffer solution (25 mM at pH 10.0, 40 μL) containing EDTA (1 mM) at 15 °C. The temperature was controlled within ±0.1 °C by using a dry thermostatic chamber. After a certain period of time, an aliquot (20 μL) was transferred into an aqueous AEMTS solution (8 mg mL−1, 200 μL) chilled at 4 °C in a micro-centrifuge tube. After 15 min, the sample solution was diluted with an aqueous TFA solution (0.1%, 800 μL) and stored at −30 °C. The samples thus obtained were analyzed by RP-HPLC under the similar conditions to those for BPIns except for an eluent gradient condition, i.e., the concentration of eluent B (0.1% TFA in MeCN) was lineally increased from 20 to 30% in 0‒10 min, from 30 to 60% in 10‒20 min. The folded Rlx isolated from HPLC was identified by ESI-MS. ESI-MS (m/z) found for [M + 4 H]4+: 1496.1, calcd for [M + 4 H]4+: 1496.2.
Data availability
The authors declare that all other data supporting the findings of this study are available within the paper and its supplementary information files, or from the authors upon reasonable request.
References
Ryle, A. P., Sanger, F., Smith, L. F. & Kitai, R. The disulphide bonds of insulin. Biochem. J. 60, 541–556 (1955).
Kemmler, W., Peterson, J. D. & Steiner, D. F. Studies on the conversion of proinsulin to insulin I. Conversion in vitro with trypsin and carboxypeptidase B. J. Biol. Chem. 246, 6786–6791 (1971).
Steiner, D. F. The proprotein convertases. Curr. Opin. Chem. Biol. 2, 31–39 (1998).
Frank, B. H. & Chance, R. E. Two routes for producing human insulin utilizing recombinant DNA technology. MMW Munch. Med. Wochenschr. Suppl 1, S14–S20 (1983).
Kjeldsen, T. Yeast secretory expression of insulin precursors. Appl. Microbiol. Biotechnol. 54, 277–286 (2000).
Mayer, J. P., Zhang, F. & DiMarchi, R. D. Insulin structure and function. Biopolym. 88, 687–713 (2007).
Dixon, G. H. & Wardlaw, A. C. Regeneration of insulin activity from the separated and inactive A and B chains. Nature 188, 721–724 (1960).
Meienhofer, J. et al. Synthesis of insulin and their combination to insulin-active preparations. Z. Naturforsch. B. 18, 1120–1121 (1963).
Katsoyannis, P. G., Fukuda, K., Tometsko, A., Suzuki, K. & Tilak, M. Insulin peptides. X. The synthesis of the B-chain of insulin and its combination with natural or synthetic A-chain to generate insulin activity. J. Am. Chem. Soc. 86, 930–932 (1964).
Kung, Y. T., Du, Y. C., Huang, W. T., Chen, C. C. & Ke, L. T. Total synthesis of crystalline bovine insulin. Sci. Sin. 14, 1710–1716 (1965).
Kauzmann, W. II. in Sulfur in Proteins (eds Benesch, R. et al.) 93–108 (Academic Press, Cambridge, 1959).
Chance, R. E. et al. in Pept.: Synth., Struct., Funct., Proc. Am. Pept. Symp., 7th, (eds. Rich, D. H. and Gross, E.) 721–728 (1981).
Tang, J. G., Wang, C. C. & Tsou, C. L. Formation of native insulin from the scrambled molecule by protein disulphide-isomerase. Biochem. J. 255, 451–455 (1988).
Wang, C.-C. & Tsou, C.-L. The insulin A and B chains contain sufficient structural information to form the native molecule. Trends Biochem. Sci. 16, 279–281 (1991).
Moroder, L., Besse, D., Musiol, H.-J., Rudolph-Böhner, S. & Siedler, F. Oxidative folding of cystine-rich peptides vs regioselective cysteine pairing strategies. Pept. Sci. 40, 207–234 (1996).
Adams, M. J. et al. Structure of rhombohedral 2 zinc insulin crystals. Nature 224, 491–495 (1969).
Wollmer, A., Brandenburg, D., Vogt, H. P. & Schermutzki, W. Reduction/reoxidation studies with cross-linked insulin derivatives. Hoppe. Seylers. Z. Physiol. Chem. 355, 1471–1476 (1974).
Obermeier, R. & Geiger, R. A new bifunctional reagent for the intramolecular crosslinking of insulin (author’s transl). Hoppe. Seylers. Z. Physiol. Chem. 356, 1631–1634 (1975).
Busse, W. D. & Carpenter, F. H. Synthesis and properties of carbonylbis(methionyl)insulin, a proinsulin analog which is convertible to insulin by cyanogen bromide cleavage. Biochemistry 15, 1649–1657 (1976).
Mutter, M. et al. Switch peptides in statu nascendi: induction of conformational transitions relevant to degenerative diseases. Angew. Chem. Int. Ed. 43, 4172–4178 (2004).
Carpino, L. A. et al. Synthesis of ‘difficult’ peptide sequences: application of a depsipeptide technique to the Jung–Redemann 10- and 26-mers and the amyloid peptide Aβ(1–42). Tetrahedron Lett. 45, 7519–7523 (2004).
Sohma, Y. et al. The ‘O-acyl isopeptide method’ for the synthesis of difficult sequence-containing peptides: application to the synthesis of Alzheimer’s disease-related amyloid β peptide (Aβ) 1–42. J. Pept. Sci. 11, 441–451 (2005).
Taniguchi, A. et al. “Click peptide” based on the “O-acyl isopeptide method”: control of Aβ1–42 production from a photo-triggered Aβ–42 analogue. J. Am. Chem. Soc. 128, 696–697 (2006).
Sohma, Y. et al. O-Acyl isopeptide method’ for the efficient synthesis of difficult sequence-containing peptides: use of ‘O-acyl isodipeptide unit’. Tetrahedron Lett. 47, 3013–3017 (2006).
Coin, I. et al. Depsipeptide methodology for solid-phase peptide synthesis: circumventing side reactions and development of an automated technique via depsidipeptide units. J. Org. Chem. 71, 6171–6177 (2006).
Liu, F., Luo, E. Y., Flora, D. B. & Mezo, A. R. A synthetic route to human insulin using isoacyl peptides. Angew. Chem. Int. Ed. 53, 3983–3987 (2014).
Shabanpoor, F., Hossain, M. A., Lin, F. & Wade, J. D. Sequential formation of regioselective disulfide bonds in synthetic peptides with multiple disulfide bonds. Methods Mol. Biol. 1047, 81–87 (2013).
Moroder, L., Musiol, H.-J., Götz, M. & Renner, C. Synthesis of single- and multiple-stranded cystine-rich peptides. Biopolym. (Pept. Sci.) 80, 85–97 (2005).
Postma, T. M. & Albericio, F. Disulfide formation strategies in peptide synthesis. Eur. J. Org. Chem. 2014, 3519–3530 (2014).
Sieber, P., Kamber, B., Hartmann, A., Jöhl, A., Riniker, B., Rittel, W. Totalsynthese von Humaninsulin. IV. Beschreibung der Endstufen. Helvetica Chimica Acta 60 27–37 (1977).
Akaji, K., Fujino, K., Tatsumi, T. & Kiso, Y. Total synthesis of human insulin by regioselective disulfide formation using the silyl chloride-sulfoxide method. J. Am. Chem. Soc. 115, 11384–11392 (1993).
Liu, F., Zaykov, A. N., Levy, J. J., DiMarchi, R. D. & Mayer, J. P. Chemical synthesis of peptides within the insulin superfamily. J. Pept. Sci. 22, 260–270 (2016).
Moroder, L. & Musiol, H.-J. Insulin–from its discovery to the industrial synthesis of modern insulin analogues. Angew. Chem. Int. Ed. 56, 10656–10669 (2017).
Sohma, Y., Hua, Q.-X., Whittaker, J., Weiss, M. A. & Kent, S. B. H. Design and folding of [GluA4(OβThrB30)]insulin (“Ester insulin”): a minimal proinsulin surrogate that can be chemically converted into human insulin. Angew. Chem. Int. Ed. 49, 5489–5493 (2010).
Zaykov, A. N., Mayer, J. P., Gelfanov, V. M. & DiMarchi, R. D. Chemical synthesis of insulin analogs through a novel precursor. Acs. Chem. Biol. 9, 683–691 (2014).
Thalluri, K. et al. Biomimetic synthesis of insulin enabled by oxime ligation and traceless “C-peptide” chemical excision. Org. Lett. 19, 706–709 (2017).
Thalluri, K. et al. Synthesis of relaxin-2 and insulin-like peptide 5 enabled by novel tethering and traceless chemical excision. J. Pept. Sci. 23, 455–465 (2017).
Dhayalan, B. et al. Scope and limitations of Fmoc chemistry SPPS-based approaches to the total synthesis of insulin lispro via ester insulin. Chem. Eur. J. 23, 1709–1716 (2017).
Musiol, H.-J. & Moroder, L. Diselenides as proxies of disulfides in cystine-rich peptides. Chem. Today 30, 14–19 (2012).
Metanis, N., Beld, J. & Hilvert, D. in PATAI’S Chemistry of Fu nctional Groups (John Wiley & Sons, New Jersey, 2009).
Arai, K. et al. Preparation of selenoinsulin as a long-lasting Insulin analogue. Angew. Chem. Int. Ed. 56, 5522–5526 (2017).
Tang, J.-G., Wang, Z.-H., Tregear, G. W. & Wade, J. D. Human gene 2 relaxin chain combination and folding. Biochemistry 42, 2731–2739 (2003).
Barlos, K. K., Gatos, D., Vasileiou, Z. & Barlos, K. An optimized chemical synthesis of human relaxin-2. J. Pept. Sci. 16, 200–211 (2010).
Liu, M. et al. Crystal structure of a “nonfoldable” insulin: impared folding effeciency despite native activity. J. Biol. Chem. 284, 35259–35272 (2009).
Bathgate, R. A. D. et al. Relaxin-3: improved synthesis strategy and demonstration of its high-affinity interaction with the relaxin receptor LGR7 both in vitro and in vivo. Biochemistry 45, 1043–1053 (2006).
Arai, K. et al. A water-soluble selenoxide reagent as a useful probe for the reactivity and folding of polythiol peptides. FEBS Open Bio 3, 55–64 (2013).
Arai, K., Dedachi, K. & Iwaoka, M. Rapid and quantitative disulfide bond formation for a polypeptide chain using a cyclic selenoxide reagent in an aqueous medium. Chem. Eur. J. 17, 481–485 (2011).
Hua, Q.-X. et al. Structure of a protein in a kinetic trap. Nat. Struct. Mol. Biol. 2, 129–138 (1995).
Hua, Q.-X., Jia, W., Frank, B. H., Phillips, N. F. B. & Weiss, M. A. A protein caught in a kinetic trap: Structures and stabilities of insulin disulfide isomers. Biochemistry 41, 14700–14715 (2002).
Qiao, Z.-S., Guo, Z.-Y. & Feng, Y.-M. Putative disulfide-forming pathway of porcine insulin precursor during its refolding in vitro. Biochemistry 40, 2662–2668 (2001).
Tang, Y., Wang, S., Chen, Y., Xu, G. & Feng, Y. In vitro insulin refolding: Characterization of the intermediates and the putative folding pathway. Sci. China C. Life. Sci. 50, 717–725 (2007).
Hua, Q., Mayer, J. P., Jia, W., Zhang, J. & Weiss, M. A. The folding nucleus of the insulin superfamily: a flexible peptide model foreshadows the native state. J. Biol. Chem. 281, 28131–28142 (2006).
Steiner, D. F. & Chan, S. J. An overview of insulin evolution. Horm. Metab. Res. 20, 443–444 (1988).
Hua, Q. et al. Mechanism of insulin chain combination: asymmetric roles of A-chain α-helices in disulfide pairing. J. Biol. Chem. 277, 43443–43453 (2002).
Carpenter, J. F., Prestrelski, S. J. & Arakawa, T. Separation of freezing- and drying-induced denaturation of lyophilized proteins using stress-specific stabilization. Arch. Biochem. Biophys. 303, 456–464 (1993).
Gekko, K. & Morikawa, T. Preferential hydration of bovine serum albumin in polyhydric alcohol-water mixtures. J. Biochem. 90, 39–50 (1981).
Gekko, K. & Morikawa, T. Thermodynamics of polyol-induced thermal stabilization of chymotrypsinogen. J. Biochem. 90, 51–60 (1981).
Wilkinson, B. & Gilbert, H. F. Protein disulfide isomerase. Biochim. Biophys. Acta. 1699, 35–44 (2004).
Schwabe, C. & Büllesbach, E. E. Relaxin: structures, functions, promises, and nonevolution. FASEB J. 8, 1152–1160 (1994).
Bathgate, R.a. D. et al. Relaxin family peptides and their receptors. Physiol. Rev. 93, 405–480 (2013).
Hossain, M. A. & Wade, J. D. Synthetic relaxins. Curr. Opin. Chem. Biol. 22, 47–55 (2014).
Bruice, T. W. & Kenyon, G. L. Novel alkyl alkanethiolsulfonate sulfhydryl reagents. J. Protein Chem. 1, 47–58 (1982).
Iwaoka, M., Takahashi, T. & Tomoda, S. Syntheses and structural characterization of water-soluble selenium reagents for the redox control of protein disulfide bonds. Heteroat. Chem. 12, 293–299 (2001).
Inaba, K. et al. Crystal structures of human Ero1α reveal the mechanisms of regulated and targeted oxidation of PDI. EMBO J. 29, 3330–3343 (2010).
Okumura, M. et al. Inhibition of the functional interplay between endoplasmic reticulum (ER) oxidoreduclin-1α (Ero1α) and protein-disulfide isomerase (PDI) by the endocrine disruptor bisphenol A. J. Biol. Chem. 289, 27004–27018 (2014).
Iwaoka, M. et al. Method for producing relaxin. JP Patent 2015160825 A (2015).
Acknowledgements
We thank Dr. Y. Asahina (Osaka University) to kindly support SPPS of the authentic SS intermediates of BPIns. We also thank T. Sato and Y. Uchida (Tokai University) for assistance of folding experiments of BPIns and Y. Kubo (Fukuoka University) for assistance of synthesis of HRlx-2 A- and B-chains. K.A. acknowledges Japan Society for the Promotion of Science (JSPS) for research fellowship for young scientists. This work was supported by KAKENHI Grant Number 26888016 for Research Activity Start-up (to K.A.), in part by Research and Study Project of Tokai University, Educational System General Research Organization (to K.A. and M.I.), Collaborative Research Program of Institute for Protein Research, Osaka University CR-15-01 (M.I. and H.H.), and Cooperative Research Program of Network Joint Research Center for Material and Devices (M.I. and K.I.).
Author information
Authors and Affiliations
Contributions
K.A. and T.T. equally contributed to this work. In addition, these authors designed and conducted the each experimental part. K.A. and M.I. developed an idea of this project and initiated. K.A., R.S., M.N., S.F., and M.I. performed the NCA experiments of BPIns, HIns, and Rlx. T.T., H.K., and, H.H. carried out the synthetic design and synthesis of SS intermediates of BPIns-A. H.K. synthesized HIns-A and HIns-B peptides. S.A. synthesized Rlx-A and Rlx-B peptides. M.O. and K.I. prepared PDI and designed the enzymatic NCA reaction. L.M. contributed in interpretation of experimental data. H.H. and M.I. jointly supervised this work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Arai, K., Takei, T., Shinozaki, R. et al. Characterization and optimization of two-chain folding pathways of insulin via native chain assembly. Commun Chem 1, 26 (2018). https://doi.org/10.1038/s42004-018-0024-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42004-018-0024-0
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.