Abstract
G-protein-coupled receptors (GPCRs) are pivotal cell membrane proteins that sense extracellular molecules and activate cellular responses. The G-protein α subunit i (Gαi) family represents the most common GPCR-coupling partner and consists of eight subunits with distinct signaling properties. However, analyzing the coupling pattern has been challenging owing to endogenous expression of the Gαi subunits in virtually all cell lines. Here, we generate a HEK293 cell line lacking all Gαi subunits, which enables the measurement of GPCR-Gαi coupling upon transient re-expression of a specific Gαi subunit. We profile Gαi-coupling selectivity across 11 GPCRs by measuring ligand-induced inhibitory activity for cAMP accumulation. The coupling profiles are then classified into three clusters, representing those preferentially coupled to Gαz, those to Gαo, and those with unapparent selectivity. These results indicate that individual Gαi-coupled GPCRs fine-tune Gαi signaling by exerting coupling preference at the Gαi-subunit level.
Similar content being viewed by others
Introduction
G-protein-coupled receptors (GPCRs), the key transducers that link extracellular stimuli to downstream intracellular signals, are the most common targets for drug development1. Upon GPCR ligand binding, the heteromeric guanine nucleotide-binding proteins (G proteins), consisting of α, β, and γ subunits, induce activation of their effector proteins. Among the three subunits, the α subunit mainly determines the key properties of individual heterotrimeric G proteins. The human genome encodes 16 α subunits, which are grouped into four subfamilies based on their sequence homology and functional similarity, namely, Gαs, Gαi, Gαq, and Gα122. The Gαi subfamily is the most common coupling partner in the family A GPCRs3,4. Furthermore, approximately 45% of GPCRs targeted by the United States Food and Drug Administration (FDA)-approved drugs couple to the Gαi subfamily5, highlighting the significance of Gαi-coupled GPCRs as drug targets.
Previous studies have shown that the Gαi subunits have both overlapping and distinct functions. The Gαi subfamily consists of eight α subunits: Gαi1, Gαi2, Gαi3, Gαt1, Gαt2, Gαt3, Gαo, and Gαz. Activation of the Gi heterotrimer induces various cellular signaling processes such as the inhibition of adenylyl cyclase and calcium channels, and the activation of potassium channels6. Despite the high sequence homology within the Gαi subfamily, several subunit-specific functions have been reported. For example, transducin (Gαt1 and Gαt2) and gustducin (Gαt3) are involved in visual and taste perception, respectively, both by activating cGMP phosphodiesterase7,8. Additionally, using Gαi subunit knockdown experiments in GH4C1 cells, Gαo and Gαi2 were found to mediate inhibition of calcium entry and cAMP accumulation, respectively9. The Gαi subunits also behave biochemically different from one another. It should be noted, however, that these biochemical properties could be modified in living cells as observed in Gq, which shows slower nucleotide dissociation in purified condition than in lipid membrane condition. For example, the rate of spontaneous GDP dissociation from Gαo is an order of magnitude faster than that of Gαi1-310, and the intrinsic GTPase activity of Gαz is 200-fold slower than that of the other Gαi subunits11. These differences between the members of the Gαi family allow for diverse Gαi-coupled GPCR signal transduction.
Although signaling properties of the Gαi subunits have been well characterized, Gαi-subunit-coupling selectivity of GPCRs has only been investigated for a limited number of receptors due to experimental difficulties. Individual GPCR-Gαi coupling cannot be assessed simply by expressing a Gαi subunit of interest because virtually all mammalian cells endogenously express multiple Gαi subunits12. To eliminate endogenous GPCR-Gαi coupling, previous studies have used pertussis toxin (PTX), which ADP-ribosylates the cysteine residue in the C-terminal tail of the Gαi subunits13. By expressing a PTX-resistant Gαi mutant with a substitution at the cysteine residue, PTX treatment enables the measurement of selective coupling of the mutant Gαi subunit14,15,16. However, altering the cysteine residue can affect the G-protein coupling abilities of the α subunits17,18. Additionally, PTX does not inhibit Gαz since it has isoleucine at the PTX-targeting site19. Recently, fluorescence or bioluminescence resonance energy transfer-based techniques have been used to evaluate coupling of individual engineered Gα subunits20,21. However, these methods require the insertion of luciferase or a fluorescent protein into the α subunit, and such modification affects its biochemical properties and resulting coupling efficiency22.
In this study, we circumvented the experimental difficulties in measuring individual intact Gαi subunits by employing genome-editing approaches to establish a Gαi-null background in human embryonic kidney (HEK) 293A cells. By transiently expressing a pair of a GPCR and a Gαi subunit of interest into the Gαi-deficient HEK293A cells, we successfully characterized Gαi-coupling profiles in 11 GPCRs and found physiologically reasonable selectivity of the Gαi subunits for these GPCRs.
Results
Generation of a HEK293A cell line lacking Gαi subunits
In order to establish a Gαi-null background for functional analysis of Gαi subunit-coupling patterns, we generated a HEK293A cell line devoid of all Gαi subunits. First, we measured expression of the Gαi subunits in HEK293A cells by quantitative RT-PCR analysis. While seven Gαi subunit-encoding genes (GNAI1, GNAI2, GNAI3, GNAO1, GNAT1, GNAT2, and GNAZ) were expressed, GNAT3 expression was undetectable (Fig. 1a). To validate the PCR primer pair for GNAT3, we measured and confirmed GNAT3 expression in a sample of small intestine tissue, in which GNAT3 is known to be expressed23 (Supplementary Fig. 1). We next targeted the seven Gαi subunit-encoding genes present in HEK293A cells using three-round CRISPR–Cas9 mutagenesis (Fig. 1b). We designed single-guide RNAs (sgRNAs) to induce double-stranded DNA cleavage within the N-terminal half of each α-subunit. Upon successful introduction of frameshift or nonsense mutations by these sgRNAs, mutated genes are expected to produce truncated Gαi subunits with the elimination of the c-terminal α5 helix, which is responsible for receptor coupling24. After transfection of the sgRNA constructs, fluorescence-activated cell sorting isolation of GFP-positive cells, and expansion of the clonal cell colonies, the clones were screened for mutations in the target genes using the restriction-enzyme method, obtaining a knock-out clone candidate (ΔGαi cells) (Fig. 1c). Sanger sequencing of the target genomic regions confirmed that the ΔGαi cells carry frameshift mutations or large insertions, including stop codons, in all targeted alleles (Supplementary Fig. 2). GNAT3 expression remained undetected in the ΔGαi cells (Supplementary Fig. 1). These results indicate that the seven Gαi subunit-encoding genes were successfully mutated in the HEK293A cells to prevent their endogenous expression.
a Quantitative real-time PCR analysis of the Gαi subunits in the parent HEK293A cells. Bars and error bars represent the mean and standard error of the mean (SEM), respectively, of three independent experiments. Each experiment was performed in duplicate. Abbreviation: N.D., not detected. b Strategy for genetic inactivation of the Gαi family members. ∆Gαi cells were obtained using three-round CRISPR–Cas9 mutagenesis. c Identification of the Gαi-mutant clone genotype using the restriction-enzyme digested-fragment method. The sgRNA targets were PCR-amplified and treated with their corresponding restriction enzymes (RE). Digested samples were subjected to capillary electrograph analysis, and pseudo-gel images of the electropherogram were visualized with a DNA marker of the pUC19/Hpa II digest. Black and red arrowheads indicate undigested and RE-digested PCR fragments of the targeted site, respectively. Note that there are two RE sites in PCR fragments of the GNAI2 and GNAT1 genes, and one of the mutated alleles in the GNAI1 gene could be digested by RE. Abbreviations: P parental, KO knock-out.
Next, we functionally examined the lack of Gαi subunits in ΔGαi cells using a GloSensor cAMP assay. We expressed µ-opioid receptor (MOR), a prototypical Gαi-coupled receptor, along with the GloSensor cAMP reporter and measured cAMP levels upon stimulation with the ligand, methionine-enkephalin (MetEnk), together with forskolin. In the parent cells, forskolin-stimulated cAMP accumulation was almost completely inhibited in a MetEnk concentration-dependent manner (Fig. 2a). In contrast, the ∆Gαi cells were completely unresponsive to MetEnk, while exogenous Gαi2 expression restored their response to the ligand (Fig. 2b). These results show that the ΔGαi cells are devoid of functional Gαi subunits, making them suitable for investigating Gαi subunit signaling.
a, b Concentration–response curves of the GloSensor cAMP assay in parent (a) and ∆Gαi cells (b). Parent and ∆Gαi cells transiently expressing Glo-22F cAMP reporter with MOR and Gαi2 were stimulated with 10 µM forskolin along with indicated concentrations of MetEnk. For each experiment, forskolin-induced cAMP accumulation was set as 100%. In both figures, symbols and error bars represent the mean and SEM, respectively, of three independent experiments. Each experiment was performed in triplicate. Note that for many data points, error bars are smaller than the size of the symbols, and thus are not visible.
To further characterize the ∆Gαi cells, we compared cell proliferation of the parent and ∆Gαi cells. We assessed cell proliferation by collecting the cells 48 h after seeding and counting the number of cells. Cell proliferation of the ∆Gαi cells in 48 h was reduced by 20% compared to that of parent cells (Supplementary Fig. 3). Similar to the ∆Gαi cells, PTX-treated parent cells also showed a 20% reduction in cell proliferation compared to that of non-treated parent cells. This result suggests that the suppression of cell proliferation in the ∆Gαi cells is not due to a clonal effect but to the on-target deficiency of Gαi proteins. In addition, the reduced proliferation in ∆Gαi cells and the PTX-treated parent cells were both recovered by culturing the cells in type I collagen-coated dishes (Supplementary Fig. 3), suggesting that the suppression of cell proliferation is due to the weakened adhesion of the ∆Gαi cells.
To examine whether the other G-protein signaling would be affected in the ΔGαi cells, Gs, Gq, and G12 signaling were individually evaluated. We expressed Gs-coupled vasopressin V2 receptor (V2R) or dopamine D1 receptor (D1R) along with the GloSensor cAMP reporter in both the ΔGαi cells and parent cells. The cells were then stimulated with their agonists or forskolin, and measured for their cAMP accumulation. Gs signaling in the ΔGαi cells was enhanced compared to that in the parent cells (Fig. 3a, b). We next measured Gq and G12 signaling via transforming growth factor (TGF) α shedding assay25 using Gq- (H1R and α1A) or G12- (EP3 and CB1) coupled GPCRs. By using G-protein-deficient cells, we have previously validated that TGFα shedding responses induced by the tested Gq-GPCR (H1R and α1A) and the tested G12-GPCR (CB1 and EP3) were solely dependent on Gq/11 and G12/13, respectively3. Each GPCR was expressed together with an alkaline phosphatase-tagged TGFα (AP-TGFα) reporter in the ∆Gαi cells and the parent cells, and subsequently measured for their ligand-induced AP-TGFα ectodomain shedding response. Gq and G12 signaling in the ΔGαi cells were comparable to that in parent cells (Fig. 3c–f). These data demonstrate that the elimination of the Gαi subunits does not affect Gq and G12 signaling but does potentiate Gs signaling, possibly due to the loss of competition in adenylyl cyclases between Gs and Gi.
a, b Concentration-response curves of Gs-mediated cAMP accumulation. Parent and ∆Gαi cells transiently expressing Glo-22F together with either D1R (a) or V2R (b) were stimulated with the indicated concentrations of dopamine or arginine vasopressin (AVP), respectively. c–f Concentration–response curve of the TGFα shedding responses induced by Gq- (H1R (c) and α1A (d)) and G12-coupled receptor (CB1 (e) and EP3 (f)) activation in parent and the ∆Gαi cells. A test GPCR was expressed in the parent and the ∆Gαi cells along with the AP-TGFα reporter, and the resulting ligand-induced response was assessed. In all figures, symbols and error bars represent the mean and SEM, respectively, of three independent experiments. Each experiment was performed in triplicate. Note that for many data points, error bars are smaller than the size of the symbols, and thus are not visible.
We additionally examined whether Gi-coupled GPCRs gain access to other Gα proteins in the absence of Gαi proteins using MOR as an example. As shown in Fig. 2b, MOR activation did not increase cAMP in the ∆Gαi cells, showing that it does not couple to Gs even in the absence of Gαi. We further examined whether MOR couples to Gq and G12 in the Gαi-deficient condition using the TGFα shedding assay. We found that MOR showed no detectable TGFα shedding response in both parent and the ∆Gαi cells (Supplementary Fig. 4), indicating that MOR fails to couple with Gq and G12 even in the absence of Gαi proteins. Of note, we validated activation of MOR in the TGFα shedding assay by using the Gαq/i1 chimeric Gα protein, which binds to Gi-coupled GPCRs and transduces Gq signaling, as a positive control (Supplementary Fig. 4). While these data support that MOR does not couple to non-preferred Gα even in the absence of preferred Gα, we cannot exclude the possibility that other Gi-coupled GPCRs will behave differently. Thus it is recommended to consider this possibility when analyzing G-protein coupling using the Gα-deficient cells.
Divergent Gαi subunit-coupling preferences across GPCRs
We evaluated GPCR-Gαi-coupling profiles by expressing a pair of GPCR and Gαi subunit of interest in the ∆Gαi cells. We chose five Gαi subunits, namely, Gαi1, Gαi2, Gαi3, Gαo, and Gαz, out of the eight Gαi subunits for the following characterization since physiological effector proteins for the rest of the Gαi subunits (Gαt1, Gαt2, and Gαt3) are known to be cGMP phosphodiesterase.
To determine suitable amounts of the plasmids in the assay, we tested multiple amounts of plasmids encoding GPCR or Gαi subunits and compared concentration-response curves. First, we measured membrane expression levels of MOR, a representative GPCR used in the study, by titrating the MOR-encoding plasmid (100 ng Gαi1 plasmid for all of the MOR conditions). From a plasmid dose of 8–200 ng (per well in a 6-well plate), expression levels increased and its level did not further elevated in the 1000 ng condition (Supplementary Fig. 5a, b). We then measured Gi-mediated cAMP response by the GloSensor assay. The concentration–response curve shifted to the left as the plasmid amount was increased, and plateaued at 200 ng volume (Supplementary Fig. 5c). Therefore, we used 200 ng of the plasmid encoding test GPCRs in the subsequent experiments. In the case of Gα, we fixed the MOR-encoding plasmid as 200 ng and titrated the plasmid amounts of all of the five Gαi subunits from 1 to 1000 ng by 10-fold. A plasmid dose of 1 ng showed negligible cAMP-suppressive effect, while a plasmid dose of 10 ng partially inhibited cAMP production (Supplementary Fig. 5d). Plasmid doses of 100 and 1000 ng almost completely inhibited cAMP production. We note that, from the plasmid volumes of 10 to 1000 ng, the concentration-response curve tended to shift to the right as the plasmid dose increased. Together, subsequent experiments were performed with a Gαi-encoding plasmid dose of 100 ng.
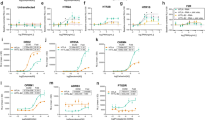
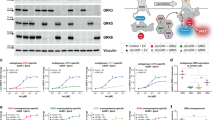
Because the GloSensor cAMP assay dominantly detects the activation of Gαs over Gαi, we selected 11 GPCRs that are coupled to Gαi but not to Gαs, which include dopamine D2 receptor (D2R), dopamine D4 receptor (D4R), mu-type opioid receptor (MOR), somatostatin receptor type 2 (SSTR2), cannabinoid receptor 1 (CB1), lysophosphatidic acid receptor 1 (LPA1), sphingosine 1-phosphate receptor 1 (S1P1), sphingosine 1-phosphate receptor 3 (S1P3), probable G-protein coupled receptor 34 (GPR34), C–C chemokine receptor type 2 (CCR2), and C-C chemokine receptor type 5 (CCR5). Endogenous ligands including dopamine, serotonin, methionine-enkephalin, somatostatin, lysophosphatidic acid, sphingosine 1-phosphate, lysophosphatidylserine, CCL2, and CCL4 were used for all respective GPCRs except for CB1, for which a high-affinity synthetic ligand (CP-55490) was used because endogenous ligands such as anandamide exhibited poor responses in the assay. For each GPCR, we plotted the cAMP responses from the five Gαi conditions and one Gαi-nontransfected condition (Fig. 4a, Supplementary Fig. 6). The inhibition of cAMP accumulation over titrated ligand concentrations was graphed and fitted with a sigmoidal concentration-response curve, from which the half maximal concentration (EC50) and maximal response (Emax) values were obtained (Supplementary Table 1). We found divergent patterns of Gαi-subunit coupling among the 11 GPCRs. In D2R, for example, all tested Gαi subunits resulted in equivalent saturating concentrations (corresponding to Emax values), but the concentration–response curve in the Gαo-expressing condition shifted leftward, which reflects the difference in EC50 values. The observed Gαo preference of D2R is consistent with previous studies using Sf9 cells and intact Gαi subunits which showed that D2R most effectively couples to Gαo26,27. In LPA1, on the other hand, the cAMP-suppressive responses in saturating concentrations were comparable between Gαi1–3 and Gαo, while the response was smaller in Gαz.
a Concentration–response curves of GloSensor cAMP assays in the ∆Gαi cells transiently expressing Glo-22F with certain combinations of GPCR and Gαi subunits. Cells were stimulated with 10 µM forskolin along with the indicated concentrations of ligands. For each experiment, forskolin-induced cAMP accumulation was set as 100%. In all figures, symbols and error bars represent the mean and SEM, respectively, of three independent experiments. Each experiment was performed in triplicate. Note that for some data points, error bars are smaller than the size of symbols. b Schematic diagram showing the process for calculating RAi values. c Heatmap of the LogRAi values for the 11 GPCRs. Colors range from white (LogRAi = −1.5) to red (LogRAi = 0). Receptors are arranged according to the dendrogram of the hierarchical clustering, which was calculated from the coupling profiles.
To compare Gαi-subunit-coupling selectivity across receptors in a uniform way, we calculated a parameter known as relative intrinsic activity (RAi)28. RAi value is a single parameter that considers both EC50 and Emax. While RAi is commonly used to evaluate ligand bias, we used it to assess receptor’s preference for Gαi subunits. For each sigmoidal curve, we divided Emax by EC50 and normalized the resulting Emax/EC50 values by the maximum Emax/EC50 value among the five tested Gαi subunits to generate dimensionless relative Emax/EC50 values (defined as RAi, Fig. 4b). We further log-transformed the RAi value (LogRAi) and used it as a Gαi-subunit-coupling index. We applied hierarchical cluster analysis to classify the GPCRs and showed their Gαi-subunit-coupling index as a heatmap with a clustering tree (Fig. 4c). We observed three groups in the clustering analysis. Cluster 1 (S1P1, CCR2, S1P3, D4R, and MOR) coupled efficiently to Gαz, followed by Gαi1–3 and Gαo. Cluster 2 (D2R) coupled to Gαo, followed by Gαi1–3, then Gαz. Cluster 3 (CB1, CCR5, GPR34, SSTR2, and LPA1) did not show selectivity for specific Gαi subunits. We compared the Gαi-coupling-based clustering with GPCR phylogenetic trees. Based on the amino-acid similarity and ligand grouping of GPCRdb (https://gpcrdb.org), the tested receptors were classified into lipid receptors (LPA1, S1P1, S1P3, CB1), orphan receptors (GPR34), aminergic receptors (D2R, D4R), chemokine receptors (CCR2 and CCR5), and peptide receptors (MOR, SSTR2). Interestingly, this classification is distinct from coupling-based clustering. These results suggest that, from an evolutionary perspective, each GPCR has individually acquired varying Gαi subunit selectivity29.
Discussion
In this study, we established a Gαi-deficient HEK293A cell line, ∆Gαi cells, in order to more effectively study the Gαi subunit-coupling preference of GPCRs. Researchers have previously used PTX to inhibit endogenously expressed Gαi family proteins. However, PTX-insensitive mutants are required to assess individual Gαi subunit signaling in PTX-treated conditions. Furthermore, PTX is incapable of inhibiting Gαz. An Escherichia coli pertussis toxin-like AB5 toxin called OZITX was recently found to inhibit all Gαi subfamily proteins, including Gαz30,31. Even with this toxin, a Gαi C-terminal mutation, which may affect coupling properties, is required to make Gαi subfamily proteins insensitive to OZITX. The ∆Gαi cells overcome these experimental problems and allow for investigation of not only GPCR-Gαi subunit coupling but also downstream signaling of intact, individual Gαi subunits.
The coupling preference of D2R for Gαo as well as D4R for Gαz provides signaling properties suitable for physiological functions. Gαz has a slow spontaneous GDP dissociation rate as well as low intrinsic GTPase activity11. D4R is expressed in the retina and regulates the circadian nature of light-adapted vision32,33. Both D4R and Gαz are expressed in photoreceptor cells and display a daily expression pattern that peaks at night34. Efficient coupling of D4R and Gαz may allow for regulation of the circadian nature of vision by taking advantage of the long-term activation properties of Gαz. Although D2R is phylogenetically the most similar to D4R and together comprise D2-like receptors, D2R is efficiently coupled to Gαo. In contrast to Gαz, Gαo has a faster spontaneous GDP dissociation rate than the other Gαi subunits10. Therefore, it is likely that D2R possesses fast-responding signaling properties through Gαo, which may allow for rapid signal transduction between neurons. Altogether, D2R and D4R likely utilize the differential coupling selectivity to allow for appropriate duration of dopaminergic signaling outcomes. Since signaling properties of GPCRs are also dependent on Gα expression profiles of the cells and their localization, which are different among cell types, signaling properties of D2R and D4R in vivo need to be further investigated in the future studies.
MOR is an important drug target for analgesics. In a previous study using PTX and PTX-insensitive Gαi subunits, MOR was coupled more effectively to Gαi3 and Gαo than to Gαi1 and Gαi214. However, in our experimental conditions, MOR couples almost comparably to Gαi1-3 and Gαo. Although difference in the amount of Gα proteins between the two studies could also affect coupling, this observed difference may be due to the use of the C-terminal mutation in the Gαi subunits in the previous study in order to confer resistance to PTX, suggesting the importance of evaluating Gαi subunit selectivity using intact Gαi subunits. Additionally, we measured the coupling of MOR for Gαz in parallel with Gαi1-3 and Gαo without modifications in the α subunits. In vivo experiments have revealed differential roles of Gαi family members in morphine-induced analgesic effects: Gαz in chronic administration35,36 and Gαi2 and Gαo in acute analgesic action37,38. In consideration of biased ligands, which induce distinct effector activation on the same receptor39,40,41, a Gαz-biased MOR agonist could potentially lead to the development of a long-acting analgesia with possibly lower tolerance. Altogether, evaluating the Gαi subunit selectivity of MOR agonizts may ultimately enhance our understanding of the differences in drug activity and contribute to the discovery of more desirable analgesics.
Along with our previously generated ∆Gαs, ∆Gαq, and ∆Gα12 cells42,43,44, we now have a full set of the four Gα knock-out cell lines. These cell lines can be used to investigate Gα signaling downstream of GPCRs individually, which will increase our understanding of GPCR signaling45 and ultimately allow for the development of drugs with more optimal therapeutic effects.
Methods
Reagents and plasmids
MetEnk (4042-v) and somatostatin (4023-v) were purchased from Peptide Institute. S1P (62570) and CP-55490 (90084) were purchased from Cayman Chemical. Dopamine (040-15433) was purchased from FUJIFILM Wako Pure Chemical. LPA (867130) and LysoPS (858143) were purchased from Avanti Polar Lipid. CCL2 (Z03292) and CCL4 (Z02831) were purchased from Genscript. Plasmids used in Glosensor cAMP assay3 and TGFα shedding assay25 were previously described.
Cell culture and transfection
HEK293A cells (Thermo Fisher Scientific) and their derivative ΔGαi cells were maintained in Dulbecco’s Modified Eagle Medium (DMEM, Nissui Pharmaceutical) supplemented with 10% fetal bovine serum (Gibco, Thermo Fisher Scientific), 0.006% (w/v) penicillin and 0.01% (w/v) streptomycin (complete DMEM), and maintained at 37 °C in humidified incubator equilibrated with 5% CO2. Transfections were performed by using polyethylenimine (PEI) solution (Polyethyleneimine “Max”, Polysciences). Typically, HEK293 cells were seeded in a 6-well culture plate at cell density of 2 × 105 cells per mL in 2 mL of the complete DMEM and cultured for one day in a CO2 incubator. A transfection solution was mixed by combining plasmid solution diluted in 100 µl of Opti-MEM (Life technologies) and 4 µl of 1 mg per mL PEI solution in 100 µl of Opti-MEM. The transfected cells were further incubated for one day before subjected to an assay as described below.
Generation of ∆Gαi-HEK293 cells
The ΔGαi cells were obtained by the iterative CRISPR/Cas9-mediated mutagenesis strategy illustrated in Fig. 1B. sgRNA constructs targeting the GNAI1, GNAI2, GNAI3, GNAO, GNAZ, GNAT1, and GNAT2 genes were designed using the CRISPR.MIT.EDU website (already shut down) so that a SpCas9-mediated DNA cleavage site (3-bp upstream of the protospacer adjacent motif (PAM) sequence) encompasses a restriction enzyme-recognizing site. Sense and antisense oligonucleotide encoding the guide RNA was synthesized (FASMAC) and inserted into the BbsI site of the pSpCas9 (BB)-2A-GFP (PX458) vector (a gift from Feng Zhang, Broad Institute, Cambridge, MA; Addgene plasmid No.48138). Correct insertion of the guide RNA sequences was verified by sequencing using the Sanger method (FASMAC). HEK293A cells were seeded into 6-well plates (1.0 × 105 cells per mL in 2 mL of complete DMEM) and 24 hours later, transfected with a combination of PX458 vectors targeting the Gαi subunit encoding genes using Lipofectamine 2000 (Thermo Fisher Scientific). Three days later, the cells were harvested and top 10–20% of GFP-positive cells were isolated by fluorescence-activated cell sorter (SH800; SONY). The isolated cells were distributed in 96-well plates for clonal expansion. Clones were analyzed for mutations in target genes by PCR and restriction enzyme digestion using capillary electrophoresis system (MultiNA, Shimadzu). The restriction enzyme-resistant PCR products were assessed for their genomic DNA alterations by direct sequencing or TA cloning. Successful targeting was confirmed by assessing GPCR-mediated cAMP inhibition, as described below. Oligonucleotide sequences encoding guide RNAs, PCR primers used to amplify the sgRNA-targeting sites, and the restriction enzymes used to digest PCR products were listed in Supplementary Table 2.
Quantitative real-time PCR analysis
Total RNA from HEK293 cells was prepared using a GenElute Mammalian Total RNA Miniprep kit (Sigma-Aldrich). Total RNA was reverse-transcribed using High-Capacity cDNA RT kits (Applied Biosystems) according to the manufacturer’s instruction. Real-time quantitative PCRs were performed with TB Green Premix Ex Taq II (Tli RNaseH Plus) (Takara Bio) and monitored by ABI Prism 7300 (Applied Biosystems). RNA expression data were normalized to the expression of GAPDH. The PCR primer sequences were listed in Supplementary Table 3.
GloSensor cAMP assay
Plasmid transfection was performed in a 6-well plate with a mixture of 1 µg (per well in a 6-well plate) Glo-22F cAMP biosensor-encoding pCAGGS plasmid (gene synthesized with codon optimization by Genscript), 200 ng of GPCR-encoding plasmid and 100 ng of Gαi subunit-encoding plasmid. After 1-day incubation, the transfected cells were harvested with 0.53 mM EDTA-containing D-PBS, centrifuged at 190 g for 5 min and suspended in 0.01% bovine serum albumin (BSA, fatty acid-free and protease-free grade, Serva) and 5 mM HEPES (pH 7.4) containing Hank’s Balanced Salt Solution (HBSS). The cells were seeded in a white 96-well plate (30 µl per well) and loaded with D-luciferin potassium solution (10 µl of 8 mM solution per well; FujiFilm Wako Pure Chemical). After 2 hour incubation in the dark at room temperature, the plate was read for its initial luminescent count (Spectramax L, Molecular Devices). The cells were treated with a ligand alone (Gs assay) or 10 µM forskolin (FujiFilm Wako Pure Chemical) together with a ligand (Gi assay) (10 µl of 5× solution per well). For kinetics measurement, the plates were measured for 20 min and expressed as fold change values. Fold-change luminescent signals from 16 min to 20 min after compound addition were averaged and normalized to those in forskolin-treated condition. Using Prism 9 software (GraphPad Prism), the cAMP signals were fitted to a four-parameter sigmoidal concentration–response curve, from which EC50 and Emax values were obtained. Hierarchical clustering of the coupling profile was performed in python using the Ward method.
TGFα shedding assay
The TGFα shedding assay was performed as described previously with minor modifications25. Plasmid transfection was performed in a 6-well plate with a mixture of 500 ng (per well in a 6-well plate) AP-TGFα-encoding plasmid and 200 ng GPCR-encoding plasmid. After 1-day culture, the transfected cells were harvested by trypsinization, pelleted by centrifugation at 190 g for 5 min and washed once with 5 mM HEPES (pH 7.4)-containing HBSS. After centrifugation, the cells were resuspended in 6 mL of the HEPES-containing HBSS. The cell suspension was seeded in a 96-well culture plate (cell plate) at a volume of 90 µl (per well hereafter) and incubated for 30 min in a CO2 incubator. The cells were treated with a GPCR ligand (10×, diluted in HBSS containing 5 mM HEPES (pH 7.4) and 0.01% (w/v) BSA. After spinning the cell plates, 80 µl of conditioned media was transferred to an empty 96-well plate (conditioned media (CM) plate). Totally, 80 µl of AP reaction solution (10 mM p-nitrophenylphosphate (p-NPP), 120 mM Tris-HCl (pH 9.5), 40 mM NaCl and 10 mM MgCl2) was dispensed into the cell plates and the CM plates. Absorbance at 405 nm of the plates was measured, using a microplate reader (SpectraMax 340 PC384, Molecular Devices), before and after 1 hr incubation at room temperature. Ligand-induced AP-TGFα release was calculated by subtracting spontaneous AP-TGFα release signal from ligand-induced AP-TGFα release signal.
Flow cytometry
Flow cytometry analysis was performed as previously described3. Plasmid transfection was performed in a 6-well plate with 1 µg (per well in a 6-well plate) of Glo-22F, 100 ng of Gαi1, and varying amount of plasmid encoding MOR harboring the N-terminal hemagglutinin signal sequence, followed by the FLAG epitope tag and a 15-amino-acid flexible linker. The transfected cells were harvested by adding 300 µl of 0.53 mM EDTA-containing D-PBS, followed by 300 µl of 5 mM HEPES (pH 7.4)-containing HBSS. The cell suspension was dispensed in a 96-well V-bottom plate (200 µl per well, two wells per sample). After centrifugation at 700g for 1 min, the cells were washed once with D-PBS and pelleted. Cell pellets were suspended in 2% goat serum- and 2 mM EDTA-containing D-PBS (blocking buffer; 100 µl per well) and incubated for 30 min on ice. After centrifugation at 700g for 1 min, the cells were stained with anti-FLAG epitope tag monoclonal antibody (Clone 1E6, FujiFilm Wako Pure Chemicals; 10 µg/mL in the blocking buffer; 50 µl per well) for 30 min on ice. After rinse with D-PBS, the cells were labeled with a goat anti-mouse IgG secondary antibody conjugated with Alexa Fluor 488 (Thermo Fisher Scientific; 10 µg/mL dilution in the blocking buffer; 25 µl per well) for 15 min on ice. The cells were washed once with D-PBS, resuspended in 100 µl of 2 mM EDTA-containing-D-PBS and filtered through a 40 µm filter. The fluorescently labeled cells (approximately 20,000 cells per sample) were analyzed by the EC800 flow cytometer (Sony). Fluorescent signal derived from Alexa Fluor 488 was recorded in an FL1 channel and flow cytometry data were analyzed by a FlowJo software (FlowJo). We used all of the recorded fluorescent signals and calculated mean fluorescence intensities (MFI).
NanoBiT-miniG recruitment assay
Plasmid transfection was performed in a 6-well plate with a mixture of 500 ng NES-LgBiT-miniGi1-encoding plasmid, whose linker length between LgBiT and miniGi1 was extended to 15-amino acids from the original construct46,47, and varying amount of C-terminally SmBiT-fused MOR plasmid (with the N-terminal hemagglutinin signal sequence, followed the FLAG epitope tag and a 15-amino-acid flexible linker). After 1-day culture, the transfected cells were harvested with 1 mL of 0.53 mM EDTA-containing D-PBS, followed by addition of 2 mL the HEPES-containing HBSS. The cells were centrifuged at 190 g for 5 min and resuspended in 2 mL of the 0.01% BSA- and 5 mM HEPES (pH 7.4)-containing HBSS (assay buffer). The cell suspension was seeded in a 96-well culture white plate (Greiner Bio-One) at a volume of 80 µl (per well hereafter) and loaded with 20 µl of 50 µM coelenterazine (Carbosynth) solution diluted in the assay buffer. After 2-hour incubation with coelenterazine at room temperature, MetEnk (6 µM, diluted in the assay buffer) was manually added to the cells (20 µl). Luminescent signals were measured every 40 seconds after ligand addition and average luminescent values of 10–15 min were plotted.
Statistics and reproducibility
For GloSensor cAMP assay and TGFα shedding assay, three independent experiments were performed in duplicate and triplicate, respectively. For quantitative RT-PCR analysis, three independent experiments were performed in duplicate. Symbols and error bars represent mean values and SEM of indicated numbers of independent experiments, respectively. Concentration–response curves were fitted to all data by the Nonlinear Regression: Variable slope (four parameter) in the Prism 9 with a constraint of the Hill Slope of absolute value less than two. For multiple comparison analysis, we tested statistical significance by two-way ANOVA, followed by the indicated post hoc test using Prism 9.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data generated or analyzed in study are provided in the Supplementary Data 1.
References
Hauser, A. S., Attwood, M. M., Rask-Andersen, M., Schiöth, H. B. & Gloriam, D. E. Trends in GPCR drug discovery: new agents, targets and indications. Nat. Rev. Drug Discov. 16, 829–842 (2017).
Hann, V. & Chazot, P. L. G-proteins. Curr. Anaesth. Crit. Care 15, 79–81 (2004).
Inoue, A. et al. Illuminating G-protein-coupling selectivity of GPCRs. Cell 177, 1933–1947.e25 (2019).
Avet, C. et al. Effector membrane translocation biosensors reveal G protein and βarrestin coupling profiles of 100 therapeutically relevant GPCRs. Elife 11, 1–34 (2022).
Sriram, K. & Insel, P. A. GPCRs as targets for approved drugs: How many targets and how many drugs? Mol. Pharmacol. Mol. 117, 111062 (2018).
Wettschureck, N. & Offermanns, S. Mammalian G proteins and their cell type specific functions. Physiol. Rev. 85, 1159–1204 (2005).
Arshavsky, V. Y., Lamb, T. D. & Pugh, E. N. G proteins and phototransduction. Annu. Rev. Physiol. 64, 153–187 (2002).
Rosenzweig, S., Yan, W., Dasso, M. & Spielman, A. I. Possible novel mechanism for bitter taste mediated through cGMP. J. Neurophysiol. 81, 1661–1665 (1999).
Liu, Y. F., Jakobs, K. H., Rasenick, M. M. & Albert, P. R. G protein specificity in receptor-effector coupling. Analysis of the roles of G(o) and G(i)2 in GH4C1 pituitary cells. J. Biol. Chem. 269, 13880–13886 (1994).
Linder, M. E., Ewald, D. A., Miller, R. J. & Gilman, A. G. Purification and characterization of G(oα) and three types of G(iα) after expression in Escherichia coli. J. Biol. Chem. 265, 8243–8251 (1990).
Casey, P. J., Fong, H. K. W., Simon, M. I. & Gilman, A. G. G(z), a guanine nucleotide-binding protein with unique biochemical properties. J. Biol. Chem. 265, 2383–2390 (1990).
Atwood, B. K., Lopez, J., Wager-Miller, J., Mackie, K. & Straiker, A. Expression of G protein-coupled receptors and related proteins in HEK293, AtT20, BV2, and N18 cell lines as revealed by microarray analysis. BMC Genomics 12, 14 (2011).
Katada, T. The inhibitory G protein Gi identified as pertussis toxin-catalyzed ADP-ribosylation. Biol. Pharm. Bull. 35, 2103–2111 (2012).
Clark, M. J., Furman, C. A., Gilson, T. D. & Traynor, J. R. Comparison of the relative efficacy and potency of mu-opioid agonists to activate Galpha(i/o) proteins containing a pertussis toxin-insensitive mutation. J. Pharm. Exp. Ther. 317, 858 (2006).
Leaney, J. L. & Tinker, A. The role of members of the pertussis toxin-sensitive family of G proteins in coupling receptors to the activation of the G protein-gated inwardly rectifying potassium channel. Proc. Natl Acad. Sci. USA 97, 5651–5656 (2000).
Anavi-Goffer, S. et al. Helix 8 Leu in the CB1cannabinoid receptor contributes to selective signal transduction mechanisms. J. Biol. Chem. 282, 25100–25113 (2007).
Moon, H. E., Bahia, D. S., Cavalli, A., Hoffmann, M. & Milligan, G. Control of the efficiency of agonist-induced information transfer and stability of the ternary complex containing the δ opioid receptor and the α subunit of Gi1 by mutation of a receptor/G protein contact interface. Neuropharmacology 41, 321–330 (2001).
Bahia, D. S. et al. Hydrophobicity of residue351 of the G protein G(il)α determines the extent of activation by the α(2A)-adrenoceptor. Biochemistry 37, 11555–11562 (1998).
Wong, Y. H., Conklin, B. R. & Bourne, H. R. Gz-mediated hormonal inhibition of cyclic AMP accumulation. Science 255, 339–342 (1992).
Olsen, R. H. J. et al. TRUPATH, an open-source biosensor platform for interrogating the GPCR transducerome. Nat. Chem. Biol. 16, 841–849 (2020).
Polit, A., Rysiewicz, B., Mystek, P., Błasiak, E. & Dziedzicka-Wasylewska, M. The Gαi protein subclass selectivity to the dopamine D2 receptor is also decided by their location at the cell membrane. Cell Commun. Signal. 18, 1–16 (2020).
Gibson, S. K. & Gilman, A. G. Giα and Gβ subunits both define selectivity of G protein activation by α2-adrenergic receptors. Proc. Natl Acad. Sci. USA 103, 212–217 (2006).
Jang, H. J. et al. Gut-expressed gustducin and taste receptors regulate secretion of glucagon-like peptide-1. Proc. Natl Acad. Sci. USA 104, 15069–15074 (2007).
Dror, R. O. et al. Structural basis for nucleotide exchange in heterotrimeric G proteins. Science 348, 1361–1365 (2015).
Inoue, A. et al. TGFα shedding assay: an accurate and versatile method for detecting GPCR activation. Nat. Methods 9, 1021–1029 (2012).
Gazi, L., Nickolls, S. A. & Strange, P. G. Functional coupling of the human dopamine D 2 receptor with Gα i1, Gα i2, Gα i3 and Gα o G proteins: evidence for agonist regulation of G protein selectivity. Br. J. Pharm. 138, 775–786 (2003).
Nickolls, S. A. & Strange, P. G. The influence of G protein subtype on agonist action at D2 dopamine receptors. Neuropharmacology 47, 860–872 (2004).
Ehlert, F. J., Griffin, M. T., Sawyer, G. W. & Bailon, R. A simple method for estimation of agonist activity at receptor subtypes: Comparison of native and cloned M3 muscarinic receptors in guinea pig ileum and transfected cells. J. Pharmacol. Exp. Ther. 289, 981–992 (1999).
Flock, T. et al. Selectivity determinants of GPCR-G-protein binding. Nature 545, 317–322 (2017).
Keen, A. C. et al. OZITX, a pertussis toxin-like protein for occluding inhibitory G protein signalling including Gαz. Commun. Biol. 5, 1–10 (2022).
Littler, D. R. et al. Structure-function analyses of a pertussis-like toxin from pathogenic Escherichia coli reveal a distinct mechanism of inhibition of trimeric G-proteins. J. Biol. Chem. 292, 15143–15158 (2017).
Jackson, C. R., Chaurasia, S. S., Hwang, C. K. & Iuvone, P. M. Dopamine D4 receptor activation controls circadian timing of the adenylyl cyclase 1/cyclic AMP signaling system in mouse retina. Eur. J. Neurosci. 34, 57–64 (2011).
Jackson, C. R. et al. Retinal dopamine mediates multiple dimensions of light-adapted vision. J. Neurosci. 32, 9359–9368 (2012).
Vancura, P. et al. Gnaz couples the circadian and dopaminergic system to G protein-mediated signaling in mouse photoreceptors. PLoS ONE 12, 1–17 (2017).
Hendry, I. A. et al. Hypertolerance to morphine in G(zα)-deficient mice. Brain Res. 870, 10–19 (2000).
Leck, K. J. et al. Deletion of guanine nucleotide binding protein αz subunit in mice induces a gene dose dependent tolerance to morphine. Neuropharmacology 46, 836–846 (2004).
Standifer, K. M., Rossi, G. C. & Pasternak, G. W. Differential blockade of opioid analgesia by antisense oligodeoxynucleotides directed against various G protein α subunits. Mol. Pharmacol. 50, 293–298 (1996).
Raffa, R. B., Martinez, R. P. & Connelly, C. D. G-protein antisense oligodeoxyribonucleotides and μ-opioid supraspinal antinociception. Eur. J. Pharmacol. 258, 5–7 (1994).
Namkung, Y. et al. Functional selectivity profiling of the angiotensin II type 1 receptor using pathway-wide BRET signaling sensors. Sci. Signal. 11, 1–21 (2018).
Corbisier, J., Huszagh, A., Galés, C., Parmentier, M. & Springael, J. Y. Partial agonist and biased signaling properties of the synthetic enantiomers J113863/UCB35625 at chemokine receptors CCR2 and CCR5. J. Biol. Chem. 292, 575–584 (2017).
Bonifazi, A. et al. Novel and potent dopamine D2 receptor go-protein biased agonists. ACS Pharmacol. Transl. Sci. 2, 52–65 (2019).
Stallaert, W. et al. Purinergic receptor transactivation by the β2-adrenergic receptor increases intracellular Ca2+ in nonexcitable cells. Mol. Pharmacol. 91, 533–544 (2017).
Devost, D. et al. Conformational profiling of the AT1 angiotensin II receptor reflects biased agonism, G protein coupling, and cellular context. J. Biol. Chem. 292, 5443–5456 (2017).
Schrage, R. et al. The experimental power of FR900359 to study Gq-regulated biological processes. Nat. Commun. 6, 1–7 (2015).
Milligan, G. & Inoue, A. Genome editing provides new insights into receptor-controlled signalling pathways. Trends Pharmacol. Sci. 39, 481–493 (2018).
Qingwen, Wan et al. Mini G protein probes for active G protein–coupled receptors (GPCRs) in live cells. J. Biol. Chem. 293, 7466–7473 (2018).
Rony, N. et al. Mini-G proteins: novel tools for studying GPCRs in their active conformation. Plos ONE 12, e0175642 (2017).
Acknowledgements
We thank Ritsuki Kuramoto at Tohoku University for his assistance in the clustering analysis; Ayaki Saito and other members of the Inoue laboratory for the manuscript editing; Kazuhiro Kobayashi and Osamu Nureki at The University of Tokyo for the miniGi1 construct. A.I. was funded by Japan Society for the Promotion of Science (JSPS) KAKENHI grants 21H04791, 21H05113, JPJSBP120213501, and JPJSBP120218801; FOREST Program JPMJFR215T and JST Moonshot Research and Development Program JPMJMS2023 from Japan Science and Technology Agency (JST); BINDS JP22ama121038 from Japan Agency for Medical Research and Development (AMED); Takeda Science Foundation; Uehara Memorial Foundation. Y.O. received JSPS KAKENHI 20J20669.
Author information
Authors and Affiliations
Contributions
Conceptualization: Y.O. and A.I.; Investigation: Y.O., G.N., K.K., and S.I.; Writing: Y.O., K.K., and A.I. with feedback from all of the coauthors. Funding acquisition: Y.O., J.A., and A.I.; Supervision: J.A. and A.I.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Bryan Roth, Ajith Karunarathne and the other, anonymous, reviewer for their contribution to the peer review of this work. Primary Handling Editor: Joao Valente.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ono, Y., Kawakami, K., Nakamura, G. et al. Generation of Gαi knock-out HEK293 cells illuminates Gαi-coupling diversity of GPCRs. Commun Biol 6, 112 (2023). https://doi.org/10.1038/s42003-023-04465-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-023-04465-2
This article is cited by
-
Identification of Gα12-vs-Gα13-coupling determinants and development of a Gα12/13-coupled designer GPCR
Scientific Reports (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.