Abstract
Peripheral tolerance prevents the initiation of damaging immune responses by autoreactive lymphocytes. While tolerogenic mechanisms are tightly regulated by antigen-dependent and independent signals, downstream pathways are incompletely understood. N-myc downstream-regulated gene 1 (NDRG1), an anti-cancer therapeutic target, has previously been implicated as a CD4+ T cell clonal anergy factor. By RNA-sequencing, we identified Ndrg1 as the third most upregulated gene in anergic, compared to naïve follicular, B cells. Ndrg1 is upregulated by B cell receptor activation (signal one) and suppressed by co-stimulation (signal two), suggesting that NDRG1 may be important in B cell tolerance. However, though Ndrg1−/− mice have a neurological defect mimicking NDRG1-associated Charcot-Marie-Tooth (CMT4d) disease, primary and secondary immune responses were normal. We find that B cell tolerance is maintained, and NDRG1 does not play a role in downstream responses during re-stimulation of in vivo antigen-experienced CD4+ T cells, demonstrating that NDGR1 is functionally redundant for lymphocyte anergy.
Similar content being viewed by others
Introduction
Balancing the sensitivity required for an effective adaptive immune response, while limiting reactivity to prevent autoimmunity, depends upon an increasingly well-defined series of checkpoints. Lymphocyte anergy is a state of cell-intrinsic functional inactivation, classically described when B or T cells become unresponsive upon antigen-receptor binding by self-antigen1,2. This is thought to occur during development when cells are exposed to self-antigen, at insufficient affinity or avidity to induce receptor editing or deletion, or in mature T and B cells when exposed to antigens in the absence of accessory signals in the periphery3,4. Central to the fate decision between induction of a tolerogenic or immunogenic response in the periphery is Bretscher and Cohn’s two signal model of cell activation5, in which full activation of B and T lymphocytes requires two distinct signals: antigen-engagement of the B cell receptor (BCR) or T cell receptor (TCR) respectively (signal one), which leads to limited initiation of intracellular signaling cascades, and co-stimulation by activated antigen-presenting cell (APC) ligands or CD4+ T cell help (signal two), which triggers the sustained, complete signaling required for productive activation6,7. If signal two is not received within a temporally-defined window of signal one, this leads to induction of an unresponsive state and cell death, limiting inappropriate responses3,7.
In the context of B cell tolerance, immature B lymphocytes that are not eliminated upon encountering low avidity forms of self-antigen in the bone marrow (BM) can undergo phenotypic and biochemical changes to produce a tolerized, functionally-inactive state3,8,9. As these anergic B lymphocytes emerge into the periphery, they have an increased threshold for activation, due to proximal blockade of BCR-signaling and selective inhibition of IgM trafficking to the cell surface, classically demonstrated in BCR Ig transgenic (tg) mouse models, including the anti-hen-egg lysozyme (HEL)-specific (IgHEL) model9,10. Continual low avidity BCR-signaling in the absence of co-stimulation induces chronic low oscillations of calcium, resulting in sustained sub-optimal activation of ERK and NFAT dependent pathways11,12,13, with proximal blockade of BCR-dependent signaling and a specific block in JNK, Card11, NF-κB13,14,15 and pathways downstream of toll-like receptors (TLRs)16. The resultant negative regulatory program diminishes both proliferative responses and capacity for cytokine production in response to stimulation. Furthermore, anergic B cells have a shortened lifespan in a mixed repertoire, due to enhanced dependence on the cytokine B cell activating factor (BAFF) and increased expression of proapoptotic Bim17,18,19. However, through high avidity stimulation of the BCR and in the presence of antigen-specific T cell help, these anergic B cells can initiate differentiation and proliferation and can be recruited into the immune response to foreign antigen20,21.
Compared to B cell tolerance, central T cell tolerance by thymic clonal deletion or anergy induction is more robust; however not all self-antigens are presented in the thymus and thus peripheral tolerance plays an important role in preventing autoimmunity4. In vivo peptide-induced CD4+ T cell anergy has been described as a tolerogenic, autonomous mechanism induced in peripheral T cells that have experienced antigen stimulation in the absence of co-stimulation4,22. Characterized by the inability to proliferate or produce cytokines upon re-stimulation, this hypo-responsive state cannot be reversed by IL-2, in contrast to in vitro antigen-experienced T cells re-stimulated with signal one alone23,24. Alongside the shared phenotype of clonally anergic CD4+ T cells and tolerized B cells, the signaling pathways that characterize the two phenomena also have parallels. In both cell types, the NFAT pathway is selectively activated, modulating negative regulators that induce further downstream anergy factors necessary for the hypo-responsive state11,12,25,26. Elucidating the shared and specific molecular mechanisms that block BCR and TCR signaling during the induction of lymphocyte tolerance may unlock new treatments for autoimmune and immunodeficient diseases.
NDRG1 was previously identified as a clonal anergy factor upregulated in naïve and anergic CD4+ T cells by signal one alone via the TCR-dependent early growth response genes, Egr2 and Egr3, and degraded in a proteasome-dependent manner by co-stimulation. While in vitro induction of NDRG1 expression led to a hypo-responsive state in T cells, the loss of NDRG1 in vivo was described as partially rescuing T cells from peptide-induced tolerance27. NDRG1 is a stress- and hypoxia-inducible 43 kDa protein regulating cell growth and differentiation28,29, and reported to possess potent anti-metastatic activity in a cell type and tissue-specific manner30,31. Though ubiquitously expressed in human tissues, the amount and subcellular localization of NDRG1 expression varies by cell type, with relatively higher staining in kidney, prostate, ovary, intestines and brain. Particular localization to the cytoplasm of Schwann cells in peripheral nerve tissue32 is important, since loss of NDRG1 causes sensorimotor neuropathy33,34. NDRG1 expression is suppressed via metabolic regulators including N-myc, and the locus is hypermethylated in many cancer tissues including gastric, colon, pancreatic, prostate and some breast cancers28,35,36,37,38. NDRG1-mediated inhibition of tumor growth and metastasis may explain the anti-cancer effects of iron chelating agents such as Dp44mT31,39,40. However positive correlation between tumor tissue NDRG1 levels and cancer progression in some human studies, suggests pleiotropic and context specific roles41,42,43,44, with silencing of NDRG1 inhibiting migration, invasion and viability of hepatocellular cancer and sarcoma cell lines45,46. The potential for pharmacological targeting of NDRG1 in cancer highlights the importance of understanding the role of NDRG1 within adaptive immunity.
In this study, we show that Ndrg1 is the third most differentially expressed gene in anergic B cells compared to naïve B cells and that the transcriptional regulation of Ndrg1 parallels that described in stimulated CD4+ T cells27. Despite this expression pattern, our experiments using a Ndrg1−/− mouse model demonstrate that NDRG1 is not required for the tolerogenic responses downstream of self-antigen engagement in B cells. In contrast to the observations of Oh et al. when we replicate the experimental protocol of in vivo peptide-induced T cell anergy, we find NDRG1 does not affect immune response or IL-2 production in antigen-experienced CD4+ T cells. These findings re-position NDRG1 as a bystander in anergy and have implications for the development of NDRG1 as a therapeutic target in cancer.
Results
Ndrg1 expression is associated with B cell anergy
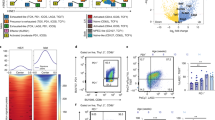
To investigate the pathways underlying B cell anergy, we performed RNA Seq on B220+CD19+IgMa+CD93−CD21midCD23hi naïve and B220+IgMloIgDa+ anergic splenic B cell subsets sorted respectively from IgHEL tg, and double tg mice expressing IgHEL with or without sHEL as a self-antigen (IgHEL/sHEL) (Fig. 1a). Differential gene expression analysis revealed significant upregulation of anergy-associated transcription factors Egr2 and Egr3, as well as Egr2 dependent targets Nab2 and Nrgn, in anergic compared to naïve B cells, a pattern of expression consistent with NFAT signaling in anergic cells (Fig. 1a, b, Supplementary Table 1)47. Of the 868 significantly differentially expressed transcripts, Ndrg1 was identified as the third most upregulated in anergic B cells by fold change (log2 7.378), and eleventh highest by p value (7.41 × 10−117 adjusted; Supplementary Table 1). NDRG1 has been described as a CD4+ T cell anergy factor downstream of Egr2/327, which together with these findings suggested it might be a common participant in the regulation of lymphocyte tolerance. Significant differential expression of Ndrg1 was verified by qPCR of mRNA extracted from naïve IgHEL or anergic IgHEL/sHEL B220+ cells (1.424 ± 0.1807 vs. 20.05 ± 2.069, p value 0.0009), confirming the association between NDRG1 and B cell anergy.
a Top panel: differential gene expression between IgHEL naïve follicular (B220+CD19+IgMa+CD93−CD21midCD23hi) and IgHEL/sHEL anergic (B220+IgMaloIgDa+) splenic B cell populations, represented in FACS plots. Lower Panel: Genes differentially expressed >1 and <−1 log2 fold change (FC) and below a significance threshold of adjusted p value, 10e−32, are shown in red, with the top 30 differentially expressed genes filtered by adjusted p value labeled. b Heatmap of the relative abundance of the top 30 differentially expressed genes by adjusted p value between naïve (left) and anergic (right) B cells. Columns represent samples from individual mice.
Ndrg1 is induced by BCR signaling but suppressed by positive co-stimulation
In CD4+ T lymphocytes, NDRG1 has been proposed as an anergy factor, induced by signal one and inhibited by signal two. To investigate whether Ndrg1 shared a similar pattern of molecular regulation in B cells, qPCR was performed on mRNA isolated from primary IgHEL B cells stimulated for 24 h ex vivo via the BCR by anti-IgM or the cognate antigen sHEL (signal one), in the presence or absence of co-stimulation by anti-CD40 or lipopolysaccharide (LPS; signal two). Consistent with the transcriptional control pattern of other anergy factors8,48, in vitro antigen-receptor signaling via signal one alone induced Ndrg1 mRNA expression in primary B cells, but addition of signal two stimulation was associated with a dose-dependent suppression of Ndrg1 mRNA (Fig. 2a). This pattern of Ndrg1 expression was also observed in vitro using A20 immortalized B cells engineered to express the IgHEL BCR (Fig. 2b), supporting the possibility that NDRG1 may play a role in the induction or maintenance of B cell anergy in central or peripheral tolerance.
a Ndrg1 mRNA expression calculated relative to Gapdh expression in IgHEL primary B lymphocytes stimulated ex vivo with the indicated concentrations of anti-IgM or sHEL, with or without anti-CD40 or LPS for 24 h at 37 °C. b Ndrg1 mRNA expression calculated relative to Gapdh expression in murine A20 IgHEL tg B cells cultured for 24 h at 37 °C with anti-IgM and with or without LPS. Lines show means with 95% confidence interval (CI) error bars. Data presented are from two to four replicate qPCR readings for each condition and is representative of 3 independent experiments using IgHEL or non-tg (a) or pooled from two A20 (b) independent experiments. Significance was determined using a two-way ANOVA with post-hoc Tukey’s testing, ***p < 0.001 and ****p < 0.0001.
A mouse model of NDRG1 deficiency
To explore the role of NDRG1 in B cell immune function, we created a Ndrg1−/− mouse model using CRISPR/Cas9 directed mutagenesis (Supplementary Fig. 1a). The gene targeting produced an 11 bp deletion within exon 4 of the Ndrg1 gene in oocytes causing a frameshift mutation Ndrg1 null allele, which was propagated further to produce homozygote Ndrg1−/− mice at expected ratios from heterozygous parents (Supplementary Fig. 1b). PCR-amplification and subsequent Sanger sequencing of cDNA reverse transcribed from RNA isolated from WT and Ndrg1−/− kidney and splenic B cell lysates demonstrated that the edited RNA transcript is expressed in both tissues (Supplementary Fig. 1c, d) with the 11 bp deletion detectable within the transcript amplified from Ndrg1−/− kidney and B cell samples with no exon skipping or repair (Supplementary Fig. 1d). NDRG1 protein is expressed at very low levels in WT splenocyte samples and thus undetectable by transitional western blot, however knockout was confirmed by immunoblot analysis of WT and Ndrg1−/− in kidney lysates using a N-terminal binding antibody (Supplementary Fig. 1e). In splenic lysates, immunoprecipitation and blotting confirmed the complete loss of NDRG1 protein expression, likely a result of nonsense-mediated decay of the altered transcript (Supplementary Fig. 1f).
In humans, nonsense Ndrg1 mutations are one of the causes of hereditary motor and sensory peripheral neuropathy-type Lom, also known as Charcot-Marie-Tooth disease type 4d (CMT4d)33. Consistent with the human disease, and previously reported neurological phenotyping of Ndrg1−/− mice34, our Ndrg1−/− mice display a shaking, tremorous phenotype with muscle weakness and abnormal clasping of the hind limbs, and reduced proportion of unsupported versus supported rearing (Supplementary Fig. 1g), indicating hind limb weakness and providing further evidence that the protein had been correctly targeted. Ndrg1−/− mice were smaller than their WT littermate controls, as previously described (Supplementary Fig. 1h)49.
Normal development of the pre-immune repertoire in the absence of NDRG1
Immunophenotyping of Ndrg1−/− non-tg mice by flow cytometry demonstrated normal development of B cells in the BM (Fig. 3a, d), and no difference in the development, distribution or absolute numbers of peripheral B cell populations within the spleen or absolute B220+ B cell numbers in the mesenteric lymph nodes (MLNs) (Fig. 3b, e). Peritoneal B1 B cell populations were also comparable (Fig. 3c), indicating that Ndrg1 deletion has no effect on development or distribution of unperturbed B cell populations; and serum IgM, IgG and IgA titers were equivalent in Ndrg1−/− and WT mice (Fig. 3f).
a Representative flow cytometry plots of adult WT (blue) and Ndrg1−/− (green) mice, showing Hardy B cell fractions in BM (a), and splenic (b) and peritoneal (c) B cell subsets. d Quantification of absolute BM B cell numbers as gated in (a). e Quantification of B cell subsets, transitional (trans), transitional populations (T1–T3), marginal zone (MZ) and follicular (Fo) numbers in the spleen and total B cells in MLNs as gated in (b). Points are individual mice, lines represent means with 95% CI error bars and are representative of three independent experiments, each with 3–6 mice per group. f Serum Ig levels from WT and Ndrg1−/− mice determined by ELISA. Points are individual mice and lines represent means with 95% CI error bars with 12–16 mice per group. Dotted line represents 1.5× interpolated background absorbance.
To investigate the requirement for NDRG1 during the response to stimulation by foreign antigens, Ndrg1−/− and WT B cells were cultured under a range of stimulatory conditions for 24 h and 72 h. The absence of NDRG1 did not alter the in vitro upregulation of activation markers CD69, CD86 or CD25, the extent of proliferation or class-switching in response to TLR ligands (Supplementary Fig. 2a–d).
Central tolerance and anergy induction during development do not require NDRG1
To explore whether loss of NDRG1 is sufficient to break tolerance to self-antigen at resting state, we tested adult WT and Ndrg1−/− mouse serum for autoantibodies to nuclear epitopes (ANAs), a sensitive hallmark of autoimmune disease in both mice and humans50. There was no increase in ANAs in the absence of NDRG1 expression (WT 1/9, 11.1% positive vs. Ndrg1−/− 1/9, 11.1% positive, P = 1, two-tailed Fisher’s exact test).
To isolate and test for a specific defect in B cell tolerance, we investigated whether the provision of T cell help might be sufficient to induce autoantibodies to endogenous self-antigen in the absence of NDRG1. Accordingly, WT or Ndrg1−/− non-tg and WT or Ndrg1−/− sHEL mice were primed with HEL-conjugated to sheep red blood cells (HEL-SRBC) and then boosted with HEL-SRBC at 30 days (Fig. 4a). There was no difference in the humoral immune response between WT or Ndrg1−/− mice in these experiments. As expected9, the sHEL expressing mice generated lower HEL-specific IgG titers than non-tg mice, despite equivalent GC formation across groups (Fig. 4b), indicating B cell tolerance to self-antigen, despite the availability of T cell help, remained intact in the absence of NDRG1 (Fig. 4c).
a Experimental schematic: WT and Ndrg1−/− non-tg or sHEL-tg recipients were immunized with HEL-SRBC at day 0 and then boosted with HEL-SRBC again at day 31 before serum was collected on day 35. b GC formation on day 35 from mice in (a) as determined by gating on GL7+CD95+ of B220+CD19+ live lymphocytes. c Serum anti-HEL IgG titers from mice in (b) on day 35. Data for (b, c) are from one experiment with 4–6 mice per group, each point is one mouse and lines are at the mean with 95% CI error bars. Dotted line represents unimmunized background titer level. d Experimental schematic: lethally irradiated sHEL or non-tg CD45.1+ recipients were reconstituted with WT or Ndrg1−/− CD45.2+ IgHEL tg BM for 8 weeks. e Representative flow cytometry plots of B220+CD43− BM cells from mice in (d). f IgMa surface expression on mature recirculating B cells in the BM (B220+CD43−IgD+), and on splenic B220+CD19+ B cells from mice in (d), quantified by flow cytometry as MFI. Results are representative of three independent experiments (2–4 mice per group), each point represents individual mice and lines show means with 95% CI error bars. g MFIs of phospho-flow intracellular staining of pSyk, pPLCy2, pBLNK and pERK in splenic B220+CD19+ IgHEL B cells from WT and Ndrg1−/− mice in (d), after stimulation ex vivo with media alone, 1 μg/ml sHEL or 10 μg/ml IgM. Results are pooled from 3 independent experiments with 2–4 mice per group. Box plots show boxes representing 25th to 75th percentile with a line at median and bars showing min to max. h Serum levels of anti-HEL IgMa in (d). Each point represents individual mice and lines show means with 95% CI error bars and are representative of two independent experiments. Dotted line shows 1.5× interpolated background. i Ex vivo proliferation of B220+ splenocytes quantified by CTV dilution in response to media alone, or 500 ng/ml LPS plus sHEL titration for 72 h at 37 °C. Results are pooled from 2 to 3 independent experiments per group, each point represents one mouse.
To investigate whether NDRG1 might play a role in tolerance and anergy induction during B cell development in the context of self-antigen, we took further advantage of the IgHEL and sHEL-tg models. Lethally irradiated CD45.1+ non-tg and sHEL mice were reconstituted with WT or Ndrg1−/− CD45.2+ IgHEL BM (Fig. 4d). Irrespective of NDRG1, flow cytometric analysis showed downmodulation of surface IgMa on mature recirculating B cells in the BM and spleens of sHEL recipients, typical of anergy (Fig. 4e, f); and ex vivo stimulation of the same cells with sHEL showed a block in BCR signaling (Fig. 4g). Anti-HEL-specific IgMa was also equivalently suppressed in sHEL recipients reconstituted with WT or Ndrg1−/− IgHEL BM (Fig. 4h). These findings show that NDRG1 is dispensable for the IgM modulation, and block in both proximal BCR signaling and plasma cell differentiation, characteristic of anergic B cells.
Persistent self-antigen stimulation of the BCR in anergic B cells also has the effect of inhibiting proliferative responses to LPS, an essential checkpoint to prevent autoimmunity during exposure to external co-stimulation such as with bacterial infection16. To exclude the possibility that NDRG1 might be induced to mediate the inhibition of co-stimulation by LPS, we stimulated CTV-labeled WT and Ndrg1−/− IgHEL naïve and IgHEL/sHEL double tg anergic splenic B cells ex vivo with LPS and increasing amounts of sHEL as self-antigen. Proliferation was equivalently inhibited in Ndrg1−/− anergic B cells compared to WT, indicating that the refractory mechanisms in place to inhibit autoreactive B cell responses to LPS do not depend upon NDRG1 expression (Fig. 4i).
NDRG1 deficiency does not provide a competitive advantage to anergic B cells within a polyclonal repertoire
The antigen-induced differential signaling pathways in anergic B cells cause a shortened lifespan in a polyclonal repertoire, due to an inability to compete with non-anergic cells for BAFF, and increased expression of the proapoptotic protein, Bim17,18,19. Since NDRG1 has been implicated in the induction of apoptosis51, we hypothesized that NDRG1 might act downstream of signal one to modulate the threshold for survival in a diverse B cell repertoire. To test this, we reconstituted lethally irradiated non-tg or sHEL CD45.1+ mice with 80:20 mixtures of Ndrg1−/− or WT CD45.2+ IgHEL BM and WT non-tg CD45.1+ BM (Fig. 5a). Analysis of the reconstituted mice showed equivalent numbers of Ndrg1−/− and WT IgHEL B cells in a mixed repertoire (Fig. 5b, Table 1) and an equivalent increase in turnover, in the presence of self-antigen, as judged by BrdU labeling in vivo (Fig. 5c). These data show that NDRG1 is not required to maintain the rapid turnover of anergic B cells that have developed in the presence of self-antigen in the context of competition with non-autoimmune cells for BAFF.
a Experimental schematic; lethally irradiated WT or sHEL transgenic CD45.1+ recipients were reconstituted with 80:20 mixtures of WT or Ndrg1−/− IgHEL CD45.2+ BM and WT non-tg CD45.1+ BM. b Percentage of total HEL-binding B220+ CD45.1− cells in the BM and spleen of mice in (a). c Percentage of HEL-binding B220+CD45.1− cells labeled with BrdU after one week. In (b, c), dots are individual mice from one experiment with 5–6 mice per group, lines show means and 95% CI error bars. d. Representative flow cytometry plots of B220+CD43− B cells from BM of sHEL mice reconstituted with 50:50 mixtures of WT IgHEL CD45.1+ BM with either WT IgHEL or Ndrg1−/− IgHEL CD45.2+ BM. e Surface IgM expression on B cell populations gated as in (d). f Intracellular Bim quantification in B220+CD19+IgDa+splenocytes from (d). Each point represents individual mice and lines represent means with 95% CI error bars from one experiment (4–5 per group).
Similarly, NDRG1 deficiency did not produce an altered anergic response in B cells exposed to self-antigen in the presence of directly competing WT IgHEL anergic B cells in a mixed chimera model. Flow cytometric analysis of CD45.1+ sHEL recipients reconstituted with equal parts WT CD45.1+ IgHEL BM and either WT IgHEL or Ndrg1−/− IgHEL CD45.2+ BM showed comparable levels of surface IgM (Fig. 5d, e), and equivalent Bim expression between Ndrg1−/− and WT IgHEL mature B cell populations in the chimeric mice (Fig. 5f).
NDRG1-deficient B cells exposed to self-antigen in the periphery are subject to follicular exclusion and cell death
Our experiments have demonstrated that NDRG1 is dispensable for the induction and maintenance of anergy when B cells develop in the presence of self-antigen and when challenged with signal two alongside chronic BCR stimulation; however, this does not exclude a role for NDRG1 unique to migration and re-localization of antigen-stimulated B cells, which is linked to their ability to find and interact with T cell help. When naïve B cells first bind antigen, they relocate to the outer periarteriolar lymphoid sheath where they require signal two in the form of T cell help for survival and entry into the follicle to initiate a productive germinal center19,52. Similarly, in the context of self-antigen and competing naïve cells, anergic B cells also relocate to the T cell rich cortical zones, where they may be rescued by T cell help; but otherwise typically die from a combination of insufficient engagement with BAFF and increased Bim-dependent apoptosis17,18,53.
To exclude a possible role for NDRG1 in these interactions, equal ratios of WT IgHEL CD45.1+ and WT or Ndrg1−/− IgHEL CD45.2+ splenocytes were labeled with CTV dye and transferred into non-tg or sHEL CD45.2+ recipients (Fig. 6a). After 48 h, the total number of transferred CTV+ HEL-binding IgHEL B cells were significantly reduced in sHEL compared to non-tg recipients, but ratios of CD45.1+:CD45.2+ surviving transferred HEL-binding cells were equivalent for WT and Ndrg1−/− donor populations (Fig. 6b). These findings show that NDRG1 does not play a role in the elimination of signal one experienced B cells in the periphery. This conclusion is supported by the equivalent induction of CD95, BAFF-R, Bcl2 and Bim proteins in the CD45.1+ WT and CD45.2+ Ndrg1−/− antigen-experienced B cells (Fig. 6c). In sHEL recipients, IgMa downmodulation and upregulation of the activation marker CD86 were similar in CD45.1+ WT and CD45.2+ Ndrg1−/− IgHEL B cells (Fig. 6c). Levels of localization markers CD23 and CXCR5 were also reduced equivalently on exposure to sHEL in CD45.2+ Ndrg1−/− antigen-experienced B cells, suggesting follicular exclusion is also unlikely to depend on NDRG1 expression (Fig. 6c).
a Representative flow cytometry plots of 50:50 mixtures of WT CD45.1+ IgHEL and WT or Ndrg1−/− CD45.2+ IgHEL splenocytes labeled with CTV dye and transferred by i.v. injection into non-tg or sHEL-tg CD45.2+ recipients for 48 h, showing live B220+CD19+ gated populations and % transferred CTV+ cells. b Percentage of total lymphocytes that were either CD45.1+ or CD45.2+ CTV+ as gated in (a), with white points showing paired CD45.1+ IgHEL cells from donor, colored points are CD45.2+. c MFI of surface marker expression CD86, CD23, CXCR5, IgMa, BAFF-R and CD95, and intracellular Bim and Bcl2, in transferred IgHEL B cells, gated as in (a). Data points are individual mice and lines show means with 95% CI error bars and data are representative from 1 to 3 independent experiments with 4–6 mice per group. d, e Ratio of B220+ cells surviving from 50:50 mixtures of WT CD45.1+ and WT or Ndrg1−/− CD45.2+ non-tg B cells stimulated with anti-IgM at the indicated concentrations for 16 h at 37 °C ex vivo (d) and CD86 surface expression as MFI on the CD45.2 + B220+ cells (e). Data are representative of 3 independent experiments with columns or circles as means with 95% CI error bars.
To further confirm that NDRG1 has no role in inducing apoptosis downstream of signal one in the absence of signal two, mixed cultures of WT CD45.1+ and WT or Ndrg1−/− CD45.2+ non-tg B cells were stimulated with a titration of plate-bound anti-IgM for 16 h in vitro18. The ratio of CD45.1+:CD45.2+ B cells was not altered by increasing concentrations of anti-IgM in cultures (Fig. 6d), despite equal antigenic engagement as measured by CD86 upregulation (Fig. 6e), supporting our conclusion that NDRG1 does not induce B cell death in response to signal one in the absence of signal two.
Antigen-experienced T cells responses are not altered in the absence of NDRG1
The finding that NDRG1 is either dispensable or acts redundantly for the induction and maintenance of B cell anergy, led us to re-examine the evidence for NDRG1 regulation of induced T cell anergy. As previously reported27, NDRG1 deletion had no effect on thymocyte development (Supplementary Fig. 3a, c) or maturation and distribution of peripheral T populations (Supplementary Fig. 3b, d).
To confirm that NDRG1 acts as a T cell-specific anergy regulator, we recapitulated the previously described in vivo peptide-induced T cell anergy assay27,54 using our mouse model of NDRG1 deficiency. In these experiments, CD45.2+ ovalbumin (OVA)-specific TCR tg CD4+ OT-II+ T cells were isolated from WT or Ndrg1−/− OT-II+ tg mice and adoptively transferred into CD45.1+ congenic recipients, with subsequent intravenous injection of endotoxin-free OVA peptide on day 1. After 5 days, naïve and challenged splenic WT and Ndrg1−/− OT-II+ CD4+ T cells were harvested and stimulated in vitro (Fig. 7a).
a Experimental schematic; OVA-specific CD4+ OT-II+ T cells were isolated from CD45.2+ WT or Ndrg1−/− OT-II+ tg mice and adoptively transferred to CD45.1+ non-tg recipients, with subsequent intravenous injection of 500μg OVA peptide. After 5 days, splenic CD4+ cells were isolated from recipient mice and naïve unstimulated WT or Ndrg1−/− OT-II+ tg mice and loaded with CTV. Equal numbers of Vα2+ CD45.2+OT-II+ cells were cultured in vitro with a titration of OVA peptide for 72 h at 37 °C with irradiated APCs. b IL-2 concentration in culture supernatants in (a). c Proliferation of CD4+Vα2+ CD45.2+OT-II+ T cells and non-tg bystander WT CD45.1+ cells (red) as measured by CTV dilution by flow cytometry. d Quantitation of intracellular IL-2 and IFNγ production by flow cytometry within OT-II+ CD4+ T cells from (a). Points represent means of technical replicates from one (naïve) or two (antigen-experienced) mice with error bars representing standard deviation. Results are representative of two (b), three (c) or one (d) independent experiment(s).
As reported previously27,54, IL-2 concentration in the culture supernatant from antigen-experienced CD4+ T cells was lower compared to that from naïve cells (Fig. 7b; p = < 0.05 at all concentrations of OVA). However, we could not replicate the reported increase in IL-2 in Ndrg1−/− relative to WT antigen-experienced T cells27 (Fig. 7b). In contrast to some previous findings using a thymidine incorporation assay, flow cytometric analysis of CTV dilution reproducibly demonstrated a significantly enhanced proliferation response in antigen-experienced OT-II+ T cells compared to naïve (Fig. 7c; p = < 0.05 between 0.1 and 5 μg/ml OVA), irrespective of NDRG1 expression. Bystander WT CD45.1+ non-tg cells demonstrated no proliferation in cultures containing either WT or Ndrg1−/− OT-II+ cells at any concentration of OVA (Fig. 7c, red points).
We therefore considered that the assay may represent a stimulatory response rather than anergy, and hypothesized that the lower IL-2 in the supernatant of antigen-experienced cells may consequently reflect a higher rate of consumption due to greater proliferative responses of these cells compared to cultures containing naïve cells. To test this, production of cytokines IL-2 and IFNγ was directly measured by intracellular staining. Ag-experienced populations produced more IL-2 and IFNγ in a dose-dependent manner than naïve CD4+ T cells, consistent with a stimulatory rather than anergic response (Fig. 7d, p = < 0.05 at 0.1–0.5 μg/ml OVA), a response that was unaffected by NDRG1 deficiency (Fig. 7d).
Discussion
The development of more effective and better tolerated treatments for cancers, autoimmune and immunodeficiency diseases relies on defining the key molecular mechanisms that distinguish productive from tolerogenic lymphocyte responses8. Despite the success of many monoclonal therapies targeting immune checkpoints, they are associated with an increased risk of autoimmune adverse events55,56,57. Pre-clinical investigation into the immune consequences of anti-cancer therapy is therefore key to preventing serious immunopathology. NDRG1 is a stress- and hypoxia-inducible regulator of cell growth and differentiation28,29, implicated in some contexts as a metastasis inhibitor due to involvement within multiple signaling pathways30,31. Consistent with this, anti-cancer iron chelators, Dp44mT and DFO may act via NDRG1-upregulation31,39,40. Pertinently for this study, in multiple human cancers increased tissue expression of NDRG1 has been associated with poor outcomes41,42,43,44, with in vitro evidence for suppression of NDRG1 as a novel anti-cancer strategy45,46. The therapeutic targeting of NDRG1 presents a promising treatment opportunity, but requires clarification of the immune effects of NDRG1 inhibition.
In 2015, Ndrg1 was described as a T cell anergy factor, regulated by the signal one/signal two axis, induced downstream of NFAT and Egr2 upon TCR signaling and repressed by co-stimulation. Ndrg1−/− CD4+ T cells were reported to be partially resistant to peptide-induced anergy in vivo27. Selective activation of the calcium-regulated transcription factor NFAT, and Egr family driven gene signatures, are known to contribute to both B and T lymphocyte unresponsiveness11,12,25,26, with defects in maintenance of T and B cell tolerance, including increased production of self-reactive Ig, in NFAT1−/− mice12,58. Our findings identified Ndrg1 as the third most upregulated gene in anergic B cells compared to naïve cells. The evidence for shared genetic regulation of Ndrg1 by the signal one and two costimulatory axis in B and T cells27 suggested that NDRG1 could represent a potential central factor regulating lymphocyte tolerance downstream of Egr2/3 and NFAT activation.
Our experiments employing a CRISPR/Cas9 generated NDRG1-deficient mouse demonstrated that NDRG1 is either dispensable or acts redundantly within tolerance mechanisms during B cell development and activation. NDRG1 deficiency did not affect induction and maintenance of B cell anergy in the IgHEL > sHEL BM chimeric model, nor did it alter the maintenance of cell-intrinsic tolerance to self-antigen at resting state, upon immunization or during direct competition. We hypothesized that NDRG1 action may be limited to specific properties of B cell tolerance such as influencing the threshold for breakdown of tolerance in response to external stimulation, as Ndrg1 expression is closely regulated by signal two in B and T lymphocytes; however, the characteristic inhibition of signal two was intact in Ndrg1−/− anergic B cells. Furthermore, the loss of NDRG1 had no effect on the response of naïve splenic B cells exposed to self-antigen, or the induction of cell death due to the absence of rescue by signal two. There was no requirement for NDRG1 in these aspects of B cell tolerance, despite clear regulation of NDRG1 by the signal one/signal two axis in lymphocytes.
To strengthen the findings from our mouse model of NDRG1 deficiency, and with the aim of demonstrating that NDRG1 functions in CD4+ T cell tolerance, we recapitulated an experimental design to show the role of NDRG1 in in vivo peptide-induced T cell anergy27. In contrast to the results reported previously, we saw increased proliferation of endotoxin-free peptide antigen-experienced in vivo pre-stimulated CD4+ T cells, as would be expected during re-stimulation rather than an anergic response. The reported reduction in IL-2 in culture supernatant upon re-stimulation of antigen-experienced CD4+ T cells was reproducible, but NDRG1 deficiency did not affect this phenomenon. On further exploration, we found that the intracellular levels of IL-2 and IFNγ were higher in the antigen-experienced cells, consistent with increased consumption of cytokines associated with a higher proliferative response on re-stimulation, explaining the reduced IL-2 detected in the culture supernatants of these cells.
The in vivo peptide-induced T cell anergy model is well described and characterized by reduced IL-2 in culture supernatant22,54,59, though the response to IV peptide may be variable22, and is not consistent in non-chimeric OT-II tg mice54. Our use of CTV viable proliferation dye in contrast to thymidine incorporation to measure proliferation accounts for any disproportionate loss or inhibition of OT-II+ CD4+ T cells during in vitro culture. Though we find that non-tg cells from the transfer recipient contribute minimally to proliferation in antigen-experienced conditions, we cannot exclude a suppressive or cytotoxic effect on antigen-experienced OT-II+ cells from non-tg peptide experienced cells in vitro in previously described experiments; such effects have been shown to inhibit proliferation in an OT-I+ CD8+ in vivo peptide model60. Together, these experiments suggest that this experimental system may not reliably test for peptide-induced CD4+ T cell unresponsiveness and call into question the proposed role of NDRG1 in vivo T cell anergy.
In contrasting our results with the previous observations in T cells, we also note that where the model of NDRG1 deficiency described here was generated using CRISPR/Cas9 on a pure C57BL/6 background, the Ndrg1−/− mouse used in previous experiments exploring T cell anergy27 was created using 129 ES cells34, a method which can be associated with co-transfer of immunomodulatory genes alongside the mutant gene61. Although the mice were backcrossed for 8 generations27, a 129 strain-specific immune effect cannot be fully excluded.
Taken together our results position NDRG1 as a bystander, which may solely represent a biomarker of lymphocytes chronically stimulated by signal one alone. An alternative possibility is that anergy is maintained in the absence of NDRG1 due to functional redundancy. Mammals have 4 NDRG1 proteins, which share 53–65% homology62,63 and transcriptional repression via myc. The preservation of central nerves despite peripheral demyelination in CMT4d has been attributed to compensation by NDRG2–464, However, it is not clear if any such compensation is relevant outside the nervous system, and notably NDRG2–4 are not upregulated in anergic B cells (Fig. 1).
When the anergic B cell engages antigen, failure to recruit Syk family kinases to the BCR65 results in Lyn mediated inhibition of the PI3K pathway via SHIP-1 and PTEN66,67. NDRG1 has been shown to exist in a positive feedback loop with PTEN, whilst inhibiting PI3K31,68. However, the lack of redundancy with loss of either SHIP-1 or PTEN suggests that NDRG1 does not act reciprocally with these proteins66,67. In addition to a role in PI3K/AKT signaling, evidence from the cancer field indicates that NDRG1 can also modulate the TGFβ/SMAD and RAS/RAF/MEK/ERK axes, inhibit NF-κB and the ErbB receptors, suppress sonic hedgehog and modulate the phosphorylation of Cbl-b31,69, Cbl-b itself associated with maintenance of B and T cell tolerance70,71. NDRG1 also inactivates canonical Wnt/β-catenin signaling in BM stromal progenitor cells and cancer cells, while promoting β-catenin plasma membrane expression8,9. Stable β-catenin expression on CD4 T cells has been linked to an unresponsive anergic like phenotype, while uncontrolled β-catenin accumulation in development induces B cell anergy10,11. Furthermore, an independent compensatory role for NDRG1 within lymphocytes in pathways such as PI3K, Src or Wnt/β-catenin cannot be fully discounted; transcriptomic or proteomic profiling of Ndrg1−/− anergic B cells in comparison to WT anergic, and Ndrg1−/− naïve B cells may identify alterations in such pathways.
Beyond tolerance, there is in vitro and in vivo evidence for immunomodulatory NDRG1 function in mast cells, where it appears to promote degranulation response to stimuli72,73. However, while NDRG1 deficiency is a well-described cause of CMT in humans, autoimmunity or immunodeficiency are not described in CMT4d33,74,75,76.
The regulation of NDRG1 points to a central role in the cellular stress response. In addition to N-myc and C-myc, NDRG1 is upregulated after DNA damage via p5351, by hypoxia via HIF1α40 and Egr177, and by PTEN68. While the modulators of NDRG1 specific to B cells have not been well characterized, several known drivers of the anergic versus activation axis in B cells are linked to NDRG1 regulation. These include transcription factors, Egr2 and Egr3, downstream of NFAT activation in B and T cell anergy47,78, of which Egr2 has been shown to regulate NDRG1 in T cell clones27. A role for HIF1α in both B cell development79 and in regulating B cell tolerance80 has been reported and hypoxia and HIF1α-regulation of NDRG1 is well characterized31. Downstream of BCR and BAFF-R stimulation, via PTEN/PI3K, the NF-κB pathway is key for B cell activation, proliferation and survival; Rictor, an essential mTOR component, is an NDRG1 regulator81, and Rictor-deficient B cells have impaired BCR and BAFF-R engagement and associated reduced AKT (273) and NDRG1 phosphorylation82. These shared regulatory pathways between B cell anergy and NDRG1 suggest possible explanations for the observed correlation between NDRG1 levels and anergy.
In conclusion, we found no evidence that NDRG1 has any functional role during antigen-receptor signaling in B or T lymphocyte populations. We have shown that, while NDRG1 is dispensable for B cell tolerance, the association of NDRG1 expression with B cell anergy represents a biomarker for anergic cells, stimulated by the BCR in the absence of signal two, and reminiscent of other cell types under conditions of cell stress29,31,34,51,83,84,85. The fact that NDRG1 expression was not required for the induction or maintenance of B cell anergy, or other processes of chronic BCR stimulation, enhances the potential for targeting NDRG1 for human anti-cancer treatment.
Methods
Mice
All procedures involving animals were performed in accordance with the Animals (Scientific Procedures) Act 1986, amended 2012, with procedures reviewed by the clinical medicine Animal Care and Ethical Review Body and conducted under Home Office Project License, P79A4C5BA. Mice were housed in individually ventilated cages, provided with food and water ad libitum and maintained on a 12 h light:12 h dark cycle (150–200 lux). The only reported positives on FELASA health screening over the entire time course of these studies were for Helicobacter spp, Chilomastix Sp, Enteromonas muris, Trichomonas Sp, mouse Norovirus, and Entamoeba spp.
C57BL/6JOlaHsd mice were purchased from Envigo. IgHEL (C57BL/6-Tg(MD4)4Ccg/J), soluble HEL (sHEL) tg (C57BL/6-Tg(ML5)5Ccg/J) and OT-II tg mice (C57BL/6-Tg(TcraTcrb) 425Cbn/Crl) have been described previously and were maintained on a C57BL/6 background9,86. Ndrg1−/− mice were generated by CRISPR/Cas9 directed mutagenesis on the IgHEL background as described in Supplementary Fig. 1a in collaboration with Dr Ben Davies, Transgenics Core, Wellcome Center for Human Genetics. CRISPR/Cas9 nuclease guide (5’GATGACAGGACGGTTGCCCTTGG) was designed against exon 4 of the Ndrg1 gene (ENSMUSG00000005125)87. Exon 4 is the most upstream exon of the Ndrg1 gene for which, if exon skipping occurred despite mutagenesis, the resulting transcript would still lead to nonsense-mediated decay. Complementary oligos for the guide sequence (5′-CACCGATGACAGGACGGTTGCCCT-3′, 5′-AAACAGGGCAACCGTCCTGTCATC-3′) were annealed, creating a compatible linker for cloning into CRISPR/Cas9 vector pX330-Puro, pre-digested with BbsI88. A template for in vitro transcription using a T7 RNA polymerase was generated from this plasmid by PCR and used for guide-RNA preparation using the EnGen sgRNA synthesis kit (NEB), followed by purification of the resulting RNA with the Megaclear Clean-up kit (Ambion). Guide-RNA efficiency and specificity was tested by lipofection into murine melanocyte, B16F10 line. C57BL/6J oocytes derived from IgHEL studs were microinjected with 20 ng/μl guide-RNA and 20 ng/μl Cas9 mRNA before reimplantation into pseudo-pregnant foster mothers at the two-cell stage. Founder mice harboring an indel deletion in Ndrg1 were identified by PCR and Sanger sequencing using the following primers, Ndrg1 genomic set 1 for PCR and Sanger Sequencing; forward 5′-GGACTGTGCTTGTATGACATTC-3′, reverse; 5′-GTGTCCATAGTCAGTGGGTCAG-3′, Ndrg1 genomic set 2 PCR; forward 5′-CCAAACTCACGGTTCATGCC-3′, reverse 5′-CAGGTGATGGGCCTCTGTCT-3′, amplifying a 522 bp region and 151 bp region of Ndrg1 exon 4, respectively. Founder mice were then backcrossed for 7 generations with wild-type (WT) C57BL6/J mice, and intercrossed to generate homozygotes, selecting both Ndrg1−/− IgHEL tg and Ndrg1−/− non-tg animals for experiments. The following primers sets were used for PCR and Sanger sequencing of Ndrg1 mRNA, mRNA set 1 spanning whole CDS; forward 5′-ATGTCCCGAGAGCTACATG-3′ and reverse 5′-TTAGCAGGACACCTCCATGG-3′ and mRNA set 2; forward 5′-ATGTCCCGAGAGCTACATG-3′ and reverse 5′-AGTTGAAGAGGGGGTTGTAG-3′.
Sheep red blood cell (SRBC) immunization
HEL-SRBC were prepared by incubating SRBC in Alsever’s solution (FirstLink) with 20 mg HEL per 10 ml 10% SRBC in conjugation buffer (0.35 M Mannitol, 0.01 M NaCl in HBSS) and later addition of 100 mg EDCI (Sigma Aldrich). After washing with HBSS, 200 μl 2 × 109 HEL-SRBC were injected intraperitoneally into each mouse.
BM chimeras
For BM chimeras, sHEL and non-tg CD45-1+/+ or CD45-1+/− recipients were irradiated with two doses of 4.5 Gy, spaced by 3 h, and intravenously injected with at least 5 × 106 Ndrg1−/− IgHEL or WT IgHEL BM cells. All chimeras were given water treated with 1% Enrofloxacin antibiotic (Baytril, Bayer) for the first 3 weeks of reconstitution. BM chimeras were allowed to reconstitute for at least 7 weeks before immunization or analysis. Specified mice received BrdU supplemented with 1% sucrose for 7 days in their drinking water at a final concentration of 0.8 mg/ml.
Hind limb strength assessment
Hind limb strength was measured by observing over 5 min the number of times an animal reared to stand, placing weight onto hind legs, either unsupported or supported, the ratio of unsupported/supported rearing was then calculated.
Ex vivo culture and stimulation
Total B cells were isolated from RBC-lysed splenocyte suspensions by positive selection using Miltenyi Biotec MACS LS columns and CD45R (B220) microbeads. B220+ or B220− splenocytes were cultured at a final density of 1–2 × 105 cells/mL in complete R10 (RPMI media with 10% FCS plus non-essential amino acids, 1 mM sodium pyruvate and 50 µM 2-β-mercaptoethanol, 20 mM hepes, 2mM L-glutamine, 10units/mL penicillin, 10 µg/mL streptomycin and 20 µg/ml neomycin) at 37 °C, 5% CO2. For RNA extraction, cells were washed with PBS after 24 h culture and pelleted for later RNA extraction as described below. For proliferation analysis, 106–107 B220+ or B220− cells were labeled with 2.5 µM CellTraceViolet (CTV) dye before culturing for 72–96 h prior to analysis by flow cytometry as below. The following stimuli were prepared as indicated in complete R10: anti-IgM, µ-chain specific (Jackson ImmunoResearch; 115-005-075), LPS from Salmonella species (Sigma Aldrich; L7770), HEL (Sigma Aldrich; L6876-5G), anti-CD40 (Biolegend; 102908), anti-CD3e (Biolegend; 100331), anti-CD28 (Biolegend; 122004) and recombinant mouse IL-4 (Biolegend; 715004).
A20 IgHEL tg cell line
A20 is a Balb/c B cell lymphoma line, originally sourced from ATCC, a gift from Professor Simon Davis, University of Oxford. HyHEL10 IgM heavy chain and kappa light chain were cloned from IgHEL mouse cDNA into pHR-SIN-CSGW vector under the control of the spleen focus forming virus promotor, then transfected into A20 B cells in which the endogenous BCR had been targeted by CRISPR-Cas9 gene editing. HyHEL10 BCR surface expression was confirmed by flow cytometry.
Flow cytometry
Cell suspensions were isolated in RPMI media containing 2% fetal calf serum (FCS), 10–20 mM Hepes and cells counted using WBC counting fluid (1.5% acetic acid, 0.5% methyl violet in water) and a haemocytometer89. If required for downstream analyses, samples were RBC-lysed. 0.5–2 × 106 cells were aliquoted and washed with 200 μl FACS buffer (PBS with 2%FCS, 20 mM Hepes, 0.05% sodium azide). Cells were stained in antibody staining cocktail, then washed for acquisition. HEL-binding cells were detected by pre-incubation with 200 ng/ml HEL, washed with FACS buffer, then counterstained with HyHEL9 conjugated to pacific blue or FITC in house89. Data were collected on a BD FACS CANTO10c. The following antibodies used during flow cytometric staining were from Biolegend; anti-B220 (1:400–500, 103236, 103232, 103212, 103243), anti-CD19 (1:400, 115528, 115530), Zombie Aqua live/dead (1:200, 423102, 423106), anti-IgM (1:400–500, 406508, 406512), anti-IgD (1:400, 405704, 405716, 405708), anti-IgDa (1:400, 406104), anti-IgMa (1:600, 408608), anti-CD23 (1:250, 101614), anti-CD93 (1:100, 136510), anti-CD21 (1:400, 123418, 123412), anti-CD24 (1:400, 101820, 101836), anti-CD5 (1:100, 100629), anti-CD86 (1:400, 105028), anti-CD44 (1:100, 103020, 103006), anti-CD25 (1:500:800, 102008), anti-CD4 (1:400, 100430), anti-CD8a (1:400, 100734), anti-CD62L (1:100, 104412), anti-CD3 (1:100, 100214, 100330, 100328), anti-CD69 (1:200, 104530), anti-CD45.1 (1:200, 110730, 110722, 110708), anti-CD45.2 (1:200, 109841, 109818, 109824), anti-BAFF-R (1:200, 134103), anti-CD95 (1:200, 152604) and anti-TCRVα2 (1:200, 127806). The following antibodies were from BD Pharmingen; anti-CD21 (1:400, 563176), anti-BP-1 (1:100, 553735), anti-IgMa (1:500, 553516), anti-CD43 (1:100, 562865), anti-IgM (1:500, 553437), phospho-PLCγ2 (1:25, 558498), phospho-BLNK (1:25, 558443), anti-BrdU (1:20, 364108), anti-CXCR5 (1:200, 145504), anti-Bcl2 (1:200, 633506), anti-Bcl2 quantification kit (1:5, 556537), anti-CD95 (1:200, 557653) and anti-IgG-1 (1:200, 563285). The following antibodies were from eBioscience anti-IgM (48-5890-82), anti-CD4 (1:400, 25-0041-82), phospho-ERK (1:50, 53-9109-42) and phospho-SYK (1:50, 12-9014-42). CellTraceViolet cell proliferation kit (C34557) was from ThermoFisher. Anti-Bim was from CST NEB (1:100, 10408S).
Phospho-FLOW and intracellular staining
2–5 × 106 splenocytes were stimulated with 1μg/ml sHEL or 10μg/ml anti-IgM F(ab)2 (Jackson Immunoresearch) at 37 °C for 5 min and subsequently fixed and permeabilized using the BD Cytofix/Cytoperm kit in combination with Cytofix and Perm Wash buffer (BD Biosciences; 554655, 554722, 557885) before staining in intracellular antibody staining cocktail.
To detect BrdU incorporation, surface marker stained cells were fixed using BD Cytofix/Cytoperm, treated with BD Cytoperm Permeabilization Buffer Plus (BD Biosciences; 561651) by protocol, and fixed again before treatment with 30μg DNase/106 cells for 1 h at 37 °C. Cells were further incubated with fluorescent anti-BrdU prior to acquisition.
Fluorescent activated cell sorting (FACS)
Splenic B220+ CD19+ CD21mid CD23hi naïve follicular B cells were isolated from IgHEL tg mice and splenic B220+ IgMalo IgDa+ anergic B cells were sorted from IgHEL/sHEL double tg mice. Cell populations were surface stained and sorted with a FACS sorter (BD-AriaIII), and collected in ice-cold medium (50% FCS).
RNA-Sequencing (RNA Seq) and Transcriptomic Analysis
RNA data90 are deposited in Gene Expression Omnibus under accession number, GSE135650. DESeq2 v.1.28.1 was used for differential expression analysis between follicular and anergic B cell populations91. Counts were transformed using variance stabilizing transformation for visualization in DESeq2. A gene was considered differentially expressed if the log2 fold change was > = ±1 and adjusted p value <0.05.
Quantitative PCR (qPCR)
RNA was extracted from homogenized murine kidney lysates or 106–107 pelleted cells using the Qiagen RNeasy Plus kit (Qiagen; 74134) or using TRIzol extraction (Life Technologies; 12183555) by protocol. Equivalent quantities of RNA were reverse transcribed to cDNA using SuperScript II Reverse Transcriptase (ThermoFisher Scientific; 18064014), priming with Oligo(dT12–18) primer (ThermoFisher; 18418012). Expression of Ndrg1 and Gapdh mRNA was quantified by qPCR using SYBR Green PowerUp Master Mix (ThermoFisher Scientific; A25776) with the following primers Ndrg1 qPCR forward 5′-ACCCTGAGATGGTAGAGGGTCTC-3′, reverse 5′-CCAATTTAGAATTGCATTCCACC-3′, Gapdh qPCR forward 5′-TGTGTCCGTCGTGGATCTGA-3′, reverse 5′-TTGCTGTTGAAGTCGCAGGAG-3′, and run on a Bio-Rad CFX96 detection system. Standard curve construction demonstrated equivalent efficiencies of Ndrg1 and Gapdh cDNA amplification, therefore changes to Ndrg1 mRNA levels were directly calculated relative to endogenous Gapdh expression using the following equation: 1/(2^(Ndrg1 Ct – Gapdh Ct))*100.
Western blot
Kidney protein lysates were produced by homogenization in lysis buffer containing 50 mM Tris-HCl pH7.4, 0.5%NP40, 150 mM NaCl, 20 mM MgCl2, protease and phosphatase inhibitors (Sigma Aldrich; 4906837001/5892970001), or in supplemented RIPA buffer (Santa Cruz; sc-24948). 1–25 μg protein lysate was reduced by addition of SDS-containing reducing buffer containing 0.1 M DTT and denatured by boiling at 95 °C for 5 min and loaded onto 4–12% Tris-HCl gels (ThermoFisher; NP0323BOX). Proteins were transferred to a nitrocellulose membrane, blocked with 5% BSA or 5% non-fat milk and probed with indicated antibodies. Antibodies; anti-NDRG1 D6C2 mAb (Cell Signaling Technologies; #9408, 1:1000) and anti-rabbit IgG, HRP-linked antibody (Cell Signaling Technology; #7074). Monoclonal anti-β-actin-peroxidase (Sigma; A3854, 1:10000) was used to detect the loading control and ECL prime (Amersham; RPN2232) was used for signal detection.
Immunoprecipitation
Immunoprecipitation of NDRG1 protein was performed using Dynabeads Co-Immunoprecipitation Kit (Invitrogen; 14321D). Following the kit specific protocol, rabbit α-NDRG1 antibody (Abcam; 196621) was conjugated to superparamagnetic Dynabeads M-270 Epoxy beads at 5μg per 1 mg of beads and incubated with protein extracted from freshly isolated tissue for 30 min at 4 °C. Purified NDRG1 was analyzed by Western blot, probed using a primary rabbit α-NDRG1 antibody (Abcam; 196621; 1:500) and secondary peroxidase conjugated α-rabbit IgG antibody (Jackson; 111-035-144, 1:500). Protein was detected with the chemiluminescent ECL Prime Western Blotting Detection Reagent (Cytiva; RPN2232).
ELISA
For HEL-binding IgMa 90 or IgG, plates were coated with 10 μg/ml HEL in carbonate buffer pH9.6 (20 mM Na2CO3, 35 mM NaHCO3), washed with wash buffer (PBS/Tween20), then blocked with PBS, 1%BSA. After washing, serially diluted serum samples were added in PBS/0.1%BSA, after incubation and washing, plates were incubated with 0.5 μg/mL biotinylated anti-IgMa (BD Pharmigen, #553515; clone DS-1) or anti-IgG-HRP (Bethyl laboratories) in 1%FCS, 1% milk powder, 0.1% Tween20 and NaN3 in PBS, then washed. Avidin-alkaline phosphate (A7294) was added to the wells at 1:3000 in PBS/0.1%BSA and incubated, then washed, then plates developed with addition of 1 mg/ml Sigma 104 phosphatase substrate (#104105) in 50 mM Na2CO3, 0.5 mM MgCl2 pH9.8. Anti-IgG-HRP (Bethyl laboratories; A90–131P) was used to detect serum HEL-specific IgG. Absorbance was measured at 405 nm using a Bio-Rad model 550 Microplate reader. Background absorbance values were subtracted from absorbance readings before interpolation to standard curve using Hyperbola (X as concentration). Bethyl laboratories mouse IgG (E90–131), IgM (E90–101) and IgA (E90–103) quantification kits were used by protocol with serum titration of samples as follows: IgG 1:4000, IgM 1:2000, IgA 1:2000, developed using TMB substrate (Life Technologies; 00-4201-56) and detected at 450 nm. IL-2 concentration in the supernatant was quantified by Mouse Ready-Set-Go IL-2 ELISA kit (eBioscience; 88–7024) and developed using TMB substrate as above.
Anti-nuclear antibody (ANA) staining
For ANA staining, serum samples were diluted at 1:100 in PBS and stored at −20 °C before incubation on Hep-2 cell coated slides (A.Menarini; 37806). Slides were washed with PBS, then water, then incubated with FITC-conjugated, goat anti-mouse IgG (ThermoFisher; 62–6511). After washing, slides were mounted and imaged on Nikon wide-field TE20000U Microscope (GFP channel: 20×, 400 ms) then analysed with automated cell segmentation and fluorescence intensity quantification with ImageJ. Positive scoring samples (2+) were manually cross-checked before scoring92.
In vivo T cell anergy
CD45.2+ CD4+ T cells were isolated from WT OT-II+ or Ndrg1−/− OT-II+ tg mice and adoptively transferred to CD45.1+ non-tg congenic recipients, with subsequent intravenous injection of 500μg endotoxin-free OVA peptide on day 1 (Insight Biotech; 21-51023-G (323–339)), as tested using Pierce LAL Chromogenic Endotoxin Quantitation kit (ThermoFisher; #88282). 5 days later, total splenic CD4+ cells were magnetically isolated from recipient mice and naïve unstimulated WT or Ndrg1−/− OT-II+ control mice, before loading with CTV dye. As determined by flow cytometry, 4 × 104 CD45.2+ Vα2+ OT-II+ CD4+ T cells were cultured in vitro with a titration of OVA peptide and 4 × 105 irradiated splenocytes. IL-2 in culture supernatant at 48 h was measured by ELISA and proliferation of CD45.2+Vα2+ OT-II+ CD4+ T cells was detected by flow cytometry at 72 h.
Statistics and reproducibility
GraphPad Prism Software was used for statistical analyses and unless otherwise specifically mentioned unpaired, two-tailed Student’s t tests were used for statistical comparison between groups, correcting for multiple comparisons using the Holm-Sidak method. Experimental groups were determined by genotype and were therefore not randomized and were not blinded. Experimental animals were not excluded from analysis except according to pre-specified experimental design on the basis of failed chimeric reconstitution or failed cell transfer. All experiments included age and sex matched controls, which were co-housed littermates wherever possible. Sample sizes were selected on the basis of previously published studies. Data shown is pooled from, or representative of replicate experiments as indicated in figure legends.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Raw RNA-Seq data are available in the Gene Expression Omnibus (https://www.ncbi.nlm.nih.gov/geo/) under accession code GSE135650. Supplementary Table 1 is provided in the Supplementary Data. Uncropped and unedited gel and blot images are available within Supplementary Fig. 4a–d. All other source data are available via the Oxford University Research Archive at https://ora.ox.ac.uk/objects/uuid:51f425ad-836f-4c3e-91d6-3321b0ab7b8b with the https://doi.org/10.5287/bodleian:gJnGGXymz.
References
Nossal, G. J. V. & Pike, B. L. Clonal anergy: persistence in tolerant mice of antigen-binding B lymphocytes incapable of responding to antigen or mitogen. Proc. Natl Acad. Sci. USA. 77, 1602–1606 (1980).
Schwartz, R. H., Mueller, D. L., Jenkins, M. K. & Quill, H. T-cell clonal anergy. Cold Spring Harb. Symp. Quant. Biol. 54, 605–610 (1989).
Ferry, H. et al. B-cell tolerance. Transplantation 81, 308–315 (2006).
Schwartz, R. H. T cell anergy. Annu. Rev. Immunol. 21, 305–334 (2003).
Bretscher, P. & Cohn, M. A theory of self-nonself discrimination. Science 169, 1042–1049 (1970).
Baxter, A. G. & Hodgkin, P. D. Activation rules: the two-signal theories of immune activation. Nat. Rev. Immunol. 2, 439–446 (2002).
Bretscher, P. A. A two-step, two-signal model for the primary activation of precursor helper T cells. Proc. Natl. Acad. Sci. USA 96, 185–190 (1999).
Yarkoni, Y., Getahun, A. & Cambier, J. C. Molecular underpinning of B-cell anergy. Immunol. Rev. 237, 249–263 (2010).
Goodnow, C. C. et al. Altered immunoglobulin expression and functional silencing of self-reactive B lymphocytes in transgenic mice. Nature 334, 676–682 (1988).
Cooke, M. P. et al. Immunoglobulin signal transduction guides the specificity of B cell-T cell interactions and is blocked in tolerant self-reactive B cells. J. Exp. Med. 179, 425–438 (1994).
Borde, M., Barrington, R. A., Heissmeyer, V., Carroll, M. C. & Rao, A. Transcriptional basis of lymphocyte tolerance. Immunol. Rev. 210, 105–119 (2006).
Barrington, R. A., Borde, M., Rao, A. & Carroll, M. C. Involvement of NFAT1 in B cell self-tolerance. J. Immunol. 177, 1510–1515 (2006).
Healy, J. I. et al. Different nuclear signals are activated by the B cell receptor during positive versus negative signaling. Immunity 6, 419–428 (1997).
Healy, J. I. & Goodnow, C. C. Positive versus negative signaling by lymphocyte antigen receptors. Annu. Rev. Immunol. 16, 645–670 (1998).
Weintraub, B. C. et al. Entry of B cell receptor into signaling domains is inhibited in tolerant B cells. J. Exp. Med. 191, 1443–1448 (2000).
Rui, L., Healy, J. I., Blasioli, J. & Goodnow, C. C. ERK signaling is a molecular switch integrating opposing inputs from B cell receptor and T cell cytokines to control TLR4-driven plasma cell differentiation. J. Immunol. 177, 5337–5346 (2006).
Lesley, R. et al. Reduced competitiveness of autoantigen-engaged B cells due to increased dependence on BAFF. Immunity 20, 441–453 (2004).
Enders, A. et al. Loss of the pro-apoptotic BH3-only Bcl-2 family member Bim inhibits BCR stimulation-induced apoptosis and deletion of autoreactive B cells. J. Exp. Med. 198, 1119–1126 (2003).
Cyster, J. G., Hartley, S. B. & Goodnow, C. C. Competition for follicular niches excludes self-reactive cells from the recirculating B-cell repertoire. Nature 371, 389–395 (1994).
Fulcher, D. A. et al. The fate of self-reactive B cells depends primarily on the degree of antigen receptor engagement and availability of T cell help. J. Exp. Med. 183, 2313–2328 (1996).
Cook, M. C., Basten, A. & Fazekas De St. Groth, B. Rescue of self-reactive B cells by provision of T cell help in vivo. Eur. J. Immunol. 28, 2549–2558 (1998).
Perez, V. L. et al. Induction of peripheral T cell tolerance in vivo requires CTLA-4 engagement. Immunity 6, 411–417 (1997).
Beverly, B., Kang, S. M. O., Lenardo, M. J. & Schwartz, R. H. Reversal of in vitro T cell clonal anergy by IL-2 stimulation. Int. Immunol. 4, 661–671 (1992).
Chiodetti, L., Choi, S., Barber, D. L. & Schwartz, R. H. Adaptive tolerance and clonal anergy are distinct biochemical states. J. Immunol. 176, 2279–2291 (2006).
Sabouri, Z. et al. IgD attenuates the IgM-induced anergy response in transitional and mature B cells. Nat. Commun. 7, 13381 (2016).
Zheng, Y., Zha, Y., Driessens, G., Locke, F. & Gajewski, T. F. Transcriptional regulator early growth response gene 2 (Egr2) is required for T cell anergy in vitro and in vivo. J. Exp. Med. 209, 2157–2163 (2012).
Oh, Y. M. et al. Ndrg1 is a T-cell clonal anergy factor negatively regulated by CD28 costimulation and interleukin-2. Nat. Commun. 6, 8698 (2015).
Van Belzen, N. et al. A novel gene which is up-regulated during colon epithelial cell differentiation and down-regulated in colorectal neoplasms. Lab. Investig. 77, 85–92 (1997).
Ellen, T. P., Ke, Q., Zhang, P. & Costa, M. NDRG1, a growth and cancer related gene: Regulation of gene expression andfunction in normal and disease states. Carcinogenesis 29, 2–8 (2008).
Fang, B. A. et al. Molecular functions of the iron-regulated metastasis suppressor, NDRG1, and its potential as a molecular target for cancer therapy. Biochim. Biophys. Acta - Rev. Cancer 1845, 1–19 (2014).
Sun, J. et al. Metastasis suppressor, NDRG1, mediates its activity through signaling pathways and molecular motors. Carcinogenesis 34, 1943–1954 (2013).
Lachat, P., Shaw, P., Gebhard, S., Nico van Belzen, P. C. & Bosman, F. T. Expression of NDRG1, a differentiation-related gene, in human tissues. Histochem. Cell Biol. 118, 399–408 (2002).
Kalaydjieva, L. et al. N-myc downstream-regulated gene 1 is mutated in hereditary motor and sensory neuropathy-Lom. Am. J. Hum. Genet. 67, 47–58 (2000).
Okuda, T. et al. Ndrg1-deficient mice exhibit a progressive demyelinating disorder of peripheral nerves. Mol. Cell. Biol. 24, 3949–3956 (2004).
Chang, X. et al. NDRG1 controls gastric cancer migration and invasion through regulating MMP-9. Pathol. Oncol. Res. 22, 789–796 (2016).
Bandyopadhyay, S. et al. Role of the putative tumor metastasis suppressor gene Drg-1 in breast cancer progression. Oncogene 23, 5675–5681 (2004).
Angst, E. et al. N-myc downstream regulated gene-1 expression correlates with reduced pancreatic cancer growth and increased apoptosis in vitro and in vivo. Surgery 149, 614–624 (2011).
Bandyopadhyay, S. et al. The Drg-1 gene suppresses tumor metastasis in prostate cancer. Cancer Res. 63, 1731–1736 (2003).
Kovacevic, Z., Chikhani, S., Lovejoy, D. B. & Richardson, D. R. Novel thiosemicarbazone iron chelators induce up-regulation and phosphorylation of the metastasis suppressor N-myc down-stream regulated gene 1: a new strategy for the treatment of pancreatic cancer. Mol. Pharmacol. 80, 598–609 (2011).
Le, N. T. V. & Richardson, D. R. Iron chelators with high antiproliferative activity up-regulate the expression of a growth inhibitory and metastasis suppressor gene: a link between iron metabolism and proliferation. Blood 104, 2967–2975 (2004).
Lu, W. J., Chua, M. S. & So, S. K. Suppressing N-Myc downstream regulated gene 1 reactivates senescence signaling and inhibits tumor growth in hepatocellular carcinoma. Carcinogenesis 35, 915–922 (2014).
Ring, B. Z. et al. Novel prognostic immunohistochemical biomarker panel for estrogen receptor-positive breast cancer. J. Clin. Oncol. 24, 3039–3047 (2006).
Li, A. et al. Upregulation of NDRG1 predicts poor outcome and facilitates disease progression by influencing the EMT process in bladder cancer. Sci. Rep. 9, 1–12 (2019).
Chua, M. S. et al. Overexpression of NDRG1 is an indicator of poor prognosis in hepatocellular carcinoma. Mod. Pathol. 20, 76–83 (2007).
Zhao, T., Meng, Y., Wang, Y. & Wang, W. NDRG1 regulates osteosarcoma cells via mediating the mitochondrial function and CSCs differentiation. J. Orthop. Surg. Res. 16, 364 (2021).
Luo, Z. et al. MiR-188-3p and miR-133b suppress cell proliferation in human hepatocellular carcinoma via post-transcriptional suppression of NDRG1. Technol. Cancer Res. Treat. 20, 1–9 (2021).
Merrell, K. T. et al. Identification of anergic B cells within a wild-type repertoire. Immunity 25, 953–962 (2006).
Powell, J. D., Ragheb, J. A., Kitagawa-Sakakida, S. & Schwartz, R. H. Molecular regulation of interleukin-2 expression by CD28 co-stimulation and anergy. Immunol. Rev. 165, 287–300 (1998).
Watari, K. et al. Impaired differentiation of macrophage lineage cells attenuates bone remodeling and inflammatory angiogenesis in Ndrg1 deficient mice. Sci. Rep. 6, (2016).
Vinuesa, C. G. & Goodnow, C. C. Illuminating autoimmune regulators through controlled variation of the mouse genome sequence. Immunity 20, 669–679 (2004).
Stein, S. et al. NDRG1 is necessary for p53-dependent apoptosis. J. Biol. Chem. 279, 48930–48940 (2004).
Cyster, J. G. & Goodnow, C. C. Antigen-induced exclusion from follicles and anergy are separate and complementary processes that influence peripheral B cell fate. Immunity 3, 691–701 (1995).
Rathmell, J. C. et al. CD95 (Fas)-dependent elimination of self-reactive B cells upon interaction with CD4+T cells. Nature 376, 181–184 (1995).
Kearney, E. R., Pape, K. A., Loh, D. Y. & Jenkins, M. K. Visualization of peptide-specific T cell immunity and peripheral tolerance induction in vivo. Immunity 1, 327–339 (1994).
Young, A., Quandt, Z. & Bluestone, J. A. The balancing act between cancer immunity and autoimmunity in response to immunotherapy. Cancer Immunol. Res. 6, 1445–1452 (2018).
Ramos-Casals, M. et al. Immune-related adverse events of checkpoint inhibitors. Nat. Rev. Dis. Prim. 6, 38 (2020).
Killestein, J. & van Oosten, B. Emerging safety issues in alemtuzumab-treated MS patients. Mult. Scler. J. 25, 1206–1208 (2019).
Macián, F. et al. Transcriptional mechanisms underlying lymphocyte tolerance. Cell 109, 719–731 (2002).
Pape, K. A., Merica, R., Mondino, A., Khoruts, A. & Jenkins, M. K. Direct evidence that functionally impaired CD4+ T cells persist in vivo following induction of peripheral tolerance. J. Immunol. 160, 4719–4729 (1998).
Koniaras, C., Heath, W. R. & Lew, A. M. T cell unresponsiveness in vitro can be due to activation in vivo. Int. Immunol. 10, 365–369 (1998).
Bygrave, A. E. et al. Spontaneous autoimmunity in 129 and C57BL/6 mice-implications for autoimmunity described in gene-targeted mice. PLoS Biol. 2, e243 (2004).
Zhou, R. H. et al. Characterization of the human NDRG gene family: a newly identified member, NDRG4, is specifically expressed in brain and heart. Genomics 73, 86–97 (2001).
Qu, X. et al. Characterization and expression of three novel differentiation-related genes belong to the human NDRG gene family. Mol. Cell. Biochem. 229, 35–44 (2002).
Okuda, T., Kokame, K. & Miyata, T. Differential expression patterns of NDRG family proteins in the central nervous system. J. Histochem. Cytochem. 56, 175–182 (2007).
Pao, L. I., Famiglietti, S. J. & Cambier, J. C. Asymmetrical phosphorylation and function of immunoreceptor tyrosine-based activation motif tyrosines in B cell antigen receptor signal transduction. J. Immunol. 160, 3305–3314 (1998).
Smith, M. J. et al. Elevated PTEN expression maintains anergy in human B cells and reveals unexpectedly high repertoire autoreactivity. JCI Insight 4, e123384 (2019).
Getahun, A., Beavers, N. A., Larson, S. R., Shlomchik, M. J. & Cambier, J. C. Continuous inhibitory signaling by both SHP-1 and SHIP-1 pathways is required to maintain unresponsiveness of anergic B cells. J. Cell Biol. 213, 2133OIA94 (2016).
Kovacevic, Z., Chikhani, S., Lui, G. Y. L., Sivagurunathan, S. & Richardson, D. R. The iron-regulated metastasis suppressor NDRG1 targets NEDD4L, PTEN, and SMAD4 and inhibits the PI3K and ras signaling pathways. Antioxid. Redox Signal. 18, 874–887 (2013).
Chung, I. C. et al. Src-family kinase-Cbl axis negatively regulates NLRP3 inflammasome activation. Cell Death Dis. 9, 1109 (2018).
Jeon, M. S. et al. Essential role of the E3 ubiquitin ligase Cbl-b in T cell anergy induction. Immunity 21, 167–177 (2004).
Kitaura, Y. et al. Control of the B cell-intrinsic tolerance programs by ubiquitin ligases Cbl and Cbl-b. Immunity 26, 567–578 (2007).
Taketomi, Y. et al. Identification of NDRG1 as an early inducible gene during in vitro maturation of cultured mast cells. Biochem. Biophys. Res. Commun. 306, 339–346 (2003).
Taketomi, Y. et al. Impaired mast cell maturation and degranulation and attenuated allergic responses in Ndrg1-deficient mice. J. Immunol. 178, 7042–7053 (2007).
Parman, Y. et al. Clinicopathological and genetic study of early-onset demyelinating neuropathy. Brain 127, 2540–2550 (2004).
Echaniz-Laguna, A. et al. NDRG1-linked Charcot-Marie-Tooth disease (CMT4D) with central nervous system involvement. Neuromuscul. Disord. 17, 163–168 (2007).
Okamoto, Y. et al. Exonic duplication CNV of NDRG1 associated with autosomal-recessive HMSN-Lom/CMT4D. Genet. Med. 16, 386–394 (2014).
Zhang, P., Tchou-Wong, K. M. & Costa, M. Egr-1 mediates hypoxia-inducible transcription of the NDRG1 gene through an overlapping Egr-1/Sp1 binding site in the promoter. Cancer Res. 67, 9125–9133 (2007).
Collins, S. et al. Opposing regulation of T cell function by Egr-1/NAB2 and Egr-2/Egr-3. Eur. J. Immunol. 38, 528–536. (2008)
Burrows, N. et al. Dynamic regulation of hypoxia-inducible factor-1α activity is essential for normal B cell development. Nat. Immunol. 21, 1408–1420 (2020).
Meng, X. et al. Hypoxia-inducible factor-1α is a critical transcription factor for IL-10-producing B cells in autoimmune disease. Nat. Commun. 9, 251 (2018).
Chen, J., Holguin, N., Shi, Y., Silva, M. J. & Long, F. mTORC2 signaling promotes skeletal growth and bone formation in mice. J. Bone Miner. Res. 30, 369–378 (2015).
Lee, K. et al. Requirement for Rictor in homeostasis and function of mature B lymphoid cells. Blood 122, 2369–2379 (2013).
Merlot, A. M. et al. The metastasis suppressor, NDRG1, differentially modulates the endoplasmic reticulum stress response. Biochim. Biophys. Acta Mol. Basis Dis. 1865, 2094–2110 (2019).
Sahni, S. et al. The metastasis suppressor, N-myc Downstream-regulated gene 1 (NDRG1), inhibits stress-induced autophagy in cancer cells. J. Biol. Chem. 289, 9692–9709 (2014).
Piquemal, D. et al. Differential expression of the RTP/Drg1/Ndr1 gene product in proliferating and growth arrested cells. Biochim. Biophys. Acta Mol. Cell Res. 1450, 364–373 (1999).
Barnden, M. J., Allison, J., Heath, W. R. & Carbone, F. R. Defective TCR expression in transgenic mice constructed using cDNA- based α- and β-chain genes under the control of heterologous regulatory elements. Immunol. Cell Biol. 76, 34–40 (1998).
Hsu, P. D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Anzilotti, C. et al. An essential role for the Zn 2+ transporter ZIP7 in B cell development. Nat. Immunol. 20, 350–361 (2019).
Xu, X. et al. An ontogenetic switch drives the positive and negative selection of B cells. Proc. Natl Acad. Sci. U.S.A. 117, 3718–3727 (2020).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Abeler-Dörner, L. et al. High-throughput phenotyping reveals expansive genetic and structural underpinnings of immune variation. Nat. Immunol. 21, 86–100 (2020).
Acknowledgements
We thank N. Ashley, P. Sopp, and C. Waugh for assistance with flow cytometry sorting and staff at the Oxford Functional Genomics Facility for animal care. This work was supported by the Medical Research Council (MRC) Human Immunology Unit. RH is supported by a MRC student fellowship and St John’s College, University of Oxford. KRB is supported by the MRC and Kidney Research, UK. JDK is supported by a Novo Nordisk postdoctoral fellowship.
Author information
Authors and Affiliations
Contributions
Experimental work was performed by: R.H., X.X., T.L.C., J.D.K., E.C., A.B., A.C.S., and M.J.W. The Ndrg1−/− mouse model was designed and created by: C.A., A.C.S., and B.D. Data analysis and interpretation were performed by: R.H., X.X., C.A., M.D.L., R.J.C., and K.R.B. Scientific writing and paper revision were performed by: R.H., C.A., M.D.L., R.J.C., and K.R.B.
Corresponding authors
Ethics declarations
Competing interests
The authors have declared that no competing interests exists, R.J.C. is a founder and shareholder in MIRO-Bio Ltd, K.R.B. is a consultant to D.J.S. antibodies Ltd.
Peer review
Peer review information
Communications Biology thanks Hong Zan, Georgios Sogkas and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Guideng Li and Manuel Breuer.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hodgson, R., Xu, X., Anzilotti, C. et al. NDRG1 is induced by antigen-receptor signaling but dispensable for B and T cell self-tolerance. Commun Biol 5, 1216 (2022). https://doi.org/10.1038/s42003-022-04118-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-022-04118-w
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.