Abstract
Simultaneous longitudinal imaging across multiple conditions and replicates has been crucial for scientific studies aiming to understand biological processes and disease. Yet, imaging systems capable of accomplishing these tasks are economically unattainable for most academic and teaching laboratories around the world. Here, we propose the Picroscope, which is the first low-cost system for simultaneous longitudinal biological imaging made primarily using off-the-shelf and 3D-printed materials. The Picroscope is compatible with standard 24-well cell culture plates and captures 3D z-stack image data. The Picroscope can be controlled remotely, allowing for automatic imaging with minimal intervention from the investigator. Here, we use this system in a range of applications. We gathered longitudinal whole organism image data for frogs, zebrafish, and planaria worms. We also gathered image data inside an incubator to observe 2D monolayers and 3D mammalian tissue culture models. Using this tool, we can measure the behavior of entire organisms or individual cells over long-time periods.
Similar content being viewed by others
Introduction
Monitoring and handling live tissues and cell cultures as well as analyzing their secreted contents are essential tasks in experimental biology and biomedicine. Advances in microscopy have revolutionized biological studies, allowing scientists to perform observations of cellular processes and organisms’ development and behaviors. Imaging has been pivotal to uncovering cellular mechanisms behind biological processes1. Several options exist on the market to perform longitudinal imaging of biological materials. These range from super-resolution microscopes, that allow the imaging of individual biomolecules2,3, to conventional benchtop microscopes, which are common in academic research3,4,5,6,7, industrial8,9, and teaching laboratories10. When deciding between the different technologies for longitudinal live tissue imaging, several factors need to be considered in the experimental design. The image acquisition speed of the microscope should be sufficient for the phenomenon being studied. The microscope should be able to acquire images without damaging or disturbing the specimen, such as photobleaching. The microscope should be capable of imaging in the environmental conditions needed for the desired experiment, including temperature, light, and humidity. The resolution of the microscope should be sufficient to view the phenomenon being studied. When scaling to simultaneous multi-well longitudinal tissue imaging it is also important that the apparatus not be bulky or expensive. It has been challenging to meet all of these criteria11.
The use of open-source technology, including 3D printers, laser cutters, and low-cost computer hardware, has democratized access to rapid prototyping tools and dramatically increased the repertoire of biomedical equipment available to laboratories around the world12,13. Through rapid prototyping and the use of open-source platforms, the technology can be replicated and quickly improved14,15. 3D printer technology has been applied to several fields in biomedicine, including biotechnology16, bioengineering17,18, and medical applications including fabrication of tissues and organs, casts, implants, and prostheses19. Existing 3D printed microscopes range in complexity from simple low-cost systems with pre-loaded imaging modules20 to portable confocal microscopes capable of imaging individual molecules21 and even 3D printed microfluidic bioreactors22.
The majority of low-cost 3D printed microscopes are not intended for longitudinal imaging of simultaneous biological cultures (e.g., multi-well, multi-week biological experiments). They usually have a single imaging unit5,17,23,24,25,26,27,28,29,30,31,32 or perform confocal,21 and even light-sheet imaging25. Other systems have taken advantage of one camera attached to a gantry system to perform imaging of multiple experimental replicates33,34,35. Few 3D-printed microscopes have been developed that perform multi-well imaging with medium throughput34,36. Several biological applications exist that would greatly benefit from multi-well, multi-week simultaneous imaging, as it allows for concurrent interrogation of different experimental conditions and the inclusion of biological replicates. These include cell culture applications, in which 2D and 3D culture models can be tracked over multi-week periods, as well as developmental and behavioral biology experiments in which multi-week tracking could be performed on whole organisms.
Here, we report a simultaneous multi-well imaging system (the Picroscope), which features a low-cost per well ($83) and performs longitudinal brightfield z-stack imaging of 24-well cell culture plates. Images are uploaded to a server as they are captured allowing the users to view the results in near real time. We used this system to longitudinally track different animal models of development and regeneration, including Xenopus tropicalis (frogs), Danio rerio (zebrafish), and planaria worms. Finally, we demonstrate this system’s versatility by imaging human embryonic stem cells and 3D cortical organoids inside a standard tissue culture incubator. We demonstrate that the Picroscope is a robust low-cost, versatile multi-well imaging system for longitudinal live imaging biological studies.
Results
System design
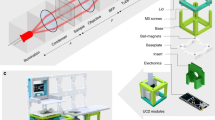
The Picroscope is a programmable, data rich, sensor-per-well simultaneous imaging system for longitudinal brightfield imaging to automate microscopy (Fig. 1a). The system simultaneously images in each one of the 24 wells multiple focal planes (the resolution of the “z-stack” can be remotely changed) several times every hour for weeks, a frequency impractical to perform manually. The instrument is made using off-the-shelf components (e.g., lenses, motors, cameras, Arduino, Raspberry Pi, and MakerBeam aluminum extrusions), and 3D printed Polylactic acid (PLA) components with 100% infill (the percentage of the internal part of the piece that is occupied by the printing material). Cost comparison with other open-source microscope projects can be found in Table 1. A cost breakdown of the materials required can be found in Table 2. The Picroscope has been used to image Planaria worms (Fig. 1b), Xenopus tropicalis (Fig. 1c), as well as zebrafish (Fig. 1d).
a The Picroscope fits a standard 24-well plate, it is controlled remotely and images can be accessed through a web browser. b–d Applications of the Picroscope to longitudinal imaging of developmental biology and regeneration. b Regeneration of planaria worms Dugesia tigrina. c Zebrafish embryonic development at oblong stage. d Zebrafish embryo at 48 hours post fertilization. In complement see Supplementary Video 1.
The system was developed to be operated remotely through the internet. Users can set and change the device settings to modify experiments on the fly. Images captured by the system are uploaded to a server where they become visible on a viewer website. We have also created several image analyses scripts that can directly access images on the server, allowing us to generate timelapse videos and composite images in an automated fashion. While the system receives commands and transmits results through the Internet, it is also capable of running on a Local Area Network (LAN) if internet access is not available. Figure 2 shows the basic workflow from control console to image viewer. Further details about the software and network architecture developed to implement these features can be found in37.
The Picroscope is designed to illuminate the samples using one or multiple lighting sources (from above or below a standard 24-well cell culture plate) (Fig. 3). Diffused illumination from below results in images that show contours and surface features, this is particularly useful when the sample is opaque. Illumination from above typically works best for samples that are sufficiently translucent and can show internal structures as the light can pass through the sample. The flexibility of using different illumination techniques emulates commercial brightfield microscopes. The difference from over and under light is best shown in Supplementary Fig. 1.
a Physical representation of the proposed imaging system. b One line of independent cameras. c An integrated rack of cameras and Raspberry Pi board computers. d The interlacing strategy of four independent racks of power distribution boards. e The Raspberry Pi Hub and Arduino Uno, Motor driver and custom relay board. f The XY adjustment stage. 1 = Over-the-plate illumination board, 2 = 3D printed Cell Culture Plate Holder and XY stage, 3 = Lenses, 4 = Illumination Board from below, 5 = 3D Printed Camera Bodies, 6 = 3D Printed Elevator, 7 = Raspberry Pi Zero W, 8 = Motors, 9 = Base, 10 = Raspberry Spy Cameras, 11 = Interface Board a. row 1, b. rows 2 and 3c. row 4, 12 = Pi Hub -- Raspberry Pi 4, 13 = Custom Relay Board, 14 = Adafruit Motor/Stepper/Servo Shield for Arduino v2, 15 = Arduino Uno, 16 = Leaf Springs, 17 = Rigid Elements, 18 = Relays, 19 = Limit switches connectors, 20 = Power distribution board connectors, 21 = Light board connectors, 22 = Motor power connector, 23 = 12 V power source, 24 = Voltage regulators, and 25 = Temperature & Humidity sensor.
The 3D printed plate holder (2 in Fig. 3) supports the biological sample during an experiment. For easy alignment, the holder is attached to a xy sliding stage that consists of two interconnected linear stages (Fig. 3f). The inner stage translating along the y-axis uses 8 leaf springs to connect a central piece holding the 24-well plate with four rigid elements surrounding it. The outer stage translating along the x-axis uses 8 additional leaf springs to connect the inner stage with the outside 4 rigid elements, two of them being connected to the Picroscope frame using 4 screws (18 in Fig. 3). While each stage is flexible along one axis (x or y), together they can slide along both, x and y axes. Each stage is actuated by two adjustment screws depicted as gray arrows in Fig. 3f.
The imaging unit consists of 24 independent objectives attached to a vertical sliding stage (“elevator piece” in 3a) using 4 makerbeam vertical columns and 2 Nema-11 stepper motors (3.175 μm Travel/Step), an example of a row can be seen in Fig. 4a. The fine threads are necessary for focusing on specific biological features and collecting z-stack imaging (Fig. 4b). With this fixed lens system, the system has a field of view of approximately 5 mm. The Picroscope is able to resolve Group 7, Element 1 targets (1951 USAF Targets), corresponding to a resolution of 7 μm (Supplementary Fig. 2). If higher resolution is needed the lens can be swapped out for more magnification (Table 1). The lens currently on the system was chosen due to our interest in imaging whole organisms. The objectives are distributed on 4-rows and 6-columns to match a standard 24-well culture plate. Each objective consists of a 3D printed camera body that hosts a 5 MegaPixel (5MP) camera (Spy Camera for Raspberry Pi Zero W, with a 1.4 μm × 1.4 μm pixel pitch) and an off-the-shelf Arducam 1/2” M12 Mount 16 mm Focal Length. Each objective is controlled by a single-board computer (Raspberry Pi Zero W), which is connected to an individual slot on one of the three custom-made power distribution boards (3c and D). All 24 single-board computers (Raspberry Pi Zero W) computers communicate to a hub board computer (Raspberry Pi 4) that manages the images and autonomously uploads them to a remote server. The hub single-board computer has the MIPI CSI-2 camera port and is connected to an Arduino Uno, which has a motor shield attachment, to control the motors and lift the elevator piece (3e). As a safety feature, the system also includes a custom-made Relay Board that is attached to the Arduino and motor driver stack. The relay board provides control of the LED boards and in the event of an overheat allows us to shut down the system, protecting the system and the biological sample. After each set of pictures, the imaging unit returns to the lowest (“park”) position, which is determined by a limiting switch attached to the elevator unit. The entire system sits on a 3D printed base, that includes a fan for heat dissipation. Supplementary Fig. 3 shows thermal images of the Picroscope to demonstrate that heat from the system does not impact the experiment. A guide on how to assemble the Picroscope and components needed can be found in Table 2 and Supplementary Note 1. During the course of an experiment, the pictures are autonomously uploaded on a remote computer/server using the ethernet connection of the hub computer board, where they can be viewed or processed in near real time (see Fig. 5).
a A single row of cameras demonstrating the z-stack function. 1.a = Over-the-plate illumination board, 1.b = Under-the-plate illumination board, 2 = Acrylic Light Diffuser, 3 = Lenses, 4 = Cell Culture Plate, 5 = LEDs, 6 = Raspberry Spy Cameras, 7 = 3D Printed Camera Bodies, 8 = Biological Sample (e.g., Frog Embryos), 9 = Individual Culture Well. b Four focal planes of a single z-stack. These photos at were taken at four planes, 0.3 mm apart. The blastopore is only in focus in Plane 1.
Longitudinal imaging of Xenopus tropicalis embryonic development
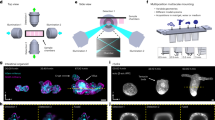
As proof of principle of the longitudinal live imaging capabilities of the Picroscope, we imaged the development of Xenopus tropicalis embryos from the onset of gastrulation through organogenesis (Figs. 5b, 6, and 7). The fertilization and development of Xenopus occur entirely externally, which allows scientists to easily observe and manipulate the process38. For decades, Xenopus have been heavily used in biology studies to model a variety of developmental processes and early onset of diseases, particularly those of the nervous system39. While several species of Xenopus are used in different laboratories around the world, Xenopus tropicalis is one of the preferred species due to its diploid genomic composition and fast sexual maturation40,41. Normal development and optimal husbandry of Xenopus tropicalis occur at 25∘–27 ∘C,42,43 closely approximating standard room temperature, which eliminates the need of special environmental control for most experiments.
a The images shown in Fig. 6 were taken hourly over a 28 h period and encompass three developmental stages: Gastrulation, neurulation, and organogenesis. Y-Axis represents the stages of frog embryonic development: 1 = Fertilization, 2 = Cleavage, 3 = Gastrulation, 4 = Neurulation, 5 = Organogenesis, and 6 = Metamorphosis. X-axis represents the timepoint at which it occurs. Each dot in the plot represents a timepoint in which the images were taken. Magenta = the beginning of each developmental process. Red = the end of the experiment at 28 h. Blue = intermediate timepoints. b Diameter of the blastopore is reduced over time from gastrulation to neurulation. Top right-hand panel shows an example of an individual blastopore. A total of 27 embryos were considered for the analyses. Error bars represent Standard Deviation (SD).
Given these convenient experimental advantages and their large size, Xenopus embryos have been used extensively to understand the development of the vertebrate body plan, with particular success in elaborating the complex cellular rearrangements that occur during gastrulation and neural tube closure44,45. These experiments rely on longitudinal imaging of developing embryos, often at single-embryo scale with dyes, fluorescent molecules, and computational tracking of single cells45,46. These studies have elucidated key cellular mechanical properties and interactions critical to vertebrate development, often replayed and co-opted during tumorigenesis. There exists an opportunity to scale these experiments to be more high-throughput with the Picroscope, as one could image hundreds of developing embryos simultaneously, rather than having to move the objective from embryo to embryo during development, or repeating the experiment many times.
We imaged Xenopus tropicalis embryos over a 28 h time period. Four embryos were placed in each of the 23 wells used in a 24-well plate, and we used an extra well as calibration (Figs. 5b and 6). The embryos were grown in simple saline solution and the experiment took place at room temperature. Imaging was performed hourly starting at gastrulation (Fig. 7). Then, we visually inspected each image and mapped the embryos to the standard stages of frog development, categorizing their development in gastrulation, neurulation, and organogenesis (Fig. 7a). Finally, we took a subset of 27 embryos and measured the diameter of the blastopore as the embryos underwent gastrulation (Fig. 7b; Supplementary Data 1). Only 27 embryos were used because those were the only embryos with their blastopores clearly visible throughout the image set. We observed a progressive reduction of blastopore diameter over a 6 h time period, consistent with progression through gastrulation and the start of neurulation. This simple experiment demonstrated that the Picroscope can be used for longitudinal sequential imaging and tracking of biological systems.
In-incubator imaging of human embryonic stem cells and brain organoids
While many biological systems including zebrafish, planaria, and frogs develop at room temperature and atmospheric gas concentrations, mammalian models require special conditions requiring an incubator enclosure. Mammalian models include 2D monolayer cell cultures, as well as 3D organoid models of development and organogenesis47. They have been used to assess molecular features and effects of drugs for a variety of phenotypes including cell proliferation48,49, morphology50,51, and activity52,53, among others.
Deploying electronics and 3D printed materials inside tissue culture incubators presents some unique challenges. The temperature and humidity conditions can cause electronics to fail and cause certain plastics to offgas toxins54. Plastics can also be prone to deformation in these conditions. A common solution for protecting electronics and preventing offgassing is to use inert protective coatings e.g., Parylene C. This requires expensive clean room equipment. Instead, we print all of the components with PLA, a non-toxic and biodegradable material, to prevent deformation we print using 100% infill and reinforce vulnerable elements with aluminum MakerBeam profiles. We coat all electronic components with Corona Super Dope Coating to protect the electronics from the conditions (heat and humidity) of an incubator.
We tested the functionality of the Picroscope inside a standard tissue culture incubator by imaging 2D-monolayers of human embryonic stem cells (hESCs) (Figs. 8a,b). To demonstrate the capacity of our system to perform longitudinal imaging across the z-axis, we imaged human cortical organoids embedded in Matrigel (Fig. 8c). Using this system, we could monitor and measure the growth of the organoids over 86 h (Fig. 8d). Tracking of individual cells within organoid outgrowths allowed us observe their migration patterns and behavior (Fig. 8e). Altogether, we show the feasibility of using our system for longitudinal imaging of mammalian cell and organoid models.
a The Picroscope inside a standard tissue culture incubator. b Imaging of human embryonic stem cells as a model of 2D-monolayer cell cultures. c Longitudinal imaging of human cortical organoids embedded in Matrigel. Zoomed images show cellular outgrowths originating in the organoids. d Tracking of cortical organoid development over 86 h. Images were taken hourly. On left. Images of the tracked organoid at timepoints 0, 43, and 86. On right. Measurement of organoid area at each timepoint analyzed. e Manual Longitudinal tracking of individual cells in embedded cortical organoids over 40 min. Images were taken every 10 min. Magenta = example of cell division, Red = example of cell migration, and Purple = example of morphological changes.
Discussion
The combination of 3D printed technology and open-source software has significantly increased the accessibility of academic and teaching laboratories to biomedical equipment55. Thermocyclers, for example, were once an expensive commodity unattainable for many laboratories around the world56,57. Now, low-cost thermocyclers have been shown to perform as well as high-end commercially available equipment58. Inexpensive thermocyclers can be used in a variety of previously unimaginable contexts, including conservation studies in the Amazon59, diagnostics of Ebola, Zika and SARS-CoV-259,60, teaching high-school students in the developing world10 and epigenetic studies onboard the International Space Station61.
Simultaneous imaging of biological systems is crucial for drug discovery, genetic screening, and high-throughput phenotyping of biological processes and disease48,49,51. This technique typically requires expensive multi-camera and robotic equipment, making it inaccessible to most. While the need for a low-cost solution has long been appreciated62, few solutions have been proposed. Currently, the low-cost solutions can be grouped in two categories: (1) those that use gantry systems that move an individual camera through multiple wells, performing “semi-simultaneous” imaging33,34,35 or (2) those that use acquisition of large fields of view encompassing multiple wells (usually with limited resolution per well, followed by post-processing of images24,63. Neither of these solutions is optimal to perform true simultaneous imaging of biological replicates across multiple conditions. To overcome these limitations, the Picroscope performs an automated image capture of a standard 24 wells cell culture plate using 24 individual objectives. The images are then transferred to a remote computer or server (using the Picroscope’s internet connection), where they can be viewed and/or processed (Fig. 5), with minimal intervention.
Commercial electronic systems for simultaneous imaging of biological samples are typically designed to image cells plated in monolayers64. Yet, significant attention has been given to longitudinal imaging-based screens using whole organisms. These have included zebrafish65,66, worms67, and plants68. Many times, the results of the screens are based on single-plane images or in maximal projections obtained from external microscopes67,68. The Picroscope was designed to overcome these limitations and image along the z-axis. This is accomplished with fine adjustment by two stepper motors that lift the elevator unit that holds all 24 camera objectives (Fig. 4).
To date, few 3D printed microscopes are designed to function inside incubators24,36. We have run the Picroscope in the incubator for three weeks. This makes the Picroscope compatible with screens in 3D mammalian models including organoids69,70. We have shown a proof of principle of this function by performing longitudinal imaging of human cortical organoids and analyzing the behavior and movement of individual cells (Fig. 8).
We anticipate many useful applications of the Picroscope and derivatives of it. Here, we demonstrated the versatility of the Picroscope across animal and cell models in different environmental conditions. The modular nature of the system, allows for new features to be easily built and added. For example, defined spectrum LED light sources and filters for fluorescent imaging would enable longitudinal studies of the appearance and fate of defined sub populations of cells in a complex culture by taking advantage of genetically encoded fluorescent reporter proteins65. Similarly, the use of fluorescent reporters or dyes that respond to dynamic cell states such as calcium sensors allow long-term imaging of cell activity52. The Picroscope, paves the way toward increased accessibility and democratization of multi-well multi-week simultaneous imaging experiments in diverse biological systems.
Materials and methods
System setup
The Picroscope uses 1 Raspberry Pi 4 as a hub Pi that is connected to an ASUS N300 Router via Ethernet. The 24 Raspberry Pi Zero W communicate via WiFi in order to transmit the photos being taken by each of the 24 Zero spy cameras. Each Raspberry Pi device, requires a micro SD card where a copy of our code can be flashed using Balena Etcher. For the lenses on the Zero Spy Cameras, we use Arducam 1/2” M12 Mount 16 mm Focal Length Camera Lenses. An Arduino Uno with the Adafruit V2 Motor Driver Shield is used to control both Nema-11 External 34 mm Stack 0.75A Lead 0.635 mm/0.025" Length 100 mm motors. To detect the bottom of the preset z-stack, we use a limit switch mounted on the 3D printed elevator component via two M2.5 fasteners.
All of the constructive elements of the Picroscope were designed using Fusion360 or AutoCAD computer aided design (CAD) packages. The 3D printed components are sliced with the Prusaslicer with 100% infill, and with 0.15 mm quality preset settings. The 3D printed components were manufactured using a Prusa MK3S 3D printer. The printing material is black PLA. MakerBeam aluminum extrusion elements were used (1) as a structural component for the elevator piece and (2) as a guide for the vertical sliding stages. The sliding stages were constructed using 4 MakerBeam Aluminum extrusions (cross section 10 x 10 mm, length 200 mm).
The custom electronics were designed on a standard 1.6 mm FR4 two-layer PCB. The Power distribution board PCB (Fig. 3c 11) is designed to power and provide structural support for the Raspberry Pi Zero W through their 5V GPIO pins. This design is modular and allows us to have a double-sided PCBs and the same design can be used for the two single-sided PCBs. The relay board (Fig. 3e) allows us to trigger the illumination boards individually as well as shut off power to the entire Picroscope in the event of temperature overheat condition.
For bright field microscopy, the over head light PCB (Fig. 3a 1) uses MEIHUA white LEDs with a brightness of 228 450MCD, and the brightness can be adjusted through a potentiometer. The PCB for lighting from below (Fig. 3a 4) are NCD063W3 Chip LEDs. All custom PCBs are manufactured by PCBWay (China); the cost, including shipping is approximately $2 per board. All electronic components were purchased from Digi-Key Electronics (MN, United States). The PCBs were assembled as shown in Supplementary Note 1. All electronic components (Raspberry Pi, Arduino, and PCBs) were coated with Corona Super Dope Coating to shield the hardware from the effects of condensation due to elevated humidity inside incubator environments. We used a Nomad883 pro to CNC a custom diffuser made out of frosted acrylic. A detailed protocol to build the Picroscope can be found in Supplementary Note 1.
Biological samples
Animal ethics statement
All animal experiments complied with the regulations of the University of California, Santa Cruz and the University of California, San Francisco. Planaria worms and zebrafish eggs are deemed exempt from IACUC protocols. Adult frogs were maintained according to approved UCSF IACUC protocols.
Frogs
Xenopus tropicalis husbandry was performed as previously described71. Adult animals were maintained and cared for according to established IACUC protocols. Animals were wild type and both sexes were used. Animals were ovulated using human chorionic gonadotropin (Sigma-Aldrich, C1063) according to Sive et al. (2000)72 and both in vitro fertilizations and natural matings were used. Embryos were maintained in 1/9 modified Ringer’s solution72 and staged according to Nieuwkoop and Faber (1958)73. Blastopore size was measured in ImageJ/FIJI (NIH) and plotted in GraphPad Prism software version 9.
Zebrafish
Fertilized Danio rerio eggs were purchased from Carolina Biological Supply Company (Catalog # 155591) and maintained in media containing 15 mM sodium chloride (Sigma-Aldrich, S9888), 0.5 mm potassium chloride (Sigma-Aldrich, P3911), 1 mM calcium chloride dihydrate (Sigma-Aldrich, 223506), 1 mM magnesium sulfate heptahydrate (Sigma-Aldrich, 1058822500), 150 μM potassium phosphate monobasic (Sigma-Aldrich, P5655), 50 μM sodium phosphate dibasic heptahydrate (Sigma-Aldrich, S9390), 0.7 mM sodium bicarbonate (Sigma-Aldrich, S5761), and 0.1% methylene blue (Sigma-Aldrich, M9140).
Planaria
Dugesia tigrina Brown planaria worms were purchased from Carolina Biological Supply Company (Catalog # 132954). Planaria were grown in bottled water (Poland Spring). Water was changed every other day.
Human ethics statement
All hESCs experiments used the H9 cell line (WiCell)74. The use of the de-identified H9 embryonic stem cell line was reviewed and approved by the UCSF Human Gamete, Embryo and Cell Research Committee, study number 12-08677.
Human embryonic stem cells and cortical organoids
hESCs were grown on vitronectin (Thermo Fisher Scientific, A14700) coated plates and cultured using StemFlex Medium (Thermo Fisher Scientific, A3349401). Passages were performed incubating the cells in 0.5 mM EDTA (Thermo Fisher Scientific, 15575020), in DPBS for 5 min.)
To generate cortical organoids, we first dissociated hESCs into single cells and re-aggregated them in Aggrewell 800 24-well plates (STEMcell Technologies) at a density of 3,000,000 cells per well with 2 mL of Aggrewell Medium (STEMcell Technologies) supplemented with Rho Kinase Inhibitor (Y-27632, 10 μM, Tocris, 1254) (Day 0). The following day (Day 1), we supplemented the aggregates with WNT inhibitor (IWR1-ε, 3 μM, Cayman Chemical, 13,659, Days 1–10) and TGF-β inhibitor (SB431542, Tocris, 1614, 5 μM, days 0–10). On Day 2, aggregates were transferred by pipetting out of the Aggrewell plate with wide bore P1000 pipette tips onto a 37 μm filter and then transferred to ultra low adhesion 6-well plates. Media was changed on Days 4, 6, 8, and 10, by replacing 2 mL of conditioned media with fresh media. On Day 11, the medium was changed to Neuronal Differentiation Medium containing Eagle Medium: Nutrient Mixture F-12 with GlutaMAX supplement (DMEM/F12, Thermo Fisher Scientific, 1,056,5018), 1X N-2 Supplement (Thermo Fisher Scientific, 17502048), 1X Chemically Defined Lipid Concentrate (Thermo Fisher Scientific, 11,905,031) and 100 U/mL Penicillin/Streptomycin supplemented with 0.1% recombinant human Fetal Growth Factor b (Alamone F-170) and 0.1% recombinant human Epidermal Growth Factor (R&D systems 236-EG). On Day 12, the organoids were transferred in 90 μL media to a custom glass-PDMS microfluidic chip for imaging/feeding containing 50 μL Matrigel hESC Qualif Matrix (BD 354277) bringing the total volume in the well to 120 μL. Partially embedding the organoid in Matrigel in this way led to 2D outgrowths on the surface of the Matrigel. Feeding occurred automatically every hour replacing 30 μL Neuronal Differentiation Medium.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All files needed for the design (CAD) of the Picroscope can be accessed in the Github Repository https://github.com/braingeneers/picroscope-supplement/tree/main/CADs. All files needed for the design of the circuit boards (PBC) can be accessed in the Github Repository https://github.com/braingeneers/picroscope-supplement/tree/main/PCBs.Source data for charts in the main figures are available as Supplementary Data 1. Any remaining information can be obtained from the corresponding author upon reasonable request.
Code availability
All code needed for running the Picroscope can be accessed in the Github Repository https://github.com/braingeneers/picroscope-supplement/tree/main/Codes.
References
Specht, E. A., Braselmann, E. & Palmer, A. E. A critical and comparative review of fluorescent tools for live-cell imaging. Annu. Rev. Physiol. 79, 93–117 (2017).
Godin, A. G., Lounis, B. & Cognet, L. Super-resolution microscopy approaches for live cell imaging. Biophys. J. 107, 1777–1784 (2014).
Miller, A. R. et al. Portable, battery-operated, low-cost, bright field and fluorescence microscope. PloS One 5, e11890 (2010).
Selinummi, J. et al. Bright field microscopy as an alternative to whole cell fluorescence in automated analysis of macrophage images. PloS One 4, e7497 (2009).
Hernández Vera, R., Schwan, E., Fatsis-Kavalopoulos, N. & Kreuger, J. A modular and affordable time-lapse imaging and incubation system based on 3d-printed parts, a smartphone, and off-the-shelf electronics. PLoS One 11, e0167583 (2016).
Savas, J., Khayatzadeh, R., Civitci, F., Gokdel, Y. D. & Ferhanoglu, O. Toward fully three-dimensional-printed miniaturized confocal imager. Opt. Eng. 57, 041402 (2018).
Wincott, M. et al. Democratising “microscopi”: a 3d printed automated xyzt fluorescence imaging system for teaching, outreach and fieldwork. Wellcome Open Res. https://doi.org/10.12688/wellcomeopenres.16536.1 (2021).
Yao, S., Mills, J. K., Ajamieh, I. A., Li, H. & Zhang, X. Automatic three-dimensional imaging for blastomere identification in early-stage embryos based on brightfield microscopy. Opt. Lasers Eng. 130, 106093 (2020).
Zamxaka, M., Pironcheva, G. & Muyima, N. Microbiological and physico-chemical assessment of the quality of domestic water sources in selected rural communities of the Eastern Cape Province, South Africa. Water Sa. 30, 333–340 (2004).
Ferreira, L. M. et al. Effective participatory science education in a diverse Latin American population. Palgrave Commun. 5, 63 (2019).
Giacomotto, J. & Ségalat, L. High-throughput screening and small animal models, where are we? Br. J. Pharmacol. 160, 204–216 (2010).
Willis, S. The maker revolution. Computer. 51, 62–65 (2018).
Barber, K. & Mostajo-Radji, M. A. Youth networks’ advances toward the sustainable development goals during the covid-19 pandemic. Front. Sociol. 5, 589539 (2020).
Coakley, M. F. et al. The NIH 3D print exchange: a public resource for bioscientific and biomedical 3D prints. 3D Print. Addit. Manuf. 1, 137–140 (2014).
Ambrose, B. et al. Democratizing single-molecule fret: an open-source microscope for measuring precise distances and biomolecular dynamics. Biophys. J. 118, 614a (2020).
Gross, B. C., Erkal, J. L., Lockwood, S. Y., Chen, C. & Spence, D. M. Evaluation of 3d printing and its potential impact on biotechnology and the chemical sciences. Anal. Chem. 7, 3240–3253 (2014).
Baden, T. et al. Open labware: 3-d printing your own lab equipment. PLoS Biol. 13, e1002086 (2015).
Alessandri, K. et al. All-in-one 3d printed microscopy chamber for multidimensional imaging, the universlide. Sci. Rep. 7, 1–10 (2017).
Ventola, C. L. Medical applications for 3d printing: current and projected uses. Pharm. Ther. 39, 704 (2014).
Beattie, R. J., Hippenmeyer, S. & Pauler, F. Scopes: sparking curiosity through open-source platforms in education and science. Front. Educ. 5, 8 (2020).
Brown, J. W. et al. Single-molecule detection on a portable 3d-printed microscope. Nat. Commun. 10, 1–7 (2019).
Khan, A., Ikram and Prabhakar, Delepine, C., Tsang, H., Pham, V. & Sur, M. A low-cost 3d printed microfluidic bioreactor and imaging chamber for live-organoid imaging. Biomicrofluidics. 15, 024105 (2021).
Chagas, A. M., Prieto-Godino, L. L., Arrenberg, A. B. & Baden, T. The €100 lab: A 3d-printable open-source platform for fluorescence microscopy, optogenetics, and accurate temperature control during behaviour of zebrafish, drosophila, and caenorhabditis elegans. PLoS Biol. 15, e2002702 (2017).
Kim, S. B. et al. A mini-microscope for in situ monitoring of cells. Lab Chip. 12, 3976–3982 (2012).
Diederich, B. et al. A versatile and customizable low-cost 3d-printed open standard for microscopic imaging. Nat. Commun. 11, 1–9 (2020).
Wang, Z. et al. A high-resolution minimicroscope system for wireless real-time monitoring. IEEE Trans. Biomed. Eng. 65, 1524–1531 (2017).
Zhang, Y. S. et al. A cost-effective fluorescence mini-microscope for biomedical applications. Lab Chip. 15, 3661–3669 (2015).
Zhang, C., Anzalone, N. C., Faria, R. P. & Pearce, J. M. Open-source 3d-printable optics equipment. PloS One 8, e59840 (2013).
Collins, J. T. et al. Robotic microscopy for everyone: the openflexure microscope. Biomed. Opt. Express 11, 2447–2460 (2020).
Cybulski, J. S., Clements, J. & Prakash, M. Foldscope: origami-based paper microscope. PloS One 9, e98781 (2014).
Kim, H. et al. Ludusscope: accessible interactive smartphone microscopy for life-science education. PloS One 11, e0162602 (2016).
Aidukas, T., Eckert, R., Harvey, A. R., Waller, L. & Konda, P. C. Low-cost, sub-micron resolution, wide-field computational microscopy using opensource hardware. Sci. Rep. 9, 1–12 (2019).
Bohm, A. An inexpensive system for imaging the contents of multi-well plates. Acta Crystallogr. F: Struct. Biol. Commun. 74, 797–802 (2018).
Merces, G. O. et al. The incubot: a 3d printer-based microscope for long-term live cell imaging within a tissue culture incubator. HardwareX 9, e00189 (2021).
Gürkan, G. & Gürkan, K. Incu-stream 1.0: an open-hardware live-cell imaging system based on inverted bright-field microscopy and automated mechanical scanning for real-time and long-term imaging of microplates in incubator. IEEE Access 7, 58764–58779 (2019).
Kim, J., Henley, B. M., Kim, C. H., Lester, H. A. & Yang, C. Incubator embedded cell culture imaging system (emsight) based on fourier ptychographic microscopy. Biomed. Opt. Express 7, 3097–3110 (2016).
Baudin, Pierre V., et al. "Low cost cloud based remote microscopy for biological sciences." Internet of Things, 100454, https://doi.org/10.1016/j.iot.2021.100454 (2021).
Goda, T. et al. Genetic screens for mutations affecting development of xenopus tropicalis. PLoS Genet. 2, e91 (2006).
Borodinsky, L. N. Xenopus laevis as a model organism for the study of spinal cord formation, development, function and regeneration. Front. Neural Circuits 11, 90 (2017).
Olmstead, A. W. et al. Reproductive maturation of the tropical clawed frog: Xenopus tropicalis. Gen. Comp. Endocrinol. 160, 117–123 (2009).
Hirsch, N., Zimmerman, L. B. & Grainger, R. M. Xenopus, the next generation: X. tropicalis genetics and genomics. Dev. Dyn. 225, 422–433 (2002).
Mcnamara, S., Wlizla, M. & Horb, M. E. Husbandry, general care, and transportation of Xenopus laevis and Xenopus tropicalis. In Xenopus, (ed. Vleminckx, K.) 1–17 (Springer, 2018).
Khokha, M. K. et al. Techniques and probes for the study of xenopus tropicalis development. Dev. Dyn. 225, 499–510 (2002).
Keller, R. & Sutherland, A. Convergent extension in the amphibian, Xenopus laevis. In Current Topics in Developmental Biology., Vol. 136, (ed. Solnica-Krezel, L.) 271–317 (Elsevier, 2020).
Baldwin, A., Kim, J. & Wallingford, J. B. Global analysis of cell behavior and protein localization dynamics reveals region-specific functions for shroom3 and n-cadherin during neural tube closure. bioRxiv (2021).
Huebner, R. J. et al. Mechanical heterogeneity along single cell-cell junctions is driven by lateral clustering of cadherins during vertebrate axis elongation. Elife 10, e65390 (2021).
Mostajo-Radji, M. A., Schmitz, M. T., Montoya, S. T. & Pollen, A. A. Reverse engineering human brain evolution using organoid models. Brain Research 1729, 146582 (2020).
Abe-Fukasawa, N., Otsuka, K., Aihara, A., Itasaki, N. & Nishino, T. Novel 3d liquid cell culture method for anchorage-independent cell growth, cell imaging and automated drug screening. Sci. Rep. 8, 1–12 (2018).
Zhao, H., Tang, C., Cui, K., Ang, B.-T. & Wong, S. T. A screening platform for glioma growth and invasion using bioluminescence imaging. J. Neurosurg.111, 238–246 (2009).
Almassalha, L. M. et al. Label-free imaging of the native, living cellular nanoarchitecture using partial-wave spectroscopic microscopy. Proc. Natl. Acad. Sci USA. 113, E6372–E6381 (2016).
Martin, H. L. et al. High-content, high-throughput screening for the identification of cytotoxic compounds based on cell morphology and cell proliferation markers. PloS One 9, e88338 (2014).
Dempsey, G. T. et al. Cardiotoxicity screening with simultaneous optogenetic pacing, voltage imaging and calcium imaging. J. Pharmacol. Toxicol. Methods 81, 240–250 (2016).
Honarnejad, K. et al. Fret-based calcium imaging: a tool for high-throughput/content phenotypic drug screening in alzheimer disease. J. Biomol. Screen.18, 1309–1320 (2013).
Park, J., Kwon, O.-h., Yoon, C. & Park, M. Estimates of particulate matter inhalation doses during three-dimensional printing: How many particles can penetrate into our body? Indoor Air 31, 392–404 (2020).
Pearce, J. M. Building research equipment with free, open-source hardware. Science 337, 1303–1304 (2012).
Mendoza-Gallegos, R. A., Rios, A. & Garcia-Cordero, J. L. An affordable and portable thermocycler for real-time pcr made of 3d-printed parts and off-the-shelf electronics. Anal. Chem. 90, 5563–5568 (2018).
Carosso, G. A., Ferreira, L. M. & Mostajo-Radji, M. A. Scientists as non-state actors of public diplomacy. Nat. Hum. Behav. 3, 1129–1130 (2019).
Kwon, H.-S. et al. Performance of minipcr tm mini8, a portable thermal cycler. Anal. Sci. Technol. 29, 79–84 (2016).
Byagathvalli, G., Pomerantz, A., Sinha, S., Standeven, J. & Bhamla, M. S. A 3d-printed hand-powered centrifuge for molecular biology. PLoS Biol. 17, e3000251 (2019).
González-González, E., Trujillo-de Santiago, G., Lara-Mayorga, I. M., Martinez-Chapa, S. O. & Alvarez, M. M. Portable and accurate diagnostics for covid-19: Combined use of the minipcr thermocycler and a well-plate reader for sars-cov-2 virus detection. PloS One 15, e0237418 (2020).
Boguraev, A.-S. et al. Successful amplification of dna aboard the international space station. NPJ Microgravity 3, 1–4 (2017).
Hossain, Z. et al. Interactive and scalable biology cloud experimentation for scientific inquiry and education. Nat. Biotech. 34, 1293–1298 (2016).
Klimaj, S. D., Licon Munoz, Y., Del Toro, K. & Hines, W. C. A high-throughput imaging and quantification pipeline for the evos imaging platform. Plos One 15, e0236397 (2020).
Ruckdäschel, S., Michaelis, S. & Wegener, J. Time lapse imaging of spheroids–zencellowl incubator. https://annualmeeting.ls2.ch/files/2021/content/364/zencellowl-incubator-microscope-time-lapse-imaging-of-spheroids.pdf (2017).
Early, J. J. et al. An automated high-resolution in vivo screen in zebrafish to identify chemical regulators of myelination. Elife 7, e35136 (2018).
Tsuji, N. et al. Whole organism high content screening identifies stimulators of pancreatic beta-cell proliferation. PloS One 9, e104112 (2014).
Lemieux, G. A. et al. A whole-organism screen identifies new regulators of fat storage. Nat. Chem. Biol. 7, 206–213 (2011).
Schreiber, K., Ckurshumova, W., Peek, J. & Desveaux, D. A high-throughput chemical screen for resistance to pseudomonas syringae in arabidopsis. Plant J. 54, 522–531 (2008).
Renner, H. et al. A fully automated high-throughput workflow for 3d-based chemical screening in human midbrain organoids. Elife 9, e52904 (2020).
Schuster, B. et al. Automated microfluidic platform for dynamic and combinatorial drug screening of tumor organoids. Nat. Commun. 11, 1–12 (2020).
Willsey, H. R. et al. The neurodevelopmental disorder risk gene dyrk1a is required for ciliogenesis and control of brain size in xenopus embryos. Development. 147, dev189290 (2020).
Sive, H. L., Grainger, R. M. & Harland, R. M. Early Development of Xenopus Laevis: A Laboratory Manual. (CSHL Press, 2000).
Nieuwkoop, P. D. & Faber, J. Normal table of Xenopus laevis (Daudin). Copeia 1958, 65–65 (1958).
Thomson, J. A. et al. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998).
Acknowledgements
We would like to thank Jeremy Linsley and Wiktoria Leks for providing us zebrafish for this study. In addition, we would like to thank Arnar Breevoort for providing experimental support. This work was supported by the Schmidt Futures Foundation SF 857 (D.H.), the National Institute of Mental Health of the National Institutes of Health under Award Number R01MH120295 (S.R.S.) and the National Science Foundation under award number NSF 2034037 (M.T.). H.R.W. was supported by grant U01MH115747-01 from NIMH to Matthew State. M.A.M.-R. was partially supported by grant TL1 TR001871 from the NIH National Center for Advancing Translational Sciences. K.V. was supported by grant T32HG008345 from the National Human Genome Research Institute (NHGRI), part of National Institutes of Health (NIH), USA. D.H. is a Howard Hughes Medical Institute Investigator.
Author information
Authors and Affiliations
Contributions
V.T.L., P.V.B., P.P., and K.V. worked on hardware, software, and assembly of the Picroscope. Y.R. and E.A.J. worked on early prototypes. V.T.L., P.V.B., H.R.W., G.L.M., S.T.S., J.A.S., S.A.C., J.M.R., A.A.P., and M.A.M.-R. performed giological experiments. H.R.W., A.A.P., T.J.N., S.R.S., M.A. M.-R., D.H., and M.T. conceived the experiments. D.H., M.T., S.R. S,A. A.P., M.A. M.-R., M.R., and T.J.N. supervised the team and secured funding. V.T.L., P.V.B.,E. A.J., M.A. M.-R., and M.T. wrote the manuscript with contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors have written patents covering the technology described in this article. A.A. P. is in the board of Herophilus. The authors declare no competing interests.
Additional information
Peer review information Communications Biology thanks Güray Gürkan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary handling editors: Chao Zhou and Anam Akhtar.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ly, V.T., Baudin, P.V., Pansodtee, P. et al. Picroscope: low-cost system for simultaneous longitudinal biological imaging. Commun Biol 4, 1261 (2021). https://doi.org/10.1038/s42003-021-02779-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-021-02779-7
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.