Abstract
Observational studies suggest smoking, cannabis use, alcohol consumption, and substance use disorders (SUDs) may impact risk for respiratory infections, including coronavirus 2019 (COVID-2019). However, causal inference is challenging due to comorbid substance use. Using summary-level European ancestry data (>1.7 million participants), we performed single-variable and multivariable Mendelian randomization (MR) to evaluate relationships between substance use behaviors, COVID-19 and other respiratory infections. Genetic liability for smoking demonstrated the strongest associations with COVID-19 infection risk, including the risk for very severe respiratory confirmed COVID-19 (odds ratio (OR) = 2.69, 95% CI, 1.42, 5.10, P-value = 0.002), and COVID-19 infections requiring hospitalization (OR = 3.49, 95% CI, 2.23, 5.44, P-value = 3.74 × 10−8); these associations generally remained robust in models accounting for other substance use and cardiometabolic risk factors. Smoking was also strongly associated with increased risk of other respiratory infections, including asthma-related pneumonia/sepsis (OR = 3.64, 95% CI, 2.16, 6.11, P-value = 1.07 × 10−6), chronic lower respiratory diseases (OR = 2.29, 95% CI, 1.80, 2.91, P-value = 1.69 × 10−11), and bacterial pneumonia (OR = 2.14, 95% CI, 1.42, 3.24, P-value = 2.84 × 10−4). We provide strong genetic evidence showing smoking increases the risk for COVID-19 and other respiratory infections even after accounting for other substance use behaviors and cardiometabolic diseases, which suggests that prevention programs aimed at reducing smoking may be important for the COVID-19 pandemic and have substantial public health benefits.
Similar content being viewed by others
Introduction
Since the first reported cases in Wuhan, China in December 20191, coronavirus disease 2019 (COVID-19) has subsequently affected more than 200 countries and continues to be a global pandemic of substantial worldwide morbidity and mortality2,3. More broadly, upper and lower respiratory infections (URIs and LRIs, respectively) and other respiratory diseases (i.e., asthma, chronic obstructive pulmonary disease (COPD), etc.) are leading causes of yearly worldwide morbidity and mortality4,5. For example, the Global Burden of Disease Study estimated that LRIs caused more than two million deaths globally in 20164, while approximately 2.3 million people died from COPD in 20155. Respiratory infection and diseases are also a large economic burden: URIs result in more than 40 million missed days of school and work per year6.
Substance use (tobacco smoking, cannabis use, and alcohol consumption) are risk factors linked with adverse lung and respiratory outcomes7,8,9. For example, observational data has shown chronic heavy alcohol consumption to be associated with increased risk for pneumonia7 and acute respiratory distress syndrome10, while cannabis smoke has been shown to contain many of the same toxins and irritants as smoke derived from tobacco11, but may differ from tobacco in its association with bronchitis and other respiratory infections12. In addition, it has been suggested that chronic alcohol abuse may compromise the ability of immune cells to destroy bacteria in the lungs, which may result in an increased vulnerability to respiratory infections like pneumonia and tuberculosis13.
Paralleling the COVID-19 pandemic have been increases in substance use14, which combined with data showing approximately 10.8% of US adults suffering from a substance use disorder (SUD)15 and recent work using electronic health records (EHRs) to show that individuals with a SUD are at increased risk for COVID-1916, suggest identifying potential causal relationships between substance use, SUD and respiratory infectious diseases would have substantial public health benefits.
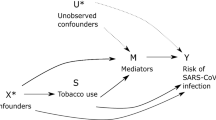
However, observational studies cannot be used to reliably identify causality due to limitations such as residual confounding and reverse causality17. For example, outcomes reached from observational studies may be subject to unmeasured confounders like comorbid disorders or underlying genetic differences that may lead to biased estimates, and consequently, may not reflect true causal relationships18,19. While randomized controlled trials (RCTs) are considered the “gold standard”, RCTs can be both unethical and impractical20,21. Constructing an RCT to examine the effect of substance use on respiratory infection risk may be further complicated by other existing comorbidities.
Mendelian randomization (MR) is a genetic approach that uses genetic variants as instrumental variables to explore causal relations between exposures (e.g., alcohol consumption, tobacco smoking, cannabis use) and health outcomes (e.g., respiratory infections and diseases). This technique takes publicly available genome wide association studies to screen for suitable genetic instrumental variables, which allows researchers to perform MR studies without the need to recruit new patients22. Because germline variants are randomly assorted at meiosis, MR may be considered conceptually equivalent to RCTs, though a more naturalized version19,22. More specifically, given genetic instruments cannot be influenced by other confounders (i.e., lifestyle, or environmental factors), MR studies, are in theory, less susceptible to confounding or reverse causality than traditional observational studies23. Therefore, MR is an important analytical approach to strengthen causal inference when RCTs are challenging due to methodological or ethical constraints24. Given the potential for confounding and limited causal inference derived from observational data, we used large, publicly available genome-wide association study (GWAS) data and two-sample MR methods to evaluate the relationships between substance use, substance use disorders (cannabis use disorder (CUD) and alcohol use disorder (AUD)) and respiratory infection and disease outcomes. Finding the genetic liability for smoking increases the risk for COVID-19 and several other respiratory infections, even after accounting for other substance use behaviors builds upon recent literature identifying modifiable risk factors for COVID-19 risk9,25,26, and also may inform research and clinical practice given the recent increase in substance use, abuse, and use disorders paralleling the COVID-19 pandemic14.
Results
Associations of substance use and SUDs with COVID-19 infection risk
COVID-19 results comparing SVMR and MVMR results are presented in Table 1. Supplementary Data 8–12 present the full COVID-19 results. Broadly, among all substance use exposures, the genetic liability for lifetime tobacco smoking consistently demonstrated the strongest associations with COVID-19 infection risk, including the risk for very severe respiratory confirmed COVID-19 (SVMR odds ratio (OR) = 2.69, 95% CI, 1.42, 5.10, P-value = 0.002), and also the risk for COVID-19 infection requiring hospitalization (hospitalized COVID-19 vs population: SVMR odds ratio (OR) = 3.49, 95% CI, 2.23, 5.44, P-value = 3.74 × 10−8; MVMR accounting for substance use disorders OR = 3.61, 95% CI, 2.19, 5.95, P-value = 4.92 × 10−7; and hospitalized vs not hospitalized COVID-19: SVMR OR = 3.44, 95% CI, 1.72, 6.87, P-value = 4.60 × 10−4; MVMR OR = 3.61, 95% CI, 1.63, 8.01, P-value = 0.002) (Table 1; Supplementary Data 8, 10, and 11). This association remained robust in secondary sensitivity analyses excluding UK Biobank participants in the COVID-19 outcome GWAS, but with reduced precision (hospitalized COVID-19 vs population: SVMR OR = 2.42,95% CI, 1.46, 4.01, P-value = 6.09 × 10−4; MVMR OR = 2.62, 95% CI, 1.46, 4.71, P-value = 0.001; and hospitalized vs not hospitalized COVID-19: SVMR OR = 3.27, 95% CI, 1.15, 9.33, P-value = 0.03; MVMR OR = 4.84, 95% CI, 1.46, 15.39, P-value = 0.008) (Supplementary Data 8, 10, and 11). Importantly, these associations were consistent across complementary SVMR and MVMR methods, including single variable GSMR (Supplementary Data 8, 10, and 11). Leave-one-out analyses highlight variants with heterogeneous causal effects that would be flagged as invalid by MR PRESSO and MV MR Lasso and removed for outlier corrected results (Supplementary Data 9).
Given the strong associations of lifetime tobacco smoking and COVID-19 risk, we further evaluated robustness by performing MVMR analyses accounting for cardiometabolic disorders (CAD, T2D, and obesity) previously reported as risk factors for COVID-19 risk27,28,29. Genetic liability for lifetime tobacco smoking generally remained associated with increased risk for COVID-19 hospitalization (e.g., accounting for CAD, hospitalized COVID-19 vs. population: MVMR OR = 3.18, 95% CI, 2.06, 4.92, P-value = 1.80 × 10−7; accounting for Type 2 diabetes, MVMR OR = 4.16, 95% CI, 2.51, 6.92, P-value = 3.76 × 10−8; accounting for obesity, MVMR OR = 3.75, 95% CI, 2.25, 6.25, P-value = 4.01 × 10−7) (Supplementary Data 12).
Associations of substance use and SUDs with other respiratory infectious disease risk
We further assessed the genetic relationships between substance use and respiratory infections. Tables 2 and 3 compares SVMR and MVMR results for asthma-related respiratory infections, bronchitis, and the common cold; Tables 4 and 5 compares SVMR and MVMR results for influenza and pneumonias. Supplementary Data 13–17 contain the full FinnGen results.
As with COVID-19 infection risk results, we found that the genetic liability of lifetime tobacco smoking was the substance use risk factor with the strongest associations, including results that were robust in MVMR models. Tobacco smoking, for example, was associated with increased risk of asthma-related infections and asthma-related pneumonia/sepsis (SVMR OR = 2.52, 95% CI, 1.59, 3.97, P-value = 7.29 × 10−7; accounting for substance use disorders, MVMR OR = 3.64, 95% CI, 2.16, 6.11, P-value = 1.07 × 10−6), but for neither bronchitis nor the common cold (Table 3; Supplementary Data 13–15). Tobacco smoking was also associated with chronic lower respiratory diseases (SVMR OR = 2.23, 95% CI, 1.73, 2.87, P-value = 5.69 × 10−10; MVMR OR = 2.29, 95% CI, 1.80, 2.91, P-value = 1.69 × 10−11) and several pneumonia-related outcomes, including bacterial pneumonia (SVMR OR = 2.22, 95% CI, 1.57, 3.15, P-value = 7.32 × 10−6; MVMR OR = 2.14, 95% CI, 1.42, 3.24, P-value = 2.84 × 10−4) (Table 5, Supplementary Data 13–15).
As with the smoking-COVID-19 findings, we tested robustness of the smoking-respiratory infection risk results using additional MVMR models that accounted for cardiometabolic disorders (CAD, T2D, and obesity) with evidence for an impact on respiratory infection risk30,31,32. Our smoking-related results were broadly robust to inclusion of cardiometabolic confounders (Supplementary Data 16). These associations were generally consistent across complementary SVMR and MVMR methods, including single variable GSMR (Supplementary Data 13–16). Leave-one-out analyses again highlight variants with heterogeneous causal effects that would be flagged as invalid by MR PRESSO and MV MR Lasso and removed for outlier corrected results (Supplementary Data 17).
Discussion
Using large summary-level GWAS data and complementary two-sample MR methods, we show that the genetic liability for tobacco smoking has potential causal relationships with several respiratory infection and disease outcomes, including COVID-19. These tobacco smoking-respiratory findings were supported by multivariable MR analyses accounting for alcohol and cannabis use and abuse, which in addition to the broadly consistent IVW results (within the IVW MR 95% confidence interval but typically less precise) with estimates from the weighted median, weighted mode, and MR Egger sensitivity analyses strengthen causal inference. Further, in single variable MR, we identify potential adverse impacts of CUD on lower respiratory infections, the common cold, and several asthma-related infections, suggesting evidence for a dose-dependent impact of cannabis use where heavy cannabis use may be harmful to the respiratory system. In parallel, we find little evidence for an alcohol-respiratory infection relationship suggesting that previous observational data may be due to confounding.
Our COVID-19 results extend recent MR studies showing adverse effects of smoking on COVID-19 risk by accounting for highly comorbid alcohol consumption, cannabis use, and SUDs, which when combined with reports suggesting smoking intensifies the severity of COVID-19 symptoms33,34, the risk for being admitted to an intensive care unit or requiring ventilation34, and recent transcriptomics-based work showing that smoking may increase the expression of angiotensin converting enzyme 2 (ACE2), the putative receptor for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) (the virus that causes COVID-19)35, suggests smoking may be an important modifiable risk factor for COVID-19 risk.
Our genetics-based findings support and extend the observational literature identifying tobacco smoking as a risk factor for respiratory infection and diseases9,25,26, and add to the recent MR literature identifying potential causal links of smoking with reduced lung function36, lung cancer37, and increased mortality due to respiratory disease38. Potential mechanisms by which smoking increases respiratory infection risk include structural changes to the respiratory tract and a dysregulated cellular and humoral immune response, including peribronchiolar inflammation, decreased levels of circulating immunoglobulins, and changes to pathogen adherence. For example, smoking has been shown to stimulate the release of catecholamine and corticosteroids, which may, in turn, increase circulating CD8+ lymphocytes and suppress the host defense against infections. Notably, many immunological effects related to smoking may resolve within six weeks of smoking cessation, which suggests that smoking cessation programs may have an important impact on reducing respiratory infections.
Regarding cannabis use, while we failed to find evidence of any relationships, smoking cannabis, like tobacco smoking, may prompt the onset of coughing, which could consequently increase viral transmission, or may possibly exacerbate respiratory symptoms.
As cannabis is the most used drug worldwide—an estimated 188 million recreational users worldwide—this aspect of cannabis use may have important implications for the spread of COVID-19. In contrast, the single-variable MR CUD results demonstrated adverse effects on several respiratory outcomes, but not COPD, which supports the existing literature39,40,41; however, accounting for lifetime tobacco smoking attenuated the CUD results, thus highlighting the complex nature of these relationships. Further, habitual cannabis smoking may have several effects on respiratory and immune systems that may impact respiratory infection susceptibility. For example, structural abnormalities in alveolar macrophages and coincident dysregulated cytokine production and antimicrobial activity have been reported. While our study provides preliminary genetic evidence suggesting potential causal relationships between heavy cannabis use and respiratory infection, additional triangulating lines of evidence (i.e., immune monitoring studies) are required to further elucidate the CUD-respiratory infection relationship. However, given that the toxin and irritant profiles of cannabis and tobacco smoke are similar11, the direct route of administration via inhalation for these substances could result in dysregulated pulmonary physiology which may, in turn, increase infection risk.
In contrast to our tobacco smoking findings, we failed to find genetic evidence of respiratory implications due to alcohol consumption not meeting the threshold of AUD, or binge drinking, suggesting that previous observational literature may be due to confounding from other comorbid behaviors—such as smoking—that may be the true causal risk factors for respiratory infections. For example, observational and genetic evidence have shown a strong association between alcohol consumption and smoking. It has been estimated that 85% of smokers consume alcohol42,43,44 and alcohol drinkers are 75% more likely than abstainers to smoke45. Therefore, it is possible that the observational study-based alcohol-respiratory infection links may be due, instead, to tobacco smoking; however, future work will be needed to confirm this hypothesis. In addition, it is important to note that our results should not be interpreted as suggesting that alcohol does not impact overall lung health and structure, which has been previously reported7. Further, while we failed to find evidence that weekly alcohol consumption impacted COVID-19 risk, the Centers for Disease Control recently showed that dining at on-site locations, such as restaurants and bars, is associated with increased COVID-19 risk; since alcohol consumption may lower inhibition and increase impulsivity, individuals consuming alcohol may take social distancing less seriously, and thereby unintentionally spread the SARS-CoV-2 virus.
This study has several strengths including the use of multiple alcohol consumption and cannabis use variables, which enabled us to evaluate various dimensions of substance use and abuse and identify possible causal relationships of substance use disorders and respiratory outcomes. In addition, our main single variable analyses included multiple MR methods, each relying on orthogonal assumptions, which provide confidence in robustness of the results and strengthen causal inference46. Our multivariable two-sample MR design, the most appropriate design given the strong correlation between tobacco smoking, alcohol consumption and cannabis use, yielded estimates that account for these correlated behaviors for each exposure on COVID-19 risk and other respiratory outcomes. Another strength is our extension of MVMR to test the robustness of the main tobacco smoking findings by incorporating other potential confounders that may impact infectious disease risk (obesity, cardiovascular disease, and T2D).
This study also has limitations. A main limitation is the possibility of collider bias—especially with regards to the COVID-19 datasets47. Collider bias may occur when analyses are controls or selects the sample based upon a collider variable that is caused by both the exposure and outcome variables and distorts the true underlying association48,49. The recent commentaries by Griffith et al. (2020) and Tattan-Birch et al. (2020) discuss in detail the potential for collider bias in COVID-19 datasets47,49, and are important for context when interpreting COVID-19 findings based upon observational data. For example, an observational study from early in the COVID-19 pandemic reported an apparent protective effect of tobacco smoking on COVID-19 risk50; however, as Tattan-Birch et al. discuss, both smoking and COVID-19 may cause coughing, which, during the COVID-19 pandemic, may increase the likelihood for smokers to be tested and their subsequent overrepresentation among clinical study participants testing negative for COVID-1949. As a result, among samples with COVID-19 tests, smoking may appear to have a protective effect49. While it is often not possible to ensure the absence of collider bias47, we aimed to design our study incorporating measures that may mitigate its impact. For example, we used the most recently released version of publicly available COVID-19 data (from January 18, 2021)51 that may include participants more representative of the general population compared to samples collected earlier in the COVID-19 pandemic. Reassuringly, we also found similar smoking effect estimates in several respiratory-related infection outcomes, which suggests a broader impact of smoking on the respiratory system that extends to COVID-19.
In addition, as with all self-reported substance use literature, these exposures may be either under- or over-reported52. Because many of the datasets included UK Biobank participants, who are more educated, lead healthier lifestyles, and have fewer health problems than the UK population53, this discrepancy may limit the applicability of our findings to other populations. Regarding our mainly null alcohol-respiratory infection results, it is possible that alcohol may have indirect impacts on infection risk through a modified immune response54, or other system dysregulations that may modulate infection risk that we were not able to directly assess using MR. However, like other recent psychiatric MR studies where the exposure instruments included a relaxed statistical threshold, our binge drinking and AUD instruments were comprised of independent SNPs associated with the respective drinking behavior (i.e., P-value < 5 × 10−6) for SNP inclusion due to the lack of conventionally GWS SNPs (P-value < 5 × 10−8)55,56, which may impact the results. Because heavy alcohol consumption and AUD have been previously linked with acute respiratory distress syndrome10—one of the most severe complications of COVID-19—future studies should re-evaluate the links between heavy alcohol consumption and AUD when better powered GWAS data become available.
Further, the included samples were comprised of primarily white individuals of European ancestry, and research has shown strong racial, ethnic, and socioeconomic disparities in COVID-19 risk and severity57,58,59. Therefore, we caution the generalization of these findings and urge future work to investigate these relationships using a genetics-based approach in other populations when the data become available. Another limitation is the overlap of the UKB participants between the alcohol consumption, lifetime smoking, and COVID-19 outcomes, which may bias resulting estimates60. However, potential bias would likely be minimal60, and it has also been shown that two-sample MR may be used in single samples provided the data is derived from large biobanks, i.e., the UKB, FinnGen, etc61. Also, results were largely unchanged when we performed analyses using the COVID-19 endpoints excluding UKB participants suggesting minimal bias.
In conclusion, our data provide genetic evidence of adverse relationships between smoking and many respiratory-related disease outcomes ranging from the common cold to severe COVID-19, which suggests prevention programs aimed at smoking cessation and prevention may have public health and clinical benefits.
Methods
Data sources and genetic instruments
Summary-level data for both modifiable risk factor instrument and infectious disease outcome data were derived from publicly available GWASs in populations of predominantly European ancestry (Fig. 1; Table 6; Supplementary Data 1). All GWASs have existing ethical permissions from their respective institutional review boards and include participant informed consent with rigorous quality control. For this study, we included all exposure SNPs associated at conventional genome-wide significance (GWAS) P < 5 × 10−8 for smoking, alcohol and cannabis use, and 5 × 10−6 for AUD and CUD due to the relatively low number of SNPs at GWS, clumped at linkage disequilibrium (LD) r2 = 0.001 and a distance of 10,000 kb, using reference samples comprised of participants of European ancestry 62.
Abbreviations: SNP: single nucleotide polymorphism; COVID-19: coronavirus disease 2019; COPD: Chronic obstructive pulmonary disease; IVW, Inverse Variance Weighted MR; SVMR; single variable Mendelian randomization; GSMR: generalized summary Mendelian randomization; MVMR: multivariable Mendelian randomization; MR PRESSO: MR pleiotropy residual sum and outlier; HEIDI: heterogeneity in dependent instruments.
Tobacco smoking
We included lifetime smoking instruments from the recent GWAS of a lifetime smoking index/score (which combined smoking initiation, duration, heaviness and cessation), conducted in a sample of 462 690 current, former and never smokers in the UKB (mean score value 0.359 (standard deviation (SD) = 0.694); sample: 54% female, mean age 56.7 years, 54% never smokers, 36% former smokers, and 11% current smokers63,64. (An SD increase in lifetime smoking index score would be equivalent to smoking 20 cigarettes per day for 15 years and stopping 17 years previously or 60 cigarettes per day for 13 years and stopping 22 years previously)63 (Supplementary Data 2).
Cannabis use
We included two cannabis-related instrument sets: cannabis use and CUD. Summary statistics for lifetime cannabis use (a yes/no variable of whether participants reported using cannabis during their lifetime) were obtained from the PGC meta-analysis GWAS of 3 cohorts (International Cannabis Consortium (35,297 respondents, 55.5 percent female, ages 16–87, mean 35.7 years; 42.8 percent had used cannabis); UKB (126 785 respondents, 56.3 percent female, ages 39–72, mean age 55.0 years, 22.3 percent had used cannabis); and 23andMe (22,683 respondents, 55.3 percent female, ages 18–94, mean age 54.0 years, 43.2% had used cannabis))65,66. CUD instruments were obtained from a recent PGC meta-analysis of three cohorts of predominantly European ancestry (PGC, Lundbeck Foundation Initiative for Integrative Psychiatric Research (iPSYCH), and deCODE cohorts, excluding related individuals from PGC family-based cohorts; demographics not available), including 14,808 cases of cannabis abuse or dependence defined as meeting DSM-IIIR, DSM-IV, DSM-5, or ICD10 codes (depending on study cohort) criteria; the 358 534 controls were defined as anyone not meeting the criteria67,68 (Supplementary Data 3).
Alcohol consumption
We included two instrument sets related to alcohol use: drinks per week69, and AUD. Drinks per week instruments were obtained from the GSCAN GWAS meta-analysis of 29 cohorts (941 280 individuals; demographics not available) of predominantly white European ancestry69,70. Given the varied cohort methods used to measure alcohol consumption (binned, normalized, etc.), the data was log transformed: thus, the effect estimate is measured in log transformed drinks per week69 (Supplementary Data 4). For the AUD instrument set, we used the Psychiatric Genomics Consortium (PGC) GWAS meta-analysis of 28 cohorts (51.6% female, 8485 cases, 20,657 controls) of predominantly European ancestry71,72. AUD was diagnosed by either clinician rating or semi-structured interview using DSM-IV criteria including the presence of at least three of seven alcohol-related symptoms (withdrawal, drinking larger amounts/drinking for longer time, tolerance, desire or attempts to cut down drinking, giving up important activities to drink, time related to drinking, or continued alcohol consumption despite psychological and/or physical problems)73 (Supplementary Data 4).
For the multivariable MR (MVMR) analyses, we concatenated independent instrument sets for alcohol use, cannabis use and lifetime smoking, and also AUD, CUD, and lifetime smoking, clumping the resulting two multivariable (MV) instrument sets to exclude intercorrelated SNPs with pairwise LD r2 > 0.001, leaving 141 and 126 MV instruments, respectively (Supplementary Data 5 and 6).
Obesity, coronary heart disease (CAD), and Type 2 Diabetes (T2D) have been identified as risk factors for COVID-1927,28,29, and other respiratory infections30,31,32. Therefore, in supplementary sensitivity analyses to further test robustness of the lifetime smoking results, we concatenated independent instrument sets for lifetime smoking and, alternatively, CAD using the CARDIoGRAMplusC4D-UK Biobank CAD (Coronary ARtery DIsease Genome wide Replication and Meta-analysis (CARDIoGRAM) plus The Coronary Artery Disease (C4D) Genetics) GWAS meta-analysis74,75; T2D, using a recent meta-analysis of three T2D studies, i.e. DIAbetes Genetics Replication and Meta-analysis (DIAGRAM), Genetic Epidemiology Research on Aging (GERA) and the full cohort release of UKB76,77; and obesity, using GWASs from GIANT (Genetic Investigation of ANthropometric Traits)78,79 (see Supplementary Data 1 for more information; Supplementary Data 7).
F statistics for the unconditional instruments were strong (>10, Supplementary Data 2–4). We were unable to calculate conditional F statistics to assess the strength of the multivariable instrument sets: SVMR statistical methods recently extended to two sample MVMR are appropriate only for non-overlapping exposure summary level data sources. When overlapping, the requisite pairwise covariances between SNP associations are only determinable by using individual level data 80.
COVID-19 outcomes
We used summary GWAS statistics from the COVID-19 Host Genetics Initiative (COVID-19 hg) meta-analysis round 5a (18 January 2021 release date) (https://www.covid19hg.org/results)81 for four COVID-19 phenotypes in cohorts of European ancestry, both including and excluding the UKB cohorts for sensitivity analyses (N cases; N controls): very severe respiratory confirmed COVID-19 versus population (4606; 702,801); very severe respiratory confirmed COVID-19 versus population excluding UKB cohorts (4297; 374,224); hospitalized versus not hospitalized COVID-19 (4829; 11,816); hospitalized versus not hospitalized COVID-19 excluding UKB cohorts (3159; 7206); hospitalized COVID-19 versus population (9373; 1,197,256); hospitalized COVID-19 versus population excluding UKB cohorts (7703; 868,679); COVID-19 versus population (29,071; 1,559,712); COVID-19 versus population excluding UKB cohorts (22,581; 1,231,135) (demographics not available) (Fig. 1; Table 6; Supplementary Data 1).
Other respiratory infection and disease outcomes
We used data from FinnGen Release 5 (released to public, 11 May 2021) for additional respiratory-related outcomes82, including acute upper respiratory infections, asthma related acute respiratory infections, pneumonia, influenza, bronchitis, chronic lower respiratory diseases, and acute nasopharyngitis (common cold) (N ≤ 218,792) (Fig. 1; Table 6; Supplementary Data 1). FinnGen is a public-private partnership incorporating genetic data for disease endpoints from Finnish biobanks and Finnish health registry EHRs82. Detailed documentation is provided on the FinnGen website (https://finngen.gitbook.io/documentation/).
Sample independence
Participant overlap in samples used to estimate genetic associations between exposures and outcomes can increase weak instrument bias (WIB) in MR analyses60,83, but to a lesser extent with large biobank samples (including UKB and deCODE). Given the large size of the overlapping cohorts (e.g., UKB, deCode) (Supplementary Data 1) and the strength of the instruments in both directions (F statistics > 10; Supplementary Data 2–4), considerable WIB would not be expected60,84. We have conducted analyses for COVID-19 outcomes using COVID-19 GWAS performed both including and excluding UKB cohorts.
Statistics and reproducibility
For SVMR analyses, we used inverse-variance weighted MR (MR IVW) as the main analyses, supplemented by MR-Egger, weighted median, and weighted mode methods. These are complementary robust methods developed to estimate consistent causal effects under weaker assumptions than MR IVW to assess evidence of causal effects for each of alcohol, cannabis and tobacco use, and use disorders on infectious disease outcomes, and evaluate the sensitivity of the analyses to different patterns of violations of IV assumptions85. Consistency of results across methods strengthens an inference of causality85. For MVMR analyses, we used the multivariable extensions of MR IVW, MR Egger, and MR median 83,86.
We used the MR Egger intercept test87, Cochran Q heterogeneity test88, and multivariable extensions thereof, to evaluate heterogeneity in instrument effects, as heterogeneity may indicate violations of IV assumptions86,87,89. The MR pleiotropy residual sum and outlier (MR PRESSO) global test, and multivariable extensions thereof90, were used to facilitate identification and removal of outlier instruments to correct potential directional horizontal pleiotropy and resolve detected heterogeneity. For SVMR, we also used, alternatively, Generalized single variable Summary-data based MR (GSMR) to identify and remove instruments with heterogeneous causal estimates suspected to be invalid instruments with apparent pleiotropic effects on both exposure and outcome disease (using the recommended default HEIDI (heterogeneity in dependent instruments) -outlier threshold (0.01) to retain sufficient power to detect heterogeneity)91. We used the SVMR Steiger directionality test to test the causal direction between the hypothesized exposure and outcomes62. We also performed a leave-one-out analysis to evaluate the potential SNPs within each instrument that may be high influence points85.
For MVMR, in addition to the multivariable extension of the MR PRESSO global test, we used the multivariable extension of the MR Lasso method, which applies lasso-type penalization to the direct effects of the instruments on the outcome disease: the so-called post-lasso estimate is obtained by performing MR IVW using only those instruments identified as valid instruments (tuning parameter specified at default heterogeneity stopping rule)89. Analyses were carried out using TwoSampleMR, version 0.5.585, MendelianRandomization, version 0.5.0, in the R environment, version 4.0.2; the GSMR method was implemented in the GCTA (Genome-wide Complex Trait Analysis) software (https://cnsgenomics.com/software/gcta/#GSMR).
Reported results and interpretation of findings
MR IVW odds ratios (OR) with 95% CI, per unit increase in the exposures (e.g., per unit increase of log-transformed alcoholic drinks per week or lifetime smoking index), with P-values derived from two-sided tests, corrected for outlier or invalid variants, are presented in Tables 1–5. For our COVID-19 analyses, we used a two-sided α of 0.0025 (based on comparing four COVID-19 outcomes and five substance use exposures) and for the other infectious disease outcomes, a threshold of 0.00071 (based on comparing 14 FinnGen infectious respiratory diseases and five substance use exposures) as a heuristic that allows for follow-up analyses for a plausible number of findings. In assessing consistency and robustness, we looked for estimates substantially agreeing in direction and magnitude (overlapping confidence intervals) across then four complementary MR methods used. We evaluate evidence strength based upon the effect magnitude and direction, the 95% confidence interval of that effect, and the P-value.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All analyses were based upon publicly available data. Single-variable MR and multivariable MR instrument datasets for each substance use behavior required to replicate the findings of this study are available in the Supplemental Data files. Full COVID-19 GWAS summary-level data is available at https://www.covid19hg.org/results/. FinnGen data are available at https://www.finngen.fi/en; lifetime smoking at https://data.bris.ac.uk/data/dataset/10i96zb8gm0j81yz0q6ztei23d; alcohol drinks per week data at: https://genome.psych.umn.edu/index.php/GSCAN; cannabis use disorder and alcohol use disorder data are available through the Psychiatric Genomics Consortium data portal: https://www.med.unc.edu/pgc/download-results/; and the cannabis use data are available through the International Cannabis Consortium at: https://www.ru.nl/bsi/research/group-pages/substance-use-addiction-food-saf/vm-saf/genetics/international-cannabis-consortium-icc/. Coronary artery disease and obesity summary statistics are available through the Cardiovascular Disease Knowledge Portal: https://cvd.hugeamp.org/. Type 2 Diabetes summary-level data is available Type 2 Diabetes Knowledge Portal: https://t2d.hugeamp.org/.
Code availability
We used TwoSampleMR and MendelianRandomization R packages to perform Mendelian Randomization analysis. These packages are publicly available at https://github.com/MRCIEU/TwoSampleMR, and https://github.com/cran/MendelianRandomization, respectively.
References
Puntmann, V. O. et al. Outcomes of cardiovascular magnetic resonance imaging in patients recently recovered from coronavirus disease 2019 (COVID-19). JAMA Cardiol. 5, 1265–1273 (2020).
Shi, S. et al. Association of cardiac injury with mortality in hospitalized patients with COVID-19 in Wuhan, China. JAMA Cardiol. 5, 802–810 (2020).
Nishiga, M., Wang, D. W., Han, Y., Lewis, D. B. & Wu, J. C. COVID-19 and cardiovascular disease: from basic mechanisms to clinical perspectives. Nat. Rev. Cardiol. 17, 543–558 (2020).
Troeger, C. et al. Estimates of the global, regional, and national morbidity, mortality, and aetiologies of lower respiratory infections in 195 countries, 1990–2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet Infect. Dis. 18, 1191–1210 (2018).
Soriano, J. B. et al. Global, regional, and national deaths, prevalence, disability-adjusted life years, and years lived with disability for chronic obstructive pulmonary disease and asthma, 1990–2015: a systematic analysis for the Global Burden of Disease Study 2015. The Lancet. Respiratory Med. 5, 691–706 (2017).
Adams, P. F., Hendershot, G. E. & Marano, M. A. Current estimates from the National Health Interview Survey, 1996. Vital Health. Stat 10, 1–203 (1999).
Simet, S. M. & Sisson, J. H. Alcohol’s effects on lung health and immunity. Alcohol Res. Curr. Rev. 37, 199–208 (2015).
Tashkin, D. P. Effects of Marijuana smoking on the lung. Ann. Am. Thorac. Soc. 10, 239–247 (2013).
Jiang, C., Chen, Q. & Xie, M. Smoking increases the risk of infectious diseases: a narrative review. Tob. Induc. Dis. 18, 60–60 (2020).
Simou, E., Leonardi-Bee, J. & Britton, J. The effect of alcohol consumption on the risk of ARDS: a systematic review and meta-analysis. Chest 154, 58–68 (2018).
Moir, D. et al. A comparison of mainstream and sidestream marijuana and tobacco cigarette smoke produced under two machine smoking conditions. Chem. Res Toxicol. 21, 494–502 (2008).
Ribeiro, L. I. G. & Ind, P. W. Effect of cannabis smoking on lung function and respiratory symptoms: a structured literature review. Respiratory Med. 26, 16071 (2016).
Trevejo-Nunez, G., Kolls, J. K. & de Wit, M. Alcohol use as a risk factor in infections and healing: a clinician’s perspective. Alcohol Res. 37, 177–184 (2015).
Farhoudian, A. et al. A global survey on changes in the supply, price and use of illicit drugs and alcohol, and related complications during the 2020 COVID-19 pandemic. Front. Psychiatry 12, 646206 https://doi.org/10.3389/fpsyt.2021.646206 (2021).
Services. TSAaMHSASotUSDoHaH. T Key Substance Use and Mental Health Indicators in the United States: results from the 2018 National Survey on Drug Use and Health. https://www.samhsa.gov/data/sites/default/files/cbhsq-reports/NSDUHNationalFindingsReport2018/NSDUHNationalFindingsReport2018.pdf. Published 2018 Accessed December 5 2020.
Wang, Q. Q., Kaelber, D. C., Xu, R. & Volkow, N. D. COVID-19 risk and outcomes in patients with substance use disorders: analyses from electronic health records in the United States. Mol. Psychiatry 26, 30–39 (2020).
Smith, G. D. & Ebrahim, S. Epidemiology—is it time to call it a day? Int. J. Epidemiol. 30, 1–11 (2001).
Smith, G. D. & Ebrahim, S. ‘Mendelian randomization’: can genetic epidemiology contribute to understanding environmental determinants of disease? Int. J. Epidemiol. 32, 1–22 (2003).
Evans, D. M., Davey & Smith, G. Mendelian randomization: new applications in the coming age of hypothesis-free causality. Annu. Rev. Genomics Hum. Genet. 16, 327–350 (2015).
Sekula, P., Del Greco, M. F., Pattaro, C. & Köttgen, A. Mendelian randomization as an approach to assess causality using observational data. J. Am. Soc. Nephrol. 27, 3253–3265 (2016).
Goldstein, C. E. et al. Ethical issues in pragmatic randomized controlled trials: a review of the recent literature identifies gaps in ethical argumentation. BMC Med. Ethics 19, 14–14 (2018).
Davey Smith, G. & Hemani, G. Mendelian randomization: genetic anchors for causal inference in epidemiological studies. Hum. Mol. Genet 23, R89–R98 (2014).
Yarmolinsky, J. et al. Association between genetically proxied inhibition of HMG-CoA reductase and epithelial ovarian cancer. JAMA323, 646–655 (2020).
Pingault, J.-B. et al. Using genetic data to strengthen causal inference in observational research. Nat. Rev. Genet. 19, 566–580 (2018).
Arcavi, L. & Benowitz, N. L. Cigarette smoking and infection. Arch. Intern. Med. 164, 2206–2216 (2004).
Jayes, L. et al. SmokeHaz: systematic reviews and meta-analyses of the effects of smoking on respiratory health. Chest 150, 164–179 (2016).
Popkin, B. M. et al. Individuals with obesity and COVID-19: a global perspective on the epidemiology and biological relationships. Obes. Rev. 21, e13128 (2020).
Liang, C., Zhang, W., Li, S. & Qin, G. Coronary heart disease and COVID-19: a meta-analysis. Med. Clin. 156, 547–554 (2021).
McGovern, A. P. et al. The disproportionate excess mortality risk of COVID-19 in younger people with diabetes warrants vaccination prioritisation. Diabetologia 64, 1184–1186 (2021).
Mancuso, P. Obesity and respiratory infections: does excess adiposity weigh down host defense? Pulm. Pharm. Ther. 26, 412–419 (2013).
Morris, A. Heart-lung interaction via infection. Ann. Am. Thoracic Soc. 11, S52–S56. (2014).
Kornum, J. B. et al. Type 2 diabetes and pneumonia outcomes. Diabetes Care 30, 2251 (2007).
Polverino, F. et al. Comorbidities, cardiovascular therapies, and COVID-19 mortality: a nationwide, italian observational study (ItaliCO). Front. Cardiovasc. Med. 7, 585866 (2020).
Vardavas, C. I. & Nikitara, K. COVID-19 and smoking: a systematic review of the evidence. Tob. Induc. Dis. 18, 20–20 (2020).
Cai, G., Bossé, Y., Xiao, F., Kheradmand, F. & Amos, C. I. Tobacco smoking increases the lung gene expression of ACE2, the receptor of SARS-CoV-2. Am. J. Respir. Crit. Care Med 201, 1557–1559 (2020).
Millard, L. A. C., Munafò, M. R., Tilling, K., Wootton, R. E. & Davey Smith, G. MR-pheWAS with stratification and interaction: Searching for the causal effects of smoking heaviness identified an effect on facial aging. PloS Genet. 15, e1008353 (2019).
Larsson, S. C. et al. Smoking, alcohol consumption, and cancer: a mendelian randomisation study in UK Biobank and international genetic consortia participants. PLoS Med. 17, e1003178–e1003178 (2020).
Vie, G. et al. The effect of smoking intensity on all-cause and cause-specific mortality-a Mendelian randomization analysis. Int. J. Epidemiol. 48, 1438–1446 (2019).
Tetrault, J. M. et al. Effects of marijuana smoking on pulmonary function and respiratory complications: a systematic review. Arch. Intern Med. 167, 221–228 (2007).
Bramness, J. G. & von Soest, T. A longitudinal study of cannabis use increasing the use of asthma medication in young Norwegian adults. BMC Pulm. Med. 19, 52 (2019).
Tashkin, D. P. Does marijuana pose risks for chronic airflow obstruction? Ann. Am. Thorac. Soc. 12, 235–236 (2015).
Marees, A. T. et al. Potential influence of socioeconomic status on genetic correlations between alcohol consumption measures and mental health. Psychol. Med. 50, 484–498 (2020).
Karlsson Linnér, R. et al. Genome-wide association analyses of risk tolerance and risky behaviors in over 1 million individuals identify hundreds of loci and shared genetic influences. Nat. Genet. 51, 245–257 (2019).
Patten, C. A., Martin, J. E. & Owen, N. Can psychiatric and chemical dependency treatment units be smoke free? J. Subst. Abus. Treat. 13, 107–118 (1996).
Touchette, J. C. & Lee, A. M. Assessing alcohol and nicotine co-consumption in mice. Oncotarget 8, 5684–5685 (2017).
Gage, S. H., Bowden, J., Smith, G. D. & Munafo, M. R. Investigating causality in associations between education and smoking: a two-sample Mendelian randomization study. Int. J. Epidemiol. 47, 1131–1140 (2018).
Griffith, G. J. et al. Collider bias undermines our understanding of COVID-19 disease risk and severity. Nat. Commun. 11, 5749 (2020).
Elwert, F. & Winship, C. Endogenous selection bias: the problem of conditioning on a collider variable. Annu. Rev. Sociol. 40, 31–53 (2014).
Tattan-Birch, H., Marsden, J., West, R. & Gage, S. H. Assessing and addressing collider bias in addiction research: the curious case of smoking and COVID-19. Addiction 116, 982–984 (2021).
Makoto Miyara, F. T. et al. Low incidence of daily active tobacco smoking in patients with symptomatic COVID-19. Qeios. https://doi.org/10.32388/WPP19W.3 (2020).
The C-HGI. The COVID-19 Host Genetics Initiative, a global initiative to elucidate the role of host genetic factors in susceptibility and severity of the SARS-CoV-2 virus pandemic. Eur. J. Hum. Genet. 28, 715–718 (2020).
Rosenman, R., Tennekoon, V. & Hill, L. G. Measuring bias in self-reported data. Int. J. Behav. Health. Res 2, 320–332 (2011).
Fry, A. et al. Comparison of sociodemographic and health-related characteristics of UK biobank participants with those of the general population. Am. J. Epidemiol. 186, 1026–1034 (2017).
Szabo, G. & Saha, B. Alcohol’s effect on host defense. Alcohol Res. 37, 159–170 (2015).
Rosoff, D. B., Smith, G. D. & Lohoff, F. W. Prescription opioid use and risk for major depressive disorder and anxiety and stress-related disorders: a multivariable Mendelian randomization analysis. JAMA Psychiatry 78, 151–160 (2020).
Hartwig, F. P. et al. Inflammatory biomarkers and risk of schizophrenia: a 2-sample Mendelian randomization study. JAMA Psychiatry 74, 1226–1233 (2017).
Price-Haywood, E. G., Burton, J., Fort, D. & Seoane, L. Hospitalization and mortality among black patients and white patients with Covid-19. N. Engl. J. Med. 382, 2534–2543 (2020).
Yancy, C. W. COVID-19 and African Americans. JAMA 323, 1891–1892 (2020).
Niedzwiedz, C. L. et al. Ethnic and socioeconomic differences in SARS-CoV-2 infection: prospective cohort study using UK Biobank. BMC Med. 18, 160 (2020).
Burgess, S., Davies, N. M. & Thompson, S. G. Bias due to participant overlap in two-sample Mendelian randomization. Genet Epidemiol. 40, 597–608 (2016).
Minelli, C. et al. The use of two-sample methods for Mendelian randomization analyses on single large datasets. Int. J. Epidemiol. https://academic.oup.com/ije/advance-article/doi/10.1093/ije/dyab084/6252978 (2021).
Hemani, G., Tilling, K. & Smith, G. D. Orienting the causal relationship between imprecisely measured traits using GWAS summary data. Plos Genet. 13, 11 (2017).
Wootton, R. E. et al. Evidence for causal effects of lifetime smoking on risk for depression and schizophrenia: a Mendelian randomisation study. Psychol. Med. 50, 2435–2443 (2020).
Wootton, R. E. et al. Evidence for causal effects of lifetime smoking on risk for depression and schizophrenia: a Mendelian randomization study. Psychol. Med. 50, 2435 (2020).
Pasman, J. A. et al. GWAS of lifetime cannabis use reveals new risk loci, genetic overlap with psychiatric traits, and a causal effect of schizophrenia liability. Nat. Neurosci. 21, 1161–1170 (2018).
Pasman, J. A. et al. GWAS of lifetime cannabis use reveals new risk loci, genetic overlap with psychiatric traits, and a causal effect of schizophrenia liability. Nat. Neurosci. 22, 1196 (2018).
Johnson, E. C. et al. A large-scale genome-wide association study meta-analysis of cannabis use disorder. Lancet Psychiatry 7, 1032–1045 (2020).
Johnson, E. C. et al. A large-scale genome-wide association study meta-analysis of cannabis use disorder. Lancet Psychiatry 7, 1032 (2020).
Liu, M. et al. Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat. Genet. 51, 237–244 (2019).
Liu, M. et al. Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat. Genet. 51, 237 (2019).
Walters, R. K. et al. Transancestral GWAS of alcohol dependence reveals common genetic underpinnings with psychiatric disorders. Nat. Neurosci. 21, 1656–1669 (2018).
Walters, R. K. et al. Transancestral GWAS of alcohol dependence reveals common genetic underpinnings with psychiatric disorders. Nat. Neurosci. 21, 1656 (2018).
Pub, A. P. Diagnostic and Statistical Manual of Mental Disorders: DSM-IV-TR. (Washington, DC, 2000).
van der Harst, P. & Verweij, N. Identification of 64 novel genetic loci provides an expanded view on the genetic architecture of coronary artery disease. Circ. Res. 122, 433–443 (2018).
van der Harst, P. & Verweij, N. Identification of 64 novel genetic loci provides an expanded view on the genetic architecture of coronary artery disease. Circ. Res. 122, 433 (2018).
Xue, A. et al. Genome-wide association analyses identify 143 risk variants and putative regulatory mechanisms for type 2 diabetes. Nat. Commun. 9, 2941 (2018).
Xue, A. et al. Genome-wide association analyses identify 143 risk variants and putative regulatory mechanisms for type 2 diabetes. Nat. Commun. 9, 2941 (2018).
Berndt, S. I. et al. Genome-wide meta-analysis identifies 11 new loci for anthropometric traits and provides insights into genetic architecture. Nat. Genet 45, 501–512 (2013).
Berndt, S. I. et al. Genome-wide meta-analysis identifies 11 new loci for anthropometric traits and provides insights into genetic architecture. Nat. Genet. 45, 501 (2013).
Sanderson, E., Spiller, W. & Bowden, J. Testing and correcting for weak and pleiotropic instruments in two-sample multivariable Mendelian randomisation. Stat. Med. 40, 5435–5452 (2021).
COVID-19 Host Genetics Initiative. Mapping the human genetic architecture of COVID-19. Nature (2021).
FinnGen. FinnGen Documentation of the R5 release. Accessed 15 April 2021. https://finngen.gitbook.io/documentation/ (2021).
Burgess, S. & Thompson, S. G. Multivariable Mendelian randomization: the use of pleiotropic genetic variants to estimate causal effects. Am. J. Epidemiol. 181, 251–260 (2015).
Minelli, C. et al. The use of two-sample methods for Mendelian randomization analyses on single large datasets. Int. J. Epidemiol. https://doi.org/10.1093/ije/dyab084 (2021).
Hemani, G. et al. The MR-Base platform supports systematic causal inference across the human phenome. Elife. 7, e34408 (2018).
Yavorska, O. O. & Burgess, S. MendelianRandomization: an R package for performing Mendelian randomization analyses using summarized data. Int. J. Epidemiol. 46, 1734–1739 (2017).
Bowden, J. et al. A framework for the investigation of pleiotropy in two-sample summary data Mendelian randomization. Stat. Med. 36, 1783–1802 (2017).
Bowden, J. et al. Improving the accuracy of two-sample summary-data Mendelian randomization: moving beyond the NOME assumption. Int. J. Epidemiol. 48, 728–742 (2019).
Rees, J. M. B., Wood, A. M. & Burgess, S. Extending the MR-Egger method for multivariable Mendelian randomization to correct for both measured and unmeasured pleiotropy. Stat. Med 36, 4705–4718 (2017).
Verbanck, M., Chen, C. Y., Neale, B. & Do, R. Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases (vol 50, 693, 2018). Nat. Genet. 50, 1196–1196 (2018).
Zhu, Z. et al. Causal associations between risk factors and common diseases inferred from GWAS summary data. Nat. Commun. 9, 224 (2018).
Acknowledgements
We want to acknowledge the participants and investigators of the many studies used in this research without whom this effort would not be possible: the COVID-19 Host Genetics Initiative and the contributors thereto specified at http://www.covid19hg.org/acknowledgments.html, the FinnGen study, and the UK Biobank. We also want to acknowledge the Medical Research Council Integrative Epidemiology Unit (MRC-IEU, University of Bristol, UK), especially the developers of the MRC-IEU UK Biobank GWAS Pipeline. We also want to acknowledge the participants and investigators of FinnGen study. This work was supported by the National Institutes of Health (NIH) intramural funding [ZIA-AA000242 to F.W.L]; Division of Intramural Clinical and Biological Research of the National Institute on Alcohol Abuse and Alcoholism (NIAAA).
Author information
Authors and Affiliations
Contributions
D.B.R. and J.Y. generated the data. All authors contributed to the analysis and interpretation. D.B.R. and F.W.L conceptualized and designed the study. F.W.L. obtained funding, provided supervision, and critical oversight to data collection and study coordination. D.B.R. and J.Y. wrote the manuscript. All authors contributed to critical editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Communications Biology thanks Zhaozhong Zhu and the other, anonymous, reviewers for their contribution to the peer review of this work. Primary Handling Editors: Chiea Chuen Khor and George Inglis.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rosoff, D.B., Yoo, J. & Lohoff, F.W. Smoking is significantly associated with increased risk of COVID-19 and other respiratory infections. Commun Biol 4, 1230 (2021). https://doi.org/10.1038/s42003-021-02685-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-021-02685-y
This article is cited by
-
Risk phenotypes of diabetes and association with COVID-19 severity and death: an update of a living systematic review and meta-analysis
Diabetologia (2023)
-
Cannabis use, abuse and dependence during the COVID-19 pandemic: a scoping review
Journal of Neural Transmission (2023)
-
Can smoking prevalence explain COVID-19 indicators (cases, mortality, and recovery)? A comparative study in OECD countries
Environmental Science and Pollution Research (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.