Abstract
Deployment of Wolbachia to mitigate dengue (DENV), Zika (ZIKV) and chikungunya (CHIKV) transmission is ongoing in 12 countries. One way to assess the efficacy of Wolbachia releases is to determine invasion rates within the wild population of Aedes aegypti following their release. Herein we evaluated the accuracy, sensitivity and specificity of the Near Infrared Spectroscopy (NIRS) in estimating the time post death, ZIKV-, CHIKV-, and Wolbachia-infection in trapped dead female Ae. aegypti mosquitoes over a period of 7 days. Regardless of the infection type, time post-death of mosquitoes was accurately predicted into four categories (fresh, 1 day old, 2–4 days old and 5–7 days old). Overall accuracies of 93.2, 97 and 90.3% were observed when NIRS was used to detect ZIKV, CHIKV and Wolbachia in dead Ae. aegypti female mosquitoes indicating NIRS could be potentially applied as a rapid and cost-effective arbovirus surveillance tool. However, field data is required to demonstrate the full capacity of NIRS for detecting these infections under field conditions.
Similar content being viewed by others
Introduction
Almost half of the global human population currently live in areas under risk of arbovirus transmission1. The incidence of mosquito-borne arboviruses, such as dengue (DENV), Zika (ZIKV), and chikungunya (CHIKV) has been on the rise globally in the past decade due to increased geographical spread of the primary vector Aedes aegypti and the secondary vector Aedes albopictus to places where they were originally absent2. Arboviruses outbreaks are linked to favorable climatic conditions, entomological, epidemiological, and immunological factors. The absence of an effective vaccine and a lack of effective and timely vector surveillance system are major contributors of rapid arbovirus spread3, which ultimately pose a high economic burden, including severe disability-adjusted life year to affected populations4,5,6.
Monitoring the spatial and temporal dynamics of vector populations through larval surveys remains a routine activity in many countries7. Although sampling adult mosquitoes provides more reliable estimates of Aedes population size and allows stratifying areas according to the risk of outbreaks8,9, estimation of transmission risk rely mostly on monitoring vector density as opposed to virus density in wild mosquitoes. Determining arbovirus infection in wild caught Aedes mosquitoes is rarely done by most vector control programs. This is because the positivity rate of captured mosquitoes is in most cases <1% and therefore setting up a large-scale routine surveillance system to detect such a natural infection in trapped mosquitoes is costly, time consuming, and unfeasible in several endemic countries10,11,12,13,14. Nonetheless, predicting when and where the next epidemic will strike is of utmost importance to facilitate vector control intensification in areas where risk is highest. Hence, surveillance methods should be capable of providing rapid detection of arboviruses required to initiate an effective response against arboviral threats15. In such a scenario, a rapid, accurate, and cost-effective tool will facilitate early arbovirus detection in trapped mosquitoes to trigger early warning systems, which will in turn initiate a timely intervention16.
One of the most promising ongoing intervention against arboviruses is releasing A. aegypti mosquitoes transinfected with the maternally inherited endosymbiotic bacterium Wolbachia pipientis in the field. This bacterium works by manipulating host reproduction through cytoplasmic incompatibility resulting in nonviable offspring, when uninfected females mate with Wolbachia-infected males17,18,19. Wolbachia-infected A. aegypti can block transmission of several arboviruses, including DENV18,20,21, ZIKV22,23, CHIKV, Mayaro, and yellow fever virus24,25. The ongoing use of Wolbachia in >14 countries is based on replacing the highly susceptible native A. aegypti population with the less susceptible Wolbachia-infected A. aegypti population to ameliorate arboviral transmission26,27,28. Population replacement is achieved after releasing hundreds of thousands of Wolbachia-infected A. aegypti mosquitoes. The prevalence of Wolbachia in the field is assessed through screening trapped mosquitoes. Such information is intended to guide subsequent releases. For example, whether to increase or reduce the number of insects in future releases or even stop the release if a desired frequency has been achieved. Traditionally, adult mosquito trapping is carried out with BG-Sentinel traps, but ovitraps are sometimes used concomitantly, usually on a weekly basis26,27,29. Collected samples (adult mosquitoes or eggs allowed to hatch in an entomological lab) are often screened with quantitative polymerase chain reaction (qPCR) technique to check for the presence of Wolbachia, but arboviruses presence in Wolbachia-infected and wild A. aegypti is rarely assessed27. Traps are inspected once a week in most cases, and thus the majority of trapped mosquitoes have died and dried out for period of time varying between 1 and 7 days27. This presents a big challenge in the detection of arboviruses and bacteria using conventional molecular techniques, such as PCR30.
The near-infrared spectroscopy (NIRS) technique has been demonstrated in a number of entomological studies as rapid, cost-effective, and high-throughput tool for characterizing biological samples based on spectral signatures. The spectral signatures mirror the amount of light reflected back following its absorption by C–H, O–H, S–H, and N–H functional groups present in those samples at specific frequencies. The type and concentration of these chemicals are unique to biological samples, and hence each unique sample can produce a specific diagnostic spectrum that can be analyzed by chemometrics or machine learning techniques to identify them. For this purpose, NIRS is: (a) non invasive, therefore the material can be used multiple times, (b) low cost, as it does not require reagents to operate and (c) rapid, as a spectrum can be collected in just 3 s, therefore allowing hundreds of samples to be analyzed daily. NIRS has been demonstrated for age prediction, species identity and for the detection of the presence of Plasmodium falciparum of the major African malaria vectors, Anopheles gambiae and Anopheles arabiensis under laboratory, semi-field, and field settings31,32,33,34. NIRS has also been used to predict the age of A. aegypti35, A. albopictus36, and to detect ZIKV in A. aegypti37, and Wolbachia in A. aegypti38 and in fruit flies39.
As a potential next-generation surveillance tool, NIRS could provide a reliable and accurate alternative to age grading, and diagnosis of arboviruses and Wolbachia in A. aegypti. Furthermore, NIRS could potentially be used to evaluate the spatiotemporal shifts in arbovirus transmission following Wolbachia deployment to allow health managers to rapidly and cost effectively assess the impacts of Wolbachia in reducing disease transmission and outbreaks. So far, the accuracy of NIRS for age grading and pathogen detection has mainly been demonstrated on fresh or preserved samples40,41. Therefore, the value of NIRS for detecting pathogens in mosquitoes that have been dead in a trap for a period of 7 days is yet to be demonstrated. To develop accurate NIRS models for predicting the presence of viruses and Wolbachia in a dead mosquito, ideally the approximate time of death should be known. The main objective of this study was to determine whether NIRS can estimate the approximate death time, and use this information to predict the presence of ZIKV, CHIKV, and Wolbachia in mosquitoes left in a BG-Sentinel trap for a period of 7 days post death.
Results
Confirmation of ZIKV, CHIKV, and Wolbachia in mosquitoes
Out of the 157, 59, and 163 mosquitoes screened for ZIKV, CHIKV, and Wolbachia, infection was confirmed in 82 (52.3%), 36 (61%), and 163 (100%) of them, respectively. At 7 days post death, 53 (54.6%) of ZIKV-infected mosquitoes were still positive for CHIKV.
NIRS prediction of days post death
The training and testing model consisted of ZIKV (n = 157), CHIKV (n = 59), Wolbachia (n = 163), and uninfected mosquitoes (n = 129). Monte Carlo simulations were performed using a 75%/25% training/testing split to validate the model. The mean absolute error (MAE; standard deviation) for prediction of days post death in the training group was 1.16 ± 0.88, 1.24 ± 0.93, 1.18 ± 0.93, and 1.15 ± 0.82 for ZIKV, CHIKV, Wolbachia, and control mosquitoes, respectively (Fig. 1a). MAE (standard deviation) for days post death in the testing group were 1.25 ± 0.98, 1.27 ± 0.95, 1.34 ± 1.71, and 1.15 ± 0.84 for ZIKV, CHIKV, Wolbachia, and control mosquitoes, respectively (Fig. 1b). When considering the accuracy of NIRS for predicting days post death of all mosquitoes regardless of their infection type, they all grouped into four categories (fresh, 1 day old, 2–4 days old and 5–7 days old). Predicted scores for each group for all infections followed a similar trend, indicating that infection type does not affect NIRS days post death prediction accuracy (Fig. 2).
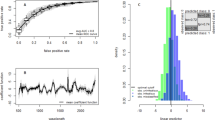
Results from training data (a) and results from testing data (b). Monte Carlo simulations were performed using a 75%/25% training/testing split to validate the model. Infection status including CHIKV (red), ZIKV (purple), Wolbachia (blue), and uninfected controls (green) is presented in both panels. Box and whisker plots follow the standard convention where the box represents the range between quartiles 1 and 3, while the whiskers represent maximum and minimum excluding outliers. Outliers are defined as either Q1 − 1.5 × IQR or Q3 + 1.5 × IQR.
NIRS prediction of CHIKV infection
Models for predicting CHIKV were the most accurate, ranging from 95.1% to 99.8% accuracy depending on days post death the mosquitoes were scanned (Fig. 3a and Table 1). The average accuracy for predicting the presence or absence of CHIKV in mosquitoes regardless of when they died was 97%. Mosquitoes that were scanned on day 1 and day 7 post death were least accurately predicted, whereas mosquitoes that were scanned 5–6 days post death were predicted with a slightly higher accuracy. However, this accuracy did not differ statistically from the accuracy of fresh mosquitoes for all groups (P = 0.261). The overall sensitivity of NIRS for predicting CHIKV in mosquitoes regardless of time of death of a mosquito was 95.8%. This sensitivity ranged from 91.6% to 99.2% depending on the time post death the mosquitoes were scanned, and it was similar to the sensitivity observed for fresh mosquitoes (P = 1 in all cases). Similarly, specificity ranged from 95.6% to 99.8% depending on time post death the mosquitoes were scanned, and it did not significantly differ from fresh mosquitoes for all groups (P = 0.269; Table 1).
Mosquitoes infected with CHIKV (a), Wolbachia (b), and ZIKV (c) vs. uninfected mosquitoes. Box and whisker plots follow the standard convention where the box represents the range between quartiles 1 and 3, while the whiskers represent maximum and minimum excluding outliers. Outliers are defined as either Q1 − 1.5*IQR or Q3 + 1.5*IQR.
NIRS prediction of Wolbachia infection
The prediction accuracy of NIRS for detecting Wolbachia in A. aegypti ranged between 85.8% and 95.3% for the testing cohort (Fig. 3b and Table 1) with the average accuracy of 90.3%. The accuracy was not statistically different for all tested groups relative to fresh mosquitoes (P = 0.722). Mosquitoes were more accurately predicted (accuracy 95.3%) when scanned 7 days post death. Similarly, sensitivity and specificity values were higher when mosquitoes were scanned 7 days post death. Although not significantly different from fresh mosquitoes, sensitivity and specificity for Wolbachia detection increased when mosquitoes were scanned 4 days post death, and the overall sensitivity and specificity regardless of when mosquitoes were scanned was 81.9% and 95.7%, respectively (Fig. 3c and Table 1).
NIRS prediction of ZIKV infection
Regardless of time post death, the average prediction accuracy of NIRS for detecting ZIKV was 93.2% for all mosquitoes scanned for the testing set. This accuracy ranged between 90.7% and 95.1% depending on the time post death mosquitoes were scanned (Fig. 3c and Table 1). The accuracy of predicting mosquitoes 5 and 7 days post death was higher and significantly different from the accuracy obtained when mosquitoes were scanned fresh (P = 0.03). The highest paired sensitivity and specificity were 92.0% and 96.5%, respectively, and they were achieved when mosquitoes were scanned 5 days post death. Overall sensitivity and specificity of 88.5% and 96.1% were achieved regardless of when the mosquitoes were scanned (Table 1).
Prediction of infection in fresh vs. mosquitoes that were scanned 7 days post death
When comparing overall accuracy, sensitivity, and specificity of ZIKV, CHIKV, and Wolbachia-infected mosquitoes scanned at 0 days post death (fresh) and those scanned 7 days post death, no difference was observed for CHIKV-infected mosquitoes. However, ZIKV and Wolbachia-infected mosquitoes were more accurately predicted 7 days post death than when they were scanned while fresh (Table 1 and Fig. 4).
Discussion
An efficient arbovirus surveillance system monitors the early signs of viral circulation in human population, which would enable a timely response to prevent future disease outbreaks. The development of a rapid and cost-effective tool to identify the presence of arboviruses in mosquitoes is crucial for control of these arboviruses16,42,43, and can minimize disease outbreaks and assist in the timely reinforcement of vector control interventions44,45,46,47. Secondly, a tool that can detect infection in mosquitoes that have been left in a trap for an extended period of time, for example, a week, will reduce the time and costs required to retrieve these samples on a daily basis. In this manuscript, we addressed (1) the capacity of NIRS to predict the time post death of female A. aegypti mosquitoes left in a BG-Sentinel trap for a period of 0–7 days, (2) the ability of NIRS to detect ZIKV, CHIKV, and Wolbachia in A. aegypti mosquitoes left in a BG-Sentinel trap for 0–7 days, and (3) whether time post death of mosquitoes affected the capacity of NIRS to detect infections in mosquitoes.
In this study, we obtained 93.2, 97.0, and 90.3% accuracy when NIRS was used to predict ZIKV, CHIKV, and Wolbachia infection in female A. aegypti mosquitoes regardless of their time of death. More importantly, NIRS prediction of these infections 7 days post death was not significantly different from fresh mosquitoes. Surprisingly, NIRS showed higher accuracy than RT-qPCR for detecting ZIKV infection in dead mosquitoes. This could be likely due to the degradation of viral RNA at insectary temperature. We hypothesize that the rate of water loss in a mosquito in the presence of an arbovirus might be slower than viral RNA degradation. This allows NIRS to detect these viruses more efficiently than molecular-based approaches, such as the RT-qPCR48,49. NIRS has previously predicted the presence of ZIKV in heads and thoraces of female A. aegypti mosquitoes with 97% accuracy37, malaria parasites in A. gambiae with 95% accuracy50, and the presence of wMel and wMelPop in female A. aegypti mosquitoes with 93% and 96% accuracy, respectively38.Previous studies used either fresh37,50 or RNAlater-preserved mosquitoes38 that were reared, infected, and maintained in a temperature-controlled environment and killed with chemicals. Herein, we simulated a more natural setting whereby A. aegypti females were killed by shaking the trap, and then left in a BG-Sentinel trap for a period of 0–7 days following their death.
Slightly higher accuracy was observed for CHIKV-infected A. aegypti than for ZIKV-infected mosquitoes (Table 1). This difference in accuracy could be attributed to specific viral changes in mosquitoes following an infection44,51,52, initial titers used to infect mosquitoes and the overall chemical composition of a mosquito following an infection. These factors ultimately affect the diagnostic signature and the accuracy. We noted no linear changes on the predicted accuracy or sensitivity for all infections over the 7-day sampling period. For instance, the highest accuracy for ZIKV, CHIKV, and Wolbachia was observed 2, 6, and 7 days post death, respectively. The authors speculate that the rate of water loss following the death of a mosquito could be relative to the type of infection carried by that mosquito because at 7 days post death, all infections were predicted with a relatively higher accuracy compared to fresh mosquitoes possibly indicating the peak of water loss regardless of the infection type. Nonetheless, the >90% accuracy of NIRS for detecting ZIKV or CHIKV in trapped A. aegypti several days post death was impressive, and indicates the potential future application of the tool for this purpose.
The release of Wolbachia-infected A. aegypti has the potential to become a key component of vector control in the future. In the past decade, Wolbachia has been released in small isolated sites and in big cities. Rapid assays to evaluate its effect on a large scale is crucial for assessing its invasion rate27,28,53. Herein, we have demonstrated the ability of NIRS in detecting Wolbachia in female A. aegypti up to 7 days post death. Although >85% accuracy was achieved over the 7-day sampling period, the accuracy of NIRS in detecting Wolbachia was slightly lower than that for CHIKV- and ZIKV-infected A. aegypti. We hypothesize this could related to the differences between immunological responses against a virus and a bacterium by a mosquito, as well as the distribution of the virus and the bacteria inside the mosquito. The wMel strain of the endosymbiont Wolbachia is present in higher densities in specific tissues, notably in the ovaries54. Spectra of infected and uninfected mosquitoes were collected from the heads and thoraces where the density of Wolbachia is still significant, but in lower densities than ovaries to mitigate the effects of abdominal contents on the spectra, which could have had an effect on the accuracy of Wolbachia-infected mosquitoes. Future work should investigate whether scanning other parts of the mosquito improves the current accuracy, and whether Wolbachia can also be detected in male mosquitoes several days post death of a mosquito.
Monitoring arboviruses circulation in trapped mosquitoes could significantly enhance the knowledge regarding silent transmission of DENV, ZIKV, and CHIKV in endemic settings45,55,56,57. We define silent transmission as infections that result in either asymptomatic cases or mild symptoms that are undetected by routine surveillance systems58. Silent infections hold a significant role in arbovirus transmission because these infections could account for >80% of DENV transmission58. Such findings could trigger the deployment of a trap-based surveillance system, whereby rapid detection of pathogens in trapped mosquitoes could inform health managers to conduct cross-sectional serosurveys on nearby households. By doing so, integrated vector management practices, such as social mobilization, empowerment of communities, intensification of vector control, and an assertive case management approaches could locally impact disease transmission.
Mosquitoes used in this experiment were killed by shaking to mimic natural death, i.e., without chemicals. The mosquitoes were added to the catching bag of a BG-Sentinel trap, a harsh microenvironment produced by the constantly blowing fan intended to retain trapped mosquitoes. NIRS predicted the time post death based on the reduction of moisture/water content over time. This can be observed in the spectra shown in Fig. 2, where fresh samples are characterized by broad overtones of water related peaks ~1450 and 1950 nm, and whereby these peaks start to diminish 7 days post death.
The results presented herein are a first step toward the development of a robust arboviruses surveillance system that could be used to predict infections in mosquitoes at different time points. Arboviruses detection under field settings would require additional experiments to account for fluctuating environmental conditions in the field, while mosquitoes are trapped. Furthermore, we acknowledge the fact that a single cohort of mosquitoes was used to develop predictive models for CHIKV. Additional experiments assessing this effect would be essential prior to utilization of NIRS within a field setting. It is expected that current models for ZIKV, CHIKV, and Wolbachia would require to be modified by including field samples in the training model to capture confounding factors, such as diet, microbiome, fluctuating temperature, and humidity prior to field application59. Therefore, a direct extrapolation of our results to a field setting must be avoided at this point. Nonetheless, these results are encouraging and will accelerate progress toward developing a rapid and high-throughput arbovirus surveillance system.
Methods
Mosquitoes
Mosquitoes were collected from two sites in Rio de Janeiro, Brazil. To represent the native field population, i.e., those with and without Wolbachia. Wolbachia-free mosquitoes were collected at Urca neighborhood (22° 56′ 43″ S; 43° 09′ 42″ W), whereas those infected with Wolbachia were collected in Tubiacanga (22° 47′ 06″ S; 43° 13′ 32″ W), an isolated village in Rio where Wolbachia was first deployed. Infection rates in Tubiacanga are close to 100% (ref. 27). To capture the local genetic diversity for both sites, collection was done using 60 ovitraps homogeneously distributed over the area60. Eggs were hatched and larvae reared at the insectarium of the Laboratório de Mosquitos Transmissores de Hematozoários, Fiocruz. Larvae were fed with TetraMin fish flakes (Tetra GmbH, Melle, Germany) until pupation. Pupae were transferred to cages measuring 40 cm3 and emerging adults received 10% sugar solution ad libitum. Adults were maintained in an insectary under 27 ± 2° C, 70 ± 5% relative humidity and a 12:12 h light:dark cycling period. For the experiment described below, four groups of mosquitoes were used: Wolbachia-infected, CHIKV-infected, ZIKV-infected, and wild-type (uninfected) A. aegypti.
Experimental blood feeding and infection
F1 female A. aegypti that were 5–6 days old were orally fed with 1 ml of ZIKV-infected or CHIKV-infected C6/36 cells supernatant mixed with 2 ml of human blood. We used the currently circulating strain of the Brazilian ZIKV [BRPE243/2015 (BRPE)]61, which was isolated from a ZIKV-infected patient in late 2015 and maintained in cell culture. The East–Central–South African CHIKV genotype deposited in the GenBank under accession nos. KP164567–KP164572, which was isolated from a patient in 2014 (ref. 62) was used. Viral titers were quantified via plaque forming assay for both ZIKV and CHIKV prior to infection.
The oral infection procedures were performed through a membrane feeding system (Hemotek, Great Harwood, UK) adapted with a pig-gut covering, which gives access to the human blood. The ZIKV viral titer used for mosquitoes was 1.9 × 106 PFU (plaque formation units)/ml and the CHIKV viral titer was 6.3 × 105 PFU/ml. Uninfected and Wolbachia-infected mosquitoes were fed similarly with uninfected blood and C6/36 cell culture. At least 40 mosquitoes were used for each experiment type. Mosquitoes were incubated for 7 days to allow the virus to replicate within mosquito body51,52 and to allow oviposition to take place. Results presented herein were obtained from three independent experiments for Wolbachia, three for ZIKV, and from a single experimental infection for CHIKV. Uninfected mosquitoes were used in every single assay.
Sample preparation and collection of NIRS spectra
Following a 7-day incubation period, mosquitoes were killed by repeated shaking to avoid use of chemicals that could influence spectra characteristics. Mosquitoes were shaken for 5 min, with a 5 min pause to observe recovery, followed by additional 5 min of shaking if needed. Mosquitoes were individually transferred to a grid with 20, 1.5 ml open ended plastic tubes covered with nylon mesh on both ends. The mesh allowed air circulation and simulated the air fan that operates continuously in a BG-Sentinel trap. Mosquitoes in the grid were placed into the catch bag of the BG-Sentinel trap for 7 days. The BG-Sentinel mosquito traps remained side-by-side in the insectary during the experiment, with standard conditions (temperature: 27 ± 2° C; relative humidity: 70 ± 5%; 12:12 h light:dark cycling period).
Spectra collection
Once a day for the 7-day period, mosquitoes remained in the trap, A. aegypti females were removed from the plastic grid for scanning with NIRS. By doing so, we were able to determine the limit of detection of NIRS for ZIKV, CHIKV, or Wolbachia infection in individually trapped A. aegypti mosquitoes. The insects were arranged sideways on a Spectralon diffuse reflection stage, and their heads and thoraces were scanned with a Labspec 4i NIRS spectrometer (Malvern Panalytical, Longmont, CO) using an external 3.2 mm diameter fiber optic probe and a 18, 6 W light source (Model 135325 Rev B, ASD Inc.) according to previously published protocols31. Spectra collection was on average 3–5 s per sample.
Confirmation of ZIKV, CHIKV, and Wolbachia in mosquitoes
Infected mosquitoes were screened by qPCR to assess their infection status. A subset of 59, 157 CHIKV-infected and ZIKV-infected mosquitoes were assessed using RT-qPCR on legs before they were placed into the BG-Sentinel. A second RT-qPCR was conducted on 97 ZIKV-infected mosquitoes following the 7-day trapping period. Viral RNA was extracted with a QIAamp Mini Viral RNA Kit (Qiagen). Detection and quantification of viral RNA in legs from each individual was performed by RT-qPCR with the SuperScript III Platinum Single-Step qRT-PCR Kit (Invitrogen), using the QuantStudio 6 Flex Real-Time PCR System (Applied Biosystems) according to published protocols with known primers and amplification conditions63,64. Viral copy numbers were calculated by absolute quantitation for each run, using a curve pattern in a six-point dilution series (101–106 copies) of in vitro ZIKV and CHIKV RNA transcripts61. The legs of Wolbachia-infected mosquitoes were not screened for Wolbachia since wMel strain is known to be found in higher densities in A. aegypti ovaries and salivary glands compared to their legs20. A subset of 43 mosquito bodies were screened for Wolbachia by RT-qPCR after the 7 days in the BG-Sentinel trap. DNA extraction from A. aegypti body involved the addition of 50 μl buffer (10 mM Tris, 1 mM EDTA, 50 mM NaCl, pH 8.2) and 2 μl proteinase K followed by maceration for 30 s. Samples were incubated at 56° C and then at 98° C for 10 min each. A mixture containing the rps primers for mosquitoes and wsp for Wolbachia was used in the PCR mixture. Each agarose gel electrophoresis reaction contained a Wolbachia positive control (mosquitoes from a lab colony whose infection has been confirmed by PCR), a PCR-confirmed negative control, and a blank sample (distilled water). Extraction and PCR protocols were performed following published materials20,27,29. The PCR data was used as the gold standard, i.e., a mosquito that received an infected blood meal with a negative PCR result, was considered negative.
Data analysis
Spectral data was analyzed in R version 3.6.2 (Dark and Stormy Night)65 using only samples confirmed positive by PCR. Partial least squares (PLS) regression was performed using the package “pls” and summary statistics were generated using the “caret” package. Data were initially encoded as reflectance and was converted to absorbance using Eq. (1).
Equation (1): conversion from reflectance to absorbance. A is the absorbance and R is the reflectance.
Death grading
PLS regression was employed to predict the number of days post death for each sample (0–7 days). A second PLS regression model was developed to predict if samples fit into four categories: (a) freshly killed (day 0), (b) 1 day, (c) 2–4 days, or (d) >4 days post death. These groups were denoted as 0, 1, 2, and 3, respectively. Data were split into training (75%) and testing (25%) groups. The split was performed on a mosquito by mosquito basis, as such data from each mosquito for all days were assigned to either the training or testing set as a group.
The optimal number of factors used was identified computationally by optimizing the accuracy within the training dataset. K-fold cross validation (k = 5, reps = 10) was used to simultaneously train the optimal factor level and parameter weights. The model was optimized by minimizing the PLS root mean squared error (RMSE) between actual days post death and predicted days post death. To facilitate interpretation of results, the MAE instead of RMSE is reported for both the training and testing cohorts.
Infectivity prediction
Partial least squares discriminative analysis (PLSDA) was employed for binary classification analyses, including infectivity prediction for ZIKV-infected vs. uninfected, Wolbachia-infected vs. uninfected, and CHIKV-infected vs. uninfected. PLSDA was also employed to develop a multi-class classification model simultaneously for ZIKV, Wolbachia, CHIKV, and uninfected samples. Data were analyzed individually day by day (i.e., one model for samples on day 0, another for day 1, etc.) and balanced for infectivity status within each cohort independently involving equal number of infected and uninfected samples per cohort. All samples not used to train the model were used to test the model. Infectivity was encoded using one-hot encoding (1 for infected, 0 for uninfected). Optimal factor level identification and parameter weight tuning was performed using the same method, as described for death grading.
One final model was generated to simultaneously differentiate between all conditions present in the study. The process for generating this model is similar to that used to generate the models for differentiating between any one disease and uninfected mosquitoes, however, instead of using one-hot encoding to specify disease as a binary outcome measure, the outcome variable was a multilevel categorical variable (ZIKV, CHIKV, Wolbachia, and uninfected). Accuracy instead of RMSE was used to optimize these models.
Accuracy, sensitivity, and specificity for the training and testing sets were reported where prediction scores >0.5 are considered infected and prediction scores <0.5 are considered uninfected.
Monte Carlo simulation
To ensure the robustness of the models, Monte Carlo simulations were employed, and the procedure described above was repeated 50 times, randomly assigning mosquitoes to the training and testing groups differently in each repetition. Results reported describe aggregated results from all runs of the Monte Carlo simulation.
Statistics and reproducibility
Statistical analysis was conducted on R (version 3.6.2—Dark and Stormy Night), using the packages “pls” and “caret” (as described above). The sample sizes for each infection treatment is described above and was based on previous experiments conducted at corresponding author’s laboratory to investigate the presence of ZIKV in freshly infected A. aegypti37. Methodology and biological materials are disclosed as much as possible, but if required, further information can be obtained by contacting from the corresponding author. The replicates of infected and control mosquitoes used in a given experimental infection were hatched at different days and raised under the same controlled conditions.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
References
Kraemer, M. U. G. et al. The global distribution of the arbovirus vectors Aedes aegypti and Ae. albopictus. Elife 4, 1–18 (2015).
Schaffner, F. & Mathis, A. Dengue and dengue vectors in the WHO European region: past, present, and scenarios for the future. Lancet Infect. Dis. 14, 1271–1280 (2014).
Yakob, L. & Walker, T. Zika virus outbreak in the Americas: the need for novel mosquito control methods. Lancet Glob. Health 4, e148–e149 (2016).
LaBeaud, A., Bashir, F. & King, C. H. Measuring the burden of arboviral diseases: the spectrum of morbidity and mortality from four prevalent infections. Popul. Health Metr. 9, 1 (2011).
Shepard, D. S., Undurraga, E. A. & Halasa, Y. A. Economic and disease burden of dengue in Southeast Asia. PLoS Negl. Trop. Dis. 7, e2055 (2013).
Bartsch, S. M. et al. What is the value of different Zika vaccination strategies to prevent and mitigate Zika Outbreaks? J. Infect. Dis. 220, 920–931 (2019).
Codeço, C. T. et al. Surveillance of Aedes aegypti: comparison of house index with four alternative traps. PLoS Negl. Trop. Dis. 9, e0003475 (2015).
Villela, D. A. M., de A Garcia, G. & Maciel-de-Freitas, R. Novel inference models for estimation of abundance, survivorship and recruitment in mosquito populations using mark-release-recapture data. PLoS Negl. Trop. Dis. 11, e0005682 (2017).
Villela, D. A. M. et al. A Bayesian hierarchical model for estimation of abundance and spatial density of Aedes aegypti. PLoS ONE 10, e0123794 (2015).
Juliano, S. A. et al. She’s a femme fatale: low-density larval development produces good disease vectors. Mem. Inst. Oswaldo Cruz 109, 1070–1077 (2014).
Pérez-Castro, R. et al. Detection of all four dengue serotypes in Aedes aegypti female mosquitoes collected in a rural area in Colombia. Mem. Inst. Oswaldo Cruz 111, 233–240 (2016).
Dutta, P. et al. First evidence of dengue virus infection in wild caught mosquitoes during an outbreak in Assam, Northeast India. J. Vector Borne Dis. 52, 293–298 (2015).
Medeiros, A. S. et al. Dengue virus in Aedes aegypti and Aedes albopictus in urban areas in the state of Rio Grande do Norte, Brazil: Importance of virological and entomological surveillance. PLoS ONE 13, e0194108 (2018).
Ritchie, S. A., Long, S., Smith, G., Pyke, A. & Knox, T. B. Entomological investigations in a focus of dengue transmission in Cairns, Queensland, Australia, by using the sticky ovitraps. J. Med. Entomol. 41, 1–4 (2004).
Hadler, J. L. et al. Assessment of arbovirus surveillance 13 years after introduction of west nile virus, United States1. Emerg. Infect. Dis. 21, 1159–1166 (2015).
Coelho, F. C. & Codeço, C. T. Precision epidemiology of arboviral diseases. J. Public Health Emerg. 3, 1–1 (2019).
Stouthamer, R., Breeuwer, J. A. J. & Hurst, G. D. D. Wolbachia pipientis: microbial manipulator of arthropod reproduction. Annu. Rev. Microbiol. 53, 71–102 (1999).
Blagrove, M. S. C., Arias-Goeta, C., Failloux, A.-B. & Sinkins, S. P. Wolbachia strain wMel induces cytoplasmic incompatibility and blocks dengue transmission in Aedes albopictus. Proc. Natl Acad. Sci. USA 109, 255–260 (2012).
Ross, P. A., Turelli, M. & Hoffmann, A. A. Evolutionary ecology of Wolbachia releases for disease control. Annu. Rev. Genet. 53, 93–116 (2019).
Walker, T. et al. The wMel Wolbachia strain blocks dengue and invades caged Aedes aegypti populations. Nature 476, 450–453 (2011).
Bian, G., Xu, Y., Lu, P., Xie, Y. & Xi, Z. The endosymbiotic bacterium Wolbachia induces resistance to dengue virus in Aedes aegypti. PLoS Pathog. 6, e1000833 (2010).
Dutra, H. L. C. et al. Wolbachia blocks currently circulating Zika virus isolates in Brazilian Aedes aegypti mosquitoes. Cell Host Microbe 19, 771–774 (2016).
Aliota, M. T., Peinado, S. A., Velez, I. D. & Osorio, J. E. The wMel strain of Wolbachia reduces transmission of Zika virus by Aedes aegypti. Sci. Rep. 6, 28792 (2016).
Aliota, M. T. et al. The wMel strain of Wolbachia reduces transmission of chikungunya virus in Aedes aegypti. PLoS Negl. Trop. Dis. 10, e0004677 (2016).
van den Hurk, A. F. et al. Impact of Wolbachia on Infection with chikungunya and yellow fever viruses in the mosquito vector Aedes aegypti. PLoS Negl. Trop. Dis. 6, e1892 (2012).
Hoffmann, A. A. et al. Successful establishment of Wolbachia in Aedes populations to suppress dengue transmission. Nature 476, 454–457 (2011).
Garcia, G. et al. Matching the genetics of released and local Aedes aegypti populations is critical to assure Wolbachia invasion. PLoS Negl. Trop. Dis. 13, e0007023 (2019).
Nguyen, T. H. et al. Field evaluation of the establishment potential of wmelpop Wolbachia in Australia and Vietnam for dengue control. Parasit. Vectors 8, 563 (2015).
de Jesus, C. P., Dias, F. B. S., Villela, D. M. A. & Maciel-de-Freitas, R. Ovitraps provide a reliable estimate of Wolbachia frequency during wMelBr strain deployment in a geographically isolated Aedes aegypti Population. Insects 11, 92 (2020).
Sylvestre, G. et al. Preliminary evaluation on the efficiency of the kit Platelia Dengue NS1 Ag-ELISA to detect dengue virus in dried Aedes aegypti: a potential tool to improve dengue surveillance. Parasit. Vectors 7, 155 (2014).
Mayagaya, V. S. et al. Non-destructive determination of age and species of Anopheles gambiae s.l. using near-infrared spectroscopy. Am. J. Trop. Med. Hyg. 81, 622–630 (2009).
Milali, M. P. et al. Age grading An. gambiae and An. arabiensis using near infrared spectra and artificial neural networks. PLoS ONE 14, e0209451 (2019).
Sikulu, M. et al. Near-infrared spectroscopy as a complementary age grading and species identification tool for African malaria vectors. Parasit. Vectors 3, 49 (2010).
Sikulu, M. T. et al. Using a near-infrared spectrometer to estimate the age of Anopheles mosquitoes exposed to pyrethroids. PLoS ONE 9, e90657 (2014).
Sikulu-Lord, M. T. et al. Near-infrared spectroscopy, a rapid method for predicting the age of male and female wild-type and Wolbachia infected Aedes aegypti. PLoS Negl. Trop. Dis. 10, 1–11 (2016).
Lambert, B. et al. Monitoring the age of mosquito populations using near-infrared spectroscopy. Sci. Rep. 8, 5274 (2018).
Fernandes, J. N. et al. Rapid, noninvasive detection of Zika virus in Aedes aegypti mosquitoes by near-infrared spectroscopy. Sci. Adv. 4, eaat0496 (2018).
Sikulu-Lord, M. T. et al. Rapid and non-destructive detection and identification of two strains of Wolbachia in Aedes aegypti by near-infrared spectroscopy. PLoS Negl. Trop. Dis. 10, e0004759 (2016).
Aw, W. C., Dowell, F. E. & Ballard, J. W. O. Using near-infrared spectroscopy to resolve the species, gender, age, and the presence of Wolbachia infection in laboratory-reared Drosophila. G3 2, 1057–1065 (2012).
Dowell, F. E., Noutcha, A. E. M. & Michel, K. The effect of preservation methods on predicting mosquito age by near infrared spectroscopy. Am. J. Trop. Med. Hyg. 85, 1093–1096 (2011).
Sikulu, M. et al. Evaluating RNAlater® as a preservative for using near-infrared spectroscopy to predict Anopheles gambiae age and species. Malar. J. 10, 186 (2011).
Taylor, L. H., Latham, S. M. & Woolhouse, M. E. J. Risk factors for human disease emergence. Philos. Trans. R. Soc. Lond. Ser. B Biol. Sci. 356, 983–989 (2001).
Ellwanger, J. H., de L Kaminski, V. & Chies, J. A. B. How to detect new viral outbreaks or epidemics? We need to survey the circulation of viruses in humans and other animals using fast, sensible, cheap, and broad-spectrum methodologies. Braz. J. Infect. Dis. 21, 211–212 (2017).
Vasconcelos, P. F. C. & Calisher, C. H. Emergence of human arboviral diseases in the Americas, 2000–2016. Vector Borne Zoonotic Dis. 16, 295–301 (2016).
Pettersson, J. H. O. et al. Re-visiting the evolution, dispersal and epidemiology of Zika virus in Asia. Emerg. Microbes Infect. 7, 1–8 (2018).
Liang, G., Gao, X. & Gould, E. A. Factors responsible for the emergence of arboviruses; strategies, challenges and limitations for their control. Emerg. Microbes Infect. 4, 1–5 (2015).
de Abreu, F. V. S. et al. Combination of surveillance tools reveals that Yellow Fever virus can remain in the same Atlantic Forest area at least for three transmission seasons. Mem. Inst. Oswaldo Cruz 114, e190076 (2019).
Sakudo, A., Suganuma, Y., Kobayashi, T., Onodera, T. & Ikuta, K. Near-infrared spectroscopy: promising diagnostic tool for viral infections. Biochem. Biophys. Res. Commun. 341, 279–284 (2006).
Sakudo, A. et al. A novel diagnostic method for human immunodeficiency virus type-1 in plasma by near-infrared spectroscopy. Microbiol. Immunol. 49, 695–701 (2005).
Maia, M. F. et al. Detection of Plasmodium falciparum infected Anopheles gambiae using near-infrared spectroscopy. Malar. J. 18, 85 (2019).
Roundy, C. M. et al. Variation in Aedes aegypti mosquito competence for Zika virus transmission. Emerg. Infect. Dis. 23, 625–632 (2017).
Dubrulle, M., Mousson, L., Moutailler, S., Vazeille, M. & Failloux, A.-B. Chikungunya virus and Aedes mosquitoes: saliva is infectious as soon as two days after oral infection. PLoS ONE 4, e5895 (2009).
O’Neill, S. L. et al. Scaled deployment of Wolbachia to protect the community from dengue and other Aedes transmitted arboviruses. Gates Open Res. 2, 36 (2018).
Dutra, H. L. C. et al. From lab to field: the influence of urban landscapes on the invasive potential of Wolbachia in Brazilian Aedes aegypti mosquitoes. PLoS Negl. Trop. Dis. 9, e0003689 (2015).
SINGH, J. et al. Silent spread of dengue and dengue haemorrhagic fever to Coimbatore and Erode districts in Tamil Nadu, India, 1998: need for effective surveillance to monitor and control the disease. Epidemiol. Infect. 125, 195–200 (2000).
Souza, T. M. L. et al. Emergence of the East-Central-South-African genotype of Chikungunya virus in Brazil and the city of Rio de Janeiro may have occurred years before surveillance detection. Sci. Rep. 9, 2760 (2019).
Torres, M. C. et al. Re-introduction of dengue virus serotype 2 in the state of Rio de Janeiro after almost a decade of epidemiological silence. PLoS ONE 14, e0225879 (2019).
ten Bosch, Q. A. et al. Contributions from the silent majority dominate dengue virus transmission. PLOS Pathog. 14, e1006965 (2018).
David, M. R., Dos Santos, L. M. B., Vicente, A. C. P. & Maciel-de-Freitas, R. Effects of environment, dietary regime and ageing on the dengue vector microbiota: evidence of a core microbiota throughout Aedes aegypti lifespan. Mem. Inst. Oswaldo Cruz 111, 577–587 (2016).
Rašić, G. et al. Contrasting genetic structure between mitochondrial and nuclear markers in the dengue fever mosquito from Rio de Janeiro: Implications for vector control. Evol. Appl. 8, 901–915 (2015).
Bonaldo, M. C. et al. Isolation of infective Zika virus from urine and saliva of patients in Brazil. PLoS Negl. Trop. Dis. 10, e0004816 (2016).
Teixeira, M. G. et al. East/Central/South African genotype chikungunya virus, Brazil, 2014. Emerg. Infect. Dis. 21, 906–907 (2015).
Ferreira-de-Brito, A. et al. First detection of natural infection of Aedes aegypti with Zika virus in Brazil and throughout South America. Mem. Inst. Oswaldo Cruz 111, 655–658 (2016).
da Silveira, I. D. et al. Zika virus infection produces a reduction on Aedes aegypti lifespan but no effects on mosquito fecundity and oviposition success. Front. Microbiol. 9, 3011 (2018).
R Development Core Team, R. Computational Many-Particle Physics. R Foundation for Statistical Computing 739 (Springer Berlin, Heidelberg, 2008).
Acknowledgements
We would like to thank the technical support of Marcelo Celestino, Renato Carvalho, Marcelo Quintela, and Mauro Muniz for field and lab assistance.
Author information
Authors and Affiliations
Contributions
M.S.-L. and R.M.d.F. conceived the study. L.M.B.S., M.M., G.A.G., M.R.D., M.G.P., and D.C.-L. processed the experimental data. L.M.B.S., M.T.P., J.C.A., and D.C.-L. conducted molecular assays and interpretation. M.G.P., D.C.-L., F.D., A.L., and M.S.-L. conducted data analysis. L.M., A.L., M.S.-L., and R.M.d.F. wrote the initial draft of the manuscript. All authors discussed the results and commented on the manuscript at all stages.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Santos, L.M.B., Mutsaers, M., Garcia, G.A. et al. High throughput estimates of Wolbachia, Zika and chikungunya infection in Aedes aegypti by near-infrared spectroscopy to improve arbovirus surveillance. Commun Biol 4, 67 (2021). https://doi.org/10.1038/s42003-020-01601-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-020-01601-0
This article is cited by
-
Age, sex, and mating status discrimination in the sand fly Lutzomyia longipalpis using near infra-red spectroscopy (NIRS)
Parasites & Vectors (2024)
-
Invasive hematophagous arthropods and associated diseases in a changing world
Parasites & Vectors (2023)
-
Key considerations, target product profiles, and research gaps in the application of infrared spectroscopy and artificial intelligence for malaria surveillance and diagnosis
Malaria Journal (2023)
-
Near infrared spectroscopy accurately detects Trypanosoma cruzi non-destructively in midguts, rectum and excreta samples of Triatoma infestans
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.