Abstract
rnf genes are widespread in bacteria and biochemical and genetic data are in line with the hypothesis that they encode a membrane-bound enzyme that oxidizes reduced ferredoxin and reduces NAD and vice versa, coupled to ion transport across the cytoplasmic membrane. The Rnf complex is of critical importance in many bacteria for energy conservation but also for reverse electron transport to drive ferredoxin reduction. However, the enzyme has never been purified and thus, ion transport could not be demonstrated yet. Here, we have purified the Rnf complex from the anaerobic, fermenting thermophilic bacterium Thermotoga maritima and show that is a primary Na+ pump. These studies provide the proof that the Rnf complex is indeed an ion (Na+) translocating, respiratory enzyme. Together with a Na+-F1FO ATP synthase it builds a simple, two-limb respiratory chain in T. maritima. The physiological role of electron transport phosphorylation in a fermenting bacterium is discussed.
Similar content being viewed by others
Introduction
Biological energy conservation in a primordial, anoxic world most likely involved separation of charges and molecules such as protons and sodium ions across membranes1 and the established electrochemical gradient as driving force for the synthesis of ATP, the universal energy currency in every living cell, by simple ATP synthases2. How these gradients have been established is still an enigma, but anaerobic microbes may shed light on this question. They often employ ferredoxin (Fd) in addition to NAD as electron carrier. The low potential electron carrier ferredoxin (EO′ = −450 to −500 mV3) can be reduced by only a few enzymes such as glycerol aldehyde 3-phosphate dehydrogenase4 or pyruvate:ferredoxin oxidoreductase5, both employed during chemoorganoheterotrophic growth (glycolysis). During the chemolithoautotrophic growth, ferredoxin must be reduced with H2 (EO′ = −414 mV) as electron donor, but this reaction is highly endergonic. The energy barrier is overcome by flavin-based electron bifurcation, a recently established mechanism widespread in anaerobes to drive endergonic ferredoxin reduction by coupling the oxidation of the same substrate to the reduction of a second, more electropositive acceptor6,7. In the electron-bifurcating hydrogenases the second electron acceptor is NAD8,9.
Fdred is an ideal electron donor in an anoxic world since, due to its low redox potential, electrons can be delivered to acceptors such as, for example, protons (EO′ = −414 mV) or NAD (EO′ = −320 mV). Reduction of protons and NAD with Fdred as donor is exergonic and nature has evolved different enzymes that store the redox energy in a transmembrane electrochemical ion potential across the cytoplasmic membrane10,11,12,13. Fdred:H+ oxidoreductase activity coupled to proton translocation is found in some anaerobic archaea11,14,15 and has been studied to a great extent in Pyrococcus furiosus12. Experiments with inverted membrane vesicles clearly demonstrated this respiratory activity, and the enzyme catalyzing this reaction, the membrane-bound hydrogenase (Mbh) has been purified and characterized16. It is a predecessor of complex I and the recently solved structure of the enzyme revealed its structural similarity to complex I17. Although the enzyme has been purified, it has not been reconstituted yet in liposomes. Therefore, the final proof that the enzyme is a proton pump is still missing.
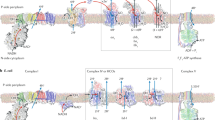
The second Fdred-dependent electron transport chain found in anaerobes uses NAD+ as electron acceptor. Fd2−:NAD+-oxidoreductase activity was demonstrated in the acetogen Acetobacterium woodii18 and inverted membrane vesicles of this bacterium coupled Fd2−:NAD+-oxidoreductase to vectorial Na+ transport across the cytoplasmic membrane10. The activity was enriched from membranes of A. woodii and the preparation contained, amongst others, proteins with similarity to proteins encoded by the rnf genes19. These genes have been firstly described in and shown to be essential for N2 fixation in Rhodobacter capsulatus20. The rnf genes are assumed to encode a membrane-bound enzyme complex with six subunits; two subunits are predicted to be membrane integral, the other four are cytosolic. RnfB is predicted to be the entry point for electrons coming from Fdred21, RnfC is the predicted NAD binding site21. Electron flow from RnfB to RnfC is most likely via RnfA and RnfG. Subunits D and E are membrane-integral subunits suggested to mediate ion (Na+) transport, driven by electron transport that involves iron sulfur cluster, monovalent iron and covalently bound flavins. Again, our view of the structure and function of Rnf complexes stems from predictions of DNA sequences and analogies to the well-studied respiratory enzyme Nqr22,23. The final proof that the purified Rnf complex is a respiratory enzyme is still missing, largely due to the fact that the complex always escaped purification, even from different microbial sources24.
We describe here a procedure to purify the Rnf complex from a thermophilic bacterium, Thermotoga maritima, in complex with the ATP synthase. The purified Rnf-ATP synthase supercomplex was reconstituted into liposomes and its catalytic properties were analyzed. The Rnf complex was further separated from the ATP synthase and shown to catalyze Na+ transport.
Results
Purification of Rnf and ATP synthase
Since attempts to purify the Rnf complex from A. woodii, Clostridium ljungdahlii or Clostridium tetanomorphum failed due to an inherent instability of the complex,24 we rationalized that the complex from a thermophilic bacterium may be more stable. Inspection of the genome sequence of T. maritima (Topt = 80 °C) revealed the presence of rnf genes in the order rnfCDGEAB, as in A. woodii. The predicted proteins are 46.9, 46.9, 29.3, 59.8, 56.1 and 49.2% similar to the corresponding proteins from A. woodii. Interestingly, RnfB is only 10.56 kDa, compared to 37.73 kDa in A. woodii. This accomodates only one predicted 4Fe-4S center, in contrast to six predicted 4Fe-4S in A. woodii. This raised the question whether the Rnf complex from T. maritima would accept ferredoxin from Clostridium pasteurianum, which is generally used as electron donor to measure Fdred:NAD-oxidoreductase activity. FdCp was purified and reduced with carbon monoxide, catalyzed by the purified carbon monooxide dehydrogenase from the mesophile A. woodii. Much to our relief, membranes of T. maritima catalyzed Fdred-dependent NAD+ reduction at 60 °C, a temperature that meets the demands of the auxilliary proteins (CO dehydrogenase (Topt = 30 °C), ferredoxin (Topt = 37 °C)) from the mesophile as well as the target enzyme, Rnf, from a thermophile. The Fd2−:NAD+-oxidoreductase activity at membranes of T. maritima was 97 mU/mg, measured at 60 °C (Supplementary Fig. 1).
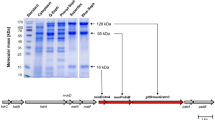
The enzyme catalyzing Fd2−:NAD+-oxidoreductase activity was purified from the cells of T. maritima under strictly anoxic conditions. Solubilization was achieved with n-Dodecyl β-d-maltoside (1 mg/mg protein), followed by anion exchange chromatography on Q-sepharose and gel filtration on Sephacryl S300. The Rnf activity eluted as a single peak from a gel filtration with an activity of 2.1 U/mg (Supplementary Fig. 2). This procedure enriched the activity 21-fold with a yield of 13% (Supplementary Table 1). When the preparation was analyzed by SDS-PAGE, more than the expected six proteins were visible (Supplementary Fig. 3a). RnfD (34.6 kDa) and RnfG (24.3 kDa) have covalently bound flavins and indeed, two proteins with apparent masses of 23 and 32 kDa were visible after exposure of the gel to UV light (Supplementary Fig. 3b). Antibodies against Rnf subunits from A. woodii did not cross react with any protein in the preparation which is plausible since the identities are only between 29.3 and 59.8%. In a native PAGE, several bands were visible. To determine the Rnf complex, we did an in-gel assay to determine the NADH:MTT-oxidoreductase activity. Two complexes of apparent masses of ≈150 and ≈290 kDa had NADH:MTT-oxidoreductase activity and the activity of the ≈290-kDa complex was higher than in the 150-kDa complex (Fig. 1a). The predicted mass of an Rnf monomer (one subunit per complex) is 160 kDa; thus the two complexes could represent a monomer and a dimer of the Rnf complex, which is within the error of the mass detection (9%) by native PAGE and the uncertainty of the subunit stochiometries. The ≈290-kDa complex was excised from the gel and applied to a SDS-PAGE. As evident from Fig. 1b, proteins with masses matching the predicting masses of the Rnf complex were visible. LC/MS/MS and MALDI-TOF analyses revealed three of six Rnf subunits (RnfCDG) in the preparation (Supplementary Table 2).
a Twenty micrograms of purified protein was separated on a native-PAGE49. The proteins were visualized by staining with Coomassie Brilliant Blue. ATPase and Rnf activity were determined as described in “Methods”. b The bands representing Rnf (dimer) and F1FO were excised from the native gel and separated on a 12.5% SDS-gel according to ref. 48 and stained with silver. The subunits were identified by western blot or by LC/MS/MS and MALDI-TOF.
The native gel contained a 550-kDa complex of high abundance. This mass corresponds to the mass of the F1FO-ATP synthase encoded in the genome of T. maritima (545–569 kDa, depending on the number of c subunits in the complex). Indeed, the complex had in-gel ATPase activity (Fig. 1a). The complex was excised and separated in a SDS-PAGE. Proteins with masses expected for subunits α, β, γ, δ, ε, a, c and b were detected by silver staining (Fig. 1b). Indeed, the 60- and 8-kDa proteins reacted with antibodies against subunits β and c of the F1FO-ATP synthase of A. woodii. Moreover all eight subunits were detected by LC/MS/MS and MALDI-TOF (Supplementary Table 2). The ATPase activity in the preparation was around 5 U/mg.
The Rnf complex requires Na+ and is inhibited by DCCD
Fd2−:NAD+-oxidoreductase activity was similar in 50 mM Tris-HCl (pH 7.7), 50 mM 3-(N-morpholino)propanesulfonic acid (MOPS) (pH 7.7) and 50 mM 1,4-Piperazinediethanesulfonic acid (PIPES) buffer (pH 7.7) and somewhat lower in 50 mM TES and 50 mM K-PO4 buffer. For the following, 50 mM Tris-HCl was used. Activity had a rather broad pH optimum from 7 to 8.5, but at pH 6 and 10.5, activity was still 50%. The temperature optimum could not be determined using the physiological activity since CODH used to reduce ferredoxin is from a mesophile and is inactive at temperatures >70 °C. Therefore, NADH:MV-oxidoreductase activity was measured. As seen in Supplementary Fig. 4, activity increased with increasing temperatures with a maximum of 51.4 ± 2.4 U/mg at the highest temperature tested, 90 °C. At 60 °C, still 47.1% of the activity was present. At 60 °C, the activity of the CODH was still 35%. Therefore, the Fd2−:NAD+-oxidoreductase activity was measured at the compromise temperature of 60 °C.
Of special importance was the effect of Na+ on Fd2−:NAD+-oxidoreductase since Na+- and H+-dependent Rnf complexes are known. In the absence of Na+ there was no Fd2−:NAD+-oxidoreductase activity. KCl did not stimulate activity, addition of LiCl stimulated activity slightly, but addition of NaCl greatly stimulated the activity. The dependence of activity on the NaCl concentration followed a Michaelis−Menten kinetic with a Km of around 0.5 mM and a vmax of 1 U/mg (Fig. 2a). These data clearly show that the Rnf complex of T. maritima is Na+-dependent. Some membrane proteins involved in ion transport are inhibited by N,N′-dicyclohexylcarbodiimide (DCCD), which binds covalently to carboxylates buried in the membrane phase25,26. DCCD also inhibited Fd2−:NAD+-oxidoreductase; maximum inhibition was observed at 5 µM DCCD. Most interestingly, DCCD inhibition was relieved by the addition of Na+ (Fig. 2b). When the enzyme was preincubated with 5 µM DCCD and 20 mM Na+, DCCD inhibition was only 20%. Li+ also protected from DCCD inhibition, but to a much smaller extent. At 5 µM DCCD and 20 mM LiCl, the protection was only 21%. KCl did not protect the enzyme from DCCD inhibition (Supplementary Fig. 5). The different starting activities are due to variations in activity of different badges of enzyme. These data indicate that, as in other Na+ translocating membrane proteins such as the Na+-F1FO-ATP synthase, Na+ and DCCD compete for a common binding site27,28.
a Purified protein (7.5 µg) was added to 1 ml buffer (20 mM Tris-HCl, 2 mM DTE, 2.2 µM resazurin, pH 7.7) and ferredoxin:NAD+ oxidoreductase was measured as described in “Methods”. Increasing amounts (0−20 mM) of NaCl (solid square), KCl (inverted solid triangle) or LiCl (solid triangle) were applied to the assays. The contaminating Na+ concentration was 0.095 mM. b Purified protein (10 µg) was added to 1 ml buffer (20 mM Tris-HCl, 2 mM DTE, 2.2 µM resazurin, pH 7.7) with increasing amounts of DCCD (0–10 µM) in the presence (solid triangle) or absence (solid square) of 20 mM NaCl. The samples were incubated for 20 min at room temperature followed by an incubation at 60 °C for 5 min before the measurement started. If NaCl was omitted during preincubation, 20 mM NaCl was added before the start of the reaction. The contaminating Na+ concentration was 0.12 mM. These data points represent two different biological replicates.
The ATP synthase of T. maritima is of the Na+-F1FO-type
ATP hydrolysis was strictly dependent on the Na+ concentration (Fig. 3) and Na+ dependence of activity followed a Michalies−Menten kinetic; the Km was 1.2 ± 0.2 mM Na+ and vmax was 4.7 U/mg. As seen before with other Na+-F1FO-ATP synthases27,28, DCCD inhibited ATP hydrolysis but inhibition was relieved by addition of Na+, indicating that Na+ and DCCD compete for a common binding site (Fig. 4).
Four micrograms of purified ATPase was added to 1200 µl ATPase buffer (100 mM Tris-HCl, 100 mM malic acid, pH 7.4) containing 0−10 mM NaCl. The sample was incubated for 5 min at 80 °C. The reaction was started by adding 3 mM K2-ATP. The contaminating Na+ concentration was 0.154 mM. All data points are mean ± SEM; N = 3 independent experiments.
Nine micrograms of purified ATPase was added to 1200 μl ATPase buffer (100 mM Tris-HCl, 100 mM malic acid, pH 7.4) containing 0−20 µM DCCD in the absence (solid square) or presence of 20 mM NaCl (solid triangle). The sample was incubated for 30 min at room temperature followed by 5 min at 80 °C. The reaction was started by adding 3 mM K2-ATP. The contaminating Na+ concentration was 0.07 mM. If NaCl was omitted during preincubation, 20 mM NaCl was added before the start of the reaction. All data points are mean ± SEM; N = 3 independent experiments.
Reconstitution of ATP synthase and Rnf in liposomes
To address ion transport by the Rnf complex and the ATP synthase, the enzymes were reconstituted into liposomes. Since the E. coli lipids used to prepare liposomes are unstable at higher temperatures, the activity assays were performed at 45 °C. At this temperature, Fd2−:NAD+-oxidoreductase and ATPase activity were 65% and 55%, respectively, of the control at 60 or 80 °C. At 45 °C the liposomes were quite stable and could hold an artificial pH created by an ammonium diffusion potential and measured by ACMA quenching for 7 min. After incorporating the enzymes by detergent removal, the activities of both were measured: ATPase was 2.6 U/mg and Fd2−:NAD+-oxidoreductase was 1.2 U/mg. To determine Na+ transport, the proteoliposomes were incubated with 22Na+. Upon addition of K2-ATP, proteoliposomes accumulated Na+ in the lumen more than twofold, which clearly demonstrates that the F1FO-ATP synthase of T. maritima uses Na+ as coupling ion (Fig. 5a). When proteoliposomes were incubated with ferredoxin and CODH under a CO atmosphere, 22Na+ was accumulated after addition of the electron acceptor, NAD (Fig. 5b). Accumulation was 2.5-fold, demonstrating that the Rnf complex also uses Na+ as coupling ion.
a Proteoliposomes (protein concentration 1.3 mg/ml) in 100 mM Tris-HCl, 100 mM malic acid, pH 7.4, containing 5 mM MgCl2, 5 mM 22NaCl, 2 mM DTE, 4 µM resazurin catalyzed 22Na+ transport upon addition of 15 µl K2-ATP (100 mM) under oxic conditions (solid square). 22Na+ transport was not observed in the absence of K2-ATP (solid triangle). b The same proteolipsomes catalyzed 22Na+ transport upon addition of 10 µl ferredoxin (3 mM), 10 µg CODH (3 mg/ml) and 30 µl NAD+ (100 mM) and a CO atmosphere (solid square). 22Na+ transport was not observed in the absence of NAD+ (solid triangle). These data points represent two different biological replicates.
Further purification and reconstitution of the Rnf
The experiments described so far clearly demonstrate a Na+ pumping Rnf and a Na+ pumping ATP synthase present in T. maritima that co-purify. Since Na+ transport with a purified Rnf complex had never been shown, it was important to unequivocally demonstrate Na+ transport by the Rnf complex. Therefore, we decided to modify the isolation procedure to remove the ATP synthase. To this end, the solubilisate was precipitated with PEG6000, the precipitate applied to a sucrose density gradient, followed by chromatography on Q-sepharose, hydroxyapatite and sephacryl S300. This procedure removed the Na+-F1FO-ATP synthase and enriched the Rnf activity by 75%. However, the yield was only 4.1% and the procedure was laborious that often led to inactivation during the purification procedure which usually takes 7−10 days. Anyway, ATPase activity could not be detected in the preparation. The preparation contained all subunits of the Rnf complex, which were confirmed by MALDI-TOF-analyses (Supplementary Table 3) and SDS-PAGE (Supplementary Fig. 6a). RnfD and RnfG contained covalently bound flavins (Supplementary Fig. 6b). In a native PAGE a rather broad band ranging from 215 to 270 kDa was observed. Whether this represent a monomer or a dimer needs to be addressed in the future. Anyway, the complex had NADH:MTT in-gel activity (Supplementary Fig. 6c). Fd2−:NAD+-oxidoreductase was strictly Na+ dependent, inhibited by DCCD and Na+ protected from DCCD inhibition (Fig. 6). The Rnf complex was then incorporated into liposomes, as before. When incubated in the presence of 22Na+, cells accumulated 22Na+ when the Fd2−:NAD+-oxidoreductase activity was started by addition of the electron acceptor NAD (Fig. 7). Accumulation of 22Na+ was inhibited by the Na+ ionophore ETH2120, but stimulated by the protonophore TCS, demonstrating that Na+ transport is a primary and electrogenic event (Fig. 8).
a Purified protein (10 µg) was added to 1 ml buffer (20 mM Tris-HCl, 2 mM DTE, 2.2 µM resazurin, pH 7.7) and ferredoxin:NAD+ oxidoreductase activity was measured as described in “Methods”. Increasing amounts (0−10 mM) of NaCl (solid square), KCl (solid triangle) or LiCl (solid inverted triangle) was applied to the assays. The contaminating Na+ concentration was 0.085 mM. b Purified protein (10 µg) was added to 1 ml buffer (20 mM Tris-HCl, 2 mM DTE, 2.2 µM resazurin, pH 7.7) with increasing amounts of DCCD (0–25 µM) in the presence (solid triangle) or absence (solid square) of 20 mM NaCl. The samples were incubated for 20 min at room temperature followed by an incubation at 60 °C for 5 min before the measurement was started. When NaCl was omitted in the preincubation, 20 mM NaCl was added with the start of the reaction. The contaminating Na+ concentration was 0.085 mM. These data points represent two different biological replicates.
Proteoliposomes (protein concentration 1.06 mg/ml) in 1 ml 100 mM Tris-HCl, 100 mM malic acid, pH 7.4, containing 5 mM MgCl2, 2 mM 22NaCl, 2 mM DTE, 4 µM resazurin catalyzed 22Na+ transport upon addition of 10 µl ferredoxin (3 mM), 10 µg CODH (3 mg/ml) and 30 µl NAD+ (100 mM) under a CO atmosphere (solid square). 22Na+ transport was not observed in the absence of NAD+ (solid triangle). These data points represent two different biological replicates.
Proteoliposomes (protein concentration 0.99 mg/ml) in 1 ml 100 mM Tris-HCl, 100 mM malic acid, pH 7.4, containing 5 mM MgCl2, 2 mM 22NaCl, 2 mM DTE and 4 μM resazurin were incubated with 40 µM TCS (solid square) or 40 µM ETH2120 (solid triangle) for 30 min at room temperature before starting 22Na+ transport by addition of 10 μl ferredoxin (3 mM), 10 μg CODH (3 mg/ml) and 30 μl NAD+ (100 mM) under a CO atmosphere. These data points represent two different biological replicates.
Discussion
rnf genes are widespread in bacteria and also present in some archaea18. Based on the homology of some subunits of the hypothetical Rnf complex to subunits of the NADH:quinone-oxidoreductase (Nqr29), it was speculated that the Rnf complex is a respiratory enzyme that oxidizes reduced ferredoxin (E′O 80 °C = −453 mV30) and reduces NAD (EO′ = −320 mV). This exergonic reaction (ΔG°′ = −34.7 kJ/mol) is supposed to be coupled to ion transport across the membrane18. The ion potential then drives the synthesis of ATP via an ATP synthase. Na+ transport coupled to Fd2−:NAD+-oxidoreductase activity was indeed demonstrated in inverted membrane vesicles of A. woodii10. As expected, the reaction is reversible and an ion gradient is also used as driving force for the endergonic reduction of ferredoxin with NADH as reductant31. This activity is essential for nitrogen reduction of R. capsulatus, but also in C. ljungdahlii20,32. In addition, growing cells of A. woodii require reduced ferredoxin for biosynthetic reactions, but during growth on several substrates, only NAD is reduced. Under these conditions the Rnf complex is used to drive the endergonic reduction of ferredoxin with NADH as reductant at the expense of the electrochemical ion gradient33.
Genetic experiments in which deletion of rnf genes led to a loss of Fd2−:NAD+-oxidoreductase activity together with the beforementioned experiments at inverted membrane vesicles are convincing circumstantial evidence that the Rnf complex is a respiratory enzyme33. However, the final proof requires the purification of the complex and its functional reconstitution into liposomes. Despite several attempts in different laboratories, the Rnf complex resisted purification. Here, we describe the purification of the membrane-integral Rnf complex from the thermophile T. maritima and it is tempting to speculate that the thermophilic nature of the target protein was advantageous for its purification. Although Rnf activity was determined under suboptimal conditions at low temperatures, due to the instability at high temperatures of the auxiliary enzymes required for Fd reduction, we could unequivocally demonstrate Na+ dependence of and, more important, Na+ transport by the purified complex. This is, to our knowledge, the first proof that the Rnf complex is indeed an ion (Na+) translocating respiratory enzyme.
Notably, the B subunit of the Rnf complex from T. maritima is much smaller than the B subunit from A. woodii or other bacteria. Inspection of RnfB subunits from different sources revealed a striking difference in the length of subunit B (Fig. 9). Apparently, the difference is in the C terminus that harbors the FeS cluster. Acetobacterium woodii has six predicted FeS centers, C. ljungdahlii, has four, R. capsulatus, Vibrio cholerae, Escherichia coli have two, whereas T. maritima has only one. Despite these differences, ferredoxin from C. pasteurianum is able to deliver electrons to the Rnf complex. Whether the number of 4Fe-4S-cluster is indeed only one remains to be established.
The demonstration of an inhibition of the Rnf complex by DCCD is unprecedented. DCCD and Na+ compete for a common binding site, as in Na+-F1Fo ATP synthases27,28. This may be a useful tool in the future to identify the Na+-binding site. Radioactively labeled DCCD is no longer commercially available and a fluorescent analog, NCD4, did not inhibit the Rnf complex specifically. For the homologous Nqr complex Juárez et al.34 speculated that the glutamate/aspartate in NqrB (E144, D397), in NqrE (E95) and in NqrD (D133) are potential Na+-binding sites. The deletion of these four amino acids resulted in a reduced stimulation of ubiquinone reduction by Na+ and mutations in NqrE-E95 resulted in an inhibition of electron transfer34,35. Except for NqrB-E144 all these amino acids are also conserved in the corresponding Rnf subunits from T. maritima, A. woodii and V. cholerae (Supplementary Figs. 7–9). However, there are more conserved acidic amino acids (NqrB-E274, NqrB-D346, NqrB-E380 NqrD-E119 and NqrD-E152) that could also be candidates for Na+ binding. From the crystal structure of the Nqr complex, it was postulated that NqrB forms an ion channel and the D346, F342, F338, V161, I164 and L168 are involved in Na+ translocation23. In the corresponding Rnf subunit of T. maritima RnfD only RnfD-D255 (Vc. NqrB-D346), RnfD-F252 (Vc. NqrB-F342) and RnfD-V79 (Vc. NqrB-V161) are conserved (Supplementary Fig. 9). This would lead to the assumption that RnfD would also form such an ion channel, but this has to await structural data.
The F1FO ATP synthase of T. maritima not only has a conserved Na+ binding motif in subunit c but its activity is Na+ dependent and ATP hydrolysis is coupled to Na+ transport. This is convincing evidence that the ATP synthase uses Na+ as coupling ion, which is expected if the Rnf complex is Na+ translocating. Apparently, the respiratory chain of T. maritima consists of Rnf and ATP synthase, which are connected by an electrochemical Na+ potential.
T. maritima is a fermenting organism that converts sugars according to36
Oxidation of glucose via the ED pathway generates NADPH that can be reoxidised via the electron-bifurcating transhydrogenase Nfn in connection with the Rnf complex (Fig. 10, ref. 37). According to the model presented in Fig. 10, 3 mol of lactate are also produced to balance out the electrons; indeed, lactate production was seen in resting cell experiments38. Altogether, 3.7 ATP are generated by substrate level phosphorylation and additional 0.035 ATP are generated by electron transport phosphorylation with Rnf and ATP synthase. This is only about 1% of the total amount of ATP produced. This underlines the notion that, during chemoorganoheterotrophic growth, the function of the Rnf complex is not primarily in energy conservation but is that of a transhydrogenase essential to balance out the electrons8. It should be mentioned in this connection that T. maritima has a membrane-bound, possibly ion translocating hydrogenase that could reduce protons (Supplementary Fig. 10) or H2S (Supplementary Fig. 11)39. The amount of ions translocated by the Rnf complex stays the same, but with protons as electron acceptors twice as much Na+ would be translocated by the hydrogenase, giving 0.1 ATP/mol glucose. Based on theoretical considerations, reduction of S to H2S at high hydrogen partial pressures is increased since the hydrogenase is limited and more electrons flow towards sulfur reduction. This could increase the ATP yield by chemiosmosis to 1.79 per mol glucose, already 46% of the yield obtained by SLP.
Glucose is oxidized via glycolysis and Entner-Doudoroff pathway and acetate is produced. NADPH generated in the glucose-6-phosphate dehydrogenase reaction is oxidized with an NADH-dependent Fd2−:NADP+-transhydrogenase (Nfn) by reducing Fd and NAD. Electrons from Fd2− are shuffled to NAD by the Rnf complex, thereby generating an Na+ gradient that is used by F1FO ATP synthase to generate ATP. NADH generated by the Rnf complex and Nfn is used to reduce pyruvate to lactate. All other reducing equivalents are used by the electron-bifurcating/confurcating hydrogenase (Hyd) to generate H2. KDPG 2-keto-3-desoxy-6-phosphogluconate, GAP glyceraldehyde 3-phosphate, BPG bisphosphoglycerate. Note that the ion/electron and ion/ATP stoichiometries are based on thermodynamics (see also ref. 39).
Methods
Growth of T. maritima
Cells of T. maritima (DSM3109) were grown at 80 °C in a 100 l fermenter under anoxic conditions as described previously with starch as substrate38. Harvested cells were frozen and stored at −80 °C.
Purification of the Rnf complex and the ATP synthase
In order to prevent protein damage through oxygen, all preparations and purification steps were performed under anoxic conditions in an anaerobic chamber (Coy Laboratory Products, Gras Lake Charter Township, MI). Additionally, all buffers contained 2 mM dithioerythritol (DTE), 4 µM resazurin and 5 µM FMN. Cells were resuspended and washed in lysis buffer (50 mM Tris-HCl, pH 8, 20 mM MgSO4). Sedimented cells were resuspended in lysis buffer containing 0.5 mM phenylmethylsulfonyl fluoride (PMSF) and 0.1 mg/ml DNaseI and disrupted by a single passage through a French Press with 1000 psig (100 bar). Cell debris were removed by a centrifugation step (24,000 × g, 20 min, 4 °C). With the following ultracentrifugation step (130,000 × g, 90 min, 4 °C), cytoplasmatic and membrane fractions were separated. Membranes were washed three times in lysis buffer and sedimented again through ultracentrifugation (130,000 × g, 30 min, 4 °C). As the yield of the Rnf complex should be maximized, cell debris were not removed before purification. An ultracentrifugation step (130,000 × g, 60 min, 4 °C) followed the cell disruption directly. Sedimented cells debris and membranes were washed three times with lysis buffer and sedimented again through ultracentrifugation (130,000 × g, 30 min, 4 °C). Incubating the sample with DDM for 16 h at 4 °C led to a complete solubilization of the Rnf complex. In order to dilute the DDM concentration, solubilized proteins were mixed with lysis buffer in a ratio of 1:1. Solubilized proteins were applied to a Q-sepharoseTM High Performance column (1.6 × 12 cm) equilibrated with buffer A (50 mM Tris-HCl, pH 8, 20 mM MgSO4, 0.02% DDM). A washing step using 100 mM NaCl using buffer B (50 mM Tris-HCl, pH 8, 20 mM MgSO4, 0.02% DDM, 1 M NaCl) followed. Elution was performed by applying a linear gradient over 200 ml from 0 mM to 350 mM NaCl. A ferredoxin-dependent NAD+ reductase eluted at approximately 230 mM NaCl. Pooled fractions were concentrated by ultrafiltration in 100-kDa Vivaspin tubes (Satorius Stedim Biotech GmbH) and applied to a SephacrylTM S-300 HP (1.6 × 60 cm) column equilibrated with buffer C (50 mM Tris-HCl, pH 8, 20 mM MgSO4, 0.02% DDM, 150 mM NaCl). The Fd2−:NAD+-oxidoreductase (Fno) eluted in a single peak and was concentrated as performed previously and stored at 4 °C. For some experiments, it was necessary to remove NaCl by using a HiTrap-Desalting column where the buffer was changed to sodium free buffer C.
Preparation and purification of the Rnf complex
All steps until the solubilization were performed as described above. Resuspended membranes were used for protein solubilization with DDM. One milligram DDM was added per 1 mg protein and incubated for 2 h at room temperature under stirring. Solubilized membranes were removed by ultracentrifugation (45,000 rpm, 45 min, 4 °C). Solubilized proteins were precipitated with polyethylene glycol 6000 (PEG6000). For the first precipitation step, 7% PEG6000 [v/v] was added and incubated for 30 min at 4 °C under stirring. Precipitated proteins were removed by ultracentrifugation (38,000 rpm, 30 min, 4 °C). For the second precipitation step, the PEG6000 concentration was increased to 12% PEG6000 [v/v]. Precipitated proteins were collected by ultracentrifugation (38,000 rpm, 30 min, 4 °C) and resuspended in buffer A. Resuspended proteins were loaded onto a sucrose density gradient (20–50%) and centrifuged for 20 h in a vertical rotor (Beckman Optima L90-K, VTi50 rotor; 38,000 rpm, 4 °C). Sucrose gradients were fractionized (Ismatec IPC, Ismatec, Wertheim, Germany) and in each fraction Fno activity was measured as described later. Fractions with the highest Fno activity were pooled and applied to an anionic exchange chromatography column using Q-sepharoseTM (GE Healthcare, Little Chalfont, UK) equilibrated with buffer A. Proteins were eluted with an NaCl gradient in buffer B at a flowrate of 2 ml/min. A ferredoxin-dependent NAD+ reductase eluted at approximately 100 mM NaCl. Fractions with the highest Fno activity were pooled and applied to a biionic exchange chromatography column using Hydroxyapatite (Bio-Rad, Hercules, CA, USA) equilibrated with buffer D (5 mM KPO4, 150 mM NaCl, 2 mM DTE, 4 µM resazurin, 5 µM FMN, 0.02% DDM [w/v], pH 7.0). Protein was eluted with a potassium phosphate gradient (153.5 to 500 mM KPO4) with buffer E (buffer D with 500 mM KPO4, pH 7.0) with a flowrate of 2 ml/min. Fractions with the highest Fno activity were pooled and concentrated (molecular mass cutoff 100 kDa) and applied to a gel filtration using Sephacryl S300 (GE Healthcare, Little Chalfont, UK) equilibrated with buffer F (buffer A containing 250 mM NaCl, pH 8.0) with a flowrate of 0.5 ml/min. The Fno activity eluted after 62 ml. The size of the protein was determined using a high molecular weight calibration kit (GE Healthcare, Little Chalfont, UK) under identical conditions. The enzyme was stable for 1 week when stored at 4 °C. For measurements in the absence of Na+ the Sephacryl S300 pool was concentrated and applied on a desalting chromatography column using HiTrap-Desalting column (GE Healthcare, Little Chalfont, UK) equilibrated with buffer A with a flowrate of 1 ml/min. Fractions with Fno activity were pooled.
Measurement of Rnf activity
Fd2−:NAD+-oxidoreductase activity was measured as described31 at 60 °C with 3 mM NAD+. NaCl concentrations were adjusted and are indicated. Membranes or purified protein were used as sample. Determination of the temperature optimum was done by measuring the NADH:MVox oxidoreductase activity of the Rnf complex since ferredoxin from C. pasteurianum and ACS/CODH from A. woodii are inactive at higher temperatures. These measurements were performed in anoxic cuvettes sealed with rubber stoppers and an atmosphere consisting of 100% N2 (0.5 × 105 Pa). They contained 1 ml buffer (20 mM Tris-HCl, pH 7.7, 2 mM DTE, 4 µM Resazurin, 20 mM NaCl) and 10 mM methylviologen. After adding the sample and an incubation time at the according temperature, the enzymatic reaction was started by adding 2 mM NADH. The formation of reduced methylviologen was measured photometrically at 604 nm (εMV = 13.9/mM/cm).
Measurement of ATPase activity
ATPase activity was measured in buffer containing 100 mM Tris-HCl, 100 mM maleic acid, 20 mM NaCl, 5 mM MgCl2 at 80 °C. The pH was adjusted to 7.4 with KOH. The sample was preincubated at the given temperature before addition of 3 mM Na2-ATP to start the reaction. ATP-dependent formation of inorganic phosphate was followed as described40. Determination of Na+-dependence was carried out using NaCl-free buffer and K2-ATP.
Inhibition assays
The samples were preincubated with DCCD at room temperature with and without 20 mM NaCl for 20 and 5 min at 60 °C before starting the reaction by adding the substrate. NaCl was added with the start of the reaction. DCCD was dissolved in ethanol; controls received the solvent only.
In-gel activity assays
Enzyme activity could also be shown in polyacrylamide gels. Rnf activity was demonstrated as an NADH:3-(4,5dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) oxidoreductase activity41. A native polyacrylamide gel was incubated in anoxic buffer (20 mM Tris-HCl, pH 7.7, 2 mM DTE, 20 mM NaCl) containing 200 µM MTT and 2 mM NADH. After short incubation MTT was reduced irreversibly and formazan accumulated in the gel41. ATPase activity was also shown in a native PAGE gel. The gel was incubated for an hour in buffer containing 35 mM Tris-HCl, pH 8.3, 270 mM glycine, 14 mM MgSO4, 0.2% (w/v) Pb(NO3)2 and 8 mM Na2-ATP at 65 °C. Released inorganic phosphate formed a white precipitate with lead nitrate. Incubating the gel in 50% methanol (v/v) stopped the reaction.
Determination of Na+ concentration
The amount of Na+ was determined with an Orion 84-ROSS sodium electrode (Thermo Electron Corp. Witchford, UK) as described42.
Preparation of proteoliposomes
Liposomes were prepared from E. coli total lipid extract as described43. Reconstitution of proteins into liposomes was carried out using a modified protocol43. Proteoliposomes containing the Rnf complex had to be prepared under anoxic condition. Liposomes with a uniform diameter were diluted in transport buffer (100 mM Tris-HCl, pH 7.4, 100 mM maleic acid, 5 mM NaCl, 5 mM MgCl2) which was purged with N2 and contained 2 mM DTE and 4 µM resazurin for measurements with the Rnf complex. After destabilization of the liposomes, protein was added in a protein to lipid ratio of 1:10 (w/w). Further steps were carried out as described43.
Measurement of 22Na+ translocation
Measurement of 22Na+ translocation by the Rnf complex had to be performed under anoxic conditions. The buffer (100 mM Tris-HCl, pH 7.4, 100 mM maleic acid, 5 mM NaCl, 5 mM MgCl2) contained additional 2 mM DTE and 4 µM resazurin and was purged with N2. All transport studies were performed at 45 °C. Rnf batches were prepared in 3.5 ml glass vials containing 500 µl sample with a protein concentration of 1.4 mg/ml. Ferredoxin purified from C. pasteurianum according to ref. 44 was added to a concentration of 30 µM. The gas phase was adjusted to 0.5 × 105 Pa CO and carrier-free 22Na+ was added to a final concentration of 1 µCi/ml. After incubation for 30 min at room temperature, ACS/CODH purified from A. woodii according to ref. 31 was added (30 µg/ml). The batch was incubated another 5 min at 45 °C to ensure ferredoxin reduction and temperature equilibration. Addition of 3 mM NAD+ started the reaction. Measurement of 22Na+ translocation catalyzed by the ATP synthase was performed under oxic conditions. 3.5 ml glass vials were filled with 500 µl sample (protein concentration corresponded to 0.66–1.4 mg/ml). Carrier-free 22Na+ was added to the batch with final concentration of 1 µCi/ml. After an incubation time of 35 min (30 min at room temperature and 5 min at 45 °C), the reaction was started with the addition of 3 mM K2-ATP. Samples (40 µl) of both batches were removed and Na+ concentration was determined as described43.
Analytical methods
The protein concentration content of non-membranous samples was determined according to ref. 45, that of membranes according to ref. 46. Determination of protein concentration in proteoliposomes was performed as described47. Proteins were separated in 12% SDS polyacrylamide gels according to ref. 48. Native PAGE was performed as described49. Proteins were stained with either Coomassie Blue50 or silver51. Western blotting was done as described52. LC-MS/MS and MALDI measurements were performed by the “Functional Genomics Center Zürich”.
Statistics and reproducibility
One biological replicate is defined as one Rnf/ATPase complex or Rnf complex purification from one batch of T. maritima cells. With n ≥ 3 biological triplicates, all data points were taken and were used to calculate their mean with standard deviation SEM.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The authors declare that all data supporting the findings of this study are available within the article, Supplementary Information, and Supplementary Data 1. Source data underlying plots shown in figures are provided in Supplementary Data 1. The data that support the findings of this study are available on request from the corresponding author V.M.
References
Martin, W. F. & Thauer, R. K. Energy in ancient metabolism. Cell 168, 953–955 (2017).
Weber, J. & Senior, A. E. ATP synthesis driven by proton transport in F1F0 ATP synthase. FEBS Lett. 545, 61–70 (2003).
Schuchmann, K. & Müller, V. Autotrophy at the thermodynamic limit of life: a model for energy conservation in acetogenic bacteria. Nat. Rev. Microbiol. 12, 809–821 (2014).
Heider, J., Ma, K. & Adams, M. W. Purification, characterization, and metabolic function of tungsten-containing aldehyde ferredoxin oxidoreductase from the hyperthermophilic and proteolytic archaeon Thermococcus strain ES-1. J. Bacteriol. 177, 4757–4764 (1995).
Blamey, J. M. & Adams, M. W. Purification and characterization of pyruvate ferredoxin oxidoreductase from the hyperthermophilic archaeon Pyrococcus furiosus. Biochim. Biophys. Acta 1161, 19–27 (1993).
Buckel, W. & Thauer, R. K. Flavin-based electron bifurcation, a new mechanism of biological energy coupling. Chem. Rev. 118, 3862–3886 (2018).
Müller, V., Chowdhury, N. P. & Basen, M. Electron bifurcation: a long-hidden energy-coupling mechanism. Annu. Rev. Microbiol. 72, 331–353 (2018).
Schut, G. J. & Adams, M. W. The iron-hydrogenase of Thermotoga maritima utilizes ferredoxin and NADH synergistically: a new perspective on anaerobic hydrogen production. J. Bacteriol. 191, 4451–4457 (2009).
Schuchmann, K. & Müller, V. A bacterial electron bifurcating hydrogenase. J. Biol. Chem. 287, 31165–31171 (2012).
Biegel, E. & Müller, V. Bacterial Na+-translocating ferredoxin:NAD+ oxidoreductase. Proc. Natl Acad. Sci. USA 107, 18138–18142 (2010).
Schölmerich, M. C. & Müller, V. Energy conservation by a hydrogenase-dependent chemiosmotic mechanism in an ancient metabolic pathway. Proc. Natl Acad. Sci. USA 116, 6329–6334 (2019).
Sapra, R., Bagramyan, K. & Adams, M. W. W. A simple energy-conserving system: proton reduction coupled to proton translocation. Proc. Natl Acad. Sci. USA 100, 7545–7550 (2003).
Imkamp, F., Biegel, E., Jayamani, E., Buckel, W. & Müller, V. Dissection of the caffeate respiratory chain in the acetogen Acetobacterium woodii: indications for a Rnf-type NADH dehydrogenase as coupling site. J. Bacteriol. 189, 8145–8153 (2007).
Hedderich, R. Energy-converting [NiFe] hydrogenases from archaea and extremophiles: ancestors of complex I. J. Bioenerg. Biomembr. 36, 65–75 (2004).
Welte, C., Krätzer, C. & Deppenmeier, U. Involvement of Ech hydrogenase in energy conservation of Methanosarcina mazei. FEBS J. 277, 3396–3403 (2010).
Sapra, R., Verhagen, M. F. & Adams, M. W. Purification and characterization of a membrane-bound hydrogenase from the hyperthermophilic archaeon Pyrococcus furiosus. J. Bacteriol. 182, 3423–3428 (2000).
Yu, H. et al. Structure of an ancient respiratory system. Cell 173, 1636–1649.e1616 (2018).
Biegel, E., Schmidt, S., González, J. M. & Müller, V. Biochemistry, evolution and physiological function of the Rnf complex, a novel ion-motive electron transport complex in prokaryotes. Cell. Mol. Life Sci. 68, 613–634 (2011).
Biegel, E., Schmidt, S. & Müller, V. Genetic, immunological and biochemical evidence of a Rnf complex in the acetogen Acetobacterium woodii. Environ. Microbiol. 11, 1438–1443 (2009).
Schmehl, M. et al. Identification of a new class of nitrogen fixation genes in Rhodobacter capsulatus: a putative membrane complex involved in electron transport to nitrogenase. Mol. Gen. Genet. 241, 602–615 (1993).
Kumagai, H., Fujiwara, T., Matsubara, H. & Saeki, K. Membrane localization, topology, and mutual stabilization of the rnfABC gene products in Rhodobacter capsulatus and implications for a new family of energy-coupling NADH oxidoreductases. Biochemistry 36, 5509–5521 (1997).
Barquera, B. et al. Purification and characterization of the recombinant Na+-translocating NADH:quinone oxidoreductase from Vibrio cholerae. Biochemistry 41, 3781–3789 (2002).
Steuber, J. et al. The structure of Na+-translocating of NADH:ubiquinone oxidoreductase of Vibrio cholerae: implications on coupling between electron transfer and Na+ transport. Biol. Chem. 396, 1015–1030 (2015).
Hess, V. et al. Occurrence of ferredoxin:NAD+ oxidoreductase activity and its ion specificity in several Gram-positive and Gram-negative bacteria. PeerJ 4, e1515 (2016).
Fillingame, R. H. Identification of the dicyclohexylcarbodiimide-reactive protein component of the adenosine 5′-triphosphate energy-transducing system of Escherichia coli. J. Bacteriol. 124, 870–883 (1975).
Altendorf, K. & Zitzmann, W. Identification of the DCCD-reactive protein of the energy transducing adenosinetriphosphatase complex from Escherichia coli. FEBS Lett. 59, 268–272 (1975).
Kluge, C. & Dimroth, P. Kinetics of inactivation of the F1FO-ATPase of Propionigenium modestum by dicyclohexylcarbodiimide in relationship to H+ and Na+ concentration: probing the binding site for the coupling ions. Biochemistry 32, 10378–10386 (1993).
Spruth, M., Reidlinger, J. & Müller, V. Sodium ion dependence of inhibition of the Na+-translocating F1FO-ATPase from Acetobacterium woodii. Probing the site (s) involved in ion transport. Biochim. Biophys. Acta 1229, 96–102 (1995).
Hreha, T. N. et al. Complete topology of the Rnf complex from Vibrio cholerae. Biochemistry 54, 2443–2455 (2015).
Smith, E. T., Blamey, J. M., Zhou, Z. H. & Adams, M. W. A variable-temperature direct electrochemical study of metalloproteins from hyperthermophilic microorganisms involved in hydrogen production from pyruvate. Biochemistry 34, 7161–7169 (1995).
Hess, V., Schuchmann, K. & Müller, V. The ferredoxin:NAD+ oxidoreductase (Rnf) from the acetogen Acetobacterium woodii requires Na+ and is reversibly coupled to the membrane potential. J. Biol. Chem. 288, 31496–31502 (2013).
Tremblay, P. L., Zhang, T., Dar, S. A., Leang, C. & Lovley, D. R. The Rnf complex of Clostridium ljungdahlii is a proton-translocating ferredoxin:NAD+ oxidoreductase essential for autotrophic growth. MBio 4, e00406-00412 (2012).
Westphal, L., Wiechmann, A., Baker, J., Minton, N. P. & Müller, V. The Rnf complex is an energy coupled transhydrogenase essential to reversibly link cellular NADH and ferredoxin pools in the acetogen Acetobacterium woodii. J. Bacteriol. 200, e00357-00318 (2018).
Juárez, O., Athearn, K., Gillespie, P. & Barquera, B. Acid residues in the transmembrane helices of the Na+-pumping NADH:quinone oxidoreductase from Vibrio cholerae involved in sodium translocation. Biochemistry 48, 9516–9524 (2009).
Mezic, K. G. et al. Glutamate 95 in NqrE is an essential residue for the translocation of cations in Na+-NQR. Biochemistry 58, 2167–2175 (2019).
Schröder, C., Selig, M. & Schönheit, P. Glucose fermentation to acetate, CO2 and H2 in the anaerobic hyperthermophilic eubacterium Thermotoga maritima—involvement of the Embden-Meyerhof pathway. Arch. Microbiol. 161, 460–470 (1994).
Demmer, J. K. et al. Insights into flavin-based electron bifurcation via the NADH-dependent reduced ferredoxin:NADP+ oxidoreductase structure. J. Biol. Chem. 290, 21985–21995 (2015).
Huber, R. et al. Thermotoga maritima sp. nov. represents a new genus of unique extremely thermophilic eubacteria growing up to 90 °C. Arch. Microbiol. 144, 324–333 (1986).
Schut, G. J., Boyd, E. S., Peters, J. W. & Adams, M. W. The modular respiratory complexes involved in hydrogen and sulfur metabolism by heterotrophic hyperthermophilic archaea and their evolutionary implications. FEMS Microbiol. Rev. 37, 182–203 (2013).
Heinonen, J. E. & Lahti, R. J. A new and convenient colorimetric determination of inorganic orthophosphate and its application to the assay of inorganic pyrophosphatase. Anal. Biochem. 113, 313–317 (1981).
Cesari, I. M., Rieber, M. & Imaeda, T. Localization and properties of enzymes involved with electron transport activity in mycobacteria. J. Bacteriol. 98, 767–773 (1969).
Pisa, K. Y., Huber, H., Thomm, M. & Müller, V. A sodium ion-dependent A1AO ATP synthase from the hyperthermophilic archaeon Pyrococcus furiosus. FEBS J. 274, 3928–3938 (2007).
Mayer, F., Lim, J. K., Langer, J. D., Kang, S. G. & Müller, V. Na+ transport by the A1AO-ATP synthase purified from Thermococcus onnurineus and reconstituted into liposomes. J. Biol. Chem. 290, 6994–7002 (2015).
Schönheit, P., Wäscher, C. & Thauer, R. K. A rapid procedure for the purification of ferredoxin from clostridia using polyethylenimine. FEBS Lett. 89, 219–222 (1978).
Bradford, M. M. A rapid and sensitive method for the quantification of microgram quantities of protein utilizing the principle of proteine-dye-binding. Anal. Biochem. 72, 248–254 (1976).
Lowry, O. H., Rosebrough, N. J., Farr, A. L. & Randall, R. J. Protein measurement with the folin-phenol reagent. J. Biol. Chem. 193, 265–275 (1951).
Kaplan, R. S. & Pedersen, P. L. Determination of microgram quantities of protein in the presence of milligram levels of lipid with amido black 10B. Anal. Biochem. 150, 97–104 (1985).
Schägger, H. & von Jagow, G. Tricine-sodium dodecylsulfate-polyacrylamide gel electrophoresis for the separation of proteins in the range from 1 to 100 kDa. Anal. Biochem. 166, 369–379 (1987).
Wittig, I., Carrozzo, R., Santorelli, F. M. & Schägger, H. Functional assays in high-resolution clear native gels to quantify mitochondrial complexes in human biopsies and cell lines. Electrophoresis 28, 3811–3820 (2007).
Weber, K. & Osborne, M. The reliability of the molecular weight determination by dodecyl sulfate polyacrylamide gel electrophoresis. J. Biol. Chem. 244, 4406–4412 (1969).
Blum, H., Beier, H. & Gross, H. J. Improved silver staining of plant proteins, RNA, and DNA in polyacrylamide gels. Electrophoresis 8, 93–98 (1987).
Hess, V., Vitt, S. & Müller, V. A caffeyl-coenzyme A synthetase initiates caffeate activation prior to caffeate reduction in the acetogenic bacterium Acetobacterium woodii. J. Bacteriol. 193, 971–978 (2011).
Acknowledgements
This work was supported by a grant (MU 801/16-1) from the Deutsche Forschungsgemeinschaft. Open access funding provided by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
M.K. planned and carried out the experiments and interpreted the data. D.T. planned and carried out the experiments, interpreted the data and prepared the manuscript. H.H. supplied the T. maritima cells for the experiments. V.M. planned the experiments, interpreted the data and prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kuhns, M., Trifunović, D., Huber, H. et al. The Rnf complex is a Na+ coupled respiratory enzyme in a fermenting bacterium, Thermotoga maritima. Commun Biol 3, 431 (2020). https://doi.org/10.1038/s42003-020-01158-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-020-01158-y
This article is cited by
-
Architecture of the RNF1 complex that drives biological nitrogen fixation
Nature Chemical Biology (2024)
-
The oxidoreductase activity of Rnf balances redox cofactors during fermentation of glucose to propionate in Prevotella
Scientific Reports (2023)
-
Life strategies for Aminicenantia in subseafloor oceanic crust
The ISME Journal (2023)
-
Physiological and stoichiometric characterization of ethanol-based chain elongation in the absence of short-chain carboxylic acids
Scientific Reports (2023)
-
Gene gain facilitated endosymbiotic evolution of Chlamydiae
Nature Microbiology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.