Abstract
Large protein families are a prominent feature of plant genomes and their size variation is a key element for adaptation. However, gene and genome duplications pose difficulties for functional characterization and translational research. Here we infer the evolutionary history of the DOMAIN OF UNKNOWN FUNCTION (DUF) 26-containing proteins. The DUF26 emerged in secreted proteins. Domain duplications and rearrangements led to the appearance of CYSTEINE-RICH RECEPTOR-LIKE PROTEIN KINASES (CRKs) and PLASMODESMATA-LOCALIZED PROTEINS (PDLPs). The DUF26 is land plant-specific but structural analyses of PDLP ectodomains revealed strong similarity to fungal lectins and thus may constitute a group of plant carbohydrate-binding proteins. CRKs expanded through tandem duplications and preferential retention of duplicates following whole genome duplications, whereas PDLPs evolved according to the dosage balance hypothesis. We propose that new gene families mainly expand through small-scale duplications, while fractionation and genetic drift after whole genome multiplications drive families towards dosage balance.
Similar content being viewed by others
Introduction
Gene duplication and loss events constitute the main factor of gene family evolution1. Duplications occur by two major processes, whole-genome multiplications (WGM) and small-scale duplications (SSD), including tandem, segmental, and transposon-mediated duplications2. They appear to be distinct modes of expansion, since gene families evolving through WGMs rarely experience SSD events3. The division is also visible on the functional level, since genes duplicated in WGMs are enriched for signal transduction, transcriptional and developmental regulation as well as signal transduction functions, whereas SSDs occur preferentially on secondary metabolism and environmental response genes3. The prevailing hypothesis for the phenomenon is dosage balance; in regulatory networks and protein complexes the stoichiometric balance between different components needs to be preserved; thus, selection acts against losses following WGMs and against duplications in SSDs4. Families retained after WGMs are typically stable across different species whereas highly variable families evolve through SSDs5, suggesting high turnover rates. These results have been obtained by analyzing the extremes, top families displaying pure WGM retention or SSD characteristics3 while most gene families likely evolve in an intermediate manner.
Plants and other eukaryotes have developed a wide range of signal transduction mechanisms for controlling cellular functions and coordinating responses on cell, tissue, organ, and organismal level. Plants in particular encode large gene families of secreted proteins6,7,8 and proteins with extracellular domains to respond to environmental and developmental cues. However, the large numbers make it difficult to dissect conserved or specialized functions4, and therefore a detailed understanding of their evolution and duplication history in different plant lineages is needed. Signaling proteins with extracellular domains include receptor-like protein kinases (RLKs)9,10 and receptor-like proteins (RLPs)11. In RLKs, extracellular domains are involved in signal perception and protein–protein interactions12 while the intracellular kinase domain transduces signals to substrate proteins. RLKs are involved in essential mechanisms including stress responses, hormone signaling, cell wall monitoring, and plant development12. The large number of secreted proteins, RLKs, and RLPs in plants may reflect their sessile lifestyle and need for meticulous monitoring of signals from other cells, tissues, or the environment. Phylogenetic relations between different groups of RLKs and RLPs have been described9,13,14,15,16 but few have been physiologically and biochemically characterized17.
The Domain of Unknown Function 26 (DUF26; Gnk2 or stress-antifungal domain; PF01657)18,19 is an extracellular domain harboring a conserved cysteine motif (C-8X-C-2X-C) in its core. It is present in three types of plant proteins. The first class is CYSTEINE-RICH RECEPTOR-LIKE SECRETED PROTEINs (CRRSPs). The best characterized CRRSP is Gnk2 from Gingko biloba with single DUF26, which acts as mannose-binding lectin in vitro with antifungal activity18,19. Two maize CRRSPs have been shown to also bind mannose and participate in defence against a fungal pathogen20. The second class, CYSTEINE-RICH RECEPTOR-LIKE PROTEIN KINASES (CRKs), has a typical configuration of two DUF26 in the extracellular region and forms a large subgroup of RLKs in plants. CRKs participate in the control of stress responses and development in Arabidopsis and in rice21,22,23,24,25,26,27,28,29,30,31. The third class of DUF26 domain-containing proteins is the PLASMODESMATA-LOCALIZED PROTEINS (PDLPs). PDLPs contain two DUF26 domains in their extracellular region and a transmembrane helix, but lack a kinase domain. They associate with plasmodesmata and are involved in symplastic intercellular signaling32, pathogen response33, systemic signaling34, control of callose deposition35 and are targets for viral movement proteins36. However, the precise biochemical functions of plant DUF26-containing proteins remain unclear.
Tandem expansions drive the evolution of, for example, F-Box proteins37, and transcription factors38, but also RLKs16 and RLPs11. Diversification processes include sub-functionalization, where paralogs retain a subset of their ancestral functions, and neofunctionalization, where duplicated proteins acquire new functions38. CRKs and CRRSPs typically exist in clusters on plant chromosomes24, suggesting relatively recent tandem expansions. This makes DUF26-containing proteins a perfect dataset for sequence-based evolutionary investigation.
Here we carry out an in-depth analysis of the DUF26-containing proteins, a protein family involved in signaling, to explore the dynamics and effect of the different duplication mechanisms on overall gene family evolution. We combine phylogenetic analyses with experimental structural biology to gain insight into the evolution of DUF26-containing proteins. While sequence analysis indicates that the DUF26 domain is specific to land plants, the domain shows structural similarity to fungal carbohydrate-binding lectins. Our results suggest that DUF26-containing proteins constitute a group of carbohydrate-binding proteins in plants. CRKs and CRRSPs experienced both ancestral and recent lineage-specific tandem duplications. In contrast to the general pattern of gene families expanding by small-scale duplication events, these gene families experienced expansion also during or after WGMs. Our work illustrates that detailed understanding of the evolution of large protein families is a prerequisite for translating findings from model plants to different species and for dissecting conserved or specialized functions of proteins.
Results
DUF26-containing proteins have diverse domain compositions
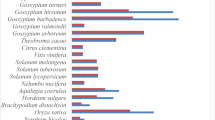
We selected 32 species with high-quality genome assemblies representing major plant lineages. After manual curation, de novo annotation, and exclusion of partial gene models and pseudogenes, we obtained 1409 high-quality gene models (Fig. 1a, Supplementary Figure 1, Supplementary Note 1, Supplementary Data 1). The PFAM protein domain database39 identifies DUF26 as specific to embryophytes. Accordingly, we identified no DUF26 or DUF26-like domains from genomes of algae, charophytes, diatoms, fungi, insects or vertebrates. DUF26-containing proteins are grouped into three categories: CRRSPs, PDLPs, and CRKs (Fig. 1b). CRRSPs consist of a signal peptide followed by one or more DUF26 domains, separated by a variable region. CRRSPs with a single DUF26 (sdCRRSPs) were identified from most land plants, including the liverwort (Marchantia polymorpha) and moss (Physcomitrella patens) lineages (Fig. 1). CRRSPs with two DUF26 domains (ddCRRSPs) were identified from vascular plants including the lycophyte Selaginella moellendorffii and represent the predominant type in vascular plants (Fig. 1). Rice as well as Brassicaceae display lineage-specific evolution with a large number of ddCRRSPs while sdCRRSPs are absent (Fig. 1a, Supplementary Figure 2).
Overview and distribution of DUF26-containing genes in plants. a DUF26-containing genes are absent from algae and charophytes but present in land plants. Marchantia polymorpha and Physcomitrella patens genomes encode sdCRRSPs. Selaginella moellendorffii possesses sdCRRSPs, sdCRKs, and canonical CRKs. Seed plant (gymnosperm and angiosperm) genomes encode the whole set of DUF26-containing genes. CRKs were defined as basal group CRKs (bCRKs) or variable group CRKs (vCRKs) based on their phylogenetic positions. Whole-genome duplication (WGD) events are presented with green circle and whole-genome triplication (WGT) events with dark blue circle. Ferns were omitted from analyses due to lack of available genome assemblies. b Overview of different domain compositions of proteins containing DUF26 in different plant lineages. The number of representative species in the analyses is given in brackets after the name of the group. Numbers in the table present the number of species in each lineage in which the domain structure was found. Abbreviations sd (single domain), dd (double domain), td (triple domain), and qd (quadruple domain) refer to the number of the DUF26 domains
CRKs contain a signal peptide, two DUF26 domains, and a transmembrane region followed by an intracellular protein kinase domain. Similar to ddCRRSPs, CRKs were identified from vascular plants but not from bryophytes (Fig. 1a). The CRKs likely originate from a fusion of sdCRRSPs with transmembrane region and kinase domain from LRR_clade_3 RLKs in the common ancestor of vascular plants15. The Selaginella genome uniquely encodes single DUF26 CRKs (sdCRKs; Fig. 1b) and only few CRKs from eudicots contain more than two DUF26 domains.
PDLPs were identified from all seed plants and are composed of a signal peptide, two DUF26, and a transmembrane region followed by short cytoplasmic extension. We also identified several angiosperm CRKs lacking signal peptide, extracellular domain, and transmembrane region. These are subsequently referred to as CYSTEINE-RICH RECEPTOR-LIKE CYTOPLASMIC KINASEs (CRCKs).
Evolution of CRKs, PDLPs, and ddCRRSPs from small sdCRRSPs
We investigated the evolution of CRRSPs, CRKs, and PDLPs by estimating phylogenetic trees (Supplementary Note 2). Overall, the tree of DUF26-containing proteins split into two distinct groups, basal group α and variable group β (Fig. 2a, b), where α is paraphyletic with respect to β. To increase the number of informative sites and obtain better resolution, we estimated separate phylogenetic trees for both groups (Supplementary Figure 2a, b). The subgroupings of the basal α- and variable β-groups remained conserved also in trees estimated for each subfamily of DUF26-containing proteins (Supplementary Figure 2c–e). Subsequently, we reconciled the gene trees with the species tree, and estimated ancestral gene contents and duplication/loss events for the subfamilies in 11 species (Supplementary Figure 3). To identify significant expansions we fitted birth–death rate models for DUF26-containing protein families and compared the rates against computationally derived gene families (orthogroups) for RLKs, all protein kinases, and plasmodesmal proteins40 using Badirate41. Finally, we assessed selective pressure by estimating amino acid conservation patterns around the main cysteine motif of the DUF26 domains for major subfamilies, and found conserved sites specific to α- or β-groups (Fig. 2, Supplementary Note 3).
Phylogenetic tree of CRRSPs, CRKs, and PDLPs. a The phylogenetic tree was estimated with the maximum-likelihood method using all high-quality full-length DUF26-containing sequences from lycophytes onwards. CRCKs and concA-CRKs were excluded while GNK2 from Gingko biloba was included. Overall, DUF26-containing genes split into basal and variable groups. Detailed phylogenetic trees with bootstrap support (1000 replicates) and filtered sequence alignments are available at http://was.bi?id=IaroP (full tree), http://was.bi?id=wpEHGt (basal group separately), and http://was.bi?id=aIJe_D (variable group separately). b The same phylogenetic tree as in panel a rooted to ancestral sdCRRSPs and sdCRKs from Selaginella moellendorffii showing that the variable group branches out from the basal group. c The MEME figures present the conservation pattern of amino acid positions around the main cysteine motif within the DUF26 domains for sdCRRSPs, bCRKs, and PDLPs from the basal group and CRRSPs and vCRKs from the variable group. The features specific only to genes either in the basal group or in the variable group are highlighted. d The DUF26-A and DUF26-B domains are clearly separated in an unrooted phylogenetic tree containing DUF26 domain sequences. The MEME figures present differences in the conservation of the AA sequence surrounding the conserved cysteines in DUF26-A and DUF26-B
The α-group is likely older, containing sequences from all vascular plants. Proteins in this group are conserved on sequence level and identification of putative orthologs from different species is frequently possible. Purifying selection, selective removal of (deleterious) variations, is likely the main force acting on this clade, as suggested by low dN/dS values (one-rate model for whole groups: bCRK-I 0.184, bCRK-II 0.192, PDLPs 0.267, sdCRRSPs 0,162, CRCK 0.134; more flexible model with branch-specific dN/dS within each group yielded similar results). The subgroups within the basal α-group evolved independently and their DUF26 domains share features distinguishing them from the β-group (Fig. 2c, Supplementary Note 3). Since the sdCRRSPs are located close to the root of the α-group (Fig. 2b) and form a monophyletic subclade at the root of the CRRSP tree (Supplementary Figure 2c), they are likely the ancestral type of DUF26 proteins in land plants. Furthermore, sdCRRSPs are present in various early diverging plant lineages such as the gymnosperm Ginkgo biloba (including Gnk218,19) and the liverwort, Marchantia polymorpha (Supplementary Figure 2f). Turnover rates of sdCRRSPs do not differ from those of all gene families and show lineage-specific expansions in early diverging species (Supplementary Figure 3a).
The placement of Selaginella sdCRKs to the root of the CRK phylogeny (Supplementary Figure 2d) and as sister to sdCRRSPs in the α-group (Fig. 2b) suggests an ancient origin. The DUF26 domain likely duplicated after fusing with transmembrane region and kinase domain, thus establishing the typical double DUF26 CRK configuration (Supplementary Figure 2d). Following duplication, the two DUF26 domains diverged into distinct, evolutionarily conserved, forms, DUF26-A and DUF26-B (Fig. 2d). All CRKs expanded significantly in the branches leading to lycophytes (P = 0.0017600) and to angiosperms (P = 0.0151412) compared to all RLKs (Fig. 3), and in the branch from lycophytes to angiosperms compared to all protein kinases (Supplementary Figure 4a).
Comparison of evolutionary rates between gene families. Analyses were carried out with Badirate for 11 species (Physcomitrella patens, Selaginella moellendorffii, Amborella trichopoda, Arabidopsis thaliana, Populus trichocarpa, Vitis vinifera, Aquilegia coerulea, Spirodela polyrhiza, Zea mays, Oryza sativa, and Brachypodium distachyon). Neutral branches are reported as bold black lines; branches involving gene family expansion are reported as bold purple lines and branches with contraction as blue dashed lines. Branches with significant differences (false discovery rate adjusted p < 0.05) to birth–death rate model estimates are marked with arrows. Node labels present the ancestral gene family sizes estimated by Badirate. Tip labels contain species abbreviations and the change in numbers compared to the most recent ancestral node. a All CRKs compared to other receptor-like kinases (RLKs). b bCRKs compared to RLKs. c PDLPs compared to other plasmodesmata-related orthogroups. d vCRKs compared to RLKs. e Phylogenetic maximum-likelihood tree showing differences in lineage-specific expansions in monocot and dicot vCRKs following the split of Amborella trichopoda. Species-specific expansions (at least two genes from same species) are marked with red and clades including sequences from only Brassicaceae or Solanaceae are marked with blue
A monophyletic group of CRKs with representatives from gymnosperms and angiosperms is located near the base of the CRK phylogeny (Supplementary Figure 2d) in the α-group (Figs 2a, b). This group, referred to as basal CRKs (bCRKs), likely represents the ancient CRKs in seed plants. Following the initial innovation in ancestral vascular plants, the group evolved at rates similar to orthogroups containing all protein kinases or all RLKs (Fig. 3b, Supplementary Figures 3b and 4b). The bCRKs split into two subgroups, bCRK-I and bCRK-II (Fig. 2b and Supplementary Figure 5), in gymnosperms and angiosperms, suggesting divergence in early seed plants. The larger bCRK-I subclade further divides into distinct branches with tandemly duplicated Amborella bCRKs at their roots (Supplementary Figures 5 and 6a, b) suggesting rapid differentiation following duplication in ancestral angiosperms42. The number of the bCRK-Is is conserved, excluding an expansion specific to Solanaceae, while the small bCRK-II subclade is absent from Brassicaceae.
PDLPs were only found in seed plants. PDLPs belong to the α-group (Fig. 2a, b) and represent the most conserved class of DUF26-containing genes. PDLPs do not display different expansion rates compared to plasmodesmata-related orthogroups40 (Fig. 3c). PDLPs split into two clades, PDLP-I and PDLP-II (Supplementary Figure 2e), which both contain Amborella trichopoda and eudicot and monocot PDLPs, suggesting divergence in ancestral angiosperms. PDLPs and ddCRRSPs originate from CRKs through loss of kinase domains and/or transmembrane regions. The loss can be a two-step process, as exemplified by an atypical PDLP from Amborella trichopoda located at the root of the main ddCRRSP clade (Supplementary Figure 7). PDLPs were possibly present already in ferns since database searches identified a partial gene model lacking a transmembrane region in Marsilea quadrifolia43 (see Methods) with similarity to PDLPs, placed at the root of a phylogenetic tree for PDLPs (Supplementary Figure 8).
In the α-group, a group of spruce-specific CRKs (spruce vCRKs) are more related to PDLPs than other CRKs (Fig. 2a, b). They form a distinct group between bCRKs and a large group of angiosperm variable CRKs (vCRKs; Fig. 2a, b, Supplementary Figures 2d and 3d). Angiosperm vCRKs form the β-group together with ddCRRSPs and atypical monocot sdCRRSPs (Fig. 2a, b). These CRRSPs likely evolved from vCRKs through loss of transmembrane regions and kinase domains and, in case of sdCRRSPs, also DUF26-B domains. The β-group is less conserved compared to the α-group and branches into two eudicot-specific and one monocot-specific group with a few Amborella trichopoda vCRKs at the root of the groups. Members of the β-group experienced several independent tandem expansions in different plant taxa (Figs 3d, e, Supplementary Figures 3d, 4c and 6c) and expanded during the diversification of monocots and dicots. CRRSPs in the β-group are not monophyletic, suggesting independent birth from partial duplications of vCRKs. Hence, expansion rates and extrapolation of ancestral gene counts for ddCRRSPs could not be reliably predicted. Lineage-specific expansions in the β-group make ortholog identification challenging.
DUF26 is related to fungal lectins and assembles as tandems
High sequence divergence of DUF26 proteins and the lineage-specific expansions raise the question whether their overall structure is conserved. The consensus DUF26 domain as defined in PFAM comprises ~90–110 amino acids. Structural information is currently available for the sdCRRSP Gnk219 but not for ddCRRSPs, CRKs, or PDLPs, proteins with a double DUF26 configuration. Mechanistic constraints restrict the evolution of protein structures, and therefore understanding structural conservation provides clues for protein function.
We defined the structural relationship of tandem DUF26 domains by determining crystal structures of AtPDLP5 (residues 26–241) and AtPDLP8 (21–253) ectodomains to 1.25 and 1.95 Å resolution, respectively (Supplementary Table 1). Individual DUF26 domains feature two small α-helices folding on top of a central antiparallel β-sheet (Fig. 4). The PDLP5 DUF26-A domain is N-glycosylated at positions Asn69 and Asn132 in our crystals (Fig. 4a). The secondary structure elements of DUF26 are covalently linked by three disulfide bridges, formed by six conserved Cys residues, part of which belong to the C-8X-C-2X-C motif (Figs. 2d and 4a). We previously suggested that tandem DUF26-containing proteins could be involved in ROS or redox sensing24,26. To assess the functional roles of the invariant disulfide bridges in PDLPs, we mutated the partially solvent exposed PDLP5Cys101, PDLP5Cys148, and PDLP5Cys191 to alanine. While the wild-type PDLP5 ectodomain behaves as a monomer in solution (Supplementary Figure 9), the mutant protein tends to aggregate in our biochemical preparations (Supplementary Figure 9) and display reduced structural stability in thermofluor assays (Supplementary Figure 10). These experiments and crystallographic data (Fig. 4a) suggest that the conserved disulfide bonds in PDLPs and potentially in other DUF26-containing proteins are involved in structural stabilization rather than redox signaling.
The crystals structures of the PDLP5 and PDLP8 ectodomains reveal a conserved tandem architecture of two lectin-like domains. a Overview of the PDLP5 ectodomain. The two DUF26 domains are shown as ribbon diagrams, colored in blue (DUF26-A) and orange (DUF26-B), respectively. N-glycans are located at Asn69 and Asn132 of DUF26-A and are depicted in bonds representation (in cyan). The DUF26-A and DUF26-B domains each contain three disulfide bridges labeled 1 (Cys89–Cys98), 2 (Cys101–Cys126), 3 (Cys36–Cys113), 4 (Cys191–Cys200), 5 (Cys203–Cys228), and 6 (Cys148–Cys215). b Close-up view of the DUF26-A–DUF26-B interface in PDLP5 (orange) and PDLP8 (blue), shown in bonds representation. c Superimposition of the Gnk2 extracellular DUF26 domain (PDB-ID 4XRE) with either PDLP5 DUF-26A (r.m.s.d. is ~1.4 Å comparing 100 aligned Cα atoms) or PDLP5 DUF26-B (r.m.s.d. is ~2.0 Å comparing 93 corresponding Cα atoms). Corresponding disulfide bridges shown in bonds representation (PDLP5 in green, Gnk2 in yellow) are highlighted in gray. Gnk2-bound mannose is shown in magenta (in bonds representation). d Close-up view of the residues involved in the binding of mannose of Gnk2 (bonds representation, in blue and magenta, respectively) and putative residues involved in ligand binding of PDLP5 DUF26-A (in orange). e The fungal LDL DUF26 domain (Cα trace in blue; PDB-ID 4NDV) and PDLP5 DUF26-A (in orange) superimposed with an r.m.s.d. of ~2.4 Å comparing 75 aligned Cα atoms). Disulfide bridges (LDL in yellow and PDLP5 in green; aligned disulfide bridges highlighted in gray) and the LDL bound globotriose (magenta) are shown in bonds representation. f Cα traces of the structural superimposition of the fungal Y3 protein (PDB-ID 5V6I) and PDLP5 DUF26-A (r.m.s.d. is ~2.6 Å comparing 78 corresponding Cα atoms). Disulfide bridges of Y3 (yellow) and PDLP5 DUF26-A (green) are shown alongside, one corresponding disulfide pair is highlighted in gray
The N-terminal DUF26-A (PDLP5 residues 30–132) and the C-terminal DUF26-B (residues 143–236) domains are connected by a structured loop (residues 133–142) and make extensive contacts with each other (Fig. 4a). The resulting ectodomain has a claw-like shape with the β-sheets of DUF26-A and B facing each other (Fig. 4a). The DUF26-A and B domains in PDLP5 and 8 closely align, with root mean square deviations (r.m.s.d.s) of 1.6 and 1.2 Å when comparing 89 corresponding Cα atoms, respectively (Supplementary Figure 11a). DUF26-A is more variable than DUF26-B on the sequence level (Fig. 2d). The DUF26-A and -B domains in PDLP5 and PDLP8 have 24% and 30% of their residues in common, most of which map to the hydrophobic core of the domain (including the six cysteine residues forming intra-molecular disulfide bonds) and to the DUF26-A–DUF26-B interface (Fig. 4b). This interface is formed by a line of aromatic and hydrophobic residues originating from the proximal face of the β-sheet in DUF26-A and -B (Fig. 4b, Supplementary Figure 12). Importantly, many of the interface residues are conserved among different PDLPs, but also among CRKs and ddCRRSPs (Supplementary Figure 12). Consistently, the ectodomains of PDLP5 and PDLP8 belonging to different phylogenetic clades (Supplementary Figure 8) closely align with an r.m.s.d. of ~1.6 Å when comparing 198 corresponding Cα atoms (Supplementary Figure 11b). These observations suggest that evolutionarily distant DUF26 tandem proteins likely share the conserved three-dimensional structure.
The physiological ligands for PDLPs are currently unknown. Therefore, we performed structural homology searches44 to obtain insights into the biochemical function of plant DUF26 domains (see Methods). Top hits include the single DUF26 domain protein Gnk219. Despite moderate sequence similarity, the overall fold of Gnk2 and PDLP5 DUF26-A and B as well as disulfide-bond arrangement is conserved (Fig. 4c). Notably, Glu130 and Arg132 implicated in mannose binding in Gnk2 are replaced by Asp131 and Lys133 in the DUF-A of PDLP5, respectively (Fig. 4d). A similar pocket is found in the DUF-A domain of PDLP8, but not in the DUF-B domains of either PDLP5 or 8. Despite these structural homologies of Gnk2, PDLP5 DUF-A, and PDLP8 DUF-A, we could not detect binding of mannose to isolated PDLP5 ectodomain in vitro (Supplementary Figure 13a). We are also unable to detect any binding of other water-soluble cell wall-derived carbohydrates to the PDLP5 ectodomain (Supplementary Figure 13b). The PDLP5 DUF26 domains share strong structural homology also with two fungal lectins, the α-galactosyl-binding Lyophyllum decastes lectin (LDL)45 and a glycan-binding Y3 lectin from Coprinus comatus46. Both proteins closely align with the plant DUF26 domain, and share one of the three disulfide bridges (Fig. 4e–f). The surface areas involved in globotriose and glycan binding, respectively, are not conserved in PDLPs, but the structural similarity of plant DUF26 domains with different eukaryotic lectins could suggest a common evolutionary origin and a role as carbohydrate recognition modules45.
We next explored potential binding sites in the two molecules by analyzing sitewise ω for orthologs of PDLP5 and PDLP8. Low ω values were observed in structural context, indicating conservation of residues buried inside the DUF26 domain, while variable residues (under more relaxed selection) appear on the surface of the structure (Fig. 5a). The variability of the PDLP5 and PDLP8 DUF26 domain surface may be central to their ability to interact with other proteins or ligands (Supplementary Figure 13c). Selection patterns may differ between young lineage-specific and evolutionarily conserved proteins. For PDLP5, high ω values on the surface could indicate fast evolution leading to sub- or neofunctionalization, as PDLP5 orthologs originate from the recent lineage-specific duplication in Brassicaceae. The different surface charge properties of related PDLPs from Arabidopsis (Fig. 5b) suggest that different PDLPs and other DUF26-containing proteins sense a diverse set of ligands. While the nature of these molecules is currently unknown, cell-wall-derived carbohydrates or small extracellular molecules represent candidate ligands. Notably, we observed typical lectin-dimers in PDLP5 and PDLP8 crystals, in which two lectin domains dimerized along an extended antiparallel β-sheet (Fig. 5c)47. In principle, this mode of dimerization could form an extended binding cleft for a carbohydrate polymer, and presents an attractive activation mechanism for PDLPs and CRKs, where a monomeric ground state forms ligand-induced oligomers, as previously seen with plant LysM-domain-containing carbohydrate receptors48.
PDLP5 and PDLP8 may have drastically different oligomerisation modes; surface charge distributions and surface-exposed residues are not widely conserved. a The conservation of amino acid residues illustrated on the molecular surface of the PDLP5 or PDPL8 crystallization dimers, respectively. Sitewise ω (dN/dS) values, indicating the intensity and direction of selection on amino acid changing mutations, illustrated on the molecular surfaces and in ribbon diagrams of PDLP5 or PDPL8. The ω-values range from 0.15 (green) to slightly over 1.0 (magenta), reflecting conserved sites under purifying selection and sites evolving close to a neutral process, respectively. b Electrostatic potential mapped onto molecular surfaces of the putative PDLP5 and PDLP8, orientation as in c dimer, respectively. c Ribbon diagrams of PDLP5 (orange) and PDLP8 (blue) crystallographic dimers. In both dimers large, antiparallel β-sheets are formed, using different protein–protein interaction surfaces
The CRK kinase domain is related to LRR and S-locus RLKs
Kinase domains transduce signals by phosphorylating substrate proteins. Typically, the kinase domain has been used to investigate phylogenetic relationships between RLKs9,15,16. The CRK kinase domain is similar to the kinase domain of S-locus lectin and LRR RLKs from LRR_clade_315 (Supplementary Data 2). Based on catalytic motifs in the kinase domains49 most CRKs seem to be active kinases and in vitro activity of several CRKs has been experimentally confirmed25,28,30. Most CRKs belong to the RD type50,51 considered capable of auto-activation but few non-RD CRKs are present in plants49.
Analyzing ectodomains and kinase domains of CRKs separately suggests that Selaginella ddCRKs share an ancestor with bCRKs, while Selaginella sdCRKs share an ancestor with vCRKs (Fig. 6a). The separation of DUF26-A and DUF26-B (Fig. 2d) and the timing of those events does not reveal whether duplication of the DUF26 domain in the CRK extracellular region occurred more than once or whether functional constraints in the kinase domain led to the similarity of Selaginella sdCRKs and vCRKs. Juxtaposition of phylogenetic trees based on ectodomains and kinase domain suggests several exchanges of kinase or extracellular regions among CRKs during evolution (Fig. 6a). Most strikingly, a group of monocot-specific CRKs separates from other CRKs in a phylogenetic tree based on the kinase domain (Fig. 6a). Those CRKs have a different exon–intron structure (Fig. 6b, Supplementary Figure 1b) and a kinase domain with high similarity to concanavalin-A-like lectin protein kinase domains (Supplementary Data 2) altogether suggestive of chimeric gene formation following tandem duplication52. This kinase domain switch is specific to grasses (Poaceae) and has likely resulted in a different set of substrates. In addition, loss of ectodomains and transmembrane regions has established CRCKs at least three times; one group is specific to angiosperms (CRCK-I clade), one is specific to Brassicaceae and one only to Arabidopsis thaliana.
CRKs experienced domain rearrangements. a Comparison of phylogenetic trees based on ectodomain region and kinase domain of 880 CRKs. Phylogenetic maximum-likelihood trees are presented as tanglegram where the tree of the CRK ectodomain region is plotted against the tree of the kinase domain. The kinase tree is rooted to atypical monocot CRKs with a Concanavalin-A type kinase domain and the ectodomain tree is rooted to CRKs from Selaginella moellendorffii. The ectodomain tree was detangled based on the kinase domain tree. Lines connect the ectodomain and kinase domain belonging to same gene, and connection are drawn in different colors for better visibility. Juxtaposition of the trees shows rearrangements and domain swaps of ecto- and kinase domains. Black circles highlight the difference between the ectodomains and kinase domains of the Selaginella sdCRKs and ddCRKs and also the group of the atypical monocot CRKs which have exchanged the kinase domain. b The exon–intron structure of the CRKs. Usually CRKs contain seven exons: one encoding DUF26 domains, one encoding transmembrane region (TMR), and five exons encoding the kinase domain. In atypical monocot CRKs with exchanged kinase domain, whole gene is encoded by one or two exons. The scale bar for each gene represents 100 bases. Regions encoding the DUF26-A are colored with blue, the DUF26-B with orange, the transmembrane region (TMR) with pink, and the kinase domain with green
Mixed-mode evolution of large gene families
For detailed analyses of gene family dynamics we analyzed the synteny, conservation of gene order between species, and tandem duplications in Amborella trichopoda, tomato (Solanum lycopersicum), Arabidopsis, rice, and maize (Zea mays; Fig. 7, Supplementary Figure 7), and estimated the timing of duplication events by reconciliation of gene trees with species trees (Supplementary Figure 14a).
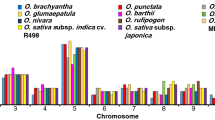
Identification of the modes of gene family evolution in DUF26-containing genes in Arabidopsis thaliana, tomato, rice, maize, and Amborella trichopoda. a Gene families that are preferentially retained after whole-genome multiplications (WGMs) are typically identified by synteny analysis. The figure illustrates syntenic regions containing DUF26 genes from Amborella trichopoda to monocots Oryza sativa and Zea mays and to eudicots Solanum lycopersicum and Arabidopsis thaliana. Within monocots and dicots, segments with at least five syntenic genes were included, whereas in comparisons to Amborella the minimum threshold was three syntenic genes. For Amborella trichopoda genomic locations of DUF26-containing genes are only known on chromosome/scaffold level based on physical mapping. b, c Gene families with a preferential retention pattern after WGMs show conserved gene counts over species. Phylogenetic tree of the five species (a) was used to reconcile the gene trees and estimate gene counts in ancestral nodes for b bCRKs and c vCRKs, using Selaginella moellendorffii as outgroup. The gains are highlighted with red and losses with blue. d Gene families with preferential retention pattern should have many orthologs. Heatmaps of the normalized transcriptional expression counts (Transcript per million [TPM]) of candidate DUF26 orthologs from four of the species: Solanum lycopersicum, Arabidopsis thaliana, Zea mays, and Oryza sativa. Coloring in heatmaps is proportional to log2 (TPM) value that represents the gene expression level. The corresponding log2 (TPM) value is displayed next to the color key. The rows represent gene models and the columns show the experiments, collected from publicly available Sequence Read Archive (SRA) database. SRA accessions are annotated to relevant stress conditions (descriptions are presented in Supplementary Data 4). Solid lines connect putative orthologs based on evidence from phylogenetic and synteny analyses; dashed lines connect putative orthologs based on evidence from either phylogenetic or synteny analyses. e Gene families evolving under dosage balance are predicted to demonstrate a high phenotypic effect in their knockouts. This can be seen by reanalysis of phenotype data from Bourdais et al.24; the bCRK T-DNA insertion mutants display a larger standard deviation (Y-axis) over different phenotyping experiments than vCRK mutants
Within the rapidly diverging β-group the vCRKs show large lineage-specific expansions. The ancestral origins for monocot and eudicot vCRKs differ, and neither synteny nor orthology can be identified (Fig. 7c, Supplementary Figure 14c). This suggests that this subfamily has a high birthrate and expands rapidly by tandem duplications. Additionally, many tandems are lost or fractionated after WGMs. Similarly, CRRSPs demonstrate little synteny between species (Fig. 7a, Supplementary Figure 14d), and CRRSPs in rice and Arabidopsis experienced lineage-specific tandem duplications (Supplementary Figure 14d). In Brassicaceae this expansion can be traced to Amborella CRRSP (AtrCRRSP2), suggesting a tandem mode of expansion.
Tandem duplications evolve through unequal crossover or homologous recombination events53. Unequal crossover produces copy number variation, whereas homologous recombination such as gene conversion plays a role in concerted evolution, which can maintain the similarity between gene copies over long periods54. Gene conversion depends on genomic distance as well as sequence homology. Accordingly, we observed several events among the lineage-specific tandem vCRK expansions (Supplementary Data 3), whereas for bCRKs gene conversion was only observed in the tandem expansion in Amborella. Thus, gene conversion is important for maintaining the similarity between recent tandem duplicates but conversion events become rare as sequences diverge over time.
The CRCK-I genes are present in most genomes as single copy genes within conserved syntenic genome segments, suggesting that duplicates from WGMs were lost during genome fractionation (Fig. 7a). Evolution follows a model where maintenance of a single copy is critical for the organism. A hallmark of gene families evolving under dosage balance is that their overall numbers should be conserved between species with similar WGM histories. In the species tree (Fig. 7a), most branches experienced one or two WGMs. Despite these events, the number of bCRKs is well conserved in angiosperms (Figs. 3b and 7b, Supplementary Figures 3b and 7b). However, in Amborella trichopoda five bCRK genes appear in tandem which form the roots of the respective orthologs (Supplementary Figure 6b), indicating an ancestral SSD origin still present in Amborella. The duplicate region experienced fractionation during evolution leading to Brassicaceae and Solanaceae lineages, resulting in scattered bCRK-I orthologs with little conserved synteny, whereas in grasses the tandem duplicate region was lost. This indicates rapid pseudogenization of the duplicated tandem blocks after WGMs, with, except for Solanaceae, no recent tandem expansions. This suggests that a gene family initially existing as a tandem duplicate may have shifted towards dosage balance evolution. Dosage balance is observed in another subfamily, PDLPs; they appear in genomic regions where synteny is conserved within eudicots and monocots (Fig. 7a), and no SSD events can be detected.
A prediction for gene families evolving under dosage balance is that retained duplicates should exhibit less functional divergence than other duplicates3. We explored functional conservation by analyzing publicly available gene expression data on stress treatments (Fig. 7d, Supplementary Data 4, Supplementary Figure 15). In agreement with previous studies24,26,31,55, pathogen treatments have the biggest impact on transcript abundance of DUF26-containing genes, in particular CRKs and CRRSPs (Supplementary Figure 15). Analysis of gene expression data suggest extensive lineage-specific functional diversification. This is visible in the correlation rank between putative orthologs; in many cases higher correlation is found with DUF26-containing genes that have less sequence similarity, indicating that closely related genes experienced sub- or neofunctionalization following duplication56,57.
Despite rearrangements and lineage-specific expansions the data provide support for seven putative orthologs, including three PDLP and three CRRSP relationships (Fig. 7d; Supplementary Data 4). Even though the synteny of bCRKs (and PDLPs) is more conserved compared to CRRSPs, bCRKs demonstrate varying responses to stimuli, whereas in CRRSPs synteny is associated with similar functions.
The second prediction from the dosage balance model is that, since the proteins encoded by the genes are highly connected and thus interact with many other proteins, disturbances in dosage balance should have large effects on an organism’s phenotype58. Reanalysis of phenotyping data crk mutants24 confirms that bCRKs indeed demonstrate a larger variance in phenotypes than vCRKs (p = 0.03; Wilcox test; Fig. 7e). Altogether the analysis suggests that PDLPs and bCRKs are evolving according to the dosage balance model, whereas the vCRKs and CRRSPs evolve by SSD mechanisms.
Discussion
Compared to animal genomes, plant genomes encode a large number of large gene families59. In particular, signal transduction components including transcription factors, protein kinases, and phosphatases have experienced drastic expansions in plants59. This might reflect adaptation to a sessile lifestyle but also different signaling strategies on the cellular level. The large, in part lineage-specific, expansions and domain rearrangements hamper the identification of orthologous proteins in different plant species. Here we used the DUF26-containing proteins to study the evolution of a large plant protein family with heterogeneous domain architecture and drastic lineage-specific expansions. We identified 1409 high-quality gene models representing CRRSPs, CRKs, and PDLPs from major plant lineages. The sdCRRSPs are the ancestral type of DUF26-containing proteins. CRKs originate from a fusion of CRRSPs with transmembrane region and kinase domain of LRR_clade_3 RLKs15 in the lineage leading to lycophytes. PDLPs and ddCRRSPs emerged subsequently through loss of kinase and transmembrane domains. Our results reveal an ancient split into two distinct groups. The α-group is conserved in size and sequence throughout embryophytes. This facilitates identification of orthologs and extrapolation of functional information from model plants to crops. The β-group evolved before the split of monocots and eudicots and contains CRKs and CRRSPs that expanded through WGMs followed by lineage-specific tandem duplications. Domain rearrangements in the β-clade led to secondary groups of ddCRRSPs and sdCRRSPs while the recruitment of a different kinase domain in grasses suggests re-routing of signaling pathways. Domain exchanges in DUF26-containing proteins highlight the importance of comparing phylogenies inferred from full-length sequences with phylogenies inferred from individual domains. It is likely that members of the β-group have been subject to sub- and neofunctionalization, which is a challenge for functional analyses. WGMs have been associated with periods of environmental upheaval and increasing biological complexity2,60. Accordingly, the evolution and radiation of DUF26-containing proteins with different domain structures co-occur with the appearance of new physiological characteristics and the adaptation to new habitats and lifestyles (Fig. 1b).
Sequence analysis suggested that DUF26 proteins could be specific to embryophytes. Crystallographic analysis of PDLP ectodomains reveals that the structure of their DUF26 domains closely matches the fold of the evolutionarily distant sdCRRSP Gnk-2. PDLPs contain two DUF26 domains and the structure of Gnk-2 is more similar to DUF26-A but despite the structural similarity the mannose-binding function of Gnk-2 is not conserved. Intriguingly, plant DUF26 domains share strong structural similarity with fungal carbohydrate-binding modules but the tandem arrangement of two lectin-like DUF26 domains appears to be plant-specific. Rapid sequence divergence61 is a limiting factor in detection of homology at the amino acid sequence level, seen e.g. in the marked differences between DUF26 from bryophytes and those from other plants. This may obscure identification of ancestral proto-DUF26 domains in charophytes and algae. Our work suggests that different tandem DUF26 domains likely recognize diverse sets of ligands which still remain to be discovered. Similar to plant malectin receptors62, DUF26 domains may have evolved novel or additional functions which might include mediation of protein–protein interactions at the cell surface20,35. The strong structural similarity between DUF26 domains and fungal lectins suggests either a common origin or convergent evolution. DUF26 proteins represent carbohydrate-binding domains in plants and identification of ligands for different DUF26 domains will provide insights into perception of cell wall status or environmental signals. However, the large number of carbohydrates and related compounds in the plant cell wall may pose a challenge for this process.
From the analyses, an overall evolutionary model emerges (Fig. 8). After introduction to the genome, gene families may initially expand through tandem duplications and experience relaxed selection63. This is supported by the fact that the tandem genes function in processes requiring fast adaptation such as adaptation to environment, pathogen responses, and secondary metabolism2,64. Furthermore, tandem duplicates show high variation across species and have high dN/dS rates5. In tandems, the main evolutionary forces are unequal crossover and concerted evolution through gene conversion, but over time genes evolve to gain specific functions. This process may be interrupted by WGM events. Since the tandem genes are not evolving under dosage balance, there is no compensatory drift65. Thus, drift and selection by dosage eventually drives some duplicates towards fixation while others turn into pseudogenes. Assuming that the elements driving tandem duplications are still present after fractionation, the remaining duplicates may in turn expand. In case of a tandem where all genes have established a unique functional role in the system, drift may drive duplicated tandems into scattered orthologs. These orthologs may assume a fixed syntenic position in the genome and a switch to a dosage balance mode of evolution results. The evolutionary mode of gene families depends on the balance between death rate after WGMs and birthrate of tandem duplications.
Model of mixed-type gene family evolution. Gene families evolve mainly through whole-genome multiplications (WGM) and small-scale duplications (SSD). Genes related to environmental responses and secondary metabolism experience SSDs, whereas highly connected genes associated with transcriptional and developmental regulation or signal transduction functions are preferentially retained after WGMs. a Prevailing hypothesis for the retention pattern is dosage balance; for highly connected genes the stoichiometric balance needs to be maintained, and therefore selection acts against gene losses after WGMs and against duplications by SSDs. b Gene family evolving through tandem duplications (b; evolution before the speciation node) has a high birthrate and therefore the number of duplicates between species can vary. After duplications the homogeneity of the duplicates may be maintained through gene conversion events occurring mainly within near-by homologous sequences. Over time, sequences eventually diverge by drift and selection. Our data suggest that tandemly expanding gene families may evolve into a dosage balance mode as a result of WGMs (b; evolution after speciation node). Following WGMs, duplicated tandems may experience extensive fractionation due to drift and selection by dosage fragmenting the tandem structure. At the same time, the connectivity of the gene family has been accumulating through sub- and neofunctionalization, increasing pressure for retention of genes. These phenomena together may result into a dosage balance model of evolution (top branch after speciation node). This does not necessarily occur across all WGM events and depends on the tandem duplication rate, as was observed for bCRKs in Solanaceae (bottom branch), where single copies and a later tandem expansion exist in the genome. Different subfamilies can be in different states of this process. c CRRSPs and PDLPs follow dosage balance mode after the paleohexaploid event, whereas bCRKs have assumed the mode in later WGM events. The overall numbers of the bCRKs are preserved but identification of orthologs between species that have experienced independent WGMs is difficult, suggesting recent convergent functionality of the members. Gene families expanding through tandem duplications such as vCRKs and CRRSPs have high birthrate and demonstrate several lineage-specific expansions. Asterisk indicates loss of sdCRRSPs in Brassicaceae and rice
Our study of DUF26-containing proteins demonstrates the challenges in analyses of large protein families and the power of combining evolutionary and structural approaches. Our analysis provides a model for future studies of similarly large protein families and facilitates detailed biochemical and physiological investigation of the mechanistic functions of CRKs, PDLPs, and CRRSPs in different plant species.
Methods
Gene identification and annotation
Altogether 32 plant and algae genomes (Supplementary Data 1) covering the major plant lineages were selected for analyses. For 27 species protein annotations (primary transcripts) and genome sequence data were retrieved from Phytozome66 and Barley (Hordeum vulgare) from Gramene (http://www.gramene.org) with the latest names for gene models from IPK server (http://webblast.ipk-gatersleben.de/barley_ibsc/)67. Silver birch (Betula pendula) was sequenced at the University of Helsinki56. Eggplant (Solanum melongena) data were retrieved from Eggplant Genome DataBase (http://eggplant.kazusa.or.jp/). Klebsormidium flaccium and Sacred lotus (Nelumbo nucifera) genome data were from NCBI (https://www.ncbi.nlm.nih.gov). Additionally the FungiDB68 (www.fungidb.org), InsectBase69 (http://www.insect-genome.com) human (Homo sapiens), chicken (Gallus gallus), and zebrafish (Danio rerio) genomes were screened for DUF26. Detailed information of the genome versions and references are given in the Supplementary Table 2.
HMMER (version 3.1b2) search70 for PFAM domain with ID PF01657 (stress-antifungal domain) was carried out among AA sequences representing gene models from different species71. Genome sequences were checked with Wise2 (version 2.4.1) software72,73. All gene models found with HMMER were manually curated, and new genes found with Wise2 were manually annotated using Fgenesh+74. Birch (Betula pendula)56 and Sacred lotus (Nelumbo nucifera) were fully manually annotated as they did not have gene models a priori. High rates of manual annotation and curation were needed for Selaginella moellendorffii, grapevine (Vitis vinifera; version Genoscope.12X75) and potato (Solanum tuberosum). Sequences from each species were further checked by carrying out a multiple sequence alignment and phylogenetic tree estimation with PASTA76. Partial gene models were identified by checking sequences individually. Genes were defined as pseudogenes if the genomic sequence was available but no full domain structure could be predicted. In cases where the prediction problem was caused by the length of the contig or a gap in the genome sequence the gene model was marked as partial. Pseudogenes and partial gene models were not included in the subsequent analyses.
For domain analyses and phylogenetic trees containing only domain sequences, the domain borders were defined with HMMER using the PFAM domain PF01657 for DUF26 and PF07714 for the kinase domain from curated dataset. The ectodomain region was defined to end at the border of the transmembrane region in the PDLPs and CRKs. The partial PDLP from Marsilea quandrifolia was identified by using pBLAST search against sequences in the NCBI database.
Phylogenetic trees
Only full gene models were used to infer phylogenetic trees. Sequence quality in alignments was checked using Guidance (version 2.01) and alignments were built using the MAFFT option77. Sequences with low-quality score were removed from datasets and alignments were built again with PASTA. For phylogenetic trees, alignments were filtered in Wasabi78 to remove residues with less than 10% coverage. Filtering was required due to the high sequence diversity (on less conserved regions) resulting in a high number of gaps in multiple sequence alignments. Maximum likelihood (ML) phylogenetic trees were inferred for filtered and also unfiltered data using RAxML (version 8.1.3)79.
ML phylogenetic trees were bootstrapped using RAxML (version 8.1.3) for 1000 bootstrap replicates. For phylogenetic trees containing full-length sequences with all domain structures bootstrapping was also carried out with partitioning (both DUF26 and kinase domains defined separately). The PROTGAMMAJTT model was used in phylogenetic analyses using RAxML. Model selection was based on a Perl script for identifying the optimal protein substitution model (available in RAxML webpage, provided by Alexandros Stamatakis). Bootstrapped trees are available on Wasabi78 (Supplementary Table 3). Comparison on phylogenetic trees based on CRK ectodomain and kinase domain regions was visualized in R using the dendextend package.
Exon–intron structure
The number of exons for all genes was estimated using Scipio (version 1.4.1)80 using default parameters (minimum identity of 90% and coverage of 60%). It internally uses BLAT to perform the initial alignment of the protein sequences against the genome followed by refinement of hits to determine the exact splicing borders and to obtain the final gene structure. The number of exons per gene was extracted from the final result.
Orthogroup generation
Eleven representative species from different clades (Arabidopsis thaliana, Amborella trichopoda, Oryza sativa, Zea mays, Vitis vinifera, Populus trichocarpa, Aquilegia coerulea, Brachypodium distachyon, Physcomitrella patens, Selaginella moellendorffii, and Spirodela polyrhiza) were chosen to study the evolution of the DUF26-containing proteins. Primary protein sequences of these 11 species were downloaded from Phytozome (version 11.0). An all-against-all BLAST was run for all the protein sequences followed by generation of orthogroups using the software OrthoMCL (version 2.0.9)81 with an inflation parameter of 1.5 for the clustering phase. Clustering yielded 34,535 orthogroups.
Species tree generation
Orthogroups containing one representative protein for each of the 11 species were chosen to generate the species tree. Multiple sequence alignment was carried out on the single copy orthogroups using PRANK82 and the output was used to infer a species tree using RAxML79.
Evolutionary rate and ancestral size estimation
The evolutionary rate and ancestral size of the orthogroups were modeled using Badirate software (version 1.35)41. The species tree and orthogroups generated from the previous steps were used as input for Badirate. The BDI (Birth, Death, Innovation) rate model was used. The Free Rates (FR) branch model was chosen which would assume every branch of the species tree to have its own turnover rates. Turnover rates of orthogroups were estimated using the ML fitting. Orthogroups were defined as protein kinases if they included sequences with PFAM domain PF00069. Orthogroups containing RLKs were defined based on known Arabidopsis RLKs15. Plasmodesmata-related orthogroups were defined based on Arabidopsis thaliana genes related to plasmodesmata40.
Nucleotide coding sequence extraction for PAML
The GFF file output from Scipio80 was pre-processed by an in-house script and processed with the gff3 module of the GenomeTools (version 1.5.4)83 software. The final GFF file along with the corresponding species genome in fasta formatted file was passed as an input to the extractfeat module of the GenomeTools software to extract the final nucleotide coding sequences.
PAML analyses
We estimated dN/dS ratios (ratio of non-synonymous and synonymous sites, ω) for conserved clades (bCRK-I, bCRK-II, CRCKs (orthologs of AtCRK43), PDLPs, and sdCRRSPs) from 11 species (Arabidopsis thaliana, Amborella trichopoda, Oryza sativa, Zea Mays, Vitis vinifera, Populus trichocarpa, Aquilegia coerulea, Brachypodium distachyon, Physcomitrella patens, Selaginella moellendorffii, and Spirodela polyrhiza) by using the codeml program from PAML (version 4.9)84. We applied the one-ratio model (M0) to estimate overall dN/dS ratios for each conserved group separately and free ratios neutral model (M1) to estimate dN/dS ratios for each branch within conserved clades85. To study the evolution of PDLP5 and PDLP8, sitewise-analyses of their homologs was carried out. As PDLP5 is specific to Brassicaceae, we added nucleotide sequences for orthologs of AtPDLP5 from NCBI, Phytozome and CoGe databases. Furthermore, additional sequences for orthologs of AtPDLP8 were included in the alignment to improve depth and reliability of the analysis. Multiple sequence alignments of coding nucleotide sequences were constructed with PRANK82 and phylogenetic trees were estimated using RAxML79 for codeml.
Syntenic vs tandem duplications
Syntenic and tandem duplications were analyzed using Synmap application in CoGe86 using default settings. Tandem duplications were defined as genome regions with at least three to five duplicate genes (Supplementary Data 4). Synteny comparisons were done between Arabidopsis thaliana and Solanum lycopersicum, S. lycopersicum and Amborella trichopoda, A. trichopoda and Oryza sativa, and Zea mays and Oryza sativa. Tandem duplication results from DAGchainer were collected for each species. The results were filtered based on annotated gene models from selected species. The currently available Amborella trichopoda genome is presented only as scaffolds, and the genes were placed to chromosomes based on physical mapping87. Scaffolds not assigned to any chromosome were added separately. Thus the location of the Amborella trichopoda genes in the genome is only a rough estimate (Fig. 7a).
Gene conversion analyses
Gene conversion events were estimated from nucleotide sequences for the same 11 species that were analyzed for dN/dS ratios with GENECONV (version 1.81a)88. Analyses were carried out for the main clades of the 11 species. For bCRKs and vCRKs separate analyses were carried out using sequences from the five species used in synteny analyses (Arabidopsis thaliana, Amborella trichopoda, Oryza sativa, Solanum lycopersicum, and Zea mays). The largest tandem region of vCRKs in A. thaliana chromosome 4 was analyzed separately to validate the results from the analysis with all vCRKs from A. thaliana.
Gene tree reconciliation
Gene tree reconciliation was carried out using DLCpar (version 1.0)89 downloaded from https://www.cs.hmc.edu/~yjw/software/dlcpar/. NCBI taxonomy was used as the species tree, downloaded in newick format from PhyloT website, http://phylot.biobyte.de/. Reconciliation was carried out using DLCpar search with 20 prescreening iterations, followed by 1000 search iterations. The solution was visualized in R, using custom scripts and “ape” package.
Phenomics data analysis
Phenotyping data of T-DNA mutant insertion lines were normalized against the Col-0 data by calculating Z-scores, see Bourdais et al.24 The standard deviation (SD) over all experiments was calculated for each allele, and in case of several insertion alleles the one with maximum SD was selected. The residuals of the bCRK vs vCRK split in the data were tested for normality using Shapiro’s test. Since the null hypothesis (normality) was rejected with p < 0.05 the difference between groups was tested with Wilcox test.
Transcriptomic analyses
Paired end RNAseq data were collected from the publicly available sequence read archive (SRA) database by fastq-dump.2 (version 2.5.7) for Arabidopsis thaliana, Oryza sativa, Solanum lycopersicum, and Zea mays. FastQC (version 0.11.4) (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/) was used to check the quality of the samples. Low-quality reads and bases were removed by Trimmomatic (version 0.36)90 with the following options: phred33, TRAILING: 20, and MINLEN: 30. Filtered reads were mapped to gene models from Phytozome version 12, by Kallisto, run in paired end mode (version 0.43.1, --bias and --bootstrap: 200)91. Bootstrap samples were averaged (custom R code) and gene expression abundance (transcript per million [TPM]) was estimated by tximport (version 1.2.0)92 followed by averaging over biological replicates. Ortholog comparison between species was carried out by grouping the experiments into seven categories, with maximum TPM among experiments representing gene response. Pearson correlation was calculated among orthologs and all other possible pairs.
Protein expression and purification
An expression construct coding for the PDLP5 ectodomain (amino acids 1–241) was codon optimized for Spodoptera frugiperda and synthesized by Geneart (Thermo Fisher). Using the PfuX7 polymerase93, the gene for the PDLP8-ECD (1–253) was amplified from Arabidopsis thaliana cDNA. The Gibson assembly method94 was employed to insert the PDLP5 and PDLP8 ectodomain coding sequences into an adapted pFAST-BAC1 vector (Geneva Biotech), providing a C-terminal 2x-STREP-9xHIS tag. PDLP5 point mutations (C101A, C148A, and C191A) were then introduced as described95. Bacmids were generated by transforming the plasmids (confirmed by sequencing) into Escherichia coli DH10MultiBac (Geneva Biotech). Virus particles were created by transfecting (Profectin, AB Vector) the bacmids into Spodoptera frugiperda SF9 cells (Thermo Fisher). For secreted protein production, Trichoplusia ni Tnao38 cells (obtained from Gary W Blissard, Boyce Thompson Institute, Tower Road, Ithaca, NY, USA)96 were infected with a viral multiplicity of 1, incubated for 3 days at 22 °C. The protein-containing supernatant was separated from the intact cells by centrifugation and subjected to Ni2+-affinity chromatography (HisTrap Excel; GE Healthcare) in buffer A (10 mM Hepes 7.5, 500 mM NaCl). Bound proteins eluted in buffer A supplemented with 500 mM imidazole. The elution fractions were pooled and further purified by StrepII-affinity purification (Strep-Tactin XT Superflow high capacity, IBA) in buffer B (20 mM Tris pH 8.0, 250 mM NaCl, 1 mM EDTA). The column was washed with 5–10 column volumes of buffer B and eluted in buffer B supplemented with 50 mM biotin. The C-terminal 2x-STREP-9xHIS tag was subsequently removed by adding tobacco etch virus (TEV)-protease to the StrepII elution in a 1:100 ratio for 16 h at 4 °C. The 2x-STREP-9xHIS-tag and the HIS-tagged TEV-protease were then separated from the respective ectodomain by an additional Ni2+-affinity chromatography step (HisTrap Excel; GE Healthcare). Cleaved PDLP5, PDLP5C101A, PDLP5C148A, PDLP5C191A, and PDLP8 ectodomains were next subjected to preparative size exclusion chromatography using either a HiLoad 26/600 Superdex 200 pg (PDLP5 and PDLP8) or HiLoad 16/600 Superdex 200 pg (PDLP5C101A, PDLP5C148A, and PDLP5C191A) column, equilibrated in 20 mM sodium citrate pH 5.0 and 150 mM NaCl. Monomeric peak fractions were collected and concentrated using an Amicon Ultra (Millipore) filter device. The concentrated monomeric peak fractions of PDLP5, PDLP5C101A, PDLP5C148A, and PDLP5C191A were additionally subjected to analytical size exclusion chromatography on a Superdex 200 Increase 10/300 GL column (GE Healthcare) equilibrated in 20 mM citrate pH 5.0 and 150 mM NaCl (Supplementary Figure 9, uncropped gel images are available in Supplementary Figure 16).
Thermostability assay
Twenty microliters reactions consisted of either PDLP5, PDLP5C101A, PDLP5C148A, and PDLP5C191A ectodomains at a concentration of 1.5 mg/ml in 20 mM citrate pH 5.0, 150 mM NaCl, 10× SYPRO Orange dye (Thermo Fisher), and were mixed in a 384-well ABI PRISM plate (Applied Biosystems). Using a 7900HT Fast Real-Time PCR system SYPRO Orange fluorescence was measured. The reactions were initially incubated for 2 min at 25 °C and then the temperature was increased to 95 °C at a heating rate of 0.5 °C/min. Resulting melting curves were fitted with a Boltzman function using GraphPad Prism and the melting temperatures, Tm, correspond to the first inflection point of the Boltzman fit.
Isothermal titration calorimetry
ITC experiments were performed at 25 °C using a Nano ITC (TA Instruments, New Castle, USA) with a 1.0 ml standard cell and a 250 μl titration syringe. The PDLP5 ectodomain was gelfiltrated into ITC buffer (20 mM sodium citrate pH 5.0, 150 mM NaCl) and all carbohydrates were resuspended into ITC buffer. The experiments were carried out by injecting 24 times 10 μl of d-+-Mannose (1 mM; Sigma), Pectic Galactan (2 mg/ml; Megazyme), Rhamnogalacturonan (2 mg/ml; Megazyme), polygalacturonic acid (2 mg/ml; Megazyme), Cellohexaose (1 mM; Megazyme), or Arabinohexaose (1 mM; Megazyme) aliquots into PDLP5 (~100 μM) in the cell at 150 s intervals. ITC data for the d-+-mannose experiment were corrected for the heat of dilution by subtracting the mixing enthalpies for titrant solution injections into protein-free ITC buffer. Data were analyzed using the NanoAnalyze program (version 3.5) as provided by the manufacturer.
Protein crystallization and crystallographic data collection
The PDLP5 ectodomain formed crystals in hanging drops composed of 1 μl of protein solution (70 mg/ml in 20 mM citrate pH 5.0 and 150 mM NaCl) and 1 μl of crystallization buffer (17.5% [w/v] polyethylene glycol 4000, 250 mM (NH4)2SO4) suspended over 800 μl of the latter as reservoir solution. Protein crystals were transferred into crystallization buffer supplemented with 25% (v/v) ethylene glycol, which served as cryoprotectant, and snap frozen in liquid N2. PDLP8 crystals (52 mg/ml in 20 mM citrate pH 5.0, 150 mM NaCl) developed in hanging drops containing 17.5 % (w/v) polyethylene glycol 4000, 0.1 M citrate pH 5.5, 20% (v/v) 2-propanol. Crystals were frozen directly in liquid N2. For PDLP5 native (λ = 1.0 Å) and redundant sulfur SAD (λ = 2.079 Å) data were collected to 1.29 Å resolution at beam line PXIII of the Swiss Light Source (SLS), Villigen, Switzerland. A 1.95 Å native dataset of PDLP8 was acquired at the same beam line. Data processing and reduction was done with XDS (version: Jan 2018)97.
Structure solution and refinement
The structure of PDLP5 was solved using the single-anomalous diffraction (SAD) method. Twenty-four S sites corresponding to the 12 disulfide bonds in the PDLP5 crystallographic dimer were located with the program SHELXD98, site-refinement and phasing was done in SHARP99, and the starting phases were used for automated model building in BUCCANEER100 and ARP/wARP101. The model was completed in alternating cycles of model correction in COOT102 and restrained refinement in Refmac5103. The structure of PDLP8 was solved using the molecular replacement methods as implemented in the program PHASER104, and using the refined PDLP5 tandem ectodomain as search model. Inspection with MolProbity105 revealed excellent stereochemistry for the final models. Structural and surface representations were done in Pymol (http://pymol.soureforge.org) and Chimera106.
Code availability
Scripts for parsing and visualizing data have been deposited in GitHub and can be retrieved from https://github.com/jsalojar/DLCpar_visualize/.
Reporting Summary
Further information on experimental design is available in the Nature Research Reporting Summary linked to this Article.
Data availability
Sequence information used in this study is available as Supplementary Data 5. Phylogenetic trees with bootstrap information for 1000 replicates and corresponding sequence alignments have been deposited on Wasabi (http://wasabiapp.org); identifiers are available in the figure legends as web links. Information on used genomic data is available in Supplementary Table 2. Publicly available gene expression data were taken from the Sequence Read Archive (SRA) database; identifiers are listed in Supplementary Data 4. Crystallographic coordinates and structure factors have been deposited with the Protein Data Bank (http://rcsb.org) with accession codes 6GRE (PDLP5) and 6GRF (PDLP8). Refinement statistics are available as Supplementary Table 1. Other materials are available from the corresponding author upon request.
References
Demuth, J. P. & Hahn, M. W. The life and death of gene families. Bioessays 31, 29–39 (2009).
Panchy, N., Lehti-Shiu, M. & Shiu, S. H. Evolution of gene duplication in plants. Plant Physiol. 171, 2294–2316 (2016).
Tasdighian, S. et al. Reciprocally retained genes in the angiosperm lineage show the hallmarks of dosage balance sensitivity. Plant Cell 29, 2766–2785 (2017).
Birchler, J. A., Bhadra, U., Bhadra, M. P. & Auger, D. L. Dosage-dependent gene regulation in multicellular eukaryotes: implications for dosage compensation, aneuploid syndromes, and quantitative traits. Dev. Biol. 234, 275–288 (2001).
Wang, P. et al. Factors influencing gene family size variation among related species in a plant family, Solanaceae. Genome Biol. Evol. 10, 2596–2613 (2018).
Nakamura, S., Suzuki, T., Kawamukai, M. & Nakagawa, T. Expression analysis of Arabidopsis thaliana small secreted protein genes. Biosci. Biotechnol. Biochem. 76, 436–446 (2012).
Agrawal, G. K., Jwa, N. S., Lebrun, M. H., Job, D. & Rakwal, R. Plant secretome: unlocking secrets of the secreted proteins. Proteomics 10, 799–827 (2010).
Tavormina, P., De Coninck, B., Nikonorova, N., De Smet, I. & Cammue, B. P. The plant peptidome: an expanding repertoire of structural features and biological functions. Plant Cell 27, 2095–2118 (2015).
Shiu, S. H. & Bleecker, A. B. Receptor-like kinases from Arabidopsis form a monophyletic gene family related to animal receptor kinases. Proc. Natl. Acad. Sci. USA 98, 10763–10768 (2001).
Shiu, S. H. & Bleecker, A. B. Expansion of the receptor-like kinase/Pelle gene family and receptor-like proteins in Arabidopsis. Plant Physiol. 132, 530–543 (2003).
Fritz-Laylin, L. K., Krishnamurthy, N., Tor, M., Sjolander, K. V. & Jones, J. D. Phylogenomic analysis of the receptor-like proteins of rice and Arabidopsis. Plant Physiol. 138, 611–623 (2005).
Smakowska-Luzan, E. et al. An extracellular network of Arabidopsis leucine-rich repeat receptor kinases. Nature 553, 342–346 (2018).
Shiu, S. H. & Bleecker, A. B. Plant receptor-like kinase gene family: diversity, function, and signaling. Sci. STKE 2001, re22 (2001).
Shiu, S. H. et al. Comparative analysis of the receptor-like kinase family in Arabidopsis and rice. Plant Cell 16, 1220–1234 (2004).
Zulawski, M., Schulze, G., Braginets, R., Hartmann, S. & Schulze, W. X. The Arabidopsis Kinome: phylogeny and evolutionary insights into functional diversification. BMC Genomics 15, 548 (2014).
Fischer, I., Dievart, A., Droc, G., Dufayard, J. F. & Chantret, N. Evolutionary dynamics of the leucine-rich repeat receptor-like kinase (LRR-RLK) subfamily in angiosperms. Plant Physiol. 170, 1595–1610 (2016).
Kimura, S., Waszczak, C., Hunter, K. & Wrzaczek, M. Bound by fate: the role of reactive oxygen species in receptor-like kinase signaling. Plant Cell 29, 638–654 (2017).
Miyakawa, T. et al. A secreted protein with plant-specific cysteine-rich motif functions as a mannose-binding lectin that exhibits antifungal activity. Plant Physiol. 166, 766–778 (2014).
Miyakawa, T., Miyazono, K., Sawano, Y., Hatano, K. & Tanokura, M. Crystal structure of ginkbilobin-2 with homology to the extracellular domain of plant cysteine-rich receptor-like kinases. Proteins 77, 247–251 (2009).
Ma, L. S. et al. The Ustilago maydis repetitive effector Rsp3 blocks the antifungal activity of mannose-binding maize proteins. Nat. Commun. 9, 1711 (2018).
Acharya, B. R. et al. Overexpression of CRK13, an Arabidopsis cysteine-rich receptor-like kinase, results in enhanced resistance to Pseudomonas syringae. Plant J. 50, 488–499 (2007).
Chen, K., Du, L. & Chen, Z. Sensitization of defense responses and activation of programmed cell death by a pathogen-induced receptor-like protein kinase in Arabidopsis. Plant Mol. Biol. 53, 61–74 (2003).
Chen, K., Fan, B., Du, L. & Chen, Z. Activation of hypersensitive cell death by pathogen-induced receptor-like protein kinases from Arabidopsis. Plant Mol. Biol. 56, 271–283 (2004).
Bourdais, G. et al. Large-scale phenomics identifies primary and fine-tuning roles for CRKs in responses related to oxidative stress. PLoS Genet. 11, e1005373 (2015).
Idänheimo, N. et al. The Arabidopsis thaliana cysteine-rich receptor-like kinases CRK6 and CRK7 protect against apoplastic oxidative stress. Biochem. Biophys. Res. Commun. 445, 457–462 (2014).
Wrzaczek, M. et al. Transcriptional regulation of the CRK/DUF26 group of receptor-like protein kinases by ozone and plant hormones in Arabidopsis. BMC Plant Biol. 10, 95 (2010).
Yeh, Y. H., Chang, Y. H., Huang, P. Y., Huang, J. B. & Zimmerli, L. Enhanced Arabidopsis pattern-triggered immunity by overexpression of cysteine-rich receptor-like kinases. Front. Plant Sci. 6, 322 (2015).
Yadeta, K. A. et al. A cysteine-rich protein kinase associates with a membrane immune complex and the cysteine residues are required for cell death. Plant Physiol. 173, 771–787 (2017).
Lee, D. S. K., Young, Cheon, Kwon, SunJae, Ryu, Choong-Min & Park, OhkmaeK. The Arabidopsis cysteine-rich receptor-like kinase CRK36 regulates immnity through interaction with the cytoplasmic kinase BIK1. Front. Plant Sci. 8, 1856 (2017).
Tanaka, H. et al. Abiotic stress-inducible receptor-like kinases negatively control ABA signaling in Arabidopsis. Plant J. 70, 599–613 (2012).
Chern, M. et al. A genetic screen identifies a requirement for cysteine-rich-receptor-like kinases in Rice NH1 (OsNPR1)-mediated immunity. PLoS Genet. 12, e1006049 (2016).
Brunkard, J. O. & Zambryski, P. C. Plasmodesmata enable multicellularity: new insights into their evolution, biogenesis, and functions in development and immunity. Curr. Opin. Plant Biol. 35, 76–83 (2017).
Caillaud, M. C. et al. The plasmodesmal protein PDLP1 localises to haustoria-associated membranes during downy mildew infection and regulates callose deposition. PLoS Pathog. 10, e1004496 (2014).
Lim, G. H. et al. Plasmodesmata localizing proteins regulate transport and signaling during systemic acquired immunity in plants. Cell Host Microbe 19, 541–549 (2016).
Cui, W. & Lee, J. Y. Arabidopsis callose synthases CalS1/8 regulate plasmodesmal permeability during stress. Nat. Plants 2, 16034 (2016).
Amari, K. et al. A family of plasmodesmal proteins with receptor-like properties for plant viral movement proteins. PLoS Pathog. 6, e1001119 (2010).
Xu, G., Ma, H., Nei, M. & Kong, H. Evolution of F-box genes in plants: different modes of sequence divergence and their relationships with functional diversification. Proc. Natl. Acad. Sci. USA 106, 835–840 (2009).
Rody, H. V., Baute, G. J., Rieseberg, L. H. & Oliveira, L. O. Both mechanism and age of duplications contribute to biased gene retention patterns in plants. BMC Genomics 18, 46 (2017).
Finn, R. D. et al. The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 44, D279–D285 (2016).
Fernandez-Calvino, L. et al. Arabidopsis plasmodesmal proteome. PLoS ONE 6, e18880 (2011).
Librado, P., Vieira, F. G. & Rozas, J. BadiRate: estimating family turnover rates by likelihood-based methods. Bioinformatics 28, 279–281 (2012).
Van de Peer, Y., Maere, S. & Meyer, A. The evolutionary significance of ancient genome duplications. Nat. Rev. Genet. 10, 725–732 (2009).
Hsu, T. C. et al. Early genes responsive to abscisic acid during heterophyllous induction in Marsilea quadrifolia. Plant Mol. Biol. 47, 703–715 (2001).
Holm, L. & Rosenstrom, P. Dali server: conservation mapping in 3D. Nucleic Acids Res. 38, W545–W549 (2010).
van Eerde, A., Grahn, E. M., Winter, H. C., Goldstein, I. J. & Krengel, U. Atomic-resolution structure of the alpha-galactosyl binding Lyophyllum decastes lectin reveals a new protein family found in both fungi and plants. Glycobiology 25, 492–501 (2015).
Zhang, P. et al. Cytotoxic protein from the mushroom Coprinus comatus possesses a unique mode for glycan binding and specificity. Proc. Natl. Acad. Sci. USA 114, 8980–8985 (2017).
Vijayan, M. & Chandra, N. Lectins. Curr. Opin. Struct. Biol. 9, 707–714 (1999).
Hohmann, U., Lau, K. & Hothorn, M. The structural basis of ligand perception and signal activation by receptor kinases. Annu. Rev. Plant Biol. 68, 109–137 (2017).
Berrabah, F. et al. A nonRD receptor-like kinase prevents nodule early senescence and defense-like reactions during symbiosis. New Phytol. 203, 1305–1314 (2014).
Dardick, C. & Ronald, P. Plant and animal pathogen recognition receptors signal through non-RD kinases. PLoS Pathog. 2, e2 (2006).
Dardick, C., Schwessinger, B. & Ronald, P. Non-arginine-aspartate (non-RD) kinases are associated with innate immune receptors that recognize conserved microbial signatures. Curr. Opin. Plant Biol. 15, 358–366 (2012).
Rogers, R. L., Shao, L. & Thornton, K. R. Tandem duplications lead to novel expression patterns through exon shuffling in Drosophila yakuba. PLoS Genet. 13, e1006795 (2017).
Hastings, P. J., Lupski, J. R., Rosenberg, S. M. & Ira, G. Mechanisms of change in gene copy number. Nat. Rev. Genet. 10, 551–564 (2009).
Chen, J. M., Cooper, D. N., Chuzhanova, N., Ferec, C. & Patrinos, G. P. Gene conversion: mechanisms, evolution and human disease. Nat. Rev. Genet. 8, 762–775 (2007).
Zou, C., Lehti-Shiu, M. D., Thomashow, M. & Shiu, S. H. Evolution of stress-regulated gene expression in duplicate genes of Arabidopsis thaliana. PLoS Genet. 5, e1000581 (2009).
Salojärvi, J. et al. Genome sequencing and population genomic analyses provide insights into the adaptive landscape of silver birch. Nat. Genet. 49, 904–912 (2017).
Fischer, I. et al. Impact of recurrent gene duplication on adaptation of plant genomes. BMC Plant Biol. 14, 151 (2014).
Veitia, R. A., Bottani, S. & Birchler, J. A. Cellular reactions to gene dosage imbalance: genomic, transcriptomic and proteomic effects. Trends Genet. 24, 390–397 (2008).
Guo, Y. L. Gene family evolution in green plants with emphasis on the origination and evolution of Arabidopsis thaliana genes. Plant J. 73, 941–951 (2013).
Van de Peer, Y., Mizrachi, E. & Marchal, K. The evolutionary significance of polyploidy. Nat. Rev. Genet. 18, 411–424 (2017).
Copley, R. R., Goodstadt, L. & Ponting, C. Eukaryotic domain evolution inferred from genome comparisons. Curr. Opin. Genet. Dev. 13, 623–628 (2003).
Franck, C. M., Westermann, J. & Boisson-Dernier, A. Plant malectin-like receptor kinases: from cell wall integrity to immunity and beyond. Annu. Rev. Plant Biol. 69, 301–328 (2018).
Innan, H. & Kondrashov, F. The evolution of gene duplications: classifying and distinguishing between models. Nat. Rev. Genet. 11, 97–108 (2010).
Moghe, G. D. & Last, R. L. Something old, something new: conserved enzymes and the evolution of novelty in plant specialized metabolism. Plant Physiol. 169, 1512–1523 (2015).
Thompson, A., Zakon, H. H. & Kirkpatrick, M. Compensatory drift and the evolutionary dynamics of dosage-sensitive duplicate genes. Genetics 202, 765–774 (2016).
Goodstein, D. M. et al. Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res. 40, D1178–D1186 (2012).
Deng, W., Nickle, D. C., Learn, G. H., Maust, B. & Mullins, J. I. ViroBLAST: a stand-alone BLAST web server for flexible queries of multiple databases and user’s datasets. Bioinformatics 23, 2334–2336 (2007).
Stajich, J. E. et al. FungiDB: an integrated functional genomics database for fungi. Nucleic Acids Res. 40, D675–D681 (2012).
Yin, C. et al. InsectBase: a resource for insect genomes and transcriptomes. Nucleic Acids Res. 44, D801–D807 (2016).
Eddy, S. R. Profile hidden Markov models. Bioinformatics 14, 755–763 (1998).
Vaattovaara, A., Salojärvi, J. & Wrzaczek, M. Extraction and curation of gene models for plant receptor kinases for phylogenetic analysis. Methods Mol. Biol. 1621, 79–91 (2017).
Birney, E., Clamp, M. & Durbin, R. Genewise and genomewise. Genome Res. 14, 988–995 (2004).
Birney, E. & Durbin, R. Using genewise in the Drosophila annotation experiment. Genome Res. 10, 547–548 (2000).
Solovyev, V. in Handbook of Statistical Genetics (eds Balding, D. J., Bishop, M. & Cannings, C.) 97–159 (John Wiley & Sons, Ltd, New York, 2008).
Jaillon, O. et al. The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449, 463–467 (2007).
Mirarab, S. et al. PASTA: Ultra-large multiple sequence alignment for nucleotide and amino-acid sequences. J. Comput. Biol. 22, (377–386 (2015).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Veidenberg, A., Medlar, A. & Löytynoja, A. Wasabi: an integrated platform for evolutionary sequence analysis and data visualization. Mol. Biol. Evol. 33, 1126–1130 (2016).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Keller, O., Odronitz, F., Stanke, M., Kollmar, M. & Waack, S. Scipio: using protein sequences to determine the precise exon/intron structures of genes and their orthologs in closely related species. BMC Bioinformatics 9, 278 (2008).
Li, L., Stoeckert, C. J. Jr & Roos, D. S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 13, 2178–2189 (2003).
Löytynoja, A. & Goldman, N. An algorithm for progressive multiple alignment of sequences with insertions. Proc. Natl. Acad. Sci. USA 102, 10557–10562 (2005).
Gremme, G., Steinbiss, S. & Kurtz, S. GenomeTools: a comprehensive software library for efficient processing of structured genome annotations. IEEE/ACM Trans. Comput. Biol. Bioinformatics 10, 645–656 (2013).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Yang, Z. N. R. Synonymous and nonsynomous rate varaition in nuclear genes of mammals. J. Mol. Evol. 46, 409–418 (1998).
Lyons, E., Pedersen, B., Kane, J. & Freeling, M. The value of nonmodel genomes and an example using SynMap within CoGe to dissect the hexaploidy that predates the rosids. Trop. Plant Biol. 1, 181–190 (2008).
Chamala, S. et al. Assembly and validation of the genome of the nonmodel basal angiosperm Amborella. Science 342, 1516–1517 (2013).
Sawyer, S. Statistical tests for detecting gene conversion. Mol. Biol. Evol. 6, 526–538 (1989).
Wu, Y. An algorithm for constructing parsimonious hybridization networks with multiple phylogenetic trees. J. Comput. Biol. 20, 792–804 (2013).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Bray, N. L., Pimentel, H., Melsted, P. & Pachter, L. Near-optimal probabilistic RNA-seq quantification. Nat. Biotechnol. 34, 525–527 (2016).
Soneson, C., Love, M. I. & Robinson, M. D. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Res. 4, 1521 (2015).
Norholm, M. H. A mutant Pfu DNA polymerase designed for advanced uracil-excision DNA engineering. BMC Biotechnol. 10, 21 (2010).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Liu, H. & Naismith, J. H. An efficient one-step site-directed deletion, insertion, single and multiple-site plasmid mutagenesis protocol. BMC Biotechnol. 8, 91 (2008).
Hashimoto, Y., Zhang, S., Chen, Y.-R. & Blissard, G. Erratum to: BTI-Tnao38, a new cell line derived from Trichoplusia ni, is permissive for AcMNPV infection and produces high levels of recombinant proteins. BMC Biotechnol. 12, https://doi.org/10.1186/1472-6750-12-12 (2012).
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993).
Sheldrick, G. M. A short history of SHELX. Acta Crystallogr. A. 64, 112–122 (2008).
Bricogne, G., Vonrhein, C., Flensburg, C., Schiltz, M. & Paciorek, W. Generation, representation and flow of phase information in structure determination: recent developments in and around SHARP 2.0. Acta Crystallogr. D Biol. Crystallogr. 59, 2023–2030 (2003).
Cowtan, K. The Buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr. D Biol. Crystallogr. 62, 1002–1011 (2006).
Cohen, S. X. et al. ARP/wARP and molecular replacement: the next generation. Acta Crystallogr. D Biol. Crystallogr. 64, 49–60 (2008).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Acknowledgements
The authors would like to thank Ms. Kerri Hunter, Drs. Julia Krasensky-Wrzaczek, Sachie Kimura, and Alexey Shapiguzov for critical comments on the manuscript. We thank Prof. David Collinge and Prof. Michael Lyngkjær (University of Copenhagen, Denmark) for help with the barley genome and Prof. Eric Schranz (Wageningen University, Netherlands) for helpful comments. This work was supported by the Doctoral Programme in Plant Sciences (DPPS) of the University of Helsinki and by the Finnish Cultural Foundation Suomen Kulttuurirahasto (to A.Va.), the Academy of Finland (grant numbers #275632, #283139, and #312498 to M.W.). and the University of Helsinki (three-year fund allocation to M.W.) and by grant 31003A_176237 from the Swiss National Science Foundation and by an International Research Scholar Award from the Howard Hughes Medical Institute (to M.H.). A.V., O.S., S.R., J.K., J.S., and M.W. are members of the Centre of Excellence in the Molecular Biology of Primary Producers (2014–2019) funded by the Academy of Finland (grant numbers #271832 and #307335). B.B. was supported by an EMBO long-term post-doctoral fellowship. We would also like to thank the staff at beam line PXIII of the Swiss Light Source (Villigen, Switzerland) for technical assistance during data collection.
Author information
Authors and Affiliations
Contributions
A.Va., B.B., J.K., J.S., M.H., and M.W. conceived and designed the project. A.Va., B.B., O.S., S.R., M.L., AVe, A.L., M.H., and J.S. carried out the analyses. A.Va., B.B., A.L., M.H., J.S., and M.W. analyzed the data. A.Va., B.B., M.H., J.S., and M.W. wrote the manuscript. All authors read and contributed to the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vaattovaara, A., Brandt, B., Rajaraman, S. et al. Mechanistic insights into the evolution of DUF26-containing proteins in land plants. Commun Biol 2, 56 (2019). https://doi.org/10.1038/s42003-019-0306-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-019-0306-9
This article is cited by
-
Overexpression of Ginkbilobin-2 homologous domain gene improves tolerance to Phytophthora cinnamomi in somatic embryos of Quercus suber
Scientific Reports (2024)
-
Evaluation of some biochemical and biomolecular indicators in rice (Oryza sativa L.) during the seedling stage under NaCl stress
Journal of Crop Science and Biotechnology (2024)
-
Plasmodesmata dynamics in bryophyte model organisms: secondary formation and developmental modifications of structure and function
Planta (2024)
-
Whole-Genome Resequencing Deciphers New Insight Into Genetic Diversity and Signatures of Resistance in Cultivated Cotton Gossypium hirsutum
Molecular Biotechnology (2023)
-
Genome wide study of cysteine rich receptor like proteins in Gossypium sp.
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.