Abstract
Kv7.1 (KCNQ1) coassembles with KCNE1 to generate the cardiac IKs-channel. Gain- and loss-of-function mutations in KCNQ1 are associated with cardiac arrhthymias, highlighting the importance of modulating IKs activity for cardiac function. Here, we report proteolysis of Kv7.1 as an irreversible posttranslational modification. The identification of two C-terminal fragments of Kv7.1 led us to identify an aspartate critical for the generation of one of the fragments and caspases as responsible for mediating proteolysis. Activating caspases reduces Kv7.1/KCNE1 currents, which is abrogated in cells expressing caspase-resistant channels. Enhanced cleavage of Kv7.1 can be detected for the LQT mutation G460S, which is located adjacent to the cleavage site, whereas a calmodulin-binding-deficient mutation impairs cleavage. Application of apoptotic stimuli or doxorubicin-induced cardiotoxicity provokes caspase-mediated cleavage of endogenous IKs in human cardiomyocytes. In summary, caspases are novel regulatory components of IKs channels that may have important implications for the molecular mechanism of doxorubicin-induced cardiotoxicity.
Similar content being viewed by others
Introduction
Voltage-gated potassium channels (Kv) form a protein class comprising 40 members in humans, which can be grouped into 12 families1. Among these, the Kv7 (KCNQ) family has attracted special attention since mutations in the five KCNQ genes cause heritable diseases, highlighting their physiological importance2,3. Dominant-negative mutations in the gene encoding Kv7.1 are associated with cardiac arrhythmias contributing to LQT syndrome4, whereas patients carrying loss-of-function mutations on both alleles additionally suffer from severe congenital hearing loss5. Gain-of-function mutations were found in patients with a form of autosomal dominant atrial fibrillation6, highlighting the important functions of Kv7.1 in the heart and inner ear. In both tissues, the α-subunit Kv7.1 coassembles with the β-subunit KCNE1, composing an ion channel conducting the slow component of the delayed rectifier potassium current, IKs, which is indispensable for shaping the cardiac action potential7,8. Another important physiological function of Kv7.1 was discovered by genome-wide association studies, in which single nucleotide polymorphisms in the KCNQ1 locus were associated with type 2 diabetes in several populations9,10. In nonexcitable (e.g., polarized thyroid, intestinal and tracheal epithelial) cells, Kv7.1 associates with the β-subunits KCNE2 and KCNE3, respectively. The latter β-subunits, in contrast to KCNE1, reduce the voltage-dependent gating of the outwardly rectifying Kv7.1 α-subunit, resulting in constitutively open channels. The Kv7.1/KCNE2 channel complex has been described to be crucial for thyroid hormone biosynthesis11, whereas Kv7.1/KCNE3 heterodimers play an important role in chloride secretion across tracheal and intestinal epithelia12. In the intestine, transposon-based forward mutagenesis genetic screens identified KCNQ1 as a cancer susceptibility gene13 and low expression of Kv7.1 was found in patients with colorectal cancer14. However, the role of Kv7.1 in cancer development has yet to be established.
Channels formed from Kv7 α-subunits share some structural features with Shaker-related Kv channels, such as a common core structure of six transmembrane domains (S1−S6) including a voltage-sensing domain (S1−S4) and a pore domain (S5−S6)2. One striking difference, however, is the presence of a large cytoplasmic C-terminal domain in Kv7 channels, which is important for gating, assembly, and intracellular trafficking of the channel and comprises four helical domains (A−D) (reviewed in Haitin and Attali15). Whereas helices A and B mediate calmodulin binding16, helices C and D form the subunit interaction domain17, which consists of a bipartite coiled-coil motif that is crucial for subunit-specific interaction and tetramerization of the Kv7 α-subunits18,19. For Kv7.1, it has also been shown that the A-kinase anchoring protein yotiao binds to helix D, leading to the recruitment of protein kinase A (PKA) and protein phosphatase 1, thereby forming a macromolecular complex important for regulating IKs activity20. In addition to PKA-mediated phosphorylation, ubiquitination and sumoylation have been reported to be important for regulation of Kv7.1 activity at the posttranslational level21,22.
Anthracyclines such as doxorubicin are well-established and effective antineoplastic agents, commonly used for the treatment of cancers. However, doxorubicin treatment has cardiotoxicity as a severe side effect, which can lead to QT prolongation and heart failure23,24. Increased production of reactive oxygen species (ROS) and the induction of mitochondrial dysfunction are well-described molecular mechanisms of doxorubicin-induced cardiomyopathy, resulting in an activation of apoptotic pathways, finally leading to caspase activation25. Dexrazoxane, a cardioprotective agent, acts in the same pathway by chelating Fe2+ ions more effectively than doxorubicin. While Fe2+ ions bound to doxorubicin are efficiently oxidized to Fe3+, resulting in release of electrons, which are rapidly transferred to produce ROS, the redox activity of dexrazoxane on Fe2+ ions is much lower, so that the Fe2+-scavenging activity and the consequently reduced ROS production by dexrazoxane have been suggested to underlie the alleviating effect on cardiotoxicity26.
Caspases are cysteine proteases, which specifically cleave their substrates at the C-terminal side of an aspartic acid and play important roles in numerous aspects of physiology such as apoptosis, aging, development, and inflammation27. Caspases are well known for their executive role in apoptosis and can be grouped into initiator (caspase 2, 8–10) and effector caspases (caspase 3, 6 and 7)28. Activation of caspases is triggered either by the extrinsic pathway, mediated by ligand binding to death receptors and activation of the initiator caspase 8, or by the intrinsic pathway28. In the latter case, mitochondrial membranes are permeabilized by the proapoptotic proteins BCL-2 and BAX, leading to a loss of mitochondrial transmembrane potentials and to the release of other proapoptotic proteins such as cytochrome C into the cytosol, which results in the activation of the initiator caspase 9. Activated caspases 8 and 9 specifically cleave effector caspases, which finally execute apoptosis28. With the exception of caspase 14, all other caspases in humans have been implicated in inflammation28. Moreover, altered caspase expression levels have been correlated with ageing29 and heart failure30,31. Recently, it has become evident that caspases also have nonapoptotic and noninflammatory functions, such as regulation of long-term depression32 or organelle removal during terminal differentiation33.
By using Kv7.1-specific antibodies directed against an epitope on the channel’s C-terminus, we detected two c-terminal fragments, leading us to the hypothesis that Kv7.1 is processed by unknown proteases. In the present study, we identify Kv7.1 as a novel substrate for caspases, which may have important implications for understanding the role of Kv7.1 in cardiac arrhythmias and its function as a tumor suppressor. Our data suggest that caspase-mediated proteolysis of Kv7.1 leads to decreased Kv7.1-mediated currents, representing a novel regulatory mechanism for modulating Kv7.1 channel activity. Furthermore, we show that Kv7.1 cleavage is induced upon administration of doxorubicin, which efficiently activates caspase 3 in human cardiomyocytes. To our knowledge, Kv7.1 is the first example of a voltage-gated potassium channel that acts as a substrate for caspases.
Results
Proteolysis of Kv7.1 produces C-terminal fragments
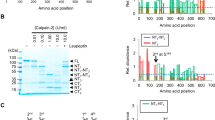
We and others34,35 have noted the occurrence of Kv7.1 C-terminal fragments in transfected cells, when antibodies directed against epitopes on the Kv7.1 C-terminus were used, which prompted us to further analyze the specificity of these fragments. We found C-terminal fragments of the full-length Kv7.1 channel in lysates derived from transiently transfected cells with human or murine Kv7.1 cDNA constructs (Fig. 1a). Two fragments with a molecular mass of about ~40 and ~28 kDa can be detected in addition to the full-length form of Kv7.1 at ~70 kDa, when an antibody directed against a C-terminally derived peptide of Kv7.1 was used (Fig. 1a and Supplementary Fig. 1A). Next, we demonstrated the specificity of the Kv7.1 antibody by the absence of signals in various tissues derived from Kv7.1-deficient mice, which do not show detectable Kv7.1 in immunoblots (Supplementary Fig. 1B). As expected, we found the highest expression of Kv7.1 in murine heart, when compared to kidney and pancreas, further supporting the specificity of the antibody (Supplementary Fig. 1B). C-terminal fragments of Kv7.1 were also detectable in cells transfected with a cDNA construct of human Kv7.1 tagged at the C-terminus with an MYC epitope (Fig. 1b). Immunoblots with either the MYC or the Kv7.1 antibodies resulted in the same pattern. Next, we analyzed neonatal rat ventricular cardiomyocytes (NRVMs) for endogenous expression of Kv7.1. Again, we found immunoreactive bands resembling full-length Kv7.1 (~70 kDa) and bands at ~40 and ~28 kDa, albeit with low intensity (Supplementary Fig. 1C). We concluded that Kv7.1 is cleaved twice at its C-terminus, resulting in three fragments: The N-terminal fragment comprising the cytoplasmic N-terminus and the membrane-embedded part of the protein and two C-terminal fragments showing immunoreactivity with the Kv7.1 antibody, which we termed CTF1 (~40 kDa) and CTF2 (~28 kDa) (Supplementary Fig. 1A).
Kv7.1 is cleaved at aspartate 459. a Western blot analysis of HeLa cell lysates overexpressing human and murine Kv7.1 constructs. Untransfected cells (Ø) served as negative control. Densitometric analysis of four independent experiments of the CTF1 or CTF2 band intensity normalized to the full-length Kv7.1 band intensity. b Immunoblot analysis of lysates derived from HeLa cells overexpressing indicated constructs. Untransfected cells (Ø) served as negative control. c Schematic illustration of different deletion constructs in the linker region between helices A and B. HeLa cell lysates overexpressing indicated constructs analyzed by western blot. Untransfected cells (Ø) served as negative control. d Alanine scan of position 457 to 462. Lysates from HeLa cells overexpressing indicated constructs were used for western blot analysis. Untransfected cells (Ø) served as negative control. Densitometric analysis of six independent experiments of CTF2 band intensity normalized to Kv7.1 full-length band intensity. Statistics were tested with one-way-ANOVA followed by Bonferroni’s Multiple Comparison test. e Cleavage site sequence logo of Kv7.1 derived from 32 different species. Logo was created with weblogo.berkley.edu and adapted accordingly. The arrow indicates the cleavage site. a Anti-Kv7.1 antibody. b−d Anti-MYC antibody. All dot blots are shown as mean and error bars as SEM
To determine the cleavage sites, we generated two Kv7.1 constructs: one comprising the complete C-terminus (CT, starting with glycine at position 348), and the other beginning at helix B (CT1 beginning with amino acid 505) (Supplementary Fig. 1A). After expression of these fragments in HeLa cells, we compared the size of the resulting bands of CT and CT1 with the C-terminal fragments CTF1 and CTF2 (Fig. 1c). CT and CTF1 migrate at the same molecular weight, whereas CTF2 runs slightly slower than the CT1 construct. From these data, we conclude that the cleavage site for the generation of CTF1 resides between the end of the transmembrane domain S6 and helix A, while the cleavage site for CTF2 likely lies within the loop region between helices A and B in the C-terminus of Kv7.1.
Since CTF2 is the most abundant C-terminal fragment (Fig. 1a, b, Supplementary Fig. 1C), we focused our analysis on the region between helices A and B. We generated five Kv7.1 constructs with varying deletions within this linker region (Fig. 1c). Immunoblot analysis of lysates derived from cells expressing these constructs revealed that a stretch of six amino acids (S457VDGYD462) was essential for the occurrence of CTF2 (Fig. 1c). Next, we performed an alanine scan over the SVDGYD region to define the precise cleavage site. Mutating the aspartate at position 459 to an alanine resulted in a complete loss of CTF2 generation, whereas mutating the other five residues (including Asp-462) appeared to have no significant effect (Fig. 1d). These data demonstrate that Kv7.1 is cleaved within the SVDGYD motif at position D459 by a protease, which requires an aspartate. However, Kv7.1 is the only member within the Kv7 family carrying an aspartate at this position (Supplementary Fig. 1D). This aspartate is highly conserved within Kv7.1 protein sequences derived from over 20 different species (Fig. 1e and Supplementary Fig. 2).
Kv7.1 is cleaved by caspases upon apoptosis
To identify the protease responsible for the generation of CTF2, we searched the Merops Database for proteases that require an aspartate in the cleavage site36. Since caspases critically depend on an aspartate at the P1 position27, we used two caspase inhibitors (Q-VD-OPH and Z-VAD(OMe)-FMK) to determine whether caspases are responsible for the generation of CTF2. Both compounds effectively inhibited the generation of CTF2 but not CTF1 (Fig. 2a and Supplementary Fig. 3). To activate caspases, we induced apoptosis in cells overexpressing wild-type Kv7.1 and the D459A mutant by applying staurosporine, a nonselective protein kinase inhibitor widely used as a proapoptotic stimulus. Whereas wild-type Kv7.1 was efficiently proteolysed, the D459A mutant appeared resistant to staurosporine treatment (Fig. 2b), demonstrating that the D459A mutant is insensitive to staurosporine-induced caspase activation and cleavage. Next, we asked whether the generation of the CTF2 was dependent on the full-length Kv7.1 α-subunit. We therefore expressed the CT construct (Supplementary Fig. 1A) under control and apoptotic conditions. Again, an increase in CTF2 generation could be observed upon staurosporine treatment (Fig. 2c) indicating that cleavage can occur independent from the membrane-embedded part of the Kv7.1 protein. To gain deeper insight into the involvement of caspases in Kv7.1 proteolysis, we applied staurosporine and the specific caspase 8 inhibitor II to cells expressing Kv7.1. As shown in Fig. 3a, inhibition of caspase 8 efficiently blocked activation of downstream caspase 3 and the generation of CTF2 dose-dependently, demonstrating that the staurosporine-induced CTF2 generation is mediated by caspases.
Cleavage of Kv7.1 occurs during apoptosis. a Western blot analysis of Cos7 cells overexpressing Kv7.1-MYC treated for 12 h with 450 nmol per L Q-VD-OPh. Untransfected (Ø), nontreated and vehicle-treated cells served as negative controls. Densitometric analysis of five independent experiments of CTF2 band intensity normalized to Kv7.1 full-length band intensity. Statistics were tested with one-way-ANOVA followed by Bonferroni’s Multiple Comparison test. b Lysates derived from HeLa cells expressing Kv7.1-MYC and Kv7.1-D459A-MYC constructs treated with 1 µmol per L staurosporine for 8 h analyzed by immunoblotting. Untransfected (Ø) and vehicle-treated cells served as negative controls. Densitometric analysis of eight independent experiments of the CTF2 band intensity normalized to Kv7.1 full-length band intensity. c Western blot analysis of HeLa cell lysates overexpressing CT-MYC construct treated for 8 h with 1 µmol per L staurosporine. Vehicle-treated cells served as negative controls. a−c Anti-MYC antibody, a, c anti-β-actin antibody. All dot blots are shown as mean and error bars as SEM
Caspases are responsible for the generation of CTF2. a Western blot analysis of lysates derived from HEK 293T cells stably expressing Kv7.1-MYC treated with either 1 µmol per L staurosporine for 6 h or treated with 1 µmol per L staurosporine for 6 h and pretreated for 2 h with 20 or 50 µmol per L of a caspase-8 inhibitor II. Untransfected (Ø) and vehicle-treated cells served as negative controls. Densitometric analysis of 4–6 independent experiments of the CTF2 band intensity normalized to Kv7.1 full-length band intensity. Statistics were tested with one-way-ANOVA followed by Bonferroni’s Multiple Comparison test. # indicates nonspecific binding of the antibody. b MCF-7 lysates expressing Kv7.1-MYC and caspase-3 or caspase-3-D28A-D175A analyzed by immunoblot. Untransfected (Ø) and eGFP-transfected cells served as negative controls. # indicates nonspecific binding of the antibody. c Lysates of HEK 293T cells stably expressing Kv7.1 and coexpressing indicated caspases analyzed by immunoblot. Untransfected (Ø) and eGFP-transfected cells served as negative controls. d Lysates of HL-1 cells treated for 6.5 h with 0.5, 1, 1.5 and 2 µmol per L of staurosporine analyzed by immunoblot. Vehicle-treated cell lysates served as negative control. Densitometric analysis of CTF2 band intensity normalized to Kv7.1 full-length band intensity of five independent experiments. Statistics were tested with one-way-ANOVA followed by Bonferroni’s Multiple Comparison test. a−c Anti-MYC antibody. Anti-β-actin antibody. a, b, d Anti-caspase-3 antibody. d Anti-Kv7.1 antibody. All dot blots are shown as mean and error bars as SEM
To address the question whether caspase 3, one of the major effector caspases, is solely responsible for Kv7.1 cleavage at position D459, we used a human breast carcinoma MCF-7 cell line, which is deficient for caspase 3 37. Overexpression of Kv7.1 in MCF-7 cells still resulted in the generation of CTF2 albeit to a lower extent (Fig. 3b, eGFP labeled lane). Whereas cotransfection of Kv7.1 together with wild-type caspase-3 restored CTF2 production, the overexpression of an inactive form of caspase-3 led to a reduction of CTF2 levels (Fig. 3b) likely by protecting Kv7.1 from endogenous caspases. To determine whether all caspases cleave Kv7.1 to the same extent, we coexpressed Kv7.1 with at least one caspase of each group, namely caspases 1, 2, 3, 7, and 8. Immunoblots revealed that all tested caspases are able to generate CTF2 (Fig. 3c). Stronger cleavage could be observed by overexpression of the initiator caspase 1, 2 and 8, which might be due to an activation of downstream effector caspases. Nevertheless, these data suggest that Kv7.1 is a substrate of all analyzed caspases.
To demonstrate that endogenous Kv7.1, embedded in the IKs channel complex undergoes caspase-mediated proteolysis, we treated murine cardiac muscle cells (HL-1 cells38) with increasing concentrations of staurosporine, confirming a dose-dependent occurrence of CTF2 (Fig. 3d). Notably, full-length Kv7.1 channel was efficiently cleaved at higher staurosporine concentrations. In summary, our data strongly suggest that Kv7.1 is cleaved by caspases at an aspartate at position 459.
Functional impact of proteolysis on Kv7.1/KCNE1 channels
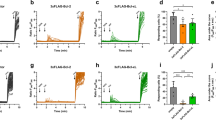
To analyze the functional impact of caspase-mediated cleavage of Kv7.1 upon induction of apoptosis, we cotransfected HEK 293T cells with wild-type Kv7.1 and the D459A mutant together with KCNE1 and measured whole-cell currents under staurosporine treatment and control conditions (Fig. 4a). Drug treatment produced a small but significant reduction of Kv7.1/KCNE1 currents, whereas currents generated by Kv7.1 D459A/KCNE1 channels remained unaffected (Fig. 4b). Cells were harvested after patch-clamp measurements and were subjected to immunoblot analysis to probe for CTF2. Again, we were able to detect CTF2 in cells expressing wild-type Kv7.1 but not the D459A mutant (Supplementary Fig. 3B). One possible explanation for the small functional effect of caspase-mediated cleavage on Kv7.1/KCNE1-mediated currents could be a protective effect of KCNE1 on Kv7.1. Since the majority of our analysis so far was done using cells expressing homomeric Kv7.1 channels, we therefore compared the generation of CTF2 in presence and absence of KCNE1. As shown in Fig. 4c, Kv7.1 is efficiently cleaved upon caspase activation even in the presence of KCNE1 indicating that the β-subunit is not shielding the heteromeric Kv7.1/KCNE1 complex from proteolysis. Subsequently, we performed surface biotinylation experiments to prove that the Kv7.1/KCNE1 heteromeric channels can be processed at the plasma membrane by caspases as demonstrated by the presence of CTF2 in the isolated fraction of cell surface proteins (Fig. 4d). Furthermore, these data suggest that CTF2 is, under this condition, still associated with the apoprotein complex. In summary, these data strongly suggest that the activity of heteromeric Kv7.1/KCNE1 channels can be modified by caspase-mediated cleavage in the C-terminus of Kv7.1.
Cleavage of Kv7.1 in physiology and pathophysiology. a Representative current traces for Kv7.1-MYC and Kv7.1-D459A-MYC, both coexpressed with KCNE1. b Mean currents amplitude was plotted versus voltage to obtain current−voltage (I−V) relationships in cells expressing Kv7.1-MYC (n = 39 for vehicle, n = 16 for staurosporine treatment) or Kv7.1-D459A-MYC (n = 27 for vehicle, n = 17 for staurosporine treatment) and KCNE1 treated with 500 nmol per L staurosporine for 10–12 h. Statistics were tested with two-way ANOVA followed by Bonferroni post-tests. c Immunoblot analysis of HeLa cells coexpressing Kv7.1 with KCNE1-MYC treated with 1 µM staurosporine for 4.5 h. Untransfected (Ø) and vehicle-treated cells served as negative controls. d Biotinylating study analyzed by immunoblots of Hek 293 cells coexpressing Kv7.1 and KCNE1-MYC treated with 1 µM staurosporine for 3 h. Untransfected (Ø) cells as well as cells not treated with biotin served as negative controls. IP Immunoprecipitation. TL total lysate. e Schematic illustration to highlight the position of G460 and A372 and calmodulin binding site in helix A. Western blot analysis of HeLa cell lysates overexpressing indicated constructs. Untransfected (Ø) and Kv7.1-D459A-transfected cells served as negative controls. Densitometric analysis of four independent experiments of CTF2 band intensity normalized to Kv7.1 full-length band intensity. Statistics were tested with one-way-ANOVA followed by Bonferroni’s Multiple Comparison test. f Coimmunoprecipitation study analyzed by immunoblots of HeLa cells overexpressing wild-type Kv7.1 and the A372D mutant with endogenous calmodulin. IP Immunoprecipitation with anti-Kv7.1 antibody, IB Immunoblot, TL total lysate. Untransfected cells (Ø) served as negative control. c Anti-KCNE1 antibody, anti-caspase 3 antibody. c, d Anti-Kv7.1 antibody, anti-GAPDH antibody. e Anti-β-actin antibody. e, f Anti-MYC antibody. f Anti-calmodulin antibody. All graphs are shown as mean and error bars as SEM
Identifying long-QT mutations modulating Kv7.1 proteolysis
Next, we tested if disease-causing mutations in Kv7.1 can interfere with the generation of CTF2 and focused on the LQT1 mutation G460S, which is located just one amino acid downstream of the aspartate residue important for Kv7.1 proteolysis (Fig. 4d). Immunoblot analysis of the G460S mutant protein demonstrated significantly increased CTF2 levels when compared to wild-type Kv7.1, indicating that this LQT1 mutation renders Kv7.1 more susceptible to caspase-mediated cleavage even under nonstaurosporine treatment conditions (Fig. 4d). This observation is in line with the finding that this mutation causes a decrease in IKs-like currents39.
The caspase cleavage site resides in an approximately 80 amino acid-long intervening loop between helices A and B (Fig. 4d). Both helices contain an IQ motif, which is important for calmodulin binding (Fig. 4d). Recently, it has been suggested by crystallography, molecular modeling, biochemical, and functional analyses that one bifunctional calmodulin molecule embraces both helices from one Kv7.1 subunit34. Given the important role of calmodulin for Kv7.1 function, we asked whether LQT1-associated mutations within the calmodulin binding sites interfere with CTF2 generation. Indeed, the analyses of the Kv7.1 A372D mutation, which is located close to the calmodulin binding motif in helix A, revealed no detectable CTF2 in immunoblots (Fig. 4d). Coimmunoprecipitations proved that the A372D mutation impaired the interaction of endogenous calmodulin with Kv7.1 (Fig. 4e), suggesting that calmodulin binding is necessary for caspase-mediated proteolysis of Kv7.1.
Doxorubicin induces Kv7.1 proteolysis in cardiomyocytes
It is well known that cancer treatment by the common antineoplastic doxorubicin is hindered by severe cardiotoxic side effects, and there is strong evidence in the literature that doxorubicin leads to caspase activation in cardiomyocytes40. In order to determine whether interference with cardiac function by doxorubicin also involves caspase-mediated cleavage of Kv7.1, we treated human-induced pluripotent stem cell-derived cardiomyocytes with staurosporine and doxorubicin. In total lysates derived from untreated human-induced pluripotent stem cell-derived cardiomyocytes, we could only detect trace amounts of CTF2, when we used the C-terminal Kv7.1 antibody to precipitate Kv7.1 (compare Fig. 5a and Supplementary Fig. 3C). We were also unable to detect CTF2 in murine cardiomyocytes (Fig. 3d) and tissue (Supplementary Fig. 1B), suggesting that baseline levels of CTF2 and likely Kv7.1 proteolysis are rather low. However, staurosporine and doxorubicin activated caspase 3 as indicated by the occurrence of cleaved-active forms and a reduction of the inactive zymogen of the protease (Fig. 5a). Doxorubicin treatment at higher concentration appeared to be more efficient in producing active caspase 3, which correlated with a higher abundance of CTF2 and a strong reduction of monomeric and tetrameric forms of Kv7.1 (Fig. 5a). Interestingly, we detected a potentially dimeric form of Kv7.1 at 150 kDa in staurosporine- and doxorubicin-treated cells, suggesting that the caspase-mediated destruction of tetramers results in Kv.7.1 dimers in human-induced pluripotent stem cell-derived cardiomyocytes (Fig. 5a). Nevertheless, lower doses of doxorubicin failed to produce similar levels of CTF2, which correlated with a decrease in caspase 3 activation (Fig. 5b).
Doxorubicin induces cleavage of Kv7.1 to CTF2. a Immunoblot analysis of human-induced pluripotent stem cell-derived cardiomyocytes treated either with vehicle, staurosporine (2 µM) for 4 h or doxorubicin (10 µM) overnight. # indicates nonspecific binding of the Kv7.1 antibody. b Densitometric analysis of 3–6 independent experiments of band intensities of CTF2 normalized to GAPDH band intensity. Statistics were tested with one-way-ANOVA followed by Bonferroni’s Multiple Comparison test. a Anti-Kv7.1 antibody, anti-caspase-3 antibody, anti-GAPDH antibody. All dot blots are shown as mean and error bars as SEM
Discussion
To date, over 1500 caspase cleavage sites and substrates have been identified41. The vast majority of caspase-mediated cleavage occurs during apoptosis, but caspase function has also been demonstrated in nonapoptotic cellular responses42, suggesting that caspases cleave a specific subset of substrates independent of apoptosis. Here, we report Kv7.1 as a novel substrate for caspases, representing to our knowledge the first example of a voltage-gated cation channel undergoing caspase-mediated proteolysis. However, the transient receptor potential melastin-like 7 (TRMP7) has been also identified as a caspase substrate, which appears to be critical for Fas-induced apoptosis43.
Cleavage of human Kv7.1 by caspases occurs after an aspartate at position 459, which is located within the intervening loop between helices A and B in the channel’s large cytoplasmic C-terminal domain that serves as a scaffold for numerous protein−protein interactions involved in cellular signaling cascades15. Both helices appear to form a two-helical bundle, which is embraced by a calmodulin molecule as revealed by X-ray crystallography of recombinantly expressed calmodulin and the proximal C-terminus of Kv7.134. In this study, the intervening loop, in which the caspase-mediated proteolysis of Kv7.1 occurs, was deleted. Thus, structural information about the caspase cleavage site is missing so far. However, our finding that the calmodulin binding-deficient Kv7.1 A372D mutant is not processed by caspases strongly suggest that calmodulin either helps to recruit caspases to the channel complex or determines the structure of the intervening loop necessary for proper caspase recognition.
Our functional analysis using patch-clamp suggests that caspase cleavage interferes with the function of Kv7.1. This finding is supported by numerous reports showing that the intracellular C-terminal domain of Kv7.1 is responsible for channel tetramerization, trafficking and modulating the biophysical properties of the channel15. The rather small effect of Kv7.1 cleavage on whole-cell currents could be explained by the continued association of CTF2 with the apoprotein complex. For example, calmodulin could, by binding to helices A and B, act as a bridging molecule. Alternatively, the interaction of CTF2 via helices C and D with uncleaved Kv7.1 subunits could keep CTF2 in the channel complex. However, our cell surface biotinylation data strongly suggest that CTF2 is still present in heteromeric Kv7.1/KCNE1 localized at the plasmalemma.
Many pathogenic LQT1 mutations have been mapped to the C-terminus of Kv7.1 emphasizing the functional importance of this particular channel region44. Our finding that the LQT1 mutation G460S, which is adjacent to the aspartate 459 residue, is more susceptible to caspase cleavage suggest a potential novel pathophysiologic mechanism for LQT1 mutations located in the C-terminus of Kv7.1.
Based on the analysis of a number of cleavage sites, a general consensus motif of DXED-A/G/S/T has been proposed for apoptosis executioner caspases such as caspases 3 and 7, whereas caspases 2, 8, 9, and 10 and caspases 1, 4, 5, 6, and 14, prefer isoleucine/ leucine or tryptophan/tyrosine/valine instead of an aspartate at the first position, respectively41. Due to the overlapping specificity of caspases and the significantly different Kv7.1 cleavage site (amino acid sequence: FSVD-G), it is difficult to predict which individual caspase cleaves Kv7.1. Our coexpression data suggest that Kv7.1 can be cleaved by all tested caspases including caspases 1, 2, 3, 7 and 8. In this group, caspase 1 was more efficient in Kv7.1 processing, which is in agreement with the larger similarity of the preferred cleavage site41.
Caspases have well-established functions in the execution of apoptosis, as well as inflammation28. However, transiently active caspases have also been detected in nonapoptotic cells. For example, in neurons, caspases 3 and 9 are critical for long-term depression and AMPA receptor internalization32. In the heart, increased expression of caspase 1 was found in murine heart failure models and in patients with end-stage heart failure31. Analysis of mice with heart-targeted overexpression of caspases 1 and 3 further supported the notion that caspases contribute to heart diseases, likely based on an overlap of apoptotic and nonapoptotic functions30,31. Our finding that IKs is sensitive to caspase-mediated cleavage uncovers a novel molecular mechanism that may contribute to cardiac arrhythmias, which is strongly supported by an increased susceptibility of the LQT1 G460S mutant to proteolytic processing by caspases. Thus, it is likely that the reported 40% smaller current density of the G460S mutant, when compared to wild-type IKs, is at least partially due to an increased cleavage of the mutant39. Although several C-terminally located LQT1 mutations have been identified which interfere with channel function by modulating calmodulin16,45 or phosphatidylinositol 4,5-bisphosphate46 binding and/or disrupt assembly of functional IKs15, for most of the C-terminal LQT mutations, the pathophysiological mechanism that leads to disease is unknown. Our data strongly suggest that susceptibility to caspase-mediated degradation should also be considered when analyzing these mutations.
Furthermore, our results demonstrate that doxorubicin treatment of human-induced pluripotent stem cell-derived cardiomyocytes efficiently induced caspase-mediated cleavage of Kv7.1, suggesting that this pathway might contribute to doxorubicin-induced cardiotoxicity. Interestingly, doxorubicin has also been shown to induce electrocardiogram abnormalities such as QT interval prolongations, which are often observed within the first day after chemotherapy25. These results have been confirmed in animal studies, showing that doxorubicin prolongs the cardiac action potential duration by specifically inactivating IKs but not IKr, which both compose the delayed rectifier potassium current IK23. In cardiomyocytes, IKs is mediated by a macromolecular complex formed by assembly of the pore-forming subunits Kv7.1 with KCNE1 β-subunits, which are linked to the scaffolding protein yotiao/A-kinase anchoring protein 9 (AKAP-9)20. Yotiao binds to the distal part of the C-terminus of Kv7.1 and recruits PKA, protein phosphatase 1 (PP1), adenylate cyclase 9 (AC9) and phosphodiesterase PDE4D3 to the complex, allowing the control of the phosphorylation state of Kv7.1, which is the molecular basis for the β-adrenergic regulation of IKs20,47. In silico sequence analyses of yotiao predict several potential caspase cleavage sites, suggesting that this scaffold protein is also processed by caspases41. Thus, it is conceivable that caspase-mediated processing of Kv7.1 and likely yotiao contributes to the prolongation of the QT interval mediated by doxorubicin. It will be important to determine the pathway by which doxorubicin treatment leads to elevated caspase activity and Kv7.1 cleavage. While it is widely accepted that doxorubicin-induced cardiotoxicity is due to the induction of mitochondrial dysfunction, resulting in an increased production of ROS in the cytoplasm and consequent activation of extrinsic and intrinsic apoptotic pathways, a more direct effect on ROS-mediated signaling by the oxidizing activity of doxorubicin on Fe2+ ions, as suggested by the alleviating effect of dexrazoxane, must also be considered. This will also clarify the issue of whether regulated C-terminal cleavage of Kv7.1 is more generally involved in ROS-mediated cardiac responses.
In summary, the present study demonstrates caspase-mediated proteolysis of Kv7.1. Posttranslational modifications such as phosphorylation, ubiquitination, sumoylation, palmitoylation, and glycosylation have been reported for potassium channels48. According to our data, proteolysis of Kv7.1 mediated by caspases is another important mechanism of posttranslational modification of Kv7.1, and we hypothesize that analysis of this regulation will advance understanding of the molecular mechanism of doxorubicin-induced cardiotoxicity.
Methods
Plasmids and antibodies
Human Kv7.1 and caspase cDNAs were subcloned in the expression vectors pFrog and pcDNA4/TO (Invitrogen, Waltham, USA), respectively. Mutations, deletions and tags for antibodies were constructed/introduced by recombinant PCR and verified by sequencing. The C-terminal anti-Kv7.1 antibody was directed against the following peptide sequence TVPRRGPDEGS. The following antibodies were used: rabbit anti-β-actin (A2066, Sigma-Aldrich, St. Louis, USA), rabbit anti-caspase 3 (8G10, Cell Signaling, Cambridge, UK), rabbit anti-calmodulin (ab45689, Abcam, Cambridge, UK), rabbit anti-Eef2 (eukaryotic translation elongation factor 2, ab33523, Abcam, Cambridge, UK), mouse anti-GAPDH (MAB374, Millipore, Billerica, USA), rabbit anti-KCNE1 (APC-163, Alomone Labs, Jerusalem, Israel), rabbit anti-Kv7.1 (ab77701, Abcam, Cambridge, UK), mouse anti-MYC (9B11, Cell Signaling, Cambridge, UK), goat anti-MYC (GTX29106, GeneTex Inc., Irvine, USA).
Cell culture, transfection, and inhibitors
HEK 293T, HeLa, Cos7, and MCF-7 (ATCC, Manassas, USA) were grown in Dulbecco’s modified Eagle’s medium (DMEM, Thermo Fisher Scientific) supplemented with 10% fetal bovine serum (FBS, Biochrome, Berlin, Germany), 100 U per mL penicillin and 100 µg per mL streptomycin (both Carl Roth, Karlsruhe, Germany). For HL-1 cells (gift from W.C. Claycomb), Claycomb medium (Sigma-Aldrich), supplemented with 10% FBS (Biochrome), 100 U per mL penicillin, 100 µg per mL streptomycin, 0.1 mM norepinephrine (Carl Roth) and 2 mM l-Glutamine (Carl Roth) was used. All cells were kept at 37 °C and 5% CO2. Transient transfections were performed using TurboFect (Thermo Fischer Scientific) according to the manufacturer’s instructions, and stable cell lines were established using G418 for selection. Inhibitors were used as follows: Z-VAD-FMK (12 h, 100 µM, Promega, Fitchberg, USA), Q-VD-OPh (12 h, 450 nM, Merck Millipore, Billerica, USA), staurosporine (1 to 8 h, 0.5–2 µM, Sigma-Aldrich), caspase-8 inhibitor II (2 h pretreatment, 20 or 50 µM, Calbiochem, Billerica, USA), Doxorubicin (overnight, 0.1 µM–10 µM, Sigma-Aldrich).
Protein extraction, immunoprecipitation, and immunoblotting
Cells were washed twice with phosphate-buffered saline (PBS) and harvested in PBS, containing a protease inhibitor cocktail (Complete, Roche, Basel, Switzerland). After centrifugation, cell pellets were lysed in PBS/Complete (1% Triton X-100, Carl Roth) by sonication. After 1 h incubation on ice, samples were centrifuged, and supernatants were analyzed by SDS-PAGE. For coimmunoprecipitations and immunoprecipitations, cells were lysed in EBC buffer/Complete (120 mM NaCl, 50 mM Tris-HCl, 0.5% NP-40, pH 7.4 (all from Carl Roth)) and treated as described above. Lysates were incubated with mouse anti-MYC antibody (coimmunoprecipitation) or treated with Kv7.1-myc antibody (immunoprecipitation) at 4 °C overnight. For precipitation, protein G agarose beads (coimmunoprecipitation) or protein G dynabeads (immunoprecipitation) were used. After thorough washing of the beads, protein complexes were released by denaturation. Samples were subjected to SDS-PAGE and transferred onto nitrocellulose membranes by tank blotting. Membranes were blocked and incubated overnight at 4 °C in primary antibody solution followed by incubation with the appropriate secondary antibodies conjugated to horseradish peroxidase. After thorough washing, bound antibodies were detected by chemiluminescence using a luminescent imager (LAS-4000, Fujifilm, GE Healthcare, Little Chalfont, UK). For quantifications, ImageJ software was used.
Biotinylating assay
After washing cells twice with PBS/CM (0.1 mM CaCl2, 1 mM MgCl2), cells were incubated with 0.5 mg NHS biotin ester (Thermo Fisher) in PBS/CM for 10 min. By adding 50 mM Glycin in PBS/CM biotinylation was stopped. After two washing steps with PBS/CM, cells were lysed as described for immunoprecipitation studies above. Incubation with Streptavidin beads for 1 h at 4 °C was used to precipitate biotinylated proteins. After through washing proteins were released by denaturation and subjected to SDS-PAGE and immunoblotting as described above.
Isolation of neonatal rat ventricular cardiomyocytes
Hearts of 1–2-day-old Wistar rats were harvested and minced in buffer (120 mmol NaCl, 20 mmol HEPES, 8 mmol NaH2PO4, 6 mmol glucose, 5 mmol KCl, 0.8 mmol MgSO4, pH = 7.4). Subsequently, up to six digestion steps were carried out with 0.6 mg per mL pancreatin (Sigma-Aldrich) and 0.5 mg per mL collagenase type II (Worthington, Lakewood, USA) in sterile ADS buffer. Cardiomyocytes were purified from contaminating fibroblasts using a Percoll gradient centrifugation step. Finally, NRVMs were resuspended and cultured in DMEM containing 10% FBS, 100 U per mL penicillin, 100 µg per mL streptomycin and 1% l-Glutamine. Protein extractions were performed as described above.
Human-induced pluripotent stem cell culture
Human-induced pluripotent stem cell-derived cardiomyocytes49 were routinely maintained in E8 medium implemented with 10 μM Rho kinase inhibitor (Y27632; Biorbyt, Cambridge, UK) for the first 24 h after passage on 1:400 reduced growth factor Matrigel (Corning, Corning, USA). Cells were passaged ~1:15 every 3–4 days using 0.5 mM EDTA in Dulbecco’s PBS (DPBS; Corning, Corning, USA) after achieving ~80% confluence. Cell lines were used between passages 20 and 85. All cultures were routinely tested for mycoplasma using a MycoAlert PLUS Kit (Lonza, Basel, Switzerland).
Cardiac differentiation from human iPS cells
Cardiac differentiation was performed as described previously49. Briefly, to initiate differentiation, medium was changed to CDM3, consisting of RPMI 1640 (Corning), 500 μg ml–1 Oryza sativa-derived recombinant human albumin (Oryzogen, Wuhan, China), and 213 μg ml–1 l–ascorbic acid 2-phosphate (Wako, Tokyo, Japan). For days 0–1, media was supplemented with 3 μM of the glycogen synthase kinase 3β inhibitor CHIR99021 (Biorbyt, San Francisco, USA)21,22 and 10 ng ml–1 BMP4 (Peprotech, Hamburg, Germany). On day 1, media was changed to CDM3 and on d2 media was changed to CDM3 supplemented with 2 μM of the Wnt inhibitor Wnt-C59 (Biorbyt). Media was changed on day 4 and every other day thereafter with CDM3. Contracting cells were noted from day 7. At days 25–29, contracting cardiomyocytes were dissociated by incubating 20 min in DPBS followed by 6 min in TrypLE (Thermo Fisher Scientific), and then 60 min with 0.5 U per mL Liberase TH (Roche) in CDM3 media. Cells were replated in 40% FBS in CDM3 by combining 2–3 wells into one well for optimal viability of the cultures. After 2 days, media was changed back to CDM3 and exchanged every 2 days until analysis.
Electrophysiology
Transfected cells were identified by using cotransfection of eGFP and imaging on an inverted fluorescence microscope (Axiovert 40, Zeiss, Jena, Germany) with a fiber optic-coupled light source (UVICO, Rapp OptoElectronic, Hamburg, Germany). Current signals were recorded in whole-cell patch-clamp mode at room temperature (22 ± 1 °C) 2 days after transfection. Recordings were started 3 min after whole-cell access was obtained. Data were sampled at 20 kHz and filtered at 5 kHz, using an Axopatch 200B amplifier in combination with a Digidata 1322A interface and pClamp10 software (all from Molecular Devices/MDS Analytical Technologies, Sunnyvale, USA). Electrodes were made from borosilicate glass (Harvard Apparatus, Edenbridge, UK or BioMedical Instruments, Zoellnitz, Germany), using a DMZ-Universal Puller (Zeitz, Munich, Germany). Pipette resistance in bath solution was 2.0–3.5 MΩ and access resistance was typically <5 MΩ before series resistance compensation (75%). External solution contained (in mM): 145 NaCl, 4 KCl, 2 CaCl2, 2 MgCl2, 10 d-glucose, 10 HEPES, adjusted to pH 7.4 with NaOH. The internal solution was composed of (in mM) 135 K-gluconate, 4 NaCl, 10 KCl, 5 Hepes, 5 EGTA, 2 Na2-ATP, 0.3 Na3-GTP (pH 7.25 with KOH). Cells were incubated with staurosporine at a concentration of 500 nM 10–12 h before recordings. Chemicals were purchased from Sigma-Aldrich.
Statistics
Data are shown as means ± SEM of n observations, as indicated. Statistical analyses were performed using unpaired two-tailed Student’s t test, one-way ANOVA and Bonferroni’s Multiple Comparison post hoc test or two-way ANOVA followed by Bonferroni post-tests, where applicable, using the GraphPad Prism 5 software (GraphPad, San Diego, USA). p ≤ 0.05 was termed significant. *0.01 ≤ p ≤ 0.05. **0.001 ≤ p < 0.01. ***p < 0.001.
Data availability
The authors declare that the data supporting the findings of this study are available within the paper and its supplementary information files.
References
Yu, F. H., Yarov-Yarovoy, V., Gutman, G. A. & Catterall, W. A. Overview of molecular relationships in the voltage-gated ion channel superfamily. Pharmacol. Rev. 57, 387–395 (2005).
Jentsch, T. J., Schroeder, B. C., Kubisch, C., Friedrich, T. & Stein, V. Pathophysiology of KCNQ channels: neonatal epilepsy and progressive deafness. Epilepsia 41, 1068–1069 (2000).
Lehman, A. et al. Loss-of-function and gain-of-function mutations in KCNQ5 cause intellectual disability or epileptic encephalopathy. Am. J. Hum. Genet. 101, 65–74 (2017).
Wang, Q. et al. Positional cloning of a novel potassium channel gene: KVLQT1 mutations cause cardiac arrhythmias. Nat. Genet. 12, 17–23 (1996).
Neyroud, N. et al. A novel mutation in the potassium channel gene KVLQT1 causes the Jervell and Lange-Nielsen cardioauditory syndrome. Nat. Genet. 15, 186–189 (1997).
Chen, Y. H. et al. KCNQ1 gain-of-function mutation in familial atrial fibrillation. Science 299, 251–254 (2003).
Barhanin, J. et al. K(V)LQT1 and lsK (minK) proteins associate to form the I(Ks) cardiac potassium current. Nature 384, 78–80 (1996).
Sanguinetti, M. C. et al. Coassembly of K(V)LQT1 and minK (IsK) proteins to form cardiac I(Ks) potassium channel. Nature 384, 80–83 (1996).
Unoki, H. et al. SNPs in KCNQ1 are associated with susceptibility to type 2 diabetes in East Asian and European populations. Nat. Genet. 40, 1098–1102 (2008).
Yasuda, K. et al. Variants in KCNQ1 are associated with susceptibility to type 2 diabetes mellitus. Nat. Genet. 40, 1092–1097 (2008).
Roepke, T. K. et al. Kcne2 deletion uncovers its crucial role in thyroid hormone biosynthesis. Nat. Med. 15, 1186–U1117 (2009).
Schroeder, B. C. et al. A constitutively open potassium channel formed by KCNQ1 and KCNE3. Nature 403, 196–199 (2000).
Than, B. L. et al. The role of KCNQ1 in mouse and human gastrointestinal cancers. Oncogene 33, 3861–3868 (2014).
den Uil, S. H. et al. Loss of KCNQ1 expression in stage II and stage III colon cancer is a strong prognostic factor for disease recurrence. Br. J. Cancer 115, 1565–1574 (2016).
Haitin, Y. & Attali, B. The C-terminus of Kv7 channels: a multifunctional module. J. Physiol. 586, 1803–1810 (2008).
Shamgar, L. et al. Calmodulin is essential for cardiac IKS channel gating and assembly: impaired function in long-QT mutations. Circ. Res. 98, 1055–1063 (2006).
Schwake, M., Jentsch, T. J. & Friedrich, T. A carboxy-terminal domain determines the subunit specificity of KCNQ K+ channel assembly. EMBO Rep. 4, 76–81 (2003).
Schmitt, N. et al. A recessive C-terminal Jervell and Lange-Nielsen mutation of the KCNQ1 channel impairs subunit assembly. EMBO J. 19, 332–340 (2000).
Schwake, M. et al. Structural determinants of M-type KCNQ (Kv7) K+channel assembly. J. Neurosci. 26, 3757–3766 (2006).
Marx, S. O. et al. Requirement of a macromolecular signaling complex for beta adrenergic receptor modulation of the KCNQ1-KCNE1 potassium channel. Science 295, 496–499 (2002).
Jespersen, T. et al. The KCNQ1 potassium channel is down-regulated by ubiquitylating enzymes of the Nedd4/Nedd4-like family. Cardiovasc. Res. 74, 64–74 (2007).
Xiong, D. et al. SUMOylation determines the voltage required to activate cardiac IKs channels. Proc. Natl. Acad. Sci. USA 114, E6686–E6694 (2017).
Ducroq, J. et al. Dexrazoxane protects the heart from acute doxorubicin-induced QT prolongation: a key role for I(Ks). Br. J. Pharmacol. 159, 93–101 (2010).
Octavia, Y. et al. Doxorubicin-induced cardiomyopathy: from molecular mechanisms to therapeutic strategies. J. Mol. Cell. Cardiol. 52, 1213–1225 (2012).
Renu, K., V, G. A., P, B. T. & Arunachalam, S. Molecular mechanism of doxorubicin-induced cardiomyopathy—an update. Eur. J. Pharmacol. 818, 241–253 (2017).
Hasinoff, B. B., Schroeder, P. E. & Patel, D. The metabolites of the cardioprotective drug dexrazoxane do not protect myocytes from doxorubicin-induced cytotoxicity. Mol. Pharmacol. 64, 670–678 (2003).
Poreba, M., Strozyk, A., Salvesen, G. S. & Drag, M. Caspase substrates and inhibitors. Cold Spring Harb. Perspect. Biol. 5, a008680 (2013).
McIlwain, D. R., Berger, T. & Mak, T. W. Caspase functions in cell death and disease. Cold Spring Harb. Perspect. Biol. 5, a008656 (2013).
Zhang, J. H., Zhang, Y. & Herman, B. Caspases, apoptosis and aging. Ageing Res. Rev. 2, 357–366 (2003).
Condorelli, G. et al. Heart-targeted overexpression of caspase3 in mice increases infarct size and depresses cardiac function. Proc. Natl. Acad. Sci. USA 98, 9977–9982 (2001).
Merkle, S. et al. A role for caspase-1 in heart failure. Circ. Res. 100, 645–653 (2007).
Li, Z. et al. Caspase-3 activation via mitochondria is required for long-term depression and AMPA receptor internalization. Cell 141, 859–871 (2010).
Aram, L. et al. A Krebs cycle component limits caspase activation rate through mitochondrial surface restriction of CRL activation. Dev. Cell 37, 15–33 (2016).
Sachyani, D. et al. Structural basis of a Kv7.1 potassium channel gating module: studies of the intracellular c-terminal domain in complex with calmodulin. Structure 22, 1582–1594 (2014).
Tommiska, J. et al. Two missense mutations in KCNQ1 cause pituitary hormone deficiency and maternally inherited gingival fibromatosis. Nat. Commun. 8, 1289 (2017).
Rawlings, N. D., Barrett, A. J. & Finn, R. Twenty years of the MEROPS database of proteolytic enzymes, their substrates and inhibitors. Nucleic Acids Res. 44, D343–D350 (2016).
Janicke, R. U., Sprengart, M. L., Wati, M. R. & Porter, A. G. Caspase-3 is required for DNA fragmentation and morphological changes associated with apoptosis. J. Biol. Chem. 273, 9357–9360 (1998).
Claycomb, W. C. et al. HL-1 cells: a cardiac muscle cell line that contracts and retains phenotypic characteristics of the adult cardiomyocyte. Proc. Natl. Acad. Sci. USA 95, 2979–2984 (1998).
Arnestad, M. et al. Prevalence of long-QT syndrome gene variants in sudden infant death syndrome. Circulation 115, 361–367 (2007).
Ueno, M. et al. Doxorubicin induces apoptosis by activation of caspase-3 in cultured cardiomyocytes in vitro and rat cardiac ventricles in vivo. J. Pharmacol. Sci. 101, 151–158 (2006).
Kumar, S., van Raam, B. J., Salvesen, G. S. & Cieplak, P. Caspase cleavage sites in the human proteome: CaspDB, a database of predicted substrates. PLoS ONE 9, e110539 (2014).
Kuranaga, E. Beyond apoptosis: caspase regulatory mechanisms and functions in vivo. Genes Cells 17, 83–97 (2012).
Desai, B. N. et al. Cleavage of TRPM7 releases the kinase domain from the ion channel and regulates its participation in Fas-induced apoptosis. Dev. Cell 22, 1149–1162 (2012).
Moss, A. J. & Kass, R. S. Long QT syndrome: from channels to cardiac arrhythmias. J. Clin. Invest. 115, 2018–2024 (2005).
Ghosh, S., Nunziato, D. A. & Pitt, G. S. KCNQ1 assembly and function is blocked by long-QT syndrome mutations that disrupt interaction with calmodulin. Circ. Res. 98, 1048–1054 (2006).
Park, K. H. et al. Impaired KCNQ1-KCNE1 and phosphatidylinositol-4,5-bisphosphate interaction underlies the long QT syndrome. Circ. Res. 96, 730–739 (2005).
Chen, L., Kurokawa, J. & Kass, R. S. Phosphorylation of the A-kinase-anchoring protein Yotiao contributes to protein kinase A regulation of a heart potassium channel. J. Biol. Chem. 280, 31347–31352 (2005).
Huang, X. & Jan, L. Y. Targeting potassium channels in cancer. J. Cell. Biol. 206, 151–162 (2014).
Burridge, P. W. et al. Chemically defined generation of human cardiomyocytes. Nat. Methods 11, 855–860 (2014).
Acknowledgements
We thank Maike Langer, Vanessa Mangels, and Marvin Murowski for excellent technical assistance; William C. Claycomb for the HL-1 cells and Jakob Völkl, Florian Lang, and Karl E. Pfeifer for Kv7.1-deficient murine tissues. This work was supported by the Deutsche Forschungsgemeinschaft (Grant SFB 877, B8 and Heisenberg Fellowship to M.S.). We acknowledge support for the Article Processing Charge by the Deutsche Forschungsgemeinschaft and the Open Access Publication Fund of Bielefeld University.
Author information
Authors and Affiliations
Contributions
A.S., C.R., and M.S. designed the study. A.S., C.R., S.H., T.H., A.J.T.S. performed experiments and data analysis. T.H., A.J.T.S., C.A., T.F., P.W.B., M.L., and M.S. interpreted the data and edited the manuscript. A.S., C.R., T.F., and M.S. wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Strigli, A., Raab, C., Hessler, S. et al. Doxorubicin induces caspase-mediated proteolysis of KV7.1. Commun Biol 1, 155 (2018). https://doi.org/10.1038/s42003-018-0162-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-018-0162-z
This article is cited by
-
Cardiac Remodelling Following Cancer Therapy: A Review
Cardiovascular Toxicology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.