Abstract
Male and female gametes differing in size—anisogamy—emerged independently from isogamous ancestors in various eukaryotic lineages, although genetic bases of this emergence are still unknown. Volvocine green algae are a model lineage for investigating the transition from isogamy to anisogamy. Here we focus on two closely related volvocine genera that bracket this transition—isogamous Yamagishiella and anisogamous Eudorina. We generated de novo nuclear genome assemblies of both sexes of Yamagishiella and Eudorina to identify the dimorphic sex-determining chromosomal region or mating-type locus (MT) from each. In contrast to the large (>1 Mb) and complex MT of oogamous Volvox, Yamagishiella and Eudorina MT are smaller (7–268 kb) and simpler with only two sex-limited genes—the minus/male-limited MID and the plus/female-limited FUS1. No prominently dimorphic gametologs were identified in either species. Thus, the first step to anisogamy in volvocine algae presumably occurred without an increase in MT size and complexity.
Similar content being viewed by others
Introduction
Male–female gamete size dimorphism is a fundamental trait found in most multicellular organisms including plants, animals, fungi, brown algae, red algae, and green algae, and has been a major topic of biological sciences since Charles Darwin first wrote on the topic of sexual selection1. However, such dimorphism is not found in many unicellular species which typically produce morphologically identical gametes—“isogametes”. Thus, male and female gametes differing in size—known as anisogamy—emerged independently from isogamous ancestors in various eukaryotic lineages2. In most lineages, emergence of anisogamy was so ancient that isogamous close relatives are long extinct, making it difficult to infer the genetic bases for steps in the transition from isogamy to anisogamy and oogamy. In contrast, the entire volvocine lineage began diverging from a single-celled, isogamous Chlamydomonas-like ancestor ~200 MYA to give rise to genera exhibiting a range of morphological and reproductive patterns including single-celled isogamous Chlamydomonas, 8- or 16-celled isogamous Gonium, 32-celled isogamous Yamagishiella, 32-celled anisogamous Eudorina, and >500-celled oogamous Volvox (Supplementary Fig. 1)3, 4; thus providing a rich source of material for comparative evolutionary studies of sexual cycles. Furthermore, sex-determining genes and sex chromosomal regions or mating-type loci (MT) have been studied extensively in both isogamous and sexually dimorphic volvocine green algae5,6,7,8,9,10,11. Therefore, this green algal group represents a unique model lineage for investigating the transition from isogamy to anisogamy based on molecular genetic data.
Charlesworth12 predicted that anisogamy could evolve from an isogamous genetic sex-determination system with two haploid mating types if an autosomally encoded gamete cell-size-determining gene with dimorphic alleles became loosely linked to the mating locus. Under conditions where anisogamy is favored, selection would act to promote closer linkage between the cell-size locus and mating locus, with an endpoint of complete linkage disequilibrium that might be achieved through suppression of recombination. A secondary outcome of generating a region of suppressed recombination near the mating locus would be potential for the MT region to capture additional genes through inversions or transposition, and to retain genes or alleles that are beneficial for their respective sex13,14,15. Thus, it is predicted that anisogamous or oogamous volvocine species will have relatively complex multi-genic MT loci compared with MT loci from isogamous species, and may contain additional genes that govern gamete size and/or other sexually selected traits. A previous comparative study of the Volvox carteri and Chlamydomonas reinhardtii MT loci was consistent with the idea of massive expansion of the mating locus in V. carteri with incorporation of many new genes, including at least one dimorphic size control gene, MAT39, a mammalian Retinoblastoma tumor suppressor (RB) homolog in the volvocine algae that regulates the cell cycle and maintains cell size in C. reinhardtii16. The V. carteri MAT3 homolog resides within MT and shows extensive divergence between male and female haplotypes9. Although this MAT3 allelic dimorphism was once hypothesized to be involved in establishing anisogamy9, a subsequent comparative analysis of the MAT3 sequences from various isogamous, anisogamous, and oogamous volvocine organisms indicated that the extensive MAT3 divergence in the V. carteri lineage appear to have occurred subsequent to the emergence of anisogamy, and therefore, MAT3 dimorphism may not be directly related to the origins of male–female gamete size dimorphism in volvocine algae17. While MT sequences of Gonium pectorale, an isogamous volvocine alga more closely related to oogamous V. carteri than to C. reinhardtii, has been also determined11, the lack of information on MT structures at intermediate isogamous and anisogamous stages in volvocine evolution leaves open important questions about how changes in MT structure and gene content relate to transitions from isogamy to anisogamy. Although Geng et al.10 showed that the V. carteri MID gene on its own can govern key aspects of gamete dimorphism, that study did not address the evolution of structural complexity in volvocine MT.
In this study, we focus on a more suitable combination for a comparative study of the transition from isogamy to anisogamy: two closely related intermediate volvocine species that bracket the transition from isogamy to anisogamy—isogamous Yamagishiella unicocca and anisogamous Eudorina sp., both of which share essentially the same vegetative (asexual) morphology (Fig. 1, Supplementary Fig. 1)18. We sequence the whole nuclear genomes of both mating types or sexes of Y. unicocca and Eudorina sp., and identify and characterize their respective MT regions. Contrary to expectations for increased MT size and gene content evolving during the transition to anisogamy, the Eudorina sp. male and female MT haplotypes are very small, 7 kb and 90 kb in size, respectively, with only two sex-specific genes and two pairs of gametologs (i.e., genes with alleles in both mating haplotypes). The highly reduced MT structures of Eudorina sp. indicates that anisogamy can evolve from isogamy without the addition of a gamete-size-control gene and without increased MT size and complexity.
Volvocine green algal phylogeny3, 4 and mating-type locus (MT) evolution. Phylogenetic relationships of volvocine algae are illustrated with vegetative morphology, gamete morphology18, and MT structures of Yamagishiella unicocca, Eudorina sp., and three other species previously studied (Chlamydomonas reinhardtii25, Gonium pectorale11, and Volvox carteri9). The bars at the flagellar bases of the isogametes indicate tubular mating structures (TMS)18. In C. reinhardtii, only the plus gametes possess TMS (red bar). In G. pectorale and Y. unicocca, both the plus and minus gametes possess TMS (red and blue bars, respectively)
Results
Mating-type loci (MT) of Yamagishiella and Eudorina
We de novo sequenced whole genomes from both sexes of isogamous Y. unicocca and anisogamous Eudorina sp., and obtained resultant assemblies as follows: plus strain of Y. unicocca (2012-1026-YU-F2-6/NIES-3982, assembly “YamagishiellaPlus_1.0”), total length 134,234,618 bp of 1461 contigs with N50 of 666,310 bp; minus strain of Y. unicocca (2012-1026-YU-F2-1/NIES-3983, assembly “YamagishiellaMinus_1.0”), total length 140,837,241 bp of 1897 contigs with N50 of 547,037 bp; male strain of Eudorina sp. (2010-623-F1-E3/NIES-4100, assembly “EudorinaMale_1.0”), total length 168,620,790 bp of 2471 contigs with N50 of 377,357 bp; and female strain of Eudorina sp. (2010-623-F1-E8/NIES-4018, assembly “EudorinaFemale_1.0”), total length 184,032,255 bp of 3180 contigs with N50 of 564,035 bp. The resultant assemblies contained dimorphic (rearranged) haplotype regions that composed MT (Supplementary Figs 2 and 3). Y. unicocca plus and minus MT regions were 268 and 165 kb in size, respectively, which is somewhat smaller than the MT regions of C. reinhardtii and G. pectorale (Fig. 1, Supplementary Table 1). Y. unicocca MT included two inverted syntenic blocs of 9 and 7 collinear gametologs, and a single gene, CRB1, that was outside these two syntenic blocs (Fig. 2a). Two conserved sex-limited genes, minus-limited MID and plus-limited FUS1 (Supplementary Figs 4–8), were also present. Eudorina sp. female and male MT haplotypes were the most highly reduced of any volvocine algal species described to date, measuring 90 and 7 kb in size, respectively (Fig. 2b, Supplementary Table 1). Only two gametologs, SPS1 (encoding a putative spermine syntase similar to ACAULIS5 required for stem elongation in flowering plants19) and UNC50 (encoding a membrane trafficking protein homolog conserved in eukaryotes20, 21), were found in Eudorina sp. MT (Fig. 2b). MID and FUS1 homologs were also found in the male and female MT, respectively, as sex-limited genes. The size and GC contents of whole genome and MT of the two algae with the other volvocine species so far analyzed5, 9, 11, 22,23,24,25,26 are summarized in Supplementary Table 1.
MT structures and molecular evolutionary analyses of MT gametologs in isogamous Y. unicocca and anisogamous Eudorina sp. a, b MT structures with sex-limited genes (FUS1, backed red; MID, blue) and gametologs of Y. unicocca (a) and Eudorina sp. (b) (accession nos. LC314412–LC314415). Red regions represent plus/female MT. Blue regions represent minus/male MT. Gray shading indicate a syntenic bloc. Open triangles in a indicate gaps between scaffolds of the de novo whole genome assembly (Supplementary Fig. 2). c Box-whisker plots comparing the distributions of synonymous (dS, green/left) and non-synonymous (dN, orange/right) substitution values for gametolog pairs found in rearranged regions of volvocine algal MT loci. Open dots are outliers from interquartile ranges except for those of Eudorina sp. which indicate two gametologs. d, e The dN/dS ratios of gametologs in rearranged (gray-shaded gene names) and flanking autosomal regions of Y. unicocca (d) and Eudorina sp. (e) MT. There are no prominently dimorphic gametologs under positive selection between sexes/mating types (dN/dS > 1)
Another striking finding in this study was the highly dynamic nature of MT haplotype structure that shows no continuity among the five volvocine MT regions (Fig. 1; Supplementary Data 1). Our data suggest relatively frequent turnover and haplotype reformation within MT, and explain the initially puzzling lack of continuity (e.g., evolutionary strata) observed between C. reinhardtii and V. carteri MT regions9, 27. Distribution of homologs of MT and MT-linked genes in the five volvocine algae (Supplementary Data 1) shows that most of these genes share the linkage to MT, except for G. pectorale in which majority of homologs of MT and MT-linked genes in other volvocine species are relocated to other genomic (autosomal) regions of G. pectorale as reported previously11. Our data further suggest that the differences in size, gene content, and structure we have observed between volvocine MT loci are largely uncoupled from the transition to anisogamy in this lineage.
Gametolog divergence rates for Y. unicocca and Eudorina sp. MT were compared to those of the other volvocine algae (Fig. 2c, d, e). Synonymous and non-synonymous substitutions (dS and dN) of gametologs between mating haplotypes in these two species were smaller than those of V. carteri. Within the Eudorina sp. and Y. unicocca gametologs, Eudorina sp. SPS1 and Y. unicocca CRB1 had relatively high dS and dN values (Fig. 2d, e). However, the dN/dS ratios of these two gametologs were low. Thus, we could not detect any strong patterns of selection or functional divergence between genes in the mating haplotypes of isogamous Y. unicocca and anisogamous Eudorina sp.
Overall, Y. unicocca and Eudorina sp. MT were smaller and simpler than MT regions from other volvocine species with only two conserved sex-limited genes (MID and FUS1) and no other clear examples of gametolog dimorphism. Some of the sex-limited gene homologs of oogamous V. carteri (VcFSI1f, VcHMG1f, and VcMTM0097) were conserved in the genomes of Y. unicocca and/or Eudorina sp. but they were autosomally encoded (Supplementary Table 2). The MTD1 gene is a minus-limited gene in C. reinhardtii and G. pectorale, while in Y. unicocca and Eudorina sp. MTD1 homologs are autosomal, but located nearby the MT region (Fig. 1, Supplementary Table 2). The presence of only two ancestral mating type genes (MID and FUS1) and lack of prominently divergent gametologs in highly reduced Eudorina sp. MT strongly disfavors the hypothesis that new sex-specific cell size-determining genes were acquired in MT during the emergence of anisogamy in the volvocine lineage12, 27.
Since female gametes of Eudorina and isogametes of Yamagishiella have no prominent morphological differentiation from their respective vegetative cells (Fig. 3a, b)18, the emergence of anisogamy from isogamy in the Yamagishiella/Eudorina-like ancestor appears to have been based on acquisition of the ability to form small male gametes. The present genome comparison demonstrated that the only male-limited gene in Eudorina MT is MID (Fig. 2b). The MID gene is present only in the minus/male MT haplotypes of heterothallic volvocine organisms28 and serves as a master regulator for mating-type determination in isogamous C. reinhardtii6. Recent molecular genetic approach in oogamous V. carteri showed that transformation of a female strain with MID enables the eggs (female gametes) to produce sperm packets (bundles of male gametes) and that a MID-knocked down male strain produces androgonidia (male reproductive initials) that can function as eggs10. Therefore, although there are an additional eight male-limited and five female-limited MT genes in the expanded MT of V. carteri9 that likely play a role in gamete fitness and early sexual development, none of them other than MID appear to have an essential role for the dimorphic gamete differentiation10. It follows that the evolution of males in volvocine algae might have resulted from altered function of the sex-determining protein MID or its target genes.
The isogamous FUS1 protein is a single-pass transmembrane protein that is present on the mating structure of plus gametes and required to recognize and adhere to minus gametes11, 29, 30. Previously, FUS1 homologs have been reported in isogamous C. reinhardtii29, 30 and G. pectorale11, but were not found in the oogamous V. carteri genome9, suggesting that FUS1 was lost at some point after the G. pectorale–V. carteri lineages split31. Our data showed that isogamous Y. unicocca and anisogamous Eudorina sp. both had a FUS1 homolog that was plus- or female-limited in MT, respectively (Fig. 2a, b; Supplementary Figs 4–8; Supplementary Note 1), and suggest that loss of FUS1 occurred subsequent to the Eudorina–V. carteri split.
Sex-related gene expressions in Yamagishiella and Eudorina
Expression programs of sex-related MT genes were evaluated for Y. unicocca and Eudorina sp. (Fig. 3). Unlike the case in C. reinhardtii and G. pectorale6, 8, but similar to V. carteri9, the MID mRNA was expressed constitutively in Y. unicocca minus cells and Eudorina sp. males. The gamete adhesion gene FUS1 from Y. unicocca plus was expressed only in gametes as is also the case for C. reinhardtii and G. pectorale FUS111, 29, but Eudorina sp. female FUS1 was most strongly expressed after mixing female with male gametes. The MTD1 gene in C. reinhardtii is found only in the minus MT haplotype, is expressed in gametes, and is required for efficient gametogenesis32, but appears to have lost its function in V. carteri where it is a pseudogene9. In Y. unicocca and Eudorina sp., MTD1 is autosomal with a copy in both sexes (Fig. 1, Supplementary Table 2). However, MTD1 expression in both Eudorina sp. and Y. unicocca was restricted to minus/male gametes (Fig. 3c, d), indicating a conserved function in gametogenesis in all genera except Volvox (Supplementary Fig. 9 and 10). The evolution of a highly reduced male MT in Eudorina sp. containing only the single male-limited MID gene might have been possible with minor changes that placed expression of MTD1 under mating-type control allowing it to reside outside of the MT locus.
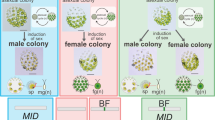
Sex induction and associated gene expression alternations in isogamous Y. unicocca and anisogamous Eudorina sp. a, b Asexual and sex-induced individuals of opposite sexes of Y. unicocca (plus/minus) (a) and Eudorina sp. (female/male) (b). Mating reactions (mixed, right panels) occurred after mixing induced cultures of the two sexes (middle panels). In Y. unicocca (a), clumping of the colonies and release of single-celled isogametes (arrowheads) were observed 1 h after mixing. In Eudorina sp. (b), sex induction treatment resulted in the formation of sperm packets and the packet dissociated into individual sperm that penetrated into a female colony (arrowheads) within 16 h after mixing. Scale bars, 20 µm. c, d Gene expression pattern of volvocine sex-limited genes in Y. unicocca (c) and Eudorina sp. (d). Semi-quantitative RT-PCR analyses were performed using the same cultures for a and b. All gels were run under the same experimental conditions44, and the cropped gel images are shown. Full-length gel images with size markers are presented in Supplementary Fig. 11
Conclusions
We have illuminated here the initial transition of the sex-determining chromosomal region or MT during the evolution of anisogamy, and shown that anisogamy can evolve without increased MT complexity. Only two sex-limited genes that encode the sex-determining factor MID (minus/male) and the gamete recognition factor FUS1 (plus/female) constitute the core of the MT haplotypes in both isogamous and anisogamous volvocine algae. Therefore, in this lineage the transition to anisogamy likely involved direct modification of the sex determination pathway controlled by MID rather than by acquisition of new gamete size control genes in MT. The presence of FUS1 not only in isogamous but also in anisogamous volvocine algae suggests that both systems share the FUS1-dependent gamete recognition mechanism. Future investigation of sex-related genes and their roles in anisogamy and gamete dialogues in volvocine algae holds great promise for understanding the dynamics of mating system evolution in eukaryotes.
Methods
Strains
For Yamagishiella, Y. unicocca strains 2012-1026-YU-F2-6 (NIES-3982, mating-type plus) and 2012-1026-YU-F2-1 (NIES-3983, mating-type minus) were used throughout the study. For the whole-genome sequencing of Eudorina, Eudorina sp. strains 2010-623-F1-E4 (NIES-3984, female) and 2010-623-F1-E2 (NIES-3985, male) were used. In other Eudorina experiments, two other sibling strains of the former two, Eudorina sp. strains 2010-623-F1-E8 (NIES-4018, female) and 2010-623-F1-E3 (NIES-4100, male), were used unless otherwise stated.
De novo whole genome assembly
Genomic DNAs were prepared according to the method of Miller et al.33. Whole-genome sequencing of plus and minus strains of Y. unicocca and male and female strains of Eudorina sp. were performed using PacBio and Illumina technologies as described previously34. Briefly, genomic DNA was sheared using a DNA shearing tube, g-TUBE (Covaris). Several 20 kb libraries for P5-C3 and P6-C4 sequencing were constructed and sequenced on SMRT cells in PacBio RS II (Pacific Biosciences). These reactions generated 2.5 M and 4.1 M sub reads (total bases: 17.2 Gb and 22.8 Gb, respectively) for plus and minus strains of Y. unicocca, respectively, and 2.6 M and 4.4 M sub reads (total bases: 15.6 Gb and 23.0 Gb, respectively) for male and female strains of Eudorina sp., respectively. Sequencing coverage was about 128x, 162x, 93x, and 125x based on the estimated genome size, respectively. In each of the four strains, PacBio reads were assembled de novo with HGAP3 assembler (Pacific Biosciences). Furthermore, genomic DNA was fragmented using a DNA Shearing System, S2 Focused-ultrasonicator (Covaris). Illumina paired-end libraries (insert sizes with 400 bp for plus and minus strains of Y. unicocca, 600 bp for male strain of Eudorina sp., and 400 bp for female strain of Eudorina sp.) were constructed using a TruSeq DNA Sample Prep Kit (Illumina) according to the manufacturer instructions. These libraries were sequenced using Illumina HiSeq 2000 and 2500 sequencers (230.5 M and 197.0 M reads with 150 bp read length for plus and minus strains of Y. unicocca, respectively, 251.2 M reads with 250 bp read length for male strain of Eudorina sp., and 165.3 M reads with 100 bp read length for female strain of Eudorina sp.). Total bases and sequencing coverage were 34.6 Gb (258x), 29.5 Gb (210x), 62.8 Gb (372x), and 16.5 Gb (89x), respectively. The Illumina data were then mapped against the PacBio assembly sequence using BWA-MEM Release 0.7.735 including error correction with the samtools/bcftools/vcfutils.pl program v0.1.19 (https://samtools.sourceforge.net/), ultimately giving a set of nuclear genome sequence. We performed a long-range scaffolding using paired-end Sanger sequences from 38,400 and 3840 fosmid clones of female and male strains of Eudorina sp., respectively (DRA: DRA004920, DRA002727, and DRA004919; whole genome assembly: BDSI01000001-BDSI01003180, BDSJ01000001-BDSJ01002471, BDSK01000001-BDSK01001897, BDSL01000001-BDSL01001461).
Sex-determining region identification
Candidate scaffolds for entire sex-determining regions (Y. unicocca plus: Scaffold0026/0199/0237; minus: Scaffold0005/0230/0253/0437/1431; Eudorina sp. Female: scaffold1024; male: scaffold1040) were screened as major significant matching subjects with more than three non-overlapping protein hits (cutoff maximum E-value: 1e−1036) by TBLASTN (NCBI) on de novo assemblies of Y. unicocca and Eudorina sp. with 80 proteins on V. carteri female MT (Genbank Acc. No. GU784915) as queries and then dotplot-analyzed between haplotypes of same species using YASS (https://bioinfo.lifl.fr/yass/index.php)37 to detect the rearranged genomic regions of MT.
Gene identification
We performed TBLASTN searches against the genome assembly databases of Y. unicocca and Eudorina sp. with the volvocine sex-limited proteins (Gonium pectorale FUS1, BAU6160711; G. pectorale MTD1, BAI4948738) as the queries, retrieved sequences with the highest similarity, and designed gene-specific primers (listed in Supplementary Table 3) based on these sequences. To identify the ORF sequences, polyadenylated mRNAs from each sample were isolated using Dynabeads Oligo (dT)25 (Thermo Fisher Scientific), reverse transcribed with Superscript III reverse transcriptase (Thermo Fisher Scientific), and amplified with KOD FX Neo DNA polymerase (Toyobo) and the gene-specific primers. To obtain the full-length cDNA sequences, 5′RACE and 3′RACE were performed using the GeneRacer kit (Thermo Fisher Scientific). The PCR products were directly sequenced, or first cloned into the pCR4Blunt-TOPO vector (Thermo Fisher Scientific) and then sequenced, using an ABI PRISM 3100 Genetic Analyzer (Thermo Fisher Scientific) with a BigDye Terminator cycle sequencing ready reaction kit, v.3.1 (Thermo Fisher Scientific). Full-length MID genes of Y. unicocca (minus strain NIES-1859) and Eudorina sp. (male strain NIES-2735) were determined using the degenerate PCR method7, 8.
Other gene models on MT scaffolds were predicted by Augustus39 with the C. reinhardtii parameter and then manually curated based on the similarity among C. reinhardtii and V. carteri gene models (JGI).
MT genome sequences harboring rearranged domains with sex-limited genes and gametologs in Y. unicocca and Eudorina sp. and the autosomal gene YuMTD1 are available under accession numbers LC314412–LC314416.
Preparation of asexual and sex-induced samples of Yamagishiella and Eudorina
Asexual samples of Y. unicocca were obtained by culturing the algae in screw-cap tubes (18 × 150 mm) containing ~11 ml AF-6 medium40, 41, on a 14-h light/10-h dark cycle (light intensity: 60–110 μmol m−2 s−1) at 23 °C for 3–5 days. To induce sexual reproduction of Y. unicocca, asexually growing algae (~0.2 ml) were inoculated into 11 ml “VTAC + soil extract medium” (VTAC medium41, 42 supplemented with 3%(v/v) soil extract (~0.5 mg of paddy soil suspended in 20 ml distilled water and autoclaved for 10 min)) in a tube, and cultured for 8 days under the same condition. The algal culture of each sex was then transferred into Petri dishes (60 mm × 15 mm), incubated for further 4 days, and used as a sex-induced Y. unicocca sample for the following analysis. “Mixed” sample of Y. unicocca were obtained by mixing the sex-induced samples (1 ml each) of both mating types in Petri dishes (30 mm × 10 mm), which were subsequently incubated for 3 h under the same condition, and used for semiquantitative RT-PCR.
Asexual samples of Eudorina sp. were obtained by culturing the algae in the screw-cap tubes containing ~10 ml SVM medium43, on a 14-h light/10-h dark cycle (light intensity: 180–320 μmol m−2 s−1) at 25 °C for 3 days. To induce sexual reproduction of Eudorina sp., ~0.4 mL of asexually grown algae in SVM medium were inoculated into Petri dishes (60 mm × 15 mm) containing ~11 ml “VTAC + soil extract medium” and cultured for 3 days under the same condition. The 11 ml culture of each sex was then transferred into Petri dishes (90 mm × 20 mm), diluted with twice volume of mating medium42, and cultured for further 8 h under the same condition to form a sex-induced male or female culture of Eudorina sp., in which formation of sperm packets was observed in the male strain (Fig. 3b). “Mixed” samples of Eudorina sp. were prepared by mixing the sex-induced female and male samples (5 ml each) in Petri dishes (60 mm × 15 mm), subsequently incubated for 16 h under the same condition, and used for the following analysis.
All microscopic images were acquired using a BX53 microscope (Olympus) equipped with differential interference contrast optics. The digital images were captured using a DP71 camera (Olympus) with DP controller software (Olympus), and their levels were adjusted with Adobe Photoshop CS6 (Adobe Systems Inc.).
Semiquantitative RT-PCR analysis
From Y. unicocca samples, polyadenylated mRNAs were isolated using Dynabeads Oligo (dT)25 and reverse transcribed as described above. From Eudorina samples, total RNAs were extracted with TRI reagent (Molecular Research Center), treated with DNase I (amplification grade; Thermo Fisher Scientific), and reverse transcribed with Superscript III reverse transcriptase and Oligo (dT)20 primer (Thermo Fisher Scientific). PCR reactions were performed with KOD FX Neo DNA polymerase (Toyobo). Primer sequences and PCR condition are listed in Supplementary Table 3 online. Under the conditions, all primer sets produced amplicons of the expected size and sequence (confirmed by direct sequencing). The PCR products were electrophoresed on 2% (w/v) agarose gels and stained with ethidium bromide44. The gel images were captured using a ChemiDoc XRS system (Bio-Rad) with Quantity One software (Bio-Rad) and their levels were adjusted as described above.
Molecular evolutionary analysis
Divergence scores of synonymous and non-synonymous substitutions between gametologs were computed using yn00 of the PAML4 package45; nonsynonymous and synonymous site divergence of aligned coding sequences of gametologs was calculated based on Yang and Nielsen46 with equal weighting between pathways, and the same codon frequency for all pairs11.
Data availability
Raw reads, genome assemblies, and annotations were deposited at DDBJ/EMBL/GenBank under the accessions as follows; DRA: DRA004920, DRA002727, and DRA004919; whole genome assembly: BDSI01000001-BDSI01003180, BDSJ01000001-BDSJ01002471, BDSK01000001-BDSK01001897, BDSL01000001-BDSL01001461; annotations: LC314412- LC314416. All the other data generated or analyzed during this study are included in this published article and its Supplementary information.
References
Darwin, C. R. The Descent of Man, and Selection in Relation to Sex 1st edn, Vol. 1 (John Murray, London, 1871).
Bell, G. The evolution of anisogamy. J. Theor. Biol. 73, 247–270 (1978).
Nozaki, H. Origin and evolution of the genera Pleodorina and Volvox (Volvocales). Biologia 58, 425–431 (2003).
Herron, M. D., Hackett, J. D., Aylward, F. O. & Michod, R. E. Triassic origin and early radiation of multicellular volvocine algae. Proc. Natl Acad. Sci. USA 106, 3254–3258 (2009).
Ferris, P. J. & Goodenough, U. W. The mating-type locus of Chlamydomonas reinhardtii contains highly rearranged DNA sequences. Cell 76, 1135–1145 (1994).
Ferris, P. J. & Goodenough, U. W. Mating type in Chlamydomonas is specified by mid, the minus-dominance gene. Genetics 146, 859–869 (1997).
Nozaki, H., Mori, T., Misumi, O., Matsunaga, S. & Kuroiwa, T. Males evolved from the dominant isogametic mating type. Curr. Biol. 16, R1018–R1020 (2006).
Hamaji, T. et al. Identification of the minus-dominance gene ortholog in the mating-type locus of Gonium pectorale. Genetics 178, 283–294 (2008).
Ferris, P. et al. Evolution of an expanded sex-determining locus in Volvox. Science 328, 351–354 (2010).
Geng, S., De Hoff, P. & Umen, J. G. Evolution of sexes from an ancestral mating-type specification pathway. PLoS Biol. 12, e1001904 (2014).
Hamaji, T. et al. Sequence of the Gonium pectorale mating locus reveals a complex and dynamic history of changes in volvocine algal mating haplotypes. G3 (Bethesda) 6, 1179–1189 (2016).
Charlesworth, B. The population genetics of anisogamy. J. Theor. Biol. 73, 347–357 (1978).
Bull, J. J. Sex chromosomes in haploid dioecy: a unique contrast to muller’s theory for diploid dioecy. Am. Nat. 112, 245–250 (1978).
Bachtrog, D. et al. Are all sex chromosomes created equal? Trends Genet. 27, 350–357 (2011).
Immler, S. & Otto, S. P. The evolution of sex chromosomes in organisms with separate haploid sexes. Evolution 69, 694–708 (2015).
Umen, J. G. & Goodenough, U. W. Control of cell division by a retinoblastoma protein homolog in Chlamydomonas. Genes Dev. 15, 1652–1661 (2001).
Hiraide, R. et al. The evolution of male-female sexual dimorphism predates the gender-based divergence of the mating locus gene MAT3/RB. Mol. Biol. Evol. 30, 1038–1040 (2013).
Nozaki, H. Morphology and evolution of sexual reproduction in the Volvocaceae (Chlorophyta). J. Plant Res. 109, 353–361 (1996).
Hanzawa, Y. et al. ACAULIS5, an Arabidopsis gene required for stem elongation, encodes a spermine synthase. EMBO J. 19, 4248–4256 (2000).
Fang, Z., Zhou, L., Jiang, S., Cao, L. & Yu, L. UNC50 prompts G1/S transition and proliferation in HCC by regulation of epidermal growth factor receptor trafficking. PLoS ONE 10, e0119338 (2015).
Selyunin, A. S., Iles, L. R., Bartholomeusz, G. & Mukhopadhyay, S. Genome-wide siRNA screen identifies UNC50 as a regulator of Shiga toxin 2 trafficking. J. Cell Biol. 216, 3249–3262 (2017).
Ferris, P. J., Armbrust, E. V. & Goodenough, U. W. Genetic structure of the mating-type locus of Chlamydomonas reinhardtii. Genetics 160, 181–200 (2002).
Merchant, S. S. et al. The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science 318, 245–250 (2007).
Prochnik, S. E. et al. Genomic analysis of organismal complexity in the multicellular green alga Volvox carteri. Science 329, 223–226 (2010).
De Hoff, P. L. et al. Species and population level molecular profiling reveals cryptic recombination and emergent asymmetry in the dimorphic mating locus of C. reinhardtii. PLoS Genet. 9, e1003724 (2013).
Hanschen, E. R. et al. The Gonium pectorale genome demonstrates co-option of cell cycle regulation during the evolution of multicellularity. Nat. Commun. 7, 11370 (2016).
Charlesworth, D. & Charlesworth, B. Evolutionary biology: the origins of two sexes. Curr. Biol. 20, R519–R521 (2010).
Yamamoto, K. et al. Molecular evolutionary analysis of a gender-limited MID ortholog from the homothallic species Volvox africanus with male and monoecious spheroids. PLoS ONE 12, e0180313 (2017).
Ferris, P. J., Woessner, J. P. & Goodenough, U. W. A sex recognition glycoprotein is encoded by the plus mating-type gene fus1 of Chlamydomonas reinhardtii. Mol. Biol. Cell 7, 1235–1248 (1996).
Misamore, M. J., Gupta, S. & Snell, W. J. The Chlamydomonas Fus1 protein is present on the mating type plus fusion organelle and required for a critical membrane adhesion event during fusion with minus gametes. Mol. Biol. Cell 14, 2530–2542 (2003).
Mori, T., Kawai-Toyooka, H., Igawa, T. & Nozaki, H. Gamete dialogs in green lineages. Mol. Plant 8, 1442–1454 (2015).
Lin, H. & Goodenough, U. W. Gametogenesis in the Chlamydomonas reinhardtii minus mating type is controlled by two genes, MID and MTD1. Genetics 176, 913–925 (2007).
Miller, S. M., Schmitt, R. & Kirk, D. L. Jordan, an active Volvox transposable element similar to higher plant transposons. Plant Cell 5, 1125–1138 (1993).
Hamaji, T. et al. Multiple independent changes in mitochondrial genome conformation in chlamydomonadalean algae. Genome Biol. Evol. 9, 993–999, https://doi.org/10.1093/gbe/evx060 (2017).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Noe, L. & Kucherov, G. YASS: enhancing the sensitivity of DNA similarity search. Nucleic Acids Res. 33, W540–W543 (2005).
Hamaji, T., Ferris, P. J., Nishii, I. & Nozaki, H. Identification of the minus mating-type specific gene MTD1 from Gonium pectorale (Volvocales, Chlorophyta). J. Phycol. 45, 1310–1314 (2009).
Hoff, K. J. & Stanke, M. WebAUGUSTUS—a web service for training AUGUSTUS and predicting genes in eukaryotes. Nucleic Acids Res. 41, W123–W128 (2013).
Kato, S. Laboratory culture and morphology of Colacium vesiculosum Ehrb. (Euglenophyceae). Jpn. J. Phycol. 30, 63–67 (1982).
Kasai, F. et al. NIES-collection list of strains, 8th edition. Jpn. J. Phycol. 57, 1–350 (2009).
Nozaki, H., Kuroiwa, H., Mita, T. & Kuroiwa, T. Pleodorina japonica sp. nov. (Volvocales, Chlorophyta) with bacteria-like endosymbionts. Phycologia 28, 252–267 (1989).
Kirk, D. L. & Kirk, M. M. Protein synthetic patterns during the asexual life cycle of Volvox carteri. Dev. Biol. 96, 493–506 (1983).
Kawai-Toyooka, H. et al. Sex-specific posttranslational regulation of the gamete fusogen GCS1 in the isogamous volvocine alga Gonium pectorale. Eukaryot. Cell 13, 648–656 (2014).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Yang, Z. & Nielsen, R. Estimating synonymous and nonsynonymous substitution rates under realistic evolutionary models. Mol. Biol. Evol. 17, 32–43 (2000).
Acknowledgements
We thank the staff of Comparative Genomics Laboratory at NIG for supporting genome sequencing. Computations were partially performed on the NIG supercomputer at ROIS National Institute of Genetics. This work was supported by a Grants-in-Aid for Scientific Research on Innovative Areas “Genome Science” (grant number 221S0002; to A.T. and A.F.), Scientific Research (A) (grant number 16H02518; to H.Nozaki), Research Activity Startup grants (grant number 16H06734 to T.H.), Scientific Research (C) (grant number 17K07510 to H.K.-T.), Grant-in-Aid for Scientific Research on Innovative Areas (grant number 17H05840 to T.H.) from MEXT/JSPS KAKENHI, and National Institutes of Health (grant number GM 078376 to J.G.U.).
Author information
Authors and Affiliations
Contributions
Conceived the study: H.Nozaki. Designed the study: T.H., H.K.-T., H.Nozaki. Prepared genomic DNA: M.S. Performed whole-genome sequencing and assembly: H.Noguchi., Y.M. A.T., A.F. Performed the experiments: T.H., H.K.-T., H.U. Analyzed the data: T.H., H.K.-T., H.Nozaki. Contributed materials: S.-y.M. Wrote and edited the manuscript: T.H., H.K.-T., A.T., J.G.U., H.Nozaki.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hamaji, T., Kawai-Toyooka, H., Uchimura, H. et al. Anisogamy evolved with a reduced sex-determining region in volvocine green algae. Commun Biol 1, 17 (2018). https://doi.org/10.1038/s42003-018-0019-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-018-0019-5
This article is cited by
-
Reorganization of the ancestral sex-determining regions during the evolution of trioecy in Pleodorina starrii
Communications Biology (2023)
-
Gene loss during a transition to multicellularity
Scientific Reports (2023)
-
Sex-linked deubiquitinase establishes uniparental transmission of chloroplast DNA
Nature Communications (2022)
-
The first draft genome of feather grasses using SMRT sequencing and its implications in molecular studies of Stipa
Scientific Reports (2021)
-
Genome sequencing of the multicellular alga Astrephomene provides insights into convergent evolution of germ-soma differentiation
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.