Abstract
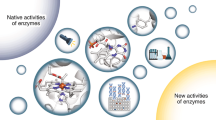
The emergence of robust methods to expand the genetic code allows incorporation of non-canonical amino acids into the polypeptide chain of proteins, thus making it possible to introduce unnatural chemical functionalities in enzymes. In this Perspective, we show how this powerful methodology is used to create enzymes with improved and novel, even new-to-nature, catalytic activities. We provide an overview of the current state of the art, and discuss the potential benefits of developing and using enzymes with genetically encoded non-canonical amino acids compared with enzymes containing only canonical amino acids.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wohlgemuth, R. Biocatalysis-key to sustainable industrial chemistry. Curr. Opin. Biotechnol. 21, 713–724 (2010).
Reetz, M. T. Biocatalysis in organic chemistry and biotechnology: past, present, and future. J. Am. Chem. Soc. 135, 12480–12496 (2013).
Arnold, F. H. Directed evolution: bringing new chemistry to life. Angew. Chem. Int. Ed. 57, 4143–4148 (2018).
Okeley, N. M. & van der Donk, W. A. Novel cofactors via post-translational modifications of enzyme active sites. Chem. Biol. 7, 159–171 (2000).
Liu, C. C. & Schultz, P. G. Adding new chemistries to the genetic code. Annu. Rev. Biochem. 79, 413–444 (2010).
Chin, J. W. Expanding and reprogramming the genetic code. Nature 550, 53–60 (2017).
Young, D. D. & Schultz, P. G. Playing with the molecules of life. ACS Chem. Biol. 13, 854–870 (2018).
Agostini, F. et al. Biocatalysis with unnatural amino acids: enzymology meets xenobiology. Angew. Chem. Int. Ed. 56, 9680–9703 (2017).
Reetz, M. T., Kahakeaw, D. & Lohmer, R. Addressing the numbers problem in directed evolution. ChemBioChem 9, 1797–1804 (2008).
Currin, A., Swainston, N., Day, P. J. & Kell, D. B. Synthetic biology for the directed evolution of protein biocatalysts: navigating sequence space intelligently. Chem. Soc. Rev. 44, 1172–1239 (2015).
Li, G., Dong, Y. & Reetz, M. T. Can machine learning revolutionize directed evolution of selective enzymes? Adv. Synth. Catal. 361, 2377–2386 (2019).
Wu, Z., Kan, S. B. J., Lewis, R. D., Wittmann, B. J. & Arnold, F. H. Machine learning-assisted directed protein evolution with combinatorial libraries. Proc. Natl Acad. Sci. USA 116, 8852–8858 (2019).
Currin, A. et al. GeneORator: an effective strategy for navigating protein sequence space more efficiently through boolean OR-type DNA libraries. ACS Synth. Biol. 8, 1371–1378 (2019).
Srinivasan, G., James, C. M. & Krzycki, J. A. Pyrrolysine encoded by UAG in Archaea: charging of a UAG-decoding specialized tRNA. Science 296, 1459–1462 (2002).
Böck, A. et al. Selenocysteine: the 21st amino acid. Mol. Microbiol. 5, 515–520 (1991).
Crick, F. H., Griffith, J. S. & Orgel, L. E. Codes without commas. Proc. Natl Acad. Sci. USA 43, 416–421 (1957).
Johnson, J. A., Lu, Y. Y., Van Deventer, J. A. & Tirrell, D. A. Residue-specific incorporation of non-canonical amino acids into proteins: recent developments and applications. Curr. Opin. Chem. Biol. 14, 774–780 (2010).
Dumas, A., Lercher, L., Spicer, C. D. & Davis, B. G. Designing logical codon reassignment-expanding the chemistry in biology. Chem. Sci. 6, 50–69 (2015).
Munier, R. & Cohen, G. N. Incorporation of structural analogues of amino acids into bacterial proteins during their synthesis in vivo. Biochim. Biophys. Acta 31, 378–391 (1959).
Wang, L., Brock, A., Herberich, B. & Schultz, P. G. Expanding the genetic code of Escherichia coli. Science 292, 498–500 (2001).
Blight, S. K. et al. Direct charging of tRNACUA with pyrrolysine in vitro and in vivo. Nature 431, 333–335 (2004).
Wan, W., Tharp, J. M. & Liu, W. R. Pyrrolysyl-tRNA synthetase: an ordinary enzyme but an outstanding genetic code expansion tool. Biochim. Biophys. Acta 1844, 1059–1070 (2014).
Li, J. C., Liu, T., Wang, Y., Mehta, A. P. & Schultz, P. G. Enhancing protein stability with genetically encoded noncanonical amino acids. J. Am. Chem. Soc. 140, 15997–16000 (2018).
Yu, Y., Hu, C., Xia, L. & Wang, J. Artificial metalloenzyme design with unnatural amino acids and non-native cofactors. ACS Catal. 8, 1851–1863 (2018).
Yang, H., Srivastava, P., Zhang, C. & Lewis, J. C. A general method for artificial metalloenzyme formation through strain-promoted azide-alkyne cycloaddition. ChemBioChem 15, 223–227 (2014).
Srivastava, P., Yang, H., Ellis-Guardiola, K. & Lewis, J. C. Engineering a dirhodium artificial metalloenzyme for selective olefin cyclopropanation. Nat. Commun. 6, 7789 (2015).
Yang, H. et al. Evolving artificial metalloenzymes via random mutagenesis. Nat. Chem. 10, 318–324 (2018).
Pauling, L. Nature of forces between large molecules of biological interest. Nature 161, 707–709 (1948).
Lee, J. & Goodey, N. M. Catalytic contributions from remote regions of enzyme structure. Chem. Rev. 111, 7595–7624 (2011).
Dominguez, M. A., Thornton, K. C., Melendez, M. G. & Dupureur, C. M. Differential effects of isomeric incorporation of fluorophenylalanines into PvuII endonuclease. Proteins Struct. Funct. Genet. 45, 55–61 (2001).
Deepankumar, K. et al. Enhancing thermostability and organic solvent tolerance of ω-transaminase through global incorporation of fluorotyrosine. Adv. Synth. Catal. 356, 993–998 (2014).
Cirino, P. C., Tang, Y., Takahashi, K., Tirrell, D. A. & Arnold, F. H. Global incorporation of norleucine in place of methionine in cytochrome P450 BM-3 heme domain increases peroxygenase activity. Biotechnol. Bioeng. 83, 729–734 (2003).
Hoesl, M. G. et al. Lipase congeners designed by genetic code engineering. ChemCatChem 3, 213–221 (2011).
Jackson, J. C., Duffy, S. P., Hess, K. R. & Mehl, R. A. Improving nature’s enzyme active site with genetically encoded unnatural amino acids. J. Am. Chem. Soc. 128, 11124–11127 (2006).
Ma, H., Yang, X., Lu, Z., Liu, N. & Chen, Y. The ‘gate keeper’ role of Trp222 determines the enantiopreference of diketoreductase toward 2-chloro-1-phenylethanone. PLoS ONE 9, e103792 (2014).
Kolev, J. N., Zaengle, J. M., Ravikumar, R. & Fasan, R. Enhancing the efficiency and regioselectivity of P450 oxidation catalysts by unnatural amino acid mutagenesis. ChemBioChem 15, 1001–1010 (2014).
Morikubo, N. et al. Cation-π interaction in the polyolefin cyclization cascade uncovered by incorporating unnatural amino acids into the catalytic sites of squalene cyclase. J. Am. Chem. Soc. 128, 13184–13194 (2006).
Ugwumba, I. N. et al. Improving a natural enzyme activity through incorporation of unnatural amino acids. J. Am. Chem. Soc. 133, 326–333 (2011).
Xiao, H. et al. Exploring the potential impact of an expanded genetic code on protein function. Proc. Natl Acad. Sci. USA 112, 6961–6966 (2015).
Stubbe, J. A., Nocera, D. G., Yee, C. S. & Chang, M. C. Y. Radical initiation in the class I ribonucleotide reductase: long-range proton-coupled electron transfer? Chem. Rev. 103, 2167–2201 (2003).
Yee, C. S., Chang, M. C. Y., Ge, J., Nocera, D. G. & Stubbe, J. 2,3-Difluorotyrosine at position 356 of ribonucleotide reductase R2: a probe of long-range proton-coupled electron transfer. J. Am. Chem. Soc. 125, 10506–10507 (2003).
Seyedsayamdost, M. R., Reece, S. Y., Nocera, D. G. & Stubbe, J. A. Mono-, di-, tri-, and tetra-substituted fluorotyrosines: new probes for enzymes that use tyrosyl radicals in catalysis. J. Am. Chem. Soc. 128, 1569–1579 (2006).
Yokoyama, K., Uhlin, U. & Stubbe, J. Site-specific incorporation of 3-nitrotyrosine as a probe of pK a perturbation of redox-active tyrosines in ribonucleotide reductase. J. Am. Chem. Soc. 132, 8385–8397 (2010).
Minnihan, E. C., Young, D. D., Schultz, P. G. & Stubbe, J. Incorporation of fluorotyrosines into ribonucleotide reductase using an evolved, polyspecific aminoacyl-tRNA synthetase. J. Am. Chem. Soc. 133, 15942–15945 (2011).
Minnihan, E. C., Nocera, D. G. & Stubbe, J. Reversible, long-range radical transfer in E. coli class Ia ribonucleotide reductase. Acc. Chem. Res. 46, 2524–2535 (2013).
Oyala, P. H. et al. Biophysical characterization of fluorotyrosine probes site-specifically incorporated into enzymes: E. coli ribonucleotide reductase as an example. J. Am. Chem. Soc. 138, 7951–7964 (2016).
Yu, Y. et al. Defining the role of tyrosine and rational tuning of oxidase activity by genetic incorporation of unnatural tyrosine analogs. J. Am. Chem. Soc. 137, 4594–4597 (2015).
Yu, Y. et al. Significant improvement of oxidase activity through the genetic incorporation of a redox-active unnatural amino acid. Chem. Sci. 6, 3881–3885 (2015).
Chen, L. et al. Use of a tyrosine analogue to modulate the two activities of a nonheme iron enzyme OvoA in ovothiol biosynthesis, cysteine oxidation versus oxidative C–S bond formation. J. Am. Chem. Soc. 140, 4604–4612 (2018).
Chen, L. et al. Mechanistic studies of a nonheme iron enzyme OvoA in ovothiol biosynthesis using a tyrosine analogue, 2-Amino-3-(4-hydroxy-3-(methoxyl) phenyl) propanoic acid (MeOTyr). ACS Catal. 9, 253–258 (2019).
Liu, X. et al. Significant increase of oxidase activity through the genetic incorporation of a tyrosine-histidine cross-link in a myoglobin model of heme-copper oxidase. Angew. Chem. Int. Ed. 51, 4312–4316 (2012).
Jones, L. H., Narayanan, A. & Hett, E. C. Understanding and applying tyrosine biochemical diversity. Mol. Biosyst. 10, 952–969 (2014).
Zhou, Q. et al. Probing the function of the Tyr-Cys cross-link in metalloenzymes by the genetic incorporation of 3-methylthiotyrosine. Angew. Chem. Int. Ed. 52, 1203–1207 (2013).
Liu, X. et al. A genetically encoded photosensitizer protein facilitates the rational design of a miniature photocatalytic CO2-reducing enzyme. Nat. Chem. 10, 1201–1206 (2018).
Romero, N. A. & Nicewicz, D. A. Organic photoredox catalysis. Chem. Rev. 116, 10075–10166 (2016).
Givens, R. S., Weber, J. F. W., Jung, A. H. & Park, C.-H. New photoprotecting groups: desyl and p-hydroxyphenacyl phosphate and carboxylate esters. Methods Enzymol. 291, 1–29 (1998).
Deiters, A., Groff, D., Ryu, Y., Xie, J. & Schultz, P. G. A genetically encoded photocaged tyrosine. Angew. Chem. Int. Ed. 45, 2728–2731 (2006).
Wu, N., Deiters, A., Cropp, T. A., King, D. & Schultz, P. G. A genetically encoded photocaged amino acid. J. Am. Chem. Soc. 126, 14306–14307 (2004).
Luo, J., Torres-Kolbus, J., Liu, J. & Deiters, A. Genetic encoding of photocaged tyrosines with improved light-activation properties for the optical control of protease function. ChemBioChem 18, 1442–1447 (2017).
Lemke, E. A., Summerer, D., Geierstanger, B. H., Brittain, S. M. & Schultz, P. G. Control of protein phosphorylation with a genetically encoded photocaged amino acid. Nat. Chem. Biol. 3, 769–772 (2007).
Nguyen, D. P. et al. Genetic encoding of photocaged cysteine allows photoactivation of TEV protease in live mammalian cells. J. Am. Chem. Soc. 136, 2240–2243 (2014).
Beharry, A. A. & Woolley, G. A. Azobenzene photoswitches for biomolecules. Chem. Soc. Rev. 40, 4422–4437 (2011).
Hoppmann, C. et al. Genetically encoding photoswitchable click amino acids in Escherichia coli and mammalian cells. Angew. Chem. Int. Ed. 53, 3932–3936 (2014).
Tsai, Y.-H., Essig, S., James, J. R., Lang, K. & Chin, J. W. Selective, rapid and optically switchable regulation of protein function in live mammalian cells. Nat. Chem. 7, 554–561 (2015).
Luo, J., Samanta, S., Convertino, M., Dokholyan, N. V. & Deiters, A. Reversible and tunable photoswitching of protein function through genetic encoding of azobenzene amino acids in mammalian cells. ChemBioChem 19, 2178–2185 (2018).
Schwizer, F. et al. Artificial metalloenzymes: reaction scope and optimization strategies. Chem. Rev. 118, 142–231 (2018).
Hayashi, T., Hilvert, D. & Green, A. P. Engineered metalloenzymes with non-canonical coordination environments. Chem. Eur. J. 24, 11821–11830 (2018).
Lee, H. S. & Schultz, P. G. Biosynthesis of a site-specific DNA cleaving protein. J. Am. Chem. Soc. 130, 13194–13195 (2008).
Drienovská, I., Rioz-Martínez, A., Draksharapu, A. & Roelfes, G. Novel artificial metalloenzymes by in vivo incorporation of metal-binding unnatural amino acids. Chem. Sci. 6, 770–776 (2015).
Roelfes, G. LmrR: a privileged scaffold for artificial metalloenzymes. Acc. Chem. Res. 52, 545–556 (2019).
Bersellini, M. & Roelfes, G. Multidrug resistance regulators (MDRs) as scaffolds for the design of artificial metalloenzymes. Org. Biomol. Chem. 15, 3069–3073 (2017).
Drienovská, I. et al. Design of an enantioselective artificial metallo-hydratase enzyme containing an unnatural metal-binding amino acid. Chem. Sci. 8, 7228–7235 (2017).
Ségaud, N., Drienovská, I., Chen, J., Browne, W. R. & Roelfes, G. Artificial metalloproteins for binding and stabilization of a semiquinone radical. Inorg. Chem. 56, 13293–13299 (2017).
Mills, J. H. et al. Computational design of an unnatural amino acid dependent metalloprotein with atomic level accuracy. J. Am. Chem. Soc. 135, 13393–13399 (2013).
Mills, J. H. et al. Computational design of a homotrimeric metalloprotein with a trisbipyridyl core. Proc. Natl Acad. Sci. USA 113, 15012–15017 (2016).
Green, A. P., Hayashi, T., Mittl, P. R. E. & Hilvert, D. A chemically programmed proximal ligand enhances the catalytic properties of a heme enzyme. J. Am. Chem. Soc. 138, 11344–11352 (2016).
Pott, M. et al. A noncanonical proximal heme ligand affords an efficient peroxidase in a globin fold. J. Am. Chem. Soc. 140, 1535–1543 (2018).
Hayashi, T. et al. Capture and characterization of a reactive haem–carbenoid complex in an artificial metalloenzyme. Nat. Catal. 1, 578–584 (2018).
Moore, E. J. & Fasan, R. Effect of proximal ligand substitutions on the carbene and nitrene transferase activity of myoglobin. Tetrahedron 75, 2357–2363 (2019).
Schmidt, B., Selmer, T., Ingendoh, A. & Figurat, Kvon A novel amino acid modification in sulfatases that is defective in multiple sulfatase deficiency. Cell 82, 271–278 (1995).
Schwede, T. F., Rétey, J. & Schulz, G. E. Crystal structure of histidine ammonia-lyase revealing a novel polypeptide modification as the catalytic electrophile. Biochemistry 38, 5355–5361 (1999).
Drienovská, I., Mayer, C., Dulson, C. & Roelfes, G. A designer enzyme for hydrazone and oxime formation featuring an unnatural catalytic aniline residue. Nat. Chem. 10, 946–952 (2018).
Mayer, C., Dulson, C., Reddem, E., Thunnissen, A.-M. W. H. & Roelfes, G. Directed evolution of a designer enzyme featuring an unnatural catalytic amino acid. Angew. Chem. Int. Ed. 58, 2083–2087 (2019).
Burke, A. J. et al. Design and evolution of an enzyme with a non-canonical organocatalytic mechanism. Nature 570, 219–223 (2019).
Lindström, U. M. Stereoselective organic reactions in water. Chem. Rev. 102, 2751–2772 (2002).
Lipshutz, B. H., Ghorai, S. & Cortes-Clerget, M. The hydrophobic effect applied to organic synthesis: recent synthetic chemistry “in water”. Chem. Eur. J. 24, 6672–6695 (2018).
van der Helm, M. P., Klemm, B. & Eelkema, R. Organocatalysis in aqueous media. Nat. Rev. Chem. 3, 491–508 (2019).
Jeschek, M., Panke, S. & Ward, T. R. Artificial metalloenzymes on the verge of new-to-nature metabolism. Trends Biotechnol. 36, 60–72 (2018).
Almhjell, P. J., Boville, C. E. & Arnold, F. H. Engineering enzymes for noncanonical amino acid synthesis. Chem. Soc. Rev. 47, 8980–8997 (2018).
Mehl, R. A. et al. Generation of a bacterium with a 21 amino acid genetic code. J. Am. Chem. Soc. 125, 935–939 (2003).
Brustad, E., Bushey, M. L., Brock, A., Chittuluru, J. & Schultz, P. G. A promiscuous aminoacyl-tRNA synthetase that incorporates cysteine, methionine, and alanine homologs into proteins. Bioorg. Med. Chem. Lett. 18, 6004–6006 (2008).
Young, D. D. et al. An evolved aminoacyl-tRNA synthetase with atypical polysubstrate specificity. Biochemistry 50, 1894–1900 (2011).
Wang, Y.-S., Fang, X., Wallace, A. L., Wu, B. & Liu, W. R. A rationally designed pyrrolysyl-tRNA synthetase mutant with a broad substrate spectrum. J. Am. Chem. Soc. 134, 2950–2953 (2012).
Mukai, T. et al. Codon reassignment in the Escherichia coli genetic code. Nucleic Acids Res. 38, 8188–8195 (2010).
Johnson, D. B. F. et al. RF1 knockout allows ribosomal incorporation of unnatural amino acids at multiple sites. Nat. Chem. Biol. 7, 779–786 (2011).
Johnson, D. B. F. et al. Release factor one is nonessential in Escherichia coli. ACS Chem. Biol. 7, 1337–1344 (2012).
Lajoie, M. J. et al. Genomically recoded organisms expand biological functions. Science 342, 357–360 (2013).
Mukai, T. et al. Highly reproductive Escherichia coli cells with no specific assignment to the UAG codon. Sci. Rep. 5, 9699 (2015).
Allen, A. E. & MacMillan, D. W. C. Synergistic catalysis: a powerful synthetic strategy for new reaction development. Chem. Sci. 3, 633–658 (2012).
Kiss, G., Çelebi-Ölçüm, N., Moretti, R., Baker, D. & Houk, K. N. Computational enzyme design. Angew. Chem. Int. Ed. 52, 5700–5725 (2013).
Acknowledgements
G.R. acknowledges financial support from the Netherlands Organisation for Scientific Research (NWO, Vici grant 724.013.003) and the Ministry of Education Culture and Science (Gravitation programme no. 024.001.035).
Author information
Authors and Affiliations
Contributions
I.D. researched data for the article. I.D. and G.R. wrote the article and both authors contributed to the discussion, reviewing and editing of the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Drienovská, I., Roelfes, G. Expanding the enzyme universe with genetically encoded unnatural amino acids. Nat Catal 3, 193–202 (2020). https://doi.org/10.1038/s41929-019-0410-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41929-019-0410-8
This article is cited by
-
Cellular decoding for non-natural peptides
Nature Chemistry (2023)
-
Unnatural activities and mechanistic insights of cytochrome P450 PikC gained from site-specific mutagenesis by non-canonical amino acids
Nature Communications (2023)
-
Rebooting life: engineering non-natural nucleic acids, proteins and metabolites in microorganisms
Microbial Cell Factories (2022)
-
Therapeutic peptides: current applications and future directions
Signal Transduction and Targeted Therapy (2022)
-
Synthetic prodrug design enables biocatalytic activation in mice to elicit tumor growth suppression
Nature Communications (2022)