Abstract
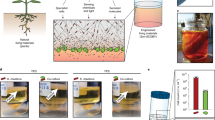
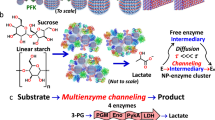
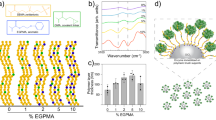
Cooperative enzyme catalysis in nature has long inspired the application of engineered multi-enzyme assemblies for industrial biocatalysis. Despite considerable interest, efforts to harness the activity of cell-surface displayed multi-enzyme assemblies have been based on trial and error rather than rational design due to a lack of quantitative tools. In this study, we have developed a quantitative approach to whole-cell biocatalyst characterization, enabling a comprehensive study of how yeast-surface displayed multi-enzyme assemblies form. Here we show that the multi-enzyme assembly efficiency is limited by molecular crowding on the yeast-cell surface, and that maximizing enzyme density is the most important parameter for enhancing cellulose hydrolytic performance. Interestingly, we also observed that proximity effects are only synergistic when the average inter-enzyme distance is greater than ~130 nm. The findings and the quantitative approach developed in this work should help to advance the field of biocatalyst engineering from trial and error to rational design.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The data that support the plots within this paper and other findings of this study are available from corresponding author F.W. upon reasonable request.
References
Alper, H. & Stephanopoulos, G. Engineering for biofuels: exploiting innate microbial capacity or importing biosynthetic potential? Nat. Rev. Microbiol. 7, 715–723 (2009).
Wheeldon, I. et al. Substrate channelling as an approach to cascade reactions. Nat. Chem. 8, 299–309 (2016).
Avalos, J. L., Fink, G. R. & Stephanopoulos, G. Compartmentalization of metabolic pathways in yeast mitochondria improves the production of branched-chain alcohols. Nat. Biotechnol. 31, 335–341 (2013).
Dodds, D. R. & Gross, R. A. Chemicals from biomass. Science 318, 1250–1251 (2007).
Gupta, N. K., Fukuoka, A. & Nakajima, K. Amorphous Nb2O5 as a selective and reusable catalyst for furfural production from xylose in biphasic water and toluene. ACS Catal. 7, 2430–2436 (2017).
Zinoviev, S. et al. Next-generation biofuels: survey of emerging technologies and sustainability issues. ChemSusChem 3, 1106–1133 (2010).
Huber, G. W., Iborra, S. & Corma, A. Synthesis of transportation fuels from biomass: chemistry, catalysts and engineering. Chem. Rev. 106, 4044–4098 (2006).
Isikgor, F. H. & Becer, C. R. Lignocellulosic biomass: a sustainable platform for the production of bio-based chemicals and polymers. Polym. Chem. 6, 4497–4559 (2015).
Caes, B. R., Teixeira, R. E., Knapp, K. G. & Raines, R. T. Biomass to furanics: renewable routes to chemicals and fuels. ACS Sustain. Chem. Eng. 3, 2591–2605 (2015).
Himmel, M. E. & Bayer, E. A. Lignocellulose conversion to biofuels: current challenges, global perspectives. Curr. Opin. Biotechnol. 20, 316–317 (2009).
Yeh, T. M. et al. Hydrothermal catalytic production of fuels and chemicals from aquatic biomass. J. Chem. Technol. Biotechnol. 88, 13–24 (2013).
Wen, F., Nair, N. U. & Zhao, H. Protein engineering in designing tailored enzymes and microorganisms for biofuels production. Curr. Opin. Biotechnol. 20, 412–419 (2009).
Horn, S. J., Vaaje-Kolstad, G., Westereng, B. & Eijsink, V. G. Novel enzymes for the degradation of cellulose. Biotechnol. Biofuels 5, 45 (2012).
Percival Zhang, Y.-H., Himmel, M. E. & Mielenz, J. R. Outlook for cellulase improvement: screening and selection strategies. Biotechnol. Adv. 24, 452–481 (2006).
Schwarz, W. H. The cellulosome and cellulose degradation by anaerobic bacteria. Appl. Microbiol. Biotechnol. 56, 634–649 (2001).
Lynd, L. R., Weimer, P. J., van Zyl, W. H. & Pretorius, I. S. Microbial cellulose utilization: fundamentals and biotechnology. Microbiol. Mol. Biol. Rev. 66, 739–739 (2002).
Shang, B. Z. & Chu, J. W. Kinetic modeling at single-molecule resolution elucidates the mechanisms of cellulase synergy. ACS Catal. 4, 2216–2225 (2014).
Percival Zhang, Y. H. et al. A transition from cellulose swelling to cellulose dissolution by o-phosphoric acid: evidence from enzymatic hydrolysis and supramolecular structure. Biomacromolecules 7, 644–648 (2006).
Artzi, L., Bayer, E. A. & Moraïs, S. Cellulosomes: bacterial nanomachines for dismantling plant polysaccharides. Nat. Rev. Microbiol. 15, 83–95 (2016).
Fierobe, H.-P. et al. Action of designer cellulosomes on homogeneous versus complex substrates: controlled incorporation of three distinct enzymes into a defined trifunctional scaffoldin. J. Biol. Chem. 280, 16325–16334 (2005).
Moraïs, S. et al. Cellulase–xylanase synergy in designer cellulosomes for enhanced degradation of a complex cellulosic substrate. MBio 1, e00285–10 (2010).
Vazana, Y., Moraïs, S., Barak, Y., Lamed, R. & Bayer, E. A. Interplay between Clostridium thermocellum family 48 and family 9 cellulases in cellulosomal versus noncellulosomal states. Appl. Environ. Microbiol. 76, 3236–3243 (2010).
Fierobe, H. P. et al. Degradation of cellulose substrates by cellulosome chimeras: substrate targeting versus proximity of enzyme components. J. Biol. Chem. 277, 49621–49630 (2002).
Sun, J., Wen, F., Si, T., Xu, J. H. & Zhao, H. Direct conversion of xylan to ethanol by recombinant Saccharomyces cerevisiae strains displaying an engineered minihemicellulosome. Appl. Environ. Microbiol. 78, 3837–3845 (2012).
Smith, M. R., Khera, E. & Wen, F. Engineering novel and improved biocatalysts by cell surface display. Ind. Eng. Chem. Res. 54, 4021–4032 (2015).
You, C., Zhang, X.-Z., Sathitsuksanoh, N., Lynd, L. R. & Zhang, Y.-H. P. Enhanced microbial utilization of recalcitrant cellulose by an ex vivo cellulosome–microbe complex. Appl. Environ. Microbiol. 78, 1437–1444 (2012).
Wieczorek, A. S. & Martin, V. J. J. Engineering the cell surface display of cohesins for assembly of cellulosome-inspired enzyme complexes on Lactococcus lactis. Microb. Cell Fact. 9, 69 (2010).
Lu, Y., Zhang, Y.-H. P. & Lynd, L. R. Enzyme–microbe synergy during cellulose hydrolysis by Clostridium thermocellum. Proc. Natl Acad. Sci. USA 103, 16165–16169 (2006).
Wen, F., Sun, J. & Zhao, H. Yeast surface display of trifunctional minicellulosomes for simultaneous saccharification and fermentation of cellulose to ethanol. Appl. Environ. Microbiol. 76, 1251–1260 (2010).
Liang, Y., Si, T., Ang, E. L. & Zhao, H. An engineered pentafunctional minicellulosome for simultaneous saccharification and ethanol fermentation in Saccharomyces cerevisiae. Appl. Environ. Microbiol. 80, 6677–6684 (2014).
Tsai, S.-L., Oh, J., Singh, S., Chen, R. & Chen, W. Functional assembly of minicellulosomes on the Saccharomyces cerevisiae cell surface for cellulose hydrolysis and ethanol production. Appl. Environ. Microbiol. 75, 6087–6093 (2009).
Bugada, L. F., Smith, M. R. & Wen, F. Engineering spatially organized multienzyme assemblies for complex chemical transformation. ACS Catal. 8, 7898–7906 (2018).
Tsai, S.-L. L., DaSilva, N. A. & Chen, W. Functional display of complex cellulosomes on the yeast surface via adaptive assembly. ACS Synth. Biol. 2, 14–21 (2013).
Fan, L.-H., Zhang, Z.-J., Yu, X.-Y., Xue, Y.-X. & Tan, T.-W. Self-surface assembly of cellulosomes with two miniscaffoldins on Saccharomyces cerevisiae for cellulosic ethanol production. Proc. Natl Acad. Sci. USA 109, 13260–13265 (2012).
Matano, Y., Hasunuma, T. & Kondo, A. Display of cellulases on the cell surface of Saccharomyces cerevisiae for high yield ethanol production from high-solid lignocellulosic biomass. Bioresour. Technol. 108, 128–133 (2012).
Meyer, A. et al. Optimization of a whole-cell biocatalyst by employing genetically encoded product sensors inside nanolitre reactors. Nat. Chem. 7, 673–678 (2015).
Bligaard, T. et al. Toward benchmarking in catalysis science: best practices, challenges and opportunities. ACS Catal. 6, 2590–2602 (2016).
Boder, E. T. & Wittrup, K. D. Yeast surface display for screening combinatorial polypeptide libraries. Nat. Biotechnol. 15, 553–557 (1997).
Bayer, E. A., Shimon, L. J. W., Shoham, Y. & Lamed, R. Cellulosomes—structure and ultrastructure. J. Struct. Biol. 124, 221–234 (1998).
Kataeva, I., Li, X. L., Chen, H., Choi, S. K. & Ljungdahl, L. G. Cloning and sequence analysis of a new cellulase gene encoding CelK, a major cellulosome component of Clostridium thermocellum: evidence for gene duplication and recombination. J. Bacteriol. 181, 5288–5295 (1999).
Béguin, P., Cornet, P. & Aubert, J. P. Sequence of a cellulase gene of the thermophilic bacterium Clostridium thermocellum. J. Bacteriol. 162, 102–105 (1985).
Reverbel-Leroy, C., Pages, S., Belaich, A., Belaich, J. P. & Tardif, C. The processive endocellulase CelF, a major component of the Clostridium cellulolyticum cellulosome: purification and characterization of the recombinant form. J. Bacteriol. 179, 46–52 (1997).
Jeng, W.-Y. et al. Structural and functional analysis of three β-glucosidases from bacterium Clostridium cellulovorans, fungus Trichoderma reesei and termite Neotermes koshunensis. J. Struct. Biol. 173, 46–56 (2011).
Bayer, E. A., Belaich, J.-P., Shoham, Y. & Lamed, R. The cellulosomes: multienzyme machines for degradation of plant cell wall polysaccharides. Annu. Rev. Microbiol. 58, 521–554 (2004).
Oliveira, C., Carvalho, V., Domingues, L. & Gama, F. M. Recombinant CBM-fusion technology—applications overview. Biotechnol. Adv. 33, 358–369 (2015).
Davis, K. A., Abrams, B., Iyer, S. B., Hoffman, R. A. & Bishop, J. E. Determination of CD4 antigen density on cells: role of antibody valency, avidity, clones and conjugation. Cytometry 33, 197–205 (1998).
Gratama, J. W. et al. Flow cytometric quantitation of immunofluorescence intensity: problems and perspectives. European working group on clinical cell analysis. Cytometry 33, 166–178 (1998).
Smith, M. R., Tolbert, S. V. & Wen, F. Protein-scaffold directed nanoscale assembly of T cell ligands: artificial antigen presentation with defined valency, density and ratio. ACS Synth. Biol. 7, 1629–1639 (2018).
Ripley, B. D. Modelling spatial patterns. J. R. Stat. Soc. Series B 39, 172–192 (1977).
Wang, X. X. & Shusta, E. V. The use of scFv-displaying yeast in mammalian cell surface selections. J. Immunol. Methods 304, 30–42 (2005).
Jindou, S. et al. Cohesin–dockerin interactions within and between Clostridium josui and Clostridium thermocellum: binding selectivity between cognate dockerin and cohesin domains and species specificity. J. Biol. Chem. 279, 9867–9874 (2004).
Bae, J., Kuroda, K. & Ueda, M. Proximity effect among cellulose-degrading enzymes displayed on the Saccharomyces cerevisiae cell surface. Appl. Environ. Microbiol. 81, 59–66 (2015).
Erickson, H. P. Size and shape of protein molecules at the nanometer level determined by sedimentation, gel filtration and electron microscopy. Biol. Proced. Online 11, 32–51 (2009).
Chen, L., Mulchandani, A. & Ge, X. Spore-displayed enzyme cascade with tunable stoichiometry. Biotechnol. Prog. 33, 383–389 (2017).
Chen, L., Holmes, M., Schaefer, E., Mulchandani, A. & Ge, X. Highly active spore biocatalyst by self-assembly of co-expressed anchoring scaffoldin and multimeric enzyme. Biotechnol. Bioeng. 115, 557–564 (2018).
Lin, J. L., Zhu, J. & Wheeldon, I. Synthetic protein scaffolds for biosynthetic pathway colocalization on lipid droplet membranes. ACS Synth. Biol. 6, 1534–1544 (2017).
Fu, J. et al. Multi-enzyme complexes on DNA scaffolds capable of substrate channelling with an artificial swinging arm. Nat. Nanotechnol. 9, 531–536 (2014).
García-Alvarez, B. et al. Molecular architecture and structural transitions of a Clostridium thermocellum mini-cellulosome. J. Mol. Biol. 407, 571–580 (2011).
Wood, T. M. & Bhat, K. M. Methods for measuring cellulase activities. Methods Enzymol. 160, 87–112 (1988).
Liu, Z. et al. Engineering of a novel cellulose-adherent cellulolytic Saccharomyces cerevisiae for cellulosic biofuel production. Sci. Rep. 6, 24550 (2016).
Acknowledgements
This work was partly supported by the National Science Foundation (NSF) under grants nos. 1511720 and 1645229 and CAREER award 1653611, by the National Institutes of Health under grants nos. CA191952 and OD020053, and by MCubed at the University of Michigan. H.G. and J.-K.L. were partly supported by the Basic Science Research Program through the National Research Foundation of Korea (2017R1A2B3011676 and 2013M3A6A8073184) and by a WTU joint research grant from Konkuk University. The authors thank L. Zhang and B.D. Hill for assistance with the confocal microscopy experiments, and C. Jackman for help with setting up the anaerobic chamber for fermentation.
Author information
Authors and Affiliations
Contributions
F.W. conceived the idea behind this work. F.W. and J.-K.L. supervised the project. F.W., M.R.S., H.G. and C.R. designed the experiments. M.R.S., H.G., P.P., C.R., D.M., L.L., C.M.Y. and L.F.B. carried out the experiments. M.R.S., R.M.Z. and F.W. carried out the modelling work. M.R.S. and F.W. analysed the data and wrote the paper with H.G.’s assistance. All authors discussed and commented on the manuscript. All authors have given approval for the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary methods, Supplementary Figs. 1–12, Supplementary Tables 1–3, Supplementary references
Rights and permissions
About this article
Cite this article
Smith, M.R., Gao, H., Prabhu, P. et al. Elucidating structure–performance relationships in whole-cell cooperative enzyme catalysis. Nat Catal 2, 809–819 (2019). https://doi.org/10.1038/s41929-019-0321-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41929-019-0321-8
This article is cited by
-
Engineered repeat proteins as scaffolds to assemble multi-enzyme systems for efficient cell-free biosynthesis
Nature Communications (2023)
-
Biodegradation of highly crystallized poly(ethylene terephthalate) through cell surface codisplay of bacterial PETase and hydrophobin
Nature Communications (2022)
-
Characterization of Cellobiohydrolases from Schizophyllum commune KMJ820
Indian Journal of Microbiology (2020)