Abstract
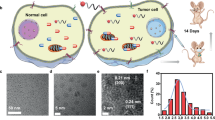
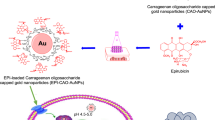
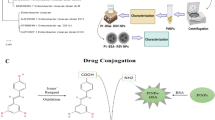
The ability of natural metalloproteins to prevent inactivation of their metal cofactors by biological metabolites, such as glutathione, is an area that has been largely ignored in the field of artificial metalloenzyme (ArM) development. Yet, for ArM research to transition into future therapeutic applications, biocompatibility remains a crucial component. The work presented here shows the creation of a human serum albumin-based ArM that can robustly protect the catalytic activity of a bound ruthenium metal, even in the presence of 20 mM glutathione under in vitro conditions. To exploit this biocompatibility, the concept of glycosylated artificial metalloenzymes (GArM) was developed, which is based on functionalizing ArMs with N-glycan targeting moieties. As a potential drug therapy, this study shows that ruthenium-bound GArM complexes could preferentially accumulate to varying cancer cell lines via glycan-based targeting for prodrug activation of the anticancer agent umbelliprenin using ring-closing metathesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The data that support the plots within this paper and other findings of this study are available from the corresponding author upon reasonable request.
References
Rebelein, J. G. & Ward, T. R. In vivo catalyzed new-to-nature reactions. Curr. Opin. Biotechnol. 53, 106–114 (2018).
Corso, C. R. & Acco, A. Glutathione system in animal model of solid tumors: from regulation to therapeutic target. Crit. Rev. Oncol. Hematol. 128, 43–57 (2018).
Wu, G. et al. Glutathione metabolism and its implications for health. J. Nutr. 134, 489–492 (2004).
Miller, M. A. et al. Nano-palladium is a cellular catalyst for in vivo chemistry. Nat. Commun. 8, 15906 (2017).
Clavadetscher, J. et al. Copper catalysis in living systems and in situ drug synthesis. Angew. Chem. Int. Ed. 55, 15662–15666 (2016).
Clavadetscher, J. et al. In-cell dual drug synthesis by cancer-targeting palladium catalysts. Angew. Chem. Int. Ed. 56, 6864–6868 (2017).
Weiss, J. T. et al. Extracellular palladium-catalysed dealkylation of 5-fluoro-1-propargyl-uracil as a bioorthogonally activated prodrug approach. Nat. Commun. 5, 3277 (2014).
Pérez-López, A. M. et al. Gold-triggered uncaging chemistry in living systems. Angew. Chem. Int. Ed. 56, 12548–12552 (2017).
Bray, T. L. et al. Bright insights into palladium-triggered local chemotherapy. Chem. Sci. 9, 7354–7361 (2018).
Liu, Y. et al. Catalytically active single-chain polymeric nanoparticles: exploring their functions in complex biological media. J. Am. Chem. Soc. 140, 3423–3433 (2018).
Li, J. et al. Palladium-triggered deprotection chemistry for protein activation in living cells. Nat. Chem. 6, 352–361 (2014).
Vidal, C. et al. Concurrent and orthogonal gold(i) and ruthenium(ii) catalysis inside living cells. Nat. Commun. 9, 1913 (2018).
Destito, P. et al. Hollow nanoreactors for Pd-catalyzed Suzuki–Miyaura coupling and O-propargyl cleavage reactions in bio-relevant aqueous media. Chem. Sci. 10, 2598–2603 (2019).
Tonga, G. Y. et al. Supramolecular regulation of bioorthogonal catalysis in cells using nanoparticle-embedded transition metal catalysts. Nat. Chem. 7, 597–603 (2015).
Streu, C. & Meggers, E. Ruthenium-induced allylcarbamate cleavage in living cells. Angew. Chem. Int. Ed. 45, 5645–5648 (2006).
Völker, T., Dempwolff, F., Graumann, P. L. & Meggers, E. Progress towards bioorthogonal catalysis with organometallic compounds. Angew. Chem. Int. Ed. 53, 10536–10540 (2014).
Tomás-Gamasa, M., Martínez-Calvo, M., Couceiro, J. R. & Mascareñas, J. L. Transition metal catalysis in the mitochondria of living cells. Nat. Commun. 7, 12538 (2016).
Yusop, R. M. et al. Palladium-mediated intracellular chemistry. Nat. Chem. 3, 239–243 (2011).
Unciti-Broceta, A. et al. Synthesis of polystyrene microspheres and functionalization with Pd(0) nanoparticles to perform bioorthogonal organometallic chemistry in living cells. Nat. Protoc. 7, 1207–1218 (2012).
Jeschek, M., Panke, S. & Ward, T. R. Artificial metalloenzymes on the verge of new-to-nature metabolism. Trends Biotechnol. 36, 60–72 (2018).
Ringenberg, M. R. & Ward, T. R. Merging the best of two worlds: artificial metalloenzymes for enantioselective catalysis. Chem. Commun. 47, 8470–8476 (2011).
Heinisch, T. & Ward, T. R. Artificial metalloenzymes based on the biotin–streptavidin technology: challenges and opportunities. Acc. Chem. Res. 49, 1711–1721 (2016).
Schwizer, F. et al. Artificial metalloenzymes: reaction scope and optimization strategies. Chem. Rev. 118, 142–231 (2018).
Lewis, J. C. Artificial metalloenzymes and metallopeptide catalysts for organic synthesis. ACS Catal. 3, 2954–2975 (2013).
Lewis, J. C. Metallopeptide catalysts and artificial metalloenzymes containing unnatural amino acids. Curr. Opin. Chem. Biol. 25, 27–35 (2015).
Liang, A. D., Serrano-Plana, J., Peterson, R. L. & Ward, T. R. Artificial metalloenzymes based on the biotin–streptavidin technology: enzymatic cascades and directed evolution. Acc. Chem. Res. 52, 585–595 (2019).
Coelho, P. S., Brustad, E. M., Kannan, A. & Arnold, F. H. Olefin cyclopropanation via carbene transfer catalyzed by engineered cytochrome P450 enzymes. Science 339, 307–310 (2013).
Coelho, P. S. et al. A serine-substituted P450 catalyzes highly efficient carbene transfer to olefins in vivo. Nat. Chem. Biol. 9, 485–487 (2013).
Prier, C. K. et al. Enantioselective, intermolecular benzylic C–H amination catalysed by an engineered iron-haem enzyme. Nat. Chem. 9, 629–634 (2017).
Kan, S. B. J., Lewis, R. D., Chen, K. & Arnold, F. H. Directed evolution of cytochrome c for carbon–silicon bond formation: bringing silicon to life. Science 354, 1048–1051 (2016).
Matthews, M. L. et al. Direct nitration and azidation of aliphatic carbons by an iron-dependent halogenase. Nat. Chem. Biol. 10, 209–215 (2014).
Zastrow, M. L., Peacock, A. F. A., Stuckey, J. A. & Pecoraro, V. L. Hydrolytic catalysis and structural stabilization in a designed metalloprotein. Nat. Chem. 4, 118–123 (2011).
Khare, S. D. et al. Computational redesign of a mononuclear zinc metalloenzyme for organophosphate hydrolysis. Nat. Chem. Biol. 8, 294–300 (2012).
Song, W. J. & Tezcan, F. A. A designed supramolecular protein assembly with in vivo enzymatic activity. Science 346, 1525–1528 (2014).
Oohora, K. et al. C(sp 3)–H bond hydroxylation catalyzed by myoglobin reconstituted with manganese porphycene. J. Am. Chem. Soc. 135, 17282–17285 (2013).
Key, H. M., Dydio, P., Clark, D. S. & Hartwig, J. F. Abiological catalysis by artificial haem proteins containing noble metals in place of iron. Nature 534, 534–537 (2016).
Dydio, P. et al. An artificial metalloenzyme with the kinetics of native enzymes. Science 354, 102–106 (2016).
Ghattas, W. et al. Receptor-based artificial metalloenzymes on living human cells. J. Am. Chem. Soc. 140, 8756–8762 (2018).
Zhao, J. et al. An artificial metalloenzyme for carbene transfer based on a biotinylated dirhodium anchored within streptavidin. Catal. Sci. Technol. 8, 2294–2298 (2018).
Hyster, T. K., Knorr, L., Ward, T. R. & Rovis, T. Biotinylated Rh(iii) complexes in engineered streptavidin for accelerated asymmetric C–H activation. Science 338, 500–503 (2012).
Yang, H., Srivastava, P., Zhang, C. & Lewis, J. C. A general method for artificial metalloenzyme formation through strain-promoted azide–alkyne cycloaddition. ChemBioChem 15, 223–227 (2014).
Yang, H. et al. Evolving artificial metalloenzymes via random mutagenesis. Nat. Chem. 10, 318–324 (2018).
Srivastava, P., Yang, H., Ellis-Guardiola, K. & Lewis, J. C. Engineering a dirhodium artificial metalloenzyme for selective olefin cyclopropanation. Nat. Commun. 6, 7789 (2015).
Grimm, A. R. et al. A whole cell E. coli display platform for artificial metalloenzymes: poly(phenylacetylene) production with a rhodium–nitrobindin metalloprotein. ACS Catal. 8, 2611–2614 (2018).
Köhler, V. et al. Synthetic cascades are enabled by combining biocatalysts with artificial metalloenzymes. Nat. Chem. 5, 93–99 (2012).
Raines, D. J. et al. Redox-switchable siderophore anchor enables reversible artificial metalloenzyme assembly. Nat. Catal. 1, 680–688 (2018).
Zhao, J. et al. Genetic engineering of an artificial metalloenzyme for transfer hydrogenation of a self-immolative substrate in Escherichia coli’s periplasm. J. Am. Chem. Soc. 140, 13171–13175 (2018).
Jeschek, M. et al. Directed evolution of artificial metalloenzymes for in vivo metathesis. Nature 537, 661–665 (2016).
Lo, C. et al. Artificial metalloenzymes for olefin metathesis based on the biotin–(strept)avidin technology. Chem. Commun. 47, 12065–12067 (2011).
Zhao, J., Kajetanowicz, A. & Ward, T. R. Carbonic anhydrase II as host protein for the creation of a biocompatible artificial metathesase. Org. Biomol. Chem. 13, 5652–5655 (2015).
Mayer, C., Gillingham, D. G., Ward, T. R. & Hilvert, D. An artificial metalloenzyme for olefin metathesis. Chem. Commun. 47, 12068–12070 (2011).
Szponarski, M., Schwizer, F., Ward, T. R. & Gademann, K. On-cell catalysis by surface engineering of live cells with an artificial metalloenzyme. Commun. Chem. 1, 84 (2018).
Heinisch, T. et al. E. coli surface display of streptavidin for directed evolution of an allylic deallylase. Chem. Sci. 9, 5383–5388 (2018).
Sauer, D. F. et al. Hybrid ruthenium ROMP catalysts based on an engineered variant of β-barrel protein FhuA ΔCVF(tev): effect of spacer length. Chem. Asian J. 10, 177–182 (2015).
Philippart, F. et al. A hybrid ring-opening metathesis polymerization catalyst based on an engineered variant of the β-barrel protein FhuA. Chem. Eur. J. 19, 13865–13871 (2013).
Matsuo, T. et al. Creation of an artificial metalloprotein with a Hoveyda–Grubbs catalyst moiety through the intrinsic inhibition mechanism of α-chymotrypsin. Chem. Commun. 48, 1662–1664 (2012).
Basauri-Molina, M. et al. Ring-closing and cross-metathesis with artificial metalloenzymes created by covalent active site-directed hybridization of a lipase. Chem. Eur. J. 21, 15676–15685 (2015).
Okamoto, Y. et al. A cell-penetrating artificial metalloenzyme regulates a gene switch in a designer mammalian cell. Nat. Commun. 9, 1943 (2018).
Peters, T. Jr. Serum albumin. Adv. Protein Chem. 37, 161–245 (1985).
Ghuman, J. et al. Structural basis of the drug-binding specificity of human serum albumin. J. Mol. Biol. 353, 38–52 (2005).
Keating, G. M. Insulin detemir. Drugs 72, 2255–2287 (2012).
Petitpas, I. et al. Crystal structure analysis of warfarin binding to human serum albumin: anatomy of drug site I. J. Biol. Chem. 276, 22804–22809 (2001).
Lissi, E., Calderón, C. & Campos, A. Evaluation of the number of binding sites in proteins from their intrinsic fluorescence: limitations and pitfalls. Photochem. Photobiol. 89, 1413–1416 (2013).
Tetko, I. V. & Tanchuk, V. Y. Application of associative neural networks for prediction of lipophilicity in ALOGPS 2.1 program. J. Chem. Inf. Comput. Sci. 42, 1136–1145 (2002).
Morris, G. M. et al. AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J. Comput. Chem. 30, 2785–2791 (2009).
Peters, T. in All About Albumin Ch. 2 (Academic Press, 1995).
Ogura, A. et al. Visualizing trimming dependence of biodistribution and kinetics with homo- and heterogeneous N-glycoclusters on fluorescent albumin. Sci. Rep. 6, 21797 (2016).
Ogura, A. et al. Glycan multivalency effects toward albumin enable N-glycan-dependent tumor targeting. Bioorg. Med. Chem. Lett. 26, 2251–2254 (2016).
Ogura, A. et al. A viable strategy for screening the effects of glycan heterogeneity on target organ adhesion and biodistribution in live mice. Chem. Commun. 54, 8693–8696 (2018).
Taichi, M. et al. In situ ligation of high- and low-affinity ligands to cell surface receptors enables highly selective recognition. Adv. Sci. 4, 1700147 (2017).
Tsubokura, K. et al. In vivo gold complex catalysis within live mice. Angew. Chem. Int. Ed. 56, 3579–3584 (2017).
Lin, Y., Vong, K., Matsuoka, K. & Tanaka, K. 2-Benzoylpyridine ligand complexation with gold critical for propargyl ester-based protein labeling. Chem. Eur. J. 24, 10595–10600 (2018).
Lahm, H. et al. Comprehensive galectin fingerprinting in a panel of 61 human tumor cell lines by RT-PCR and its implications for diagnostic and therapeutic procedures. J. Cancer Res. Clin. Oncol. 127, 375–386 (2001).
Carlsson, S. et al. Affinity of galectin-8 and its carbohydrate recognition domains for ligands in solution and at the cell surface. Glycobiology 17, 663–676 (2007).
Shakeri, A., Iranshahy, M. & Iranshahi, M. Biological properties and molecular targets of umbelliprenin—a mini-review. J. Asian Nat. Prod. Res. 16, 884–889 (2014).
Rashidi, M. et al. Umbelliprenin shows antitumor, antiangiogenesis, antimetastatic, anti-inflammatory and immunostimulatory activities in 4T1 tumor-bearing Balb/c mice. J. Cell. Physiol. 233, 8908–8918 (2018).
Jun, M. et al. Synthesis and biological evaluation of isoprenylated coumarins as potential anti-pancreatic cancer agents. Bioorg. Med. Chem. Lett. 24, 4654–4658 (2014).
Barthomeuf, C., Lim, S., Iranshahi, M. & Chollet, P. Umbelliprenin from Ferula szowitsiana inhibits the growth of human M4Beu metastatic pigmented malignant melanoma cells through cell-cycle arrest in G1 and induction of caspase-dependent apoptosis. Phytomedicine 15, 103–111 (2008).
Gholami, O. et al. Umbelliprenin from Ferula szowitsiana activates both intrinsic and extrinsic pathways of apoptosis in Jurkat T-CLL cell line. Iran. J. Pharm. Res. 12, 371–376 (2013).
Gholami, O. et al. Mcl-1 is up regulated by prenylated coumarin, umbelliprenin in Jurkat cells. Iran. J. Pharm. Res. 13, 1387–1392 (2014).
Sibgatullina, R. et al. Highly reactive ‘RIKEN click’ probe for glycoconjugation on lysines. Tetrahedron Lett. 58, 1929–1933 (2017).
Fery-Forgues, S. & Lavabre, D. Are fluorescence quantum yields so tricky to measure? A demonstration using familiar stationery products. J. Chem. Educ. 76, 1260–1264 (1999).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Piraux, G. et al. New ruthenium-based probes for selective G-quadruplex targeting. Chem. Eur. J. 23, 11872–11880 (2017).
Acknowledgements
In memory of Professor Koji Nakanishi. The authors thank Glytech Inc. for supplying the complex N-glycans. CD spectral measurements were supported by the Molecular Structure Characterization Unit, RIKEN Center for Sustainable Resourse Science (CSRS). This work was supported financially by JSPS KAKENHI grants nos. JP16H03287, JP18K19154, JP18K14347 and JP15H05843 for Middle Molecular Strategy. This work was also performed with the support of the Russian Government Program for Competitive Growth, granted to Kazan Federal University.
Author information
Authors and Affiliations
Contributions
Preparation of reagents was carried out by S.E. and I.N. Reactivity and kinetic studies were performed by S.E., I.N. and K.V. Binding studies were performed by K.V. Prodrug activation and biological studies were carried out by I.N. and K.V. Modelling studies were carried out by K.V. and N.K. The manuscript was written by K.V. and K.T. and checked by M.Y. and A.K. The research was directed and supervised by K.T.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–60, Supplementary Tables 1–5, Supplementary methods, Supplementary discussion, Supplementary references
Rights and permissions
About this article
Cite this article
Eda, S., Nasibullin, I., Vong, K. et al. Biocompatibility and therapeutic potential of glycosylated albumin artificial metalloenzymes. Nat Catal 2, 780–792 (2019). https://doi.org/10.1038/s41929-019-0317-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41929-019-0317-4
This article is cited by
-
Metal complex catalysts broaden bioorthogonal reactions
Science China Chemistry (2024)
-
Synthetic prodrug design enables biocatalytic activation in mice to elicit tumor growth suppression
Nature Communications (2022)
-
Engineering and emerging applications of artificial metalloenzymes with whole cells
Nature Catalysis (2021)
-
Bioorthogonal catalytic patch
Nature Nanotechnology (2021)
-
Design and evolution of chimeric streptavidin for protein-enabled dual gold catalysis
Nature Catalysis (2021)