Abstract
The COVID-19 pandemic has severely impacted public health and the worldwide economy. Converging evidence from the current pandemic, previous outbreaks and controlled experiments indicates that SARS-CoVs are present in wastewater for several days, leading to potential health risks via waterborne and aerosolized wastewater pathways. Conventional wastewater treatment provides only partial removal of SARS-CoVs, thus safe disposal or reuse will depend on the efficacy of final disinfection. This underscores the need for a risk assessment and management framework tailored to SARS-CoV-2 transmission via wastewater, including new tools for environmental surveillance, ensuring adequate disinfection as a component of overall COVID-19 pandemic containment.
Similar content being viewed by others
Main
The ongoing COVID-19 pandemic is caused by a new human coronavirus (SARS-CoV-2). Infections can result in severe acute respiratory syndrome (SARS) as well as a growing number of other severe medical conditions. This global pandemic has already led to more than 600,000 deaths globally within seven months1. Over the past century, outbreaks of viral origin have increasingly evolved into worldwide pandemics (Fig. 1a,b). In the last 20 years, two coronaviruses have caused infections with respiratory syndrome: SARS-CoV-1 in 2003 and Middle East respiratory syndrome coronavirus (MERS-CoV) in 20122,3.
a,b, Approximate number of infected and deceased individuals in extensive outbreaks (a) and the COVID-19 pandemic in recent months (b). c, Overview of potential SARS-CoV-2 dissemination via waterborne pathways in industrialized countries. Included outbreaks were caused by enveloped single-stranded RNA viruses. Asterisks depict a confirmed or putative dissemination via wastewater. Letters next to each pandemic indicate specific references: a125,126; b127; c128; d106,127; e126,129; f130,131; g132; h127; i133,134; j1 (data sources accessed 28 July 2020).
SARS-CoV-2 is highly contagious and spreads much like SARS-CoV-1, by close personal contact through respiratory droplets or mutual interactions with surfaces, as well as by aerosols3,4,5,6. Currently, it is not clear if additional transmission pathways are important for the spread of this disease locally or globally7,8. Investigation of a SARS-CoV-1 outbreak within an apartment building during the 2003 pandemic indicated that the virus can spread via wastewater systems9,10. It was suggested that the SARS-CoV-1 infection occurred through respiration of aerosols created by toilet flushing or faulty plumbing systems9,10. Similar to SARS-CoV-1, RNA of the newly emerged SARS-CoV-2 has been detected in stool samples as well as wastewater11,12,13,14. Therefore, it has been postulated that wastewater, a sustainable source of freshwater15,16,17, could be an indirect infection pathway during SARS-CoVs outbreaks (Fig. 1c)2,13,18,19.
Entry of the virus into the sewer system results in a variety of potential transport pathways (Fig. 1c) that must be considered in the context of faecal–oral transmission2,18. In industrialized countries, most of the collected domestic wastewater and viral load is treated in centralized wastewater treatment plants (WWTPs). However, conventional WWTPs generally do not remove virions completely and high influent viral loads during pandemics can lead to insufficient reduction of viruses before discharge20,21,22. Furthermore, freshwater scarcity results in the reuse of an increasing volume of treated wastewater for a variety of purposes, such as groundwater recharge, recreation and irrigation of food crops, thus creating other potential routes for SARS-CoV-2 transmission. An additional and potentially serious health risk is faecal–oral transmission in low-income countries where communities with inadequate sanitation infrastructure (for example, open sewers and direct discharge into the environment) could be infected by untreated wastewater or faecal waste23,24,25.
Overall, it is critical to evaluate the potential of wastewater as a transmission pathway of SARS-CoV-2. Information on the survival and dissemination of enveloped viruses in general, and SARS-CoV-2 in particular during wastewater collection, treatment and reuse, is limited. Herein we critically review and synthesize existing knowledge on the health risks as well as the potential spread of SARS-CoVs in waterborne, waterborne–aerosolized and waterborne–foodborne pathways during a pandemic, when the probability of viral infection is substantially higher than in non-outbreak scenarios. Specifically, we focus on centralized wastewater systems that are commonly implemented in regions with moderate to high socio–economic capacity, including those regions currently experiencing the largest numbers of COVID-19 cases. These conditions indicate an urgent need for monitoring programmes and risk assessments tailored to SARS-CoV-2 in wastewater, which may aid in the early detection and containment of future viral disease outbreaks.
SARS-CoVs
Viruses are ubiquitous nanoscale infectious agents that exist as extracellular particles between intracellular reproduction cycles, and infect specific host cells. Viruses vary in structure (from simple helical capsids to complex constructs), size (20–300 nm), as well as replication mechanisms (that is, budding to lytic) and life-cycle durations (minutes to hours). In some viruses, the capsid is surrounded by an envelope consisting of a lipid-bilayer membrane with embedded proteins that play a crucial role for attachment to the host cell. Enveloped viruses are often more readily inactivated than non-enveloped viruses since the envelope is less robust against environmental conditions and disinfecting agents26. The viral genome can consist of either DNA or RNA, with RNA viruses showing a higher propensity for mutations26.
Coronaviruses (CoVs) are enveloped, single-stranded, positive-sense RNA viruses, ranging from 60 to 220 nm in size. The name derives from the spike glycoproteins that decorate the envelope giving the virions a crown-like shape (‘corona’ in Latin) (Fig. 2a,b)3,27. The spike glycoproteins mediate receptor recognition and membrane fusion. Recent studies have suggested that the spike glycoprotein of SARS-CoV-2 binds the human receptors with a higher affinity than SARS-CoV-1, leading to a more infectious virion28. SARS-CoV-1 and SARS-CoV-2 are members of lineage B of the genus Betacoronavirus3,27 (Fig. 2c); these viruses are both highly adaptive and able to infect different human tissue. They easily transfer between new host species, and adjust to diverse ecological conditions through accumulation of point mutations and homologous recombination29.
a, Transmission electron micrograph of a negatively stained SARS-CoV-2 virus that was slightly damaged revealing enclosed RNA, as indicated by the white arrow. b, Schematic illustration of the main structural features of SARS-CoV-2. c, Phylogenetic analysis based on the RNA-dependent RNA polymerase (RdRP) amino acid sequences of SARS-CoV-2 and representative enveloped and non-enveloped viruses.
Sources of SARS-CoVs in wastewater
Although SARS-CoVs are primarily respiratory viruses, SARS-CoV-2 may infect and replicate in the gastrointestinal tract30,31. Additionally, it has been observed that SARS-CoV-2 (detected via reverse transcription quantitative polymerase chain reaction (RT-qPCR)) enters the wastewater system via human excretions (that is, stool and urine)13,14,32. The frequency of gastrointestinal disease manifestations, including diarrhoea and vomiting, ranges between 2% to 80% of confirmed patients12,33,34. Up to 67% of stool samples test positive for SARS-CoV-2 RNA with counts nearly reaching the maximum in sputum (108 copies ml–1 stool versus 109 copies ml–1 sputum)35,36. SARS-CoV-2 RNA shedding in stool has also been observed in cases without gastrointestinal but other symptoms, as well as in pre- and asymptomatic cases, and for up to four weeks after symptoms cease34,35,36,37. Interestingly, SARS-CoV-2 RNA is often present in stool after respiratory infection resolves and respiratory samples are found to be negative35,38.
Although only a few studies have succeeded in isolating infective SARS-CoV-2 from stool and urine samples32,39,40, many studies have detected SARS-CoV-2 RNA in stool and wastewater13,14,20,41,42,43. To date it is unclear if SARS-CoV-2 RNA in the stool originates from swallowed sputum or active replication within the gastrointestinal tract; the latter case would strongly influence dissemination of infective virions via wastewater. A bioinformatics modelling study and an experimental study indicated infection and replication of virions in the gastrointestinal tract30,31. Additionally, a compartmental epidemiological model based on data from the Wuhan epidemic indicated that the faecal–oral route is putatively important for transmission of the virus44. These results suggest that replication of the virus in the intestinal tract is highly likely. Thus, wastewater should be assumed to contain a considerable number of infective virions2,13,35,45. The unsuccessful isolation of infective SARS-CoV-2 from stool and wastewater samples may be due to the difficulty of isolating intact enveloped virions, rather than the absence of infective virions.
Greywater (that is, water discharged from sinks, showers and drains; Fig. 1c) is not expected to be a major SARS-CoV-2 transmission vehicle despite containing body fluids such as saliva and sputum with potentially high viral concentrations35,39. Low virus concentration is expected since greywater often contains detergents, soaps and other disinfectants, to which SARS-CoV-2 is sensitive5,46.
Detection and infectivity of SARS-CoVs
Current detection and monitoring of SARS-CoVs in wastewater can be divided into three categories: (1) qualitative and (2) quantitative molecular approaches as well as (3) in vitro counts by plaque-forming units (PFU). Molecular approaches target the SARS-CoVs RNA and can provide estimates of the presence and abundance of RNA copies (or fragments) in a water sample, but do not measure viral infectivity18. PFU can provide a quantitative estimation of infective virions, but this method is slow and difficult as in vitro cultivation requires an appropriate host18,26. It should be noted that the sensitivity of plaque assays for viral detection can be further limited by the cytotoxicity of toxins often found in wastewater samples47,48,49. Additionally, virus concentrations need to be even higher to isolate infective virions compared to RNA detection (>106 copies ml–1)35. Therefore, it is not surprising that wastewater of hospitals tested positive for SARS-CoVs RNA, but not for infective virions13,35. However, it is also possible that the concentration of infective virions in hospital wastewater was under the detection limit due to the intense use of disinfectants and various surfactants46,50.
Regardless of the methods used, detection and enumeration of SARS-CoVs in wastewater is extremely challenging due to their low abundance after dilution (101 to 106 copies l–1) relative to direct assays on human excretions11,14,20,45. Thus, concentrating the sample with high recovery rates is required51 and promising methods have recently been reviewed19,52. Intact virions can be concentrated on cell-free substrate coated with corresponding receptors after enzyme treatment to remove broken virions53. Then, bound virions can be quantified by RT-qPCR53. Recently, this method has been shown to detect and quantify SARS-CoV-2 in wastewater after dedicated extraction methods13,45,54. A transcriptome (RNA) analysis from activated sludge uncovered a wide range of uncultivated bacterial viruses that could not be discovered in other molecular techniques55. Although low viral concentrations will require various concentration approaches, this technique, in addition to metagenomic virome studies56, may provide a novel template for the detection of single-stranded RNA viruses such as SARS-CoV-2 in wastewater.
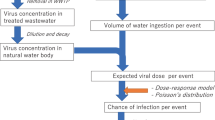
Both abundance and viral infectivity are critical factors for disease transmission. However, extraction protocols often lead to low recovery rates for intact viruses (on the order of a few percent), since the lipid-bilayer membrane surrounding the capsid is easily disrupted18,57. Thus, other approaches must be developed to determine infectivity in wastewater. Currently, the minimal infectious dose (MID) of SARS-CoV-2 (that is, the number of viral particles that causes an infection) for humans is unknown19. Yet, the rapid spread of the disease suggests that the MID is low and similar to other enveloped viruses (Table 1)27,58,59.
Survival of SARS-CoVs in water and wastewater
Survival time distributions of viruses are often exponential and reported as a half-life, number of log removal or the time required to reach 90% inactivation (T90; Fig. 3). Under numerous environmental conditions, virions of SARS-CoVs and other enveloped viruses remain infective for several days (Fig. 3). Factors that were found to affect SARS-CoVs infectivity in water and wastewater include temperature, organic content and solution pH46,60,61. However, the way this translates into risk of infection is yet unknown, especially since human activities and water exposure differ across seasons and regions.
Survival time of SARS-CoVs (squares) and other enveloped viruses (circles) is approximated by the time required for 90% inactivation (T90). Water samples include: deionized and sterilized tap water (RO + UV tap); tap water after chlorine removal (Dechlor. tap); phosphate buffered saline solution (PBS); lake water and human excretions (urine and stool); pasteurized (Past.) wastewater (WW) after settling; past. raw WW; primary or secondary effluent (Pri., sec. eff); and raw WW46,57,61,62,64,66,135,136.
Temperature is an important variable for survival of virions in general and SARS-CoVs in particular46,60,61. Longer retention of SARS-CoVs infectivity has been observed at lower temperatures (for example, 14 days at 4 °C versus two days at 25 °C in wastewater)61. This implies that in cold seasons and temperate climate zones, the environmental survival of SARS-CoV-2 may be increased. Temperatures above 56 °C reliably inactivate SARS-CoV-1 and SARS-CoV-2 after 90 minutes and 30 minutes, respectively, most likely due to denaturation of proteins and lipid bilayers46,62,63.
Organic matter at increasing concentration was reported to reduce the survival time of spiked CoVs in various water samples (Fig. 3; for example, ten days in lake water versus two days in raw wastewater). This may be due to the presence of antagonist bacteria that can inactivate the viruses via extracellular enzymatic activity57,64,65. Differently, organic matter in the context of wastewater treatment can non-specifically adsorb to the envelope of SARS-CoV virions, protecting them from oxidative damage, chlorination, ultraviolet (UV) radiation and protozoan or metazoan predation57,65. Additionally, viruses shed by infected patients are often already associated with organic material (for example, faeces and sputum) and are thus protected from some inactivation mechanisms57,60.
The pH of faeces had a considerable impact on SARS-CoV-1 survival ranging from three hours in slightly acidic faeces of a new-born to four days in diarrheal faeces of an adult with a pH of up to 966. In contrast, SARS-CoV-2 in suspension does not show substantial reduction in infective titre after 60 minutes over a wide range of pH (3–10)46.
Dissemination of SARS-CoVs from sewer systems
Wastewater in regions with moderate to high socio–economic capacity, which is the focus of this Review, is mostly collected from large municipal areas. The size of the population connected to the sewer system has a direct impact on the concentration of SARS-CoVs in wastewater and thus the potential for dissemination. Extensive sewer systems in large cities effectively mix wastewater from large areas, resulting in a rather homogenous viral dispersion, and thus lower concentration67. However, larger populations inherently have greater likelihood of virus importation, and COVID-19 outbreaks in large population centres naturally produce high virus concentrations that increase transmission risk68,69. Survival time of SARS-CoVs in wastewater is sufficiently long for infective viruses to reach WWTPs and to be further disseminated by several transmission pathways18,57,61. SARS-CoVs, similarly to other microbial pathogens, can reach natural water bodies used for recreation such as ponds, rivers and lakes via leakage or combined sewer overflows during storm events18,57. The high infectivity of SARS-CoV-2 could lead to transmission of COVID-19 in such environments. A compartmental epidemiological model suggests that contaminated natural water bodies could become environmental reservoirs of SARS-CoVs, which would require the enforcement of strict post-epidemic measures to prevent re-infection44.
Surveillance of SARS-CoV-2 via wastewater monitoring has recently been highlighted as a powerful tool for wastewater-based epidemiology and public health interventions complementary to individual testing11,19,41,45,54,70. Collecting data from central sewer systems in addition to individual testing can provide real-time information on the distribution of SARS-CoV-2 in related communities19,20,70 at reduced costs relative to personal testing71. Moreover, this information can be used as an early warning signal for COVID-19 outbreaks in specific communities with pre- and asymptomatic infected individuals14,19,54,72. These early signs of outbreak can be detected as development of SARS-CoV-2 concentration in wastewater precedes changes in confirmed COVID-19 cases by at least several days20,54. Wastewater monitoring is particularly useful to provide an early indication of re-emergence of SARS-CoV-2 in communities that contained an initial outbreak and subsequently relaxed containment measures. This information provides the ability to reinstate containment measures and allocate healthcare resources before COVID-19 infections become highly prevalent in specific communities. The approach of relaxing containment measures and reopening economies with ongoing community surveillance could be a cost-effective means for pandemic containment.
Low-income regions often lack wastewater sanitation, with partial to no sewer systems. Over 0.5 billion people still practice open defaecation, while another 3.5 billion people use unsafe sanitation73,74. These circumstances may facilitate transmission of viral diseases such as SARS-CoV-2 via the incidental faecal–oral route, as people are likely to come in contact with infected waste or wastewater75,76. Unsafe sanitation is often combined with inadequate drinking water infrastructure (for example, exposed sewer systems that contaminate freshwater sources), and lack of basic hygiene services such as clean water and soap for hand washing73. The spread of the pandemic in low-income countries is likely to be further accelerated by high population density in cities together with limited implementation of COVID-19 control measures23,77,78. Additionally, tropical and/or monsoonal weather with large volumes of rainwater flushing streets further increases viral contamination of water bodies. We stress that the COVID-19 pandemic is likely to be especially precarious for the 4 billion people who lack access to safe sanitation, frequently come in direct contact with faecally contaminated water and consume crops irrigated with contaminated wastewater23,77. Although these infection pathways have been confirmed for other viral diseases75,76, no such studies have been reported to date for COVID-19. Thus, investigation of the faecal–oral route for SARS-CoV-2 in low-income regions is of paramount importance.
SARS-CoVs dispersion in irrigated agriculture
Recent evidence of SARS-CoV-2 RNA in treated wastewater20 indicates possible risks associated with wastewater reuse for agriculture. Irrigation of fruit and vegetables with contaminated wastewater effluent may serve as an indirect transmission pathway for SARS-CoVs through handling or consumption of contaminated food79,80. This may be especially relevant for technologies that do not apply the water directly to the root zone (for example, drip irrigation), such as surface or sprinkler irrigation. Although foodborne transmission of SARS-CoV-2 has not been documented, similar viruses are known to be transmitted by foodborne pathways following irrigation with treated wastewater81,82. Bovine CoV, which is very similar to SARS-CoVs, remains infective on lettuce leaves for the entire shelf-life of the lettuce (at least 14 days)81 and human CoV 229E on lettuce only declined by 0.2 log after two days of storage at 4 °C82. Moreover, washing produce does not completely eliminate virions81. Thus, especially during a large SARS-CoV outbreak in areas without adequate sanitation, the connection of faecal–waterborne–foodborne transmission through irrigation with wastewater can be an important dissemination pathway. Additionally, sprinkler irrigation with wastewater and fertilization with wastewater solids generates considerable aerosols. These aerosols are often dispersed at regional scales79,80, which is especially important for farm workers and potentially relevant where agricultural and populated areas are in relatively close proximity. Transmission of SARS-CoV-2 via waterborne–foodborne or waterborne–aerosolized pathways during an outbreak can be minimized by disinfection prior to wastewater reuse, emphasizing the importance of standards for safe wastewater reuse.
Wastewater aerosols and SARS-CoVs exposure
Aerosolized viruses may be generated and transported locally in buildings as well as at larger scales by winds during wastewater treatment, from recreational water bodies (for example, urban rivers and ponds) fed by treated wastewater, or during irrigation and fertilization80,83,84,85. The formation of wastewater aerosols and droplets was confirmed as a key mechanism for faecal–droplet–respiration transmission during the SARS-CoV-1 outbreak, and is suspected in the current SARS-CoV-2 outbreak10,83,86. Aerosolized human CoV (HCoV 229E) has been found to be infective for up to six days at 25 °C in 50% humidity, and is suspected to be infective for even longer periods at 6 °C87. SARS-CoV-2 remains viable in aerosols for up to 16 hours with a median half-life of approximately one hour88,89. Although dispersal of larger droplets is limited, as they deposit close to the source, larger droplets cause local contamination of surfaces due to their enhanced capacity to carry pathogens, and are a major vector for pathogen transmission, including SARS-CoV-286,90.
WWTP operators must follow standard practices to limit exposure to wastewater and contaminated surfaces, thus reducing the risk of exposure to pathogens, such as SARS-CoV-219,91. Although no analysis of aerosolized SARS-CoV-2 within WWTPs has been reported, aerosol formation during the treatment process could pose a risk to WWTP operators and facilitate dissemination, especially for WWTPs in densely populated areas84,92. This pathway has been observed for multiple enteric viruses and bacteria84,92. Non-potable reuse of treated wastewater or greywater systems that generate aerosols (for example, cooling towers and sprinklers) as well as decorative structures such as fountains must ensure sufficient treatment to avoid infection pathways beyond WWTPs.
Dissemination into surface water and groundwater
SARS-CoVs may be disseminated to aquatic ecosystems during an outbreak due to leaking sewers or insufficient removal following wastewater treatment. Leakage of wastewater from septic tanks, pipe failure or lack of proper infrastructure can result in direct discharge of SARS-CoVs into receiving water bodies (for example, streams, rivers, ponds, estuaries, lakes and groundwater). Additionally, treated wastewater such as secondary effluent that is discharged may also carry viruses into the environment20,22. Except for two studies that reported SARS-CoV-2 RNA in an Italian and a Japanese river, but did not succeed to isolate infective virions93,94, there has not been direct detection of SARS-CoVs in aquatic environments52. However, several previous studies found that enveloped viruses travel considerable distances, and survive for a prolonged time in aquatic environments95,96. Further, rain events can increase virus concentrations in natural water systems via combined sewer overflows or failure of wastewater infrastructure95,97, which raises the probability of SARS-CoVs dissemination.
In the subsurface, viruses can be highly mobile due to the steric interaction of their outer spike glycoproteins with the porous media98,99, especially through preferential flow pathways and fractures100. Based on the size of SARS-CoV-2 (~100 nm) as well as the relatively long survival time in water (Fig. 3) and on surfaces5,88, SARS-CoV-2 could potentially travel considerable distances in the subsurface leading to contamination of aquifers used as freshwater sources for potable use. However, a recent study indicated that many enteric viruses are completely removed from secondary effluent during infiltration through a 30–40-m-thick vadose zone, leading to zero virus counts in the monitored groundwater wells101. This indicates that long infiltration times drastically reduce the risk of groundwater contamination of viruses, including SARS-CoV-2101.
Wastewater treatment to reduce dissemination
In WWTPs, virions can potentially be removed through physical, biological and chemical processes (Fig. 4). Wastewater first undergoes primary treatment where removal of viruses by sedimentation alone is low102,103. Secondary (biological) treatment combines aeration tanks with secondary sedimentation to retain the activated sludge. Virus sorption to organic particulates and removal by settling is thought to play an essential role in these secondary treatment steps103,104. Treatment approaches that maximize retention and removal of solids (for example, membrane bioreactors) have been suggested as a particularly efficacious means to remove viral loads from wastewater104,105. Although no specific data for SARS-CoV-2 are yet available, enveloped viruses are more likely to be removed together with particles than non-enveloped viruses57,65. Additionally, extracellular enzymes such as hydrolases and proteases present in the concentrated bacterial consortia characteristic of secondary bioreactors are also likely to inactivate SARS-CoVs, similarly to other viruses57,104,105.
Secondary biological treatment supports enzymatic degradation (Enz. deg.) of organic material (OM), including viruses. SARS-CoVs that were not removed by settling will likely aggregate with suspended organic material and be transported in primary and secondary effluent. a,b, Secondary effluent can be directly discharged to surface waters (a) or disinfected prior to discharge (b). c,d, Alternatively, secondary effluent can be further treated with advanced oxidation processes (AOP) (c) and/or filtration via different membrane systems such as ultrafiltration (for example, in a membrane bioreactor, MBR) (d) for reuse applications.
Concentrating SARS-CoVs in the sludge may pose the subsequent problem of sludge treatment and disposal102. In a metagenomic study on sludge from wastewater treatment processes, a high diversity of viruses, including those associated with respiratory diseases, was detected106. CoV genes were found in 80% of untreated wastewater sludge samples, and the second-most common RNA virus was CoV HKU1106. Data on the survival of enveloped viruses during sludge treatment are scarce and non-existent for SARS-CoVs18. Based on the fate of non-enveloped viruses, treatment of sludge by thermophilic digestion, lime addition, drying and composting is most promising for SARS-CoVs inactivation102,107. However, aerosol control should still be applied during fertilization with sludge in agricultural settings102,108.
The inactivation or removal of SARS-CoVs during primary and secondary treatment has not been studied in detail. SARS-CoV-2 RNA has been detected in treated wastewater with only 2-log removal of viruses compared to raw wastewater20, yet complete removal after secondary treatment was observed in a different study14. These mixed results indicate that, similar to other viruses, sufficient inactivation of SARS-CoV-2 is not ensured21,22. In some countries (for example, Israel), secondary effluent is further disinfected prior to reuse or before discharge into the environment to minimize viral dissemination (Fig. 4). Yet, disinfection measures are not compulsory in many countries (for example, in the United States, reuse without disinfection is allowed for irrigation of vineyards and wetlands, as well as stream augmentation), increasing the potential for SARS-CoV-2 dissemination. Disinfection of treated wastewater may currently be the most important step to ensure reliable SARS-CoV-2 inactivation2,18. While the mechanisms are unclear, enveloped viruses like SARS-CoVs tend to be more susceptible to chlorine-based disinfectants than non-enveloped viruses60,109. Although not tested in real wastewater, enveloped viruses such as SARS-CoV-2 are often found covered in organic material that provides a physical barrier against disinfection57,60. Hence, it is likely that in a complex medium rich in organic matter such as secondary effluent, SARS-CoV-2 would be less sensitive to disinfectants. In addition, chemical disinfectants are scavenged by organic matter and nitrogen-containing compounds in secondary effluent110, resulting in lower concentration of active chlorine. Consequently, infective enteric viruses have been detected even in disinfected secondary effluent21,22,111. During a pandemic outbreak, when viral loads in raw wastewater would be higher than normal, insufficient viral removal (particularly if disinfectant doses are not increased) may result in viral transmission via reuse.
Many industrialized countries apply tertiary treatment (that is, advanced particle removal and disinfection) before wastewater reuse. Tertiary treatment can include sand filtration, managed aquifer recharge, UV radiation, advanced oxidation processes (AOP) and/or membrane technologies to ensure enhanced removal of microbial pathogens (Fig. 4). UV irradiation at 254 nm is known to be effective against SARS-CoV-162,112 through reactions with the viral genome109. However, the required dose (a function of irradiance and time) is highly dependent on many virus- and media-related factors (that is, concentration of organic matter) and ranges widely112. Ozone-based treatment of secondary effluents effectively inactivates viruses through DNA or RNA attack by ozone113 or by forming free radicals114. Similar to chlorination, considerable scavenging of oxidation capacity by background constituents and formation of harmful disinfection by-products are possible. To date, the effectiveness of these disinfection processes for SARS-CoV-2 inactivation is unknown, and research is urgently needed.
Membrane technology to avert dissemination
Low-pressure membrane filtration, which includes microfiltration (MF) and ultrafiltration (UF), is an advanced technology used in wastewater treatment with potential to provide a complete barrier to SARS-CoV-2 dissemination. Additionally, the modular structure of membrane systems could facilitate upgrading of existing WWTPs to reduce effluent concentrations of SARS-CoV-2. Removal of virions by these porous membranes (that is, MF > 50 nm and UF 2–50 nm) is feasible, albeit highly dependent on the pore size distribution in relation to the size of the target virus110,115. Thus, SARS-CoV-2 with a diameter of ~100 nm should be removed reliably by UF. Virion removal may be further enhanced, depending on surface characteristics of both membranes and SARS-CoVs (that is, hydrophobic and charged regions on the envelope), which can lead to removal beyond size exclusion due to electrostatic and hydrophobic interactions104,115. Application of UF in membrane bioreactors (MBRs) further increases viral removal (not specific to SARS-CoVs) via a combination of three mechanisms: steric removal, adsorption and inactivation during biological treatment104,105,115. Consequently, MBRs have shown increased removal of enteric viruses (up to 6.8-log removal) in comparison to conventional WWTPs (up to 3.6-log removal)115. Although less common in wastewater treatment, high-pressure membrane systems using denser and tighter membranes (pore size <2 nm), such as nanofiltration (NF), reverse osmosis (RO) and forward osmosis (FO) membranes should accomplish complete removal of SARS-CoVs110,116.
Critical knowledge gaps and recommendations
The health risks of COVID-19 via waterborne transmission may be greater than initially assumed, and wastewater should be further studied as a potential pathway for COVID-19 transmission. Evidence for the presence of SARS-CoV-2 RNA in wastewater systems is accumulating around the world. The large number of infected individuals in the current pandemic together with the high infectivity of SARS-CoV-2 could present a new challenge for wastewater treatment and calls for future assessment of the risk for transmission via wastewater reuse. Such risks might be expected to be highest in areas with high population density, direct exposure to aerosolized wastewater as well as regions that lack adequate wastewater collection, treatment and disinfection.
Extensive research into the frequency of detecting the infectious SARS-CoV-2 in wastewater is urgently needed to gain: (1) critical information on the abundance of viruses in raw wastewater, treated wastewater and the receiving environment, which can be used to generate a quantitative risk assessment; (2) information on removal efficiencies through the wastewater treatment train; (3) disinfection requirements according to virus loading and transmission via WWTPs to ensure complete removal of SARS-CoV-2 for wastewater reuse; and (4) epidemic surveillance for policymakers on the outbreak, extent and prevalence of the COVID-19 pandemic within the community.
Characterizing and mitigating any identified risks of SARS-CoV-2 transmission via waterborne pathways will require addressing the following knowledge gaps: (1) SARS-CoV-2 concentrations in wastewater are currently estimated by molecular approaches that quantify viral RNA rather than infective virions. Whether these approaches predominantly quantify fully functional virions rather than viral RNA fragments remains to be determined. (2) The minimal infectious dose of SARS-CoV-2 from water and aerosols is currently unknown. (3) The extent of SARS-CoV-2 dissemination via waterborne pathways is not clear, and neither is the associated SARS-CoV-2 loading of freshwater environments and water reuse systems (for example, for recreation, cooling and agriculture).
Overall, this Review highlights the urgent need for enhanced monitoring, risk assessment and new risk management strategies for COVID-19 in wastewater. Developing innovative tools for environmental monitoring will provide necessary scientific evidence for policymakers, while optimized disinfection strategies will mitigate COVID-19 transmission associated with wastewater reuse. Beyond COVID-19, these approaches will also improve detection, response and containment of future viral disease outbreaks.
References
Coronavirus Disease Situation Reports (World Health Organization, accessed 28 July 2020); https://go.nature.com/3aahjbg
Naddeo, V. & Liu, H. Editorial perspectives: 2019 novel coronavirus (SARS-CoV-2): what is its fate in urban water cycle and how can the water research community respond? Environ. Sci. Water Res. Technol. 6, 1213–1216 (2020).
Ashour, H. M., Elkhatib, W. F., Rahman, M. M. & Elshabrawy, H. A. Insights into the recent 2019 novel Coronavirus (SARS-CoV-2) in light of past human coronavirus outbreaks. Pathogens 9, 186 (2020).
Kim, Y.-I. et al. Infection and rapid transmission of SARS-CoV-2 in ferrets. Cell Host Microbe 27, 704–709 (2020).
Kampf, G., Todt, D., Pfaender, S. & Steinmann, E. Persistence of coronaviruses on inanimate surfaces and their inactivation with biocidal agents. J. Hosp. Infect. 104, 246–251 (2020).
Chan, J. F.-W. et al. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster. Lancet 395, 514–523 (2020).
Water, Sanitation, Hygiene and Waste Management for the COVID-19 Virus: Interim Guidance (World Health Organization, 2020).
Modes of Transmission of Virus causing COVID-19: Implications for IPC Precaution Recommendations. Scientific Brief (World Health Organization, 2020).
McKinney, K. R., Yang Gong, Y. & Lewis, T. G. Environmental transmission of SARS at Amoy Gardens. J. Environ. Health 68, 26–30 (2006).
Yu, I. T. S. et al. Evidence of airborne transmission of the severe acute respiratory syndrome virus. N. Engl. J. Med. 350, 1731–1739 (2004).
Ahmed, W. et al. First confirmed detection of SARS-CoV-2 in untreated wastewater in Australia: a proof of concept for the wastewater surveillance of COVID-19 in the community. Sci. Total Environ. 728, 138764 (2020).
Cheung, K. S. et al. Gastrointestinal manifestations of SARS-CoV-2 infection and virus load in fecal samples from the Hong Kong cohort and systematic review and meta-analysis. Gastroenterology 159, 81–95 (2020).
Medema, G., Heijnen, L., Elsinga, G., Italiaander, R. & Brouwer, A. Presence of SARS-Coronavirus-2 in sewage and correlation with reported COVID-19 prevalence in the early stage of the epidemic in the Netherlands. Environ. Sci. Technol. Lett. 7, 511–516 (2020).
Randazzo, W. et al. SARS-CoV-2 RNA titers in wastewater anticipated COVID-19 occurrence in a low prevalence area. Water Res. 181, 115942 (2020).
Mauter, M. S. et al. The role of nanotechnology in tackling global water challenges. Nat. Sustain. 1, 166–175 (2018).
Greve, P. et al. Global assessment of water challenges under uncertainty in water scarcity projections. Nat. Sustain. 1, 486–494 (2018).
Liu, L. et al. The importance of system configuration for distributed direct potable water reuse. Nat. Sustain. 3, 548–555 (2020).
Wigginton, K. R., Ye, Y. & Ellenberg, R. M. Emerging investigators series: the source and fate of pandemic viruses in the urban water cycle. Environ. Sci. Water Res. Technol. 1, 735–746 (2015).
Kitajima, M. et al. SARS-CoV-2 in wastewater: state of the knowledge and research needs. Sci. Total Environ. 739, 139076 (2020).
Wurtzer, S. et al. Evaluation of lockdown impact on SARS-CoV-2 dynamics through viral genome quantification in Paris wastewaters. Preprint at medRxiv https://doi.org/10.1101/2020.04.12.20062679 (2020).
Carducci, A., Battistini, R., Rovini, E. & Verani, M. Viral removal by wastewater treatment: Monitoring of indicators and pathogens. Food Environ. Virol. 1, 85–91 (2009).
Qiu, Y. et al. Assessment of human virus removal during municipal wastewater treatment in Edmonton, Canada. J. Appl. Microbiol. 119, 1729–1739 (2015).
Ramia, S. Transmission of viral infections by the water route: implications for developing countries. Rev. Infect. Dis. 7, 180–188 (1985).
Progress on Drinking Water, Sanitation and Hygiene: 2017 Update and SDG Baselines (WASH, WHO & UNICEF, 2017).
Xu, Z. et al. Urban river pollution control in developing countries. Nat. Sustain. 2, 158–160 (2019).
Madigan, M. T., Martinko, J. M., Parker, J. & Brock, T. D. Brock Biology of Microorganisms (Pearson Education, 2012).
Bar-On, Y. M., Flamholz, A., Phillips, R. & Milo, R. SARS-CoV-2 (COVID-19) by the numbers. eLife 9, e57309 (2020).
Yan, R. et al. Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 367, 1444–1448 (2020).
Decaro, N. & Buonavoglia, C. An update on canine coronaviruses: viral evolution and pathobiology. Vet. Microbiol. 132, 221–234 (2008).
Zang, R. et al. TMPRSS2 and TMPRSS4 mediate SARS-CoV-2 infection of human small intestinal enterocytes. Sci. Immunol. 5, eabc3582 (2020).
Zhang, H. et al. The digestive system is a potential route of 2019-nCoV infection: a bioinformatics analysis based on single-cell transcriptomes. Preprint at bioRxiv https://doi.org/10.1101/2020.01.30.927806 (2020).
Sun, J. et al. Isolation of infectious SARS-CoV-2 from urine of a COVID-19 patient. Emerg. Microbes Infect. 9, 991–993 (2020).
Chen, N. et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet 395, 507–513 (2020).
Lo, I. L. et al. Evaluation of SARS-CoV-2 RNA shedding in clinical specimens and clinical characteristics of 10 patients with COVID-19 in Macau. Int. J. Biol. Sci. 16, 1698–1707 (2020).
Wölfel, R. et al. Virological assessment of hospitalized patients with COVID-2019. Nature 581, 465–469 (2020).
Chen, Y. et al. The presence of SARS-CoV-2 RNA in feces of COVID-19 patients. J. Med. Virol. 92, 833–840 (2020).
Xing, Y. H. et al. Prolonged viral shedding in feces of pediatric patients with coronavirus disease 2019. J. Microbiol. Immunol. Infect. 53, 473–480 (2020).
Xiao, F. et al. Evidence for gastrointestinal infection of SARS-CoV-2. Gastroenterology 158, 1831–1833 (2020).
Wang, W. et al. Detection of SARS-CoV-2 in different types of clinical specimens. J. Am. Med. Assoc. 323, 1843–1844 (2020).
Zhang, Y. et al. Isolation of 2019-nCoV from a stool specimen of a laboratory-confirmed case of the coronavirus disease 2019 (COVID-19). China CDC Weekly 2, 123–124 (2020).
Randazzo, W., Ferrando, E. C., Sanjuán, R., Domingo-Calap, P. & Sánchez, G. Metropolitan wastewater analysis for COVID-19 epidemiological surveillance. Preprint at medRxiv https://doi.org/10.1101/2020.04.23.20076679 (2020).
La Rosa, G. et al. First detection of SARS-CoV-2 in untreated wastewaters in Italy. Sci. Total Environ. 736, 139652 (2020).
Nemudryi, A. et al. Temporal detection and phylogenetic assessment of SARS-CoV-2 in municipal wastewater. Preprint at medRxiv https://doi.org/10.1101/2020.04.15.20066746 (2020).
Danchin, A., Ng, P. T. W. & Turinici, G. A new transmission route for the propagation of the SARS-CoV-2 coronavirus. Preprint at medRxiv https://doi.org/10.1101/2020.02.14.20022939 (2020).
Wu, F. Q. et al. SARS-CoV-2 titers in wastewater are higher than expected from clinically confirmed cases. Preprint at medRxiv https://doi.org/10.1101/2020.04.05.20051540 (2020).
Chin, A. W. H. et al. Stability of SARS-CoV-2 in different environmental conditions. Lancet Microbe 1, e10 (2020).
Schmidt, N. J., Ho, H. H., Riggs, J. L. & Lennette, E. H. Comparative sensitivity of various cell culture systems for isolation of viruses from wastewater and fecal samples. Appl. Environ. Microbiol. 36, 480–486 (1978).
Xagoraraki, I., Yin, Z. & Svambayev, Z. Fate of viruses in water systems. J. Environ. Eng. 140, 04014020 (2014).
Greening, G. E., Hewitt, J. & Lewis, G. D. Evaluation of integrated cell culture-PCR (C-PCR) for virological analysis of environmental samples. J. Appl. Microbiol. 93, 745–750 (2002).
Wang, X. W. et al. Concentration and detection of SARS coronavirus in sewage from Xiao Tang Shan Hospital and the 309th Hospital. J. Virol. Methods 128, 156–161 (2005).
Bofill-Mas, S. & Rusiñol, M. Recent trends on methods for the concentration of viruses from water samples. Curr. Opin. Environ. Sci. Heal. 16, 7–13 (2020).
La Rosa, G., Bonadonna, L., Lucentini, L., Kenmoe, S. & Suffredini, E. Coronavirus in water environments: pccurrence, persistence and concentration methods - a scoping review. Water Res. 179, 115899 (2020).
Araud, E., Fuzawa, M., Shisler, J. L., Li, J. & Nguyen, T. H. UV inactivation of rotavirus and Tulane virus targets different components of the virions. Appl. Environ. Microbiol. 86, e02436–19 (2020).
Peccia, J. et al. SARS-CoV-2 RNA concentrations in primary municipal sewage sludge as a leading indicator of COVID-19 outbreak dynamics. Preprint at medRxiv https://doi.org/10.1101/2020.05.19.20105999 (2020).
Callanan, J. et al. Expansion of known ssRNA phage genomes: from tens to over a thousand. Sci. Adv 6, eaay5981 (2020).
Martínez-Puchol, S. et al. Characterisation of the sewage virome: comparison of NGS tools and occurrence of significant pathogens. Sci. Total Environ. 713, 136604 (2020).
Ye, Y., Ellenberg, R. M., Graham, K. E. & Wigginton, K. R. Survivability, partitioning, and recovery of enveloped viruses in untreated municipal wastewater. Environ. Sci. Technol. 50, 5077–5085 (2016).
Watanabe, T., Bartrand, T. A., Weir, M. H., Omura, T. & Haas, C. N. Development of a dose-response model for SARS coronavirus. Risk Anal. 30, 1129–1138 (2010).
Lindsley, W. G. et al. Quantity and size distribution of cough-generated aerosol particles produced by influenza patients during and after illness. J. Occup. Environ. Hyg. 9, 443–449 (2012).
Geller, C., Varbanov, M. & Duval, R. E. Human coronaviruses: insights into environmental resistance and its influence on the development of new antiseptic strategies. Viruses 4, 3044–3068 (2012).
Wang, X.-W. et al. Study on the resistance of severe acute respiratory syndrome-associated coronavirus. J. Virol. Methods 126, 171–177 (2005).
Duan, S.-M. et al. Stability of SARS coronavirus in human specimens and environment and its sensitivity to heating and UV irradiation. Biomed. Environ. Sci. 16, 246–255 (2003).
John, D. E. & Rose, J. B. Review of factors affecting microbial survival in groundwater. Environ. Sci. Technol. 39, 7345–7356 (2005).
Casanova, L., Rutala, W. A., Weber, D. J. & Sobsey, M. D. Survival of surrogate coronaviruses in water. Water Res. 43, 1893–1898 (2009).
Gundy, P. M., Gerba, C. P. & Pepper, I. L. Survival of coronaviruses in water and wastewater. Food Env. Virol. 1, 10–14 (2009).
Lai, M. Y. Y., Cheng, P. K. C. & Lim, W. W. L. Survival of severe acute respiratory syndrome coronavirus. Clin. Infect. Dis. 41, 67–71 (2005).
Berchenko, Y. et al. Estimation of polio infection prevalence from environmental surveillance data. Sci. Transl. Med. 9, eaaf6786 (2017).
Bettencourt, L. M. A., Lobo, J., Helbing, D., Kühnert, C. & West, G. B. Growth, innovation, scaling, and the pace of life in cities. Proc. Natl Acad. Sci. USA 104, 7301–7306 (2007).
Yang, S. et al. Functional topology of evolving urban drainage networks. Water Resour. Res. 53, 8966–8979 (2017).
Mallapaty, S. How sewage could reveal true scale of coronavirus outbreak. Nature 580, 176–177 (2020).
Hart, O. E. & Halden, R. U. Computational analysis of SARS-CoV-2/COVID-19 surveillance by wastewater-based epidemiology locally and globally: feasibility, economy, opportunities and challenges. Sci. Total Environ. 730, 138875 (2020).
Lodder, W. & de Roda Husman, A. M. SARS-CoV-2 in wastewater: potential health risk, but also data. Lancet Gastroenetrol. Hepatol. 5, 533–534 (2020).
Progress on Household Drinking Water, Sanitation and Hygiene 2000-2017 (WHO/UNICEF, 2019).
Berendes, D. M., Sumner, T. A. & Brown, J. M. Safely managed sanitation for all means fecal sludge management for at least 1.8 billion people in low and middle income countries. Environ. Sci. Technol. 51, 3074–3083 (2017).
Heller, L., Mota, C. R. & Greco, D. B. COVID-19 faecal-oral transmission: are we asking the right questions? Sci. Total Environ. 729, 138919 (2020).
Usman, M., Farooq, M. & Hanna, K. Existence of SARS-CoV-2 in wastewater: implications for its environmental transmission in developing communities. Environ. Sci. Technol. 54, 7758–7759 (2020).
The Global Risks Report 2020 (World Health Organization, 2020).
Water, Sanitation, Hygiene and Waste Management for COVID-19: Technical Brief (World Health Organization, 2020).
Adegoke, A. A., Amoah, I. D., Stenström, T. A., Verbyla, M. E. & Mihelcic, J. R. Epidemiological evidence and health risks associated with agricultural reuse of partially treated and untreated wastewater: a review. Front. Public Health 6, 337 (2018).
Dickin, S. K., Schuster-Wallace, C. J., Qadir, M. & Pizzacalla, K. A review of health risks and pathways for exposure to wastewater use in agriculture. Environ. Health Perspect. 124, 900–909 (2016).
Mullis, L., Saif, L. J., Zhang, Y., Zhang, X. & Azevedo, M. S. P. Stability of bovine coronavirus on lettuce surfaces under household refrigeration conditions. Food Microbiol. 30, 180–186 (2012).
Yépiz-Gómez, M. S., Gerba, C. P. & Bright, K. R. Survival of respiratory viruses on fresh produce. Food Environ. Virol. 5, 150–156 (2013).
Gormley, M., Aspray, T. J. & Kelly, D. A. COVID-19: mitigating transmission via wastewater plumbing systems. Lancet Glob. Health 8, E643 (2020).
Brisebois, E. et al. Human viral pathogens are pervasive in wastewater treatment center aerosols. J. Environ. Sci. 67, 45–53 (2018).
Courault, D. et al. Assessment and risk modeling of airborne enteric viruses emitted from wastewater reused for irrigation. Sci. Total Environ. 592, 512–526 (2017).
Ding, Z. et al. Toilets dominate environmental detection of SARS-CoV-2 virus in a hospital. Preprint at medRxiv https://doi.org/10.1101/2020.04.03.20052175 (2020).
Ijaz, M. K., Brunner, A. H., Sattar, S. A., Nair, R. C. & Johnson-Lussenburg, C. M. Survival characteristics of airborne human coronavirus 229E. J. Gen. Virol. 66, 2743–2748 (1985).
van Doremalen, N. et al. Aerosol and surface stability of SARS-CoV-2 as compared with SARS-CoV-1. N. Engl. J. Med. 382, 1564–1567 (2020).
Fears, A. C. et al. Comparative dynamic aerosol efficiencies of three emergent coronaviruses and the unusual persistence of SARS-CoV-2 in aerosol suspensions. Preprint at medRxiv https://doi.org/10.1101/2020.04.13.20063784 (2020).
Barker, J. & Jones, M. V. The potential spread of infection caused by aerosol contamination of surfaces after flushing a domestic toilet. J. Appl. Microbiol. 99, 339–347 (2005).
Zabinski, J. W., Pieper, K. J. & MacDonald Gibson, J. A Bayesian belief network model assessing the risk to wastewater workers of contracting Ebola virus disease during an outbreak. Risk Anal. 38, 376–391 (2018).
Lin, K. & Marr, L. C. Aerosolization of Ebola virus surrogates in wastewater systems. Environ. Res. Lett. 51, 2669–2675 (2017).
Rimoldi, S. G. et al. Presence and infectivity of SARS-CoV-2 virus in wastewaters and rivers. Sci. Total Environ. 744, 140911 (2020).
Haramoto, E., Malla, B., Thakali, O. & Kitajima, M. First environmental surveillance for the presence of SARS-CoV-2 RNA in wastewater and river water in Japan. Sci. Total Environ. 737, 140405 (2020).
Hata, A. et al. Effects of rainfall events on the occurrence and detection efficiency of viruses in river water impacted by combined sewer overflows. Sci. Total Environ. 468–469, 757–763 (2014).
Katz, A. et al. Heteroaggregation of an enveloped bacteriophage with colloidal sediments and effect on virus viability. Sci. Total Environ. 637–638, 104–111 (2018).
Kim, J. Y., Lee, H., Lee, J. E., Chung, M.-S. & Ko, G. P. Identification of human and animal fecal contamination after rainfall in the Han river, Korea. Microbes Environ. 28, 187–194 (2013).
Bhattacharjee, S., Ryan, J. N. & Elimelech, M. Virus transport in physically and geochemically heterogeneous subsurface porous media. J. Contam. Hydrol. 57, 161–187 (2002).
Gutierrez, L. & Nguyen, T. H. Interactions between rotavirus and natural organic matter isolates with different physicochemical characteristics. Langmuir 29, 14460–14468 (2013).
Weisbrod, N., Meron, H., Walker, S. & Gitis, V. Virus transport in a discrete fracture. Water Res. 47, 1888–1898 (2013).
Elkayam, R. et al. Viral and microbial pathogens, indicator microorganisms, microbial source tracking indicators, and antibiotic resistance genes in a confined managed effluent recharge system. J. Environ. Eng. 144, 05017011 (2018).
Hurst, C. J. & Gerba, C. P. Fate of viruses during wastewater sludge treatment processes. Crit. Rev. Environ. Sci. Technol. 18, 317–343 (1989).
Clarke, N. A., Stevenson, R. E., Lu Chang, S. & Kabler, P. W. Removal of enteric viruses from sewage by activated sludge treatment. Am. J. Public Health 51, 1118–1129 (1961).
Chaudhry, R. M., Nelson, K. L. & Drewes, J. E. Mechanisms of pathogenic virus removal in a full-scale membrane bioreactor. Environ. Sci. Technol. 49, 2815–2822 (2015).
Lv, W. et al. Virus removal performance and mechanism of a submerged membrane bioreactor. Process Biochem. 41, 299–304 (2006).
Bibby, K. & Peccia, J. Identification of viral pathogen diversity in sewage sludge by metagenome analysis. Environ. Sci. Technol 47, 1945–1951 (2013).
Goddard, M. R. R., Bates, J. & Butler, M. Recovery of indigenous enteroviruses from raw and digested sewage sludges. Appl. Environ. Microbiol. 42, 1023–1028 (1981).
Viau, E., Bibby, K., Paez-Rubio, T. & Peccia, J. Toward a consensus view on the infectious risks associated with land application of sewage sludge. Environ. Sci. Technol. 45, 5459–5469 (2011).
Ye, Y., Chang, P. H., Hartert, J. & Wigginton, K. R. Reactivity of enveloped virus genome, proteins, and lipids with free chlorine and UV 254. Environ. Sci. Technol. 52, 7698–7708 (2018).
Tchobanoglous, G., Burton, F. L. & Stensel, H. D. Wastewater Engineering, Treatment and Reuse (McGraw-Hill, 2004).
Simmons, F. J. & Xagoraraki, I. Release of infectious human enteric viruses by full-scale wastewater utilities. Water Res. 45, 3590–3598 (2011).
Derraik, J. G. B., Anderson, W. A., Connelly, E. A. & Anderson, Y. C. Rapid evidence summary on SARS-CoV-2 survivorship and disinfection, and a reusable PPE protocol using a double-hit process. Preprint at medRxiv https://doi.org/10.1101/2020.04.02.20051409 (2020).
von Gunten, U. & von Sonntag, C. Chemistry of Ozone in Water and Wastewater Treatment from Basic Principles (IWA Publishing, 2012).
Sonntag, C. Free-Radical-Induced DNA Damage and Its Repair (Springer, 2006).
Bodzek, M., Konieczny, K. & Rajca, M. Membranes in water and wastewater disinfection – review. Arch. Environ. Prot. 45, 3–18 (2019).
Pendergast, M. M. & Hoek, E. M. V. A review of water treatment membrane nanotechnologies. Energy Environ. Sci. 4, 1946–1971 (2011).
Martina, B. E. E. et al. SARS virus infection of cats and ferrets. Nature 425, 915 (2003).
DeDiego, M. L. et al. Pathogenicity of severe acute respiratory coronavirus deletion mutants in hACE-2 transgenic mice. Virology 376, 379–389 (2008).
Deng, W. et al. Ocular conjunctival inoculation of SARS-CoV-2 can cause mild COVID-19 in rhesus macaques. Preprint at bioRxiv https://doi.org/10.1101/2020.03.13.990036 (2020).
Munster, V. J. et al. Respiratory disease in rhesus macaques inoculated with SARS-CoV-2. Nature https://doi.org/10.1038/s41586-020-2324-7 (2020).
Bao, L. et al. The pathogenicity of SARS-CoV-2 in hACE2 transgenic mice. Nature 583, 830–833 (2020).
Douglas, M. G., Kocher, J. F., Scobey, T., Baric, R. S. & Cockrell, A. S. Adaptive evolution influences the infectious dose of MERS-CoV necessary to achieve severe respiratory disease. Virology 517, 98–107 (2018).
Yezli, S. & Otter, J. A. Minimum infective dose of the major human respiratory and enteric viruses transmitted through food and the environment. Food Environ. Virol. 3, 1–30 (2011).
Converting TCID50 to plaque forming units PFU-124 (ATCC, accessed 19 April 2020); https://go.nature.com/3aesQpX
Taubenberger, J. K. & Morens, D. M. 1918 Influenza: the mother of all pandemics. Emerg. Infect. Dis. 12, 15–22 (2006).
Arena, C. et al. Simultaneous investigation of influenza and enteric viruses in the stools of adult patients consulting in general practice for acute diarrhea. Virol. J. 9, 1–8 (2012).
Wong, G. et al. MERS, SARS, and Ebola: the role of super-spreaders in infectious disease. Cell Host Microbe 18, 398–401 (2015).
Lai, S. et al. Global epidemiology of avian influenza A H5N1 virus infection in humans, 1997-2015: a systematic review of individual case data. Lancet Infect. Dis. 16, E108–E118 (2016).
Dawood, F. S. et al. Estimated global mortality associated with the first 12 months of 2009 pandemic influenza A H1N1 virus circulation: a modelling study. Lancet Infect. Dis. 12, 687–695 (2012).
Durai, P., Batool, M., Shah, M. & Choi, S. Middle East respiratory syndrome coronavirus: transmission, virology and therapeutic targeting to aid in outbreak control. Exp. Mol. Med. 47, e181 (2015).
Yeo, C., Kaushal, S. & Yeo, D. Enteric involvement of coronaviruses: is faecal-oral transmission of SARS-CoV-2 possible? Lancet 5, 335–337 (2020).
Su, S. et al. Epidemiology, evolution, and pathogenesis of H7N9 influenza viruses in five epidemic waves since 2013 in China. Trends Microbiol. 25, 713–728 (2017).
Zika Cases 2015 - 2018 (Pan American Health Organzation & World Health Organization, 2018).
Li, C. et al. Zika virus shedding in the stool and infection through the anorectal mucosa in mice. Emerg. Microbes Infect. 7, 1–10 (2018).
Rabenau, H. F. et al. Stability and inactivation of SARS coronavirus. Med. Microbiol. Immunol. 194, 1–6 (2005).
Sizun, J., Yu, M. W. N. & Talbot, P. J. Survival of human coronaviruses 229E and OC43 in suspension and after drying on surfaces: a possible source of hospital-acquired infections. J. Hosp. Infect. 46, 55–60 (2000).
Acknowledgements
We acknowledge the assistance of the Dangerous Pathogens team (Health & Biosecurity, CSIRO Australia) for virus propagation within Physical Containment Level 4 facilities. S.Y. received funding from the European Union Programme Horizon 2020 (no. 776816) for the project PROJECT Ô: ‘Demonstration of planning and technology tools for a circular, integrated and symbiotic use of water’. A.P. was supported by US National Science Foundation award no. CBET-1848683.
Author information
Authors and Affiliations
Contributions
E.B.-Z. and A.B. conceptualized the initial structure of this Review, with guidance from A.P., P.J.A., M.E. and E.M. All authors were involved in writing and editing the manuscript, and provided data interpretation and critical insights. J.L.B. provided the phylogenetic analysis of SARS-CoVs and S.C. generated the transmission electron microscopy (TEM) image of SARS-CoV-2.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Bogler, A., Packman, A., Furman, A. et al. Rethinking wastewater risks and monitoring in light of the COVID-19 pandemic. Nat Sustain 3, 981–990 (2020). https://doi.org/10.1038/s41893-020-00605-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41893-020-00605-2
This article is cited by
-
Destiny of emerging viruses: a systematic review
International Journal of Environmental Science and Technology (2024)
-
Solar-driven efficient heterogeneous subminute water disinfection nanosystem assembled with fingerprint MoS2
Nature Water (2023)
-
Enhanced detection of viruses for improved water safety
Scientific Reports (2023)
-
Sustainable water captured from air for fulfilling the SDGs
Environment, Development and Sustainability (2023)
-
Hybrid energy harvesting systems for self-powered sustainable water purification by harnessing ambient energy
Frontiers of Environmental Science & Engineering (2023)