Abstract
Sleep is essential to life. Accurate measurement and classification of sleep/wake and sleep stages is important in clinical studies for sleep disorder diagnoses and in the interpretation of data from consumer devices for monitoring physical and mental well-being. Existing non-polysomnography sleep classification techniques mainly rely on heuristic methods developed in relatively small cohorts. Thus, we aimed to establish the accuracy of wrist-worn accelerometers for sleep stage classification and subsequently describe the association between sleep duration and efficiency (proportion of total time asleep when in bed) with mortality outcomes. We developed a self-supervised deep neural network for sleep stage classification using concurrent laboratory-based polysomnography and accelerometry. After exclusion, 1113 participant nights of data were used for training. The difference between polysomnography and the model classifications on the external validation was 48.2 min (95% limits of agreement (LoA): −50.3 to 146.8 min) for total sleep duration, −17.1 min for REM duration (95% LoA: −56.7 to 91.0 min) and 31.1 min (95% LoA: −67.3 to 129.5 min) for NREM duration. The sleep classifier was deployed in the UK Biobank with ~100,000 participants to study the association of sleep duration and sleep efficiency with all-cause mortality. Among 66,262 UK Biobank participants, 1644 mortality events were observed. Short sleepers (<6 h) had a higher risk of mortality compared to participants with normal sleep duration 6–7.9 h, regardless of whether they had low sleep efficiency (Hazard ratios (HRs): 1.36; 95% confidence intervals (CIs): 1.18 to 1.58) or high sleep efficiency (HRs: 1.29; 95% CIs: 1.04–1.61). Deep-learning-based sleep classification using accelerometers has a fair to moderate agreement with polysomnography. Our findings suggest that having short overnight sleep confers mortality risk irrespective of sleep continuity.
Similar content being viewed by others
Introduction
Sleep is essential to life and is structurally complex. Humans spend approximately one third of their lives asleep, yet sleep is hard to assess in free-living environments1. Our understanding of how sleep is associated with health and morbidity primarily draws on studies that use self-report sleep diaries, which capture the subjective experience2. However, sleep diaries have a low correlation with objective device-measured sleep parameters3,4. The accepted standard for sleep measurement is laboratory-based polysomnography, which monitors sleep using a range of physical and physiological signals. However, polysomnography is not feasible for use at scale due to its high cost and technical complexity. Instead, wrist-worn accelerometers are more viable to deploy in large-scale epidemiological studies because of their portability and low user burden.
Despite the popularity of sleep monitoring in consumer and research-grade wrist-worn devices, sleep assessment algorithms are frequently proprietary and validated in small populations, making their measurement validity unclear5,6,7,8. Methods for Sleep classification (i.e. defining periods of wake, NREM and REM sleep) primarily rely on hand-crafted spatiotemporal features such as device angle, which may not make full use of all the information in the signals. Hence, data-driven methods like deep learning could be advantageous. Furthermore, existing actigraphy-based sleep studies on large health datasets have only focused on the differentiation between sleep and wakefulness4,9,10,11 without evaluating variations in the stages of sleep.
We therefore set out to: (1) develop and internally validate an open-source novel deep learning method to infer sleep stages from wrist-worn accelerometers, (2) externally validate our proposed algorithm together with existing sleep staging benchmarks and (3) investigate the association between device-measured overnight sleep duration and efficiency with all-cause mortality.
Results
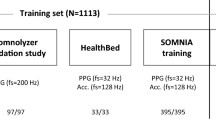
In our multicentre cohort study, we developed and tested a sleep staging model for accelerometers (SleepNet) using a self-supervised deep recurrent neural network. We designed the model to classify each 30-s window of accelerometry data into one of the three sleep stages, wake, rapid-eye-movement sleep (REM) and non-rapid-eye movement sleep (NREM). Figure 1 illustrates the three main steps in our study: (1) feature extraction from unlabelled free-living data, (2) sleep staging model development and (3) face validity assessment and health association analysis using the machine learning-estimated sleep parameters.
1. We use multi-task self-supervised learning to obtain a feature extractor by learning from 700,000 person-days of tri-axial accelerometry data in the UK Biobank. 2. The pre-trained feature extractor was then fine-tuned with a deep recurrent network to train a sleep-stage classifier using polysomnography as the ground truth. 3. We deploy the sleep prediction model on the UK Biobank and investigate the association between device-measured sleep and mortality outcomes.
Comparison to polysomnography
After preprocessing, 1113 participants were included in the internal validation and 53 participants were included in the external validation. Our proposed deep recurrent neural network (SleepNet) pre-trained with self-supervision achieved the best performance when compared with other baseline models that used hand-crafted features (Supplementary Table 6).
On the internal validation, SleepNet had a mean bias of 9.9 min (95% limits of agreement (LoA): −100.5–120.4 min) for total sleep duration, −24.4 min (95% LoA: −136.7–87.8 min) for REM duration and 34.4 min (95% LoA: −106.4–175.1 min) for NREM duration (Fig. 2). In comparison, on the external validation, the mean bias was 48.2 min (95% LoA: −50.3–146.8 min) for total sleep duration, −17.1 min (95% LoA: −56.7–91.0 min) for REM duration and 31.1 min (95% LoA: −67.3–129.5 min) for NREM duration. Overall, our model tends to underestimate REM and short sleep and overestimate NREM and long sleep. Supplementary Figs. 5–10 depict the agreement assessments for other sleep parameters on the individual cohorts.
a the agreement for the internal validation; b the agreement for the external validation. The internal validation consists of 1113 polysomnography nights from the Raine Study and the Newcastle cohort, whereas the external validation consists of 53 polysomnography nights from the Leicester and Pennsylvania cohorts.
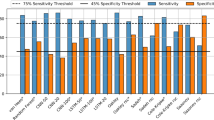
The subject-wise performance for both the internal and external validation using the pre-trained SleepNet is shown in Supplementary Table 7. On the pooled internal validation, our model obtained an F1 of 0.75 ± 0.1 in the two-class setting (sleep/wake) and an F1 of 0.57 ± 0.11 in the three-class setting (wake/REM/NREM). The agreement decreased slightly on the external validation with an F1 of 0.66 ± 0.12 in the two-class setting (sleep/wake) and an F1 of 0.49 ± 0.10 in the three-class setting (wake/REM/NREM). In the Newcastle cohort, for the sleep/wake classification, sensitivity decreased and specificity increased in participants with sleep disorders. No obvious difference was observed in both Raine Gen1 and Gen2 cohorts when the participants were stratified by sex, BMI, AHI and sleep disorder conditions. (Supplementary Tables 8–10).
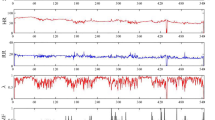
To classify any given window in an epoch-by-epoch fashion, the SleepNet achieved a Kappa score of 0.39 on the internal validation set and a Kappa score of 0.32 on the external validation set in the three-class setting (Supplementary Fig. 11). Cohort-specific confusion matrices can be found in Supplementary Figs. 12–15. Supplementary Fig. 16 visualises a one-night sample actigram, its ground-truth polysomnography labels and SleepNet predictions. We used SleepNet to generate all the sleep parameters for the rest of the paper.
Face validity in the UK Biobank
Before deploying the SleepNet on the UK Biobank, we excluded participants with unusable accelerometer data and participants with missing covariates in the descriptive analysis. We further excluded participants with any prior hospitalisation for cardiovascular disease or cancer in the association analysis (Supplementary Fig. 17). In sum, 66,262 participants were included in the final analysis.
Table 1 describes the variations in overnight sleep duration, REM and NREM durations and sleep efficiency across population subgroups in the UK Biobank. Older participants generally slept longer with higher sleep efficiency. Females had a longer overnight sleep duration, REM and NREM durations. Participants with better self-rated health had longer sleep duration and higher sleep efficiency than those with poor self-rated health. Sleep efficiency was relatively stable across different seasons and days of the week. The correlation coefficients between device-measured sleep parameters during accelerometer wear and self-reported total sleep duration at baseline assessment were all below 0.25 (Supplementary Fig. 18). The distributions of device-measured overnight sleep duration tend to have a greater variability for participants who self-reported to have less than 5 or greater than 10 h of total sleep duration (Supplementary Fig. 19). Overall, older participants have a shorter REM sleep than younger participants (Supplementary Fig. 20). No major differences were seen between females and males.
We found expected sleep-wake patterns in population subgroups. For example, timing of the sleep opportunity for participants with a self-reported ‘morning’ chronotype was about 1 h earlier when compared with those that had a self-reported ‘evening’ chronotype (Fig. 3a). We saw similar but shorter phase advance (~30 min) in participants who were most physically active compared to the participants that were least physically active (Fig. 3b). When comparing groups that had a history of self-reported insomnia symptoms versus those who did not, we found that participants with a history of insomnia symptoms were more likely to be in REM sleep on average during the overnight sleep window (Fig. 3c, d). Participants with a history of self-reported insomnia symptoms tended to have a longer overnight sleep duration but with a lower sleep efficiency (Supplementary Fig. 21). The sleep architecture for different population subgroups were similar between weekdays and weekends, with a slight phase delay over the weekend (Supplementary Fig. 22).
Top: variations of the average overnight sleep probability for the participants with self-reported ‘morning’ and ‘evening’ chronotype (a) and the overnight sleep distributions across thirds of device-measured physical activity level (b). Bottom: variations of the average REM (c) and NREM (d) probability in participants with a history of self-reported insomnia symptoms versus those without. REM rapid-eye-movement sleep, NREM non-rapid-eye-movement sleep.
Association with all-cause mortality
Over 452,652 years of the follow-up, 1644 mortality events among 66,262 participants were observed. Short sleepers (<6 h) had a higher risk of mortality in groups of low sleep efficiency (Hazard ratios (HRs): 1.36; 95% confidence intervals (CIs): 1.18–1.58) and high sleep efficiency (HRs: 1.29; 95% CIs: 1.04–1.61) compared to participants with normal sleep duration (6–7.9 h, Fig. 4). The risk of all-cause mortality appeared to decrease linearly as sleep efficiency increased. However, a non-linear association was observed in the association for overnight sleep duration (Supplementary Fig. 23). When further adjusted for BMI, associations of overnight sleep duration and sleep efficiency with all-cause mortality were slightly attenuated (Supplementary Figs. 24–25). Longer overnight sleep duration was not found to have a higher risk than the reference group (Supplementary Fig. 23).
a participants with less than 85% sleep efficiency; b participants with greater or equal to 85% sleep efficiency. The model used 1644 events among 66,262 participants. We used age as the timescale and adjusted for sex, ethnicity, Townsend Deprivation Index of baseline address (split by quarter in the study population), educational qualifications, smoking status, alcohol consumption (Never, <3 times/week, 3+ times/week), overall activity (measured in milli-gravity units). The median was used to separate groups with low and high sleep efficiency. Areas of squares represent the inverse of the variance of the log risk. The I bars denote the 95% confidence interval for the floated risks.
Discussion
We have developed, and internally and externally validated a deep learning method to characterise sleep architecture from a wrist-worn accelerometer with competitive performance against 1113 nights of laboratory-based polysomnography recordings. When applying our developed method in the UK Biobank in an epidemiological analysis of 66,214 participants, we found that shorter sleep time was associated with an increased risk of all-cause mortality individually regardless of sleep continuity, indexed by sleep efficiency. Our open-source algorithm and the inferred sleep parameters will open the door to future studies on sleep and sleep architecture using large-scale accelerometer databases.
Our novel self-supervised deep learning sleep staging method outperformed existing baseline methods that rely on hand-crafted features. The inferred sleep architecture estimates had a fair agreement (κ = 0.37) with the polysomnography ground truth on the internal validation12. Unlike previous work in sleep classification methods that depended on hand-crafted features13,14, our proposed method automatically extracted the features using self-supervision, hence removing the need for manual engineering. Even for sleep/wake classification, SleepNet achieved comparable results to a systematic evaluation of eight state-of-the-art sleep algorithms8 in the Newcastle dataset. However, our work offers a more robust evaluation and identifies the upper limit of using accelerometry for sleep classification by developing a model with one of the largest multicentre datasets with polysomnography ground truth, at least ten times the size of existing studies.
In the subsequent epidemiological analysis, we found a clear association between short overnight sleep duration with increased risk of all-cause mortality in both good and poor sleepers defined by sleep efficiency. Short overnight sleep duration has been linked with mortality outcomes in self-report and actigraphy-based studies15,16. However, few studies have investigated the joint effect of sleep duration and efficiency. One recent study has suggested that participants with short and long total sleep time had an increased risk after accounting for sleep efficiency17. However, our analysis did not find that long overnight sleep duration was associated with increased risk, potentially because we did not include daytime naps in our measurement of overnight sleep duration. Daytime napping has been found to be associated with an increased risk of cardiovascular events and deaths in those with longer nighttime sleep18. We did not find a U-shape association between device-measured sleep and mortality that has been suggested by other smaller studies15. Instead, our data are supportive of adverse associations with short sleep duration only, which is concordant with pre-clinical human and animal studies19.
This study has several strengths, including the analysis of sleep architecture in a large, prospective Biobank with longitudinal follow-up. Compared with self-reported sleep questionnaires that only captured sleep duration to the nearest hour, actigraphy-based methods like ours can provide more fine-grained sleep duration and efficiency estimates. The extensive multicentre evaluation of the sleep classification allowed for the characterisation of the measurement uncertainty and a less biased interpretation of the health association analysis. Sleep stage identification from actigraphy is highly challenging, especially for wake periods in bed that are not characterised by wrist movement. With the proposed SleepNet, we could obtain sleep architecture estimates for population health inference after evaluating the face validity of the sleep parameters in the UK Biobank. While future work might improve sleep staging performance by incorporating additional physiological signals, such as electrocardiogram, to improve sleep staging performance, multi-modal sensor signals are not yet available for population-scale studies with longitudinal follow-up beyond a few years20. Despite our best efforts to include diverse validation cohorts from different centres, the included datasets mainly consist of healthy populations from a Caucasian ethnic background. Validation in populations with chronic diseases and different ethnic backgrounds would aid in quantifying the measurement uncertainty. Laboratory-based polysomnography is known to suffer from the first-night effect consisting of a reduction in sleep duration, quality and continuity21. Future validation studies could also assess the within-person variability using multi-night polysomnography.
In this work, we have developed and validated an open-source sleep staging method that substantially improves the ability to measure sleep characteristics with wrist-worn accelerometers in large biomedical datasets. Using the sleep parameters generated by our model, we demonstrated that shorter overnight sleep was associated with a higher risk of all-cause mortality in both good and poor sleepers. Our proposed method provides the community with a rich set of new measurements to study how sleep parameters are longitudinally associated with clinical outcomes.
Methods
Study participants
We used the UK Biobank accelerometry dataset22 for two purposes: learning health-relevant accelerometer features to support the training of the sleep staging model and conducting the downstream health association analyses using the developed sleep staging model.
For sleep staging model development, internal validation consisted of two generations of participants from the Raine Study23,24 and a sleep patient population from the Newcastle cohort25. The Raine Study has followed up roughly 2900 children since 1989 in Australia. A subset of children (Raine Generation 2, Gen2) at the age of 22 and their parents (Raine Generation 1, Gen1) were invited to undergo one night of laboratory-based polysomnography at Western Australia’s Center for Sleep Science. The external validation consisted of two general populations from Leicester26 and Pennsylvania27. Detailed population characteristics and inclusion criteria are listed in Supplementary Section 1.1.
Accelerometer devices and data preprocessing
Three different devices were used to collect the accelerometry for the included datasets, ActiGraph GT3X, Axivity AX3 and GENEActive Original accelerometers. The devices used have been shown to have a high inter-instrument agreement (>80%) in derived sedentary and sleep-related time estimates in free-living environments28. As for device placement, we selected data from the dominant wrist where possible to be consistent with the UK Biobank protocol.
We used the Biobank Accelerometer Analysis Tool29,30 to preprocess all the data. The raw tri-axial accelerometry was first resampled into 30 Hz and clipped to ±3 g. The accelerometry sequence was then divided into consecutive 30-s windows. We considered stationary periods (x/y/z sd < 13 mg) with a duration greater than 60 min as non-wear22. We further excluded the data that could not be parsed, had unrealistic high values (>200 mg), or were poorly calibrated.
Ascertainment of sleep stages via polysomnography
The gold-standard, laboratory-based polysomnography sleep label was aligned with its concurrent accelerometer data as the model ground truth. The polysomnography labels were scored according to the American Academy of Sleep Medicine (AASM) protocol31, which divided sleep into five categories: wake, REM and NREM I, II and III. In total, 1,157,913 (~10,000 h) sleep windows were used to train the network. The sleep stage distributions were similar across all the datasets except for the Newcastle cohort, which had a greater proportion of wakefulness than the others (Supplementary Fig. 1).
Deep learning analysis of sleep stages from wrist-worn accelerometers
A deep recurrent neural network (SleepNet) was trained to classify the sleep stages for every 30-s window of tri-axial accelerometry data. The SleepNet has three components: a ResNet-17 V232 with 1D convolution for feature extraction, a bi-directional Long-Short-Term-Memory (LSTM) network for temporal dependencies learning33 and two fully-connected layers for sleep stage prediction. During training, we provided the SleepNet with five-stage polysomnography labels (wake, REM and NREM I, II, III). When evaluating the model, we collapsed all the NREM stages into one class for classification (wake/REM/NREM). Similarly, we collapsed all the REM and NREM stages together to classify wake vs sleep.
The SleepNet was pre-trained using multi-task self-supervision on the UK Biobank to learn features of human motion dynamics34. Multi-task self-supervision automatically extracts the features relevant to motion by learning to discriminate different spatiotemporal transformations applied to the unlabelled 700,000 person-days of data. Self-supervised pre-training has been shown to help classify human activity recognition not just in healthy but clinical populations35. See Supplementary Section 1.2 for further details of the model development.
For internal validation, we used subject-wise five-fold cross-validation on the Raine Gen2, Raine Gen1 and Newcastle cohorts. For external validation, we trained the SleepNet on all the internal datasets and then evaluated its performance on the Leicester and Pennsylvania cohorts. We compared the SleepNet performance with a random forest model that used the hand-crafted spatiotemporal features13,30. The random forest feature definitions are listed in Supplementary Table 2.
We reported the staging performance in both subject-wise and epoch-to-epoch fashion. Three-class and five-class confusion matrices were plotted for both internal and external validation. Since Cohen Kappa, F1 scores and balanced accuracies (Supplementary Table 3) are less influenced by class imbalance, they were used to evaluate the overall model. To assess the relationship between the model performance and population characteristics, we stratified the subject-wise sleep staging performance by age, sex, employment status, income level, body mass index (BMI), presence and severity of sleep apnoea using the apnoea-hypopnea index (AHI), existing sleep disorders and neurological disorders where available.
Finally, we evaluated the agreement between summary sleep parameters per each night derived from our deep learning method and polysomnography via Bland-Altman plots for the following sleep parameters: total sleep duration, sleep efficiency (proportion of total time asleep when in bed), time awake after sleep onset (WASO), REM duration, NREM duration, REM ratio, NREM ratio. Supplementary Table 4 entails the sleep parameter definitions and their calculations.
Measurements of sleep in 100,000 UK Biobank participants
We obtained the sleep architecture estimates on the UK Biobank by applying SleepNet on the longest overnight sleep windows. Since no concurrent sleep diaries were collected in the UK Biobank, we used a random forest model trained on sleep diaries with Hidden Markov Models smoothing to first obtain time in bed29,30. The random forest model achieved 90%+ precision and recall for detecting sleep windows in 152 free-living participants with sleep diaries that asked two questions: ‘What time did you first fall asleep last night?’ and ‘What time did you wake up (eyes open, ready to get up)?’30. We used the sleep window output from the random forest model as a proxy for the time in bed. We then merged any time in bed windows within 60 min of one another36. Finally, we applied the SleepNet on the longest window over each noon-to-noon interval to estimate the overnight sleep duration. The difference between overnight and total sleep duration is that total sleep duration is a sleep parameter used to assess the agreement between our SleepNet output and polysomnography for model validation. In a single night of polysomnography, the total sleep duration refers to the total time spent in sleep, whereas in a free-living environment, total sleep duration consists of both napping and overnight sleep duration. Overnight sleep duration refers to the estimate for the amount of sleep one obtains for a noon-to-noon interval in a free-living environment using a random forest model for sleep window detection and the SleepNet for sleep stage identification.
We simulated the effects of random missing data on the participants that had no missing data across 7-days to determine the minimum wear time required for stable weekly sleep parameter estimates (Supplementary Section 1.3.2). We found that a minimum of 22 h of wear time per day for at least 3 days were required to ensure the intra-class correlation was greater than 0.75 between the weekly average sleep duration from incomplete and perfect wear data. Moreover, we tried to mitigate the weekend effect by only including the participants who had at least one weekday and one weekend day during the device wear. Shift workers and participants whose data had daylight saving cross-overs were also excluded, as circadian disruption is not the focus of our paper.
Descriptive analyses were performed on the device-measured sleep parameters in the UK Biobank to quantify variations by age, sex, device-measured physical activity level, self-reported chronotype and insomnia symptoms. Estimated marginal means, adjusted for age and sex, were also calculated for different self-rated health groups and self-reported insomnia symptoms.
This research has been conducted using the UK Biobank Resource under Application Number 59070. The UK Biobank received ethical approval from the National Health Service National Research Service (Ref 21/NW/0157). Written informed consent was obtained from all the participants.
Health association analysis
The associations of overnight sleep duration and sleep efficiency with incident mortality were assessed using Cox proportional hazards regression. All-cause mortality was determined using death registry data (obtained by UK Biobank from NHS Digital for participants in England and Wales and from the NHS Central Register, National Records of Scotland, for participants in Scotland). Participants were censored at the earliest of UK Biobank’s record censoring date for mortality data (2021-09-30 for participants in England and Wales and 2021-10-31 for participants in Scotland, with country assigned based on baseline assessment centre). Cox models used age as the timescale, and the main analysis was adjusted for sex, ethnicity, Townsend Deprivation Index, educational qualifications, smoking status, alcohol consumption and overall activity. See Supplementary Section 1.3.1 for the full specification of the analysis.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data for the Newcastle cohort is available from direct download via https://zenodo.org/record/1160410#.Y-O65i-l1qs. The data for other cohorts can be requested by contacting the corresponding host institute.
Code availability
All the sleep staging models and analysis scripts are freely available for academic use on GitHub: https://github.com/OxWearables/asleep.
Change history
01 July 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41746-024-01148-y
References
Meyer, N., Harvey, A. G., Lockley, S. W. & Dijk, D.-J. Circadian rhythms and disorders of the timing of sleep. Lancet 400, 1061–7078 (2022).
Ferrie, J. E., Kumari, M., Salo, P., Singh-Manoux, A. & Kivimäki, M. Sleep epidemiology—a rapidly growing field. Int. J. Epidemiol. 40, 1431–1437 (2011).
Short, M. A., Gradisar, M., Lack, L. C., Wright, H. & Carskadon, M. A. The discrepancy between actigraphic and sleep diary measures of sleep in adolescents. Sleep. Med. 13, 378–384 (2012).
Wainberg, M. et al. Association of accelerometer-derived sleep measures with lifetime psychiatric diagnoses: a cross-sectional study of 89,205 participants from the UK Biobank. PLoS Med. 18, e1003782 (2021).
Mantua, J., Gravel, N. & Spencer, R. Reliability of sleep measures from four personal health monitoring devices compared to research-based actigraphy and polysomnography. Sensors 16, 646 (2016).
Boe, A. J. et al. Automating sleep stage classification using wireless, wearable sensors. NPJ Digit. Med. 2, 1–9 (2019).
Devine, J. K., Chinoy, E. D., Markwald, R. R., Schwartz, L. P. & Hursh, S. R. Validation of Zulu watch against polysomnography and actigraphy for on-wrist sleep-wake determination and sleep-depth estimation. Sensors 21, 76 (2020).
Patterson, M. R. et al. 40 years of actigraphy in sleep medicine and current state of the art algorithms. NPJ Digit. Med. 6, 51 (2023).
Doherty, A. et al. GWAS identifies 14 loci for device-measured physical activity and sleep duration. Nat. Commun. 9, 1–8 (2018).
Jones, S. E. et al. Genetic studies of accelerometer-based sleep measures yield new insights into human sleep behaviour. Nat. Commun. 10, 1–12 (2019).
Katori, M., Shi, S., Ode, K. L., Tomita, Y. & Ueda, H. R. The 103,200-arm acceleration dataset in the UK biobank revealed a landscape of human sleep phenotypes. Proc. Natl Acad. Sci. 119, e2116729119 (2022).
McHugh, M. L. Interrater reliability: the kappa statistic. Biochem. Med. 22, 276–282 (2012).
Sundararajan, K. et al. Sleep classification from wrist-worn accelerometer data using random forests. Sci. Rep. 11, 1–10 (2021).
Trevenen, M. L., Turlach, B. A., Eastwood, P. R., Straker, L. M. & Murray, K. Using hidden Markov models with raw, triaxial wrist accelerometry data to determine sleep stages. Aust. N. Z. J. Stat. 61, 273–298 (2019).
Yin, J. et al. Relationship of sleep duration with all-cause mortality and cardiovascular events: a systematic review and dose-response meta-analysis of prospective cohort studies. J. Am. Heart Assoc. 6, e005947 (2017).
Itani, O., Jike, M., Watanabe, N. & Kaneita, Y. Short sleep duration and health outcomes: a systematic review, meta-analysis, and meta-regression. Sleep Med. 32, 246–256 (2017).
Liang, Y. Y. et al. Joint associations of device-measured sleep duration and efficiency with all-cause and cause-specific mortality: a prospective cohort study of 90 398 UK biobank participants. J. Gerontol. Ser. A 78, 1717–1724 (2023).
Wang, C. et al. Association of estimated sleep duration and naps with mortality and cardiovascular events: a study of 116 632 people from 21 countries. Eur. Heart J. 40, 1620–1629 (2019).
Taheri, S. Sleep and cardiometabolic health—not so strange bedfellows. Lancet Diabetes Endocrinol. 11, 532–534 (2023).
Golbus, J. R., Pescatore, N. A., Nallamothu, B. K., Shah, N. & Kheterpal, S. Wearable device signals and home blood pressure data across age, sex, race, ethnicity, and clinical phenotypes in the Michigan predictive activity & clinical trajectories in health (MIPACT) study: a prospective, community-based observational study. Lancet Digit. Health 3, e707–e715 (2021).
Agnew Jr, H., Webb, W. B. & Williams, R. L. The first night effect: An eeg studyof sleep. Psychophysiology 2, 263–266 (1966).
Doherty, A. et al. Large scale population assessment of physical activity using wrist worn accelerometers: the UK biobank study. PloS One 12, e0169649 (2017).
Straker, L. et al. Cohort profile: the western Australian pregnancy cohort (raine) study–generation 2. Int. J. Epidemiol. 46, 1384–1385j (2017).
Dontje, M. L., Eastwood, P. & Straker, L. Western Australian pregnancy cohort (raine) study: generation 1. BMJ open 9, e026276 (2019).
van Hees, V., Charman, S. & Anderson, K. Newcastle polysomnography and accelerometer data. https://doi.org/10.5281/zenodo.1160410 (2018).
Plekhanova, T. et al. Validation of an automated sleep detection algorithm using data from multiple accelerometer brands. J. Sleep Res. e13760 (2022).
Byrne, E. M., Gehrman, P. R., Trzaskowski, M., Tiemeier, H. & Pack, A. I. Genetic correlation analysis suggests association between increased self-reported sleep duration in adults and schizophrenia and type 2 diabetes. Sleep 39, 1853–1857 (2016).
Migueles, J. H. et al. Equivalency of four research-grade movement sensors to assess movement behaviors and its implications for population surveillance. Sci. Rep. 12, 1–9 (2022).
Willetts, M., Hollowell, S., Aslett, L., Holmes, C. & Doherty, A. Statistical machine learning of sleep and physical activity phenotypes from sensor data in 96,220 uk biobank participants. Sci. Rep. 8, 1–10 (2018).
Walmsley, R. et al. Reallocation of time between device-measured movement behaviours and risk of incident cardiovascular disease. Br. J. Sports Med. 56, 1008–1017 (2022).
Berry, R. B. et al. Rules for scoring respiratory events in sleep: update of the 2007 AASM manual for the scoring of sleep and associated events: deliberations of the sleep apnea definitions task force of the American Academy of Sleep Medicine. J. Clin. Sleep. Med. 8, 597–619 (2012).
He, K., Zhang, X., Ren, S. & Sun, J. Identity mappings in deep residual networks. In European Conference on Computer Vision, 630–645 (Springer, 2016).
Huang, Z., Xu, W. & Yu, K. Bidirectional lstm-crf models for sequence tagging. arXiv preprint arXiv:1508.01991 (2015).
Yuan, H. et al. Self-supervised learning for human activity recognition using 700,000 person-days of wearable data. npj Digit. Med. 7, 91 (2024).
Creagh, A. P. et al. Digital health technologies and machine learning augment patient reported outcomes to remotely characterise rheumatoid arthritis. NPJ Digit. Med. 7, 33 (2024).
van Hees, V. T. et al. Estimating sleep parameters using an accelerometer without sleep diary. Sci. Rep. 8, 12975 (2018).
Acknowledgements
We would like to acknowledge the Raine Study participants and their families for their ongoing participation in the study and the Raine Study team for study coordination and data collection. We also thank the NHMRC for their long-term contribution to funding the study over the last 30 years. The core management of the Raine Study is funded by The University of Western Australia, Curtin University, Telethon Kids Institute, Women and Infants Research Foundation, Edith Cowan University, Murdoch University, The University of Notre Dame Australia and the Raine Medical Research Foundation. The 22-year Gen2 Raine Study follow-up was funded by NHMRC project grants 1027449 & 1044840. The data collection for the Pennsylvania dataset is funded, in part, by US National Institute of Health (NIMH) grant R21 MH103963 (M.B.). H.Y., D.B. and A.D. are supported by Novo Nordisk. R.W. and A.D. are supported by Health Data Research UK, an initiative funded by UK Research and Innovation, Department of Health and Social Care (England) and the devolved administrations and leading medical research charities. A.D. is additionally supported by Swiss Re, Wellcome Trust [223100/Z/21/Z] and the British Heart Foundation Centre of Research Excellence (grant number RE/18/3/34214). D.W.R. is supported by MRC programme grant MR/P023576/1; Wellcome Trust (107849/Z/15/Z). T.P. and A.R. are supported by the National Institute for Health Research (NIHR) Leicester Biomedical Research Centre and NIHR Applied Research Collaboration East Midlands (ARC EM). S.D.K. is supported by the National Institute of Health and Care Research (NIHR) Oxford Health Biomedical Research Centre (NIHR203316), NIHR Efficacy and Mechanisms Evaluation Programme (Ref: 131789), NIHR Programme Grants for Applied Research (Ref: 203667) and the Wellcome Trust (226784/Z/22/Z and 227093/Z/23/Z). Computational aspects of this research were funded from the National Institute for Health Research (NIHR) Oxford Biomedical Research Centre (BRC) with additional support from Health Data Research (HDR) UK and the Wellcome Trust Core Award [grant number 203141/Z/16/Z]. The views expressed are those of the authors and not necessarily those of the NHS, the NIHR or the Department of Health. For the purpose of open access, the author has applied a CC-BY public copyright licence to any author accepted manuscript version arising from this submission. We would like to thank Andrew Creagh, Angel Wong, Scott Small and Alaina Shreves for their input on the revision of this manuscript. We would also like to thank Andrew Creagh for his feedback in creating the graphic illustrations.
Author information
Authors and Affiliations
Contributions
H.Y., K.J.M., J.M., L.S., P.E., S.D.K. and A.D. conceptualised and designed the study. T.P., M.B., P.G., A.R., J.M., L.S. and P.S. did the data curation of the accelerometers and polysomnography data. H.Y., T.P. and R.W. did the formal analysis and validation. D.B., S.D.K. and A.D. provided supervision to H.Y. and R.W. H.Y. wrote the manuscript, and all the authors contributed to the review & editing process. H.Y. and R.W. had direct access to the summary statistics and verified the data.
Corresponding author
Ethics declarations
Competing interests
P.G. reports receiving funding from NIH/NIMH. A.D. receives funding from Wellcome Trust, Novo Nordisk, Swiss Re, the British Heart Foundation Centre of Research Excellence and Health Data Research UK, an initiative funded by UK Research and Innovation, Department of Health and Social Care (England) and the devolved administrations and leading medical research charities. A.C.R. is supported by Flinders Foundation, Hospital Research Foundation, Compumedics, Vanda Pharmaceuticals, Teva Pharmaceuticals and Sleep Health Foundation. D.R. receives funding from Wellcome Trust and Medical Research Council. D.R. also received lecture fees from Pfizer for an education programme. K.J.M. is supported by Sir Charles Gairdner Hospital Research Advisory Council Funding, Early to Mid-Career Researchers Grant Medical Research Future Fund, Chevron Australia, CHC Helicopter Australia, Incannex Healthcare Limited, Nyxoah Pty Ltd. K.J.M. receives consulting fees from Melius Consulting, HIF, Invicta Medical and lecture fees from Sleep Health Foundation, WA Dental Association and Shire of Cannington. Maja receives funding from the National Institute of Health. The remaining authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yuan, H., Plekhanova, T., Walmsley, R. et al. Self-supervised learning of accelerometer data provides new insights for sleep and its association with mortality. npj Digit. Med. 7, 86 (2024). https://doi.org/10.1038/s41746-024-01065-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41746-024-01065-0