Abstract
Recent studies have reported numerous predictors for adverse outcomes in COVID-19 disease. However, there have been few simple clinical risk scores available for prompt risk stratification. The objective is to develop a simple risk score for predicting severe COVID-19 disease using territory-wide data based on simple clinical and laboratory variables. Consecutive patients admitted to Hong Kong’s public hospitals between 1 January and 22 August 2020 and diagnosed with COVID-19, as confirmed by RT-PCR, were included. The primary outcome was composite intensive care unit admission, need for intubation or death with follow-up until 8 September 2020. An external independent cohort from Wuhan was used for model validation. COVID-19 testing was performed in 237,493 patients and 4442 patients (median age 44.8 years old, 95% confidence interval (CI): [28.9, 60.8]); 50% males) were tested positive. Of these, 209 patients (4.8%) met the primary outcome. A risk score including the following components was derived from Cox regression: gender, age, diabetes mellitus, hypertension, atrial fibrillation, heart failure, ischemic heart disease, peripheral vascular disease, stroke, dementia, liver diseases, gastrointestinal bleeding, cancer, increases in neutrophil count, potassium, urea, creatinine, aspartate transaminase, alanine transaminase, bilirubin, D-dimer, high sensitive troponin-I, lactate dehydrogenase, activated partial thromboplastin time, prothrombin time, and C-reactive protein, as well as decreases in lymphocyte count, platelet, hematocrit, albumin, sodium, low-density lipoprotein, high-density lipoprotein, cholesterol, glucose, and base excess. The model based on test results taken on the day of admission demonstrated an excellent predictive value. Incorporation of test results on successive time points did not further improve risk prediction. The derived score system was evaluated with out-of-sample five-cross-validation (AUC: 0.86, 95% CI: 0.82–0.91) and external validation (N = 202, AUC: 0.89, 95% CI: 0.85–0.93). A simple clinical score accurately predicted severe COVID-19 disease, even without including symptoms, blood pressure or oxygen status on presentation, or chest radiograph results.
Similar content being viewed by others
Introduction
The coronavirus disease 2019 has a wide clinical spectrum, with disease severities ranging from completely asymptomatic to the need for intubation and death1,2,3,4. For example, those with existing cardiac problems are more likely to suffer from more severe disease life courses5,6,7,8,9,10,11, with potential modifier effects from different medication classes12,13,14. Aside from comorbidities, numerous risk factors such as high D-dimer15, neutrophil16, and liver damage17 and deranged clotting18 have been associated with disease severity. Such patients may benefit from early aggressive treatment19,20,21,22,23. However, to date, there are only a few easy-for-use risk models that can be used for early identification of such at-risk individuals in clinical practice24,25. The aim of the study is to extend these previous findings and develop a predictive risk score based on demographic, comorbidity, medication record, and laboratory data using territory-wide electronic health records, without clinical parameters or imaging results. We hypothesized that incorporation of test results on successive time points would improve risk prediction. The model was validated internally, and externally using a single-center cohort from Wuhan.
Results
Basic characteristics
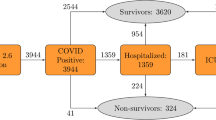
A total of 4442 patients (median age 44.8 years old, 95% CI: [28.9, 60.8]); 50% males) were diagnosed with the COVID-19 infection between 1 January 2020 and 22 August 2020 in Hong Kong public hospitals or their associated ambulatory/outpatient facilities (Table 1). On follow-up until 8 September 2020, a total of 212 patients (4.77%) met the primary outcome of need for intensive care admission or intubation, or death. The survival curve is presented in Fig. 1. The sudden inflexion point at 200 days likely reflects the surge of new cases around this period. The baseline and clinical characteristics of male and female COVID-19 patients are provided in Supplementary Table 3.
Development of a clinical risk score and validation
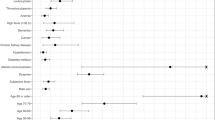
Univariate logistic regression analyses are shown in Table 2, which identified the significant risk predictors for the composite outcome. However, for clinical practice, it is impractical to precisely input the values of all variables assessed from the different domains of the health records. Three different models were developed (Tables 3–5), as detailed in the “Methods” section. The easy-to-use score system is shown in Table 6.
Patients meeting the primary outcome (n = 212) have significantly higher risk score (median: 5.13, 95% CI: 3.13–7.43, max: 18.6) than those who did not (median: 1.41, 95% CI: 0.65–5.94, max: 18.2) (Table 7), indicating the significant risk stratification performance of the clinical risk score (OR: 17.1, 95% CI: 11–26.6) (Table 8). Survival curves stratified by the dichotomized risk score are shown in Fig. 2, where yellow and blue curves represent the survival analysis for patients with a clinical risk score is larger and smaller than the cut-off, respectively.
For external validation, a total of 202 patients (48% males) from the Wuhan Heart Hospital were included. Comparisons of different performance measures for the clinical risk score for the Hong Kong cohort (fivefold cross-validation) and Wuhan cohort are detailed in Table 9. Receiver operating characteristic curve (ROC) of predicting adverse composite outcome of COVID-19 patients with the dichotomized risk score cut-off is shown in Fig. 3 (Hong Kong cohort: top panel; Wuhan cohort: bottom panel). As the Wuhan cohort did not routinely have AST tested, this variable was excluded for the performance comparisons. The AUC of 0.86 for the Hong Kong cohort (fivefold cross-validation) and 0.89 for the Wuhan cohort.
Discussion
In this study, we developed a simple clinical score to predict severe COVID-19 disease based on age, gender, medical comorbidities, medication records, and laboratory examination results. We compared the prediction strengths of different criteria for the clinical risk score for out-of-sample validation for the Hong Kong cohort (fivefold cross-validation) and external validation for the Wuhan cohort, with AUC as 0.86 (95% CI: 0.82–0.91) and 0.89 [0.85–0.93], respectively. The derived score system achieved good predictions even without the consideration of clinical parameters such as symptoms, blood pressure, oxygen status on presentation, or chest radiograph results.
COVID-19 disease has placed significant pressures on healthcare systems worldwide. Early risk stratification may better direct the use of limited resources and allow clinicians to triage patients and make clinical decisions based on limited evidence objectively. For example, low-risk patients may require simple monitoring only, while patients that are likely to deteriorate may benefit from intensive drug treatment or intensive care. Currently, the availability of simple clinical risk scores for risk stratification is limited. The COVID-GRAM predicts development of critical illness, based on symptoms, radiograph results, clinical and laboratory details24. Similarly, the 4C Mortality Score included eight variables readily available at initial hospital assessment: age, sex, number of comorbidities, respiratory rate, peripheral oxygen saturation, level of consciousness, urea level, and C-reactive protein (score range 0–21 points)25. These scores produced moderately accurate predictions with C-index values of 0.86 and 0.61–0.76, respectively. A systematic review and meta-analysis have recently summarized different risk scores that have been developed by investigators from different countries26. As reported, the most frequently reported predictors were age, clinical status such as temperature, imaging results from chest radiography, and lymphocyte count. Recently, a study including 3927 patients from 33 hospitals developed the COVID-19 Mortality Risk (CMR) tool using the XGBoost algorithm27. This score is based on age, blood urea nitrogen, CRP, creatinine, glucose, AST, and platelet counts. Different teams in our country have already used a data-driven approach to develop predictive risk models for COVID-19 to predict viral transmission28,29, adverse outcomes30,31 and even to determine effects of risk perceptions on behaviors in response to the outbreak32. For example, our team recently developed a risk model based on non-linear interactions between different variables to predict intensive care unit admission using a tree-based machine learning model30. The above models are based on individual-level patient data. Where these are not available, investigators have successfully developed a useful model by using aggregate epidemiological reports of COVID-19 case fatality events33.
In this study, with an expanded cohort, we developed a simple and easy-to-use model was based on past comorbidity and laboratory data only, without needing clinical assessment details or chest imaging interpretation. The model based on test results taken on the day of admission already demonstrated an excellent predictive value with a C-statistic of 0.89. Incorporation of test results on successive time points did not further improve risk prediction, indicating that initial data are sufficient to produce accurate predictions of severe disease. Our model can aid clinical decision making as early intervention may be associated with better outcomes19,20,21,22,23.
The major limitation of this study is that it is based on a territory-wide cohort from a single city in China (Hong Kong). However, the risk score was independently validated using an external cohort from another city (Wuhan). We recognize that the baseline demographic and clinical characteristics of COVID-19 patients may differ in other countries. The model should be further externally validated using patient data involving from other geographical regions to allow further generalization.
In conclusion, simple clinical score based on only demographics, comorbidities, medication records, and laboratory tests accurately predicted severe COVID-19 disease, even without including symptoms on presentation, blood pressure, oxygen status, or chest radiograph results. The model based on test results taken on the day of admission showed an excellent predictive value. Incorporation of test results on successive time points did not further improve risk prediction. Both out-of-sample fivefold cross-validation on Hong Kong cohort and independent external validation on Wuhan cohort demonstrated the significant risk stratification performance of the derived score system for severe COVID-19 disease. The presented score system tool used commonly available clinical and laboratory results and does not require imaging results or advanced testing, and therefore can be particularly useful in facilities with constrained resources or remote hospitals with limited diagnostic capabilities such as computed tomography scans.
Methods
Study design and population
This study was approved by the Institutional Review Board of the University of Hong Kong/Hospital Authority Hong Kong West Cluster. The need for informed consent was waived by the Ethics Committee owing to the retrospective and observational nature of this study. This was a retrospective, territory-wide cohort study of patients undergoing COVID-19 RT-PCR testing between 1 January 2020 and 22 August 2020 in Hong Kong, China. The patients were identified from the Clinical Data Analysis and Reporting System (CDARS), a territory-wide database that centralizes patient information from 43 local hospitals and their associated ambulatory and outpatient facilities to establish comprehensive medical data, including clinical characteristics, disease diagnosis, laboratory results, and drug treatment details. The system has been previously used by both our team and other teams in Hong Kong34,35, including recently COVID-19 research36,37. This system captures PCR tests performed in Accident and Emergency, outpatient and inpatient settings. Patients demographics, prior comorbidities, hospitalization characteristics before admission due to COVID-19, medication prescriptions, laboratory examinations of complete blood counts, biochemical tests, diabetes mellitus tests, cardiac function tests, c-reactive protein, and blood gas tests were extracted. The list of ICD-9 codes for comorbidities and intubation procedures are detailed in the Supplementary Tables 1 and 2.
Outcomes and statistical analysis
The primary outcome was a composite of need for intensive care admission, intubation, or all-cause mortality with follow-up until 8 September 2020. Mortality data were obtained from the Hong Kong Death Registry, a population-based official government registry with the registered death records of all Hong Kong citizens linked to CDARS. Patients who passed away 30 days later or longer after discharge were excluded. The need for ICU admission and intubation were extracted directly from CDARS. There was no adjudication of the outcomes as this relied on the ICD-9 coding or a record in the death registry. However, the coding was performed by the clinicians or administrative staff, who were not involved in the mode development. Descriptive statistics are used to summarize baseline clinical characteristics of all patients with COVID-19 and based on the occurrence of the primary outcome. Continuous variables were presented as median (95% confidence interval [CI] or interquartile range [IQR]) and categorical variables were presented as count (%). The Mann–Whitney U test was used to compare continuous variables. The χ2 test with Yates’ correction was used for 2 × 2 contingency data. Univariate logistic regression identifies significant mortality risk predictors. Odds ratios (ORs) with corresponding 95% CIs and P values were reported. There was no imputation performed for missing data. An easy-for-use predictive model was developed using the beta coefficients for different predictors identified from logistic regression. Successive laboratory tests at least 24 h apart were used. No blinding was performed for the predictor as the values were obtained from the electronic health records automatically.
Development of different scoring systems
Three different models were developed.
Model 1: optimum cut-off values of different variables at baseline were obtained from receiver operating characteristic (ROC) analysis. Laboratory examinations on for each successive 24 h was compared to cut-off to determine whether the criterion was met at each time point.
Model 2: the criterion was met if the value was abnormal by standard laboratory criteria, without consideration of optimal cut-off values.
Model 3: laboratory test results are compared to the criteria without cut-off values, to determine if they were met on successive testing. For example, if a particular criterion is met on day 1, then they will automatically fulfill the criteria for subsequent days.
A simple and easy-to-use score system was built based on beta coefficients using logistic regression analysis. The risk score of each COVID-19 patient was then calculated. The derived score system was evaluated within-sample fivefold cross testing set and out-of-sample dataset from Wuhan for external validation. The model was not recalibrated after validation.
External validation
For external validation, patients admitted to the Wuhan Asia General Hospital38, Wuhan, China, between 10 February and 10 March 2020, were included. Diagnosis of COVID-19 was based on positive PCR test and ground glass shadows in the lungs on computed tomography scan, with follow-up 2 weeks post-discharge. Lipid and aspartate aminotransferase were not routinely collected and therefore not included for validation.
Performance of the score
Performance of the score system was evaluated based on its ability to discriminate the composite outcome for each population. The results for in-sample testing set, and for external out-of-sample validation cohort were reported, with the corresponding CIs. The area under the curve (AUC), accuracy, specificity, and precision were computed for all patient subpopulations. Receiver operating characteristic (ROC) curves were created for each of the cohorts with the derive score system to predict the adverse composite outcome. All statistical tests were two-tailed and considered significant if p value < 0.001. They were performed using RStudio software (Version: 1.1.456) and Python (Version: 3.6).
Reporting summary
Further information on experimental design is available in the Nature Research Reporting Summary linked to this paper.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Change history
28 March 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41746-022-00586-w
References
Mody, A. et al. The clinical course of COVID-19 disease in a US hospital system: a multi-state analysis. Am. J. Epidemiol. https://doi.org/10.1093/aje/kwaa286 (2020).
Thomson, R. J. et al. Clinical characteristics and outcomes of critically ill patients with COVID-19 admitted to an intensive care unit in London: a prospective observational cohort study. PLoS ONE 15, e0243710 (2020).
Grasselli, G., Cattaneo, E. & Scaravilli, V. Ventilation of coronavirus disease 2019 patients. Curr. Opin. Crit. Care 27, 6–12 (2021).
Coromilas, E. J. et al. Worldwide survey of COVID-19 associated arrhythmias. Circ. Arrhythm. Electrophysiol. https://doi.org/10.1161/CIRCEP.120.009458 (2021).
Guo, T. et al. Cardiovascular implications of fatal outcomes of patients with coronavirus disease 2019 (COVID-19). JAMA Cardiol. https://doi.org/10.1001/jamacardio.2020.1017 (2020).
Shi, S. et al. Association of cardiac injury with mortality in hospitalized patients with COVID-19 in Wuhan, China. JAMA Cardiol. https://doi.org/10.1001/jamacardio.2020.0950 (2020).
Wang, Y., Roever, L., Tse, G. & Liu, T. 2019-novel coronavirus-related acute cardiac injury cannot be ignored. Curr. Atheroscler. Rep. 22, 14 (2020).
Li, X. et al. Impact of cardiovascular disease and cardiac injury on in-hospital mortality in patients with COVID-19: a systematic review and meta-analysis. Heart https://doi.org/10.1136/heartjnl-2020-317062 (2020).
Tan, E., Song, J., Deane, A. M. & Plummer, M. P. Global impact of coronavirus disease 2019 infection requiring admission to the ICU: a systematic review and meta-analysis. Chest https://doi.org/10.1016/j.chest.2020.10.014 (2020).
Wang, Y. et al. Cardiac arrhythmias in patients with COVID-19. J. Arrhythm. 36, 827–836 (2020).
Hui, Y. et al. The risk factors for mortality of diabetic patients with severe COVID-19: a retrospective study of 167 severe COVID-19 cases in Wuhan. PLoS ONE 15, e0243602 (2020).
Baral, R., White, M. & Vassiliou, V. S. Effect of renin-angiotensin-aldosterone system inhibitors in patients with COVID-19: a systematic review and meta-analysis of 28,872 patients. Curr. Atheroscler. Rep. 22, 61 (2020).
Zhou, J. et al. Proton pump inhibitor or famotidine use and severe COVID-19 disease: a propensity score-matched territory-wide study. Gut. https://doi.org/10.1136/gutjnl-2020-323668 (2020).
Wang, Y., Tse, G., Li, G., Lip, G. Y. H. & Liu, T. ACE inhibitors and angiotensin II receptor blockers may have different impact on prognosis of COVID-19. J. Am. Coll. Cardiol. 76, 2041 (2020).
Yao, Y. et al. D-dimer as a biomarker for disease severity and mortality in COVID-19 patients: a case control study. J. Intensive Care 8, 49 (2020).
Huang, C. et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 395, 497–506 (2020).
Cai, Q. et al. COVID-19: abnormal liver function tests. J. Hepatol. 73, 566–574 (2020).
Tang, N., Li, D., Wang, X. & Sun, Z. Abnormal coagulation parameters are associated with poor prognosis in patients with novel coronavirus pneumonia. J. Thromb. Haemost. 18, 844–847 (2020).
Forrest, J. I., Rayner, C. R., Park, J. J. H. & Mills, E. J. Early treatment of COVID-19 disease: a missed opportunity. Infect. Dis. Ther. 9, 715–720 (2020).
Zhang, Q., Wei, Y., Chen, M., Wan, Q. & Chen, X. Clinical analysis of risk factors for severe COVID-19 patients with type 2 diabetes. J. Diabetes Complications 34, 107666 (2020).
Bose, S. et al. Medical management of COVID-19: evidence and experience. J. Clin. Med. Res. 12, 329–343 (2020).
Million, M. et al. Early treatment of COVID-19 patients with hydroxychloroquine and azithromycin: a retrospective analysis of 1061 cases in Marseille, France. Travel Med. Infect. Dis. 35, 101738 (2020).
de Simone, G. & Mancusi, C. COVID-19: timing is important. Eur. J. Intern. Med. 77, 134–135 (2020).
Liang, W. et al. Development and validation of a clinical risk score to predict the occurrence of critical illness in hospitalized patients with COVID-19. JAMA Intern. Med. 180, 1081–1089 (2020).
Knight, S. R. et al. Risk stratification of patients admitted to hospital with covid-19 using the ISARIC WHO Clinical Characterisation Protocol: development and validation of the 4C Mortality Score. BMJ 370, m3339 (2020).
Wynants, L. et al. Prediction models for diagnosis and prognosis of covid-19 infection: systematic review and critical appraisal. BMJ 369, m1328 (2020).
Bertsimas, D. et al. COVID-19 mortality risk assessment: an international multi-center study. PLoS ONE 15, e0243262 (2020).
Jia, J. S. et al. Population flow drives spatio-temporal distribution of COVID-19 in China. Nature 582, 389–394 (2020).
Wan, H., Cui, J.-A. & Yang, G.-J. Risk estimation and prediction of the transmission of coronavirus disease-2019 (COVID-19) in the mainland of China excluding Hubei province. Infect. Dis. Poverty 9, 116 (2020).
Zhou, J. et al. Identifying main and interaction effects of risk factors to predict intensive care admission in patients hospitalized with COVID-19: a retrospective cohort study in Hong Kong. Preprint at medRxiv. https://doi.org/10.1101/2020.06.30.20143651 (2020).
Cao, G. et al. A risk prediction model for evaluating the disease progression of covid-19 pneumonia. Front. Med. (Lausanne) 7, 556886 (2020).
Ye, Y. et al. Effect of heterogeneous risk perception on information diffusion, behavior change, and disease transmission. Phys. Rev. E 102, 042314 (2020).
Barda, N. et al. Developing a COVID-19 mortality risk prediction model when individual-level data are not available. Nat. Commun. 11, 4439 (2020).
Li, C. K. et al. Association of NPAC score with survival after acute myocardial infarction. Atherosclerosis 301, 30–36 (2020).
Ju, C. et al. Comparative cardiovascular risk in users versus non-users of xanthine oxidase inhibitors and febuxostat versus allopurinol users. Rheumatology 59, 2340–2349 (2020).
Zhou, J. et al. Anticoagulant or antiplatelet use and severe COVID-19 disease: a propensity score-matched territory-wide study. Pharmacol. Res. https://doi.org/10.1016/j.phrs.2021.105473 (2021).
Zhou, J. et al. Interaction effects between angiotensin-converting enzyme inhibitors or angiotensin receptor blockers and steroid or anti-viral therapies in COVID-19: a population-based study. J. Med. Virol. https://doi.org/10.1002/jmv.26904 (2021). Epub ahead of print.
Li, Y. et al. Electrocardiograhic characteristics in patients with coronavirus infection: a single-center observational study. Ann. Noninvasive Electrocardiol. 25, e12805 (2020).
Acknowledgements
Q.Z. acknowledges the following funding: National Natural Science Foundation of China (NSFC): 71972164 and 72042018; Health and Medical Research Fund of the Food and Health Bureau of Hong Kong: 16171991; Innovation and Technology Fund of Innovation and Technology Commission of Hong Kong: MHP/081/19; National Key Research and Development Program of China, Ministry of Science and Technology of China: 2019YFE0198600; I.C.K.W. acknowledges the following funding: Collaborative Research Fund (CRF) of Research Grants Council of Hong Kong: C7154-20G.
Author information
Authors and Affiliations
Contributions
J.Z. and S.L.: data analysis, data interpretation, statistical analysis, manuscript drafting, and critical revision of manuscript. K.S.K.L. and A.K.C.W.: data acquisition and interpretation, critical revision of manuscript. X.W., Y.L., W.K.K.W., T.L., Z.C., D.D.Z., I.C.K.W., B.M.Y.C.: project planning, data acquisition, data interpretation, critical revision of manuscript. Q.Z. and G.T.: study conception, study supervision, project planning, data interpretation, statistical analysis, manuscript drafting, critical revision of manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhou, J., Lee, S., Wang, X. et al. Development of a multivariable prediction model for severe COVID-19 disease: a population-based study from Hong Kong. npj Digit. Med. 4, 66 (2021). https://doi.org/10.1038/s41746-021-00433-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41746-021-00433-4
This article is cited by
-
Machine learning-based prediction of COVID-19 mortality using immunological and metabolic biomarkers
BMC Digital Health (2023)
-
Prognostic models in COVID-19 infection that predict severity: a systematic review
European Journal of Epidemiology (2023)
-
Comparisons of the risk of myopericarditis between COVID-19 patients and individuals receiving COVID-19 vaccines: a population-based study
Clinical Research in Cardiology (2022)