Abstract
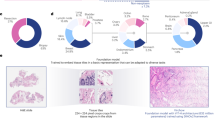
The emergence of digital pathology has opened new horizons for histopathology. Artificial intelligence (AI) algorithms are able to operate on digitized slides to assist pathologists with different tasks. Whereas AI-involving classification and segmentation methods have obvious benefits for image analysis, image search represents a fundamental shift in computational pathology. Matching the pathology of new patients with already diagnosed and curated cases offers pathologists a new approach to improve diagnostic accuracy through visual inspection of similar cases and computational majority vote for consensus building. In this study, we report the results from searching the largest public repository (The Cancer Genome Atlas, TCGA) of whole-slide images from almost 11,000 patients. We successfully indexed and searched almost 30,000 high-resolution digitized slides constituting 16 terabytes of data comprised of 20 million 1000 × 1000 pixels image patches. The TCGA image database covers 25 anatomic sites and contains 32 cancer subtypes. High-performance storage and GPU power were employed for experimentation. The results were assessed with conservative “majority voting” to build consensus for subtype diagnosis through vertical search and demonstrated high accuracy values for both frozen section slides (e.g., bladder urothelial carcinoma 93%, kidney renal clear cell carcinoma 97%, and ovarian serous cystadenocarcinoma 99%) and permanent histopathology slides (e.g., prostate adenocarcinoma 98%, skin cutaneous melanoma 99%, and thymoma 100%). The key finding of this validation study was that computational consensus appears to be possible for rendering diagnoses if a sufficiently large number of searchable cases are available for each cancer subtype.
Similar content being viewed by others
Introduction
Digital pathology is the virtual version of conventional microscopy utilized for the examination of glass pathology slides. In recent years, there has been accelerated adoption of digital pathology, whereby pathology laboratories around the world are slowly beginning to trade in their light microscopes for digital scanners, computers, and monitors. As a result, the pathology community has begun to scan many slides resulting in the creation of large databases of whole-slide images (WSIs). The emergence of deep learning and other artificial intelligence (AI) methods and their impressive pattern-recognition capabilities when applied to these digital databases has immensely added to the value proposition of digital pathology1,2,3. Computerized operations, such as segmentation of tissue fragments and cell nuclei, and classification of diseases and their grades become possible after pathology slides are digitized. These operations could assist with many diagnostic and research tasks with expert-like accuracy when trained with the proper level of labeled data4. The majority of recent studies in digital pathology have reported the success of supervised AI algorithms for classification and segmentation4,5,6,7. This overrepresentation compared with other AI algorithms is related to the ease of design and in-lab validation to generate highly accurate results. However, compared with other methods of computer-vision algorithms, AI-based image search and retrieval offers a new approach to computational pathology.
Content-based image search8,9,10,11 implies that the input for search software is not text (e.g., disease description in a pathology report), but rather the input is an image such that the search and retrieval can be performed based on image pixels (visual content). Content-based image search is inherently unsupervised, which means that its design and implementation may not need manual delineation of a region of interest in the images12,13,14. More importantly, image search does not make any direct diagnostic decision on behalf of the pathologist; instead, it searches for similar images and retrieves them along with the corresponding metadata (i.e., pathology reports), and displays them to the pathologist as decision support.
Variability in the visual inspection of medical images is a well-known problem15,16,17. Both inter- and intra-observer variability may affect image assessment and subsequently the ensuing diagnosis18,19,20,21. A large body of work have reported high rates of diagnostic inaccuracy as a result of major discordance among participating physicians with respect to case target diagnoses, and propose a combination of “routine second opinions” and “directed retrospective peer review”22,23,24. As most proposed AI-driven solutions for digital pathology mainly focus on the concept of classification, it appears that algorithmic decision-making may not necessarily contribute to supporting concordance by providing a framework for consensus building. Most capable classification schemes trained with immense effort are supposed to be used for triaging cases in the pathology laboratory, and not for direct assistance in the pathologist’s office4. In contrast, instantly retrieving multiple diagnosed cases with histopathologic similarity to the patient’s biopsy about to be diagnosed offers a new generation of decision support that may even enable “virtual” peer review.
Content-based image retrieval (CBIR) systems have been under investigation for more than two decades25,26,27. Recently, deep learning has gained a lot of attention for image search28,29,30. While CBIR systems of medical images have been well researched11,31,32,33, only with the emergence of digital pathology34,35 and deep learning3,36,37 has research begun to focus on image search and analysis in histopathology2,38,39,40. In the past 3 years, an image search engine called Yottixel has been designed and developed for application in pathology32,41,42,43. Yottixel is a portmanteau for one yotta pixel alluding to the big-data nature of pathology images. The underlying technology behind Yottixel consists of a series of AI algorithms, including clustering techniques, deep networks, and gradient barcoding. By generating a “bunch of barcodes” (BoB) for each WSI, digitized pathology slides can be indexed for real-time search. In other words, the tissue patterns of a WSI are converted into barcodes, a process that is both storage-friendly and computationally efficient. In this paper, we report the outcome of a comprehensive validation of the Yottixel search engine. We used WSI data from The Cancer Genome Atlas (TCGA) repository provided by the National Cancer Institute (NCI)/National Institutes of Health (NIH). Almost 30,000 WSI files of 25 primary anatomic sites and 32 cancer subtypes were processed by dismantling these large slides into almost 20,000,000 image patches (also called tiles) that were then individually indexed employing ~3,000,000 barcodes. We employ the largest publicly available archive of WSIs to verify the performance of an image search engine for digital pathology.
Results
Performance measurement of search engine
In two major series of experiments, we calculated the “accuracy” of image search through “leave-one-patient-out” samplings. Whereas the literature of computer vision focuses on top-n accuracy (if any one of the n search results is correct, then the search is considered be to be successful), we calculated the majority-n accuracy (only if the majority among n search results were correct, the search was considered correct). Specifically, “correct” means that the tumor type (horizontal search) or tumor subtype within a specific diagnostic category (vertical search) was recognized correctly and matched by the majority of identified and retrieved cases. In order to avoid falsification of results through anatomic duplicates, we excluded all WSIs of the patient when one of the WSIs was the query.
Horizontal search: cancer-type recognition
The first series of experiments undertaken for all anatomic sites was horizontal search. The query WSI is compared against all other cases in the repository, regardless of anatomic site categorization. Of course, the primary anatomic site is generally known, and, in many cases, the cancer type may also be known to the pathologist. Thus, the purpose of the horizontal search (which is for either organ or cancer-type recognition) is principally a fundamental algorithmic validation that may also have applications like searching for origin of malignancy in case of metastatic cancer.
The results of the horizontal search are depicted in Fig. 1 (see Appendix for details with Table 1 showing results for frozen section and Table 2 for permanent diagnostic slides). All experiments were conducted via “leave-one-patient-out” validation.
The following observations can be made from the results:
-
Provided there are sufficient number of patients, we observed that the more we retrieve the more likely it was to achieve the right diagnosis: top-10 is better than top-5, and top-5 is better than top-3.
-
General top-n accuracy that is common in the computer-vision literature (top-3, top-5 and top-10 column in Tables 1 and 2) show high values, but may not be suitable in the medical domain as it considers the search to be a success if at least one of the search results has the same cancer type as the query image.
-
The majority vote among top-n search results appears to be much more conservative and perhaps more appropriate, as it only considers a search task as successful if the majority of top-n search results show the same cancer type as the query image (majority-5 and majority-10 columns in Tables 1 and 2).
-
With some exceptions, a general trend is observable that the more images/patients are available the higher the search-based consensus accuracy. The number of cases positively correlated with the majority-vote accuracy for both frozen sections and permanent diagnostic slides.
Vertical search: correctly subtyping cancer
In the second series of experiments, we performed vertical search. Given the primary site of the query slide we confined the search only to WSIs from that organ. Hence, the goal of the vertical search was to recognize the cancer subtype. For this purpose, only those primary anatomic sites in the data set with at least two possible subtypes were selected. Sample retrievals are illustrated in Appendix Fig. 2. The results for “leave-one-patient-out” validation are depicted in Figs 3 and 4 (details in Appendix, Table 3 for frozen sections and Table 4 for diagnostic slides).
The top four slides are of permanent diagnostic slides whereas the bottom three slides are of frozen section slides. The misclassified and successful queries are marked with red and green boundaries, respectively (for abbreviations, see Table 5).
Looking at the results of Figs. 3 and 4 (Tables 3 and 4), we can observe the following:
-
For both frozen sections and permanent diagnostic slides, we continue to see a general trend whereby “the more patients the better” with both positive exceptions (KICH with 196 patients, and PCPG with 179 patients in Table 3) and negative exceptions (LUAD with 520 patients in Table 4).
-
With majority-vote accuracy values for frozen sections (Table 3) in excess of 90% (KIRC, GBM, COAD, UCEC, PCPG), a search-based computational consensus appear to be possible when a large number of evidently diagnosed patients are available.
-
With majority-vote accuracy values for diagnostic slides (Table 4) in excess of 90% (GBM, LGG, UCEC, KIRC, COAD, ACC, PCPG), a search-based computational consensus appear to be possible when a large number of evidently diagnosed patients are available.
-
In most cases, it appeared that taking the majority of the top-7 search results provided the highest accuracy in most cases. However, the accuracy dropped drastically for subtypes with a small number of patients as we retrieved more and more images beyond six slides, as the majority in such cases were taken from incorrect cases (we do not filter any result; no threshold is used; hence, all search results are considered as valid results).
-
Based on all observations, it seems that there is a direct relationship between the number of diagnosed WSIs in the data set and achievable consensus accuracy. For vertical search, we calculated positive correlations of \(0.5456\) for frozen sections (Table 3) and \(0.5974\) for permanent diagnostic slides (Table 4). This trend was more pronounced for horizontal search with positive correlation of \(0.7780\) for frozen sections slides (Table 1), and \(0.7201\) for permanent diagnostic slides (Table 2).
-
In addition, the Cox-Stuart trend test44 was used to check the upward monotonic trend of accuracy with respect to patients number. Having an increasing trend is considered as the null hypothesis for this test. The \(p\)-values for the horizontal (vertical) search are 1 (0.9991) and 0.9844 (0.9713) for frozen and diagnostic slides, respectively. Since the \(p\)-values are greater than the significance level (0.05), the null hypothesis is accepted. Consequently, there is a strong evidence of an upward monotonic trend.
Visualization of search results
Examining best, average, and worst cases for diagnostic slides, we randomly selected 3000 slides and visualized them using the T-distributed Stochastic Neighbor Embedding (t-SNE) method45 (see Fig. 5). From this visualization, we can observe that several subtype groups have been correctly extracted through search (see groups a to f). We can also observe the presence of outliers (e.g., DLBC in groups a and b). The outliers may be a product of the resolution of these scans, at least in part. At 20× magnification, for example, recognizing a diffuse large B-cell lymphoma (DLBC) from other large cell, undifferentiated non-hematopoietic tumors may not always be immediately possible for pathologists. This typically requires serial sections examined at multiple magnifications with ancillary studies such as immunohistochemistry.
These primary sites are selected to contain top, average, worst accuracy from the Table 2—lung, brain (top-2), kidney, liver (middle-2), lymph nodes, and pleura (bottom-2). Six different areas containing majority of the points from the same cancer subtype are assigned with unique alphabets—a, b, c, d, e, f. The random slides from the majority cancer subtype within each of the assigned areas are shown in Samples box (gray background). The outliers (not belonging to majority the cancer subtype or the primary site) are shown in the outliers box (red outline). For example, area a contains majority of scans from brain with glioblastoma multiforme (GBM), whereas its outliers are from lymph nodes with diffuse large B-cell lymphoma (DLBC). Without any explicit training, our technique maintains the semantic categories within the diagnostic slides as shows by the t-SNE plot of the pairwise distances. The kidney, liver, and brain form different isolated groups whereas lung, pleura, and lymph nodes are intermixed with each other.
The challenge of validating histologic similarity
One of the major benefits of using classification methods is that they can easily be validated; every image belongs to a class or not, a binary concept that can be conveniently quantified by counting the number of correctly/incorrectly categorized cases. It should be noted that through treating the image search as a classifier, we have not only used the primary diagnosis for “objective” evaluation of search results but also we are most likely ignoring some performance aspects of image search as search is a technology inherently suitable for looking at border cases and fuzziness of histologic similarity. The concept of similarity in image search is intrinsically a gradual concept (i.e., cannot be answered with a simple yes/no in many cases) and mostly a matter of degree (very similar, quite dissimilar, etc.). In addition, the similarity (or dissimilarity) between images is generally calculated using a distance metric/measure (in our case the Hamming distance46). The histologic similarity as perceived by pathologists may not correspond to tests where we used distance as a classification criterion. In other words, the classification-based tests that we run may be too harsh for search results and ignorant toward anatomic similarities among different organs.
One of the possible ways of examining the performance of the search is to look at the heatmap47 of the confusion matrix. The values to construct the heatmap can be derived from the relative frequency of every subtype among the top ten search results for a given subtype. A perfect heatmap would exhibit a pronounced diagonal with other cells being insignificant. Figure 6 shows the generated heatmap for all diagnostic subtypes in the data set. The ordering of subtypes along the \(y\)-axis was done manually. It should be noted that our matching heatmap is not symmetrical like a correlation-based heatmap.
Analysis of the heatmap
The pronounced diagonal in Fig. 6 shows that most disease subtypes have been correctly classified as they were very frequently retrieved among the top ten horizontal search results. Other obvious observations:
-
MESO is a difficult diagnosis with almost absent diagonal values.
-
READ and COAD build a confusion region of four squares; they are confused with each other frequently.
-
The same observation can be made for LUAD and LUSC. The vertical values for LUAD and LUSC also show that they are present in many other searches, for instance, when we search for UESC, HNSC, and ESCA.
-
LIHC is frequently among the search results for CHOL.
-
For PRAD and BRCA we predominantly found PRAD and BRCA images, respectively.
Of note, the observational analysis of the heatmap alone may be limited. If we cluster (group) the search result frequencies and construct the dendrograms for the relationships in order to create an advanced heatmap, we might more easily discover the benefits of the search (see Fig. 7). From there, we can observe:
-
LGG and GBM are both glial tumors of the central nervous system.
-
Rectum and colon cancer are gland forming tumors of the colon.
-
Both uterine and ovarian carcinoma are grouped under gynecological.
-
Gallbladder, stomach, and esophagus are upper gastrointestinal tumors.
-
Adenocarcinoma and squamous cell carcinoma are both subtypes of lung tumors.
-
Three kidney tumors appear close together.
The errors (i.e., misclassifications) identified were still within the general grouping that the tumor originated from. Hence, from an image search perspective, it suggests that is it good at being close to the site of origin when it makes “classification” errors.
Chord diagram of image search
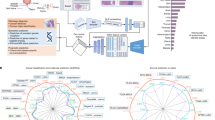
We used a chord diagram to further explore retrieved results. A chord diagram is the graphic display of the inter-relationships between numbers in a matrix. The numbers are arranged radially around a circle with the relationships between the data points generally visualized as arcs connecting the numbers/labels48. In Fig. 8a, the chord diagram of horizontal search (cancer-type recognition) for 11,579 permanent diagnostic slides of the TCGA data set is illustrated. We can observe the following:
-
Adenocarcinomas from several disparate organ systems match (e.g., colon, lung, stomach, and breast). This is not surprising, as adenocarcinomas formed by glandular structures of equivalent grade in most organs are morphologically similar.
-
Certain tumors derived from the same organ are related (e.g., LGG and GBM, UCEC and CESC, and kidney RCC and KIRP).
-
High-grade tumors from different anatomic locations appear to match (e.g., GBM and sarcoma). This may be attributed to the fact that such high-grade tumors likely display similar morphologic findings (e.g., necrosis).
-
Squamous tumors from the head and neck and lung resemble urothelial carcinoma from the urinary bladder. In clinical practice, this differential diagnosis can be morphologically challenging to diagnose, and thus warrants the use of ancillary studies such as immunohistochemistry to determine tumor origin.
-
Hepatocellular carcinoma and thyroid carcinoma appear to exhibit the greatest number of matches (eight to nine) to other tumor subtypes. The significance of this finding is unclear.
-
The broad relationship demonstrated among certain tumor subtypes is unexpected (e.g., cutaneous melanoma to sarcoma, LUSC, and adenocarcinoma from several organs). Indeed, melanoma is known as the great mimicker in pathology given that these melanocytic tumors can take on many morphological appearances.
a Chord diagram of horizontal image search for diagnostic slides of the TCGA data set. Sample relations for (b) brain (LGG and GBM), (c) pulmonary (LAUD, LUSC, and MESO), and (d) gynecological (UCEC, UCS, and CESC). The chord diagram can be interactively viewed online: https://bit.ly/2k6g3k1.
One has to emphasize that some relationships depicted in the chord diagram may disappear if distances are normalized and threshold applied. We did not filter any search results. No threshold was used. Hence, all search results were considered. The interactive version of TSNE plot is available online at http://dev1-kimia.uwaterloo.ca:5001/.
Discussion
The accelerated adoption of digital pathology is coinciding with and probably partly attributed to recent progress in AI applications in the field of pathology. This disruption in the field of pathology offers a historic chance to find novel solutions for major challenges in diagnostic histopathology and adjacent fields, including biodiscovery. In this study, we indexed and searched the largest publicly available data set of histopathology WSIs provided by the NIH/NCI. The question was whether one can build a computational consensus to potentially remedy the high intra- and inter-observer variability seen with diagnosing certain pathology tumors through search in a large archive of previously (and evidently) diagnosed cases. We performed a horizontal search to verify basic recognition capabilities of the image search engine. Furthermore, we performed leave-one-patient-out vertical searches to examine the accuracy of top \(n\) search results for establishing a diagnostic majority for cancer subtypes.
The results of this validation study show that building a computational consensus to assist pathologists with “virtual peer review” is possible if large and representative archives of well-characterized and evidently diagnosed cases are available. The ideal size of the data set appears to be in excess of several thousand patients for each primary diagnosis, and is most likely directly related to the anatomic complexity and intrinsic polymorphism of individual tissue types.
Whereas one may need substantial computational power (i.e., a set of high-performance GPUs) to index a large existing repository from scratch, the usage of bunch-of-barcodes idea makes the continuous indexing and search quite feasible for any laboratory, clinic, and hospital.
Since we used a mosaic (a set of patches) to represent and to retrieve WSIs, the search was guided to look for features present in multiple patches to classify the entire WSI. For detailed search, such as mitotic rates and grading applications, one needs a different data set and should also apply single-patch search to look for details. As well, regardless of implementation (e.g., onsite versus cloud), the validated search technology is completely safe toward patient-sensitive information as the barcodes do not contain any reversible information that could compromise patient privacy.
Future research should look into subtype consensus for individual primary diagnoses in more details for carefully curated data sets. As well, the need for much larger curated archives in the pathology community is clearly evident, which includes additional tissue types such as hematological. Lastly, comprehensive discordance measurement for subtypes with and without computational consensus should be planned and carried out as the ultimate evidence for the efficacy of the image search as a supportive diagnostic tool.
The intellectual property as well as the financial implications for related works emerging from sharing image repositories are certainly significant issues that need elaboration in future works.
Methods
Data collection
We used the publicly available data set of 30,072 WSIs from the TCGA project49,50 (Genomic Data Commons GDC). Due to the retrospective nature of this study using only publicly available data, ethics approval was not required. All WSIs are tagged with a primary diagnosis. We removed 952 WSIs due to the following reasons: poor staining, low resolution, lack of all magnification levels in the WSI pyramid, large presence of out-of-focus regions, and/or presence of unreadable regions within an image. Most WSIs had a magnification of 20\(\times\) or 40\(\times\), some at lower magnifications. In total, we processed 29,120 WSIs at 20\(\times\) magnification (approximately six terabytes in compressed form) for this study. The data set contains 25 anatomic sites with 32 cancer subtypes. Ten tumor types (brain, endocrine, gastrointestinal tract, gynecological, hematopoietic, liver/pancreaticobiliary, melanocytic, prostate/testis, pulmonary, and urinary tract) had more than one primary diagnoses. From the 29,120 WSIs, 26,564 specimens were neoplasms, and 2556 were non-neoplastic. A total of 17,425 files comprised frozen section digital slides, and 11,579 files were of permanent hematoxylin and eosin (H&E) sections. For the remaining 116 WSIs, the tissue section preparation was unspecified. We did not remove manual pen markings from the slides when present. The TCGA codes for all 32 cancer subtypes are provided in Table 5 in the appendix. The TCGA data set has a number of shortcomings50. Many of the cases are of frozen section in which tissue morphology may be compromised by frozen artifacts. Available cases may also reflect research bias in institutional biorepository collections. Furthermore, “tumors routinely subjected to neoadjuvant therapy may not have been able to be included in TCGA, because of limited availability of untreated specimens”50. Moreover, hematopathology is conspicuously absent from the TCGA data"set with just a few lymph nodes included. In spite of the shortcomings, the TCGA is the largest public data set that can support a pan-cancer validation of AI solutions for digital pathology.
The search algorithm
The Yottixel image search engine incorporates clustering, transfer learning, and barcodes and was used to conduct all experiments30,32,41,42,43,51,52,53,54. Before any search can be performed, all images in the repository have to be “indexed”, i.e., every WSI is catalogued utilizing a “bunch of barcodes” (BoB indexing). These barcodes are stored for later use and generally not visible to the user. This process contains several steps (Fig. 9):
-
Tissue extraction—Every WSI contains a bright (white) background that generally contains irrelevant (non-tissue) pixel information. In order to process the tissue, we need to segment the tissue region(s), and generate a black and white image (binary mask) that provides the location of all tissue pixels as “1” (white). Such a binary mask is depicted in the top row of Fig. 9.
-
Mosaicking—Segmented tissue now gets patched (divided into patches/tiles). These patches have a fixed size at a fixed magnification (e.g., 500 × 500 µm2 at 20\(\times\) scan resolution). All patches of the WSI get grouped into a pre-set number of categories (classes) via a clustering method (we used \(k\)-means algorithm55). A clustering algorithm is an unsupervised method that automatically groups WSI patches into clusters (i.e., groups) that contain similar tissue patterns. A small percentage (5–20%) of all clustered patches are selected uniformly distributed within each class to assemble a mosaic. This mosaic represents the entire tissue region within the WSI. A sample mosaic consisting of four patches is depicted in the second row of Fig. 9. Most WSIs we processed had a mosaic with around 70–100 patches.
-
Feature mining—All patches of the mosaic of each WSI are now pushed through pretrained artificial neural networks (generally trained with natural images using data sets such as ImageNet56). The output of the network is ignored and the last pooling layers or the first connected layers are generally used as “features” to represent each mosaic patch. There could be ~1000–4000 features. The third row of Fig. 9 shows this process where the features (colored squares) are passed on to the next stage, namely BoB indexing.
-
Bunch of barcodes—All feature vectors of each mosaic are subsequently converted into binary vectors using the MinMax algorithm43. This bunch of barcodes is the final index information for every query/input WSI that will be stored in the Yottixel index for future or immediate search. This is illustrated at the bottom of Fig. 9.
A mosaic of representative patches (tiles) is assembled through grouping of all patches of the tissue region using an unsupervised clustering algorithm (second block from the top). All patches of the mosaic are fed into a pretrained artificial neural network for feature mining (third block from the top). Finally, a bunch of barcodes is generated and added to the index of all WSI files in the archive (bottom block).
In summary, Yottixel assigns “a bunch of barcodes” to each WSI to index the entire digital slide. The BoB indexing enables Yottixel to search a large archive of histopathology images very efficiently. The index can be easily shared among institutions if necessary. Technical details of Yottixel algorithms are described in a separate paper where its performance was tested with 2300 WSIs41.
Reproducibility
Does image search generate the same results for the same WSI if fed into the Yottixel engine again? We ran indexing several times and the results did not change significantly. We observed slight changes in the order of search results affecting neither the hit rate nor the majority vote. The only component of our approach with some non-deterministic behavior is the K-means clustering algorithm. However, the K-means is run for as many iterations until it converges to a stable solution when we index WSIs. After a new WSI has been indexed its “bunch of barcodes” do not change anymore, and hence the same WSI as input (with unique patient ID) will generate the same results.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Code availability
The deep models used in this study were implemented using TensorFlow. The weights for pretrained models are openly provided by Keras libray. Statistical analysis was performed using Python libraries—Scikit Learn, Numpy, and Pandas. The Dask library was used as distributed computing framework for running large-scale computation jobs.
References
Janowczyk, A. & Madabhushi, A. Deep learning for digital pathology image analysis: a comprehensive tutorial with selected use cases. J. Pathol. Inform. 7, 29 (2016).
Madabhushi, A. & Lee, G. Image analysis and machine learning in digital pathology: challenges and opportunities. Med. Image Anal. 33, 170–175 (2016).
Tizhoosh, H. R. & Pantanowitz, L. Artificial intelligence and digital pathology: challenges and opportunities. J. Pathol. Inform. 9, 38 (2018).
Campanella, G. et al. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images. Nat. Med. 25, 1301–1309 (2019).
Guo, Z. et al. A fast and refined cancer regions segmentation framework in whole-slide breast pathological images. Sci. Rep. 9, 882 (2019).
Niazi, M. K. K., Parwani, A. V. & Gurcan, M. N. Digital pathology and artificial intelligence. Lancet Oncol. 20, e253–e261 (2019).
Xing, F., Xie, Y., Su, H., Liu, F. & Yang, L. Deep learning in microscopy image analysis: a survey. IEEE Trans. Neural Netw. Learn. Syst. 29, 4550–4568 (2017).
Lehmann, T. M. et al. Content-based image retrieval in medical applications. Methods Inf. Med. 43, 354–361 (2004).
Long, L. R., Antani, S., Deserno, T. M. & Thoma, G. R. Content-based image retrieval in medicine: retrospective assessment, state of the art, and future directions. Int. J. Healthcare Inf. Syst. Inform. 4, 1–16 (2009).
Markonis, D. et al. A survey on visual information search behavior and requirements of radiologists. Methods Inform. Med. 51, 539–548 (2012).
Müller, H., Michoux, N., Bandon, D. & Geissbuhler, A. A review of content-based image retrieval systems in medical applications—clinical benefits and future directions. Int. J. Med. Inform. 73, 1–23 (2004).
Sathya, R. & Abraham, A. Comparison of supervised and unsupervised learning algorithms for pattern classification. Int. J. Adv. Res. Artif. Intell. 2, 34–38 (2013).
LeCun, Y., Kavukcuoglu, K. & Farabet, C. Convolutional networks and applications in vision. in Proceedings of 2010 IEEE International Symposium on Circuits and Systems, 253–256 (IEEE, 2010).
Onder, D., Sarioglu, S. & Karacali, B. Automated labelling of cancer textures in colorectal histopathology slides using quasi-supervised learning. Micron 47, 33–42 (2013).
Elmore, J. G., Wells, C. K., Lee, C. H., Howard, D. H. & Feinstein, A. R. Variability in radiologists’ interpretations of mammograms. New Eng. J. Med. 331, 1493–1499 (1994).
Mussurakis, S., Buckley, D., Coady, A., Turnbull, L. & Horsman, A. Observer variability in the interpretation of contrast enhanced mri of the breast. Br. J. Radiol. 69, 1009–1016 (1996).
Burnett, R. et al. Observer variability in histopathological reporting of malignant bronchial biopsy specimens. J. Clin. Pathol. 47, 711–713 (1994).
Winkfield, B., Aubé, C., Burtin, P. & Calès, P. Inter-observer and intra-observer variability in hepatology. Eur. J. Gastroenterol. Hepatol. 15, 959–966 (2003).
Louie, A. V. et al. Inter-observer and intra-observer reliability for lung cancer target volume delineation in the 4d-ct era. Radiother. Oncol. 95, 166–171 (2010).
Cooper, W. A. et al. Intra-and interobserver reproducibility assessment of pd-l1 biomarker in non-small cell lung cancer. Clin. Cancer Res. 23, 4569–4577 (2017).
Lewis, J. S. Jr. et al. Inter-and intra-observer variability in the classification of extracapsular extension in p16 positive oropharyngeal squamous cell carcinoma nodal metastases. Oral Oncol. 51, 985–990 (2015).
Peck, M., Moffat, D., Latham, B. & Badrick, T. Review of diagnostic error in anatomical pathology and the role and value of second opinions in error prevention. J. Clin. Pathol. 71, 995–1000 (2018).
Strosberg, C. et al. Second opinion reviews for cancer diagnoses in anatomic pathology: a comprehensive cancer center’s experience. Anticancer Res. 38, 2989–2994 (2018).
Sasada, K. et al. Inter-observer variance and the need for standardization in the morphological classification of myelodysplastic syndrome. Leuk. Res. 69, 54–59 (2018).
Veltkamp, R. C. & Tanase, M. Content-Based Image Retrieval Systems: A Survey. A Report at the Department of Computing Science, 1–62 (Utrecht University, 2002).
Singhai, N. & Shandilya, S. K. A survey on: content based image retrieval systems. Int. J. Comput. Appl. 4, 22–26 (2010).
Zheng, L., Yang, Y. & Tian, Q. Sift meets cnn: a decade survey of instance retrieval. IEEE Trans. Pattern Anal. Machine Intell. 40, 1224–1244 (2017).
Babenko, A. & Lempitsky, V. Aggregating local deep features for image retrieval. in Proceedings of the IEEE International Conference on Computer Vision, 1269–1277 (IEEE, 2015).
Liu, H., Wang, R., Shan, S. & Chen, X. Deep supervised hashing for fast image retrieval. in Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 2064–2072 (IEEE, 2016).
Kieffer, B., Babaie, M., Kalra, S. & Tizhoosh, H. R. Convolutional neural networks for histopathology image classification: Training vs. using pre-trained networks. in 2017 Seventh International Conference on Image Processing Theory, Tools and Applications (IPTA), 1–6 (IEEE, 2017).
Rahman, M. M., Bhattacharya, P. & Desai, B. C. A framework for medical image retrieval using machine learning and statistical similarity matching techniques with relevance feedback. IEEE Trans. Inf. Technol. Biomed. 11, 58–69 (2007).
Tizhoosh, H. R. Barcode annotations for medical image retrieval: a preliminary investigation. in 2015 IEEE International Conference on Image Processing (ICIP), 818–822 (IEEE, 2015).
Qayyum, A., Anwar, S. M., Awais, M. & Majid, M. Medical image retrieval using deep convolutional neural network. Neurocomputing 266, 8–20 (2017).
Farahani, N., Parwani, A. V. & Pantanowitz, L. Whole slide imaging in pathology: advantages, limitations, and emerging perspectives. Pathol. Lab. Med. Int. 7, 23–33 (2015).
Liu, Y. & Pantanowitz, L. Digital pathology: review of current opportunities and challenges for oral pathologists. J. Oral Pathol. Med. 48, 263–269 (2019).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436 (2015).
Goodfellow, I., Bengio, Y. & Courville, A. Deep Learning (MIT Press, 2016).
Komura, D. & Ishikawa, S. Machine learning methods for histopathological image analysis. Comput. Struct. Biotechnol. J. 16, 34–42 (2018).
Shi, X. et al. Supervised graph hashing for histopathology image retrieval and classification. Med. Image Anal. 42, 117–128 (2017).
Komura, D. et al. Luigi: Large-scale histopathological image retrieval system using deep texture representations. Preprint at https://www.biorxiv.org/content/10.1101/345785v2 (2018).
Kalra, S., Choi, C., Shah, S., Pantanowitz, L. & Tizhoosh, H. R. Yottixel—an image search engine for large archives of histopathology whole slide images. Preprint at https://arxiv.org/abs/1911.08748 (2019).
Kumar, M. D., Babaie, M. & Tizhoosh, H. R. Deep barcodes for fast retrieval of histopathology scans. in 2018 International Joint Conference on Neural Networks (IJCNN) 1–8 (IEEE, 2018).
Tizhoosh, H. R., Zhu, S., Lo, H., Chaudhari, V. & Mehdi, T. Minmax radon barcodes for medical image retrieval. in International Symposium on Visual Computing, 617–627 (Springer, 2016).
Cox, D. R. & Stuart, A. Some quick sign tests for trend in location and dispersion. Biometrika 42, 80–95 (1955).
Maaten, L. v. d. & Hinton, G. Visualizing data using t-sne. J. Machine Learn. Res. 9, 2579–2605 (2008).
Bookstein, A., Kulyukin, V. A. & Raita, T. Generalized hamming distance. Inf. Retriev. 5, 353–375 (2002).
Wilkinson, L. & Friendly, M. The history of the cluster heat map. Am. Stat. 63, 179–184 (2009).
Holten, D. Hierarchical edge bundles: visualization of adjacency relations in hierarchical data. IEEE Trans. Vis. Comput. Graph. 12, 741–748 (2006).
Tomczak, K., Czerwińska, P. & Wiznerowicz, M. The cancer genome atlas (tcga): an immeasurable source of knowledge. Contemp. Oncol. 19, A68 (2015).
Cooper, L. A. et al. Pancancer insights from the cancer genome atlas: the pathologist’s perspective. J. Pathol. 244, 512–524 (2018).
Chenni, W., Herbi, H., Babaie, M. & Tizhoosh, H. R. Patch clustering for representation of histopathology images. in European Congress on Digital Pathology, 28–37 (Springer, Cham, 2019).
Tizhoosh, H. R. & Babaie, M. Representing medical images with encoded local projections. IEEE Trans. Biomed. Eng. 65, 2267–2277 (2018).
Tizhoosh, H. R. & Czarnota, G. J. Fast barcode retrieval for consensus contouring. Preprint at https://arxiv.org/abs/1709.10197 (2017).
Tizhoosh, H. R., Mitcheltree, C., Zhu, S. & Dutta, S. Barcodes for medical image retrieval using autoencoded radon transform. in 2016 23rd International Conference on Pattern Recognition (ICPR), 3150–3155 (IEEE, 2016).
Jain, A. K. Data clustering: 50 years beyond k-means. Pattern Recogn. Lett. 31, 651–666 (2010).
Deng, J. et al. Imagenet: a large-scale hierarchical image database. in 2009 IEEE Conference on Computer Vision and Pattern Recognition, 248–255 (IEEE, 2009).
Acknowledgements
We would like to thank the Ontario Government for awarding an ORF-RE grant for this project (Ontario Research Fund—Research Excellence). The first author is supported by an MITACS internship. The basic research of the corresponding author leading to this work has been supported by The Natural Sciences and Engineering Research Council of Canada (NSERC) through multiple Discovery Grants. For access to computational and storage facilities, we like to thank Dell EMC and Teknicor for their generous support.
Author information
Authors and Affiliations
Contributions
S.K. conducted most experiments and collected all results. H.R.T. designed the indexing and search algorithms developed the first prototype in Matlab, supervised Python conversion and the validation process, analyzed the results, and conceptualized/wrote most of the paper. S.S. developed the indexing engine in C++ and run many experiments. C.C. prototyped most algorithms in Python with contributions to the patch grouping. S.D. supervised parts of the validation and critically reviewed the paper. S. Shafiei, A.S., and M.B. contributed to many critical discussions at different stages of algorithm development. P.D., C.J.V.C., and L.P. critically reviewed the paper and analyzed/validated the results. L.P. provided advice over a period of several months to assist in development of use cases.
Corresponding author
Ethics declarations
Competing interests
S.K. is a PhD intern at Huron Digital Pathology. H.R.T. is an advisor to Huron Digital Pathology. H.R.T. is the principal investigator of the ORF-RE grant (Ontario Research Fund—Research Excellence) sponsored by Huron Digital Pathology. L.P. is on the medical advisory board for Ibex and a Consultant for Hamamatsu. The remaining authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kalra, S., Tizhoosh, H.R., Shah, S. et al. Pan-cancer diagnostic consensus through searching archival histopathology images using artificial intelligence. npj Digit. Med. 3, 31 (2020). https://doi.org/10.1038/s41746-020-0238-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41746-020-0238-2