Abstract
Transcriptional regulators include a superfamily of nuclear proteins referred to as co-activators and co-repressors, both of which are involved in controlling the functions of several nuclear receptors (NRs). The Nuclear Receptor Signaling Atlas (NURSA) has cataloged the composition of NRs, co-regulators, and ligands present in the human cell and their effort has been identified in more than 600 potential molecules. Given the importance of co-regulators in steroid, retinoid, and thyroid hormone signaling networks, hypothesizing that NRs/co-regulators are implicated in a wide range of pathologies are tempting. The co-activators known as peroxisome proliferator-activated receptor gamma co-activator 1 (PGC-1) and their key nuclear partner, the estrogen-related receptor (ERR), are emerging as pivotal transcriptional signatures that regulate an extremely broad repertoire of mitochondrial and metabolic genes, making them very attractive drug targets for cancer. Several studies have provided an increased understanding of the functional and structural biology of nuclear complexes. However, more comprehensive work is needed to create different avenues to explore the therapeutic potential of NRs/co-activators in precision oncology. Here, we discuss the emerging data associated with the structure, function, and molecular biology of the PGC-1/ERR network and address how the concepts evolving from these studies have deepened our understanding of how to develop more effective treatment strategies. We present an overview that underscores new biological insights into PGC-1/ERR to improve cancer outcomes against therapeutic resistance. Finally, we discuss the importance of exploiting new technologies such as single-particle cryo-electron microscopy (cryo-EM) to develop a high-resolution biological structure of PGC-1/ERR, focusing on novel drug discovery for precision oncology.
Similar content being viewed by others
Introduction
Transcriptional regulators comprise of nuclear proteins known as co-activators and co-repressors, which bind and control the functions of nuclear receptors (NRs) and transcription factors (TFs).1,2,3,4 The essential role of NRs and their co-factors in many aspects of mammalian development and physiology raises the possibility that dysfunctions in biological signaling networks controlled by receptors or co-activators, which could be associated with metabolic diseases.5,6
Under normal physiological conditions, the proliferator-activated receptor gamma co-activator 1/estrogen-related receptor (PGC-1/ERR) transcriptional axis is involved in the control of mitochondrial biogenesis.7,8 Mitochondria are considered to be key regulatory organelles that control cellular survival and death mechanisms, including biomass and energy production for rapid cell growth and apoptosis, respectively.9 Hence, many reports support the concept that the PGC-1/ERR pathway plays a dual role in cancer, depending on the specific cellular or tissue context and the environmental stimuli.10,11,12,13,14,15 Notably, the PGC-1/ERR axis has been shown to be essential for functional cancer cell motility and metastasis, leading to malignant transformation in breast and melanoma cancer progression.14,16,17 In contrast, this pathway has also been shown to suppress prostate cancer progression and metastasis.13,18,19 Although substantial progress has been made in increasing the understanding of the function and molecular biology of NRs and their co-activators, a considerable gap still exists in comprehending how the PGC-1/ERR axis integrates mitochondrial activity through oxidative phosphorylation (OxPhos) leading to cell survival or cell death and how this regulatory function is connected to its dual role in cancer progression.
Additional studies have been conducted that have led to a more comprehensive knowledge of the structure, function, and molecular biology of PGC-1/ERR signaling in cancer biology.20,21,22,23,24 Notably, accumulating evidence supports the importance of the PGC-1/ERR transcriptional axis in the context of metabolic-addicted cancer cells.12,25,26,27,28,29 This increases the significance of exploiting the therapeutic potential of these targets in an effort to predict the efficacy of therapeutic resistance, the mechanism of which relies on mitochondrial metabolic plasticity (Fig. 1).12,20,30,31,32 This therapeutic potential depends on the function of PGC-1/ERR in coordinating the activity of a broad repertoire of target gene expression associated with mitochondrial biogenesis, OxPhos, energy homeostasis, and glucose, glutamine and lipid metabolism (Table 1).22,33 Notably, targeting the PGC-1/ERR network could be accessed by exploring the potential of mitochondrial-linked weak spots, where selective inhibitors of the PGC-1/ERR axis and mitochondrial metabolism might have to be used in combination to target the metabolic addiction of specific cancer cells (Table 1 and Fig. 2).34
The PGC-1/ERR network as a potential onco-metabolic target in cancer treatment. a Cancer translational research methodology based on the patient-derived xenograft (PDX) model highlighting drug resistance in cancer. The survival mechanism of residual cells after conventional chemotherapy relies on metabolic plasticity. b The concept of metabolic vulnerability associated with cancer progression can be exploited by targeting a combination of the PGC-1/ERR network and several other mitochondrial weak spots, such as respiratory chain defects, TCA cycle enzymes, including citrate synthase (CS), isocitrate dehydrogenase (IDH), succinate dehydrogenase (SDH) and fumarate hydratase (FH). These enzymes might be exploited as potential onco-metabolic targets, depending on the specific type of cancer
Cryo-EM technology as an important tool for precision oncology. a The new cyro-EM technology could provide a better understanding of the native/physiological state of protein complexes for the development of promising therapeutic targets and drug discovery for precision oncology. Reprinted from publication: Merk, A. et al., Breaking Cryo-EM Resolution Barriers to Facilitate Drug Discovery, 1698-707, 2016, with permission from Elsevier.126 b Mitochondrial inhibitors, like metformin, enasidenib or etomoxir, have been used to treat cancer patients. An ideal scenario to exploit the resistance mechanism in cancer cells that seems to rely on OxPhos activity could be achieved using the concept of metabolic vulnerability, when a combination of targetable genes leads to a lethal phenotype. The PGC-1/ERR axis has great potential to be included as a potential metabolic target for precision oncology for the treatment of non-responder patients. Thus, the combination of mitochondrial inhibitors and inverse agonists of the PGC-1/ERR axis, like compound 29, might provide new hope for treating non-responder patients whose cancer cell survival mechanism relies on mitochondrial metabolism
In an effort to provide new insights into understanding the role of the PGC-1/ERR network, we suggest that new technologies, such as cryo-electron microscopy (cryo-EM), could provide mechanistic comprehension of the biological processes of these protein complexes. The use of cryo-EM could explain the conflicting observations of biochemistry and crystallography that focused only on individual domains or peptides of some of the molecules involved. Moreover, using cryo-EM will allow the investigation of the native state of the PGC-1/ERR complex, which might lead to the identification of novel mechanistic insights into the PGC-1/ERR biology, as well as the discovery of new binding-partners, which could open new therapeutic windows for targeting the PGC-1/ERR complex and its metabolism in precision oncology. For example, Yi et al., used cryo-EM and observed a novel biological insight in the transcriptional activity of the estrogen-receptor (ER) co-activator complex on DNA.35 The authors suggested that ER recruits two steroid receptor co-activator 3 proteins and one p300 protein from a DNA-bound complex. The structure of the ER-co-activator complex provided an initial step toward the understanding of the assembly of a full transcriptionally active NR-co-activator complex. Thus, studying the PGC-1/ERR complex in its native state that could shed new light on mechanisms of cancer resistance, which could be better exploited as a therapeutic strategy.
Also, applications of novel technologies that can shed new light on high-resolution biological structures are urgently needed as potential tools to fully elucidate the function and molecular biology of the PGC-1/ERR network in order to be able to further develop promising therapeutic targets and drug discovery for precision oncology (Fig. 2).36 Advances in cryo-EM are enabling structure determination of smaller protein complexes without additional modifications such as those required for crystallization that restricts the ability to fully access the mechanistic basis of how cancer metabolism can be orchestrated by these NRs and co-factors.
The main purpose of this review is to provide a critical understanding of the structural biology and function of the PGC-1/ERR network derived from work over the past decade. Moreover, focusing on promising therapeutic targets for precision oncology, this review will explore the underlying potential mechanisms of mitochondrial metabolic targets that could be exploited in combination with the PGC-1/ERR network to improve patient care against therapeutic resistance (Table 1).30,37 Understanding this signaling axis could yield crucial insights for the development of novel drugs and therapeutic strategies. This knowledge could lead to a better understanding of the specific type of cancer and patients who are responders and who would benefit from the pharmacological targeting of the PGC-1/ERR network. Thus, the PGC-1/ERR transcriptional axis fits into a novel category of targets that could be useful for exploitation in future research in personalized cancer medicine, so called precision oncology (Fig. 2).
Structure, function, and molecular biology of PGC-1 and ERR
The PGC-1 co-activator family comprises three different members, PGC-1α, PGC-1β, and PGC-1 related co-activator (PRC). Peroxisome proliferator-activated receptor gamma co-activator 1 alpha (PGC-1α) was first reported to regulate thermogenesis, interacting with the nuclear receptor peroxisome proliferator-activated receptor gamma (PPARγ) in brown adipose tissue (BAT).38 The other two members of the PGC-1 family, PGC-1β (PERC) and PRC were described by using sequence homology against PGC-1α.39,40
All members of the PGC-1 co-activator family share extensive protein sequence similarity and distinct domains, which could explain their similar physiological role and protein binding-patterns that have already been described.22 The human PPARGC1A gene on chromosome 4 encodes a 798-amino acid protein also known as PGC-1α. Its homologues, PGC-1β and PRC, comprise 1023 amino acids and 1664 amino acids, respectively. They are encoded by the human PPARGC1B gene on chromosome 5 and the human PPRC1 gene on chromosome 10. Notably, several truncated variants of PGC-1α with distinct transcripts and protein structures have been described. Basically, these variants are generated by alternative splicing and/or differential promoter usage.41,42 The presence of different variants of PGC-1α suggests that different protein variants might have distinct transcripts and protein structures with diverse functions, expression levels, and protein–protein interactions, depending on tissue-type or specific disease-context.41 More structural and functional studies are needed to address the mechanisms of the regulation of PGC-1α variants and to determine the presence of different variants associated with PGC-1β and PRC.
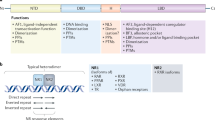
Structurally, all three members of the PGC-1 co-activator family and all nine variants of PGC-1α have one activation domain localized at their N-terminal region, which contains at least two LXXLL nuclear receptor box motifs.38,43 These specific leucine-rich repeats can bind with several NRs, especially the estrogen-related receptor family (ERRα, ERRβ, and ERRγ).44 The C-terminal region of all full-length PGC-1s contains a well-conserved RNA-binding domain, including short serine/arginine-rich stretches (RS), a nuclear localization signal (NLS), and the RNA recognition motif (RRM). The RS domain is present only in PGC-1α and PRC, suggesting that both members regulate RNA splicing and processing of mRNAs. However, further investigation is needed for determining whether the RS domain could be involved in processing PGC-1α variants by alternative splicing.45 The NLS domain plays a role in the maintenance of PGC-1s inside the nucleus. This domain is missing in alternative PGC-1α variants, which might be found in different cellular compartments, such as the cytosol or mitochondria.46,47 The RRM motif also regulates RNA splicing processing of mRNAs, whose function still needs to be substantiated. In addition, other conserved domains have been described in the co-activator family. For example, the aspartic acid (D), histidine (H), aspartic acid (D), and tyrosine (Y) tetrapeptide has been reported to be a binding-partner of the host cell factor (HCF) protein, a transcription factor that regulates gene expression during cell cycle progression. Another motif, including three threonine and four proline amino acids (TPPTTPP), is present in all full-length protein members but the function still needs to be determined.48
Functionally, PGC-1α was described as a docking platform for the assembly of transcriptional machinery, forming a macromolecular complex at specific DNA sequences to drive target gene expression.38 The same group that first described its activity reported that PGC-1α, even without any histone acetyl transferase activity, promotes gene transcription activity through the formation of a multi-protein complex encompassed by histone acetyltransferase proteins, such as cAMP response element-binding protein-binding protein/p300 and steroid receptor co-activator 1 (SRC-1).49 Later, PGC-1α was reported to bind with protein acetyl transferase p300 and the TRAP/mediator complex, mediator of RNA polymerase II transcription subunit 1, leading to the coordination of an important mechanism of chromatin remodeling.50 The idea that the interaction between a co-activator and NRs could recruit proteins responsible for chromatin remodeling, histone acetyl transferase activity, and transcriptional activity has emphasized the complex biological network involved in the PGC-1 axis. In vivo studies using double knockout (PGC-1α and PGC-1β) mice suggested that both members of the family share a similar role in the maintenance of mitochondrial function and energetic metabolic demand in many tissues.51,52 Conversely, the attempt to generate PRC-knockout mice failed because deleting this gene resulted in embryonic lethality.53 Nonetheless, in vitro data have shown that PRC plays an important role in mitochondrial biogenesis, but responds to proliferative signals leading to increased cell growth.54,55
The molecular biology of PGC-1 has been extensively exploited in different fields of health-related research, including cancer, diabetes, cardiovascular disease, and obesity.56 The expression of PGC-1α is characterized by high expression levels in tissues, including kidney, skeletal muscle, liver, heart, neural tissue, and blood mononuclear cells, which exhibit greater energy demand caused by increased mitochondrial activity.57,58,59 The vast number of different tissues or physiological contexts in which PGC-1α is expressed reflects the large number of different NRs and TFs that are regulated by PGC-1α, possibly including all three estrogen-related receptors (ERRα, ERRβ, and ERRγ), SRC-1, and glucocorticoid receptors (GR), as well as the tumor suppressor p53, PPARγ, forkhead box protein O1 (FoxO1), hepatocyte nuclear factor 4α (HNF-4α), nuclear respiratory factor 1 (NRF-1), the cAMP response element binding protein (CREB), and the signal transducer and activator of transcription 6 (STAT-6).44,60,61,62
Most intriguing, the activity of PGC-1α can also be regulated by post-translational modification (PTM) mechanisms.63 In fact, most of these regulatory mechanisms are dictated by PTMs, including phosphorylation, acetylation, methylation, ubiquitylation, and O-glycosylation. Clearly, this emphasizes the complex molecular biology and function of the PGC-1 family of proteins. For example, PGC-1α acts as a master regulator of mitochondrial metabolism mediating the entire demand of acetyl-groups and methyl-groups through the tri-carboxylic acid (TCA) cycle and amino acid metabolism, leading to feedback responses of PTM mechanisms, such as acetylation and methylation.64,65 Moreover, PGC-1α contributes to ATP production through the OxPhos process to supply the demand of ATP necessary for phosphorylation processes.66 Thus, PGC-1α is emerging as a fascinating transcriptional metabolic co-regulator playing a role in the maintenance of a tight equilibrium between metabolic precursors and energy production for sustaining PTM mechanisms.67 Besides the complex molecular biology of PGC-1α, more studies need to be conducted to characterize the function of other members of the family.
The molecular biology of PGC-1α is also associated with oncogenes and tumor suppressors. Several studies have demonstrated the interplay between PGC-1α and several oncogenes, including hypoxia-inducible factor 1-alpha (HIF-1α), oncogene carried by the Avian virus, Myelocytomatosis (c-Myc), vascular endothelial growth factor, protein kinase B, and B-Raf proto-oncogene, serine/threonine kinase/microphthalmia-associated transcription factor (MITF), as well as tumor suppressor p53 and 5’ AMP-activated protein kinase (AMPK).16,62,68,69,70 For example, higher PGC-1α expression predicts poor outcome in human melanoma, when the expression of PGC-1α is regulated by the MITF increasing mitochondrial function and resistance to oxidative stress.16 Moreover, in wild-type p53 lung adenocarcinoma, PGC-1α binds with p53, promoting cell survival in the presence of metabolic stress.62,71 However, the mechanism by which PGC-1α directly modulates oncogenic and tumor suppressor signaling in cancer cells is still unclear.
Different members of the PGC-1 family can act through similar or different molecular mechanisms depending on the cancer type and the stage of disease.22 Both PGC-1α and PGC-1β exhibit a similar tissue-specific expression pattern.44 As indicated above for PGC-1α, the molecular biology of PGC-1β in cancer is also associated with oncogenes, such as HIF-1α and c-Myc, and the tumor suppressor gene known as Von Hippel-Lindau (VHL).72 c-Myc is known to control the transcription of the gene encoding PGC-1β73 and the VHL/HIF-1 pathway can act as a repressor of c-Myc.72 For instance, the loss of PGC-1β and PGC-1α expression is a major factor contributing to impaired mitochondrial respiration in VHL-deficient renal carcinoma cells.72,74 Interestingly, the molecular biology of PGC-1β and PGC-1α seems to be modulated toward a metabolic rewiring through the VHL/HIF-1 pathway, leading to “the Warburg effect”, instead of mitochondrial OxPhos metabolism, through the inhibition of the transcriptional c-Myc/PGC-1β axis. Therefore, targeting PGC-1β and PGC-1α and rescuing the mitochondrial metabolic phenotype could be exploited as a therapeutic approach for the treatment of VHL-deficient renal carcinoma cells.74 Furthermore, a genomic study designed to assess the activity of ERRα in eight hundred breast tumor samples suggested that the molecular biology of PGC-1β in breast cancer progression relies on the c-Myc pathway. The authors had shown that the insulin-like growth factor 1 receptor pathway controls the stabilization of the c-Myc protein, leading to the up-regulation of PGC-1β.75
PGC-1α and PGC-1β were also reported to play a role in the resistance of ER-positive/tamoxifen-sensitive breast tumors.76 The interaction between PGC-1β and ERRα mediates a positive transcriptional regulation of receptor tyrosine-protein kinase (ERBB2) expression and co-amplifies genes associated with the ERBB2 amplicon. This biological mechanism was reported as a major factor contributing to tamoxifen resistance in a breast cancer model.76 Moreover, PGC-1β mediates adaptive resistance to genotoxic stress in lung cancer associated with mitochondrial DNA mutations.77
Taken together, these findings highlight a similar oncogenic network and resistance-related mechanisms in the cancer biology of both members of the PGC-1 family. On the other hand, the third member of the PGC-1 family, called PRC, seems to be restricted to the regulation of the expression of the mitochondrial biogenesis genes in proliferating cells. In contrast to both PGC-1α and PGC-1β, more studies are still needed to determine whether PRC plays a role in cancer progression by associating with oncogenic pathways.44 However, one study has recently suggested that the molecular biology of PRC could be associated with the c-Myc pathway in response to mitochondrial stress.78
Progress has been made in understanding the signaling network between PGC-1α and ERRα. This network has emerged as an important nuclear transcriptional axis and metabolic signaling pathway in regulating metabolic adaptation in specific cancer types, including breast cancer, prostate cancer, and melanoma.12 A majority of the literature has described the function of the founding members of the co-activator family, PGC-1α and ERRα, as a pivotal axis that might be linked to metabolic addiction of specific cancer cells that rely on mitochondrial metabolism for survival.
ERR
The ERR family encompasses three different members, ERRα (NR3B1), ERRβ (NR3B2), and ERRγ (NR3B3).79,80 All three members belong to a subfamily of orphan NRs, sharing sequence homology with the estrogen receptor (ER), but do not require endogenous ligands for activation. In fact, ERRs were first discovered by cDNA library screening using the ERα homology sequence to identify novel steroid receptors.80 The DNA-binding domains (DBD) and the ligand-binding domains (LBD) of ERRα and ERα are present in both classical estrogen receptor and orphan receptor families of NRs, but display different molecular biology and function, such as hormone-independent transcriptional activation and ERRα binding with co-activators.81 For instance, the ERR-LBD is only 36% similar with the ERs, which could explain why some ERα ligands, such as 17β-estradiol, estrone, and estriol do not activate ERRα.
Structurally, the N-terminal region of ERRs contains a DNA-binding domain and a ligand-independent transcriptional activation function (AF-1) that is poorly conserved within the ERR family members. The N-terminal AF-1 domain can weakly bind several TFs. The recent advances in structural biology have shown that this domain is subjected to various post-translational modifications (e.g., sumoylation and phosphorylation). Only a few groups have described the high-resolution structure of AF-1, which is at least partially due to the very flexible structure and low-affinity nuclear co-activator-binding.82,83 The DBD domain can also be acetylated, which controls the affinity by which ERR can bind with its own element responsive sequence (ERRE).84 Interestingly, the DBD domain is well-conserved across the members of this subfamily of NRs and contains two highly conserved zinc finger motifs with a specific-DNA binding sequence, TCAAGGTCA. The ERRE is exclusive for the DNA-binding sequence of all three ERR members. However, ERR members have been shown to bind with the responsive element sequence of ER (ERE), suggesting that ERRs play an important role in regulating similar downstream genes controlled by the ER pathway.82 The C-terminal region of ERRs comprises an LBD containing a conserved AF-2 helix motif. The LBD of ERR is required for the physical protein interaction with co-activator and co-repressor proteins, such as PGC-1α and PGC-1β or receptor-interacting protein 140 and nuclear receptor co-repressor 1, respectively. The conformation of the LBD of ERR, even in the absence of ligand, is responsible for constitutive transcriptional activation of ERR, due to its unique conformation that facilitates the recruitment of nuclear co-activators.79 The LBD domain of ERR binds with the nuclear receptor box motif LXXLL of PGC-1α, forming a binary complex, the stoichiometry of which is defined as two molecules of ERR (dimer) and one molecule of PGC-1α.21 Despite the importance of co-repressors in the context of ERR function, we focus herein on the structure, function, and molecular biology of ERR and nuclear co-activators, concentrating on how the PGC-1/ERR network can be exploited as a promising therapeutic target to improve patient outcome. The role of co-repressors and ERRs has been extensively reviewed.82
Previously, the observation of a largely occluded ligand-binding pocket in the transcriptionally active conformation of ERRα that has led some to the conclusion that ERRα does not lend itself to direct activation by small molecule agents. However, the crystal structure of the ERRα–LBD has presented an opportunity for generating selective inverse agonists.85,86 Currently, a series of diaryl ether-based thioazolidinediones has been screened resulting in the identification of specific inverse agonists of ERRα.87 Interestingly, compound 29 was obtained for the ERRα–LBD by solving the X-ray crystal structure.85 Compound 29 acts as a ligand of ERRα through a covalent interaction leading to conformational changes (in the amino acid Phe328) that disrupts the interaction between ERRα and PGC-1α.87 The biological consequence of this binding has shown growth-inhibitory therapeutic effects in certain cancers such as breast cancer and melanoma.17,29,88 Furthermore, several other compounds have been developed as inverse agonists of ERRα, such as compound 1a, compound 3, N-arylindole, XCT790, and GSK0903. However, more biological study is needed to determine the specificity of these compounds in targeting only the PGC-1/ERR network. For instance, XCT790 was previously developed as a specific inverse agonist of ERRα with the capacity to disrupt the interaction between ERRα and PGC-1α,89 leading to growth-inhibitory therapeutic effects in breast cancer.29 However, XCT790 does not seem to be a very specific inverse agonist of ERRα because at nanomolar concentrations (10-fold lower than the concentration required to inhibit ERRα), XCT790 is a potent mitochondrial uncoupler, leading to a rapid depletion of ATP and activation of AMPK. The authors suggested that XCT790 is a potent, fast-acting, mitochondrial uncoupler that acts independent of its inhibition of ERRα.90
Notably, Kallen and co-workers showed for the first time the X-ray crystal structure of the ERRα-LBD with one co-activator peptide derived from PGC-1α, leading to a ligand-independent transcriptional activation by ERR.91 In 2007, the same group used an inverse agonist called compound 1a to show that the binding interface of this compound with the ERRα-LBD comprised the helix H12 together with helices H3 and H4. Interestingly, the compound imposed dramatic conformational changes in the amino acid Phe328 located at H3, moving away the amino acid Phe510 of H12, which contains the co-activator groove of the ERRα-LBD. Based on this evidence, they proposed a novel molecular mechanism supporting the idea that the helix H12 binds with the co-activator peptide or compound 1a filling the co-activator groove of the AF-2 domain of ERRα. Hence, no co-activators and co-repressors are allowed to interact with ERRα.85 The X-ray crystal structure of the ERRα-LBD with compound 29 displayed similar dramatic conformational changes and side chain rotation of the amino acid Phe328 (H3). The significant change in structure was observed in the loop between helices H11 and H12. Hence, the C-terminal AF-2 domain that is responsible for transducing the constitutive activity for ERRα is no longer functional.87
As indicated earlier, the function and molecular biology of ERRs under normal physiological conditions are associated with the regulation of metabolic genes that are involved in glycolysis, the TCA cycle, and mitochondrial metabolism. These receptors also influence enzymes participating in OxPhos (e.g., several components of mitochondrial respiratory complexes), amino acid metabolism, and lipid synthesis (Table 1).
However, in the context of cancer, the different members of the ERR family seem to exhibit distinct functions in cancer progression. ERRα expression is associated with poor prognosis in breast tumors because it appears to drive lapatinib-resistance and tamoxifen-resistance in those patients.27,92 Several studies have shown that ERRα expression is associated with an increased risk of recurrence and worse prognosis, as well as drug resistance in patients with breast cancer.92,93,94 ERRα has been shown to increase the expression of ERBB2, mediating endocrine-resistant ERα-positive cells.76,95 Interestingly, ERRα has been shown to mediate pro-survival functions and represents a novel therapeutic target in a particularly aggressive melanoma phenotype, known as PGC-1α-positive melanomas.17
In contrast, the role of the ERRγ isoform in cancer biology seems to be paradoxical, whether this NR functions as an oncogene or as a tumor suppressor. Recently, genomic analysis in gastric cancer revealed that ERRγ acts as a tumor suppressor by directly targeting the Wnt signaling pathway. In fact, activating ERRγ expression by a specific agonist, DY131, inhibits gastric cancer cell growth and improved patient prognosis.96 Furthermore, ERRγ was described as an anti-proliferative target in androgen-sensitive and androgen-insensitive prostate cancer cells.97 Conversely, ERRγ is up-regulated in liver cancer and its inhibition suppresses cancer cell survival through the p21 and p27 proteins.98
Most intriguing is the paradoxical function of ERRγ in breast cancer progression. Recently, several studies reported that the hyper-activation of ERRγ induces a pro-survival transcriptional program in tamoxifen-treated breast cancer, as reported to the isoform ERRα.99,100 Conversely, in breast tumors co-expressing ER and PR, ERRγ induces E-cadherin expression and promotes the mesenchymal-to-epithelial transition (MET), resulting in the inhibition of tumor growth.101,102 In spite of the inconsistencies, the expression of ERRα and ERRγ in breast cancer and prostate cancer seems to be inversely correlated, where the ERRα is associated with a more aggressive disease and the expression of ERRγ is associated with a favorable prognosis of patients with breast and skin cancer.81,103 For instance, in androgen-dependent and castration-resistant prostate cancer, the progression of the disease is associated with a loss of ERRγ expression, whereas strategies to reactivate ERRγ expression could be exploited as a generalized therapeutic approach to manage prostate cancer.19
Overall, ERRα and ERRγ are considered key regulators of metabolic reprogramming in breast and prostate cancer. However, how this nuclear receptors network influences the metabolic state of cancer cells seems to be very complex and diverse, depending on the cancer type. Regarding the third isoform of this family, in prostate cancer the expression of ERRβ transactivates a promoter upstream of the cyclin-dependent kinase inhibitor, p21 gene, resulting in the inhibition of cell cycle progression, whereas the potential role of ERRβ in breast cancer remains unclear.104
Clearly a close relationship exists between PGC-1/ERR activity and cancer therapeutic resistance. New insights into the PGC-1α/ERRα network in cancer will be discussed next.
New biological insights into the PGC-1α co-activator and ERRα
Reprogramming of energy metabolism and evading immune destruction have been recently included in the select list of biological capabilities or hallmarks acquired during the development of cancer.105 Notably, the plasticity of cancer cells toward metabolic reprogramming has gained attention in the mechanisms of drug resistance.30,106,107,108 The entire molecular network that orchestrates the inherent ability of tumor cells to switch between different metabolic profiles, depending on the microenvironment stimuli, still needs to be fully elucidated.
The most important metabolic plasticity mechanisms in cancer rely on glycolysis-dependent or mitochondrial OxPhos-addiction activities.109,110 Both are considered metabolic hallmarks of cancer cells because they are involved in the direct activation of many oncogenic pathways.107,111,112,113 However, not all reprogrammed metabolic activities contribute equally to cancer progression. Yet, deregulation of mitochondrial metabolism could be considered a potential therapeutic target in tumor resistance.114 Because the PGC-1α/ERRα network is a master regulator of mitochondrial biogenesis, it could be considered a nodal regulatory step capable of controlling the entire cellular metabolism and, at least in part, in modulating this resistance-related mechanism in cancer, leading to cancer recurrence (Fig. 1).115,116
Recent studies have provided a rationale for therapeutically targeting mitochondria in certain types of cancer.11,16,117,118 Different modulators of mitochondrial activity that have been approved by The Food and Drug Administration (FDA) to treat chronic diseases, such as type 2 diabetes, cardiovascular disease, obesity, and acute myelogenous leukemia (AML), are currently being explored in clinical trials to determine potential efficacy to treat cancer. For example, anti-diabetic drugs like metformin and phenformin, the anti-obesity drug, etomoxir, or the anti-AML drug, enasidenib (IDHIFA), might provide new hope for treating responder patients whose cancer cell survival mechanism relies on mitochondrial metabolism.119 Although the clinical use of these drugs in combination with conventional chemotherapeutic drugs has led to clinical improvement outcomes, the drug-resistance mechanisms of cancer cells still remain unclear.
From metabolism to precision oncology
To focus on precision oncology and the development of effective drug design to eradicate drug-resistant cancer cells, the signaling network associated with PGC-1α/ERRα must be considered a novel targetable vulnerability in cancer cells. In examining the concept of metabolic rewiring, targeting this particular transcriptional/mitochondrial metabolic network might expose other vulnerabilities to oxidative stress in tumors (Table 1).34 Thus, targeting PGC-1α/ERRα in combination with additional metabolic vulnerabilities such as the respiratory chain defects, antioxidant programs, and TCA cycle enzymes might lead to the disruption of the nutrient sensing pathways responsible for survival of residual cells (Fig. 2b).120
A thorough analysis of the literature shows that many of the human NRs and their co-activators have been extensively studied using traditional structural analysis. However, only studies with partial protein structure and limited protein–protein interaction have been purposed, which has led to an incomplete understanding of the entire functional mechanism of the PGC-1/ERR network.21,85,91,121,122 Therefore, new insights into the overall structure of the PGC-1/ERR complex could provide insights for effectively targeting cancer resistance mechanisms and answering important questions associated with the function of this nodal transcriptional signaling network. Essential questions that could be answered include whether the full-length protein structure of both components would impose particular changes in the protein–protein interaction model that could affect the discovery and development of new therapies targeting this complex. In addition, structural insights could assist in determining whether the PGC-1α and ERRα proteins interact with specific oncogenes or tumor suppressors and thus play different roles in metabolic plasticity favoring drug resistance or decreasing cancer progression.
In this context, recent advances have been made in successfully determining high-resolution biological structures. Solving the PGC-1/ERR complex structure in its physiological state might lead to the discovery of novel mechanistic insights into the biology of the PGC-1/ERR axis, as well as the identification of novel binding partners that might have clinical relevance to treat cancers that rely on mitochondrial activity.17,123 Therefore, more studies must be pursued to predict how this complex might be exploited in basic and clinical research, leading to the elucidation of dynamic biological processes in their native states and drug discovery for personalized medicine.
Cryo-EM as a potential tool for the visualization of protein complexes
Cryo-EM is an outstanding new technology that is based on transmission electron microscopy in which a protein sample is examined in its native state at cryogenic temperatures, which can lead to successful resolution of the protein’s structure at the subatomic or atomic level. This technology has several advantages over X-ray crystallography because the protein is frozen in its native state, which can overcome technical problems with proteins that are refractory to crystallization or are just difficult to crystallize. Cryo-EM has been used successfully to resolve proteins of greater than 300 kDa to produce images with resolution as good as 2.2 Å. For example, the large β-galactosidase protein (465 kDa) has been successfully resolved by cryo-EM.124 Notably, high resolution cryo-EM images could reveal protein–protein interactions, conformational changes, and interactions between proteins and drug targets at an atomic level of precision.125 Despite the lower size limitation of protein structures that can be resolved by cryo-EM, one study showed that cryo-EM is suitable to solve the structure of small metabolic enzymes at near-atomic resolution.126 The authors presented the structure of a known therapeutic cancer target, isocitrate dehydrogenase (IDH, 93 kDa), with a resolution of 3.8 Å, which could facilitate research toward therapeutic targets and drug discovery. This is possible because crossing the 3 Å resolution level and obtaining protein structures with sizes <100 kDa might allow scientists to investigate drug-target interactions and dynamic conformational states of protein complexes (Fig. 2a).126,127 This could even benefit patients already treated with conventional chemotherapies, such as lapatinib and tamoxifen in metastatic breast cancer, epidermal growth factor inhibitors (EGFRi) in EGFR-driven lung adenocarcinoma, mitogen-activated protein kinase inhibitors (MAPKi) in melanoma, and 5-fluorouracil in Myc/PGC-1α-driven pancreatic cancer.30
The PGC-1/ERR network as a promising therapeutic target for precision oncology
Panomics data, including genomics, transcriptomics, proteomics and metabolomics, in combination with patient-matched data, are currently being used for designing treatments for personalized medicine.128 New technologies, including the clustered regularly interspaced short palindromic repeats-associated 9 system for targeted genome editing and cryo-EM, can provide mechanistic understanding of complex biological processes and are the potential tools for identifying promising therapeutic targets for precision oncology.36
The relevance of deciphering the role of the PGC-1/ERR signaling network and the therapeutic implication for precision oncology relies on several factors. First, the PGC-1/ERR transcriptional network is responsible for metabolic plasticity, which corresponds well with therapeutic resistance.42 For example, like all cancers, breast cancer is considered a heterogeneous disease and currently the approach to treat breast cancer is still based on histopathological markers that rely on tumor subtypes to evaluate and treat each patient.129,130,131 In this era of precision oncology, seeking clinically relevant biomarkers that might be exploited for therapeutic purposes is highly pertinent and could be combined with conventional therapies to generate synthetic lethality in breast cancer.132,133 Synthetic lethality is defined as any genetic mutation, chemical or drug perturbation, and environmental conditions that have a unique effect on cell viability but when exploited in combination results in cell death.134 For instance, the concept of synthetic lethality is used to treat breast cancer based on the treatment of BRCA1-deficient patients with PARP inhibitors.135 However, the means by which the PGC-1/ERR axis can be targeted to interfere with the metabolic synthetic lethality of mitochondrial enzymes still remains largely unknown.34 Second, the PGC-1/ERR complex is an important nuclear transcriptional axis that orchestrates the mitochondrial bioenergetic requirements of tumors and thereby it could be therapeutically exploited in metabolic-addictive cancers as a new metabolic vulnerability.136 Third, preventing or bypassing drug resistance is arguably the most important medical need in cancer research.114 Cleary, the identification of biomarker-defined patient populations that will most likely respond to specific drugs is critical.114 Thus, silencing the PGC-1/ERR axis in drug-resistant cancers with a high level of OxPhos might culminate in specific elimination of these cells. Finally, the attempt to access the native and physiological state of PGC-1/ERR axis will allow us to a greater understanding of the PGC-1/ERR axis in cancer biology. Moreover, the potential discovery of important oncogenes or tumor suppressors that interact with this complex will lead to the ability to fully access the mechanistic basis of how these NRs and co-activators orchestrate metabolic plasticity toward drug-resistance in cancer treatment.
Concluding remarks
Despite the progress that has been made in using structural biology for potential drug discovery to increase patient outcomes underlying precision oncology, new therapies that effectively eradicate drug-resistant cancer cells are an immediate clinical necessity. The PGC-1/ERR network holds promise as a therapeutic target for precision medicine, because this transcriptional axis orchestrates the expression of several genes involved in mitochondrial biogenesis and cell metabolism. Bosc et al. have suggested that the resistance mechanism in cancer might be associated with a shift toward an increased OxPhos status that should be considered a distinctive characteristic of drug resistance.30 The master PGC-1/ERR axis that controls mitochondrial OxPhos activity should be considered as a new pathway that drives resistance in tumor progression. The metabolic vulnerability concept that explores potential mitochondrial targets to treat cancer, rather than conventional chemotherapy, extends the concept that the up-stream mitochondrial biogenesis PGC-1/ERR network must be included as a novel targetable metabolic vulnerability (Fig. 2b). Furthermore, a native-state high-resolution structure of this nuclear complex is urgently needed.89,137,138,139 Overall, further studies are needed to determine the role of PGC-1/ERR network as a key metabolic vulnerability associated with cancer cell progression by using cryo-EM as a promising tool for drug discovery in precision oncology. Such refinements could provide opportunities to be exploited in therapeutic resistance.
Reporting summary
Further information on experimental design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Data sharing not applicable to this article as no datasets were generated or analyzed for the current study.
References
Dasgupta, S., Lonard, D. M. & O’Malley, B. W. Nuclear receptor coactivators: master regulators of human health and disease. Annu Rev. Med. 65, 279–292 (2014).
Lonard, D. M. & O’Malley, B. W. Molecular pathways: targeting steroid receptor coactivators in cancer. Clin. Cancer Res. 22, 5403–5407 (2016).
Evans, R. M. & Mangelsdorf, D. J. Nuclear receptors, RXR, and the big bang. Cell 157, 255–266 (2014).
Baek, S. H. & Kim, K. I. Emerging roles of orphan nuclear receptors in cancer. Annu Rev. Physiol. 76, 177–195 (2014).
Sonoda, J., Pei, L. & Evans, R. M. Nuclear receptors: decoding metabolic disease. FEBS Lett. 582, 2–9 (2008).
Sever, R. & Glass, C. K. Signaling by nuclear receptors. Cold Spring Harb. Perspect. Biol. 5, a016709 (2013).
Schreiber, S. N., Knutti, D., Brogli, K., Uhlmann, T. & Kralli, A. The transcriptional coactivator PGC-1 regulates the expression and activity of the orphan nuclear receptor estrogen-related receptor alpha (ERRalpha). J. Biol. Chem. 278, 9013–9018 (2003).
Fan, W. et al. ERRγ promotes angiogenesis, mitochondrial biogenesis, and oxidative remodeling in PGC1α/β-deficient muscle. Cell Rep. 22, 2521–2529 (2018).
Duchen, M. R. Mitochondria in health and disease: perspectives on a new mitochondrial biology. Mol. Asp. Med. 25, 365–451 (2004).
Gravel, S. P. Deciphering the dichotomous effects of PGC-1α on tumorigenesis and metastasis. Front. Oncol. 8, 75 (2018).
LeBleu, V. S. et al. PGC-1α mediates mitochondrial biogenesis and oxidative phosphorylation in cancer cells to promote metastasis. Nat. Cell Biol. 16, 1001–1015 (2014).
Tan, Z. et al. The role of PGC1α in cancer metabolism and its therapeutic implications. Mol. Cancer Ther. 15, 774–782 (2016).
Torrano, V. et al. The metabolic co-regulator PGC1α suppresses prostate cancer metastasis. Nat. Cell Biol. 18, 645–656 (2016).
Deblois, G., St-Pierre, J. & Giguere, V. The PGC-1/ERR signaling axis in cancer. Oncogene 32, 3483–3490 (2013).
Yun, S. H., Han, S. H. & Park, J. I. Peroxisome proliferator-activated receptor. PPAR Res. 2018, 6727421 (2018).
Vazquez, F. et al. PGC1alpha expression defines a subset of human melanoma tumors with increased mitochondrial capacity and resistance to oxidative stress. Cancer Cell 23, 287–301 (2013).
Luo, C. et al. ERRα maintains mitochondrial oxidative metabolism and constitutes an actionable target in PGC1α-elevated melanomas. Mol. Cancer Res. 15, 1366–1375 (2017).
Wallace, M. & Metallo, C. M. PGC1α drives a metabolic block on prostate cancer progression. Nat. Cell Biol. 18, 589–590 (2016).
Audet-Walsh, É. Androgen-dependent repression of ERRγ reprograms metabolism in prostate cancer. Cancer Res. 77, 378–389 (2017).
Deblois, G. & Giguere, V. Oestrogen-related receptors in breast cancer: control of cellular metabolism and beyond. Nat. Rev. Cancer 13, 27–36 (2013).
Devarakonda, S. et al. Disorder-to-order transition underlies the structural basis for the assembly of a transcriptionally active PGC-1alpha/ERRgamma complex. Proc. Natl Acad. Sci. USA 108, 18678–18683 (2011).
Luo, C., Widlund, H. R. & Puigserver, P. PGC-1 coactivators: shepherding the mitochondrial biogenesis of tumors. Trends Cancer 2, 619–631 (2016).
Tam, I. S. & Giguère, V. There and back again: the journey of the estrogen-related receptors in the cancer realm. J. Steroid Biochem Mol. Biol. 157, 13–19 (2016).
Mastropasqua, F., Girolimetti, G. & Shoshan, M. PGC1α: friend or foe in cancer? Genes 9, 48 (2018).
Chang, C. Y. & McDonnell, D. P. Molecular pathways: the metabolic regulator estrogen-related receptor α as a therapeutic target in cancer. Clin. Cancer Res. 18, 6089–6095 (2012).
Martin-Martin, N., Carracedo, A. & Torrano, V. Metabolism and transcription in cancer: merging two classic tales. Front. Cell Dev. Biol. 5, 119 (2017).
Deblois, G. et al. ERRα mediates metabolic adaptations driving lapatinib resistance in breast cancer. Nat. Commun. 7, 12156 (2016).
McGuirk, S. et al. PGC-1α supports glutamine metabolism in breast cancer. Cancer Metab. 1, 22 (2013).
Park, S. et al. ERRα-regulated lactate metabolism contributes to resistance to targeted therapies in breast cancer. Cell Rep. 15, 323–335 (2016).
Bosc, C., Selak, M. A. & Sarry, J. E. Resistance is futile: targeting mitochondrial energetics and metabolism to overcome drug resistance in cancer treatment. Cell Metab. 26, 705–707 (2017).
Audet-Walsh, É. et al. The PGC-1α/ERRα axis represses one-carbon metabolism and promotes sensitivity to anti-folate therapy in breast cancer. Cell Rep. 14, 920–931 (2016).
Gentric, G., Mieulet, V. & Mechta-Grigoriou, F. Heterogeneity in cancer metabolism: new concepts in an old field. Antioxid. Redox Signal. 26, 462–485 (2017).
Valcarcel-Jimenez, L., Gaude, E., Torrano, V., Frezza, C. & Carracedo, A. Mitochondrial metabolism: yin and yang for tumor progression. Trends Endocrinol. Metab. 28, 748–757 (2017).
Zecchini, V. & Frezza, C. Metabolic synthetic lethality in cancer therapy. Biochim Biophys. Acta 1858, 723–731 (2017).
Yi, P. et al. Structure of a biologically active estrogen receptor-coactivator complex on DNA. Mol. Cell 57, 1047–1058 (2015).
Shin, S. H., Bode, A. M. & Dong, Z. G. Addressing the challenges of applying precision oncology. Npj Precision. NPJ Precision Oncol. 1, 10 (2017).
Zong, W. X., Rabinowitz, J. D. & White, E. Mitochondria and cancer. Mol. Cell 61, 667–676 (2016).
Puigserver, P. et al. A cold-inducible coactivator of nuclear receptors linked to adaptive thermogenesis. Cell 92, 829–839 (1998).
Andersson, U. & Scarpulla, R. C. Pgc-1-related coactivator, a novel, serum-inducible coactivator of nuclear respiratory factor 1-dependent transcription in mammalian cells. Mol. Cell Biol. 21, 3738–3749 (2001).
Kressler, D., Schreiber, S. N., Knutti, D. & Kralli, A. The PGC-1-related protein PERC is a selective coactivator of estrogen receptor alpha. J. Biol. Chem. 277, 13918–13925 (2002).
Martínez-Redondo, V. et al. Peroxisome proliferator-activated receptor γ coactivator-1 α isoforms selectively regulate multiple splicing events on target genes. J. Biol. Chem. 291, 15169–15184 (2016).
Zhang, G. et al. Targeting mitochondrial biogenesis to overcome drug resistance to MAPK inhibitors. J. Clin. Invest 126, 1834–1856 (2016).
Knutti, D., Kaul, A. & Kralli, A. A tissue-specific coactivator of steroid receptors, identified in a functional genetic screen. Mol. Cell Biol. 20, 2411–2422 (2000).
Villena, J. A. New insights into PGC-1 coactivators: redefining their role in the regulation of mitochondrial function and beyond. FEBS J. 282, 647–672 (2015).
Monsalve, M. et al. Direct coupling of transcription and mRNA processing through the thermogenic coactivator PGC-1. Mol. Cell 6, 307–316 (2000).
Aquilano, K. et al. Peroxisome proliferator-activated receptor gamma co-activator 1alpha (PGC-1alpha) and sirtuin 1 (SIRT1) reside in mitochondria: possible direct function in mitochondrial biogenesis. J. Biol. Chem. 285, 21590–21599 (2010).
Chang, J. S. et al. Regulation of NT-PGC-1alpha subcellular localization and function by protein kinase A-dependent modulation of nuclear export by CRM1. J. Biol. Chem. 285, 18039–18050 (2010).
Lin, J. et al. Transcriptional co-activator PGC-1 alpha drives the formation of slow-twitch muscle fibres. Nature 418, 797–801 (2002).
Puigserver, P. et al. Activation of PPARgamma coactivator-1 through transcription factor docking. Science 286, 1368–1371 (1999).
Wallberg, A. E., Yamamura, S., Malik, S., Spiegelman, B. M. & Roeder, R. G. Coordination of p300-mediated chromatin remodeling and TRAP/mediator function through coactivator PGC-1alpha. Mol. Cell 12, 1137–1149 (2003).
Lai, L. et al. Transcriptional coactivators PGC-1alpha and PGC-lbeta control overlapping programs required for perinatal maturation of the heart. Genes Dev. 22, 1948–1961 (2008).
Zechner, C. et al. Total skeletal muscle PGC-1 deficiency uncouples mitochondrial derangements from fiber type determination and insulin sensitivity. Cell Metab. 12, 633–642 (2010).
He, X. et al. Peri-implantation lethality in mice lacking the PGC-1-related coactivator protein. Dev. Dyn. 241, 975–983 (2012).
Gleyzer, N. & Scarpulla, R. C. PGC-1-related coactivator (PRC), a sensor of metabolic stress, orchestrates a redox-sensitive program of inflammatory gene expression. J. Biol. Chem. 286, 39715–39725 (2011).
Gleyzer, N. & Scarpulla, R. C. Activation of a PGC-1-related coactivator (PRC)-dependent inflammatory stress program linked to apoptosis and premature senescence. J. Biol. Chem. 288, 8004–8015 (2013).
Handschin, C. & Spiegelman, B. M. Peroxisome proliferator-activated receptor gamma coactivator 1 coactivators, energy homeostasis, and metabolism. Endocr. Rev. 27, 728–735 (2006).
Lin, J., Handschin, C. & Spiegelman, B. M. Metabolic control through the PGC-1 family of transcription coactivators. Cell Metab. 1, 361–370 (2005).
Puigserver, P. Tissue-specific regulation of metabolic pathways through the transcriptional coactivator PGC1-alpha. Int J. Obes. 29, S5–S9 (2005).
Fabregat-Andrés, Ó. Induction of PGC-1α expression can be detected in blood samples of patients with ST-segment elevation acute myocardial infarction. PLoS ONE 6, e26913 (2011).
Huss, J. M., Kopp, R. P. & Kelly, D. P. Peroxisome proliferator-activated receptor coactivator-1alpha (PGC-1alpha) coactivates the cardiac-enriched nuclear receptors estrogen-related receptor-alpha and -gamma. Identification of novel leucine-rich interaction motif within PGC-1alpha. J. Biol. Chem. 277, 40265–40274 (2002).
Kamei, Y. et al. PPARgamma coactivator 1beta/ERR ligand 1 is an ERR protein ligand, whose expression induces a high-energy expenditure and antagonizes obesity. Proc. Natl Acad. Sci. USA 100, 12378–12383 (2003).
Sen, N., Satija, Y. K. & Das, S. PGC-1α, a key modulator of p53, promotes cell survival upon metabolic stress. Mol. Cell 44, 621–634 (2011).
Fernandez-Marcos, P. J. & Auwerx, J. Regulation of PGC-1α, a nodal regulator of mitochondrial biogenesis. Am. J. Clin. Nutr. 93, 884S–890S (2011).
Martinez-Pastor, B., Cosentino, C. & Mostoslavsky, R. A tale of metabolites: the cross-talk between chromatin and energy metabolism. Cancer Discov. 3, 497–501 (2013).
Wong, C. C., Qian, Y. & Yu, J. Interplay between epigenetics and metabolism in oncogenesis: mechanisms and therapeutic approaches. Oncogene 36, 3359–3374 (2017).
Ardito, F., Giuliani, M., Perrone, D., Troiano, G. & Lo Muzio, L. The crucial role of protein phosphorylation in cell signaling and its use as targeted therapy (Review). Int J. Mol. Med. 40, 271–280 (2017).
Fan, J., Krautkramer, K. A., Feldman, J. L. & Denu, J. M. Metabolic regulation of histone post-translational modifications. ACS Chem. Biol. 10, 95–108 (2015).
Arany, Z. et al. HIF-independent regulation of VEGF and angiogenesis by the transcriptional coactivator PGC-1alpha. Nature 451, 1008–1012 (2008).
Li, X., Monks, B., Ge, Q. & Birnbaum, M. J. Akt/PKB regulates hepatic metabolism by directly inhibiting PGC-1alpha transcription coactivator. Nature 447, 1012–1016 (2007).
Sancho, P. et al. MYC/PGC-1α balance determines the metabolic phenotype and plasticity of pancreatic cancer stem cells. Cell Metab. 22, 590–605 (2015).
Cioce, M. & Blandino, G. PGC1α confers specificity-metabolic stress and p53-dependent transcription. Mol. Cell 44, 515–516 (2011).
Zhang, H. et al. HIF-1 inhibits mitochondrial biogenesis and cellular respiration in VHL-deficient renal cell carcinoma by repression of C-MYC activity. Cancer Cell 11, 407–420 (2007).
Zeller, K. I. et al. Global mapping of c-Myc binding sites and target gene networks in human B cells. Proc. Natl Acad. Sci. USA 103, 17834–17839 (2006).
LaGory, E. L. et al. Suppression of PGC-1α is critical for reprogramming oxidative metabolism in renal cell carcinoma. Cell Rep. 12, 116–127 (2015).
Chang, C. Y. et al. The metabolic regulator ERRα, a downstream target of HER2/IGF-1R, as a therapeutic target in breast cancer. Cancer Cell 20, 500–510 (2011).
Deblois, G. et al. Transcriptional control of the ERBB2 amplicon by ERRalpha and PGC-1beta promotes mammary gland tumorigenesis. Cancer Res. 70, 10277–10287 (2010).
Yao, Z. et al. PGC-1β mediates adaptive chemoresistance associated with mitochondrial DNA mutations. Oncogene 32, 2592–2600 (2013).
Gleyzer, N. & Scarpulla, R. C. Concerted Action of PGC-1-related Coactivator (PRC) and c-MYC in the stress response to mitochondrial dysfunction. J. Biol. Chem. 291, 25529–25541 (2016).
Giguere, V. To ERR in the estrogen pathway. Trends Endocrinol. Metab. 13, 220–225 (2002).
Giguere, V., Tremblay, A. & Tremblay, G. B. Estrogen receptor beta: re-evaluation of estrogen and antiestrogen signaling. Steroids 63, 335–339 (1998).
Misawa, A. & Inoue, S. Estrogen-related receptors in breast cancer and prostate cancer. Front Endocrinol. 6, 83 (2015).
Huss, J. M., Garbacz, W. G. & Xie, W. Constitutive activities of estrogen-related receptors: transcriptional regulation of metabolism by the ERR pathways in health and disease. Biochim Biophys. Acta 1852, 1912–1927 (2015).
Misra, J., Kim, D. K. & Choi, H. S. ERRγ: a junior orphan with a senior role in metabolism. Trends Endocrinol. Metab. 28, 261–272 (2017).
Wilson, B. J., Tremblay, A. M., Deblois, G., Sylvain-Drolet, G. & Giguere, V. An acetylation switch modulates the transcriptional activity of estrogen-related receptor alpha. Mol. Endocrinol. 24, 1349–1358 (2010).
Kallen, J. et al. Crystal structure of human estrogen-related receptor alpha in complex with a synthetic inverse agonist reveals its novel molecular mechanism. J. Biol. Chem. 282, 23231–23239 (2007).
Abad, M. C. et al. Structural determination of estrogen-related receptor gamma in the presence of phenol derivative compounds. J. Steroid Biochem Mol. Biol. 108, 44–54 (2008).
Patch, R. J. et al. Identification of diaryl ether-based ligands for estrogen-related receptor α as potential antidiabetic agents. J. Med. Chem. 54, 788–808 (2011).
Riggins, R. B., Mazzotta, M. M., Maniya, O. Z. & Clarke, R. Orphan nuclear receptors in breast cancer pathogenesis and therapeutic response. Endocr. Relat. Cancer 17, R213–R231 (2010).
Willy, P. J. et al. Regulation of PPARgamma coactivator 1alpha (PGC-1alpha) signaling by an estrogen-related receptor alpha (ERRalpha) ligand. Proc. Natl Acad. Sci. USA 101, 8912–8917 (2004).
Eskiocak, B., Ali, A. & White, M. A. The estrogen-related receptor α inverse agonist XCT 790 is a nanomolar mitochondrial uncoupler. Biochemistry 53, 4839–4846 (2014).
Kallen, J. et al. Evidence for ligand-independent transcriptional activation of the human estrogen-related receptor alpha (ERRalpha): crystal structure of ERRalpha ligand binding domain in complex with peroxisome proliferator-activated receptor coactivator-1alpha. J. Biol. Chem. 279, 49330–49337 (2004).
Manna, S. et al. ERRα is a marker of tamoxifen response and survival in triple-negative breast cancer. Clin. Cancer Res. 22, 1421–1431 (2016).
Suzuki, T. et al. Estrogen-related receptor alpha in human breast carcinoma as a potent prognostic factor. Cancer Res. 64, 4670–4676 (2004).
Berman, A. Y. et al. ERRα regulates the growth of triple-negative breast cancer cells via S6K1-dependent mechanism. Signal. Transduct. Target Ther. 2, e17035 (2017).
Thewes, V. et al. Reprogramming of the ERRα and ERα target gene landscape triggers tamoxifen resistance in breast cancer. Cancer Res. 75, 720–731 (2015).
Kang, M. H. et al. Estrogen-related receptor gamma functions as a tumor suppressor in gastric cancer. Nat. Commun. 9, 1920 (2018).
Yu, S., Wang, X., Ng, C. F., Chen, S. & Chan, F. L. ERRgamma suppresses cell proliferation and tumor growth of androgen-sensitive and androgen-insensitive prostate cancer cells and its implication as a therapeutic target for prostate cancer. Cancer Res. 67, 4904–4914 (2007).
Kim, J. H. et al. Estrogen-related receptor γ is upregulated in liver cancer and its inhibition suppresses liver cancer cell proliferation via induction of p21 and p27. Exp. Mol. Med. 48, e213 (2016).
Heckler, M. M., Thakor, H., Schafer, C. C. & Riggins, R. B. ERK/MAPK regulates ERRγ expression, transcriptional activity and receptor-mediated tamoxifen resistance in ER + breast cancer. FEBS J. 281, 2431–2442 (2014).
Madhavan, S., Gusev, Y., Singh, S. & Riggins, R. B. ERRγ target genes are poor prognostic factors in Tamoxifen-treated breast cancer. J. Exp. Clin. Cancer Res. 34, 45 (2015).
Ariazi, E. A., Clark, G. M. & Mertz, J. E. Estrogen-related receptor alpha and estrogen-related receptor gamma associate with unfavorable and favorable biomarkers, respectively, in human breast cancer. Cancer Res. 62, 6510–6518 (2002).
Tiraby, C., Hazen, B. C., Gantner, M. L. & Kralli, A. Estrogen-related receptor gamma promotes mesenchymal-to-epithelial transition and suppresses breast tumor growth. Cancer Res. 71, 2518–2528 (2011).
Audet-Walsh, É. & Giguére, V. The multiple universes of estrogen-related receptor α and γ in metabolic control and related diseases. Acta Pharmacol. Sin. 36, 51–61 (2015).
Yu, S. et al. Orphan nuclear receptor estrogen-related receptor-beta suppresses in vitro and in vivo growth of prostate cancer cells via p21(WAF1/CIP1) induction and as a potential therapeutic target in prostate cancer. Oncogene 27, 3313–3328 (2008).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Sancho, P., Barneda, D. & Heeschen, C. Hallmarks of cancer stem cell metabolism. Br. J. Cancer 114, 1305–1312 (2016).
Ward, P. S. & Thompson, C. B. Metabolic reprogramming: a cancer hallmark even warburg did not anticipate. Cancer Cell 21, 297–308 (2012).
Zhao, Y., Butler, E. B. & Tan, M. Targeting cellular metabolism to improve cancer therapeutics. Cell Death Dis. 4, e532 (2013).
DeBerardinis, R. J. & Chandel, N. S. Fundamentals of cancer metabolism. Sci. Adv. 2, e1600200 (2016).
Vander Heiden, M. G. & DeBerardinis, R. J. Understanding the intersections between metabolism and cancer biology. Cell 168, 657–669 (2017).
Denko, N. C. Hypoxia, HIF1 and glucose metabolism in the solid tumour. Nat. Rev. Cancer 8, 705–713 (2008).
Nagarajan, A., Malvi, P. & Wajapeyee, N. Oncogene-directed alterations in cancer cell metabolism. Trends Cancer 2, 365–377 (2016).
Viale, A. et al. Oncogene ablation-resistant pancreatic cancer cells depend on mitochondrial function. Nature 514, 628–632 (2014).
Morandi, A. & Indraccolo, S. Linking metabolic reprogramming to therapy resistance in cancer. Biochim Biophys. Acta 1868, 1–6 (2017).
Kuntz, E. M. et al. Targeting mitochondrial oxidative phosphorylation eradicates therapy-resistant chronic myeloid leukemia stem cells. Nat. Med. 23, 1234–1240 (2017).
Lee, K. M. et al. MYC and MCL1 cooperatively promote chemotherapy-resistant breast cancer stem cells via regulation of mitochondrial oxidative phosphorylation. Cell Metab. 26, 633–647 e637 (2017).
Hensley, C. T. et al. Metabolic heterogeneity in human lung tumors. Cell 164, 681–694 (2016).
Porporato, P. E., Filigheddu, N., Pedro, J. M. B., Kroemer, G. & Galluzzi, L. Mitochondrial metabolism and cancer. Cell Res. 28, 265–280 (2018).
Trousil, S. et al. Phenformin enhances the efficacy of ERK inhibition in NF1-mutant melanoma. J. Invest. Dermatol. 137, 1135–1143 (2017).
Efeyan, A., Comb, W. C. & Sabatini, D. M. Nutrient-sensing mechanisms and pathways. Nature 517, 302–310 (2015).
Takacs, M. et al. The asymmetric binding of PGC-1α to the ERRα and ERRγ nuclear receptor homodimers involves a similar recognition mechanism. PLoS ONE 8, e67810 (2013).
Wang, L. et al. X-ray crystal structures of the estrogen-related receptor-gamma ligand binding domain in three functional states reveal the molecular basis of small molecule regulation. J. Biol. Chem. 281, 37773–37781 (2006).
Weinberg, S. E. & Chandel, N. S. Targeting mitochondria metabolism for cancer therapy. Nat. Chem. Biol. 11, 9–15 (2015).
Bartesaghi, A. et al. 2.2A resolution cryo-EM structure of beta-galactosidase in complex with a cell-permeant inhibitor. Science 348, 1147–1151 (2015).
Kuhlbrandt, W. Cryo-EM enters a new era. Elife 3, e03678 (2014).
Merk, A. et al. Breaking Cryo-EM resolution barriers to facilitate drug discovery. Cell 165, 1698–1707 (2016).
Banerjee, S. et al. 2.3 Å resolution cryo-EM structure of human p97 and mechanism of allosteric inhibition. Science 351, 871–875 (2016).
Sandhu, C., Qureshi, A. & Emili, A. Panomics for precision medicine. Trends Mol. Med. 24, 85–101 (2018).
Polyak, K. Heterogeneity in breast cancer. J. Clin. Invest. 121, 3786–3788 (2011).
Yersal, O. & Barutca, S. Biological subtypes of breast cancer: prognostic and therapeutic implications. World J. Clin. Oncol. 5, 412–424 (2014).
Koren, S. & Bentires-Alj, M. Breast tumor heterogeneity: source of fitness, hurdle for therapy. Mol. Cell 60, 537–546 (2015).
Rehman, F. L., Lord, C. J. & Ashworth, A. Synthetic lethal approaches to breast cancer therapy. Nat. Rev. Clin. Oncol. 7, 718–724 (2010).
La Thangue, N. B. & Kerr, D. J. Predictive biomarkers: a paradigm shift towards personalized cancer medicine. Nat. Rev. Clin. Oncol. 8, 587–596 (2011).
Lord, C. J. & Ashworth, A. PARP inhibitors: synthetic lethality in the clinic. Science 355, 1152–1158 (2017).
Farmer, H. et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434, 917–921 (2005).
Wolpaw, A. J. & Dang, C. V. Exploiting metabolic vulnerabilities of cancer with precision and accuracy. Trends Cell Biol. 28, 201–212 (2018).
Busch, B. B. et al. Identification of a selective inverse agonist for the orphan nuclear receptor estrogen-related receptor alpha. J. Med. Chem. 47, 5593–5596 (2004).
Lanvin, O., Bianco, S., Kersual, N., Chalbos, D. & Vanacker, J. M. Potentiation of ICI182,780 (Fulvestrant)-induced estrogen receptor-alpha degradation by the estrogen receptor-related receptor-alpha inverse agonist XCT790. J. Biol. Chem. 282, 28328–28334 (2007).
Sharabi, K. et al. Selective chemical inhibition of PGC-1α gluconeogenic activity ameliorates type 2 diabetes. Cell 169, 148–160.e115 (2017).
Giguère, V. Transcriptional control of energy homeostasis by the estrogen-related receptors. Endocr. Rev. 29, 677–696 (2008).
Simons, A. L., Ahmad, I. M., Mattson, D. M., Dornfeld, K. J. & Spitz, D. R. 2-Deoxy-D-glucose combined with cisplatin enhances cytotoxicity via metabolic oxidative stress in human head and neck cancer cells. Cancer Res. 67, 3364–3370 (2007).
Michalek, R. D. et al. Estrogen-related receptor-α is a metabolic regulator of effector T-cell activation and differentiation. Proc. Natl Acad. Sci. USA 108, 18348–18353 (2011).
Yu, Y. et al. Selective active site inhibitors of human lactate dehydrogenases A4, B4, and C4. Biochem. Pharmacol. 62, 81–89 (2001).
Cai, Q., Lin, T., Kamarajugadda, S. & Lu, J. Regulation of glycolysis and the Warburg effect by estrogen-related receptors. Oncogene 32, 2079–2086 (2013).
Porporato, P. E., Dhup, S., Dadhich, R. K., Copetti, T. & Sonveaux, P. Anticancer targets in the glycolytic metabolism of tumors: a comprehensive review. Front. Pharmacol. 2, 49 (2011).
Pereira da Silva, A. P. et al. Inhibition of energy-producing pathways of HepG2 cells by 3-bromopyruvate. Biochem. J. 417, 717–726 (2009).
Casaburi, I. et al. Estrogen related receptor α (ERRα) a promising target for the therapy of adrenocortical carcinoma (ACC). Oncotarget 6, 25135–25148 (2015).
Mullard, A. FDA approves first-in-class cancer metabolism drug. Nat. Rev. Drug Discov. 16, 593 (2017).
Brown, E. L. et al. Estrogen-related receptors mediate the adaptive response of brown adipose tissue to adrenergic stimulation. iScience 2, 221–237 (2018).
Janzer, A. et al. Metformin and phenformin deplete tricarboxylic acid cycle and glycolytic intermediates during cell transformation and NTPs in cancer stem cells. Proc. Natl Acad. Sci. USA 111, 10574–10579 (2014).
Hao, Y. et al. Oncogenic PIK3CA mutations reprogram glutamine metabolism in colorectal cancer. Nat. Commun. 7, 11971 (2016).
Reaven, G. M., Chang, H. & Hoffman, B. B. Additive hypoglycemic effects of drugs that modify free-fatty acid metabolism by different mechanisms in rats with streptozocin-induced diabetes. Diabetes 37, 28–32 (1988).
Acknowledgements
This work was supported by The Hormel Foundation and National Institutes of Health grants CA166011, CA187027, and CA196639, and CAPES 9281-13-4. We thank Dr. Ferdinando Chiaradonna for important discussion regarding this manuscript.
Author information
Authors and Affiliations
Contributions
A.M.B. and Z.D. supervised the conception of the work and participated in discussion and revision of the manuscript. H.D.V. wrote the manuscript and produced figures. H.D.V., A.M.B., and Z.D. revised the manuscript. All authors approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
De Vitto, H., Bode, A.M. & Dong, Z. The PGC-1/ERR network and its role in precision oncology. npj Precis. Onc. 3, 9 (2019). https://doi.org/10.1038/s41698-019-0081-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-019-0081-6
This article is cited by
-
The lncRNA lnc_AABR07044470.1 promotes the mitochondrial-damaged inflammatory response to neuronal injury via miR-214-3p/PERM1 axis in acute ischemic stroke
Molecular Biology Reports (2024)
-
Genomic and epigenomic integrative subtypes of renal cell carcinoma in a Japanese cohort
Nature Communications (2023)
-
Transcriptional co-activators: emerging roles in signaling pathways and potential therapeutic targets for diseases
Signal Transduction and Targeted Therapy (2023)
-
Circulating proteins as predictive and prognostic biomarkers in breast cancer
Clinical Proteomics (2022)
-
Regulatory chromatin rewiring promotes metabolic switching during adaptation to oncogenic receptor tyrosine kinase inhibition
Oncogene (2022)