Abstract
The aim of the present study was to determine if colostrum and the equipment for harvesting and feeding colostrum are sources of fecal ESBL/AmpC-producing Escherichia coli (ESBL/AmpC-E. coli) in calves. Therefore, 15 male calves fed with pooled colostrum on a dairy farm and held individually in an experimental barn, the colostrum pool and the equipment for harvesting and feeding colostrum were sampled and analyzed for the occurrence of ESBL/AmpC-E. coli. The ESBL-AmpC-E. coli suspicious isolates were subjected to whole-genome sequence analysis. Forty-three of 45 fecal samples were tested positive for ESBL/AmpC-E. coli. In the colostrum sample and in the milking pot, we also found ESBL/AmpC-E. coli. All 45 E. coli isolates were ESBL-producers, mainly commensal sequence type (ST) 10, but also human-extraintestinal pathogenic E. coli ST131 and ST117 were found. The clonal identity of six fecal isolates with the ESBL-E. coli isolate from the colostrum and of five fecal isolates with the strain from the milking pot demonstrates that the hygiene of colostrum or the colostrum equipment can play a significant role in the spread of ESBL-E. coli. Effective sanitation procedures for colostrum harvesting and feeding equipment are crucial to reduce the ESBL-E. coli shedding of neonatal dairy calves.
Similar content being viewed by others
Introduction
In 2015, the World Health Organization (WHO) designated antimicrobial resistance (AMR) as a major threat to global health, food security, and development. Extended-spectrum β-lactamase (ESBL)-producing bacteria are resistant to broad-spectrum β-lactams, such as 3rd generation cephalosporins. Recognizing the severity of this resistant phenotype in terms of increasing mortality rates in humans and animals, ESBL-producing Enterobacterales have been included in the WHO global priority list of antibiotic-resistant bacteria, which guides research for the discovery and development of new antibiotics1,2. Apart from ESBL, resistance to 3rd generation-cephalosporins can also be mediated through AmpC β-lactamases (AmpC)3.
ESBL/AmpC-producing Enterobacterales, especially Escherichia coli (E. coli), are frequently found in livestock (e.g., dairy cattle), as well as in food products4,5,6. Although commensal E. coli strains rarely cause infections, they can, however, transfer resistance genes horizontally to pathogenic E. coli strains or other Enterobacterales that can be transmitted to humans via the food chain or environmental effluents7,8,9.
Usually, the prevalence of ESBL/AmpC-producing E. coli (ESBL/AmpC-E. coli) in dairy cattle is typically age-dependent with a higher prevalence and abundance in pre-weaning calves10. A recent study in Germany showed that 63.5% of the young calves in large dairies shed ESBL/AmpC-E. coli although most of them had never been treated with antibiotics11. With increasing age and dietary transformation from a monogastric animal to a ruminant, ESBL/AmpC-E. coli excretion decreases12. This indicates that diet is a strong contributing factor of the fecal release of antibiotic-resistant bacteria13.
There are several studies investigating risk factors for the occurrence of ESBL/AmpC-E. coli in calves11,14,15,16,17. Concerning diet, some of them could reveal that feeding of waste milk containing antibiotic residues increases ESBL/AmpC-E. coli prevalence in calves11,14,18. Another study associated the antibiotic dry-off therapy of the cows with higher fecal ESBL/AmpC-E. coli shedding in calves, suggesting an influence of antibiotic residues in colostrum on ESBL/AmpC-E. coli occurrence19. However, in addition to antibiotic contamination of colostrum or milk rations, vertical transmission of ESBL/AmpC-E. coli from the dam, horizontal transmission from other animals in the herd, or transmission from the housing environment, colostrum/feed, or feeding equipment may also play an important role11,13,16,20.
As calves are the main contributors of ESBL/AmpC-E. coli release in dairy production, understanding the early colonization of ESBL/AmpC-E. coli in calves during the milk-feeding period is of great importance. As part of a concurrent study, in which calves were fed with pooled colostrum, we could sample feces, colostrum and the equipment for harvesting and feeding colostrum to get a deeper insight in the sources for the early acquisition of ESBL/AmpC-E. coli. According to the research of Liu et al.13 and of He et al.21 detecting the same genes for AMR in colostrum and calves’ feces, we hypothesized that colostrum or the equipment may serve as the first vectors for ESBL/AmpC-E. coli in young calves.
Results
Bacteriological examination
On the first and fifth day of life, 14 out of 15 fecal samples of the calves were tested positive for ESBL/AmpC-E. coli. On day 8 all calves showed ESBL/AmpC-E. coli excretion. Furthermore, we detected ESBL/AmpC-E. coli in the colostrum sample and in the swab of the milking pot. In total, we obtained 45 ESBL-E. coli isolates (43 from feces and one from colostrum and one from equipment for harvesting colostrum).
Antimicrobial susceptibility test (AST)
The results of the AST are displayed in Table 1. All 45 ESBL/AmpC-suspicious E. coli isolates showed ESBL- and not AmpC-phenotype, i.e., were resistant against ampicillin, piperacillin, cefuroxime, cefotaxime and ceftazidime, but were less resistant against combinations of β-lactam antibiotics with inhibitors of β-lactamases (tazobactam, avibactam). Twenty-four (= 53.3%) of the isolates fulfilled the definition of multidrug-resistant (MDR, resistant to at least three antibiotic classes) organisms. Of the MDR isolates, 14 (= 31% of all isolates) showed phenotypic resistance against ciprofloxacin. All isolates were sensitive to ertapenem, imipenem, meropenem, gentamicin, eravacycline, tigecycline, and colistin.
Whole genome sequencing (WGS)
Details of the WGS are shown in Table 2. MLS typing revealed 13 different sequence types (STs), belonging to ST10, ST88, ST117, ST131, ST354, ST362, ST540, ST744, ST761, ST1122, ST1429, ST1431, and ST2325, with ST10 being dominant in the fecal swabs (21/43 = 49%). Moreover, the isolates of colostrum and milking pot belonged to ST10. The most frequent STs besides ST10 were ST88, ST117 and ST362, each occurring three times. Based on sequencing, ESBL genes (CTX-M type) were detected in all 45 isolates. Most isolates harbored blaCTX-M-1 (35/45 = 78%), whereas we could also detect blaCTX-M-15 (8/45) and blaCTX-M-27 (5/45), thus, three isolates were positive for both, blaCTX-M1 and blaCTX-M27. Other β-lactamase genes present in the samples were: blaoxa-1 (8/45), blalap-2 (9/45) and blaTEM-1 (25/45). qnrS1 gene, which is associated with quinolone resistance, was determined in 19 isolates including the ESBL-E. coli isolated from colostrum and milking pot. The most common resistance gene was mdf (41/45), which belongs to a transport protein conveying resistance to a broad spectrum of toxic substances and antibiotics. Some of the isolates additionally harbored phenicol resistance genes (floR, catA and cmlA). In addition, several aminoglycoside, tetracycline, sulfonamide and trimethoprim, and macrolide resistance genes were also found in the WGS analysis.
Clonal expansion
Phylogeny analysis of ST10 isolates based on WGS data revealed three phylogenetic clusters. The size of the core genome of the ST 10 isolates studied was 5,140,627 bp and within each cluster core genomes of the isolates differed only in 0–4 SNPs or in 0–55 SNPs, respectively (see Fig. 1). The isolate No. 2012 (swab of milking pot) belonged to one cluster, in which only the fecal isolate No. 2009 differed in a single SNP from the other five fecal isolates. All isolates of that milking pot-cluster harbored the CTX-M-15 gene. To the second cluster the isolate No. 996 (colostrum sample) was allocated differing only in 2–4 SNPs to six fecal isolates. Three additional ESBL-E. coli isolates of that cluster exhibited more varying SNPs (52–55) compared to the other seven isolates. CTX-M-1 was detected in all isolates of the colostrum-cluster. The third cluster only contained fecal isolates which differed in 0–4 SNPs from each other. In the feces-cluster CTX-M-1 gene was present. The fecal isolate No. 2005 could not be allocated to one of the clusters. The difference to all other isolates of ST10 was 187–699 SNPs. In contrast to the isolates nearby in the phylogenetic tree, this isolate was CTX-M-15 positive.
Core SNP phylogeny of ESBL-producing ST10 isolates originating from calf feces, pooled colostrum, and milking pot. Core genome SNPs were called using snippy v. 4.4.1. SNP distances were calculated and depicted in a heat diagram (SNP distance ranged from 0 to 760 sites). The phylogenetic tree is based on a core SNP alignment and visualized in iTOL. Additionally, presence of CTX-M-1 and CTX-M-15 were depicted in the phylogenetic tree.
Biofilm formation
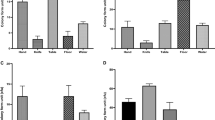
To test the ability to form biofilms, one representative was selected from each of the three different clusters we identified in the clonal expansion analysis: No. 2012 (milking pot isolate), No. 996 (colostrum isolate), and No. 461 (fecal isolate). The test showed that the ability to form specific biofilms is high in the isolates from milking pot and colostrum and low in the fecal isolate from the feces-cluster (see Fig. 2).
Discussion
Today, livestock production is known to be a significant source of AMR. In the past decades, the occurrence of ESBL-carrying pathogens increased worldwide22. ESBL/AmpC-E. coli are less prevalent in dairy cattle than other livestock species, with the exception of suckling calves, which have a high ESBL/AmpC-E. coli prevalence, e.g., above 50% in Germany11,17.
Due to their immature immune system, newborn calves depend on passive immunization with maternal antibodies from the colostrum23. As it is common practice in conventional dairy farms to separate calf and dam right after birth, the colostrum is usually administered to the calf via bottle, bucket, or drench. A major advantage of this method compared to natural intake is the controllability of the amount and timing of colostrum intake.
Already on the first day of life, 14 out of 15 calves in our study were tested positive for ESBL-E. coli, although they had neither spatial nor temporal contact with each other and did not get any antibiotics. Before the next calf could enter the experimental barn, it was cleaned, disinfected and unoccupied for at least 1 week to prevent nosocomial transmission. Therefore, horizontal transmission between the calves and from the equipment of the experimental barn seemed unlikely. Moreover, the transport vehicle has also been cleaned and disinfected, so that transmission of ESBL-E. coli during transportation is also unlikely. Moreover, according to previous research, there is little chance that transmission from the dams to the calves is the main reason for ESBL-E. coli occurrence in the calves as the prevalence of the corresponding dams is quite low11 and the ESBL-E. coli strains of calves differ from that of older animals10. Therefore, a sample of the colostrum pool and swabs of the equipment for harvesting and administering colostrum were analyzed for the occurrence of ESBL/AmpC-E. coli. In colostrum and in the milking pot for harvesting colostrum, we could detect ESBL-E. coli belonging to commensal ST10.
WGS revealed that six of 43 fecal isolates were nearly identical to the colostrum strain. Since the number of core genome SNPs was below 17, these isolates fulfil the clone definition of Ludden et al.24 who developed this SNP cut-off to demonstrate transmission between patients. An even higher degree of core genomic identity was also demonstrated between five fecal isolates and the milking pot strain, demonstrating direct transmission of ESBL E. coli via colostrum or equipment in a total of 11 of the 43 ESBL E. coli isolates (25.6%). Our findings are underlined by two recent studies, which assumed that the feeding of colostrum contaminated with ESBL-E. coli can be responsible for intestinal colonization of calves with ESBL-E. coli, as the same bacterial resistome (i.e., all AMR genes and their precursors) was found in colostrum and in feces of neonatal calves13,21. In addition to the suggestion that antibiotic residues in colostrum may lead to increased occurrence of ESBL-E. coli in suckling calves19, our results prove that colostrum serves as an early vector for resistant bacteria in calves.
It remains unknown how exactly the ESBL-E. coli contamination of the colostrum occurred. However, based on the detection of an ESBL-E. coli in the milking pot, we assume that the presence of ESBL E. coli in the colostrum is due to poor hygiene of the milking equipment used to collect colostrum, particularly since both strains found in colostrum and milking pot have a high ability to form specific biofilms. The ability of biofilm formation allows ESBL E. coli to withstand standard cleaning procedures8 and to persist in the milking equipment. In this way, washing out bacteria results in contamination of colostrum and colonization of calves. According to two German studies, hygiene management of the calf’s feeding equipment is associated with the occurrence of ESBL-E. coli in dairy farms. Heinemann and colleagues found ESBL-E. coli in the inner surface of nipples of feeding buckets and concluded that sanitation measures in dairy farms are inadequate, maybe leading to ESBL-E. coli infection of the calves20. In addition, the cleaning procedure of feeding buckets was associated with the ESBL-E. coli prevalence of dairy calves11.
Acidification or pasteurization of colostrum, which are common to optimize hygiene in milk rations for calves, are possible methods to sanitize colostrum25,26. However, if the equipment for colostrum administration, particularly the nipple, is contaminated with biofilm forming ESBL-E. coli, transmission of resistant bacteria is still possible. According to the data of Heinemann et al. 20 nipples for calves are regularly contaminated with ESBL-E. coli and other bacteria.
In addition to ST10, we found other STs of ESBL-producing E. coli, including ST131, a globally distributed clonal lineage known to cause severe extraintestinal infections in humans and animals22,27. This single lineage is mainly responsible for the increase in urinary tract and bloodstream infections with ESBL-E. coli worldwide. Strains of ST131 usually carry blaCTX-M-15 on a plasmid26,28. Deviating from that, our isolate of ST131 is blaCTX-M-1-positive. ESBL-E. coli carrying blaCTX-M1 are primarily isolated from livestock in Europe29,30. CTX-M15 is predominant in ST13129,31, therefore, horizontal gene transfer of a blaCTX-M-1 encoding plasmid seems certainly probable for our isolate. In studies with companion animals, ESBL-E. coli producing CTX-M-15 and other variants (e.g., CTX-M-14, CTX-M-55), which were frequently found in humans, could be detected, indicating a direct transfer of viable bacteria between companion animals and humans30,32. Transmission of ESBL-E. coli from animals to humans is often postulated. Regarding livestock production, contaminated slurry and waste water from abattoirs and animal rendering plants can harbor a significant risk for human health8,33,34.
Among our study, we also detected ST88, ST117 and ST362, which seem to be associated with calves as they were also found in a farm in Mecklenburg Western Pomerania10. ST362 is also known to be an efficient biofilm former33,35, therefore, is able to survive at surfaces of milking or feeding equipment or at calf barns/igloos10,20. Just like ST131, ST117 is also related to extraintestinal infections in humans and results of five studies indicate that poultry may be the reservoir, but it could also be found in calves34,36. With the present study being at least the third detecting ST117 in calves, it cannot be denied that calves may also play a role as a vector for human or avian ST117 colonization.
To our concern, > 50% of all isolates fulfilled the definition of being MDR and 31% of the ESBL-E. coli showed phenotypic quinolone resistance. Fortunately, none of the strains associated with human infections harbored multiple AMR.
In the present study, we were able to detect ESBL E. coli in a large number of different STs, which underlines the fundamental need for improved hygiene in calf husbandry. The transmission modes of the individual STs must be the subject of further investigations. With regard to colostrum as a possible source of ESBL E. coli, we were able to demonstrate the direct transmission of viable ESBL-E. coli via contaminated colostrum. Besides lowering the use of antibiotic agents in dairy herds, excellent sanitation procedures of the equipment for harvesting and feeding colostrum is crucial to reduce the prevalence of ESBL-E. coli in neonatal dairy calves, consequently minimizing the spread of AMR. Therefore, more attention should be paid on the improvement of farm hygiene, as it remains to be a simple and efficient method to prevent the environmental contamination with ESBL-E. coli.
Material and methods
Animals
Fecal, colostrum and equipment samples were collected as part of another study37 in which calves were initially fed with pooled colostrum at birth and thereafter moved to an experimental barn where they were individually housed and fed with milk replacer within the first eight days of life. Briefly, 15 male Holstein–Friesian calves from a German dairy farm were fed 3 L of colostrum right after birth at the farm. The colostrum was pooled prior to the beginning of the experiments and stored frozen in portions so that all calves received the same colostrum. The calves were then transported to the University of Leipzig within their first 24 h of life. There, they were stalled in one of two separate barns (control and infected with Cryptosporidium parvum, see below) with only one calf at a time. The calves were fed 3 × 2 L of milk replacer (Union A50 S, Arla Foods, Viby J, Denmark) daily and had free access to water in a bucket. Additionally, from day 4 onward they received 2 L of electrolytes (Ursolyt G oral, Serumwerk Bernburg AG, Bernburg, Germany) daily.
Upon arrival at the University, the animals were examined clinically, and blood and fecal samples were collected. Afterwards, one group of calves was infected by oral application of 2 × 107 Cryptosporidium parvum oocysts (n = 5) whereas the control group received only water. In the following days, the calves underwent several blood and fecal samplings for study purposes according to the aims of the concurrent study37.
The experiments, including all animal sampling, were conducted in accordance with the German legislation on the protection of animals and were approved by the Landesdirektion Leipzig as TVV 19/20. All authors comply with the ARRIVE guidelines.
Sampling, bacteriological examination
Fecal swabs (Sigma Transwab, MWE, United Kingdom) for the determination of ESBL/AmpC-E. coli carriage of the calves were taken at the 1st day (before infection with C. parvum), day 5 and day 8 of life, preserved in Amies medium and stored at 5 °C. A sample of the stored colostrum pool was additionally analyzed, and subsequently the dairy farm was visited and the equipment for harvesting and administering colostrum including liner, tube and milking pot and the nipple of the colostrum bottle was sampled using swabs.
Fecal, colostrum and equipment samples were cultured on CHROM ID agar plates (Mast Group, Reinfeld, Germany) supplemented with 2 µg/mL cefotaxime (Alfa Aesar by Thermo Fisher Scientific, Kandel, Germany) and incubated at 37 °C overnight. Moreover, the pooled colostrum was enriched in LB-broth supplemented with 2 µg/mL cefotaxime before cultivation. According to the manufacturer’s protocol, pink-violet colored, shiny colonies represent presumptive ESBL/AmpC-E. coli-positive results. Positive colonies were picked and sub-cultivated until a pure culture was achieved. All isolates were stored at − 80 °C until further use.
Antimicrobial susceptibility testing
AST was carried out using VITEK2 (bioMérieux, Nürtingen, Germany). Testing was performed using software version 9.02 and AST-N428 and AST-XN24 card, according to the manufacturer’s instructions. The AST card used for the VITEK2 included an ESBL confirmation test. Second and 3rd generation cephalosporins (ceftazidime, cefotaxime and cefuroxime) were used alone or in combination with tazobactam/avibactam. A reduction of growth in the presence of inhibitor of β-lactamases was considered indicative of ESBL production.
Minimal inhibitory concentration (MIC) breakpoints were set according to the European Committee on Antimicrobial Susceptibility Testing (EUCAST) breakpoint tables for interpretation of MICs and zone diameters (Version 13.1, 2023. http://www.eucast.org).
Whole genome sequencing and analysis
Whole genome sequencing of all isolates was applied (n = 45). DNA extraction was performed using the MasterPure™ DNA Purification Kit for Blood, Version II (Lucigen, Middleton, USA) and subsequently quantified using a Qubit 4 fluorometer (Thermofisher Scientific, Waltham, USA). DNA samples were then shipped to the Microbial Genome Sequencing Center (MiGS, Pittsburgh, PA, USA). Sample libraries were prepared using the Illumina DNA Prep kit and IDT 10 bp UDI indices, and sequenced on an Illumina NextSeq 2000, producing 2 × 151 bp reads. Demultiplexing, quality control and adapter trimming was performed with bcl-convert (v3.9.3) (Illumina, Inc.; https://support-docs.illumina.com/SW/BCL_Convert/Content/SW/FrontPages/BCL_Convert.htm).
The sequence analysis is described elsewhere38,39. In brief: We used BBDuk from BBTools v. 38.89 (http://sourceforge.net/projects/bbmap/) for (i) adapter-trimming, (ii) filtering for contaminants, and (iii) quality-trimming. For de novo genome assembly we used the shovill v. 1.1.0 assembly pipeline (https://github.com/tseemann/shovill) in combination with SPAdes v. 3.15.040. Thereafter, assemblies were analyzed for multilocus sequence type (MLST) determination and antibiotic resistance/virulence gene detection using the tools mlst v. 2.19.0 (https://github.com/tseemann/mlst) and ABRicate v. 1.0.0 (https://github.com/tseemann/abricate), respectively. Third-party databases (e.g., PubMLST41, VFDB42, ResFinder43, PlasmidFinder44, BacMet45, ARG-ANNOT46, and Ecoli_VF (https://github.com/phac-nml/ecoli_vf)) were used for the analyses of both tools.
Clonal expansion
To create the core SNP phylogeny of the 25 ESBL-producing ST10 isolates, the reads were mapped against 919 as reference genome using Snippy v. 4.6.0 (https://github.com/tseemann/snippy, accessed on 12.08.2022) to generate an alignment of these sequences. In the next step, the alignment was processed using Gubbins v.2.4.147, snp-sites v. 2.5.148 and FastTree v. 2.1.11 (http://www.microbesonline.org/fasttree/) (in detail described elsewhere5). The final alignment was midpoint-rooted in iTOL v. 6.7.249 and vizualized with the CTX-M types.
Snp-dists v. 0.8.2. (https://github.com/tseemann/snp-dists, accessed on 05.06.2023) was used to convert the FASTA alignment to a SNP distance matrix. (The distance matrix was depicted as a heat map diagram beside the core SNP phylogeny.)
Biofilm formation
Biofilm formation on polystyrene surfaces was assessed using crystal violet (CV) staining, as previously described5. The strength of biofilm formation was determined as specific biofilm formation (SBF), which was calculated using the formula: SBF = (B − NC)/G, where B is the OD570 of the stained bacteria, NC is the OD570 of the stained control wells to account for CV adhering to the polystyrene surface due to non-biological factors, and G is the OD600 representing the cell density in the culture medium. To evaluate biofilm formation, the test was also performed with two strains which are weak (PBIO72950) or strong (W3110) biofilm formers, respectively.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Tacconelli, E. et al. Discovery, research, and development of new antibiotics: The WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect. Dis. 18, 318–327 (2017).
GBDAR Collaborators. Global mortality associated with 33 bacterial pathogens in 2019: A systematic analysis for the Global Burden of Disease Study 2019. Lancet 400, 2221–2248. https://doi.org/10.1016/S0140-6736(22)02185-7 (2022).
Grover, N., Sahni, A. K. & Bhattacharya, S. Therapeutic challenges of ESBLS and AmpC beta-lactamase producers in a tertiary care center. Med. J. Armed Forces India 69, 4–10. https://doi.org/10.1016/j.mjafi.2012.02.001 (2013).
Dahms, C. et al. Occurrence of ESBL-producing Escherichia coli in livestock and farm workers in Mecklenburg-Western Pomerania, Germany. PLoS One https://doi.org/10.1371/journal.pone.0143326 (2015).
Eger, E. et al. Highly virulent and multidrug-resistant Escherichia coli sequence type 58 from a sausage in Germany. Antibiotics (Basel) https://doi.org/10.3390/antibiotics11081006 (2022).
Odenthal, S., Akineden, O. & Usleber, E. Extended-spectrum beta-lactamase producing Enterobacteriaceae in bulk tank milk from German dairy farms. Int. J. Food Microbiol. 238, 72–78. https://doi.org/10.1016/j.ijfoodmicro.2016.08.036 (2016).
Marshall, B., Petrowski, D. & Levy, S. B. Inter- and intraspecies spread of Escherichia coli in a farm environment in the absence of antibiotic usage. Proc. Natl. Acad. Sci. USA 87, 6609–6613. https://doi.org/10.1073/pnas.87.17.6609 (1990).
Homeier-Bachmann, T. et al. Antibiotic-resistant Enterobacteriaceae in wastewater of Abattoirs. Antibiotics (Basel) https://doi.org/10.3390/antibiotics10050568 (2021).
Schaumburg, F. et al. The risk to import ESBL-producing Enterobacteriaceae and Staphylococcus aureus through chicken meat trade in Gabon. BMC Microbiol. 14, 286. https://doi.org/10.1186/s12866-014-0286-3 (2014).
Homeier-Bachmann, T., Kleist, J. F., Schutz, A. K. & Bachmann, L. Distribution of ESBL/AmpC-Escherichia coli on a dairy farm. Antibiotics (Basel) https://doi.org/10.3390/antibiotics11070940 (2022).
Weber, L. P. et al. Prevalence and risk factors for ESBL/AmpC-E. coli in pre-weaned dairy calves on dairy farms in Germany. Microorganisms 9, 2135 (2021).
Springer, H. R. et al. Antimicrobial resistance in fecal Escherichia coli and Salmonella enterica from dairy calves: A systematic review. Foodborne Pathog. Dis. 16, 23–34. https://doi.org/10.1089/fpd.2018.2529 (2019).
Liu, J. et al. The fecal resistome of dairy cattle is associated with diet during nursing. Nat. Commun. 10, 4406. https://doi.org/10.1038/s41467-019-12111-x (2019).
Duse, A. et al. Risk factors for antimicrobial resistance in fecal Escherichia coli from preweaned dairy calves. J. Dairy Sci. 98, 500–516. https://doi.org/10.3168/jds.2014-8432 (2015).
Gonggrijp, M. A. et al. Prevalence and risk factors for extended-spectrum beta-lactamase- and AmpC-producing Escherichia coli in dairy farms. J. Dairy Sci. 99, 9001–9013. https://doi.org/10.3168/jds.2016-11134 (2016).
Hordijk, J. et al. Dynamics of faecal shedding of ESBL- or AmpC-producing Escherichia coli on dairy farms. J. Antimicrob. Chemother. 74, 1531–1538. https://doi.org/10.1093/jac/dkz035 (2019).
Schmid, A. et al. Prevalence of extended-spectrum β-lactamase-producing Escherichia coli on Bavarian dairy and beef cattle farms. Appl. Environ. Microbiol. 79, 3027–3032. https://doi.org/10.1128/aem.00204-13 (2013).
Brunton, L. A., Reeves, H. E., Snow, L. C. & Jones, J. R. A longitudinal field trial assesing the impact of feeding waste milk containing antibiotic residues on the prevalence of ESBL-producing Escherichia coli in calves. Prev. Vet. Med. 117, 403–412. https://doi.org/10.1016/j.prevetmed.2014.08.005 (2014).
Tetens, J. L., Billerbeck, S., Schwenker, J. A. & Holzel, C. S. Short communication: Selection of extended-spectrum beta-lactamase-producing Escherichia coli in dairy calves associated with antibiotic dry cow therapy—A cohort study. J. Dairy Sci. 102, 11449–11452. https://doi.org/10.3168/jds.2019-16659 (2019).
Heinemann, C., Leubner, C. D., Hayer, J. J. & Steinhoff-Wagner, J. Hygiene management in newborn individually housed dairy calves focusing on housing and feeding practices. J. Anim. Sci. https://doi.org/10.1093/jas/skaa391 (2021).
He, Z., Yang, S., Ma, Y., Zhang, S. & Cao, Z. Detection of CTX-M-15 extended-spectrum beta-lactamases producing Escherichia coli isolates from colostrum and faeces of newborn dairy calves in China. Pathogens https://doi.org/10.3390/pathogens10091162 (2021).
Mills, J. P. et al. Clinical and molecular epidemiology of extended-spectrum beta-lactamase-producing Escherichia coli infections in metro Detroit: Early dominance of the ST-131 clone. Infect. Dis. Ther. 9, 683–690. https://doi.org/10.1007/s40121-020-00321-6 (2020).
Godden, S. M., Lombard, J. E. & Woolums, A. R. Colostrum management for dairy calves. Vet. Clin. North Am. Food Anim. Pract. 35, 535–556. https://doi.org/10.1016/j.cvfa.2019.07.005 (2019).
Ludden, C. et al. Defining nosocomial transmission of Escherichia coli and antimicrobial resistance genes: A genomic surveillance study. Lancet Microbe 2, e472–e480. https://doi.org/10.1016/S2666-5247(21)00117-8 (2021).
Armengol, R. & Fraile, L. Colostrum and milk pasteurization improve health status and decrease mortality in neonatal calves receiving appropriate colostrum ingestion. J. Dairy Sci. 99, 4718–4725. https://doi.org/10.3168/jds.2015-10728 (2016).
Smith, B. I., Cady, S. V. & Aceto, H. W. Effect of formic acid treatment on colostrum quality, and on absorption and function of immunoglobulins: A randomized controlled trial in Holstein dairy calves. BMC Vet. Res. 18, 318. https://doi.org/10.1186/s12917-022-03418-x (2022).
Platell, J. L., Johnson, J. R., Cobbold, R. N. & Trott, D. J. Multidrug-resistant extraintestinal pathogenic Escherichia coli of sequence type ST131 in animals and foods. Vet. Microbiol. 153, 99–108. https://doi.org/10.1016/j.vetmic.2011.05.007 (2011).
Riley, L. W. Pandemic lineages of extraintestinal pathogenic Escherichia coli. Clin. Microbiol. Infect. 20, 380–390. https://doi.org/10.1111/1469-0691.12646 (2014).
Dahmen, S., Métayer, V., Gay, E., Madec, J. Y. & Haenni, M. Characterization of extended-spectrum beta-lactamase (ESBL)-carrying plasmids and clones of Enterobacteriaceae causing cattle mastitis in France. Vet. Microbiol. 162, 793–799. https://doi.org/10.1016/j.vetmic.2012.10.015 (2013).
Irrgang, A. et al. Diversity of CTX-M-1-producing E. coli from German food samples and genetic diversity of the bla(CTX-M-1) region on IncI1 ST3 plasmids. Vet. Microbiol. 221, 98–104. https://doi.org/10.1016/j.vetmic.2018.06.003 (2018).
Gerhold, G., Schulze, M. H., Gross, U. & Bohne, W. Multilocus sequence typing and CTX-M characterization of ESBL-producing E. coli: A prospective single-centre study in Lower Saxony, Germany. Epidemiol. Infect. 144, 3300–3304. https://doi.org/10.1017/s0950268816001412 (2016).
Chen, J. W. et al. Antibiotic-resistant Escherichia coli and sequence type 131 in fecal colonization in dogs in Taiwan. Microorganisms https://doi.org/10.3390/microorganisms8091439 (2020).
Gregova, G. & Kmet, V. Antibiotic resistance and virulence of Escherichia coli strains isolated from animal rendering plant. Sci. Rep. 10, 17108. https://doi.org/10.1038/s41598-020-72851-5 (2020).
Friese, A. et al. Faecal occurrence and emissions of livestock-associated methicillin-resistant Staphylococcus aureus (laMRSA) and ESBL/AmpC-producing E. coli from animal farms in Germany. Berl. Munch. Tierarztl. Wochenschr. 126, 175–180 (2013).
Zhuge, X. et al. Chicken-source Escherichia coli within phylogroup F shares virulence genotypes and is closely related to extraintestinal pathogenic E. coli causing human infections. Transbound. Emerg. Dis. 68, 880–895. https://doi.org/10.1111/tbed.13755 (2021).
Kim, S. W., Karns, J. S., Van Kessel, J. A. S. & Haley, B. J. Genome sequences of five multidrug-resistant Escherichia coli sequence type 117 isolates recovered from dairy calves. Genome Announc. https://doi.org/10.1128/genomeA.00732-17 (2017).
Dengler, F. et al. Cryptosporidium parvum competes with the intestinal epithelial cells for glucose and impairs systemic glucose supply in neonatal calves. Vet. Res. 54, 40. https://doi.org/10.1186/s13567-023-01172-y (2023).
Homeier-Bachmann, T. et al. Antibiotic-resistant Enterobacteriaceae in wastewater of abattoirs. Antibiotics 10, 568. https://doi.org/10.3390/antibiotics10050568 (2021).
Homeier-Bachmann, T. et al. Genomic analysis of ESBL-producing E. coli in wildlife from North-Eastern Germany. Antibiotics 11, 123 (2022).
Bankevich, A. et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477. https://doi.org/10.1089/cmb.2012.0021 (2012).
Jolley, K. A., Bray, J. E. & Maiden, M. C. J. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications [version 1; peer review: 2 approved]. Wellcome Open Res. https://doi.org/10.12688/wellcomeopenres.14826.1 (2018).
Liu, B., Zheng, D. D., Jin, Q., Chen, L. H. & Yang, J. VFDB 2019: A comparative pathogenomic platform with an interactive web interface. Nucleic Acids Res. 47, D687–D692. https://doi.org/10.1093/nar/gky1080 (2019).
Zankari, E. et al. Identification of acquired antimicrobial resistance genes. J. Antimicrob. Chemother. 67, 2640–2644. https://doi.org/10.1093/jac/dks261 (2012).
Carattoli, A. et al. In silico detection and typing of plasmids using plasmid finder and plasmid multilocus sequence typing. Antimicrob. Agents Chemother. 58, 3895–3903. https://doi.org/10.1128/Aac.02412-14 (2014).
Pal, C., Bengtsson-Palme, J., Rensing, C., Kristiansson, E. & Larsson, D. G. J. BacMet: Antibacterial biocide and metal resistance genes database. Nucleic Acids Res. 42, D737–D743. https://doi.org/10.1093/nar/gkt1252 (2014).
Gupta, S. K. et al. ARG-ANNOT, a new bioinformatic tool to discover antibiotic resistance genes in bacterial genomes. Antimicrob. Agents Chemother. 58, 212–220. https://doi.org/10.1128/aac.01310-13 (2014).
Croucher, N. J. et al. Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res. 43, e15. https://doi.org/10.1093/nar/gku1196 (2015).
Page, A. J. et al. SNP-sites: Rapid efficient extraction of SNPs from multi-FASTA alignments. Microb. Genom. 2, e000056. https://doi.org/10.1099/mgen.0.000056 (2016).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296. https://doi.org/10.1093/nar/gkab301 (2021).
Schaufler, K. et al. Carriage of extended-spectrum beta-lactamase-plasmids does not reduce fitness but enhances virulence in some strains of pandemic E. coli lineages. Front. Microbiol. 7, 336. https://doi.org/10.3389/fmicb.2016.00336 (2016).
Acknowledgements
We thank the participating farm for the eager willingness to be a part of the study. We furthermore thank the “Freundeskreis Tiermedizin” of the Faculty of Veterinary Medicine and Ceva Santé Animale for financing the experimental setting of the calves at the University of Leipzig and the Federal Ministry of Education and Research (BMBF) to support the position of MS by a grant entitled “Disarming pathogens as a different strategy to fight antimicrobial-resistant Gram-negatives” (01KI2015).
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
Planning and design of the study, LB, WL, HMH, FD, CD and THB; conducting and sampling, LB, WL, HMH, FD and CD; sample analysis, THB, AS, EE and MS; manuscript drafting, LB, LW, AS and THB; manuscript editing and finalizing, EE, MS, KB, KS, LB, AS and THB; All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bachmann, L., Weber, L., Liermann, W. et al. Colostrum as a source of ESBL-Escherichia coli in feces of newborn calves. Sci Rep 14, 9929 (2024). https://doi.org/10.1038/s41598-024-60461-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-60461-4
This article is cited by
-
Foodborne bacteria in milk and milk products along the water buffalo milk chain in Bangladesh
Scientific Reports (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.