Abstract
A synbiotic is a combination of live microorganisms and specific substrates that are selectively utilized by host microorganisms, resulting in health benefits for the host. Previous studies have demonstrated the protective effects of L. reuteri KUB-AC5 against Salmonella infection in chicken and mouse models. The probiotic activity of L. reuteri KUB-AC5 in these hosts was influenced by nutritional supplements. Water-based plants contain significant amounts of carbohydrates, particularly dietary fiber and proteins, making them potential prebiotic substrates. In this study, four water-based plants (Ulva rigida, Caulerpa lentillifera, Wolffia globosa, and Gracillaria fisheri) were screened for their ability to support the growth of L. reuteri KUB-AC5. Under monoculture testing, U. rigida exhibited the highest capacity to support the growth of L. reuteri KUB-AC5 and the production of organic acids, including acetic acid, lactic acid, and propionic acid (p ≤ 0.05). In co-culture experiments, the synbiotic combination of U. rigida and L. reuteri KUB-AC5 demonstrated the potential to eliminate Salmonella Typhimurium DMST 48437 when inoculated at 104 CFU/mL within 9 h. The synbiotic activities of U. rigida and L. reuteri KUB-AC5 were further investigated using an in vitro human gut model. Compared to the probiotic treatment, the synbiotic combination of L. reuteri KUB-AC5 and U. rigida showed significantly higher levels of L. reuteri KUB-AC5 (5.1 log copies/mL) and a reduction of S. Typhimurium by 0.8 log (CFU/ml) after 24 h (p ≤ 0.05). Synbiotic treatment also significantly promoted the production of short-chain fatty acids (SCFAs), including butyric acid, propionic acid, and acetic acid, compared to prebiotic and probiotic treatments alone (p ≤ 0.05). Furthermore, the synbiotic formulation modulated the in vitro simulated gut microbiome, enhancing putatively beneficial gut microbes, including lactobacilli, Faecalibacterium, and Blautia. Our findings demonstrated that L. reuteri KUB-AC5, in combination with U. rigida, exhibited synergistic activity, as indicated by increased viability, higher anti-pathogenicity toward Salmonella, and the ability to modulate the gut microbiome.
Similar content being viewed by others
Introduction
Salmonella infections cause significant problems, especially in developing countries, resulting in thousands of fatalities each year1,2. There is also a fear that the rising prevalence of multidrug-resistant Salmonella in the food supply chain will exacerbate this problem in the future3,4. Therefore, it is crucial to consider ways to prevent and minimize Salmonella infection in humans and livestock5,6. Limosilactobacillus reuteri KUB-AC5 has been shown to reduce salmonellosis in the poultry industry7,8,9 by producing antimicrobial substances that are effective against various Salmonella strains10. A previous study also reported that KUB-AC5 reduced Salmonella infection in mouse colitis models with attenuated gut inflammation in a cell density-dependent manner5,8.
A prebiotic is described as a substrate that is selectively utilized by host microorganisms to confer health benefits11. Water-based plants, particularly seaweed and duckweed, are rich sources of dietary fiber, proteins, polyunsaturated fatty acids, and minerals12. They have become attractive sources for commercial applications because of their fast growth rates and lack of arable land requirements compared to terrestrial plants13, with high dietary fiber content making them potential prebiotic candidates14. Several studies have shown that seaweed and duckweed are metabolizable by gut microbiota, thereby generating beneficial short-chain fatty acids (SCFAs)12,15,16,17.
A mixture of live microorganisms and a substrate that is selectively utilized by host microorganisms to promote host health is described as a synbiotic18. In recent years, the benefits of combining probiotics and prebiotics compared to their individual use have been increasingly recognized, with synbiotics demonstrating enhanced efficacy in various aspects of gut health. Studies have shown that synbiotics are more effective in reducing intestinal NH4+ levels, which is highly beneficial because it helps reduce the potential for gut dysbiosis. Moreover, lower levels of ammonium have been shown to create a conducive environment for the growth of butyric acid-producing bacteria19.
In an in vivo study involving mice with induced inflammatory bowel disease (IBD), the combination of Bacillus coagulans MTCC 5856 with sugarcane cane fiber exhibited a higher anti-inflammatory effect, reduced disease severity, and demonstrated superior modulation of metabolite and short-chain fatty acid (SCFA) profiles compared to probiotic or prebiotic treatment alone20.
A recent consensus by the International Scientific Association for Probiotics and Prebiotics (ISAPP)18 determined two main approaches for synbiotic formulation. The first is a complementary synbiotic with a formulation consisting of a recognized probiotic strain and a prebiotic substrate, where the probiotic and prebiotic do not necessarily have synergistic effects. The second is a synergistic synbiotic, where the substrate is designed to be selectively utilized by the probiotic; however, the prebiotic might also stimulate other beneficial members of the gastrointestinal microbiota21.
Addressing Salmonella infections has encountered challenges, including the emergence of drug-resistant strains and limitations in existing preventive strategies. Synbiotic represent a novel approach that harnesses the combined strength of probiotics and prebiotics. The choice of water-based plants as potential prebiotic substrates exhibits major advantages, with their attributes extending beyond mere high dietary fiber content12. These plants harbor an array of components and a diverse nutritional profile, including proteins, polyunsaturated fatty acids, and essential minerals. Moreover, with their fast growth rates and minimal arable land requirements, these plants provide a rich array of components that could foster a conducive environment for beneficial gut microorganisms.
In this study, we hypothesized that combining water-based plants with L. reuteri KUB-AC5 might enhance the overall functional capabilities, such as supporting the bacterial strain, stimulating specific groups of desirable gut microbes, and increasing the production of SCFAs. To prove or disprove this hypothesis, we investigated the efficacy of a synbiotic combination of L. reuteri KUB-AC5 and water-based plants during Salmonella challenge in an in vitro simulated human gut model. The gut model replicated the human gut’s environment and contained a human-like gut microbiome, enabling to explore the interaction between the human microbiome, synbiotics, and Salmonella.
Results
Nutrient composition of potential prebiotics from water-based plants
To formulate a functional synbiotic, the substrate must provide sufficient nutrients to support the growth of the co-administered bacteria. Here, we examined the fat, protein, ash, carbohydrate, and dietary fiber contents of water-based plants using proximate analysis (Table 1). More than 80% of the dry matter from all samples was carbohydrate and protein, except C. lentilifera, where ash and carbohydrate were the major compounds. Gracillaria fisheri had the highest carbohydrate content among the four tested water plants, followed by C. lentilifera and U. rigida, with the highest protein content found in W. globosa and U. rigida (Table 1).
Among the four water-based plants examined in this study, C. lentillifera and G. fisherii exhibited the highest carbohydrate and protein ratios, while U. rigida and W. globosa displayed balanced carbohydrate and protein ratios approaching a 1:1 proportion. These findings highlight the varying nutritional composition of the four plant species and provide insights into their suitability as substrates for microbial metabolism and growth (Table 1).
Survival of Limosilactobacillus reuteri KUB-AC5 during in vitro simulated passage through the upper gastrointestinal tract
The ability of L. reuteri KUB-AC5 to survive gastric conditions was tested, and a modest 0.7 log CFU/mL reduction after 2.5 h in simulated gastric juice was found, with a further reduction of 2 log CFU/mL observed after 4 h of exposure to artificial small intestinal fluid (Fig. 1).
Determination of the ability of water-based plants to support the growth of Limosilactobacillus reuteri KUB-AC5
The ability of L. reuteri KUB-AC5 to utilize water-based plant powders as carbon sources was investigated by substituting glucose in MRS broth with four different water-based plant powders (4% w/v) (Fig. 2a). Among the water-based plant powders tested, only U. rigida and W. globosa demonstrated significant growth-promoting effects compared to the negative control (MRS without added carbohydrate), reaching 8.29 and 8.96 log CFU/mL, respectively. The production of propionic acid, acetic acid, and lactic acid was also analyzed, and the results indicated that the levels of these organic acids produced by L. reuteri KUB-AC5 in the presence of U. rigida were the highest compared to those in other water-based plants (Fig. 2b). Therefore, U. rigida was selected to form a synbiotic with L. reuteri KUB-AC5.
Anti-Salmonella activity of Limosilactobacillus reuteri KUB-AC5 grown in the presence of Ulva rigida
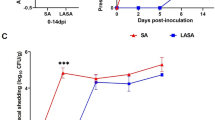
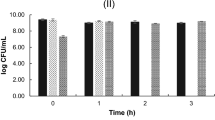
The anti-Salmonella activity of L. reuteri KUB-AC5 with U. rigida was investigated by co-cultivation with S. Typhimurium DMST 48437. The growth of S. Typhimurium DMST 48437 increased from 4 log CFU/mL to 5 log CFU/mL after 9 h and then suddenly decreased from 5 log CFU/mL to below the detection limit (102 CFU/mL) after 18 h. In contrast, the growth of S. Typhimurium DMST 48437 grown as a single culture increased from 4 to 8 log CFU/mL after 12 h. The growth curves of L. reuteri KUB-AC5 in both single and co-cultivation were similar and stable at 9 log CFU/mL for 24 h. This indicates that the combination of L. reuteri KUB-AC5 with U. rigida showed enhanced inhibitory activity against S. Typhimurium DMST 48437 (Fig. 3) relative to the individual constituents.
Growth of Limosilactobacillus reuteri KUB-AC5 (red circle) and Salmonella Typhimurium DMST 48437 (red triangle) co-cultured in gut basal medium with Ulva rigida supplement. Single cultures of L. reuteri KUB-AC5 (black circle) and S. Typhimurium DMST 48437 (black triangle) in gut basal medium with Ulva rigida supplement were used as controls.
Effect of Limosilactobacillus reuteri KUB-AC5 and Ulva rigida synbiotic on an in vitro simulated human colon microbiome exposed to a Salmonella challenge
As shown in Fig. S1, the richness and diversity indices of the in vitro simulated colon microbiome did not differ between the negative control and the prebiotic group, while the probiotic and synbiotic fermentations led to significantly lower values (p ≤ 0.05). The linear discriminant analysis (LDA) effect size (LEfSe) approach was applied to identify differences in the gut microbiota composition between treatments at 24 h. Distinctive phylotypes were found to be linked to the treatment (Fig. 4). The gut microbiome in each treatment group exhibited a different pattern. Salmonella, Clostridium, and Bifidobacterium constituted a large fraction of the bacteria present in the negative control, and the probiotic treatment showed an abundance of lactobacilli, whereas Bacteroides and Parabacteroides were significantly more abundant in the prebiotic and synbiotic treatments. Faecalibacterium and Blautia were also highest in the synbiotic treatments (Fig. 4).
Anti-Salmonella activity of Limosilactobacillus reuteri KUB-AC5 and Ulva rigida in an in vitro simulated human gut model
The Salmonella inhibitory activity of L. reuteri KUB-AC5 and U. rigida was investigated during in vitro simulation of the human colon model. The level of S. Typhimurium was quantified using quantitative real-time PCR (Fig. 5). Interestingly, S. Typhimurium constituted between 6.59 and 6.71 log (copies/mL) in the negative control, probiotic, and prebiotic treatments. This was significantly higher than the synbiotic treatment, resulting in 5.76 log (copies/mL) (p ≤ 0.05). This indicated that only the combination of L. reuteri KUB-AC5 and U. rigida inhibited Salmonella.
The level of L. reuteri KUB-AC5 in the human gut model was quantified using species-specific primers and quantitative real-time PCR. L. reuteri KUB-AC5 was detected only in the probiotic and synbiotic treatments, suggesting that donors in the gut microbiome did not contain L. reuteri KUB-AC5. The concentration of L. reuteri KUB-AC5 in the synbiotic treatment was significantly higher than that in the probiotic treatment (p ≤ 0.05), suggesting that L. reuteri KUB-AC5 utilized U. rigida during in vitro simulated colon passage.
The effect of Limosilactobacillus reuteri KUB-AC5 and Ulva rigida on SCFA production in an in vitro simulated colon model
Acetic, propionic, and butyric acids were identified in all treatments. The level of SCFAs in the synbiotic treatment was the highest, followed by the prebiotic, probiotic, and negative control (p ≤ 0.05). Interestingly, SCFA production in the synbiotic treatment was 1.6-fold higher than that in the prebiotic treatment, possibly because the synbiotic treatment enriched some bacterial taxa that could produce organic acids (Fig. 6a).
(a) Short-chain fatty acid (SCFA) profiles after 24 h of in vitro simulated colon passage. Different letters (A, B) beside the bar graph indicate significantly different concentrations of the SCFA in question (p < 0.05). (b) Pearson correlation analysis between SCFA and bacterial taxa (genus level). Only correlations with p < 0.05 are shown.
Pearson’s correlation analysis (Pearson) was used to identify bacterial taxa that correlated well with organic acid secretion. Overall, SCFA production was positively correlated with bacteria enriched in the prebiotic and synbiotic groups but inversely correlated with abundance in the negative control treatment (Fig. 6b). The strongest positive correlation in the synbiotic treatment was observed for Blautia with propionic and butyric acids and Faecalibacterium with butyric acid. A strong negative correlation was also observed between Enterobacteriaceae family members (Escherichia, Salmonella, and Klebsiella) and overall SCFA production.
Discussion
Prebiotics play a crucial role in promoting the growth and colonization of probiotic bacteria in the gut. By providing a favorable environment, synbiotics increase the chances of probiotics surviving and thriving in the gastrointestinal tract, which can lead to greater health benefits. To enhance the activity of L. reuteri KUB-AC5 in the human gut environment, four water-based plants were screened. Ulva rigida and W. globosa showed the best growth-promoting effects, while in in vitro co-culture, U. rigida also demonstrated a higher capacity for supporting the production of organic acids compared to other water-based plants (Fig. 2). Limosilactobacillus reuteri KUB-AC5 displayed robust growth in U. rigida and W. globosa, which had similar carbohydrate-to-protein ratios of approximately 1:1. In contrast, the other two water-based plants had carbohydrate–protein ratios ranging from 4 to 6. Interestingly, the 1:1 carbohydrate-to-protein ratio is comparable to that of MRS media, which is commonly used for culturing Lactobacillus spp.22. Furthermore, a recent study reported that Lactobacillus spp. utilize U. rigida and enhance the production of SCFAs23.
Our previous report showed that L. reuteri KUB-AC5 produced antimicrobial substances including short peptide and lactic acid against Salmonella. However, the inhibitory activity varies depending on the gut host environment5,8,24. The variation in L. reuteri KUB-AC5 against Salmonella in diverse gut environments emphasizes the importance of optimizing working conditions through synbiotic combinations. The addition of prebiotics can significantly contribute to lowering pH levels by offering a readily accessible substrate. This addition also might help to induce the production of SCFA, thus increasing the antimicrobial capabilities of L. reuteri. Therefore, a synbiotic combination of L. reuteri KUB-AC5 and U. rigida was formulated to evaluate its effectiveness in modulating gut microbiota composition and inhibiting the growth of Salmonella using an in vitro simulated human gut model. Our results showed that U. rigida was selectively utilized by L. reuteri KUB-AC5, leading to significantly higher levels of L. reuteri KUB-AC5 than probiotic treatment alone.
Elevated growth of L. reuteri KUB-AC5 in synbiotic treatment led to the inhibition of Salmonella during in vitro simulated colon passage. However, the inhibitory activity of L. reuteri KUB-AC5 in this model was lower than that of L. reuteri KUB-AC5 grown under co-culture conditions. Salmonella challenge disrupted the microbiome balance in the human gut model by promoting the proliferation of pathogenic bacteria within the Enterobacteriaceae family. Notably, the increased abundance of Enterobacteriaceae and Rikenellaceae was negatively correlated with the production of SCFAs, such as acetic acid, propionic acid, and butyric acid. Low levels of SCFAs in the gut have been linked to an increased risk of colorectal cancer, ulcerative colitis, and Crohn's disease25,26,27. Conversely, SCFAs enhance gut health by maintaining intestinal barrier integrity and protecting against inflammation28.
Both probiotic and synbiotic treatments affect the composition of the gut microbiota by increasing the abundance of Lactobacillaceae family members. Synbiotic treatment also influenced other beneficial gut microbes, including Faecalibacterium and Blautia, while prebiotic treatment encouraged the growth of Bacteroides and Parabacteroidetes. Pearson’s correlation analysis revealed a strong association between SCFAs and Faecalibacterium and Blautia, which were enriched in the synbiotic treatment. These findings suggest that synbiotic interventions might assist in providing favorable conditions for modulating the gut microbiome and contribute to the suppression of Salmonella in the gut.
Previous research has shown that SCFA depletion is associated with Salmonella infections in pigs 29. Consistent with this finding, reduced fecal SCFA levels in children with salmonellosis have also been observed30. Both studies used probiotics to ameliorate infection and recorded increased levels of SCFAs. Higher SCFA levels were also linked to milder symptoms in children with salmonellosis, and probiotic administration demonstrated protective properties against future infection in piglets. These results indicate that SCFA-producing probiotics may play a prominent role in the treatment of Salmonella infections.
Materials and methods
Bacterial cultures and potential prebiotic powder origin
Limosilactobacillus reuteri KUB-AC5 was supplied by the Specialized Research Unit of Probiotics and Prebiotics for Health, Department of Biotechnology, Kasetsart University, Thailand, and Salmonella enterica serovar Typhimurium was provided by the Research and Development Center, Betagro Agro-Group Public Co., Ltd., Thailand. Limosilactobacillus reuteri KUB-AC5 was maintained on MRS agar (Difco Laboratories, Detroit, MI, USA) supplemented with 0.5% CaCO3, whereas S. Typhimurium DMST 48437 was maintained on nutrient agar (Oxoid, Basingstoke, UK). All isolates were preserved in nutrient broth or MRS broth supplemented with 40% glycerol and stored at -80 °C.
Green seaweed (U. rigida and C. lentillifera) and red seaweed (G. fisheri) were collected from a seaweed farm in southern Thailand between May and June 2021, with support from the Phetchaburi Coastal Fisheries Research and Development Center, Department of Fisheries, Ministry of Agriculture and Cooperatives. Duckweed (W. globosa) was provided by the Advanced GreenFarm Co., Ltd. (Nakhon Pathom, Thailand). After harvesting, the seaweeds were washed with tap water to remove any visible surface contaminants and dried in an oven at 60 °C for 8 h to obtain a moisture content of approximately 10%. The seaweeds were then finely ground using a hammer mill and sieved through a 0.2 mm mesh. All seaweed and duckweed were collected and prepared simultaneously to obtain consistent samples for the entire study. The ground sample powders were stored at room temperature in a desiccator until further use.
Proximate analysis of water-based plants
The chemical composition of the ground samples was determined by proximate analysis, following the methods described by the Association of Official Analytical Chemists31. Moisture, protein, ash, crude fiber, and fat contents were analyzed using a hot air oven, N-Kjeldahl × 6.25, ignition at 550 °C, Fibertec™, and Soxhlet extraction, respectively. The carbohydrate content was calculated using the available carbohydrate by the difference (CHOAVDF) method. The total dietary fiber content, including soluble and insoluble fiber, was also analyzed according to the AOAC32.
Preparation of bacterial cultures
Limosilactobacillus reuteri KUB-AC5 and S. Typhimurium inocula were cultivated in MRS broth (Difco Laboratories, Detroit, MI, USA) and nutrient broth (Oxoid, Basingstoke, UK), respectively. Salmonella Typhimurium was grown with agitation (120 rpm) for 18 h, whereas L. reuteri was cultured under static conditions for 15 h. Both strains were incubated at 37 °C. After centrifugation at 13,000×g for 3 min at 4 °C, the supernatant was discarded, and the pelleted cells were resuspended in an equal volume of phosphate buffer pH 7 before each experiment.
Screening of water-based plants as substrates for the growth of Limosilactobacillus reuteri KUB-AC5
Modified MRS without glucose33 supplemented with 4% w/v water-based plant extract in sterile 50 mL Duran bottles and 2% v/v L. reuteri KUB-AC5 culture (adjusted to 108 CFU/mL) was used to test the ability of the extracts to support L. reuteri KUB-AC5 growth during incubation at 37 °C for 15 h without agitation. Growth was determined by plating on MRS agar (Difco Laboratories, Detroit, MI, USA) and is presented as log CFU/mL. All experiments were performed in duplicate. For SCFA analysis, the culture broth was centrifuged twice at 13,000×g for 5 min, and the supernatants were stored at − 20 °C until analysis by high-performance liquid chromatography.
Survival of Limosilactobacillus reuteri KUB-AC5 under in vitro simulated upper gastrointestinal tract conditions
To examine the ability of L. reuteri KUB-AC5 to survive the harsh conditions of the human gastrointestinal tract, the bacterium was exposed to in vitro simulated stomach and small intestine conditions. For the stomach phase, artificial gastric fluids consisting of a pH 2.5 solution of pepsin (Oxoid, Basingstoke, UK) (2000 UA/mL) in phosphate buffer were used, while for the small intestine phase, artificial intestinal fluids consisting of a pH 8 solution of pancreatin (200 UA/mL) and 0.3% bile salt were used 34,35,36. The number of viable cells was assessed sequentially in each phase of the artificial gastrointestinal tract (baseline, stomach, and small intestine).
The artificial stomach condition was initiated by mixing L. reuteri KUB-AC5 suspensions (109 CFU/mL) with 9 mL of artificial gastric fluid. The mixture was then incubated at 37 °C with homogenization every 20 min for 2.5 h. To simulate passage to the small intestine, the artificial gastric fluid was removed by centrifugation, and the pelleted cells were resuspended in 10 mL of artificial intestinal fluid. The mixture was incubated at 37 °C for 4 h, with homogenization every 40 min. After each of the two stages, the artificial gastrointestinal fluids were removed by centrifugation, and the cells were resuspended and serially diluted with the maximum recovery diluent (Oxoid, Basingstoke, UK) before plating on MRS agar (Difco Laboratories, Detroit, MI, USA).
Co‑cultivation of Limosilactobacillus reuteri KUB-AC5 and Salmonella Typhimurium supplemented with Ulva rigida powder
The effect of U. rigida supplementation on the ability of L. reuteri KUB-AC5 to inhibit the growth of S. Typhimurium was investigated in co-culture using gut basal medium (0.2% w/v peptone, 0.2% w/v yeast extract, 0.01% w/v NaCl, 0.004% w/v K2HPO4, 0.004% w/v KH2PO4, 0.001% w/v MgSO4, 0.001% CaCl2, 0.2% w/v NaHCO3, 0.2% v/v Tween 80, 0.5% w/v bile salt, 0.001% v/v Antifoam SE15, and 0.001% w/v resazurin) with 4% w/v U. rigida powder.
For the assay, L. reuteri KUB-AC5 and Salmonella Typhimurium were seeded into gut basal broth supplemented with U. rigida at final concentrations of 109 and 104 CFU/mL, respectively. The adjustment process was performed by measuring the OD600 of each strain with a spectrophotometer and was confirmed by plating each inoculum onto its respective medium. Monocultures of each strain in the same medium were used as controls. All tubes were then incubated under static conditions at 37 °C for 24 h, followed by bacterial enumeration in selective media, MRS (Difco Laboratories, Detroit, MI, USA), and nutrient agar (Oxoid, Basingstoke, UK) for L. reuteri KUB-AC5 and S. Typhimurium DMST 48437, respectively. The results were reported as log CFU/mL. Each co-culture was performed in duplicate.
Inhibition of Salmonella Typhimurium by Limosilactobacillus reuteri KUB-AC5 during in vitro simulated colon passage
The ability of L. reuteri KUB-AC5 to inhibit Salmonella Typhimurium during in vitro simulated colon passage with and without U. rigida supplementation was investigated using the CoMiniGut system, with experimental conditions designed to simulate colon transit for 24 h, as previously described 37. Fecal inocula were collected from three healthy adults who had not received antibiotic treatment or probiotics during the preceding 3 months. Informed consent from the volunteers were obtained before sample collection and the process was conducted according to the Helsinki Declaration. All protocol involving fecal sample collection from volunteers were approved by the Ethical Committee of the Capital Region of Denmark registration number H-20028549. The fecal slurries were individually homogenized in a 1:1 ratio with 1 M PBS and 20% glycerol (v/v) for 2 × 60 s using a Stomacher (Stomacher 400; Seward, Worthing, UK) at normal speed. Fecal slurries were then aliquoted and stored at − 80 °C until further use. Fecal slurry stocks were thawed and diluted with 0.1 M PBS pH 5.6 (1:4) on the day of the experiment.
Fecal batch fermentations were separated into four treatments: negative control, probiotic (L. reuteri KUB-AC5 at 8 log CFU/mL)), prebiotic (4% w/v of U. rigida), and synbiotic (L. reuteri KUB-AC5 at 8 log CFU/mL and 4% w/v of U. rigida). All treatments were spiked with S. Typhimurium DMST 48437 at 4 log CFU/mL. Each treatment was fermented in triplicate in a random order to avoid potential run effects. Samples were collected at 0 and 24 h for microbiome determination and metabolite analysis.
Bacterial DNA extraction
Bacterial DNA was extracted according to the DNeasy PowerSoil Pro Kit protocol (Qiagen) from CoMiniGut fermentates at 0 h and 24 h of fermentation, with the cell pellet obtained from 125 to 200 µL of fermentation by centrifugation at 13,000×g for 10 min. The FastPrep bead-beating step was performed in two cycles of 30 s each at a speed of 2000 rpm in a FastPrep-24™ homogenizer (MP). DNA quantity and quality were measured using a QubitTM4 Fluorometer (Thermo Scientific, Waltham, MA, USA).
Microbiota profile analysis
The fecal microbiota composition in the gut model was determined using MinION (Oxford Nanopore Technologies, Oxford, UK), with PCR and library preparation of the V1–V8 hypervariable region of the 16S rRNA gene, as described previously38.
Data generated by MinION were collected using MinKnow software v19.06.8 (https://nanoporetech.com). The Guppy v3.2.2 base-calling toolkit was used to base raw fast5 to fastq (https://nanoporetech.com). Porechop v0.2.2 was used for adapter trimming and sample demultiplexing (https://github.com/rrwick/Porechop). Sequences containing quality scores (fastq files) were quality corrected using NanoFilt (q ≥ 10; read length > 1 kb). Taxonomy assignment of quality-corrected reads against the Greengenes (13.8) database was conducted using the uclust method implemented in parallel_assign_taxonomy_uclust.py (QIIME v1.9.1). The uclust settings were tuned to mock communities (similarity 0.8; min_consensus_fraction 0.51), assuring annotations to the lowest taxonomic level with no false positive annotations. The settings allowed the program to treat individual amplicon sequence variants as individual “seeds”. Reads classified at the minimum phylum level were subjected to further analysis. Alpha diversity matrices (Chao1 richness, Pielou’s evenness, and Shannon index) and Bray‒Curtis beta diversity were calculated based on the amplicon sequence variance (ASVs) table using USEARCH v11.0.66739.
Quantification of Limosilactobacillus reuteri KUB-AC5 and Salmonella Typhimurium by quantitative PCR (qPCR)
Limosilactobacillus reuteri KUB-AC5 and S. Typhimurium were quantified by qPCR using the LightCycler480 platform (Roche, Basel, Switzerland). An oligonucleotide primer for L. reuteri KUB-AC5 was developed in this study based on the protein sequence of the antimicrobial peptide from L. reuteri KUB-AC5, which has a molecular weight of active AMP of 4.7 kDa as determined by MALDI-TOF mass spectrometry10,24. The protein sequence was converted to a DNA sequence and checked for specificity using the NCBI BLAST database (http://www.ncbi.nlm.nih.gov/BLAST, accessed on 23rd October 2019). Primer quality was verified using the PrimerQuest™ tool (IDT, Coralville, IA, USA). The unique putative primers of L. reuteri KUB-AC5 (AC-1792F: 5′-CGAAAATGGGAGTAATTAACTATGG-3′ and AC-1792R: 5′-ATTTCCTGCAGCTAAACTTCCA-3′) were validated for specificity against several Lactobacillus species and Escherichia coli. To obtain absolute quantification of S. Typhimurium, the STM4497 primer set was used, including STM4497F:5′-AACAACGGCTCCGGTAATGA and STM4497R3:5′-TGACAAACTCTTGATTCTGA40. The reaction mixture contained 10 μL of SsoAdvanced Universal SYBR Green Supermix (Bio-Rad, Hercules, CA, USA), 0.5 μL of 10 pmol/μL of the forward and reverse primers, 2 μL of DNA template (30–50 ng), and nuclease-free water added to obtain a final volume of 20 μL. The amplification conditions were as follows: initial denaturation at 95 °C for 5 min, 30 cycles of amplification with denaturation at 95 °C for 1 min, annealing at the primer specified temperature (56 °C for AC5-1792 and 53 °C for STM4497) for 15 s, extension at 72 °C for 25 s, and a final extension at 72 °C for 5 min. To confirm the specific amplification of the target DNA, a dissociation curve was created using a denaturation step at 95 °C for 5 s, decreased to 65 °C for 1 min, and continuously increased from 65 to 97 °C, with signal measurement every 12 s. Specific sizes of the PCR products were determined by gel electrophoresis.
Standard curves were constructed using specific primers to amplify plasmids containing DNA from L. reuteri KUB-AC5 and S. Typhimurium DMST 48437. A standard plasmid was constructed following a previously described method41. Ten-fold serial dilutions of the plasmid, ranging from 101 to 108 copies/μL, were used to create a standard curve.
Metabolite analysis of fecal fermentation broths
Fatty acids in in vitro colon-simulated fecal samples were quantified by high-performance liquid chromatography (Waters, Milford, MA, USA) using a UV detector at a wavelength of 210 nm. The sample supernatant was passed through a 0.22 µM filter (Vertical, Santa Clara, CA, USA) and mixed with 0.2% v/v tartaric acid (2:1). For SCFA separation, an Aminex HPX-87H column (300 × 7.8 mm) (Bio-Rad, Hercules, CA, USA) served as the static phase, with 8 mM sulfuric acid as the mobile phase. The flow rate was set at 0.6 mL/minute, and the injector and detector temperatures were maintained at 55 °C. Standard curves were constructed by injecting solutions of lactic acid, acetic acid, propionic acid, and butyric acid at concentrations between 0.9375 and 83 mmol/mL.
Statistical analysis
All data used for statistical purposes were checked using the Shapiro‒Wilk normality test in GraphPad Prism version 8.4.3. ANOVA was used for parametric data, and the Kruskal‒Wallis test was used for non-parametric data. LEfSe analysis was performed to determine OTU biomarkers in each group based on previously described methods42,43. In brief, significant differences in relative abundance between each treatment were determined using two-tailed non-parametric Kruskal‒Wallis and unpaired Wilcoxon tests. LDA was performed to estimate the effect size for significantly different taxa. Only gut bacteria with a p value ≤ 0.05 and LDA scores higher than two were considered significantly different. Correlation analyses between SCFA and bacteria at the family and genus levels were performed using MetScape3 correlation calculator v1.0144 and visualized as a heatmap on ImageGP43.
Data availability
The raw 16 s rRNA amplicon sequences used in this study have been deposited in the NCBI shorts read archive (SRA) under the Bio Project Accession Number PRJNA906335.
References
Akil, L., Ahmad, H. A. & Reddy, R. S. Effects of climate change on Salmonella infections. Foodborne Pathog. Dis. 11, 974–980. https://doi.org/10.1089/fpd.2014.1802 (2014).
Marchello, C. S., Birkhold, M. & Crump, J. A. Complications and mortality of non-typhoidal Salmonella invasive disease: A global systematic review and meta-analysis. Lancet Infect. Dis. 22, 692–705. https://doi.org/10.1016/s1473-3099(21)00615-0 (2022).
Chonsin, K. et al. Prevalence and multidrug resistance of Salmonella in swine production chain in a Central Province, Thailand. J. Food Prot. 84, 2174–2184. https://doi.org/10.4315/jfp-21-003 (2021).
Nair, D. V. T., Venkitanarayanan, K. & Kollanoor Johny, A. Antibiotic-resistant Salmonella in the food supply and the potential role of antibiotic alternatives for control. Foods 7, 167. https://doi.org/10.3390/foods7100167 (2018).
Buddhasiri, S. et al. Anti-inflammatory effect of probiotic Limosilactobacillus reuteri KUB-AC5 against Salmonella infection in a mouse colitis model. Front. Microbiol. 12, 716761. https://doi.org/10.3389/fmicb.2021.716761 (2021).
Buddhasiri, S. et al. Probiotic Limosilactobacillus reuteri KUB-AC5 inhibits growth of clinical Salmonella enterica typhimurium isolates. Asia-Pac. J. Sci. Technol. 27, 28. https://doi.org/10.14456/apst.2022.30 (2022).
Nakphaichit, M. et al. The effect of including Lactobacillus reuteri KUB-AC5 during post-hatch feeding on the growth and ileum microbiota of broiler chickens. Poult. Sci. 90, 2753–2765. https://doi.org/10.3382/ps.2011-01637 (2011).
Nakphaichit, M. et al. Protective effect of Lactobacillus reuteri KUB-AC5 against Salmonella enteritidis challenge in chickens. Benef. Microbes 10, 43–54. https://doi.org/10.3920/BM2018.0034 (2019).
Jatuponwiphat, T. et al. Comparative genome analysis reveals metabolic traits associated with probiotics properties in Lactobacillus reuteri KUB-AC5. Gene Rep. 17, 100536. https://doi.org/10.1016/j.genrep.2019.100536 (2019).
Sobanbua, S. et al. Cloning and expression of the antimicrobial peptide from Lactobacillus reuteri KUB-AC5 and its characterization. Int. J. Agric. Technol. 16, 1013–1036 (2020).
Gibson, G. R. et al. Expert consensus document: The International Scientific Association for Probiotics and Prebiotics (ISAPP) consensus statement on the definition and scope of prebiotics. Nat. Rev. Gastroenterol. Hepatol. 14, 491–502. https://doi.org/10.1038/nrgastro.2017.75 (2017).
Charoensiddhi, S. et al. Gut microbiome modulation and gastrointestinal digestibility in vitro of polysaccharide-enriched extracts and seaweeds from Ulva rigida and Gracilaria fisheri. J. Funct. Foods 96, 105204. https://doi.org/10.1016/j.jff.2022.105204 (2022).
Charoensiddhi, S., Conlon, M., Franco, C. & Zhang, W. The development of seaweed-derived bioactive compounds for use as prebiotics and nutraceuticals using enzyme technologies. Trends Food Sci. Technol. 70, 20–33. https://doi.org/10.1016/j.tifs.2017.10.002 (2017).
Charoensiddhi, S., Abraham, R. E., Su, P. & Zhang, W. Seaweed and seaweed-derived metabolites as prebiotics. Adv. Food Nutr. Res. 91, 97–156. https://doi.org/10.1016/bs.afnr.2019.10.001 (2020).
Shannon, E., Conlon, M. & Hayes, M. Seaweed components as potential modulators of the gut microbiota. Marine Drugs 19, 358 (2021).
Gonçalves, A. T., Simões, M., Costa, C., Passos, R. & Baptista, T. Modulatory effect of Gracilaria gracilis on European seabass gut microbiota community and its functionality. Sci. Rep. 12, 14836. https://doi.org/10.1038/s41598-022-17891-9 (2022).
Diotallevi, C. et al. Measuring the effect of Mankai® (Wolffia globosa) on the gut microbiota and its metabolic output using an in vitro colon model. J. Funct. Foods 84, 104597. https://doi.org/10.1016/j.jff.2021.104597 (2021).
Swanson, K. S. et al. The International Scientific Association for Probiotics and Prebiotics (ISAPP) consensus statement on the definition and scope of synbiotics. Nat. Rev. Gastroenterol. Hepatol. 17, 687–701. https://doi.org/10.1038/s41575-020-0344-2 (2020).
Bianchi, F. et al. In vitro modulation of human gut microbiota composition and metabolites by Bifidobacterium longum BB-46 and a citric pectin. Food Res. Int. 120, 595–602. https://doi.org/10.1016/j.foodres.2018.11.010 (2019).
Shinde, T. et al. Synbiotic supplementation containing whole plant sugar cane fibre and probiotic spores potentiates protective synergistic effects in mouse model of IBD. Nutrients 11, 818 (2019).
Kolida, S. & Gibson, G. R. Synbiotics in health and disease. Annu. Rev. Food Sci. Technol. 2, 373–393. https://doi.org/10.1146/annurev-food-022510-133739 (2011).
Reuter, G. Elective and selective media for lactic acid bacteria. Int. J. Food Microbiol. 2, 55–68. https://doi.org/10.1016/0168-1605(85)90057-1 (1985).
Guan, Z. W., Yu, E. Z. & Feng, Q. Soluble dietary fiber, one of the most important nutrients for the gut microbiota. Molecules 26, 6802. https://doi.org/10.3390/molecules26226802 (2021).
Sobanbua, S., Dolkittikul, S., Nakphaichit, M., Keawsompong, S. & Nitisinprasert, S. Antimicrobial peptide presenting potential strain-specific real time polymerase chain reaction assay for detecting the probiotic Lactobacillus reuteri KUB-AC5 in chicken intestine. Poult. Sci. 99, 526. https://doi.org/10.3382/ps/pez549 (2019).
Kumari, R., Ahuja, V. & Paul, J. Fluctuations in butyrate-producing bacteria in ulcerative colitis patients of North India. World J. Gastroenterol. 19, 3404–3414. https://doi.org/10.3748/wjg.v19.i22.3404 (2013).
Parada Venegas, D. et al. Short chain fatty acids (SCFAs)-mediated gut epithelial and immune regulation and its relevance for inflammatory bowel diseases. Front. Immunol. https://doi.org/10.3389/fimmu.2019.00277 (2019).
O’Keefe, S. J. D. Diet, microorganisms and their metabolites, and colon cancer. Nat. Rev. Gastroenterol. Hepatol. 13, 691–706. https://doi.org/10.1038/nrgastro.2016.165 (2016).
Silva, Y. P., Bernardi, A. & Frozza, R. L. The role of short-chain fatty acids from gut microbiota in gut-brain communication. Front. Endocrinol. https://doi.org/10.3389/fendo.2020.00025 (2020).
He, T. et al. Lactobacillus johnsonii L531 reduces pathogen load and helps maintain short-chain fatty acid levels in the intestines of pigs challenged with Salmonella enterica Infantis. Vet. Microbiol. 230, 187–194. https://doi.org/10.1016/j.vetmic.2019.02.003 (2019).
Siigur, U. et al. Effect of bacterial infection and administration of a probiotic on faecal short-chain fatty acids. Microb. Ecol. Health Dis. 9, 271–277. https://doi.org/10.3109/08910609609166467 (2009).
AOAC & Latimer, G. W. Official Methods of Analysis of AOAC International. (AOAC International, 2012).
AOAC. Official Methods of Analysis of AOAC International. 17 edn, (AOAC International, 2010).
Namrak, T. et al. Probing genome-scale model reveals metabolic capability and essential nutrients for growth of probiotic Limosilactobacillus reuteri KUB-AC5. Biology 11, 294 (2022).
Minekus, M. et al. A standardised static in vitro digestion method suitable for food: An international consensus. Food Funct. 5, 1113–1124. https://doi.org/10.1039/C3FO60702J (2014).
Naissinger da Silva, M., Tagliapietra, B. L., Flores, Vd. A. & Pereira dos Santos Richards, N. S. In vitro test to evaluate survival in the gastrointestinal tract of commercial probiotics. Current Res. Food Sci. 4, 320–325. https://doi.org/10.1016/j.crfs.2021.04.006 (2021).
Campana, R., van Hemert, S. & Baffone, W. Strain-specific probiotic properties of lactic acid bacteria and their interference with human intestinal pathogens invasion. Gut Pathog. 9, 12. https://doi.org/10.1186/s13099-017-0162-4 (2017).
Wiese, M. et al. CoMiniGut-a small volume in vitro colon model for the screening of gut microbial fermentation processes. PeerJ 6, e4268. https://doi.org/10.7717/peerj.4268 (2018).
Hui, Y. et al. Supplementation of a lacto-fermented rapeseed-seaweed blend promotes gut microbial- and gut immune-modulation in weaner piglets. J. Anim. Sci. Biotechnol. 12, 85. https://doi.org/10.1186/s40104-021-00601-2 (2021).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Kim, H. J. et al. Identification of Salmonella enterica Serovar Typhimurium using specific PCR primers obtained by comparative genomics in Salmonella Serovars. J. Food Prot. 69, 1653–1661. https://doi.org/10.4315/0362-028X-69.7.1653 (2006).
Mok, K. et al. ITS2 sequencing and targeted meta-proteomics of infant gut mycobiome reveal the functional role of Rhodotorula sp. during atopic dermatitis manifestation. J. Fungi 7, 748 (2021).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60. https://doi.org/10.1186/gb-2011-12-6-r60 (2011).
Chen, T., Liu, Y.-X. & Huang, L. ImageGP: An easy-to-use data visualization web server for scientific researchers. iMeta 1, e5. https://doi.org/10.1002/imt2.5 (2022).
Basu, S. et al. Sparse network modeling and metscape-based visualization methods for the analysis of large-scale metabolomics data. Bioinformatics 33, 1545–1553. https://doi.org/10.1093/bioinformatics/btx012 (2017).
Acknowledgements
This research was funded by Kasetsart University through the Graduate School Fellowship Program and an Agro-Industrial Scholarship for International Students at Kasetsart University. The authors also wish to thank the Research and Development Center, Betagro Agro-Group Public Co., Ltd., Thailand, for providing the Salmonella strain DMST 48437.
Funding
This work was financially supported by The Office of the Ministry of Higher Education, Science, Research and Innovation; The Thailand Science Research and Innovation through the Kasetsart University Reinventing University Program 2021; Kasetsart University through the Graduate School Fellowship Program; and Agro-Industrial Scholarship for International Students, Kasetsart University, and the Department of Biotechnology, Faculty of Agro-Industry, Kasetsart University.
Author information
Authors and Affiliations
Contributions
Conceptualization M.N., S.C, D.S.N.; data curation K.M., O.H., T.F.; methodology K.M., S.C., D.S.N., M.N.; investigation K.M., O.H., M.N.; formal analysis K.M., O.H., T.F., with input from S.N., D.S.N.; visualization K.M., T.F.; supervision S.N., D.S.N., M.N.; writing—original Draft K.M., S.C., M.N.; writing—review and editing K.M., S.C., S.N., D.S.N., M.N.; funding Acquisition M.N., D.S.N.; project administration S.C., D.S.N., M.N.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mok, K., Honwichit, O., Funnuam, T. et al. Synergistic activity of Limosilactobacillus reuteri KUB-AC5 and water-based plants against Salmonella challenge in a human in vitro gut model. Sci Rep 14, 4730 (2024). https://doi.org/10.1038/s41598-024-53912-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-53912-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.