Abstract
Human malaria, transmitted by Anopheles mosquitoes, is the most predominant mosquito-borne disease that is responsible for hundreds of thousands of deaths worldwide each year. In the Republic of Korea (ROK), there are currently several hundred malaria cases annually, mostly near the demilitarized zone (DMZ). Eight species of Anopheles mosquitoes are currently known to be present in the ROK. Similar to other major malaria vectors in Africa and India, it is very challenging to morphologically differentiate Anopheles mosquitoes in the ROK due to their extremely similar morphology. In this study, wing geometric morphometrics (WGM) were used to differentiate the eight Anopheles species collected at six locations near the DMZ, Seoul and Pyeongtaek from April–October 2021. Phylogenetic analysis was also performed using cytochrome c oxidase subunit 1 (COI), internal transcribed spacer 2 (ITS2), and tyrosine hydroxylase (TH) genes for comparison with WGM analysis and to infer evolutionary relationships. The results of cross-validation (overall accuracy = 74.8%) demonstrated that species identification using WGM alone was not possible with a high accuracy for all eight species. While phylogenetic analyses based on the COI region could not clearly distinguish some species, the analysis based on ITS2 and TH was more useful for resolving the phylogenetic correlation of the eight species. Our results may improve Anopheles species identification strategies for effective identification and control of malaria vectors in the ROK.
Similar content being viewed by others
Introduction
Malaria, a mosquito-borne disease transmitted by Anopheles spp. causes more than 600,000 deaths annually1. The primary mosquito vectors of the five human malarias belong to the genus Anopheles (Diptera: Culicidae). Malaria is caused by protozoan parasites (Plasmodium spp.) that enter the host’s bloodstream when female mosquitoes blood feed for egg development2. In the Republic of Korea (ROK), there are currently approximately 200–300 locally acquired malaria cases annually resulting from infections of Plasmodium vivax3,4. While there are imported cases of Plasmodium falciparum, autochthonous transmission has not been documented5.
Eight Anopheles species are found in the ROK, including six members belonging to the Hyrcanus Group (Anopheles pullus Yamada, Anopheles belenrae Rueda, Anopheles sineroides Yamada, Anopheles lesteri Baisas & Hu, Anopheles kleini Rueda, Anopheles sinensis sensu stricto Wiedemann; and one each belonging to the Barbirostris Group (Anopheles koreicus Yamada & Watanabe) and Lindesayi Group (Anopheles lindesayi Giles)6,7. Two species, An. lesteri and An. kleini are suspected to be the primary malaria vectors. However, there is much controversy concerning which species are the primary vectors of vivax malaria in the ROK8,9,10,11. Nevertheless, studies in the ROK have detected P. vivax in all species, except An. koreicus that is infrequently collected by mosquito traps12,13,14. Some members of the Barbirostris Group are major malaria vectors in some tropical regions15, such as Thailand16, suggesting the need to continue to monitor and control all Anopheles mosquitoes in the ROK. In the ROK, An. sinensis seems to have the widest distribution range among Anopheles mosquitoes7,17, and there are differences in seasonal occurrence of the species belonging to the Hyrcanus Group14. A survey of the larval habitat range conducted by Rueda et al.18 also found that An. sinensis was collected in the most diverse environments, while An. koreicus and An. lindesayi were only found primarily in forested areas7,18,19.
Effective malaria control is contingent on accurate surveillance of suspected malaria vector species. Anopheles mosquitoes are highly susceptible to the loss of primary morphological taxonomic key characteristics, e.g., wing scale patterns and leg spots, during their collection. In addition, Anopheles mosquitoes are morphologically extremely similar, making them difficult to identify to the species level20. This challenge is not limited to ROK Anopheles but also other Anopheles species in other continents, e.g., Africa21 and South America22. While molecular markers are currently being used for species identification of Anopheles spp., the process is time consuming and costly23,24. Therefore, more affordable and streamlined surveillance tools are necessary for timely monitoring Anopheles species.
The mitochondrial genome is a commonly used region for species identification. However, studies of the African malaria vector, An. gambiae complex, reported that mitochondrial markers are not sufficient to distinguish species within the complex25. Similarly, phylogenetic studies of mosquitoes belonging to the Hyrcanus Group have been performed using the cytochrome c oxidase subunit I (COI), cytochrome c oxidase subunit II (COII), and internal transcribed spacer 2 (ITS2) regions. However, the mitochondrial DNA (mtDNA) COI and COII regions are not useful for resolving phylogenetic relationships between An. sinensis, An. kleini, and An. belenrae26,27. The ITS2-based PCR method has been used for distinguishing species in the Maculipennis Group28, e.g., the An. crucians complex29 and others30. Although interspecific differences exist in the ITS2 region in the Hyrcanus Group, differences among An. sinensis, An. kleini, and An. belenrae are not substantial24,27,31,32.
The use of multiple nuclear DNA markers for species identification has an added benefit for investigating hybridization of closely related species33,34. Studies of African malaria vectors utilizing species-specific markers on three different chromosomes reveal a spectrum of degree in hybridization and adaptive introgression accelerating insecticide resistance selective sweep33,34,35. Anopheles kleini and An. belenrae were designated as new species in 20056, and hybridization among An. kleini and An. sinensis are still known to occur36,37. Each genetic region is likely to diverge at different rates, and hence, comparing multiple genetic regions with varying functions will be useful for determining accurate evolutionary relationships38. Methods using multi-locus markers have been successfully used to accurately identify Anopheles species, e.g., members of the An. gambiae complex that is known to be capable of hybridization33,39,40.
Wing Geometric Morphometrics (WGM) is currently being used for the accurate species identification and ecological investigations of insects, and various tools have been developed that can be used by personnel unfamiliar with taxonomy41,42. In particular, the two-dimensional structure of the mosquito wing enables easy collection of landmarks (LMs), and the vein structure of the wing is species-specific making it cost effective and time-efficient compared with traditional methods of identification or using molecular markers42.
Phylogenetic signals are the tendency for closely related species to resemble each other more than evolutionarily distant species. Phylogenetic signals are now widely accepted and used in various ecological and evolutionary studies43. This analysis is being applied to test the congruence of morphometric data and molecular trees and is actively used in wing patterns of various insects44,45.
In this study, we investigated multiple approaches for Anopheles species identification, e.g., the potential use of wing shape and size for Anopheles species identification, in the ROK. The DNA barcoding region, COI, for comparison with WGM data was sequenced. Moreover, the partial sequence of tyrosine hydroxylase (TH) was obtained. This is a nuclear gene associated with the wing melanin pattern in Drosophila (Diptera: Drosophilidae)46, which also has important physiological functions, e.g., as cuticle tanning and innate immunity in Anopheles mosquitoes47. A molecular phylogenetic study of all Anopheles spp. in the ROK using the TH and COI regions in this study together with ITS2 obtained from GenBank was conducted and compared with the results of WGM analysis.
Methods
Sample collection and identification
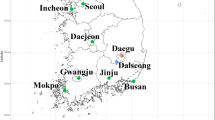
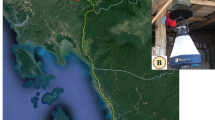
Eight species of Anopheles mosquitoes were collected at six sites [(1) Neutral Nations Supervisory Commission camp (NNSC) (37° 57ʹ17.19″ N, 126° 40ʹ47.91″ E); (2) Daeseong-dong (37° 56ʹ28.31″ N, 126° 40ʹ37.38″ E); (3) South gate entrance to the DMZ (37° 56ʹ03.53″ N, 126° 43ʹ15.46″ E); (4) Camp Bonifas (37° 55ʹ55.25″ N, 126° 43ʹ21.73″ E), (5) Warrior Base training area (37° 55ʹ03.96″ N, 126° 44ʹ29.74″ E); and (6) Dagmar North training area (37° 58ʹ29.85″ N, 126° 50ʹ40.88″ E)] from April to October 2021 in/near the demilitarized zone (DMZ), a malaria high-risk area in the ROK, to reduce seasonal variation among individuals for WGM analysis. Additional collections were also conducted at Yongsan US Army Garrison (USAG) (Seoul) (37° 31ʹ56.2″ N, 126° 58ʹ53.4″ E) and Humphreys USAG (Pyeongtaek) (36° 57ʹ19.9″ N, 127° 01ʹ41.4″ E) (Fig. 1).
Collection sites of Anopheles mosquitoes in the ROK. 1. Neutral Nations Supervisory Commission camp (< 10 m from the DMZ), 2. Daeseong-dong (village inside the DMZ), 3. South gate entrance to the DMZ, 4. Camp Bonifas, 5. Warrior Base training area, 6. Dagmar North training area, 7. Yongsan USAG, 8. Humphreys USAG. This map was generated using Google earth Pro v.9.1.9 (https://earth.google.com) and QGIS 3.26.3 (https://www.qgis.org/ko/site).
Mosquitoes were collected using Mosquito Magnets (Woodstream Corp., Lititz, PA, USA), returned to the central laboratory at Humphreys USAG where they were identified to species (An. lindesayi, An. koreicus, and An. sineroides) or genera, and then stored at − 70 °C. Mosquitoes were then transported on dry ice to Kyungpook National University, Daegu where they were stored at − 70 °C until used, molecular identification was performed using the hind leg24 (Table S1).
Geometric morphometrics analysis
WGM analysis was conducted using the right wings of eight Anopheles species that were identified using molecular methods. The right wing was removed scales removed using a paintbrush, then placed on a slide and then covered with a cover slip using Canada balsam (Duksan, Seoul, ROK). The wings of each Anopheles species were then photographed under magnification (× 20) using an Olympus SZ61 Stereo Microscope (Olympus Corp., Tokyo, Japan). For WGM analysis, 18 landmarks (LMs) were selected using TPSdig2 v.2.3148 (Fig. 2), and the LM data analyzed using R v.4.2.149.
The thin plate spline (TPS) file with LM data was first analyzed using the geomorph v.4.0.5 package50. To generate Procrustes coordinates, the “gpagen” function was used to transform differences in the position and orientation of each data point, and the coordinates were superimposed. In this process, the centroid size (CS), defined as the square root of the sum of squared distances of all the LMs of an object from their centroid, was calculated as a proxy for the wing size of each species51. The allometric effect, which indicates the relationship between the CS and wing shape, was calculated using the “procD.lm” function with 1000 permutations. For statistical comparisons of the mean values of CS for each measured species, an analysis of variance (ANOVA) was performed, and multiple pairwise comparisons applied for the eight Anopheles species using the “LSD.test” function of the agricolae v.1.3-6 package, followed by t-tests (Bonferroni-adjusted p values)52. To determine the accuracy of the grouping of the eight Anopheles species based on wing shape, a linear discriminant analysis (LDA) using the “lda” function in the MASS v.7.3-60 package was performed53. Furthermore, the convex hull algorithm was applied, which is a method for including all data for each species in the scatterplot resulting from LDA.
Canonical variate analysis (CVA) was conducted using the “CVA” function in the Morpho v.2.11 package, and the Mahalanobis distance calculated using a statistical measure of morphological differences in each group54. To verify the accuracy of the reclassification based on wing shape, a pairwise cross-validation test (leave-one-cut method) based on canonical variate analysis was performed using the “CVA” and “typprobClass” function of the Morpho v.2.11 package54. Based on the Mahalanobis distance, a hierarchical clustering dendrogram was constructed using the “pvclust” function in the pvclust v.2.2-0 package55. To check whether the set number of LMs was oversampled, the “lasec” function of the LaMBDA v.0.1.1 package was used56. The morphological differences in the wings of Anopheles mosquitoes were determined by first evaluating the average wing shape of the eight species and then evaluating the average shape of each species, followed by visualization using the “plotRefToTarget” (method = points) function of the geomorph v.4.0.5package50. The results of the analysis using WGM were visualized using the ggplot2 v.3.4.2 package57.
DNA sequencing
Genomic DNA was extracted from the entire body, except the right wing and hind legs, of the eight Anopheles species for COI and TH. DNA extraction was performed using the Clear-S™ Quick DNA Extraction Kit (InVirusTech, Gwangju, ROK) according to the manufacturer’s protocol. Universal primer pairs (LCO1490: 5′-GGT CAA CAA ATC ATA AAG ATA TTG G-3′/HCO2198: 5′-TAA ACT TCA GGG TGA CCA AAA AAT CA-3′) were used to amplify the COI region58. The reaction mixture for PCR amplification (total: 25 μL) consisted of 1× PCR buffer, 0.2 mM dNTPs, 1.5 mM MgCl2, 0.4 μM of each primer, 0.5 units Taq DNA polymerase (TaKaRa, Shiga, Japan), and 1 μL extracted genomic DNA. The PCR cycling conditions were as follows: initial denaturation at 94 °C for 5 min followed by 30 cycles at 94 °C for 30 s, 56 °C for 30 s, 72 °C for 1 min, and then a final extension at 72 °C for 5 min.
Universal primers for the TH gene region were constructed using the sequences of An. sinensis (Genbank accession numbers: KU886220, AXCK02010477) and An. stephensi (Genbank accession number: CP032302) registered in GenBank. Primers were designed to enable the combined amplification of the exon and intron regions (Kan_th_F: 5ʹ-GGT TCA ACA GAC AGT CGA GG-3′/Kan_th_R: 5ʹ-GTT GAC GTT GTT CTC CGA CA-3′; Kan_th_F2: 5ʹ-CTG CCC CAG AAG CCA GAG-3′/Kan_th_R2: 5ʹ-CTG CAA CAC AAC CTC CTC CT-3′). The reaction mixture for PCR amplification (total: 25 μL) consisted of 1 × PCR buffer, 0.2 mM dNTPs, 1.5 mM MgCl2, 0.4 μM of each primer, 0.5 units Taq hotstart DNA polymerase (TaKaRa, Shiga, Japan), and 1 μL extracted genomic DNA. The PCR cycling conditions were as follows: initial denaturation at 94 °C for 5 min followed by 35 cycles at 94 °C for 30 s, 58 ~ 60 °C for 40 s, 72 °C for 1 min, and then a final extension at 72 °C for 10 min.
To confirm whether the PCR amplification was successful, the amplification products were identified by electrophoresis on a 1.5% agarose gel, and the successfully amplified samples were subjected to Sanger dideoxy sequencing in both directions by Macrogen (Daejeon, ROK).
The sequences for each specimen were aligned and edited using BioEdit59 and then compared and confirmed using NCBI’s Basic Local Alignment Search Tool. The sequences for the COI and TH genes of the eight Anopheles species obtained in this study were deposited in the GenBank (GenBank accession numbers (COI): OR150355-OR150394; GenBank accession numbers (TH): OR187248–OR187297).
Phylogenetic analysis
Three regions (COI, TH, and ITS2) were analyzed for the molecular phylogenetic analysis (COI: 615 bp; TH: 656 bp; ITS2: 538 bp). The registered sequences in GenBank were used for the analysis of the ITS2 region (Table S2). Sequence alignments for each region of the eight Anopheles species were made using the L-INS-i method in MAFFT v.760. Next, a maximum likelihood (ML) tree was generated for the phylogenetic analysis of each sequence. The R’s phangorn v.2.11.1 and ape v. 5.7-1 packages were used for nucleotide substitution model selection and bootstrap analysis for each locus61,62. After checking with the “modelTest” function, COI: GTR + G (4) + I; ITS2, TH: GTR + G (4) were found to be the best fit models. The “bootstrap.pml” function with 1000 replications were applied to check the support for a node in the tree. Consensus sequences for each species were generated using the “ConsensusSequence” function in the DECIPHER v.2.24.0 package63. The relative nucleotide diversity (π) for the exon and intron of the TH gene using the “nuc.div” function of the pegas v.1.2 package was also calculated64. The results obtained from this phylogenetic analysis were visualized using the ggtree v.3.4.4 and ggmsa v.1.3.4 packages65,66.
In addition, evolutionary phylogenetic inferences about the CS and wing shape of the eight Anopheles species analyzed by WGM were drawn using the geomorph v.4.0.5 and phytools v.1.5-1 packages50,67, for which a phylogenetic tree based on the TH gene region of this study was used. According to the Brownian model of evolutionary change, species diverge over time, and the similarity or dissimilarity of variation between species can be deduced by the phylogenetic signal68. To identify these phylogenetic signals, the “physignal” function in geomorph v.4.0.5 package with 10,000 permutations was used to determine whether a phylogenetic signal existed in the Procrustes shape variables50. To visualize the quantitative morphometric data in a phylogenetic context, the phylogenetic signals for CS and wing shape were projected onto a phylogenetic tree using the “contMap” function in phytools v.1.5.-1 package and “gm.prcomp” function in geomorph v.4.0.5 package, respectively50,67.
Results
Mosquitoes were collected over several months for WGM analysis to minimize seasonal variation in their wings. First, the wing shape and CS exhibited small but significant relationships (R2 = 1.8%, P = 0.0006). However, because wing size can also be useful for species identification, the data obtained were used in further analyses. Comparisons of wing size as a proxy for CS demonstrated differences in mean CS between groups of Anopheles mosquitoes of each species (ANOVA, F = 8.234, P < 0.001). Among the eight Anopheles species, An. lesteri had the largest CS, whereas An. lindesayi had the smallest CS (Fig. 3). However, when analyzed by pairwise comparisons, no significant size differences were observed in CS for most species (Table S3).
Boxplot showing the results of centroid size comparisons for each Anopheles species present in the ROK. Lin, Hyr, and Bar refer to the Lindesayi Group, Hyrcanus Group, and Barbirostris Group, respectively. The box represents the first and third quartiles, and the black line in the middle represents the median. Black stars indicate outliers.
LDA analysis of wing shape data based on each group confirmed that the Hyrcanus and Barbirostris Groups, which are known to be more closely related than the Lindesayi Group, were in close proximity based on the scatterplot (Fig. 4a). In the scatterplot representing the results of the analysis at the species level, a strong overlap between An. sinensis, An. belenrae, and An. kleini was observed (Fig. 4b). When the average wing shape of the eight species was evaluated and compared (Fig. 4c), six mosquito species belonging to the Hyrcanus Group showed variations primarily between LMs 17 and 18. However, the differences among members of the Hyrcanus Group were not large compared with the average morphological differences of all eight species, whereas An. koreicus (Barbirostris Group) and An. lindesayi (Lindesayi Group) exhibited large differences in LMs 10 and 11, LMs 15 and 16, and LMs 17 and 18. The pairwise cross-validation test (leave-one-cut method) was performed to evaluate the reclassification accuracy, and the results showed a high level of accuracy at the group level [Overall accuracy: 94.4% (Hyrcanus Group: 94.9%; Barbirostris Group: 88.2%; Lindesayi Group: 100%)]. In contrast to the comparison at the group level, the pairwise cross-validation test at the species level showed a lower reclassification rate (Overall accuracy: 75.8%). Results of the analysis at the species level, An. pullus could be recognized with 88.2% accuracy, and An. sineroides could be identified with 80.9% accuracy. However, An. belenrae and An. lesteri, which belong to the Hyrcanus Group, showed the lowest classification accuracy with 57.6% and 68.1% accuracy, respectively (Table 1). This was observed especially for An. belenrae where many specimens were misidentified as An. sinensis or An. kleini. The LMs for potential oversampling were checked and the 18 LMs established for WGM analysis were not oversampled (fit = 0.90: 14 LMs; fit = 0.95: 16 LMs; fit = 0.99: 17 LMs) (Fig. S1).
(a) Scatterplot of Linear Discriminant Analysis (LDA) results based on wing shape variation from the three groups (LD1: 65%, LD2: 35%). (b) Scatterplot of LDA results based on wing shape variation from the eight Anopheles species (LD1: 38%, LD2: 25%). (c) Comparison of mean shapes for Anopheles mosquitoes. The gray line is the average shape for all eight species, and the black line is the average shape for each species (magnified by 3×).
An ML tree was generated for molecular phylogenetic inference using the COI regions of the eight Anopheles species in the ROK obtained during this study (Fig. 5a). Consistent with previous results26, analyses using COI reaffirmed that the relationships of An. sinensis, An. belenrae, and An. kleini in the Hyrcanus Group were unclear to resolve. However, using the ITS2 region, all eight species were clearly separated, including An. sinensis, An. belenrae, and An. kleini, which were not accurately identified in the analysis using the COI region (Fig. 5b). For the tree based on the TH region, the topology did not match the tree constructed based on the ITS2 region, but all eight species were separated (Fig. 5c). In particular, interspecific differences were evident primarily in the intron region of the TH gene (Fig. 6a), whereas no substantial sequence differences were observed in the exon region (Fig. 6b). The nucleotide diversity for the exon and intron TH gene was evaluated, respectively, and found that the value of π in the exon was 0.0204 (n = 50), whereas that in the intron was 0.1699 (n = 50), indicating much larger nucleotide diversity in the intron.
Simplified maximum likelihood (ML) phylogenetic trees constructed in this study using COI, ITS2, and TH gene fragments (SIN = An. sinensis, BEL = An. belenrae, KLE = An. kleini, LES = An. lesteri, SNR = An. sineroides, PUL = An. pullus, KOR = An. koreicus, LIN = An. lindesayi). (a) ML tree based on COI regions [GTR + G (4) + I)]. (b) ML tree based on ITS2 regions [GTR + G (4)]. (c) ML tree based on TH gene [GTR + G (4)]. Bootstrap was performed with 1000 replications, and the value is represented on the node in red. Unresolved clustering is highlighted in red. Hyr, Lin and Bar refer to the Lindesayi Group, Hyrcanus Group, and Barbirostris Group, respectively.
(a) Multiple nucleotide sequence alignments of the TH gene fragment (red: A, blue: C, green: T, yellow: G, black: ambiguous nucleotide). Red boxes represent intron regions, and blue boxes represent exon regions. (b) Multiple amino acid sequence alignments of TH gene. Red circles indicate areas of variation.
After analysis of the phylogenetic signal for wing shape, we found a strong phylogenetic signal in shape (K = 1.4, P = 0.01). A strong phylogenetic signal indicates that the wing shape evolved from a shared evolutionary history, rather than environmental factors43,68,69. As illustrated by the phylomorphospace formed based on wing shape (PC1: 61.94%, PC2: 22.85%), the six species belonging to the Hyrcanus Group were closely positioned to each other, whereas An. koreicus and An. lindesayi, which belong to different groups, are far diverged from each other in the phylomorphospace (Fig. 7a). In contrast to wing shape, no significant phylogenetic signal was observed in the CS (K = 0.41, P = 0.31) (Fig. 7b). In the hierarchical clustering dendrogram, An. lesteri and An. pullus clustered together, and the other four species in the Hyrcanus Group clustered together (Table S4, Fig. 7c). For An. koreicus and An. lindesayi, which do not belong to the Hyrcanus Group, a separate branch was formed.
(a) Phylomorphospace based on wing shapes of the eight Anopheles species present in the ROK (PC1: 61.94%, PC2: 22.85%). (b) CS evolution of Anopheles mosquitoes in the ROK. Wing shape and CS were projected onto the TH gene tree constructed in this study. (c) Dendrogram based on Mahalanobis distance between the eight Anopheles species in the ROK. Values at branches are approximately unbiased (au) p-value (red), bootstrap probabilities (bp) (green) with 1000 bootstrap replicates.
Discussion
Anopheles specimens collected by Mosquito Magnet traps during the mosquito season from April–October were used for WGM analysis. Our results showed that CS was not a useful phenotypical trait for the identification of Anopheles species in the ROK. The CS did not demonstrate any statistically significant differences for most of the Anopheles species present in the ROK. However, wing size is highly influenced by environmental factors, e.g., temperature and humidity. Therefore, caution is necessary when interpreting CS results using WGM analysis70,71.
LDA analysis based on wing shape confirmed that each Anopheles mosquito demonstrates species-specific venation at the group level (Hyrcanus Group vs. Barbirostris Group vs. Lindesayi Group). Each mosquito species showed a considerable amount of variation, primarily in LM 15–18. In this study, LM15 is located in radius 4 plus 5, LM16, 17 are located in radius 2 plus 3, and LM18 is located in media 1 plus 272. A study comparing the wings of An. dirus and An. baimaii73 and the An. barbirostris complex74 found differences primarily in radius 2 plus 3 and media 1 plus 2, similar to our results. Based on the results of this study and previous studies73,74, it appears that the variation in LM is mainly found in radius 2 plus 3 and media 1 plus 2, which will need to be evaluated among more Anopheles species in the future. Wing venation is an important marker for species identification and has been shown to exert potentially important aerodynamic effects in insect wings75,76,77. In the case of mosquitoes, each species exhibits different ecological characteristics (endophilic or exophilic; stenogamous or eurygamous) with large differences in host preference (e.g., mammal- or avian-feeding). Different ecological characteristics of each species may have exerted selective pressure on wing venation, which was reflected in the eight Anopheles spp. that exhibited differences in venation. However, it is difficult to determine whether the variation in wing shape at the group and species level is due to ecological features since there is a lack of ecological research data on Anopheles mosquitoes present in the ROK. Future studies that link morphological data, such as WGM, to ecological features will provide more information. At the present time, only one species each of the Lindesayi and Baribirostris Groups of mosquitoes is known to occur in the ROK7,12,13,14. Therefore, further studies should include more mosquitoes from the Lindesayi and Barbirostris Groups.
In this study, data were obtained for the TH and COI gene fragments and a molecular phylogeny was inferred using the three regions (TH, COI, and ITS2). Based on COI, the mosquito species in each of the three groups were monophyletic, but it was difficult to determine the phylogenetic relationships for An. sinensis, An. kleini, and An. belenrae in the Hyrcanus Group. In the case of An. sinensis and An. kleini, hybridization occurred under the experimental conditions, and similarly, An. sinensis and An. kleini, as well as An. belenrae, might have experienced ancient hybridization36,37. Unlike the analysis based on the mtDNA COI region, the analysis based on nuclear DNA ITS2 and TH gene fragments resolved all three species (An. sinensis, An. kleini, and An. belenrae). The COI region is actively used as a barcoding region for species identification78, and it is generally believed that the mitochondrial genome diverges at a faster rate than the nuclear genome79. However, the analysis based on TH + ITS2 (nuclear DNA) and COI (mtDNA) in this study and previous studies for members of the Hyrcanus Group suggests a contradiction in some species belonging to the Hyrcanus Group. Some reports suggest that these results are due to male-biased dispersal26,27,32,80, but whether the lower diversity of the mitochondrial genome than that of the nuclear genome is due to male-biased dispersal needs verification using various genetic tools and field observations81. Moreover, Anopheles mosquitoes within the Hyrcanus Group present in the ROK had an inconsistent topology between the TH and ITS2 trees, which is likely due to different rates of divergence for different genes that are dependent upon selection pressures38,82. For closely related species, such as An. sinensis, An. kleini, and An. belenrae, genome-wide data will be required to establish true phylogenetic tree.
For the TH gene fragment used for phylogenetic analysis, there was much greater nucleotide diversity in the intron regions than in exon regions. Currently, various nuclear genes, except COI and ITS2, are considered genetic markers to distinguish Anopheles species83,84. However, the TH gene fragment may also be used as a molecular marker for members of the Hyrcanus Group in the ROK. Conserved regions of exon are ideal for PCR primers, and the high divergence of intron sites may be useful for determining the phylogenetic relationships.
The hierarchical clustering dendrogram constructed using Mahalanobis distance based on wing shape, demonstrated similar results compared to molecular phylogeny. This was especially observed in the clustering of closely related species, e.g., An. sinensis, An. kleini, and An. belenrae, in the dendrogram and similar to the tree created using the ITS2 and TH gene fragments. Phylogenetic analysis using morphometric data is generally recognized to have some limitations compared to phylogenetic trees using molecular data, as studies have demonstrated that there is incongruence between molecular and morphometric analyses caused by homoplasy85. The analysis of wing shape used in our study demonstrated similar results as the molecular analysis for some species. Whether this pattern extends to other morphological features, as well as wings, will need to be compared with additional morphometrics data. No significant phylogenetic signals were identified in the CS of the eight Anopheles spp. used in this study. However, there was a strong phylogenetic signal identified using wing shape, which was confirmed by clustering of each group using phylomorphospace. These data indicate that closely related species have similar wing shapes compared with their distant relatives68,86. However, the presence of a strong phylogenetic signals in wing shapes does not mean that morphological comparisons using wing shapes can be used to make accurate phylogenetic inferences85,87,88. Consequently, further analysis of various anatomical features is necessary to validate the congruence between morphometric data and molecular phylogenetic analysis.
Conclusion
Currently, there are more than 200 annual cases of vivax malaria in the ROK, mostly near the DMZ in Gyeonggi and Gangwon provinces adjacent to the Democratic People’s Republic of Korea (DPRK)89,90. Anopheles kleini is more frequently collected in human environments89. Successful disease vector mosquito control requires accurate species identification. For this purpose, a comparative analysis of all Anopheles mosquito species in the ROK through quantitative morphological and molecular phylogenetic analysis was conducted. This study confirmed that WGM can be used to identify with high accuracy at the group level but not sufficient for species identification. These results may assist a wide variety of personnel who are unfamiliar with traditional mosquito taxonomy or have difficulty with molecular approaches. The study also identified the TH gene as a candidate for a new genetic marker in addition to previously used COI and ITS2 molecular markers. TH gene genealogy was different from ITS2 genealogy and may be useful for phylogenetic studies for various Anopheles mosquito species where it is difficult to analyze the COI and ITS2 regions alone. The inclusion of more Anopheles species, not included in this study, would allow for more rigorous evolutionary comparisons and inferences between morphometric and molecular data.
Data availability
All sequencing data used in this research were deposited in the GenBank https://www.ncbi.nlm.nih.gov/under the accession numbers (COI: OR150355-OR150394; TH: OR187248-OR187297).
References
World Health Organization (WHO). WHO malaria report. 2022 https://www.who.int/publications/i/item/9789240064898 (2022).
Savi, M. K. An overview of malaria transmission mechanisms, control, and modeling. Med. Sci. 11, 3 (2023).
Bahk, Y. Y. et al. Epidemiological characteristics of re-emerging vivax malaria in the Republic of Korea (1993–2017). Korean J. Parasitol. 56, 53 (2018).
Korea Disease Control and Prevention Agency (KDCA). Malaria management guideline. https://kdca.go.kr/board/board.es?mid=a20507020000&bid=0019&act=view&list_no=721868&tag=&nPage=1 (2023).
Kim, Y. H. et al. Recent spatial and temporal trends of malaria in Korea. Korean J. Parasitol. 59, 585–593 (2021).
Rueda, L. M. Two new species of Anopheles (Anopheles) Hyrcanus group (Diptera: Culicidae) from the Republic of South Korea. Zootaxa 941, 1–26 (2005).
Foley, D. H. et al. Geographic distribution and ecology of potential malaria vectors in the Republic of Korea. J. Med. Entomol. 46, 680–692 (2009).
Ree, H. I. Studies on Anopheles sinensis, the vector species of vivax malaria in Korea. Korean J. Parasitol. 43, 75–92 (2005).
Joshi, D. et al. The susceptibility of Anopheles lesteri to infection with Korean strain of Plasmodium vivax. Malar. J. 8, 42 (2009).
Joshi, D., Kim, J. Y., Choochote, W., Park, M. H. & Min, G. S. Preliminary vivax malaria vector competence for three members of the Anopheles Hyrcanus Group in the Republic of Korea. J. Am. Mosq. Control Assoc. 27, 312–314 (2011).
Ubalee, R. et al. Vector competence of Anopheles kleini and Anopheles sinensis (Diptera: Culicidae) from the Republic of Korea to vivax malaria-infected blood from patients from Thailand. J. Med. Entomol. 53, 1425–1432 (2016).
Foley, D. H. et al. Mosquito species composition and Plasmodium vivax infection rates on Baengnyeong-do (Island), Republic of Korea. Korean J. Parasitol. 49, 313–316 (2011).
Yoo, D. H. et al. Mosquito species composition and Plasmodium vivax infection rates for Korean Army bases near the demilitarized zone in the Republic of Korea, 2011. Am. J. Trop. Med. Hyg. 88, 24–28 (2013).
Lee, S. Y. et al. Species diversity of Anopheles mosquitoes and Plasmodium vivax infection rates, Gyeonggi Province, Republic of Korea during 2020. J. Med. Entomol. 59, 1778–1786 (2022).
Tainchum, K., Kongmee, M., Manguin, S., Bangs, M. J. & Chareonviriyaphap, T. Anopheles species diversity and distribution of the malaria vectors of Thailand. Trends Parasitol. 31, 109–119 (2015).
Apiwathnasor, C., Prommongkol, S., Samung, Y., Limrat, D. & Rojruthai, B. Potential for Anopheles campestris (Diptera: Culicidae) to transmit malaria parasites in Pa Rai subdistrict (Aranyaprathet, Sa Kaeo Province), Thailand. J. Med. Entomol. 39, 583–586 (2002).
Kang, S., Jung, J., Lee, S., Hwang, H. & Kim, W. The polymorphism and the geographical distribution of the knockdown resistance (kdr) of Anopheles sinensis in the Republic of Korea. Malar. J. 11, 151 (2012).
Rueda, L. M. et al. Species composition, larval habitats, seasonal occurrence and distribution of potential malaria vectors and associated species of Anopheles (Diptera: Culicidae) from the Republic of Korea. Malar. J. 9, 55 (2010).
Kim, H. C. et al. Overwintering of Anopheles lindesayi japonicus larvae in the Republic of Korea. J. Am. Mosq. Control Assoc. 25, 32–37. https://doi.org/10.2987/08-5781.1 (2009).
Ree, H.-I. Taxonomic review and revised keys of the Korean mosquitoes (Diptera: Culicidae). Entomol. Res. 33, 39–52 (2003).
Dahan-Moss, Y. et al. Member species of the Anopheles gambiae complex can be misidentified as Anopheles leesoni. Malar. J. 19, 89 (2020).
Ruiz-Lopez, F. et al. Systematics of the oswaldoi complex (Anopheles, Nyssorhynchus) in South America. Parasites Vectors 6, 324 (2013).
Joshi, D., Park, M. H., Saeung, A., Choochote, W. & Min, G. S. Multiplex assay to identify Korean vectors of malaria. Mol. Ecol. Resour. 10, 748–750 (2010).
Bang, W. J. et al. Multiplex PCR assay for the identification of eight Anopheles species belonging to the Hyrcanus, Barbirostris and Lindesayi groups. Malar. J. 20, 287 (2021).
Hanemaaijer, M. J. et al. Mitochondrial genomes of Anopheles arabiensis, An. gambiae and An. coluzzii show no clear species division. F1000Research 7, 347 (2018).
Fang, Y., Shi, W. Q. & Zhang, Y. Molecular phylogeny of Anopheles hyrcanus group (Diptera: Culicidae) based on mtDNA COI. Infect. Dis. Poverty 6, 1–10 (2017).
Zhang, C. L. et al. Molecular phylogeny of the Anopheles Hyrcanus group (Diptera: Culicidae) based on rDNA-ITS2 and mtDNA-COII. Parasites Vectors 14, 454 (2021).
Hodge, J. M. et al. The new internal transcribed spacer 2 diagnostic tool clarifies the taxonomic position and geographic distribution of the North American malaria vector Anopheles punctipennis. Malar. J. 20, 141 (2021).
Wilkerson, R. C., Reinert, J. F. & Li, C. Ribosomal DNA ITS2 sequences differentiate six species in the Anopheles crucians complex (Diptera: Culicidae). J. Med. Entomol. 41, 392–401 (2004).
Hackett, B. J. et al. Ribosomal DNA internal transcribed spacer (ITS2) sequences differentiate Anopheles funestus and An. rivulorum, and uncover a cryptic taxon. Insect Mol. Biol. 9, 369–374 (2000).
Li, C. et al. A newly recognized species in the Anopheles Hyrcanus Group and molecular identification of related species from the Republic of South Korea (Diptera: Culicidae). Zootaxa 939, 1–8 (2005).
Fang, Y., Shi, W. Q. & Zhang, Y. Molecular phylogeny of Anopheles hyrcanus group members based on ITS2 rDNA. Parasites Vectors 10, 1–11 (2017).
Lee, Y. et al. Spatiotemporal dynamics of gene flow and hybrid fitness between the M and S forms of the malaria mosquito, Anopheles gambiae. Proc. Natl. Acad. Sci. U. S. A. 110, 19854–19859 (2013).
Norris, L. C. & Norris, D. E. Phylogeny of anopheline (Diptera: Culicidae) species in southern Africa, based on nuclear and mitochondrial genes. J. Vector. Ecol. 40, 16–27 (2015).
Main, B. J. et al. Complex genome evolution in Anopheles coluzzii associated with increased insecticide usage in Mali. Mol. Ecol. 24, 5145–5157 (2015).
Joshi, D., Choochote, W. & Min, G. S. Short report: Natural hybrid between Anopheles kleini and Anopheles sinensis. Am. J. Trop. Med. Hyg. 81, 1020–1022 (2009).
Choochote, W. et al. Evidence to support natural hybridization between Anopheles sinensis and Anopheles kleini (Diptera: Culicidae): Possibly a significant mechanism for gene introgression in sympatric populations. Parasites Vectors 7, 36 (2014).
Graur, D., Sater, A. K. & Cooper, T. F. Molecular and Genome Evolution (Sinauer Associates Inc., 2016).
Weetman, D. et al. Contemporary gene flow between wild An. gambiae s.s. and An. arabiensis. Parasite Vectors 7, 345 (2014).
Norris, L. C. et al. Adaptive introgression in an African malaria mosquito coincident with the increased usage of insecticide-treated bed nets. Proc. Natl. Acad. Sci. U. S. A. 112, 815–820 (2015).
Bookstein, F. L. Foundations of morphometrics. Annu. Rev. Ecol. Syst. 13, 451–470 (1982).
Lorenz, C. et al. Geometric morphometrics in mosquitoes: What has been measured?. Infect. Genet. Evol. 54, 205–215 (2017).
Revell, L. J., Harmon, L. J. & Collar, D. C. Phylogenetic signal, evolutionary process, and rate. Syst. Biol. 57, 591–601 (2008).
Muñoz-muñoz, F. et al. Phenotypic differentiation and phylogenetic signal of wing shape in western European biting midges, Culicoides spp., of the subgenus Avaritia. Med. Vet. Entomol. 28, 319–329 (2014).
dos Santos, C. F., Souza dos Santos, P. D., Marques, D. M., da-Costa, T. & Blochtein, B. Geometric morphometrics of the forewing shape and size discriminate Plebeia species (Hymenoptera: Apidae) nesting in different substrates. Syst. Entomol. 44, 787–796 (2019).
True, J. R., Edwards, K. A., Yamamoto, D. & Carroll, S. B. Drosophila wing melanin patterns form by vein-dependent elaboration of enzymatic prepatterns. Curr. Biol. 9, 1382–1391 (1999).
Qiao, L. et al. Tyrosine hydroxylase is crucial for maintaining pupal tanning and immunity in Anopheles sinensis. Sci. Rep. 6, 29835 (2016).
Rohlf, F. J. The tps series of software. Hystrix 26, 9–12 (2015).
Ihaka, R. & Gentleman, R. R: A language for data analysis and graphics. J. Comput. Graph. Stat. 5, 299–314 (1996).
Adams, D. C. & Otárola-Castillo, E. geomorph: An R package for the collection and analysis of geometric morphometric shape data. Methods Ecol. Evol. 4, 393–399 (2013).
Bookstein, F. L. Morphometric Tools for Landmark Data: Geometry and Biology (Cambridge University Press, 1992).
De Mendiburu, F. & Simon, R. Agricolae: Ten years of an open source statistical tool for experiments in breeding, agriculture and biology. PeerJ 3, e1404v1401 (2015).
Venables, W. N. & Ripley, B. D. Modern Applied Statistics with S-PLUS (Springer Science & Business Media, 2013).
Schlager, S. Morpho and Rvcg-shape analysis in R: R-packages for geometric morphometrics, shape analysis and surface manipulations. In Statistical Shape and Deformation Analysis 217–256 (Elsevier, 2017).
Suzuki, R. & Shimodaira, H. Pvclust: An R package for assessing the uncertainty in hierarchical clustering. Bioinformatics 22, 1540–1542 (2006).
Watanabe, A. How many landmarks are enough to characterize shape and size variation?. PLoS ONE 13, e0198341 (2018).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2016).
Folmer, O., Black, M., Hoeh, W., Lutz, R. & Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 3, 294–299 (1994).
Hall, T. A. Bioedit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/ NT. Nucleic Acids Symp. Ser. 41, 95–98 (1999).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Schliep, K. P. Phangorn: Phylogenetic analysis in R. Bioinformatics 27, 592–593 (2011).
Paradis, E. & Schliep, K. Ape 5.0: An environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35, 526–528 (2019).
Wright, E. S. Using DECIPHER v2.0 to analyze big biological sequence data in R. R J. 8(1), 352–359 (2016).
Paradis, E. Pegas: An R package for population genetics with an integrated–modular approach. Bioinformatics 26, 419–420 (2010).
Yu, G., Smith, D. K., Zhu, H., Guan, Y. & Lam, T. T. Y. ggtree: An R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 8, 28–36 (2017).
Zhou, L. et al. Ggmsa: A visual exploration tool for multiple sequence alignment and associated data. Brief. Bioinformatics 23, bbac222 (2022).
Revell, L. J. Phytools: An R package for phylogenetic comparative biology (and other things). Methods Ecol. Evol. 2, 217–223 (2012).
Revell, L. J. & Harmon, L. J. Phylogenetic Comparative Methods in R (Princeton University Press, 2022).
Blomberg, S. P. & Garland, T. Jr. Tempo and mode in evolution: Phylogenetic inertia, adaptation and comparative methods. J. Evol. Biol. 15, 899–910 (2002).
Gómez, G. F., Márquez, E. J., Gutiérrez, L. A., Conn, J. E. & Correa, M. M. Geometric morphometric analysis of Colombian Anopheles albimanus (Diptera: Culicidae) reveals significant effect of environmental factors on wing traits and presence of a metapopulation. Acta Trop. 135, 75–85 (2014).
Phanitchat, T. et al. Geometric morphometric analysis of the effect of temperature on wing size and shape in Aedes albopictus. Med. Vet. Entomol. 33, 476–484 (2019).
Harbach, R. E. & Knight, K. L. Taxonomists’ Glossary of Mosquito Anatomy (Plexus Publishing Inc., 1980).
Chaiphongpachara, T. et al. Geometric morphometrics versus DNA barcoding for the identification of malaria vectors Anopheles dirus and An. baimaii in the Thai-Cambodia border. Sci. Rep. 12, 13236 (2022).
Chaiphongpachara, T. et al. Geometric morphometric and molecular techniques for discriminating among three cryptic species of the Anopheles barbirostris complex (Diptera: Culicidae) in Thailand. Heliyon 8, e11261 (2022).
Mountcastle, A. M. & Combes, S. A. Wing flexibility enhances load-lifting capacity in bumblebees. Proc. R. Soc. B 280, 20130531 (2013).
Perrard, A., Baylac, M., Carpenter, J. M. & Villemant, C. Evolution of wing shape in hornets: Why is the wing venation efficient for species identification?. J. Evolut. Biol. 27, 2665–2675 (2014).
Salcedo, M. K. & Socha, J. J. Circulation in insect wings. Integr. Comp. Biol. 60, 1208–1220 (2020).
Hebert, P. D. N. et al. Biological identifications through DNA barcodes. Proc. R. Soc. B 270, 313–321 (2003).
Moore, W. S. Inferring phylogenies from mtdna variation: Mitochondrial-gene trees versus nuclear-gene trees. Evolution 49, 718–726 (1995).
Petit, R. J. & Excoffier, L. Gene flow and species delimitation. Trends Ecol. Evol. 24, 386–393 (2009).
Prugnolle, F. & de Meeus, T. Inferring sex-biased dispersal from population genetic tools: A review. Heredity 88, 161–165 (2002).
Weedall, G. D. et al. An Africa-wide genomic evolution of insecticide resistance in the malaria vector Anopheles funestus involves selective sweeps, copy number variations, gene conversion and transposons. PLoS Genet. 16, e1008822 (2020).
Besansky, N. J. & Fahey, G. T. Utility of the white gene in estimating phylogenetic relationships among mosquitoes (Diptera: Culicidae). Mol. Biol. Evol. 14, 442–454 (1997).
Gholizadeh, S. et al. The Anopheles stephensi odorant binding protein 1 (AsteObp1) gene: A new molecular marker for biological forms diagnosis. Acta Trop. 146, 101–113 (2015).
Klingenberg, C. P. & Gidaszewski, N. A. Testing and quantifying phylogenetic signals and homoplasy in morphometric data. Syst. Biol. 59, 245–261 (2010).
Adams, D. C. A generalized K statistic for estimating phylogenetic signal from shape and other high-dimensional multivariate data. Syst. Biol. 63, 685–697 (2014).
Klingenberg, C. P. & Monteiro, L. R. Distances and directions in multidimensional shape spaces: Implications for morphometric applications. Syst. Biol. 54, 678–688 (2005).
Klingenberg, C. P. Morphological integration and developmental modularity. Annu. Rev. Ecol. Evol. Syst. 39, 115–132 (2008).
Chang, K. S. et al. Distribution of malaria vectors and incidence of vivax malaria at Korean army installations near the demilitarized zone, Republic of Korea. Malar. J. 15, 259 (2016).
Chai, J. Y. History and current status of malaria in Korea. Infect Chemother. 52, 441–452 (2020).
Acknowledgements
The authors thank the Government-wide R&D Fund project for infectious disease research (GFID), Republic of Korea (grant number: HG18C0046), and the Armed Forces Health Surveillance Division, Global Emerging Infections Surveillance (GEIS) Branch, ProMIS ID P0016_21_ME. We thank the members of the Force Health Protection and Preventive Medicine, Medical Department Activity-Korea (Mr Sung-Tae Chong and Ms Myung-Sun Kim) for their support.
Disclaimer
The opinions or assertions contained herein are the private views of the authors and are not to be construed as official or reflecting the true views of the U.S. Department of the Army, U.S. Department of Defense, or the U.S. Government. This work was prepared in accordance with Title 17, U.S.C., §101 that defines a U.S. Government work as work prepared by a military service member or employee of the U.S. Government (TA Klein and HC Kim) as part of that person’s official duties.
Author information
Authors and Affiliations
Contributions
J.J. and K.S.C. designed the study. J.J. analyzed the data and drafted the manuscript. H.C.K. and T.A.K. collected mosquitoes and helped draft the manuscript. K.S.C. helped draft the manuscript with analysis of the data. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jeon, J., Kim, H.C., Klein, T.A. et al. Analysis of geometric morphometrics and molecular phylogeny for Anopheles species in the Republic of Korea. Sci Rep 13, 22009 (2023). https://doi.org/10.1038/s41598-023-49536-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-49536-w
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.