Abstract
Otoliths are calcium carbonate components of the stato-acoustical organ responsible for hearing and maintenance of the body balance in teleost fish. During their formation, control over, e.g., morphology and carbonate polymorph is influenced by complex insoluble collagen-like protein and soluble non-collagenous protein assemblages; many of these proteins are incorporated into their aragonite crystal structure. However, in the fossil record these proteins are considered lost through diagenetic processes, hampering studies of past biomineralization mechanisms. Here we report the presence of 11 fish-specific proteins (and several isoforms) in Miocene (ca. 14.8–14.6 Ma) phycid hake otoliths. These fossil otoliths were preserved in water-impermeable clays and exhibit microscopic and crystallographic features indistinguishable from modern representatives, consistent with an exceptionally pristine state of preservation. Indeed, these fossil otoliths retain ca. 10% of the proteins sequenced from modern counterparts, including proteins specific to inner ear development, such as otolin-1-like proteins involved in the arrangement of the otoliths into the sensory epithelium and otogelin/otogelin-like proteins that are located in the acellular membranes of the inner ear in modern fish. The specificity of these proteins excludes the possibility of external contamination. Identification of a fraction of identical proteins in modern and fossil phycid hake otoliths implies a highly conserved inner ear biomineralization process through time.
Similar content being viewed by others
Introduction
Paleoproteomics is an accelerating research field providing new perspectives on, e.g., the evolution of biomineralization processes through time and refining our understanding of fossil remains1. While studies of ancient DNA are limited to few million years because DNA degrades relatively fast after cell death2, the study of more stable protein remains in the fossil record offers an opportunity to explore protein function and their evolution over geological time scales; several to hundreds millions of years3. Paleoproteomic studies of biominerals such as bones, teeth, and shells are particularly promising because such structures have high potential of preserving protein residues embedded within well-preserved fossil specimens. Here, we explore this potential in fossilized calcium carbonate structures of the inner ear of teleost fish (otoliths). Fish otoliths are, in contrast to osteological fish remains, frequently found in the fossil record during the Mesozoic, and with increasing abundance in Cenozoic strata4. Due to their taxon-specific morphology, these fossils are key to the interpretation of palaeobiodiversity of fish and to palaeoenvironmental reconstructions based on their isotope- and trace element compositions5. Although otolith calcium carbonate mineral structures can resemble inorganically precipitated aggregates of crystals, they are in fact complex organic-mineral composites, akin to many other biogenic carbonates6. Studies of the modern otolith biomineralization process have shown that the organic macromolecules, proteins in particular, control key aspects of otolith formation7, including regulation of calcium transport, nucleation, and saturation state at the crystallization site, thus actively modulating aragonite crystal growth8. Indeed, the strict selection of a specific calcium carbonate polymorph (aragonite as opposed to, e.g., calcite or vaterite) has also been shown to be controlled by proteins, such as aspartic acid and serine residues, that attract calcium cations to the growing crystal surface and favor denser (i.e., aragonitic) packing of ions9. To date, several hundred proteins have been identified in modern fish otoliths, many of which are thought to be directly involved in biomineralization10. Proteins known to be involved in otolith biomineralization can be divided into two main groups: (1) complexes of structural, insoluble collagen-like proteins and (2) soluble, non-collagenous proteins (NCPs). The collagen-like proteins, such as otolin-1, create a scaffold for the growing biomineral; otolin-1 has homology of sequences to collagen X, a protein also involved in endochondral ossification and bone fracture repair11. The soluble NCPs are usually highly acidic and intrinsically disordered proteins (IDPs) that directly regulate nucleation, orientation, and crystal growth. Such IDPs were identified in several fish taxa, e.g., Starmaker (Stm) in zebrafish7, Starmaker-like (Stm-l) in medaka12, and Otolith Matrix Macromolecule-64 (OMM-64) in rainbow trout13, and their role in the biomineralization has been thoroughly characterized14,15,16,17.
Finding evidence of proteins embedded into fossil otoliths would enhance our understanding of the evolution of an important aquatic biomineralization process, but such residues have not yet been identified and it is generally thought that these organic components are broken down and lost due to diagenetic alteration of the aragonite polymorph, which is metastable and normally transforms into a more stable calcite, e.g., via dissolution–precipitation processes in the presence of active solutions18. Such diagenetic processes also strongly affect the preservation of many inter/intra-crystalline proteins—in particular the highly labile NCPs19. In order to succeed in the detection and identification of remains of such proteins in fossil otoliths, we hypothesized that only specimens preserved in water-impermeable deposits and still composed entirely of aragonite with ultrastructural features similar to modern counterparts, have the potential to preserve organic components, most likely as inclusions embedded inside the aragonite crystals. Some easy-to-find sagittal Miocene otolith fossils from teleost fish fulfil these criteria. Here we report the results of a search for proteins embedded in fossil otoliths from phycid hake fish found in Miocene (about 14 million years old) water-impermeable clays exposed in Korytnica (Holy Cross Mountains, Central Poland; see Material)20.
Results
Mineral phase characteristics
Modern (white) and fossil (brownish color) saggital otoliths of Phycis spp. are slim and elongated calcium carbonate biomineral structures; wider at the anterior and gradually narrowing towards the posterior parts (Fig. 1a,h). The inner face (proximal surface) is convex without distinct sulcus acusticus (the area where the sensory tissue comes into contact with the otolith). The outer face (distal surface) is most commonly composed of irregular thickenings/protuberances and grooves (Fig. 1a,h). Longitudinal thin-sections (i.e., in the sagittal plane) observed with both polarized and normal transmitted light exhibit ca. 500 µm thick columnar units that correspond in size to protuberances on the surface (Fig. 1c,j). Occasionally, spindle-shaped voids occur between columnar units that may correspond to the grooves between neighboring surface protuberances (white arrows in Fig. 1c,j). The columnar units and other parts (proximal) of modern and fossil otoliths in longitudinal sections show numerous layers (alternating dark-brown and colorless zones, ca. 5–7 µm thick; Fig. 1c,d,j,k. Layers are composed of crystals that in crystallographic orientation images (EBSD) are present with similar size in modern and fossil samples (Fig. 1e,l). Phase maps have confirmed the strictly aragonitic mineralogy of both modern and fossil samples (Fig. 1f,m) and the resulting pole figures are fully comparable between modern (Fig. 1g) and fossil specimens (Fig. 1n). A high degree of similarity between modern and fossil specimens was also observed for crystal size, inclination, azimuthal dispersion, and turbostratic distribution in the plane (222) (Fig. 1g,n). The aragonite fibres of modern and fossil otoliths consist of slender units ca. 200–300 nm wide that show nanogranular organization (Fig. 2a,b,g,h), with nanograins ca. 50–100 nm in diameter clearly visible in AFM height- and peak force error-mode images (Fig. 2c,d,i,j). TEM observations of skeletal lamellae (extracted by focused ion beam) show aragonite, fine-scale fibres comparable in size with those observed in FESEM cut either obliquely (Fig. 2e) or longitudinally (Fig. 2k) to the growth direction. The fibres include numerous intracrystalline low density defects (inclusions) ranging in size between 2 and 25 nm, which are clearly seen as bright spots in TEM images (white arrows in Fig. 2f,l). These inclusions are aligned perpendicularly with the crystal growth direction as seen in longitudinal cuts (Fig. 2f,l). Material of similar low density is also located at the grain boundaries (Fig. 2e,k). The modern and fossil samples show overall similar weight loss profiles, but the derivative curves suggest differences in decomposition. Modern otoliths decompose over broader temperature range in comparison to modern ones (ca. 200–430 °C vs. 250–400 °C) and show several weight-loss steps (at ca. 210 °C, 360 °C, and 410 °C) in comparison to the one relatively large weight-loss (at ca. 340 °C) in fossil otoliths (Supplementary Fig. S1).
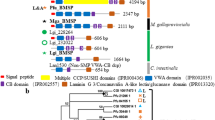
Morphological, microstructural and crystallographic similarity of modern Phycis phycis (a–g) and fossil Phycis tenuis (h–n) saggital otoliths. Modern (a) and fossil (h) sagittas are slim and elongated; the outer face (distal surface) most commonly is composed of irregular thickenings/protuberances and grooves. Thin-sections in polarized (b,c,i,j) and normal transmitted light (d,k) show numerous growth rings and point to ontogenetic continuity of protuberances (columns in longitudinal cut; occasionally separated by spindle-shaped voids, white arrows). Crystallographic orientation images (EBSD) show aragonite crystals of similar size in modern (e) and fossil (l) samples (phase maps (f,m) confirm aragonite mineralogy of both samples); pole figures are fully comparable between modern (g) and fossil samples (n): same crystal size, distribution, inclination, azimuthal dispersion and turbostratic distribution in the plane (222). ZPAL P.21/R-OTH-242/001 (a–g); ZPAL P.21/ C-OTH-07/006 (h–n). The BungeColorKey palette (more accessible for color-blind users) from MTEX was used to create orientation images and the pole figures.
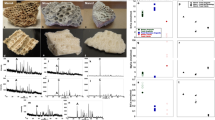
Composite, organic-mineral structure of Phycis phycis (a–f) and fossil Phycis tenuis (g–l) saggital otoliths. In FESEM the aragonite fibers of modern and fossil otoliths consist of slender units ca. 200–300 nm wide (a,g) that in higher magnification (b,h) show nanogranular organization; nanograins ca. 50–100 nm in diameter are visible in AFM height-mode (c,i) and peak force error-mode (d,j) images. TEM observations of skeletal lamellae (extracted by focused ion beam) show aragonite, fine-scale fibers (comparable in size with those observed in FESEM) that include numerous organic inclusions (arrows in f,l). Fibers in (e) cut obliquely or longitudinally (k) to the growth direction. ZPAL P.21/R-OTH-187/002 (a–f); ZPAL P.21/C-OTH-07/007 (g–l).
Protein preservation: amino acid racemization test
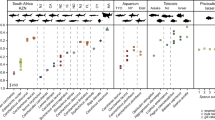
The amino acid racemization and their relative content in fossil otolith samples was used as a proxy of protein degradation. The analyses were performed using fossil P. tenuis otoliths compared directly with modern P. phycis. The measurements included free amino acids originally present within biomineral (FAAs) as well as those formed during hydrolysis of complete peptides into individual amino acids (so called total hydrolyzable amino acids, THAA). FAAs tend to result from the hydrolysis of highly-racemized N-terminal amino acids, so that the D/L ratio of FAA should be higher than that of the THAA for a given amino acid. As expected, the distribution of amino acids between pools of free amino acids (FAA) versus polymerized amino acids (THAA-FAA; as pmol amino acids per mg of starting otolith) shows that most amino acids in modern specimens are part of intact peptides whereas the majority of fossil peptides have broken down to individual amino acids. A dramatic loss of both Asx and Glx—which include the acidic amino acids aspartic and glutamic acid—as well as of Ser was observed in fossil specimens (Table 1).
Proteome analysis
In total, peptides for 132 fish-specific proteins and several isoforms were detected in modern P. phycis otoliths (Supplementary Table S2), while peptides for 11 fish-specific proteins were found in fossil P. tenuis otoliths (Table 2, Supplementary Tables S3–S5), which is a return ratio comparable to that of modern vs. fossil coral (aragonite) skeletons 21,22. Proteins were observed across both acid solubility fractions in all specimens and GluC improved protein detection in modern samples, but only tryptic digest peptides were detected in fossil samples (Supplementary Table S6). Additionally, deamidation of asparagine and glutamine and oxidation of methionine were detected (Supplementary Table S6)23. Five of 11 proteins sequenced from fossil otoliths represent proteins encoded by genes expressed in the inner ear (i.e., alpha-tectorin, beta-tectorin, otolin-1-like, otogelin/otogelin-like and otogelin-like). The other six proteins occur in modern fish otoliths, but are not specific to inner ear function. Proteins found in modern otoliths but not fossils include, e.g., usherin, a protein important to the development and homeostasis of the inner ear. Other modern otolith proteins include several types of collagens, contactin, low-density receptor-related lipoprotein, carbonic anhydrase that interconverts CO2 and bicarbonate (Supplementary Table S2).
Discussion
One of the most distinct features of biominerals, carbonates included, is that they are invariably organic-mineral composites in which the organic components, such as polysaccharides, lipids, and proteins are incorporated/embedded into the inorganic mineral phase, forming meso- to nanoscale intra- and intercrystalline inclusions and networks 24,25. These organic components participate in the physiologically mediated process of biomineral formation. Because the proteomic profiles can be linked with transcriptomic resources/expression, studies of the proteome of modern biomineral structures provide insights into molecular mechanisms of biomineralization. Identification of proteins in fossil biomineral structures therefore raises hope to gain indirect access to genome-related information in the absence of preserved DNA in fossils much older than ca. 2 My, the age beyond which DNA is generally no longer surviving in the fossil record 26,27.
Increasingly, paleoproteomic data have been extracted from various fossil biomineral structures 21,28,29,30,31. However, to date no paleoproteomic information exists from fossil otoliths that represent the most abundant fish remains in Mesozoic and Cenozoic deposits. This study is the first report on protein identification in fossil otoliths conducted in direct comparison with proteome of congeneric modern otoliths (phycid hakes). The following discussion focuses on two key aspects: structural criteria of fossil biominerals that preserve pristine paleoproteomic information and comparative analysis of protein content in modern and fossil phycid hakes otoliths.
Mineral phase vs. paleoproteome information preservation
The fossil material selected for this study came from Korytnica, a locality well known for exceptional preservation of aragonitic biominerals (see also “Material” section)32,33. Indeed, in all fossil otolith samples studied here, only the aragonite carbonate polymorph was detected; i.e., we observed no evidence of the presence of secondary calcite or other secondary phases. The exceptional preservation of these fossil otoliths is further supported by their crystallographic and ultrastructural features, which are indistinguishable from those characterizing the modern counterparts in terms of their distribution of crystal sizes, orientation/inclination, azimuthal dispersion, and turbostratic distribution (plane (222) (Figs. 1, 2). Further evidence of the extremely pristine preservation state of these fossil otoliths is provided by the occurrence of their nanogranular texture, typical of otoliths and most other biogenic minerals6,34,35. The nodular nanograins (ca. 100 nm in diameter), which are typically visualized with atomic force microscopy (Fig. 2c,d,i,j) are considered the product of a biomineralization process that involved amorphous precursors36,37. In this process, the organic molecules become incorporated into the crystallizing biomineral, e.g., as inclusions, and as organic-rich ‘envelopes’ around the resulting nanograins (Fig. 2e,f,k,l)25. Individual proteins involved in the carbonate biomineralization process have masses up to hundreds kDa and sizes from few to several nanometers in diameter/radius of random coil (structured/IDPs proteins). Their embedment into the crystal structure was interpreted as occurrence of intra- and intercrystalline inclusions38,39. Such inclusions are consistently present in modern and fossil otolith samples analysed herein, and we assume that they are the primary source of the proteinaceous material in this study. Prior (paleo)proteomic analyses examined amino acid racemization and their relative content measurements to assess the preservation potential of the samples40. Protein degradation is clearly suggested by the distribution of amino acids between pools of free amino acids (FAAs) versus polymerized amino acids (THAA-FAA, as pmol amino acids per mg of starting otolith material); most fossil peptides have broken down to individual amino acids. The observed bias toward acidic amino acids in fossil specimens suggests that highly acidic proteins (common to biominerals in general) are part of more soluble portions of the biomineral, which were preferentially degraded; this is reflected in the types of proteins that were sequenced by LC–MS/MS (Table 2). Protein degradation is also supported by the thermogravimetric data. The thermograms of Recent samples exhibit broader thermal decomposition range and a higher number of weight-loss steps in comparison to fossil ones. The Recent samples contain organic compounds that differ in their susceptibility to thermal decomposition, thus it is not surprising that the temperature range of their decay is relatively wide. It appears therefore that modern otoliths contain a greater diversity of organic (proteinaceous) components, which is manifested by higher thermal decomposition range (number of weight-loss steps) in comparison to fossil otoliths, for which the organic diversity is smaller.
Comparative (paleo)proteome analysis of modern and fossil phycid hake otoliths
More than 130 proteins were identified in otoliths from modern adult Phycis phycis. These include otolins, otogelins, usherins, and a cochlin, which have been previously identified in otoliths41,42. Of the detected modern otolith proteins, 11 were also observed in the fossil P. tenuis otoliths. These include alpha and a beta tectorin, two otogelin-like proteins, otolin-1-like, and a neuroserpin, all of which have been suggested to be directly involved in calcium carbonate biomineralization. We also identified in the fossil otoliths additional proteins not previously detailed from fish otoliths, such as collagenase; transforming growth factor b-inducing protein, splicing factor; arginine/serine-rich 19-like protein; protocadherin FAT 4-like proteins; and thrombospondin. All of these additional proteins except the splicing factor have GO terms designating them as extracellular or associated with a membrane. Several are calcium binding and at least one, protocadherin, has been detected in biogenic carbonates from other organisms43.
There are several possible reasons why this particular suite of proteins previously established as otolith matrix proteins were preserved in the fossil otoliths studied here. Co-preservation of some proteins may result from their intimate interactions during biomineralization. Both otogelin and alpha tectorins are necessary for tethering of the otolith membrane to sensory structures in the ear44, and otogelin is crucial in early larval development of the initial seeding of the otolith10. Beta tectorins likely sequester calcium as biomineralization proceeds10, and may polymerize with otolin45, possibly enhancing their co-preservation. Similarly, alpha tectorins possess an N-terminal Nidogen domain that has been proposed to allow it to also interact with otolin46. The proteins detected in fossil otoliths are some of the highest scoring proteins in modern otoliths (higher scores indicate a more confident match between combined scores of all observed mass spectra and amino acid sequences within examined protein), which has been shown to be linearly related to relative protein abundance47. Lastly, some proteins with acidic residues that provide an organic scaffold for biomineral formation (e.g., otolin-1 like48) are strongly stabilized by calcium ions; such proteins may firmly adhere to the biomineral surface, which may enhance their preservation potential in fossil record29.
Our observations refine the structural criteria for exceptional preservation of carbonate biominerals in the fossil record, and the paleoprotein sequences indicate highly conserved inner ear biomineralization processes in fish through geological time.
Material
Modern samples
Forkbeard (Phycis phycis, Phycidae family) modern otoliths were collected from fish caught off by fishermen along the mainland Portuguese west coast between 2011 and 2012. Sagittal otoliths were removed with ventral cranium section through gills, rinsed with water, air dried and stored in labelled plastic tubes at Lisbon Sciences Faculty (Portugal) until analyses49,50. Two large specimens of Phycis phycis were selected for biomineral structure analyses (ZPAL P.21/R-OTH-242/001, ZPAL P.21/R-OTH-187/002) and three specimens (ca. 2.5 g in weight) were selected for proteome analysis (ZPAL P.21/R-OTH-196/003, ZPAL P.21/R-OTH-197/004, ZPAL P.21/R-OTH-198/005).
Fossil samples
The fossil sagittal otolith samples of P. tenuis were collected from the Korytnica Clays [GPS position: 50°39′50′′ to 50°40′50′′ N and 20°31′20′′ to 20°33′00′′ E; three sites: Korytnica-Plebania, Korytnica-Forest, and Mt. Lysa33], a unique facies deposited in the terminal part of the bay (Korytnica Basin) developed in the Miocene along the rocky shore on the southern slopes of the Holy Cross Mountains, Central Poland20. The Korytnica Basin is filled by a shallowing-up sedimentary sequence composed of ca. 30–60 m thick sequence of clays, locally interfingering with oyster shellbeds. The absolute age of the Korytnica sequence is estimated as 14.8–14.6 Ma51,52 Korytnica Clays are renown from pristine preservation of Miocene fossils. Such exceptional preservation is supported by aragonite mineralogy (a metastable CaCO3 polymorph in normal conditions) of skeletons of e.g., scleractinian corals, gastropods, and fish otoliths and their distinct and fully comparable to modern counterparts micro- and nanostructural features6. The unusually favorable conditions of fossilization are further supported by exceptional, residual color patterns of some gastropod and barnacle shells20,53,54. Such preservation implies that fossils embedded in impermeable clays were virtually sealed off from extremal environment. After withdrawal of the Paratethys seas from the southern outskirts of the Holy Cross Mountains, these sediments were not covered by additional thick sedimentary cover that could cause any geothermal gradient heat effect on fossil material. The preset-day annual mean-temperature for Świętokrzyskie region in Poland ranges from 5.61 °C (in 1940) to 10.10 °C (in 2019); observations from 1901 till 202155. Considering the very fine grained nature of Korytnica-clays and consequently, typical to such sediments, extremely low thermal conductivity56 it can be reliable suggested that the examined otolith samples (retrieved from sediments found today at 1–2 m depth) have not experienced any significant temperature fluctuations during their burial history.
The clay samples were washed and sieved through standard sieves (500/250/125 μm) and dried at 40 °C. Of ca. 300 otolith specimens of P. tenuis, 2 specimens were selected for biomineral structure analyses (ZPAL P.21/C-OTH-07/006, and ZPAL P.21/C-OTH-07/006); 20 specimens (ca. 1.6 g in weight) were selected for proteome analysis (collective number ZPAL P.21/C-OTH-07/008-027). Specimens selected for (paleo)proteome analyses were soaked in sodium hypochlorite (5%) for 3 h, rinsed and ultrasonicated with deionized water and dried at 40 °C overnight.
Material of modern and fossil otoliths is housed at the Institute of Paleobiology, Polish Academy of Sciences, Warsaw (abbreviation ZPAL). Detailed sample identification information is provided in Supplementary Table S1.
Experimental
Otolith biomineral structure
Structural features of the otolith were studied and photographed using a transmitted light microscope Nikon Eclipse 80i at Institute of Paleobiology, Polish Academy of Sciences, Field–Emission Scanning Electron Microscopy (FESEM, Zeiss Merlin) at the Department of Chemistry, University of Warsaw, Thermo Fisher Tecnai Osiris microscope at Central facility in electron microscopy (CIME) of Swiss Federal Institute of Technology in Lausanne (EPFL), and Atomic Force Microscopy (AFM) using Multimode 5 instrument (Veeco) upgraded to Multimode 8 version (Bruker), at the Department of Chemistry, University of Warsaw. Light microscope images (in polarized light) were taken from ultra-thin (2–12 μm thick) sections made in sagittal plane of the otolith. FE-SEM images were taken of transverse broken sections of otoliths mounted on stubs with double-sided adhesive tape and sputter coated with a conductive platinum film; the accelerating voltage was of 5 kV, working distance 4–6 mm. Atomic Force Microscopy imaging was acquired in ScanAsyst mode using dedicated silicone cantilevers. Two signals (height and peak force error) were simultaneously collected during each scan. Otolith polished sagittal sections (Buehler Topol 3 final polishing suspension with particle size 0.25 µm) were rinsed in Milli-Q water, washed in an ultrasonic cleaner for 10 s, and then etched with a Milli-Q water solution for 7 h. The images were processed with WSxM v5.0 Develop 10.2 software from Nanotec57. Samples for Transmitted Electron Microscopy (TEM) were prepared as cross-sectional TEM lamellae extracted and then milled using a dual-beam Gemini NVision 40 Focused Ion Beam machine. The initial chunks are milled with gallium ions at 30 kV, 6.5 nA and then thinned down with lower currents step by step until using 80 pA, and finally smoothed at 5 kV, 80 pA. TEM analyses were performed at 200 kV accelerating voltage. High angle annular dark field (HAADF) images in scanning transmission electron microscopy (STEM) mode were recorded with a spot size of 0.5 nm and camera length of 115 mm.
Electron backscatter diffraction (EBSD)
The surface sample (of thick slides) was polished with alumina of 1 µm, 0.3 µm, and 0.05 µm and finally polished with colloidal silica (0.05 µm). Before analysis, samples were coated with a thin layer (ca. 2 nm) of carbon using a high vacuum coater. The EBSD study was carried out with Oxford NordlysMax detector mounted on a scanning electron microscope JEOL JSM-6610LV at the Institute of Materials Engineering, Łódź University of Technology. EBSD data were collected with AztecHKL software at high vacuum, 20 kV, large probe current, and 20 mm of working distance. EBSD patterns were collected at a resolution of 0.22 μm step size for crystallographic maps using the unit cell settings characteristic of aragonite and calcite as follows58,59: “Pmcn” symmetry and a = 4.96 Å, b = 7.97 Å, and c = 5.75 Å estimated for Favia coral using X-ray powder diffraction with synchrotron radiation (43) and a = b = 4.99 Å, and c = 17.06 Å, respectively. The EBSD data are represented in this study by crystallographic maps, phase images, and the pole figures, which represent the stereographic projection of crystallographic planes in reference to the (100), (010), (001) and (222) aragonite planes. Orientation images and the pole figures were created using MTEX open source plugin for Matlab program (https://mtex-toolbox.github.io/). To eliminate combination of red and green colors and create images more accessible for color-blind users we selected BungeColorKey palette from MTEX (the outcome was tested using Coblis, the Color Blindness Simulator at https://www.color-blindness.com/coblis-color-blindness-simulator/).
Thermogravimetry
Thermogravimetric analysis was performed with a TGA Q50 apparatus (TA Instruments) at the Department of Chemistry, University of Warsaw. Otolith samples (37.48 and 37.16 mg for fossil and Recent samples, respectively) were heated in nitrogen environment under linear gradient (10 °C min–1) from ambient 20 °C to 550 °C.
Amino acid racemization analysis
The first test of the assumption that original organic material remains in the otolith included amino acid racemization analysis (AAR)60,61. The AAR analysis provided information that the samples have been appropriately cleaned (D/L of fossil specimens approaching (1), that original protein material was still embedded (D/L of fossil specimens approaching (1), and gave an idea of the state of degradation (i.e., amino acid relative concentrations and abundance similar to those in modern otolith samples should suggest minimal degradation).
Amino acids for racemization analysis, both free and total hydrolysable amino acids, were extracted, hydrolyzed, and evaporated to dryness from cleaned skeleton powders (described below) by standard methods61. All samples were prepared in duplicate and analyzed at the Northern Arizona University Amino Acid Geochronology Laboratory using standard methods with modifications for microfossils60,62. Rehydrated samples were spiked with L-homo-arginine as an internal standard and then injected into an HPLC fitted with a reverse-phase C18-packed column. ‘Blank’ samples were included. We have previously shown that our ‘clean space’ set up and handling protocol are sufficient to prevent exogenous protein contamination in the laboratory21.
Proteome analysis
The fossil and modern otoliths were powdered by mortar and pestle to 125 μm, oxidized in 50:50 concentrated bleach/H2O2 for 1 h while sonicating following modified methods of Stoll et al.63, rinsed five times with MilliQ, and dried. Oxidation and rinses were repeated two more times. Cleaned powders of approximately 0.5 g per sample were decalcified in 0.5 M acetic acid with all handling occurring in a laminar flow hood to minimize contamination. Soluble organic matrix (SOM) was concentrated by centrifugal filtration (Amicon, 3 kDa cutoff) and rinsed with filtered phosphate buffered saline. Insoluble organic matrix (IOM; material that pelleted at 43,000×g for 5 min) was three times washed washed with 80% acetone. Fossil proteins were prepared as single samples for each solubility fraction (sample 07); modern samples were extracted as biological triplicates (samples 196–198). Samples were solubilized in SDS buffer and then digested using the MED-FASP protocol on a 30 kDa Microcon Centrifugal Unit (Sigma Aldrich) after rinsing out the SDS buffer with 8 M urea 64 by sequential applications of trypsin and then Glu-C enzymes. Samples were sequenced by liquid chromatography tandem mass spectrometry (LC–MS/MS) at the UCLA Semel Institute Proteomics Facility. Each fraction was analyzed separately on a nano-liquid-chromatography system coupled to a benchtop high-resolution orbitrap mass spectrometer (QE-Plus; Thermo Fisher) and operated in positive ion mode with data-dependent acquisition. MS1 was performed at resolution of 70,000 (at 400 m/z) and MS2 at 17,500. Instrumental blanks were run between all samples to minimize carryover. Transformed mass spectra were analyzed in Mascot against the UniProt-Human database, a common contaminants database, and the Phycis phycis genome’s 65 predicted protein database. The P. phycis protein database (Supplementary Table S5) was generated using the BRAKER pipeline, which includes the use of the GeneMark-ES/ET and Augustus programs66,67,68,69, to predict protein-coding regions of the unannotated P. phycis genome. The Atlantic cod (Gadus morhua) predicted protein database (NCBI assembly GCA_902167405.1 gadMor3.0) (predicted protein file available) was used to input “hints” to guide CDS prediction. All samples allowed a fixed modification of carbamidomethylation on C and variable oxidation on MW, deamidation on NQ and protein N-terminal acetylation; fossil samples also allowed Phospho K & T and MOD. For each sample, a first decoy search was carried out to determine p-values for a 1% false discovery rate. Then an error tolerant search was conducted, with the p-value adjusted if necessary. Cutoff scores were applied at the value recommended by the Mascot algorithm. Returned sequences were annotated in Blast2GO software and further run in Blast2GO against the NCBI nr primates database to test for potential human contaminants not picked up by the UniProt-Human database in Mascot; likely human contaminant proteins were excluded from the final list if they were > 90% similar to primates with an e-value within 20 units of the original annotation, within 10 e-value units and 10% similarity for e-50 and lower, or within 5 e-value units and 10% similarity for e-50 and higher22. Duplicate sequences were checked in CD-HIT with > 90% similarity; duplicates are noted separately but counted together in total protein counts. Several proteins were predicted as separate peptides; those peptides were BLASTed against the Atlantic cod (Gadus morhua) predicted proteome (NCBI assembly GCA_902167405.1 gadMor3.0) and concatenated, with strings of XXs denoting regions of unknown sequence between known peptides.
Data availability
All data generated or analyzed during this study are included in this published article (and its Supplementary Information). The mass spectrometry proteomics data have been deposited to the Proteomic Xchange Consortium via the Pride partner repository70 (https://www.ebi.ac.uk/pride/login) under the dataset identifier PXD036742 and https://doi.org/10.6019/PXD036742. No ethical approval or guidance was required because this study only analyzed specimens were collected from commercial landings of fishing vessels and fossil material in museum collections.
References
Thomas, B. & Taylor, S. Proteomes of the past: The pursuit of proteins in paleontology. Exp. Rev. Proteom. 16, 881–895. https://doi.org/10.1080/14789450.2019.1700114 (2019).
Green, E. J. & Speller, C. F. Novel substrates as sources of ancient DNA: Prospects and hurdles. Genes (Basel) https://doi.org/10.3390/genes8070180 (2017).
Cappellini, E. et al. Ancient biomolecules and evolutionary inference. Annu. Rev. Biochem. 87, 1029–1060. https://doi.org/10.1146/annurev-biochem-062917-012002 (2018).
Schwarzhans, W., Beckett, H. T., Schein, J. D., Friedman, M. & Rahman, I. Computed tomography scanning as a tool for linking the skeletal and otolith-based fossil records of teleost fishes. Palaeontology 61, 511–541. https://doi.org/10.1111/pala.12349 (2018).
Campana, S. E. Otolith science entering the 21st century. Mar. Freshw. Res. https://doi.org/10.1071/mf04147 (2005).
Stolarski, J. & Mazur, M. Nanostructure of biogenic versus abiogenic calcium carbonate crystals. Acta Palaeontol. Pol. 50, 847–865 (2005).
Sollner, C. et al. Control of crystal size and lattice formation by starmaker in otolith biomineralization. Science 302, 282–286. https://doi.org/10.1126/science.1088443 (2003).
Payan, P., De Pontual, H., Bœuf, G. & Mayer-Gostan, N. Endolymph chemistry and otolith growth in fish. C.R. Palevol 3, 535–547. https://doi.org/10.1016/j.crpv.2004.07.013 (2004).
Thompson, J. B. et al. Direct observation of the transition from calcite to aragonite growth as induced by Abalone shell proteins. Biophys. J . 79, 3307–3312. https://doi.org/10.1016/s0006-3495(00)76562-3 (2000).
Thomas, O. R. B. & Swearer, S. E. Otolith biochemistry—A review. Rev. Fish. Sci. Aquacult. 27, 458–489. https://doi.org/10.1080/23308249.2019.1627285 (2019).
Murayama, E. et al. Fish otolith contains a unique structural protein, otolin-1. Eur. J. Biochem. 269(2), 688–696 (2002).
Bajoghli, B., Ramialison, M., Aghaallaei, N., Czerny, T. & Wittbrodt, J. Identification of starmaker-like in medaka as a putative target gene of Pax2 in the otic vesicle. Dev. Dyn. 238, 2860–2866. https://doi.org/10.1002/dvdy.22093 (2009).
Tohse, H., Takagi, Y. & Nagasawa, H. Identification of a novel matrix protein contained in a protein aggregate associated with collagen in fish otoliths. FEBS J. 275, 2512–2523. https://doi.org/10.1111/j.1742-4658.2008.06400.x (2008).
Kapłon, T. et al. Biomacromolecules. Vol. 9.8. 2118–2125 (2008).
Wojtas, M., Marek, W., Andrzej, O. & Piotr, D. Phosphorylation of intrinsically disordered starmaker protein increases its ability to control the formation of calcium carbonate crystals. Cryst. Growth Des. 12, 158–168. https://doi.org/10.1021/cg200905f (2012).
Różycka, M. et al. Intrinsically disordered and pliable starmaker-like protein from Medaka (Oryzias latipes) controls the formation of calcium carbonate crystals. PLoS ONE 9, 114308 (2014).
Kalka, M. et al. In vivo and in vitro analysis of starmaker activity in zebrafish otolith biomineralization. FASEB J 33, 6877–6886. https://doi.org/10.1096/fj.201802268R (2019).
Hashim, M. S. & Kaczmarek, S. E. The transformation of aragonite to calcite in the presence of magnesium: Implications for marine diagenesis. Earth Planet. Sci. Lett. https://doi.org/10.1016/j.epsl.2021.117166 (2021).
Mitterer, R. M. Organic Geochemistry (eds. Macko Micheael, S.A., Engel, H.). 739–753 (Plenum Press, 1993).
Bałuk, W. & Radwański, A. Organic communities and facies development of the Korytnica Basin (Middle Miocene; Holy Cross Mountains, Central Poland). Acta Geol. Pol. 27, 85–123 (1977).
Drake, J. L., Whitelegge, J. P. & Jacobs, D. K. First sequencing of ancient coral skeletal proteins. Sci. Rep. https://doi.org/10.1038/s41598-020-75846-4 (2020).
Peled, Y. et al. Optimization of skeletal protein preparation for LC-MS/MS sequencing yields additional coral skeletal proteins in Stylophora pistillata. BMC Mater 2, 8. https://doi.org/10.1186/s42833-020-00014-x (2020).
Hendy, J. et al. A guide to ancient protein studies. Nat. Ecol. Evol. 2, 791–799. https://doi.org/10.1038/s41559-018-0510-x (2018).
Mann, S. Biomineralization: Principles and Concepts in Bioinorganic Materials Chemistry. 1–210 (Oxford University Press, 2001).
Cuif, J.-P., Dauphin, Y. & Sorauf, J. E. Biominerals and Fossils Through Time. (2011).
van der Valk, T. et al. Million-year-old DNA sheds light on the genomic history of mammoths. Nature 591, 265–269. https://doi.org/10.1038/s41586-021-03224-9 (2021).
Kjaer, K. H. et al. A 2-million-year-old ecosystem in Greenland uncovered by environmental DNA. Nature 612, 283–291. https://doi.org/10.1038/s41586-022-05453-y (2022).
Bada, J. L., Wang, X. S. & Hamilton, H. Preservation of key biomolecules in the fossil record: Current knowledge and future challenges. Philos. Trans. R Soc. Lond. B Biol. Sci. 354, 77–86. https://doi.org/10.1098/rstb.1999.0361 (1999).
Demarchi, B. et al. Protein sequences bound to mineral surfaces persist into deep time. Elife https://doi.org/10.7554/eLife.17092 (2016).
Schroeter, E. R. et al. Expansion for the Brachylophosaurus canadensis collagen I sequence and additional evidence of the preservation of cretaceous protein. J. Proteome Res. 16, 920–932. https://doi.org/10.1021/acs.jproteome.6b00873 (2017).
Warinner, C., Korzow Richter, K. & Collins, M. Paleoproteomics. J. Chem Rev 122, 13401–13446. https://doi.org/10.1021/acs.chemrev.1c00703 (2022).
Radwańska, U. Fish otoliths in the Middle Miocene (Badenian) deposits of southern Poland. Acta Geol. Pol. 42, 141–327 (1992).
Stolarski, J. Miocene Scleractinia from the Holy Cross Mountains, Poland; Part 1—Caryophylliidae, Flabellidae, Dendrophylliidae, and Micrabaciidae. Acta Geol. Pol. 41, 37–67 (1991).
Dauphin, Y. & Dufour, E. Nanostructures of the aragonitic otolith of cod (Gadus morhua). Micron 39, 891–896. https://doi.org/10.1016/j.micron.2007.11.007 (2008).
Gilbert, P. U. P. A. et al. Biomineralization by particle attachment in early animals. Proc. Natl. Acad. Sci. 116, 17659. https://doi.org/10.1073/pnas.1902273116 (2019).
Mastropietro, F. et al. Revealing crystalline domains in a mollusc shell single-crystalline prism. Nat. Mater. 16, 946–952. https://doi.org/10.1038/nmat4937 (2017).
De Yoreo, J. J. et al. Crystal growth. Crystallization by particle attachment in synthetic, biogenic, and geologic environments. Science 349, aaa6760. https://doi.org/10.1126/science.aaa6760 (2015).
Younis, S., Kauffmann, Y., Bloch, L. & Zolotoyabko, E. Inhomogeneity of nacre lamellae on the nanometer length scale. Cryst. Growth Des. 12, 4574–4579. https://doi.org/10.1021/cg3007734 (2012).
Frankowiak, K. et al. Fine-scale skeletal banding can distinguish symbiotic from asymbiotic species among modern and fossil Scleractinian corals. PLoS ONE 11, e0147066. https://doi.org/10.1371/journal.pone.0147066 (2016).
Drake, J. L., Guillermic, M., Eagle, R. A. & Jacobs, D. K. Fossil corals with various degrees of preservation can retain information about biomineralization-related organic material. Front. Earth Sci. https://doi.org/10.3389/feart.2021.643864 (2021).
Thomas, O. R. B. et al. The inner ear proteome of fish. FEBS J. 286, 66–81. https://doi.org/10.1111/febs.14715 (2019).
Leventea, E. et al. Ciliopathy genes are required for apical secretion of Cochlin, an otolith crystallization factor. Proc. Natl. Acad. Sci. USA https://doi.org/10.1073/pnas.2102562118 (2021).
Drake, J. L., Mass, T. & Falkowski, P. G. The evolution and future of carbonate precipitation in marine invertebrates: Witnessing extinction or documenting resilience in the Anthropocene?. Elementa Sci. Anthropocene https://doi.org/10.12952/journal.elementa.000026 (2014).
Stooke-Vaughan, G. A., Obholzer, N. D., Baxendale, S., Megason, S. G. & Whitfield, T. T. Otolith tethering in the zebrafish otic vesicle requires Otogelin and alpha-Tectorin. Development 142, 1137–1145. https://doi.org/10.1242/dev.116632 (2015).
Yang, C. H. et al. Zona pellucida domain-containing protein beta-tectorin is crucial for zebrafish proper inner ear development. PLoS ONE 6, e23078. https://doi.org/10.1371/journal.pone.0023078 (2011).
Lundberg, Y. W., Zhao, X. & Yamoah, E. N. Assembly of the otoconia complex to the macular sensory epithelium of the vestibule. Brain Res. 1091, 47–57. https://doi.org/10.1016/j.brainres.2006.02.083 (2006).
Liu, H., Sadygov, R. G. & Yates, J. R. 3rd. A model for random sampling and estimation of relative protein abundance in shotgun proteomics. Anal. Chem. 76, 4193–4201. https://doi.org/10.1021/ac0498563 (2004).
Holubowicz, R. et al. Effect of calcium ions on structure and stability of the C1q-like domain of otolin-1 from human and zebrafish. FEBS J. 284, 4278–4297. https://doi.org/10.1111/febs.14308 (2017).
Vieira, A. R., Neves, A., Sequeira, V., Paiva, R. B. & Gordo, L. S. Age and growth of forkbeard, Phycis phycis, in Portuguese continental waters. J. Mar. Biol. Assoc. U.K. 94, 623–630. https://doi.org/10.1017/s0025315413001549 (2013).
Vieira, A. R., Neves, A., Sequeira, V., Paiva, R. B. & Gordo, L. S. Otolith shape analysis as a tool for stock discrimination of forkbeard (Phycis phycis) in the Northeast Atlantic. Hydrobiologia 728, 103–110. https://doi.org/10.1007/s10750-014-1809-5 (2014).
Rögl, F. H. M. et al. The Middle Miocene Badenian stratotype at Baden-Sooss (Lower Austria). Geol. Carpathica 59, 367–374 (2008).
Zágoršek, K., Radwańska, U. & Radwański, A. Bryozoa from the Korytnica Basin (Middle Miocene; Holy Cross Mountains, Central Poland). Bull. Geosci. https://doi.org/10.3140/bull.geosci.1249 (2012).
Bałuk, W. Lower Tortonian gastropods from Korytnica. Poland. PartI. Palaeontol. Polon. 32, 1–186 (1975).
Bałuk, W. R. A. Miocene cirripeds domiciled in corals. Acta Palaeontol. Polon. 12, 457–513 (1967).
Climate Change Knowledge Portal. https://climateknowledgeportal.worldbank.org/country/poland/climate-data-historical (2023).
Midttømme, K. R. E. & Aagaard, P. Thermal conductivity of selected claystones and mudstones from England. Clay Miner. 33, 131–145 (1998).
Horcas, I. et al. WSXM: A software for scanning probe microscopy and a tool for nanotechnology. Rev. Sci. Instrum. 78, 013705. https://doi.org/10.1063/1.2432410 (2007).
Stolarski, J., Przeniosło, R., Mazur, M. & Brunelli, M. High-resolution synchrotron radiation studies on natural and thermally annealed scleractinian coral biominerals. J. Appl. Crystallogr. 40, 2–9. https://doi.org/10.1107/s002188980604489x (2007).
Coronado, I., Fine, M., Bosellini, F. R. & Stolarski, J. Impact of ocean acidification on crystallographic vital effect of the coral skeleton. Nat. Commun. 10, 2896. https://doi.org/10.1038/s41467-019-10833-6 (2019).
Kaufman, D. S. & Manley, W. F. A new procedure for determining dl amino acid ratios in fossils using reverse phase liquid chromatography. Quatern. Sci. Rev. 17, 987–1000. https://doi.org/10.1016/s0277-3791(97)00086-3 (1998).
Penkman, K. E., Kaufman, D. S., Maddy, D. & Collins, M. J. Closed-system behaviour of the intra-crystalline fraction of amino acids in mollusc shells. Quat. Geochronol. 3, 2–25. https://doi.org/10.1016/j.quageo.2007.07.001 (2008).
Kaufman, D. Dating deep-lake sediments by using amino acid racemization in fossil ostracodes. Geology https://doi.org/10.1130/g20004.1 (2003).
Stoll, H. M. et al. A first look at paleotemperature prospects from Mg in coccolith carbonate: Cleaning techniques and culture measurements. Geochem. Geophys. Geosyst. https://doi.org/10.1029/2000gc000144 (2001).
Wisniewski, J. R. Filter-aided sample preparation: The versatile and efficient method for proteomic analysis. Methods Enzymol. 585, 15–27. https://doi.org/10.1016/bs.mie.2016.09.013 (2017).
Malmstrom, M., Matschiner, M., Torresen, O. K., Jakobsen, K. S. & Jentoft, S. Whole genome sequencing data and de novo draft assemblies for 66 teleost species. Sci. Data 4, 160132. https://doi.org/10.1038/sdata.2016.132 (2017).
Hoff, K. J., Lange, S., Lomsadze, A., Borodovsky, M. & Stanke, M. BRAKER1: Unsupervised RNA-seq-based genome annotation with GeneMark-ET and AUGUSTUS. Bioinformatics 32, 767–769. https://doi.org/10.1093/bioinformatics/btv661 (2016).
Hoff, K. J., Lomsadze, A., Borodovsky, M. & Stanke, M. Whole-genome annotation with BRAKER. Methods Mol. Biol. 65–95, 2019. https://doi.org/10.1007/978-1-4939-9173-0_5 (1962).
Stanke, M., Schoffmann, O., Morgenstern, B. & Waack, S. Gene prediction in eukaryotes with a generalized hidden Markov model that uses hints from external sources. BMC Bioinform. 7, 62. https://doi.org/10.1186/1471-2105-7-62 (2006).
Lomsadze, A., Burns, P. D. & Borodovsky, M. Integration of mapped RNA-Seq reads into automatic training of eukaryotic gene finding algorithm. Nucleic Acids Res. 42, e119. https://doi.org/10.1093/nar/gku557 (2014).
Perez-Riverol, Y. et al. The PRIDE database and related tools and resources in 2019: Improving support for quantification data. Nucleic Acids Res. 47, D442–D450. https://doi.org/10.1093/nar/gky1106 (2019).
Acknowledgements
This work was supported by National Science Centre (Poland) research grants 2017/25/B/ST10/02221 and 2020/39/B/ST10/01253 to JS, NSF Postdoctoral Research Fellowship in Biology award #1611943 and Zuckerman STEM postdoctoral leadership fellowship to JD, and Swiss National Science Foundation grant 205321_212614 to AM. ARV was supported by Fundação para a Ciência e a Tecnologia (FCT, Portugal) through the research contract CEECIND/01528/2017. We thank T. Bernat, the Amino Acid Geochronology Laboratory at Northern Arizona University and the NPI-Semel Institute Proteomics Laboratory at UCLA for sample processing, amino acid and protein analysis, respectively.
Author information
Authors and Affiliations
Contributions
J.S. and J.D. designed the study, analysed the data and drafted the manuscript. A.R.V. and U.R. provided and identified material of modern and some fossil otoliths. I.C. made EBSD maps and interpreted crystallographic measurements. M.M. made AFM images and thermogravimetric analysis. E.A.C.H. conducted the P. phycis protein prediction using BRAKER. G.J. made TEM analyses. J.S., J.D., I.C., E.A.C.H., G.J., A.M. contributed to the interpretation of the results and writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Stolarski, J., Drake, J., Coronado, I. et al. First paleoproteome study of fossil fish otoliths and the pristine preservation of the biomineral crystal host. Sci Rep 13, 3822 (2023). https://doi.org/10.1038/s41598-023-30537-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-30537-8
This article is cited by
-
Preservation of proteins in the geosphere
Nature Ecology & Evolution (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.