Abstract
Microbial rewilding, whereby exposure to naturalistic environments can modulate or augment gut microbiomes and improve host-microbe symbiosis, is being harnessed as an innovative approach to human health, one that may also have significant value to animal care and conservation. To test for microbial rewilding in animal microbiomes, we used a unique population of wild-born ring-tailed lemurs (Lemur catta) that were initially held as illegal pets in unnatural settings and, subsequently, relocated to a rescue center in Madagascar where they live in naturalistic environments. Using amplicon and shotgun metagenomic sequencing of lemur and environmental microbiomes, we found multiple lines of evidence for microbial rewilding in lemurs that were transitioned from unnatural to naturalistic environments: A lemur’s duration of exposure to naturalistic settings significantly correlated with (a) increased compositional similarly to the gut communities of wild lemurs, (b) decreased proportions of antibiotic resistance genes that were likely acquired via human contact during pethood, and (c) greater covariation with soil microbiomes from natural habitats. Beyond the inherent psychosocial value of naturalistic environments, we find that actions, such as providing appropriate diets, minimizing contact with humans, and increasing exposure to natural environmental consortia, may assist in maximizing host-microbe symbiosis in animals under human care.
Similar content being viewed by others
Introduction
Gut microbiomes (GMBs), critical to animal health1, are shaped by various environmental factors, such that altered or unnatural ecosystems (e.g., degraded habitats) have perturbative effects on host-associated communities, with negative health implications for hosts2,3. Exposure to key environmental factors has the potential to augment or restore native host-associated microfauna4 via an understudied, presumably gradual process known as microbial ‘rewilding.’ The Microbiome Rewilding Hypothesis posits that the restoration of ‘green’ habitats and promotion of diverse environmental microbiomes in urban settings can improve human GMBs and health5. If the exposure to or introduction of certain microbial inhabitants can improve host-microbe symbiosis and the host’s ability to adapt to new environments, then rewilding could benefit captive animals transitioning between settings or ecosystems, such as during transfers between captivity facilities, translocations, or reintroductions6. Here, we expand the hypothesis to nonhuman primates and test for microbial rewilding in wild-born, captive ring-tailed lemurs (Lemur catta) transitioning from highly unnatural settings during illegal pethood to a more natural setting after their surrender to the Lemur Rescue Center (LRC) in Madagascar (Table 1). We ask if, with exposure to naturalistic environments, the GMBs of LRC lemurs better resemble those of pet lemurs or their wild counterparts.
Belying traditional dichotomization, both wild and captive settings represent a range of variation known to influence animal GMB structure and function7. The GMBs of ring-tailed lemurs, for instance, vary within and between captive and wild settings, such that there is not a universal signal of captivity nor is there a specific, core microbiome that is representative of all of the wild animals8 (Supplementary Fig. S1). Here, we focus on three factors known to impact GMB structure and variation across settings: diet, human contact, and exposure to natural environments (Table 1). Notably, the degree of evolutionary mismatch between the diets of wild and captive counterparts is thought to underlie significant variation in GMB diversity and composition9,10. In addition, contact with humans can facilitate transmission of microbes and antibiotic resistance genes (ARGs) between humans and other animals11. Lastly, exposure to natural environments can mediate the acquisition of environmental microbes and ARGs that can impact host-associated communities and animal health8,12. Transitions between settings with different types or degrees of these factors could precipitate changes in multiple aspects of the microbiome, whether via a detrimental perturbation or a beneficial microbial rewilding.
The wild-born lemurs at the LRC have experienced at least two drastic environmental transitions within their lifetime. The first is a perturbative transition when removed from the wild to be kept as pets13, and the second is a potentially rewilding transition from pethood to life at the LRC. We use cross-sectional data to first address if time in residency at the LRC correlates with the (a) diversity, (b) phylogenetic composition, and (c) abundance of bacterial taxa in lemur GMBs. We focus on the genera Bacteroides, Prevotella, and Ruminococcus, as these may serve as biomarkers of host diet type and gut health14. Notably, despite the absence of a diverse core GMB among wild and captive ring-tailed lemurs, these microbes are shared and abundant across populations8, are also present in the GMBs of other wild and captive primates, and are linked to distinct enterotypes in human GMBs. Investigating variation in these ubiquitous microbes, in combination with broader attributes of microbial communities (e.g., diversity and composition), affords a holistic view of lemur GMB structure, as well as potential insights into changes in functional potential. Next, we also ask if residency at the LRC influences ARG abundance and covariation between lemur GMBs and soil microbiomes from natural habitats. Microbial rewilding in LRC lemurs predicts (i) greater compositional similarity to the GMBs of wild lemurs, (ii) decreased ARG abundance, and (iii) greater covariation with soil microbiomes.
Methods
Subjects and samples
The subjects included ring-tailed lemurs living (a) in the wild (n = 139), (b) as pets in Malagasy households (n = 8), and (c) at the LRC in Mangily, Madagascar (n = 25)8. Their diets and exposure to humans and environmental microbiomes are summarized in Table 1. Wild lemurs inhabited protected areas (e.g., national parks, community-managed reserves) that varied in habitat type from dry spiny forest to riverine forest. They relied entirely on naturally foraged diets and were constantly exposed to natural environmental microbiomes. Pet lemurs lived in human dwellings in townships located around Toliara, Madagascar. Two of the pet lemurs had limited access to outdoor areas. Their diets were ‘humanized,’ consisting of commercial grains and produce, and they had limited exposure to natural environmental microbiomes. The LRC lemurs were wild-born, former pets that had known dates of surrender to the LRC, where they were socially housed in outdoor enclosures, with access to shelter. They thus could forage freely, obtaining a partial natural diet, supplemented with seasonally available produce, and were exposed to natural environmental microbiomes. Exposure to humans and to ARGs (from combined environmental exposure and/or direct antibiotic administration) was least in the natural populations, maximal in pets, and relatively limited in LRC animals.
We opportunistically collected fresh fecal samples upon observing lemur defecation. To avoid soil contamination of the fecal samples, we removed the outer layer of each fecal pellet. We also collected samples of topsoil (n = 22) from the wild lemurs’ natural habitats, including spiny, dry, and riverine forests in southern Madagascar. When collecting soil, we avoided high-defecation areas (e.g., under sleeping trees) and areas with significant organic matter (e.g., dead vegetation), focusing instead on areas with bare soil, where the lemurs most commonly spent time on the ground. Within these areas, we demarcated a 2–3 m2 area and collected topsoil (the top 2–3 cm of soil) from each of five evenly spaced locations. For each area, we pooled the five aliquots of topsoil in a single tube to create a representative soil sample. All fecal and soil samples were preserved in Omnigene.Gut tubes (DNAgenotek, Ontario, Canada)15 and, within 8 weeks of collection, were transported to the U.S. and stored at − 80 °C until analysis. No animal handling, manipulation, or experimentation was performed for this study.
Microbial DNA extraction and sequencing
Following the manufacturer’s protocols for the DNeasy Powersoil kit (QIAGAN, Frederick, MD), we extracted bacterial genomic DNA from fecal and soil samples. We sent aliquots of extracted DNA to Argonne National Laboratory’s Environmental Sequencing facility (Lemont, IL) for library preparation and amplicon sequencing of the V4 region of the 16S rRNA gene. Amplicons were sequenced on a 151 × 151 base pair Illumina MiSeq run16.
We sent a subset of the extracted DNA aliquots (wild lemurs, n = 7; pet lemurs, n = 7; LRC lemurs, n = 9) to CosmosID Inc. (Rockville, MD) for shotgun metagenomic sequencing to identify antibiotic resistance genes. DNA libraries were prepared using the Illumina Nextera XT library preparation kit, with a modified protocol17. Libraries were then sequenced on an Illumina HiSeq platform 2 × 150 bp. On average, the sequencing yielded approximately 17 million total sequence reads per sample, with an average of 18 million and 10 million reads for fecal and soil samples, respectively. Samples with fewer than 5 million reads (n = 2 samples) were omitted from downstream analyses.
Bioinformatics and statistical analyses
We processed the 16S rRNA sequence data using a bioinformatics pipeline generated in QIIME218,19. We used the pipeline to join forward and reverse reads, demultiplex, quality filter joined reads and remove chimeras (DADA2 plugin; PHRED scores indicated no quality trimming was needed)20, omit non-bacterial sequences (Mitochondria, but not chloroplasts as they can serve as a valuable proxy for diet and environmental exposure18,21,22), and generate a phylogenetic tree (mafft program23 and fasttree224). To assign taxonomy to our sequence features and generate amplicon sequence variants (ASVs), we de novo trained the Naive Bayes classifier using the SILVA database (ver. 138.1) at 99% sequence similarity25,26 and tested the classifier using our representative sequences. After quality filtering, all samples had > 10,000 reads and were retained for downstream analysis. Using QIIME2, we calculated metrics of alpha diversity (Shannon and Faith’s Phylogenetic diversity metric) and beta diversity (weighted and unweighted UniFrac distances) on a rarefied ASV feature table subsampled to 15,000 reads per sample (Supplementary Fig. S2). To examine variation in the abundance of specific microbial taxa, we used R Studio (ver. 4.2.0) to perform a center log-ratio (CLR) transformation on the unrarefied ASV feature table (package ‘compositions’)27,28. CLR abundances reflect log-transformed ratios of the raw sequence counts of each taxon over the geometric mean of all other taxa in the sample29.
For shotgun metagenomic data, unassembled sequencing reads were directly analyzed using CosmosID’s bioinformatics platform for identifying and profiling ARGs17,30,31. The system uses multiple genome databases and a high-performance, data-mining algorithm that disambiguates metagenomic sequence reads. To identify ARGs, we queried the unassembled sequence reads against the CosmosID curated ARG database, which was compiled through assimilation of ARG sequences collected from the published literature, as well as from different open-source databases, including the following: NCBI, CARD, ResFinder, ARDB, ARG-ANNOT, and SEEC. If annotation of a gene conferring resistance was not included in their database, the CosmosID team performed literature searches to determine the class or relevant mechanisms of resistance.
Briefly, and without revealing proprietary information, the CosmosID system uses a high-performance, data-mining k-mer algorithm and highly curated dynamic comparator databases (GenBook®) that rapidly disambiguate millions of short reads into the discrete genomes or genes engendering the particular sequences. The pipeline has two separable comparators: the first consists of a pre-computation phase for reference database and a per-sample computation. The input to the pre-computation phase is a reference microbial genome or antibiotic resistance and virulence gene database, and its output is phylogeny trees, together with sets of variable length k-mer fingerprints (biomarkers) that are uniquely identified with distinct nodes, branches and leaves of the tree. The second per-sample, computational phase searches the hundreds of millions of short sequence reads or contigs from draft assembly against the fingerprint sets. The resulting statistics are analyzed to give fine-grain composition and relative abundance estimates. The second comparator uses edit distance-scoring techniques to compare a target genome or gene with a reference set. The algorithm provides similar functionality to BLAST, but sacrifices some recall precision for a one- or two-order-of-magnitude processing gain. Overall classification precision is maintained through aggregation statistics. Enhanced detection specificity is achieved by running the comparators in sequence. The first comparator finds reads in which there is an exact match with a k-mer uniquely identified with an ARG; the second comparator then statistically scores the entire read against the reference to verify that the read is indeed uniquely identified with that reference. For each sample, the reads from a species are assigned to the strain with the highest aggregation statistics. Outputs include the identity and family, percent gene coverage, and frequency counts of ARGs within each sample. To calculate the proportion of ARGs within a fecal sample, we divided the frequency count of all ARGs or specific gene families by the sample’s total read count.
To calculate covariation between lemur GMBs and soil microbiomes, we used FEAST32, a tool that uses fast expectation–maximization, multinomial distributions, and machine-learning classification to model microbial source tracking. FEAST provides “source proportions” of the scaled proportion of each LRC lemur’s GMB community that could be attributed to soil communities from natural habitats or to a default ‘unknown source’ that accounts for microbes not relevant to soil microbiota32.
For all LRC lemurs, we calculated time in residency at the LRC as the number of days between surrender date and the date of sample collection (range = 248–2,537 days, standard deviation = 617.7, median = 1,736). Using linear models in R Studio (package ‘stats’), we tested for effects of time in residency at the LRC on lemur GMB diversity, composition, membership, ARGs, and covariation with soil microbiomes. The model included the duration of residency at the LRC as a fixed effect.
Ethics
Sampling in Madagascar occurred with approval from Madagascar National Parks and appropriate governmental agencies (Ministry of Environment, Ecology, and Forests; permit #s 147/18/MEEF/SG/DGF/DSAP/SCB.Re, 152/19/MEDD/SG/DGEF/DGRNE, 159/16/MEEF/SG/DGF/DSAP/SCB.Re, 154/17/ MEEF/SG/DGF/DSAP/SCB.Re, 156/19/MEEF/SG/DGF/DSAP/SCB.Re). At the time of collection, samples did not require CDC, USDA, or CITES permits. All samples were declared, permits presented, and cleared through U.S. Customs and Border Protection.
Results
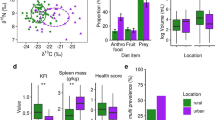
We observed a negative trend in alpha diversity with time in residence at the LRC; nevertheless, the patterns did not reach statistical significance for any metric. In contrast, both compositional measures (or beta diversity) of lemur GMBs significantly correlated with time in residence (Table 2). Specifically, the longer animals resided at the LRC, the more similar their GMB composition was to that of their wild counterparts (Fig. 1a,b; Table 2), consistent with rewilding.
The center log-ratio (CLR)-transformed abundance of the Bacteroides genus increased significantly with increasing time at the LRC (Fig. 1c). In contrast, the CLR abundances of both the genera Prevotella and Ruminococcus decreased significantly with increasing time at the LRC (Fig. 1d,e; Table 2).
Compositional patterns in the gut microbiomes (GMBs) of three categories of ring-tailed lemurs (Lemur catta) in Madagascar. (a) ‘Population signatures’ as revealed by principal coordinate plots of unweighted UniFrac distances for wild lemurs (blue), pet lemurs (yellow), and lemurs in semi-natural conditions at the Lemur Rescue Center (LRC; color-graded in relation to duration in residency). (b) Rewilding, as revealed by pairwise comparisons, using unweighted UniFrac distance, between the GMBs of pet vs. wild lemurs, LRC vs. wild lemurs, and within wild lemurs. (c–e) Center log-ratio (CLR)-transformed abundances of Bacteroides, Prevotella, and Ruminococcus in the GMBs of LRC lemurs. Shown are linear trend lines and 95% confidence intervals.; See Table 2 for full statistical results from linear mixed models.
The total relative abundance of ARGs in the GMBs of LRC lemurs ranged from 0.16–0.59% (mean = 0.29% ± 0.14%). As predicted by rewilding, the relative abundance of total ARGs and of tetracycline ARGs (i.e., the most abundant class of ARGs) decreased significantly with time spent at the LRC (Fig. 2a,b; Table 2).
Environmental influences on the gut microbiomes (GMBs) of three categories of ring-tailed lemurs (Lemur catta) in Madagascar. Relative abundances of (a) total antibiotic resistance genes (ARGs) in wild lemurs (blue), pet lemurs (yellow), and lemurs in semi-natural conditions at the Lemur Rescue Center (LRC; color-graded in relation to duration in residency) and (b) tetracycline ARGs in the GMBs of LRC lemurs. (c) Total source proportion of soil microbes from natural habitats in the GMBs of LRC lemurs. Shown are linear trend lines and 95% confidence intervals. See Table 2 for full statistical results from linear mixed models.
The source proportion of soil microbes from natural habitats in the GMBs of LRC lemurs—a proxy for covariation between lemur fecal and soil microbiomes – was also significantly and positively correlated with longer residency at the LRC (Fig. 2c; Table 2), again consistent with rewilding.
Discussion
The present study provides multiple lines of evidence that the Microbiome Rewilding Hypothesis applies not only to humans, but also to wildlife, suggesting that rewilding can serve as a tool to promote animal wellbeing in captivity or during transitional periods, including to ease the microbial reintegration of reintroduced or translocated, endangered species. Notably, for animals that fell victim to the illegal pet trade, but were then relinquished to the LRC, longer periods of exposure to naturalistic environments were strongly linked to more ‘native’ or ‘wild-type’ GMBs, as revealed by microbial community structure, resistance genes, and their covariation with environmental microbiomes. Despite clear patterns in the composition of lemur GMBs, alpha diversity was not significantly correlated with the host’s time spent in naturalistic environments; however, there was a nonsignificant trend for all alpha diversity metrics to decrease with residency at the LRC. Alpha diversity, alone, is increasingly proving to be an inconsistent metric for assessing the influences of environmental factors on host-associated microbiomes and relevant health outcomes8,33,34,35. Although data on animal health would further solidify the relevance of microbial rewilding to animal wellbeing, these results emphasize the importance of incorporating multifaceted microbiome science into animal care and conservation.
Metrics of community composition (i.e., beta diversity) well reflected the predicted and nuanced patterns of environmentally mediated microbial variation8. Specifically, longer residency at the LRC was associated with a GMB composition that was more similar to the gut communities of wild lemurs than to those of pet lemurs. The increased similarity was evidenced in both the presence-absence and the abundance-weighted metrics of phylogenetic compositions (i.e., unweighted and weighted UniFrac), indicating that both rare and abundant microbes were driving the pattern of rewilding. We thus explored specific patterns in Bacteroides, Prevotella, and Ruminococcus—three dominant members of primate GMBs8,36,37,38.
Bacteroides is a ubiquitous, diverse, and functionally relevant genus in lemur GMBs35,39, linked to polysaccharide breakdown and decreased intestinal disease in humans and animal models40,41. It is negatively influenced by the common food additives, monosaccharide fructose and glucose42. Our evidence of increased Bacteroides in the GMBs of LRC lemurs, relative to pet lemurs, could reflect the more appropriate diet provided at the LRC and, in turn, entail decreased disease risk relative to the disease-prone, pet lemurs43. Although Prevotella has saccharolytic function44 similar to Bacteroides, Prevotella was significantly decreased in LRC lemurs that had longer residency at the LRC. Both genera rely on similar nutritional resources in the gut, leading to competitive inhibition and contrasting patterns of abundance between the two genera45. This competitive relationship has led many to consider abundances of Prevotella and Bacteroides to be mutually exclusive (i.e., for these genera to be distinct enterotypes), such that the ratio of the two genera may be a proxy for microbial function, host metabolism, and gut health46,47. In humans, a lower Prevotella to Bacteroides ratio—as we see with increased residency at the LRC—has been linked to maintaining or gaining weight when consuming a high-fiber diet48. This pattern suggests that the ‘terminal’ microbiomes of LRC lemurs may facilitate or reflect a metabolic shift from malnourishment to improving body condition, achieved by allowing the animals to forage on natural vegetation while being supplemented with the produce-rich LRC diet.
The genus Ruminococcus, which was negatively correlated with longer residency at the LRC, is linked to the degradation of resistant dietary starches49, including those found in grains, such as rice50. Rice is the most widely consumed food in Madagascar and the food most commonly fed to pet lemurs. By contrast, the diets of LRC lemurs do not include rice and are not rich in starch. Importantly, the diets of LRC lemurs include natural forage, which has been shown to dramatically impact GMB diversity and function in folivorous lemurs51. Together, the changes in these three dominant taxa—Bacteroides, Prevotella, and Ruminococcus—suggest that the transition from diets associated with pethood to more natural diets at the LRC can facilitate the microbial rewilding process.
Regarding antibiotic resistance, recent studies show that ARG enrichment and propagation can occur in wildlife in the absence of direct clinical treatment with antibiotics35,52, namely through the transmission of ARGs between hosts and their social or physical environment52. Although pet lemurs in Madagascar almost never receive antibiotics, they have markedly high proportions of ARGs in their GMBs. LRC lemurs, however, are treated with antibiotics in cases of injury or disease. Despite the increased likelihood of LRC lemurs, relative to pets, receiving antibiotic treatment during veterinary care, we found that residency at the LRC, under diminished human contact, significantly correlated with lower proportions of total and tetracycline ARGs. These results suggest a potent role for human contact (or exposure to domesticated animals and their excreta) in ARG transmission to animals, such that minimizing human contact and anthropogenic disturbance would be an important step in the rewilding process.
In terms of the physical environment, beyond acquisition of environmental pathogens53, acquisition of commensal or symbiotic microbes is gaining recognition as a component of GMB assembly54. The functional relevance of these environmental microbes remains to be determined; yet, there is clear and longstanding evidence that exposure to environmental microbes, or lack thereof, plays a role in shaping animal (including human) immune responses and determining overall health outcomes5,55,56,57. In support of our previous finding that exposure to natural environments dictates environmental acquisition in lemur GMBs8, longer residency at the LRC, which equated to greater exposure to naturalistic environments, correlated with greater covariation between lemur GMBs and soil microbiomes from natural habitats. In addition to the inherent psychological and behavioral value of providing naturalistic environments for wildlife under human care, we find that exposure to rich, natural microbial landscapes has the potential to augment host-associated communities.
Together, our results suggest that microbial rewilding is a multi-faceted and gradual process that includes host-associated and environmental microbial communities. Moreover, we suggest that providing appropriate diets, minimizing contact with humans, and increasing exposure to natural environmental consortia are actionable steps that can promote microbial rewilding in captive animals. These actions may be particularly valuable for animals slated to undergo environmental transitions or reintroduction6,58. By rewilding host GMBs prior to the transition, we may be able to prime animals for success in their new environments. Going forward, the collection of longitudinal data on the GMBs and overall health of animals undergoing environmental transitions will be essential for understanding the microbial dynamics that drive microbial rewilding and their ultimate relevance to the animal host.
Data availability
The 16S sequencing reads are available in the National Center for Biotechnology Information's Sequence Read Archive (BioProject ID #PRJNA821395). Data on antibiotic resistance genes are deposited in the Open Science Framework repository, link: https://osf.io/vkr2f/, https://doi.org/10.17605/OSF.IO/VKR2F. The full metagenomic library is available upon reasonable request from the corresponding author (SLB).
References
Peixoto, R. S., Harkins, D. M. & Nelson, K. E. Advances in microbiome research for animal health. Annu. Rev. Anim. Biosci. 9, 289–311 (2021).
Amato, K. R. et al. Habitat degradation impacts black howler monkey (Alouatta pigra) gastrointestinal microbiomes. ISME J. 7, 1344 (2013).
West, A. G. et al. The microbiome in threatened species conservation. Biol. Conserv. 229, 85–98 (2019).
Robinson, J. M., Mills, J. G. & Breed, M. F. Walking ecosystems in microbiome-inspired green infrastructure: An ecological perspective on enhancing personal and planetary health. Challenges 9, 40 (2018).
Mills, J. G. et al. Urban habitat restoration provides a human health benefit through microbiome rewilding: The Microbiome Rewilding Hypothesis. Restor. Ecol. 25, 866–872 (2017).
Trevelline, B. K., Fontaine, S. S., Hartup, B. K. & Kohl, K. D. Conservation biology needs a microbial renaissance: A call for the consideration of host-associated microbiota in wildlife management practices. Proc. R. Soc. B 286, 20182448 (2019).
Dallas, J. W. & Warne, R. W. Captivity and animal microbiomes: Potential roles of microbiota for influencing animal conservation. Microb. Ecol. https://doi.org/10.1007/s00248-022-01991-0 (2022).
Bornbusch, S. L. et al. Gut microbiota of ring-tailed lemurs (Lemur catta) vary across natural and captive populations and correlate with environmental microbiota. Anim. Microbiome 4, 1–19 (2022).
Greene, L. K. et al. Gut microbiota of frugo-folivorous sifakas across environments. Anim. Microbiome 3, 39 (2021).
McKenzie, V. J. et al. The effects of captivity on the mammalian gut microbiome. Integr. Comp. Biol. 57, 690–704 (2017).
Bornbusch, S. L. & Drea, C. M. Antibiotic resistance genes in lemur gut and soil microbiota along a gradient of anthropogenic disturbance. Front. Ecol. Evol. 9, 514 (2021).
Hyde, E. R. et al. The oral and skin microbiomes of captive komodo dragons are significantly shared with their habitat. mSystems 1, e00046-e116 (2016).
LaFleur, M., Clarke, T. A., Reuter, K. E. & Schaefer, M. S. Illegal trade of wild-captured Lemur catta within Madagascar. Folia Primatol. 90, 199–214 (2019).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180 (2011).
Choo, J. M., Leong, L. E. & Rogers, G. B. Sample storage conditions significantly influence faecal microbiome profiles. Sci. Rep. 5, 16350 (2015).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–1624 (2012).
Hasan, N. A. et al. Microbial community profiling of human saliva using shotgun metagenomic sequencing. PLoS ONE 9, e97699 (2014).
Bornbusch, S. L. et al. Stable and transient structural variation in lemur vaginal, labial and axillary microbiomes: Patterns by species, body site, ovarian hormones and forest access. FEMS Microbiol. Ecol. 96, fiaa090 (2020).
Bolyen, E. et al. QIIME 2: Reproducible, interactive, scalable, and extensible microbiome data science. PeerJ 37, 852–857 (2018).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581 (2016).
Trosvik, P., Rueness, E. K., de Muinck, E. J., Moges, A. & Mekonnen, A. Ecological plasticity in the gastrointestinal microbiomes of Ethiopian Chlorocebus monkeys. Sci. Rep. 8, 1–10 (2018).
Wills, M. O. et al. Host species and captivity distinguish the microbiome compositions of a diverse zoo-resident non-human primate population. Diversity 14, 715 (2022).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2–approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Quast, C. et al. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2012).
Yarza, P. et al. Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nat. Rev. Microbiol. 12, 635 (2014).
Aitchison, J. The statistical analysis of compositional data. J. R. Stat. Soc. Ser. B 44, 139–160 (1982).
Quinn, T. P., Erb, I., Richardson, M. F. & Crowley, T. M. Understanding sequencing data as compositions: An outlook and review. Bioinformatics 34, 2870–2878 (2018).
Gloor, G. B., Macklaim, J. M., Pawlowsky-Glahn, V. & Egozcue, J. J. Microbiome datasets are compositional: And this is not optional. Front. Microbiol. 8, 2224 (2017).
Ottesen, A. et al. Enrichment dynamics of Listeria monocytogenes and the associated microbiome from naturally contaminated ice cream linked to a listeriosis outbreak. BMC Microbiol. 16, 1–11 (2016).
Lax, S. et al. Longitudinal analysis of microbial interaction between humans and the indoor environment. Science 345, 1048–1052 (2014).
Shenhav, L. et al. FEAST: Fast expectation-maximization for microbial source tracking. Nat. Methods 16, 627 (2019).
Barelli, C. et al. The gut microbiota communities of wild arboreal and ground-feeding tropical primates are affected differently by habitat disturbance. mSystems 5, e00061 (2020).
Frankel, J. S., Mallott, E. K., Hopper, L. M., Ross, S. R. & Amato, K. R. The effect of captivity on the primate gut microbiome varies with host dietary niche. Am. J. Primatol. 81, e23061 (2019).
Bornbusch, S. L. et al. Antibiotics and fecal transfaunation differentially affect microbiota recovery, associations, and antibiotic resistance in lemur guts. Anim. Microbiome 3, 65 (2021).
Amato, K. R. et al. Evolutionary trends in host physiology outweigh dietary niche in structuring primate gut microbiomes. ISME J. 13, 576–587 (2019).
Bornbusch, S. L. et al. A comparative study of gut microbiomes in captive nocturnal strepsirrhines. Am. J. Primatol. 81, e22986 (2019).
Nishida, A. H. & Ochman, H. A great-ape view of the gut microbiome. Nat. Rev. Genet. 20, 195–206 (2019).
Nagpal, R. et al. Gut microbiome composition in non-human primates consuming a Western or Mediterranean diet. Front. Nutr. 5, 28 (2018).
Deng, H. et al. Bacteroides fragilis prevents Clostridium difficile infection in a mouse model by restoring gut barrier and microbiome regulation. Front. Microbiol. 9, 2976 (2018).
Wang, C. et al. Roles of intestinal bacteroides in human health and diseases. Crit. Rev. Food Sci. Nutr. 61, 3518–3536 (2021).
Townsend, G. E. et al. Dietary sugar silences a colonization factor in a mammalian gut symbiont. Proc. Natl. Acad. Sci. USA 116, 233–238 (2019).
LaFleur, M. et al. Drug-resistant tuberculosis in pet ring-tailed lemur, Madagascar. Emerg. Infect. Dis. 27, 977 (2021).
Gálvez, E. J. C. et al. Distinct polysaccharide utilization determines interspecies competition between intestinal Prevotella spp. Cell Host Microbe 28, 838–852 (2020).
Costea, P. I. et al. Enterotypes in the landscape of gut microbial community composition. Nat. Microbiol. 3, 8–16 (2018).
Roager, H. M., Licht, T. R., Poulsen, S. K., Larsen, T. M. & Bahl, M. I. Microbial enterotypes, inferred by the prevotella-to-bacteroides ratio, remained stable during a 6-month randomized controlled diet intervention with the new nordic diet. Appl. Environ. Microbiol. 80, 1142–1149 (2014).
Hjorth, M. F. et al. Pretreatment Prevotella-to-Bacteroides ratio and markers of glucose metabolism as prognostic markers for dietary weight loss maintenance. Eur. J. Clin. Nutr. 74, 338–347 (2020).
Hjorth, M. F. et al. Pre-treatment microbial Prevotella-to-Bacteroides ratio, determines body fat loss success during a 6-month randomized controlled diet intervention. Int. J. Obes. 42, 580–583 (2018).
DeMartino, P. & Cockburn, D. W. Resistant starch: Impact on the gut microbiome and health. Curr. Opin. Biotechnol. 61, 66–71 (2020).
Wang, K. et al. Diet with a high proportion of rice alters profiles and potential function of digesta-associated microbiota in the ileum of goats. Animals 10, 1261 (2020).
Greene, L. K., McKenney, E. A., O’Connell, T. M. & Drea, C. M. The critical role of dietary foliage in maintaining the gut microbiome and metabolome of folivorous sifakas. Sci. Rep. 8, 14482 (2018).
Allen, H. K. et al. Call of the wild: Antibiotic resistance genes in natural environments. Nat. Rev. Microbiol. 8, 251–259 (2010).
Daszak, P., Cunningham, A. A. & Hyatt, A. D. Anthropogenic environmental change and the emergence of infectious diseases in wildlife. Acta Trop. 78, 103–116 (2001).
Shapira, M. Gut microbiotas and host evolution: Scaling up symbiosis. Trends Ecol. Evol. 31, 539–549 (2016).
Sbihi, H. et al. Thinking bigger: How early-life environmental exposures shape the gut microbiome and influence the development of asthma and allergic disease. Allergy 74, 2103–2115 (2019).
Bendiks, M. & Kopp, M. V. The relationship between advances in understanding the microbiome and the maturing hygiene hypothesis. Curr. Allergy Asthma Rep. 13, 487–494 (2013).
Alexandre-Silva, G. M. et al. The hygiene hypothesis at a glance: Early exposures, immune mechanism and novel therapies. Acta Trop. 188, 16–26 (2018).
Yao, R. et al. The, “wildness” of the giant panda gut microbiome and its relevance to effective translocation. Glob. Ecol. Conserv. 18, e00644 (2019).
Acknowledgements
For their assistance in field collection and logistics, we thank Lydia Greene, Marina Blanco, Samantha Calkins, Ryan Rothman, Laurent ‘Raleso’ Randrianasolo, Remi Rakotovao, Georges René Rakotonirina, Chelsea Southworth, Melina Nolas, Lauren Petronaci, Michelle Sauther, and Patricia Wright. We are also grateful to management and staff members at the LRC. We thank Karlis Graubics and Brian Fanelli at CosmosID and Sarah Owens at Argonne National Laboratory for guidance and sequencing services.
Funding
Funding was provided by an NSF Doctoral Dissertation Research Improvement Grant to SLB and CMD (award #1945776), an NSF Behavioral and Cognitive Sciences Grant to CMD (award #1749465), a Triangle Center for Evolutionary Medicine Graduate Student Research Grant to SLB, and a Research fellowship from The Kenan Institute for Ethics (Duke University) to SLB. During collections, TAC and ML were funded by the Margot Marsh Biodiversity Fund and private donations to Lemur Love (San Diego, C.A.).
Author information
Authors and Affiliations
Contributions
S.L.B.: Conceptualization, Data curation, Formal analysis, Funding acquisition, Investigation, Methodology, Project administration, Resources, Visualization, Writing—original draft, Writing—review & editing. T.A.C.: Data curation, Resources, Writing—review & editing. S.H.: Data curation, Methodology, Resources, Writing—review & editing. S.H.R.: Methodology, Resources, Writing—review & editing. M.L.: Data curation, Resources, Writing—review & editing. C.M.D.: Conceptualization, Funding acquisition, Investigation, Methodology, Project administration, Resources, Supervision, Visualization, Writing—original draft, Writing—review & editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bornbusch, S.L., Clarke, T.A., Hobilalaina, S. et al. Microbial rewilding in the gut microbiomes of captive ring-tailed lemurs (Lemur catta) in Madagascar. Sci Rep 12, 22388 (2022). https://doi.org/10.1038/s41598-022-26861-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-26861-0
This article is cited by
-
Markers of fertility in reproductive microbiomes of male and female endangered black-footed ferrets (Mustela nigripes)
Communications Biology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.