Abstract
Juvenile hormone (JH) is synthesized by the corpora allata (CA) and controls development and reproduction in insects. Therefore, achieving tissue-specific expression of transgenes in the CA would be beneficial for mosquito research and control. Different CA promoters have been used to drive transgene expression in Drosophila, but mosquito CA-specific promoters have not been identified. Using the CRISPR/Cas9 system, we integrated transgenes encoding the reporter green fluorescent protein (GFP) close to the transcription start site of juvenile hormone acid methyl transferase (JHAMT), a locus encoding a JH biosynthetic enzyme, specifically and highly expressed in the CA of Aedes aegypti mosquitoes. Transgenic individuals showed specific GFP expression in the CA but failed to reproduce the full pattern of jhamt spatiotemporal expression. In addition, we created GeneSwitch driver and responder mosquito lines expressing an inducible fluorescent marker, enabling the temporal regulation of the transgene via the presence or absence of an inducer drug. The use of the GeneSwitch system has not previously been reported in mosquitoes and provides a new inducible binary system that can control transgene expression in Aedes aegypti.
Similar content being viewed by others
Introduction
The corpora allata (CA) is the site of synthesis of juvenile hormone (JH), an essential sesquiterpenoid that controls development and reproduction in insects1. In adult female mosquitoes, the biosynthetic activity of the CA is regulated by the interaction of multiple factors that stimulate (allatotropins) or inhibit (allatostatins) CA activity1,2,3. Developing a system to drive inducible CA-specific expression in mosquitoes would facilitate the study of modulatory factors of JH synthesis, as well as contribute to the identification of targets for designing new, specific, and affordable strategies suitable for mosquito control4,5. For instance, mosquito lines that ectopically express endogenous/foreign genes or dsRNAs could be generated, resulting in developmental or reproductive changes. Dissecting the roles of these critical genes requires manipulation of their functions in different tissues and at different developmental stages, which can be technically challenging.
An extensive collection of genetic tools supports the activity of researchers working with Drosophila6,7. Among them, the GAL4 enhancer trap system was used to identify the Aug21 locus, described later as a CA-specific driver in the larval stages of Drosophila8. Many of these genetic resources are now available for non-model organisms such as mosquitoes9. Spatial control of gene expression in mosquitoes has been implemented in tissues such as olfactory neurons, midgut, and fat-body10,11,12,13. However, approaches to achieving mosquito CA-specific expression have not yet been described.
Binary systems have been previously used in mosquitoes, such as the GAL4/UAS and QF2/QUAS11,13. Still, more are needed for intersectional strategies that require the expression of multiple transgenes in distinct or overlapping groups of cell types. In addition to cell-specificity, achieving temporal control of transgenes adds another level of complexity over constitutive expression. Tetracycline-controlled Tet-Off and Tet-On gene expression systems have been used in several insect systems, including Drosophila and mosquitoes14,15. Due to the potential "off-target" effects of tetracycline and its analogs16, and a certain level of promoter "leakiness," the inducible GeneSwitch system could offer improvements. This technique is a modified version of the classic GAL4/UAS system, which allows inducible gene expression in Drosophila under the control of the progesterone analog RU-48617,18.
In the present study, we tested the hypothesis that we could link the activity of a strong CA promoter to drive the CA-specific expression of GFP in Aedes aegypti mosquitoes. We targeted the transcription start site of juvenile hormone acid methyl transferase (JHAMT), a JH biosynthetic enzyme that is specifically and highly expressed in the adult female CA19,20. Using the CRISPR/Cas9 system, we integrated a promoter-less transgene encoding eGFP (enhanced GFP) close to the transcription start site of the jhamt locus. Transgenic mosquitoes showed specific eGFP expression in the CA. In addition, CRISPR/Cas9 was also used to develop GeneSwitch driver and responder mosquito lines expressing an inducible fluorescent marker, enabling the temporal regulation of GFP expression via the presence or absence of RU-486.
Our lines offer a model to investigate the spatial and temporal patterns of the jhamt promoter. In addition to basic mosquito physiology research, the lines provide valuable genetic tools to investigate CRISPR-based technologies for mosquito population control. Overall, the findings and resources reported here expand the genetic toolkit for mosquito research.
Results
Construction of a plasmid vector for CRISPR/Cas9-mediated mutagenesis
We first tested the hypothesis that we could link the activity of a robust CA-specific promoter to drive the CA-specific expression of a transgene. We employed CRISPR/Cas9-mediated homology-dependent repair (HDR) to introduce an eGFP marker into the first exon of the Ae. aegypti jhamt locus as previously described21. Subsequently, the expression of eGFP was monitored to evaluate if the transgene would display the spatiotemporal features of expression of the jhamt gene.
The plasmid pSL1180 (Amersham, UK) was used as a backbone to make the pSL_jhamt_eGFP construct. Integration of the transgene was guided into the first exon of the jhamt locus with two homologous sequences flanking the CRISPR target sites, a 1573 bp jhamt left arm and a 1511 bp jhamt right arm. The inserted cassette comprised a 1008-bp eGFPSV40 promoter-less green fluorescent reporter and a 1241-bp fragment containing 3xP3dsRedSV40, a fluorescent red eye-specific marker used for visualization and selection of transformants11 (Fig. 1A). A synthetic intron IVS8 sequence was included in the first exon of jhamt prior to the start codon to enhance mRNA nuclear export.
Integration of a promoter-less transgene encoding eGFP into the jhamt locus. (A) Diagram of the pSL_jhamt_eGFP construct. eGFP was flanked by the synthetic intron IVS8. dsRed (RFP) was used as a selection marker under the control of 3xP3, which drives expression in the eye. The insert was surrounded by two 1.5 kb jhamt locus homologous arms to facilitate recombination. (B) Image of head and thorax of a first instar larvae showing expression of dsRed in eyes and eGFP in the CA. Scale bar, 100 µm.
To generate jhamt mutants, embryos of Ae. aegypti (Orlando strain) were injected with the pSL_jhamt_eGFP construct21. jhamt−/− mutants never survived to adulthood but died as L4 instar just before pupation21. A stable heterozygous jhamt+/− line was established after six backcrosses to wild type (WT) mosquitoes21. Transformants expressed dsRed in the eyes and eGFP in the CA (Fig. 1B).
Tissue-specific expression of eGFP recapitulates jhamt expression
Real-time quantitative PCR (RT-qPCR) was used to analyze if eGFP expression recapitulates the transcript tissue specificity expression of jhamt19,20 (Fig. 2). eGFP and jhamt mRNA titers were evaluated in different tissues of WT and heterozygous jhamt adult female mutants exhibiting either eGFP+ or eGFP- corpora allata. As previously described, jhamt mRNA expression was very high in the CA, with trace amounts present in the brain, ovaries, and midgut19,20; likewise, substantial amounts of eGFP mRNA were only detected in CA, although transcripts levels were less than 1% of those detected for jhamt (Fig. 2).
Tissue-specific expression of jhamt and eGFP. Real-time qPCR measurement of jhamt and eGFP mRNA levels in tissues of 3–5 days old sugar-fed WT and jhamteGFP+/− adult female exhibiting either eGFP+ or eGFP− CA. CA WT: corpora allata wild-type; CA R+G−: heterozygote with red eyes and CA not green; CA R+G+: heterozygote with red eyes and green CA; OV ovaries, BR brain, MG midgut (all tissues dissected from insects with green CA). Levels of mRNAs are expressed as copy numbers per 10,000 copies of RpL32 mRNA. Each data point is an average of 3 independent biological replicates of tissues from 5 to 10 insects.
To further evaluate the spatial expression of eGFP, the offspring of twenty-five eGFP+ females of G7–G10 generations (n = 1922, first instar larvae) were analyzed. Surprisingly, we detected four different "bilaterally asymmetric" phenotypes: insects (1) with both CA eGFP− (77%), (2) with only a left CA eGFP+ (10%), (3) with only a right CA eGFP+ (10%) and (4) with both CA eGFP+ (3%) (Fig. 3).
Asymmetric eGFP expression in CA: (A) Images of the four phenotypes of dsRed+ first instar larvae expressing eGFP in the CA. −/−: both CA GFP negative; +/−: only right CA eGFP positive; −/+: only left CA eGFP positive; +/+: both CA eGFP positive. Scale bar, 100 µm. (B) Percentages of individuals expressing eGFP in the CA. −/−: both CA eGFP negative, +/−: only right CA eGFP positive, −/+: only left CA eGFP positive. +/+: both CA eGFP positive. Scale bar, 100 µm.
We subsequently evaluated the relationship between the expression of eGFP by each individual member of a CA pair, and each member's ability to synthesize JH in our ex vivo assay22. eGFP positive and eGFP negative CAs were dissected (Fig. 4a), and the amount of JH III synthesized by each individual gland ex vivo was analyzed by LC–MS/MS23. There was no correlation between the ability of the CA to synthesize JH and the eGFP expression in an individual CA (Fig. 4b).
Lack of correlation between expression of eGFP and JH biosynthetic activity. Individual CAs were dissected from three insects expressing eGFP in only one CA, and JH biosynthesis was evaluated individually in each individual member of a CA pair. (A) Images of dissected CA, (a) before and (b) after separation of individual CA. (B) Amount of synthesized JH expressed as fmol of JH III/CA/h. Green bars: CA from GFP+, Black bars: CA from GFP-. Scale bar, 100 µm.
Temporal profile of eGFP expression
Levels of jhamt mRNA in WT are high in early 4th instar larvae, undetectable in late 4th instar and pupa, elevated in sugar-fed adult females, and low again in blood-fed females19,20,21. Using a fluorescent microscope, we evaluated the expression of eGFP during the life cycle of jhamt+/− mosquitoes. We detected expression of eGFP in all the stages examined, including late 4th instar larvae and pupae, two periods where jhamt mRNA expression is never detected (Fig. 5). Theoretically, the fluorescent protein reporter should exhibit complete concordance with the expression of the endogenous gene, except perhaps in cases in which reporter visualization is due to eGFP perdurance24. To evaluate this possibility, we selected pupae expressing GFP and evaluated eGFP mRNA titers. We did not detect eGFP mRNA in early pupae that were GFP positive, confirming that the reporter visualization was due to GFP perdurance (Supp. Fig. 1).
Stage-specific expression of eGFP. Fluorescent microscopy was employed to detect eGFP in the CA at all stages of the mosquito life cycle. Individual eGFP-positive CA were observed through the cuticle of larvae. Afterward, CA were dissected from late 4th instar larva, pupa, or adults to analyze in detail the presence of eGFP. eGFP was present only in some cells of the CA. Autofluorescence was detected neither in the CA of WT nor in most cells of the CA of mutants. Scale bar, 50 µm.
To evaluate if the "asymmetric" expression of eGFP changed during development, we followed the expression of eGFP in 12 individual mosquitoes during their entire life cycle (n = 3 for each of the four phenotypes). We established that the phenotype of eGFP expression remained constant within but not across mosquitoes; a particular insect expressed eGFP in one (left or right), both or none of the CA from larvae to adult. In addition, the expression of eGFP was sporadic within the CA. Fluorescence was detected only in some cells of the CA, with the number of eGFP-positive cells varying significantly among individual transgenic animals (Fig. 5).
Development of an inducible GeneSwitch system (GS)
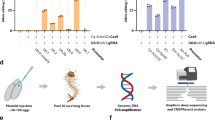
CRISPR/Cas9 was also used to develop driver and responder mosquito lines expressing a transgene that could be activated by providing an inducer drug, RU486. The driver line was transformed with a pSL_jhamt_GS plasmid, which was also integrated into the first exon of the jhamt locus using the identical jhamt homologous sequences employed for the pSL_jhamt_eGFP construct, but with the eGFP sequence replaced by a GeneSwitch Gal4 driver sequence (Fig. 6A). Transformants expressed dsRed in the eyes using the 3xP3 promoter.
Development of an inducible GeneSwitch system. (A) Diagram of the driver and responder constructs. Driver: GenSwitch locus. dsRed (red eyes) was used as a selection marker under the control of 3xP3. The insert was surrounded by two 1.5 kb jhamt locus homologous arms to facilitate recombination. Responder: 2xUAS locus. GFP with a membrane localization signal (mCD8). eCFP (blue eyes) was used as a selection marker under the control of 3xP3. The insert was surrounded by two 1.5 kb epox locus homologous arms to facilitate recombination. (B) Crossing schemes to generate a hybrid. jhamtGS+/− females were crossed with epoxUASGFP+/− males to generate hybrids containing both the effector and responder genes (jhamtGS+/−; epoxUASGFP+/−). Hybrids were used to test inducibility by RU486.
To build the responder line, we integrated a transgene into the fourth exon of the epox locus, which encodes the methyl farnesoate epoxidase (EPOX). EPOX is another JH biosynthetic enzyme that is specifically and highly expressed in the CA20,25. The pSL_2xUAS_mCD8GFP_3xP3eCFP plasmid was modified to include epox homologous sequences flanking the CRISPR target sites, a 1540 bp (left) epox arm and a 1508 bp (right) epox arm21 (Fig. 6A). A stable heterozygous epoxUASGFP+/− line was established after six backcrosses to wild-type (WT) mosquitoes. Transformants expressed CFP in the eyes.
A homozygous driver line could not be established since jhamtGS/GS insects died as 4th instar larvae. In addition, the homozygous responder line epoxUASGFP/UASGFP had severe physiological defects21; therefore, heterozygous driver and responder lines were used to evaluate the inducibility of UAS-driven GFP by RU486. Since the expression of eGFP in the CA of jhamt eGFP+/− insects was sporadic, and only about 25% of the insects displayed eGFP+, RT-qPCR could not be used to assess the inducibility of UASGFP by RU486. Instead, the ability to induce GFP in 4th instar larvae, pupae, and adult mosquitoes was qualitatively evaluated using a fluorescent microscope. jhamtGS+/− females were crossed with epoxUASGFP+/− males and F1 1st instar larvae were screened for dsRed, and eCFP fluorescence in their eyes (Fig. 6B). Individual dsRed+ and eCFP+ insects were exposed to RU486 as either 4th instar larvae, pupae, or adults. We detected a strong GFP inducibility in 4th instar larvae and pupae with little background before induction. On the other hand, in adults, we also observed a weak GFP signal in non-induced animals (Fig. 7), indicating that the GS inducible expression system in these adult mosquitoes can be leaky. In all cases, we only observed GFP expression in a few adult female CA cells, a sporadic pattern similar to that observed in the jhamteGFP+/− line, confirming that the expression of GS inserted in the jhamt transcription start locus was regulated similarly to the expression of eGFP in the jhamt eGFP+/− line.
Discussion
A few tissue-specific promoters have been described in mosquitoes to drive the expression of transgenes in tissues such as the midgut10, salivary glands26, fat body27, and testis28; nevertheless, promoters that drive mosquito CA-specific expression have not been reported. In contrast, different promoters have been used to drive CA-specific expression in Drosophila, with the jhamt promoter displaying more robust CA-specific expression than the Aug21 locus29,30. The present series of experiments used the CRISPR/Cas9 approach to assess the ability of the jhamt promoter to drive the specific expression of eGFP in the mosquito CA gland. We observed that the introduced transgene exhibited some of the features of endogenous gene expression. The tissue-specificity represented wild-type patterns of jhamt expression. Initially, we thought the temporal-specificity was lacking, as we observed the presence of GFP in pupae. Subsequently, we uncovered that the presence of GFP in the pupa stage resulted from eGFP perdurance22, because no de novo transcription of jhamt or GFP occurred at this stage. Perdurance is a common problem with in vivo reporters, and in some cases, the GFP signal lasted for about 4–5 days after the disappearance of the mRNA signal31. In summary, GFP perdurance could have confounded our assessment of the transgene expression in some stages.
In addition, the eGFP mRNA expression was much lower than jhamt expression, although the transgene expression was not particularly low. The strength of the native jhamt promoter is extremely high20. The amount of jhamt mRNA in the CA is seven times larger than the amount of ribosomal protein L32 mRNA (the expression level of jhamt is around 70,000 copies/10,000 copies of rpL32). The eGFP transgene was expressed at 500 copies/10,000 copies of rpL32. To provide some context, half of the JH synthetic enzyme pathway transcripts were expressed at the same or lower level than the eGFP transgene in the CA20. Nevertheless, the intensity of the epifluorescent signal allowed the visualization of the "green CA" through the cuticle of live larvae from first to fourth-instar and dissected CA from pupae and adult mosquitoes.
In the locust Schistocerca gregaria, JH biosynthesis by the paired CA gland was described as remarkably "asymmetric" with respect to the relative JH synthetic activity of either member of the pair32. This apparently random pattern was observed at all ages studied32. We also observed a bilateral asymmetry of eGFP expression in the two individual CA of a single insect, as well as an unequal distribution among the four "asymmetric phenotypes." The vast majority of larvae (~ 80%) did not show any fluorescent signal in either of the two glands, and only about 3% of the larvae were fluorescent in both glands. Remarkably, a particular phenotype remained invariable during the entire life cycle of an individual mosquito. We cannot explain the molecular basis or significance of this asymmetry. There was no correlation between the expression of jhamt and GFP in individual glands, which suggests a complex mechanism for jhamt expression, perhaps involving the spatial dynamics of cis-acting elements within the jhamt gene itself.
Expression of GFP was characteristically observed in a random cellular distribution within the CA that normally express jhamt. The local chromatin environment at the site of transgene insertion can alter both the pattern and the level of transgene expression33,34. Although integration sites in the genome might result in uniform expression in a particular cell type; often transgene expression is stochastic; that is, not all cells of a given type express the transgene, and the precise cells showing expression vary from individual to individual, frequently because of the difficulty in overcoming chromatin blocks to initiating transcription33,34. Mechanisms involving chromatin modification may be responsible for the silencing of genes in position effects resulting from the insertion of transgenes. The results presented here suggest the action of silencer-like elements affecting GFP expression at the chromosomal insertion site in a cell-specific manner in transformed mosquitoes.
It is also possible that the low and unpredictable levels of eGFP expression we observed were due to technical features of the construct and that the potential strength of the jhamt promoter was underestimated as a result. For example, a synthetic IVS8 sequence was included in the first exon of jhamt prior to the start codon because it is a standard component of the pSwitch system (Invitrogen). Short introns are routinely added to transgenes to enhance mRNA nuclear export in many eukaryotes, including Drosophila and Aedes albopictus C6/36 cells34,35,36. The IVS8 fragment included canonical eukaryotic donor and acceptor splice sites and their respective consensus flanking sequences37. The sequence of this fragment was verified in G4 mosquito lines, and no mutations were detected. However, proteins such as transcription factors or splicing factors that normally interact with the jhamt 5′-UTR could have conceivably interfered with the expected splicing of the IVS8 intron in such a way as to excise eGFP from the mature transcript, using an aberrant and perhaps unconventional 3′ splice site38. We tested if inappropriate splicing due to the presence of the artificial intron was reducing the observed transgene expression. Quite the opposite, the amount of transcribed 5'UTR upstream of IVS8 (Supl. Fig. 2) corresponded well to the amount of transcribed eGFP or transcribed jhamt (Fig. 2), and it was in agreement with the visual presence or absence of eGFP, detected using the fluorescence microscope. We concluded that the transgene expression was defined primarily by the transcription efficiency and not by incorrect splicing due to the presence of the IVS8 intron.
Many experiments in biological research rely critically on the ability to express exogenous proteins or RNAs in transgenic animals in a manner that is regulated for level, timing, and cell type. Enhancer trapping has been widely used to identify tissue-specific promoters in Drosophila39, often resulting in the reporter having spatial and temporal expression identical to the patterns of the associated enhancer region. A PiggyBac transposon-based enhancer-trap system was developed for the mosquito Anopheles stephensi40 and used to generate enhancer-trap lines expressing eGFP in adult female salivary glands, midgut, and fat body40. While Drosophila and Anopheles reporter genes have frequently been shown to respond to neighboring transcriptional regulatory elements, gene regulatory mechanisms may be more complex in Ae. Aegypti—with a genome size five times larger than either model system41,42. The longer intronic and intergenic sequences in Aedes may interfere with the ability of the transcriptional apparatus to co-opt the precise spatial and temporal dynamics of gene expression that are required for successful enhancer traps.
By inserting our eGFP construct within the open reading frame of the jhamt gene, we disrupted the structure of the locus. Although some fundamental elements of the jhamt promoter drove the transgene expression in the correct tissue (CA) at the expected developmental stages (adult females), other critical regulatory components were not preserved in our design.
Unexpectedly, the expression of eGFP in the CA of an individual female mosquito did not predict the expression of the reporter in her progeny. Having an eGFP+ or eGFP- female parent did not change the proportion of eGFP+ in the offspring, suggesting that the pattern of GFP expression is defined de novo in each generation and may be influenced by epigenetic factors. The role of epigenetic factors in the regulation of the mosquito CA is currently unknown. Our mosquito lines could provide a model system to investigate epigenetic modifications that could underlie intergenerational differences in eGFP expression in Ae. aegypti CA.
The ultimate aim of these experiments was to drive CA-specific expression in mosquitoes with defined temporal and spatial patterns. For that reason, it would be desirable to develop a conditional expression system that is activated only by the presence of an inducer. GeneSwitch has been demonstrated to function in Drosophila17, albeit the system can be leaky depending on the nature of the promoter and responder transgene18. In our case, the expression of GFP in the absence of the inducer RU-486 was low in larvae and pupae, and the inducer clearly activated the reporter gene. In adults, the expression of GFP in the absence of an inducer was more noticeable. Thus, in cases where tight regulation is required, it might be advisable to explore a different system that has been validated in dipterans, including mosquitoes, such as the Q-system43,44,45 and T2A46,47.
In summary, we report a set of experiments in which we empirically tested a strategy to direct exogenous gene expression in the CA of mosquitoes. Our data suggest that we have not developed an unequivocally useful general tool to direct the expression of transgenes. Whether this is due to the approach, the construct, or a peculiarity of this particular locus would require additional studies. Nevertheless, given the widespread need for these methods, we expect our results will be a useful guide to generate improved approaches in the future. If the immediate goal were to use the jhamt promoter to drive the expression of transgenes in wild populations, the lines we generated are a first step in this process and require additional optimization. However, we have generated stable mosquito lines that provide tools for investigating the spatial and temporal patterns of the jhamt promoter. Although the concept of harnessing the tissue-specific regulation by inserting a promoter-less transgene inside a locus of a highly expressed CA gene did not render the expected spatiotemporal expression, our intriguing expression patterns revealed that jhamt is regulated by a complex interaction of promoter and enhancers that cannot be "captured" by approaches that drive specific expression in other mosquito tissues10,25,26,27. Nevertheless, our findings extend the genetic toolkit in Aedes aegypti by demonstrating the efficacy of the GeneSwitch binary system to control gene expression spatially and temporally. This technique can enhance our understanding of this important disease vector and can be applied to develop genetic mosquito control methods.
Methods
Insects
Ae. aegypti of the Orlando strain were reared at 28 °C and 80% humidity as previously described48. Adult mosquitoes were offered a cotton pad soaked in a 10% sucrose solution. Four-day-old female mosquitoes were fed pig blood equilibrated to 37 °C, and ATP was added to the blood meal to a final concentration of 1 mM immediately before use, as previously described48.
CRISPR/Cas9 reagents and generation of jhamt dsRed and epox eCFP mutant lines
Homology-directed repair (HDR) was utilized to introduce gene cassettes encoding fluorescent markers into the first exon of jhamt and the fourth exon of epox genes, as previously described21. Stable germline integrations were generated by injecting CRISPR/Cas9 reagents into the posterior end of preblastoderm Ae. aegypti embryos21. One line with a verified insertion site was selected for each transgene, backcrossed to the WT strain for five generations, and used for experiments starting at G621. Detailed steps of construct design are included as Supplemental Materials.
Induction of GeneSwitch with RU-486
RU-486 was diluted in ethanol (1 mg/ml) and directly added to the water, where early 4th instar larvae were raised at a final concentration of 1% ethanol (10 µg/ml RU-486).
To induce GeneSwitch in the adult mosquitoes, RU-486 was mixed with the sugar meal to get a final concentration of 1% ethanol (5 µg/ml RU-486) and provided ad-libitum to insects from eclosion. Control insects received the same concentration of ethanol (1%).
Quantitative PCR (RT-qPCR)
Total RNA was isolated using the Norgen Biotek Total RNA purification kit, treated with DNase I, and reverse-transcribed using the Verso cDNA Synthesis Kit (Thermo Fisher). The number of mRNA copies normalized to rpL32 mRNA was quantified in triplicate reactions in a 7300 Real-Time PCR System using the TaqMan Universal PCR MasterMix (Applied Biosystems)21. Gene accession numbers and primer and probe sequences are listed in Table S1.
JH biosynthesis assay
Mosquitoes were cold-anesthetized, and the CA complexes were dissected and incubated at 32 °C for 4 h in 150 μl of M-199 tissue culture medium (Gibco) containing 2% Ficoll, 25 mM Hepes (pH 6.5), and 100 μM methionine22. JH III in the tissue culture medium was analyzed by LC–MS/MS, as previously described23.
Fluorescent microscopy
The patterns of eGFP, dsRed and eCFP expression were determined by microscopic observations of larvae, pupae, and adults using a fluorescent dissecting microscope equipped with optical filters for Cy3, eGFP, and simultaneous visualization of both colors. The evaluation of eGFP expression in CA was performed on unfixed preparations using a DM 5500 B Leica fluorescence microscope with a Leica DFC 310 FX mounted camera and Leica LAS imaging software. Autofluorescence was detected neither in the CA of WT nor in most cells of the CA of mutants.
Data availability
All data generated or analyzed during this study are included in this published article (and its Supplementary Information files).
References
Rivera-Pérez, C., Clifton, M. E., Noriega, F. G. & Jindra, M. Juvenile hormone regulation and action. Adv. Invertebr. Endocrinol. 2, 1–76 (2020).
Noriega, F. G. Juvenile hormone biosynthesis in insects: What is new, what do we know, what questions remain?. ISRN. https://doi.org/10.1155/2014/967361 (2014).
Zhu, J. & Noriega, F. G. The role of juvenile hormone in mosquito development and reproduction. Adv. Insect Physiol. 51, 93–113 (2016).
Dong, S., Dong, Y., Simões, M. L. & Dimopoulos, G. Mosquito transgenesis for malaria control. Trends Parasitol. 38, 54–66 (2021).
Wang, G. H. et al. Combating mosquito-borne diseases using genetic control technologies. Nat. Commun. 12, 4388 (2021).
Venken, K. J. & Bellen, H. J. Emerging technologies for gene manipulation in Drosophila melanogaster. Nat. Rev. Genet. 6, 167–178 (2005).
Xu, R. G. et al. Perspectives on gene expression regulation techniques in Drosophila. J. Genet. Genomics. 46, 213–220 (2019).
Mirth, C., Truman, J. W. & Riddiford, L. M. The role of the prothoracic gland in determining critical weight for metamorphosis in Drosophila melanogaster. Curr. Biol. 15, 1796–1807 (2005).
Adolfi, A. & Lycett, G. J. Opening the toolkit for genetic analysis and control of Anopheles mosquito vectors. Curr. Opin. Insect Sci. 30, 8–18 (2018).
Moreira, L. A. et al. Robust gut-specific gene expression in transgenic Aedes aegypti mosquitoes. Proc. Natl. Acad. Sci. USA 97, 10895–10898 (2000).
Kokoza, V. A. & Raikhel, A. S. Targeted gene expression in the transgenic Aedes aegypti using the binary Gal4-UAS system. Insect Biochem. Mol. Biol. 41, 637–644 (2011).
Lynd, A. & Lycett, G. J. Development of the bi-partite Gal4-UAS system in the African malaria mosquito, Anopheles gambiae. PLoS ONE 7, e31552 (2012).
Riabinina, O. et al. Organization of olfactory centres in the malaria mosquito Anopheles gambiae. Nat. Commun. 7, 13010 (2016).
Stebbins, M. J. et al. Tetracycline-inducible systems for Drosophila. Proc. Natl. Acad. Sci. USA 98, 10775–10780 (2001).
Lycett, G. J., Kafatos, F. C. & Loukeris, T. G. Conditional expression in the malaria mosquito Anopheles stephensi with Tet-On and Tet-Off systems. Genetics 167, 1781–1790 (2004).
Moullan, N. et al. Tetracyclines disturb mitochondrial function across Eukaryotic models: A call for caution in biomedical research. Cell Rep. 10, 1681–1691 (2015).
Osterwalder, T., Yoon, K. S., White, B. H. & Keshishian, H. A. Conditional tissue-specific transgene expression system using inducible GAL4. Proc. Natl. Acad. Sci. USA 98, 12596–12601 (2001).
Scialo, F., Sriram, A., Stefanatos, R. & Sanz, A. Practical recommendations for the use of the GeneSwitch Gal4 system to knock-down genes in Drosophila melanogaster. PLoS ONE 11(8), e0161817 (2016).
Mayoral, J. G. et al. Molecular and functional characterization of a juvenile hormone acid methyltransferase expressed in the corpora allata of mosquitoes. Insect Biochem. Mol. Biol. 39, 31–37 (2009).
Nouzova, M., Edwards, M. J., Mayoral, J. G. & Noriega, F. G. A coordinated expression of biosynthetic enzyme controls the flux of juvenile hormone precursors in the corpora allata of mosquitoes. Insect Biochem. Mol. Biol. 41, 660–669 (2011).
Nouzova, M. et al. Epoxidation of juvenile hormone was a key innovation improving insect reproductive fitness. Proc. Natl. Acad. Sci. USA 118(45), e2109381118 (2021).
Li, Y., Hernández-Martínez, S., Unnithan, G. C., Feyereisen, R. & Noriega, F. G. Activity of the corpora allata of adult female Aedes aegypti: Effects of mating and feeding. Insect Biochem. Mol. Biol. 33, 1307–1315 (2003).
Ramirez, C. E., Nouzova, M., Michalkova, V., Fernandez-Lima, F. & Noriega, F. G. Common structural features facilitate the simultaneous identification and quantification of the five most common juvenile hormones by liquid chromatography-tandem mass spectrometry. Insect Biochem. Mol. Biol. 116, 103287 (2020).
Kwon, S. G. & Hadjantonakis, A. K. Eomes::GFP: A tool for live imaging cells of the trophoblast, primitive streak, and telencephalon in the mouse embryo. Genesis 45, 208–217 (2007).
Dermauw, W., Van Leeuwen, T. & Feyereisen, R. Diversity and evolution of the P450 family in arthropods. Insect Biochem. Mol. Biol. 127, 103490 (2020).
Yoshida, S. & Watanabe, H. Robust salivary gland-specific transgene expression in Anopheles stephensi mosquito. Insect Mol. Biol. 15, 403–410 (2006).
Nirmala, X. et al. Functional characterization of the promoter of the vitellogenin gene, AsVg1, of the malaria vector, Anopheles stephensi. Insect Biochem. Mol. Biol. 36, 694–700 (2006).
Smith, R., Walter, M., Hice, R., O’Brochta, D. & Atkinson, P. Testis- specific expression of the b2 tubulin promoter of Aedes aegypti and its application as a genetic sex-separation marker. Insect Mol. Biol. 16, 61–71 (2007).
Abdou, M. et al. Drosophila Met and Gce are partially redundant in transducing juvenile hormone action. Insect Biochem. Mol. Biol. 41, 938–945 (2011).
Wen, D. et al. Methyl farnesoate plays a dual role in regulating Drosophila metamorphosis. PLoS Genet. 11, e1005038 (2015).
Figure, T. E. et al. 5HTR3A-driven GFP labels immature olfactory sensory neurons. Comp. Neurol. 525, 1743–1755 (2017).
Tobe, S. S. Asymmetry in hormone biosynthesis by insect endocrine glands. Can. J. Zool. 55, 1509–1514 (1977).
Kirkpatrick, R. B., Parveen, Z. & Martin, P. F. Isolation of silencer-containing sequences causing a tissue-specific position effect on alcohol dehydrogenase expression in Drosophila melanogaster. Dev. Genet. 15, 188–200 (1994).
Pfeiffer, B. D. et al. Refinement of tools for targeted gene expression in Drosophila. Genetics 186, 735–755 (2010).
Duncker, B. P. et al. Introns boost transgene expression in Drosophila melanogaster. Mol. Gen. Genet. 254, 291–296 (1997).
Zieler, H. & Huynh, C. Q. Intron-dependent stimulation of marker gene expression in cultured insect cells. Insect Mol. Biol. 11, 87–95 (2002).
Baten, A. K. M. A. et al. Splice site identification using probabilistic parameters and SVM classification. BMC Bioinform. 7(5), 15 (2006).
Sibley, C. R., Blazquez, L. & Ule, J. Lessons from non-canonical splicing. Nat. Rev. Genet. 17, 407–421 (2016).
Brand, A. H. & Perrimon, N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118, 401–415 (1993).
O’Brochta, D. A., Pilitt, K. L., Harrell, R. A., Aluvihare, C. & Alford, R. T. Gal4-based enhancer-trapping in the malaria mosquito Anopheles stephensi. G3 2(11), 1305–1315 (2012).
Nene, V. et al. Genome sequence of Aedes aegypti, a major arbovirus vector. Science 316, 1718–1723 (2007).
Matthews, B. J. et al. Improved reference genome of Aedes aegypti informs arbovirus vector control. Nature 563, 501–507 (2018).
Potter, C. J., Tasic, B., Russler, E. V., Liang, L. & Luo, L. The Q system: A repressible binary system for transgene expression, lineage tracing, and mosaic analysis. Cell 141, 536–548 (2010).
del Valle Rodriguez, A., Didiano, D. & Desplan, C. Power tools for gene expression and clonal analysis in Drosophila. Nat. Methods. 9, 47–55 (2012).
Riabinina, O. & Potter, C. J. The Q-system: a versatile expression system for Drosophila. Methods Mol. Biol. 1478, 53–78 (2016).
Diao, F. & White, B. H. A novel approach for directing transgene expression in Drosophila: T2A-Gal4 in-frame fusion. Genetics 190, 1139–1144 (2012).
Matthews, B. J., Younger, M. A. & Vosshall, L. B. The ion channel ppk301 controls freshwater egg-laying in the mosquito Aedes aegypti. Elife 8, e43963 (2019).
Rivera-Perez, C., Nouzova, M., Lamboglia, I. & Noriega, F. G. Metabolic analysis reveals changes in the mevalonate and juvenile hormone synthesis pathways linked to the mosquito reproductive physiology. Insect Biochem. Mol. Biol. 51, 1–9 (2014).
Acknowledgements
This research was funded by Project 22-21244S from the Czech Science Foundation, Czech Republic to MN, and Grants R21AI167849 to FGN and R21AI153689 to MN from the National Institutes of Health-NIAID, USA to FGN. We thank Dr. René Feyereisen for his critical comments.
Author information
Authors and Affiliations
Contributions
M.N. designed research; M.N., M.J.E., M.D., D.L., L.V.T., F.F.-L. performed research; M.N. and F.G.N. analyzed data; and M.N., M.J.E., M.D. and F.G.N. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nouzova, M., Edwards, M.J., DeGennaro, M. et al. Genetics tools for corpora allata specific gene expression in Aedes aegypti mosquitoes. Sci Rep 12, 20426 (2022). https://doi.org/10.1038/s41598-022-25009-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-25009-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.