Abstract
Honeybee health and the species’ gut microbiota are interconnected. Also noteworthy are the multiple niches present within hives, each with distinct microbiotas and all coexisting, which we termed “apibiome”. External stressors (e.g. anthropization) can compromise microbial balance and bee resilience. We hypothesised that (1) the bacterial communities of hives located in areas with different degrees of anthropization differ in composition, and (2) due to interactions between the multiple microbiomes within the apibiome, changes in the community of a niche would impact the bacteria present in other hive sections. We characterised the bacterial consortia of different niches (bee gut, bee bread, hive entrance and internal hive air) of 43 hives from 3 different environments (agricultural, semi-natural and natural) through 16S rRNA amplicon sequencing. Agricultural samples presented lower community evenness, depletion of beneficial bacteria, and increased recruitment of stress related pathways (predicted via PICRUSt2). The taxonomic and functional composition of gut and hive entrance followed an environmental gradient. Arsenophonus emerged as a possible indicator of anthropization, gradually decreasing in abundance from agriculture to the natural environment in multiple niches. Importantly, after 16 days of exposure to a semi-natural landscape hives showed intermediate profiles, suggesting alleviation of microbial dysbiosis through reduction of anthropization.
Similar content being viewed by others
Introduction
The Western honey bee (Apis mellifera) is at risk1,2 due to multiple stressors, including but not limited to poor nutrition, pesticides, and pathogens3. Though not unique to agricultural areas, these stressors are typical of anthropized lands (i.e. areas under human influence), where most beekeeping practices are performed. Thus, anthropized areas are an ideal place to study honey bee microbiota dysbiosis and its recovery. The microbial realm associated with honey bees and their beehives is essential to maintain host and community homeostasis4,5, and the host immune response is intrinsically associated with native gut microbiota4,6. Symbiotic interactions between gut microbiota and bees can be highly specialised, whereby some bacteria and fungi prevent pathogen colonisation, clear toxic metabolites, or metabolise substances otherwise unavailable to bees4,6,7,8,9,10,11,12. The core gut microbiota of honey bee workers is composed of 8–10 bacterial species13 and can be disrupted by external microbes, nutrient shortage or chemicals14,15,16, with the latter harming honey bees at all stages of development17,18,19,20. Dysbiosis of the intestinal flora can compromise honey bee resilience and cause disease onset, while positive reinforcement induces recovery of beneficial microbial profiles and strengthens host health6. Recuperation of the core microbiota could be achieved through probiotic supplementation of core gut bacteria21. However, there is no consensus on what causes gut microbiota dysbiosis, its regulation and recovery, and how it affects (and is affected by) other hive microbial communities.
Within the hive, multiple microenvironments or niches (e.g. food reserves), each with their unique microbiotas, coexist alongside bees and their gut. These niche microbiotas are non-isolated and can undergo alterations when encountering other microbial communities or stressors. Habitat and bee diet can also affect some of these niche communities, as seen in corbicular bee pollen22 and in bee bread, where the microbial community composition changes from initial environmental profiles towards bee bread-specific communities23. It is equally important to consider the effect of stressors, habitat and diet, in other niches such as airborne particles or hive surfaces. Pesticides not only directly affect honey bees but can also accumulate to hazardous concentrations within hive components20,24, disrupting the microbiota of these niches and weakening colonies18. Social interactions among colony integrants could also facilitate distribution of microbes within hives25, as seen with the Arsenophonus genus26, or enable intercolony pathogen transmission27,28. Due to the close relationship between bees, the colony and hive components, stressor-mediated dysbiosis of hive niche microbiotas could be transmitted to the honey bee gut microbiota. Indeed, Anderson et al.29 found gut-specific bacteria in hive samples, which suggested bacterial transmission between the honey bee gastrointestinal tract and hive internal microenvironments. We propose the concurrent study of multiple niche-specific microbiotas, to better understand their role in this beehive-bee interconnectedness. We nominate the term "apibiome" to refer to the microbial community formed by all beehive niches, including the bees.
Herein, we characterised the in-field bacterial consortia and the predictive functional profile of the bacterial fraction of the apibiome, particularly of the honey bee gut, bee bread, hive entrance and internal hive air, of beehives in contrasting environments, through V4 16S rRNA Illumina amplicon sequencing. Functional characterization was performed using predictive PICRUSt2. Based on the results obtained by Muñoz-Colmenero et al.30, wherein contrasting anthropic habitats were compared, we expected higher anthropization levels to increase opportunistic/pathogenic bacterial loads and deplete beneficial bacteria, whilst lower anthropization would result in more balanced gut microbiotas. The contrasting environments studied here consisted of an agricultural, a semi-natural and a natural area. Agricultural and semi-natural areas contained newly formed colonies of the same origin, while colonies located in the natural area had survived over 10 years without human interference and have remained tolerant to the presence of Varroa destructor infections. To the best of our knowledge, this is the first study testing the effect of anthropization in multiple beehive microbiomes (i.e. the apibiome).
Results
Measurement of colony strength traits and Varroa destructor load
Hives located in the natural area (henceforth natural hives or colonies) weighed the most (hive weight, Kruskal–Wallis test, p < 0.0001) (Fig. 1a) yet had the smallest bee population (ANOVA test, p < 0.0001) (Fig. 1d), excluding one colony with higher bee loads (i.e. outlier). Pollen was nearly absent in hives situated in the semi-natural area (i.e. semi-natural hives or colonies) compared with the pollen-rich agricultural (henceforth agricultural hives or colonies) and natural colonies (pairwise Dunn test, p < 0.0001) (Fig. 1b). Brood presence was highest in agriculture (ANOVA test, p < 0.0001) and very similar for natural and semi-natural ambients (pairwise Tukey test, p > 0.05) (Fig. 1e). Mean Varroa load was null and equal in all apiaries (Kruskal–Wallis test, p > 0.05) but two natural colonies had high mite loads (Fig. 1c).
Statistical comparison between environments for colony strength traits and Varroa destructor loads. The top plots (a–c) show pairwise Dunn tests with Benjamini–Hochberg correction, with significance expressed as 0 (****), 0.001 (***), 0.01 (**), 0.05 (*), no-significant (ns). The bottom plots (d,e) show pairwise post-hoc Tukey’s test, with different letters indicating statistically significant differences. (a) Hive weight measured in kilograms (Kruskal–Wallis test, χ2(2) = 26.58, p < 0.0001). (b) Estimated pollen cells per apiary (Kruskal–Wallis test, χ2(2) = 24.08, p < 0.0001). (c) Varroa destructor load measured by the Powdered Sugar method (Kruskal–Wallis test, χ2(2) = 4.72, p > 0.05). (d) Bee population measured as the number of bees per colony (ANOVA, F = 13.64, p < 0.0001; pairwise Tukey, for A vs N p < 0.001, for SN vs N p < 0.05). (e) Brood population measured by quantification of brood cells (ANOVA, F = 23.35, p < 0.0001; pairwise Tukey, for A vs SN p < 0.0001, for A vs N p < 0.0001).
Comparison of bacterial community richness and taxonomic composition between hive niches
In total, 158 samples were adequately amplified and sequenced. After pair-end sequence assembly, quality filtering and singletons removal, our final dataset consisted of 7,962,468 reads. The total frequency of reads was 4,040,931 for gut (mean = 93,975.139 ± 29,760), 1,089,501 for bee bread (mean = 26,573.195 ± 13,298), 2,471,615 for hive entrance (mean = 58,847.976 ± 19,523), and 360,421 for internal air (mean = 11,263.156 ± 9,867).
Hive niche had a significant impact on bacterial community biodiversity (Faith Phylogenetic Diversity, PD), Shannon’s diversity index (H) and Pielou’s evenness index (J′) (Kruskal–Wallis p < 0.0001). Gut samples presented the least diverse (lowest PD and H, p < 0.0001) but most evenly distributed bacterial microbiome of all sample types (highest J′ p < 0.05) (Supplementary Fig. S1).
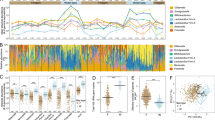
Proteobacteria were the most abundant taxa (Fig. 2a), being Alphaproteobacteria the most abundant class (in all niches except for gut), followed by Gammaproteobacteria, Bacilli and Actinobacteria (Fig. 2b). Internal hive air samples were overrun by Alphaproteobacteria [86%, represented by genera Sphingomonas (54.9%), Methylobacterium (13.2%), Bradyrhizobium (2.66%), and Phyllobacterium (1.82%)] (Fig. 2c, Supplementary Fig. S2), and only Gammaproteobacteria (5.66%), Bacilli (2.2%) and Actinobacteria (1.4%) classes had relative abundances over 1% (Fig. 2b). Bee bread microbiotas were rich in Acidobacteria (0.83%), Verrucomicrobiae (0.80%), Thermoleophilia (0.69%) and Oxynphotobacteria (0.27%) families (Fig. 2b), and Methylobacterium (15.06%), Acinetobacter (6.79%) and Pseudomonas (1.06%), and shared similar abundances of Sphingomonas, Bradyrhizobium and Phyllobacterium with air samples (Fig. 2c, Supplementary Fig. S2). Hive entrance presented the highest relative abundances of the Actinobacteria (6.81%), Bacteroidia (3.69%) and Blastocatellia (0.21%) families among all hive niches, as well as unknown bacteria (2.92%) (Fig. 2b) and the Arsenophonus (15.64%), Curtobacterium (1.81%) and Hymenobacter (1.21%) genera (Fig. 2c, Supplementary Fig. S2). The bacterial microbiome composition of gut samples was skewed towards Gammaproteobacteria (55.7%) class and the Firmicutes phylum (31.47% Bacilli) presence, and was rich in classes with < 0.1% relative abundance (Fig. 2b). At genus level, over 70% of the gut microbiome was formed by the genera Gilliamella, Lactobacillus and Snodgrasella (Fig. 2c, Supplementary Fig. S2).
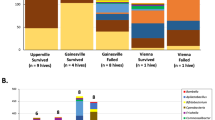
Bacterial communities of the studied hive niches at class and phylum levels. (a) Bar plots of the relative frequencies of the bacterial phyla present in each hive niche. Proteobacteria were the most abundant, excluding some natural and semi-natural gut samples rich in Firmicutes, as well as several natural hive entrances enriched for Actinobacteria and Bacteroidetes (A = agricultural apiary, SN = semi-natural apiary, N = natural apiary). (b) Pie charts of the bacterial classes presenting relative abundances ≥ 0.1%, in percentages. Some ASVs were only classified up to Domain level, and grouped as “Bacteria”. The group “Others” includes all additional taxa. Gut samples displayed a distinct microbial profile. Internal air, bee bread and hive entrance had similar abundances, with enrichment of Alphaproteobacteria, Actinobacteria and Bacteroidia; and numerous bacteria less than 0.1% abundant. (c) Most abundant bacterial genera (≥ 1.0%) per sample type, in percentages. Internal air was not sampled in the semi-natural apiary due to methodological constraints.
Environmental and anthropic effects on diversity in different hive niches
Microbial communities were in general more evenly composed in the natural environment (Pielou’s index, Kruskal–Wallis test, p < 0.05 for gut, p < 0.01 for hive entrance), with semi-natural samples showing intermediate values (Supplementary Fig. S3a). Phylogenetic diversity changed for hive entrance samples, with natural hives having the highest values and anthropic the lowest (Kruskal–Wallis p < 0.05) (Supplementary Fig. S3b). There were also significant differences in Shannon’s diversity when comparing hive entrances located in agricultural versus natural landscapes (Kruskal–Wallis p < 0.01) (Supplementary Fig. S3c).
Bacterial community composition significantly differed by environment for bee bread, hive entrance and gut samples (PERMANOVA, p ≤ 0.01 for all, pseudo-F > 4, 11, 14 respectively) but not for internal air (Supplementary Table S1). Bee bread community differences between environments were significant but weak, with a slight clustering in the PCoA (Supplementary Fig. S4a) and a lack of clustering in the UPGMA (Supplementary Fig. S4b). On the contrary, hive entrance samples showed a differentiated cluster formed by natural samples and a significant division between anthropic and semi-natural environments (pairwise PERMANOVA, pseudo-F = 7.63, p = 0.002) (Supplementary Table S1) reflected in clustering of environments on UPGMA, but not clearly visible in the PCoA (Fig. 3a). However, heterogeneous dispersion by environment was found to be significant (betadisper, F = 9.941, p = 0.001) with the natural group showing intra compositional variance. Worker guts also clustered separately for natural samples and were the only sample type showing a clear microbial clade divergence between agricultural and semi-natural environments (PCo2 = 12.81% and PERMANOVA pseudo-F = 8.86, p = 0.001) (Fig. 3b) (Supplementary Table S1). However, the largest Bray–Curtis distance (an index that measures between samples microbial compositional dissimilarity) was found when comparing either anthropic or semi-natural against natural samples (PCo1 = 42.87% and PERMANOVA pseudo-F > 10, p = 0.001) (Supplementary Table S1).
Beta diversity analysis of bacterial microbiota for hive entrance and gut samples. (a) Bray–Curtis based Principal Coordinate Analysis (PCoA) and UPGMA tree of hive entrance samples. Natural hives form an isolated cluster in the PCo1 axis. (b) Bray–Curtis based Principal Coordinate Analysis (PCoA) and UPGMA tree of adult worker gut samples, with clustering and isolation of all environments, and special differentiation for natural samples. Plotting: PCoAs were plotted using Vega editor (v5.22.1, https://vega.github.io/editor/#/). UPGMA trees were visualized in iTOL (v6.5.8, https://itol.embl.de/) and internal colors added via INKSCAPE (v0.92.3-1, https://inkscape.org/).
Functional and bacterial community profiles across environments
Honey bee worker gut samples
All three environments had a strong representation of Firmicutes (Lactobacillus), Gammaproteobacteria (the Orbaceae Gilliamella and Frischella), Betaproteobacteria (Snodgrasella), Actinobacteria (Bifidobacterium) and Alphaproteobacteria (Bartonella, Methylobacterium, Sphingomonas and Bradyrhizobium). However, there were clear differences between natural and agricultural environments. Enrichment of the Enterobacteriaceae (Gammaproteobacteria) and Rhizobiaceae (Alphaproteobacteria) families was detected in the agricultural environment (LDA > 3.6) (Fig. 4a). The Gammaproteobacteria genera Pantoea and Arsenophonus were also enriched in the agricultural environment compared to the other environments, albeit not significantly (Supplementary Table S2). The genera Lactobacillus (Bacilli), Commensalibacter (Alphaproteobacteria) and Snodgrasella (Gammaproteobacteria) represented the natural environment (LDA > 4.6) (Fig. 4a). Interestingly, the Gilliamella genus was underrepresented in natural samples, with overall lower relative frequencies than in the other two environments (median 16.94% in natural, 34.28% in semi-natural and 37.16% in agricultural) (Supplementary Table S2). Semi-natural samples were characterised by a higher presence of Bartonella (sparse in the other environments) and Frischella (median 8.24% versus agricultural 2.91% and natural 3.54%) (Supplementary Table S2). No differently enriched taxa were found in the semi-natural habitat (Fig. 4a), but importantly most agricultural and natural representatives had intermediate abundances in this environment (Fig. 6a).
Characterization of bacterial communities of worker gut samples. (a) Cladogram of significantly enriched bacteria in each environment, from phylum to genus level, according to LEfSe. LEfSe cladogram results are plotted according to phylogeny. The outermost circle corresponds to the lowest taxonomic rank (genera, level to which ASVs were collapsed). From there, each circle equals a higher taxonomic rank, with phyla being the inner circle. Only significant taxa are plotted. (b) Spearman correlation analysis of the honey bee gut bacterial communities at p < 0.05. (c) Principal Coordinate Analysis (PCoA) of samples according to the predictive functional profile. (d) Significantly expressed MetaCyc pathways (predicted functional profile) according to LEfSe. The bigger the LDA value obtained for a feature, the more significant. Only significant features are plotted in the histogram. (d) Nonoxipent: Pentose phosphate pathway I (non-oxidative branch), UMP syn I: UMP biosynthesis I, Pyr syn III: Pyrimidine deoxyribonucleotides de novo biosynthesis III, Glucurocat: β-d-glucuronosides degradation, L-Arg syn III: l-arginine biosynthesis III (via N-acetyl-l-citrulline). Plotting: Cladograms and histograms of LEfSe results were plotted in Galaxy (web application, https://huttenhower.sph.harvard.edu/galaxy/) and taxa names cleaned with INKSCAPE (v0.92.3-1, https://inkscape.org/). PCoAs were plotted using Vega editor (v5.22.1, https://vega.github.io/editor/#/).
Concerning bacteria-bacteria interactions, the natural representatives Commensalibacter and Lactobacillus were positively correlated (R = 0.50, p < 0.01). Several genera from the Enterobacteriaceae family (Hafnia-Obesumbacterium, Pantoea and one non-assigned) possessed high negative correlations with Commensalibacter (R < − 0.62, p < 0.001) and more moderate correlations with Bartonella (R < − 0.46, p < 0.05), which was more abundant in semi-natural colonies. Other Enterobacteriaceae (Morganella, Serratia) presented moderate negative correlations with Commensalibacter alone (R < − 0.48, p < 0.05). Overall, Enterobacteriaceae bacteria promoted the presence of other Enterobacteriaceae, especially between bacteria negatively correlated with Commensalibacter. Two Rhizobiaceae were also positively correlated with several Enterobacteriaceae (R > 0.4, p < 0.05). High positive correlations were also observed for several Acetobacteraceae genera, with the highest correlations present among Asaia versus Gluconacetobacter (R = 1, p < 0.0001); and Gluconobacter versus Komagataeibacter (R = 0.70, p < 0.001) (Fig. 4b).
In the predictive functional profile, natural and agricultural environments presented either the lowest or highest recruitment of features. Semi-natural samples generally had intermediate values, indicating a possible “intermediate microbiome” found in semi-natural gut samples. Similarity analysis via PCoA demonstrated clustering of environments along the PCo1 axis (Fig. 4c). All environments had similar biodiversity values, with agricultural samples having the highest H values against both natural and semi-natural (p < 0.001) and no significant differences found between semi-natural and natural samples (Supplementary Fig. S5a). Natural samples indicated significant recruitment of several anabolic reactions for the generation of precursor metabolites, nucleosides and nucleotides, while Arginine biosynthesis and β-D-glucuronoside degradation pathways were more representative of agricultural samples (Fig. 4d). The semi-natural environment retained no significant pathways, even though it had intermediate abundances for every agricultural- and natural-significant pathway detected.
Hive entrance samples
Environmental effects were clear, with only Sphingomonas, Bradyrhizobium and Methylobacterium abundantly present in all environments. Agricultural and semi-natural apiaries were overrun by Proteobacteria and more enriched than natural samples for Firmicutes (Lactobacillus, Staphylococcus, Streptococcus, Paenibacillus). The non-natural apiaries differed in bacterial abundances, rather than presence/absence of bacteria. Gammaproteobacteria (mainly Arsenophonus, Stenotrophomonas and Pseudomonas) were representative of agricultural samples, as was Lactococcus (Firmicutes) (Fig. 5a). Natural samples had a divergent microbial profile, with abundance of both Actinobacteria and Bacteroidia classes. The Aureimonas and Deinococcus genera were significantly enriched in these natural colonies (LDA > 3.5) (Fig. 5a) while Massilia was slightly enriched (Supplementary Table S2), Arsenophonus was absent and Sphingomonas, Phyllobacterium and Bradyrhizobium had overall lower abundances (Supplementary Table S3). The Sphingomonas genus (LDA > 5.0), the Bacilli class, several Proteobacteria genera, as well as the phyla Firmicutes, Gemmatimonadetes and Actinobacteria were the most significant for semi-natural samples (Fig. 5a). Two genera significant for agriculture, Arsenophonus and Lactococcus, showed intermediate abundances in semi-natural samples (Fig. 6c). Species wise, Paenibacillus larvae and Lactobacillus kunkeei (both Bacilli) plus Corynebacterium afermentans sub. sp. afermentans (Actinobacteria) were semi-natural representatives (LDA > 3.0), with P. larvae present in natural samples and practically absent in agricultural hives, while L. kunkeei and C. afermentans were present in agricultural hives but absent in natural hives (Supplementary Table S4).
Characterization of the bacterial communities in hive entrance samples. (a) Significantly enriched bacteria in each environment, according to LEfSe. Agricultural hives were rich in Gammaproteobacteria and Lactococcus. The classes Actinobacteria and Bacteroidia were prevalent in natural samples. Semi-natural samples were enriched in the Sphingomonas genus (LDA > 5.0), the Bacilli class, genera from the Alphaproteobacteria (Bradyrhizobium, Phyllobacterium) and Gammaproteobacteria (Enhydrobacter) classes, as well as genera from the Firmicutes, Gemmatimonadetes (Gemmatimonas and an uncultured genus), and Actinobacteria phyla. (b) Spearman correlation analysis at p < 0.05. Positive values were particularly high for Curtobacterium/Hymenobacter, Phyllobacterium/Sphingomonas and Phyllobacterium/Bradyrhizobium interactions (R > 0.80, p < 0.0001). The most negative interactions were found between Arsenophonus/Spirosoma and Arsenophonus/Nocardioides (R ⋍ − 0.6, p < 0.001). (c) Principal Coordinate Analysis (PCoA) of samples according to the predictive functional profile (MetaCyc pathways). (d) Significantly recruited functions according to LEfSe. (e) Significantly enriched enzymes according to LEfSe. The enzymes Endo X3 (EC 3.1.22.4) and coenzyme Q reductase (EC 7.1.1.2, formerly EC 1.6.5.3) were agricultural representatives, while tryptophan 7-halogenase (EC 1.14.19.9) was enriched in semi-natural samples. Bifido shunt: Bifidobacterium shunt, L-Met transS: l-methionine biosynthesis (transsulfuration), TCA VII: TCA cycle VII (acetate-producers), L-Met syn I: l-methionine biosynthesis I, S-Adenosyl-l-Met: S-adenosyl-l-methionine biosynthesis, Gondoate syn: gondoate biosynthesis (anaerobic), Denovopurine II: purine nucleotides de novo biosynthesis II, Pyrimidine syn II: pyrimidine deoxyribonucleotides de novo biosynthesis II, Pyridoxal syn I: pyridoxal 5′-phosphate biosynthesis I, 8-amino-7-oxo: 8-amino-7-oxononanoate biosynthesis I, tRNA processing: tRNA processing, Biotin syn: biotin biosynthesis I, PRPP: histidine, purine, and pyrimidine biosynthesis, KDO Lipid A syn: (Kdo)2-lipid A biosynthesis, Pyridoxal sal: pyridoxal 5′-phosphate biosynthesis and salvage, NAD sal III: NAD salvage pathway III (to nicotinamide riboside), ppGpp: ppGpp metabolism, LPS syn: lipopolysaccharide biosynthesis, Mycolate syn: mycolate biosynthesis, Oleate syn IV: oleate biosynthesis IV (anaerobic), (5Z)-Dode syn I: (5Z)-dodecenoate biosynthesis I. Plotting: Cladograms and histograms of LEfSe results were plotted in Galaxy (web application, https://huttenhower.sph.harvard.edu/galaxy/) and taxa names cleaned with INKSCAPE (v0.92.3-1, https://inkscape.org/). PCoAs were plotted using Vega editor (v5.22.1, https://vega.github.io/editor/#/).
Median relative abundances of bacterial and functional biomarkers showing intermediate values in the semi-natural location for gut, hive entrance and bee bread samples. The scales of the axes vary according to the relative abundance of the plotted feature. Biomarkers were defined as features with relative abundances significantly different across environments, according to LEfSe analysis (Kruskal–Wallis test p < 0.05, LDA > 3.0). (a) Bacterial biomarkers of guts, plus schematic of Spearman correlations (p < 0.05). (b) Predicted functional biomarkers in the gut. (c) Bacterial biomarkers of hive entrances, plus schematic of Spearman correlations (p < 0.05) for all environments. (d) Predicted functional biomarkers in the hive entrance. (e) Bacterial biomarkers in bee bread. Nonoxipent: Pentose phosphate pathway I (non-oxidative branch), L-Arg syn III: l-arginine biosynthesis III (via N-acetyl-l-citrulline), UMP syn I: UMP biosynthesis I, Pyr syn III: Pyrimidine deoxyribonucleotides de novo biosynthesis III, Glucurocat: β-d-glucuronosides degradation. (d) Bifido shunt: Bifidobacterium shunt (hexose catabolism), TCA VII: TCA cycle VII (acetate-producers), L-Met syn I: l-methionine biosynthesis I, L-Met transS: l-methionine biosynthesis (transsulfuration), S-Adenosyl-l-Met: S-adenosyl-l-methionine biosynthesis, KDO Lipid A syn: (Kdo)2-lipid A biosynthesis, (5Z)-Dode syn I: (5Z)-dodecenoate biosynthesis I, Oleate syn IV: oleate biosynthesis IV (anaerobic), Mycolate syn: mycolate biosynthesis, LPS syn: lipopolysaccharide biosynthesis, Denovopurine II: superpathway of purine nucleotides de novo biosynthesis II, Pyrimidine syn II: pyrimidine deoxyribonucleotides de novo biosynthesis II, NAD sal III: NAD salvage pathway III (to nicotinamide riboside), Biotin syn: biotin biosynthesis I, Pyridoxal sal: pyridoxal 5′-phosphate biosynthesis and salvage, Pyridoxal syn I: pyridoxal 5'-phosphate biosynthesis I, tRNA processing: tRNA processing, 8-amino-7-oxo: 8-amino-7-oxononanoate biosynthesis I, PRPP: histidine, purine, and pyrimidine biosynthesis, ppGpp: ppGpp metabolism, gondoate syn: gondoate biosynthesis (anaerobic). Plotting: Schematics were done in INKSCAPE (v0.92.3-1, https://inkscape.org/) considering the results of Hmisc and corrplot packages.
Concerning interactions among bacteria, correlations were mostly grouped by taxa. Actinobacteria and Bacteroidia such as Jatrophihabitans, Curtobacterium or Hymenobacter promoted mutual presence and vice versa, while the presence of Alphaproteobacteria was promoted by different Firmicutes (Spearman correlation, R > 0, p < 0.05). Exclusion, marked by negative correlation, was detected between several Actinobacteria or Bacteroidia versus genera such as Arsenophonus, Corynebacterium 1, Micrococcus and Gaiella (Fig. 5b).
In the predictive functional profile, natural hives clustered together (explaining 19.53% of dissimilarities) while agricultural samples scattered across the PCo1 axis (Fig. 5c). Diversity of functional profiles was highest in natural hives and lowest in semi-natural (Shannon’s index, p < 0.05) (Supplementary Fig. S5b). None of the significantly recruited functions were exclusive to one environment. The only non-ubiquitous function was lipopolysaccharide (LPS) biosynthesis (absent in natural colonies) (Fig. 6d), although certain pathways were also scarce in the other environments (pyrimidine biosynthesis III in agricultural gut and (Kdo)2-lipid A biosynthesis in natural hive entrance) (Fig. 6b,d). Natural colonies exhibited increased frequency of the Bifidobacterium shunt, l-methionine biosynthesis (mostly mediated by transsulfuration occurring from oxaloacetate), and a tricarboxylic acid cycle specific to acetate-producers (TCA cycle VII) (Fig. 5d). Agricultural hives possessed enriched synthesis of nucleotides, cofactors, nicotinamide adenine dinucleotide (NAD), membrane components (Kdo2 lipid A, LPS, mycolate) and fatty acids. The tRNA processing pathway resulting in tRNA activation, and the stringent specific ppGpp metabolism were also significantly expressed (Fig. 5d). Semi-natural samples had significant recruitment of the tryptophan 7-halogenase enzyme (Fig. 5e) and of anaerobic gondoate biosynthesis (Fig. 5d) and intermediate values for other pathways enriched in the more extreme environments (Fig. 6b,d).
Bee bread samples
Contrasting environments shared similar microbiomes, with differences primarily found in abundances of scarce taxa. Sphingomonas and Methylobacterium (Alphaproteobacteria) were overall the most abundant genera, followed by Acinetobacter in the natural environment and Bradyrhizobium in the other two (Supplementary Table S2). The gut core genus Bombella was enriched in natural hives, Arsenophonus represented the agricultural environment, and no taxa was augmented in semi-natural samples (LDA > 3.0) (Supplementary Fig. S4c).
Internal hive air
Environmental effects were scarce for internal air, and only seen in the enrichment of Enterobacteriaceae (mostly Arsenophonus), Curtobacterium and Massilia in agricultural samples (Supplementary Fig. S6).
Intermediate functional and bacterial community profiles in semi-natural hives
Beta-Diversity results of gut bacterial communities and predicted functions showed that semi-natural samples clustered between natural and agricultural hives (Figs. 3b, 4c). This effect was also apparent for predicted functions of hive entrance (Figs. 3a, 5c) samples. In concordance, semi-natural hives showed intermediate relative abundances for several taxa and functional pathways, while natural and agricultural environments exhibited either lowest or highest relative abundances (Fig. 6). This trend was more evident at a functional (Fig. 6b,d) than at the taxonomic level (Fig. 6a,c). Bacterial representatives showing intermediate abundances for gut samples included Comensalibacter, which was enriched in natural, scarce in semi-natural and mostly absent in agricultural samples. An unknown Rhizobiaceae genus exhibited the opposite tendency (enrichment in agriculture and absence or nearly absence in natural samples) (Fig. 6a). Same pattern was observed for Lactococcus in hive entrance (Fig. 6b) and Arsenophonus in hive entrance (Fig. 6b), bee bread (Fig. 6e) and gut samples (Supplementary Table S2). A less pronounced transition was detected for Lactobacillus in gut samples and Pseudomonas in the hive entrance. Both were present in all environments but augmented in natural and agricultural environments, respectively, while the semi-natural environment had intermediate abundances. Bee bread and gut samples also showed overall intermediate abundances for some non-environmentally relevant bacteria, such as Bradyrhizobium in bee bread and Gilliamella in gut (Supplementary Table S2). Besides the aforementioned bacteria showing intermediate abundances, all significantly recruited functions in gut and hive entrance samples had highest or lowest recruitment in natural and agricultural colonies, and intermediate values in the semi-natural environment (Fig. 6b,d). This behaviour was clear in the agricultural recruitment of NAD and Kdo2-lipid A synthesis, both displaying relative mean abundances under 0.001% in natural colonies (Fig. 6d). The exceptions to the rule were the gondoate anaerobic synthesis enriched in semi-natural hive entrance and the Bifidobacterium shunt with equally low relative abundances in both anthropized locations (Fig. 6d).
Discussion
Several studies have revealed anthropization-induced bacterial shifts in the honey bee gut microbiota14,16,17,19,30, disturbing gut microbial abundances, composition and functions. In this study, environmental anthropization resulted in the enrichment of potential pathogens and bacteria capable of surviving in contaminated landscapes, as well as recruitment of stress response-related functional pathways in bee gut and hive entrance samples. Importantly, semi-natural hives, despite being genetically identical to the agricultural hives (formed from same origin bees, queens and food reserves) but unrelated to the natural ones, showed intermediate values at taxonomic and functional levels. This mixing of agricultural and natural traits likely stemmed from environmental factors (e.g. pollen diversity or pollution).
Overall, bacterial communities associated with natural hives did not differ from the expected core profile for gut samples13, and were devoted to performing essential functions. Bacteria adapted to the natural environment were mainly found in the hive entrance. These natural hives were heavier but less populated than the non-natural beehives. Reduced colony sizes have been associated with Varroa-surviving colonies31.
Worker guts from the natural environment displayed bacterial profiles associated with good health, and were enriched in Acetobacteraceae and the gut core members Snodgrasella, Lactobacillus and Commensalibacter (involved in nutrient acquisition and immune responses). Enrichment of Acetobacteraceae and Lactobacillaceae has been associated with apiaries found > 1 km away from crops14, while Commensalibacter enrichment has been related to natural environmental conditions30, winter honey bees32, and resilience against European foulbrood28. Some Lactobacilli are core members of the honey bee gut microbiome, ferment bee-diet byproducts9,10,11 and can inhibit the pathogen Paenibacillus larvae in the gut of larvae7. Interestingly, Paenibacillus appeared in the natural apiary at low abundances across all the apibiome, except for beebread where its abundances equalled the ones found in agricultural samples. P. larvae was also present in natural hive entrance samples, yet no clinical symptoms of American Foulbrood were seen, suggesting spore inactivation or suppression of the pathogen by the bee gut bacterial community. Lactobacillus impoverishment (as detected in our agricultural hives) has been detected following antibiotic application33. Snodgrasella further contributes to improved honey bee gut equilibrium by facilitating biofilm formation in the gut34, maintaining anaerobiosis for gut symbionts9, and expressing immune genes after Escherichia coli infection35. Protozoan inhibition in natural honey bee gut through a combined enrichment of Lactobacillus, Commensalibacter and Snodgrasella was previously hypothesised as likely30, and known to occur in bumblebees36. Consistently, Commensalibater and Lactobacillus were positively correlated in our study and were dominant in the natural environment. All in all, the consortium of genera found in guts of natural colonies maintained gut environment homeostasis (anaerobiosis and biofilms) and possibly hindered infectious or opportunistic colonizations, supporting honey bee welfare by both suppressing pathogen infections or allowing its tolerance (e.g. P. larvae), and by favouring the intake of essential molecules and nutrients. Taken together, this natural taxonomic profile represented a balanced honey bee microbiome. In contrast, agricultural gut bacterial communities were enriched in non-core bacteria, similar to the findings of Muñoz-Colmenero et al.30, who also stipulated that anthropic factors could lead to microbial shifts and benefit the proliferation of opportunistic bacteria. The agricultural guts were rich in Enterobacteriaceae and Rhizobiaceae, both anthropization-related, with Rhizobiaceae having been linked to sugar syrup feeding15 and Enterobacteriaceae to Coumaphos and Chlorothalonil pesticide usage18 as well as to anthropization30. Several Enterobacteriaceae species are also opportunistic environmental bacteria and their presence in the honey bee gut has been associated with colony health weakening37 and illness38. Various Enterobacteriaceae within our samples were negatively correlated with Commensalibacter and Bartonella. Augmentation of Bartonella was detected in winter bees32 and in newly emerged bees after nutritional stress and Nosema ceranae infection16. Thus, whilst Commensalibacter maintained homeostasis of natural bacterial communities, Bartonella might have performed a similar function in semi-natural colonies.
In summary, enrichment of opportunistic bacteria and reduction of beneficial taxa was evident in agricultural colonies, confirming the detrimental impact of environmental anthropization. In between both extremes, semi-natural profiles were shifting towards natural communities despite genetic similarities and geographic proximity to agricultural colonies. The combined reduction of Enterobacteraceae and enrichment of beneficial Commensalibacter and Lactobacillus (natural > semi-natural > agriculture), supported the reduced anthropization and more balanced state of semi-natural colonies.
Honey bee gut bacteria show a diminished metabolism39 specialised in the usage of recalcitrant compounds (sugars, flavonoid glycosides, etc.) derived from the bee diet11. In this sense, most gut organisms conduct anaerobic carbohydrate fermentation11,39 while a few take part in biofilm formation or cell adhesion, such as Gilliamella apicola, Snodgrasella alvi, Lactobacillus and Bifidobacterium12,39. In concordance, the natural environment displayed an inherent metabolism supporting the synthesis of indispensable metabolites (UMP and pyrimidine deoxyribonucleotides) and the recruitment of ubiquitous anabolic reactions (non-oxidative branch of the pentose phosphate pathway, PPP). These inherent pathways, depleted in agricultural guts, showed intermediated recruitment in semi-natural samples, suggesting a weakened metabolism in both anthropized locations. Pyrimidine synthesis was especially low in agriculture, which could reflect the diminished Lactobacillus abundances in that location as Lactobacillus synthesise pyrimidine exclusively11. In combination with a less active anabolism, agricultural samples exhibited stress-related pathways. They had increased arginine (Arg) biosynthesis, linked to the aftermath of cold stress in the diptera Bactrocera dorsalis40, and increased β-d-glucuronoside degradation. Glucuronosides are formed in mammals as byproducts of hepatic glucuronidation, enabling detoxification of unwanted and toxic compounds41, and are later excreted into the gut. Recruitment of β-d-glucuronoside degradation within bee guts suggests glucuronoside presence following glucuronidation of toxic molecules by honey bees. The presence of xenobiotics might force the gut bacterial microbiome to invest most of its energy into defence mechanisms, thus neglecting pathways needed for the community to thrive (i.e. functions seen in natural guts). Agricultural honey bees endure under these conditions, but are unable to achieve a sound health state.
Aside from the honey bee gut, other niches within the apibiome also showed a strong environmental impact. The hive entrance, being directly in contact with the hive external area, appeared as a valuable indicator of the landscape. The coastal location of natural colonies (in Unije island) promoted enrichment of mostly aquatic and salt-tolerant bacteria. Bacteroidetes, for instance, are often enhanced in haloalkaline habitats42 such as our natural shoreline (i.e. high salinity and humidity). More relevant to our study was the decreased presence of the contamination-resistant Sphingomonas43, and of the potential pathogen Arsenophonus37,38. The bacteria enriched in agricultural and semi-natural landscapes are common in environments associated with plants (e.g. soils and roots). Resistance to environmental contamination has been reported or suspected for some of these bacteria. Gammaproteobacteria, enriched in agricultural colonies, are highly adaptable chemotrophs suggested to resist unfavourable conditions42, while the Sphingomonas and Gemmatimonas genera in semi-natural hives (enriched and present respectively) are key bacteria in cadmium-contaminated and saline-alkaline stressed soils42. Increased Gemmatimonas abundance has also been linked to Pyraclostrobin fungicide application44 and to long-term use of organic and inorganic fertilisers45. Sphingomonas are often associated with plant microbiomes, capable of degrading several recalcitrant substances and common helpers of fungi and plants during metal-degradation of soils43,46. Sphingomonas proliferation in leaf microbiomes has been reported after anti-pathogen treatment of plants47. A similar Sphingomonas enrichment to the one observed here in semi-natural hives was reported for petroleum-contaminated soils48. Indeed, semi-natural and agricultural apiaries were situated near traditional oil and natural gas exploitations49.
Enrichment of potential pathogens was another common trait of semi-natural and agricultural hives. Agricultural samples were particularly enriched in Pseudomonas50 and Arsenophonus. In bees, Arsenophonus has been linked to increased death rates26 and weakened colony health37, whilst enrichment of one Arsenophonus candidate was associated with increased incidence of Colony Collapse Disorder38. Interestingly, one study showed that both environment and social interactions play an important role in honey bee Arsenophonus acquisition26, and Arsenophonus was enriched in all agricultural hive niches. Arsenophonus could be a biomarker of anthropization, transmitted through social activities. However, the same study26 demonstrated that Arsenophonus abundances within honey bee guts were very location-dependent, with nearby hives sharing similar abundances. Thus, we cannot discard the influence of apiary in the differences detected for Arsenophonus abundances. Semi-natural hives also revealed several potential honey bee pathogens, including the human-affecting Streptococcus51, Anaerococcus52 and Paenibacillus. The latter genus posed a great risk to honey bees, since Paenibacillus larvae is the causative agent of American Foulbrood (AFB)53. If transmitted from entrance to bees, Paenibacillus could infect the brood and threaten semi-natural colonies. Likewise, the Lactobacillus kunkeei present in these semi-natural beehives, if transmitted to the brood, could protect semi-natural colonies by increasing brood resistance against both P. larvae and Nosema ceranae7,54. Further studies would be needed to determine P. larvae transmission within beehives. On the contrary, the hive entrances of natural hives exhibited an adapted and overall more pathogen-poor bacterial community, with the exception of P. larvae (more abundant than in agriculture). Agricultural hive entrances, compared to semi-natural, had less positive reinforcements against pathogens and contamination (i.e. less abundance of Lactobacillus kunkeei and bioremediators). Whilst natural hive entrances recruited functions prevalent among balanced metabolisms (e.g. methionine synthesis), agricultural hive entrances had more active stress-related pathways. They recruited Gram negative bacterial pathways for synthesis of outer membrane components (e.g. Kdo2-lipid A, LPS) as well as the stringent response-inducing ppGpp metabolism, which enables bacterial persistence and pathogenicity55,56. The semi-natural profiles shifted towards natural abundances.
Hive internal air and bee bread were the least influenced niches in the apibiome. Internal air, formed by floating abiotic and biotic particles found within hives, was expected to act as an indicator of beehive fitness. Indeed, shifts in airborne microbiome composition have been linked to soil, flora and possibly pollution57, as well as urbanisation58. Consequently, we expected differences between anthropic and natural landscapes, but the internal hive air microbiota turned out to be mostly stable and largely unaffected by environmental factors. This could be because free floating particles within hives, as happens with pollen granules, might stick to bee bodies and reduce the pool of available bacteria that can be sampled from the in-hive air. Anthropization had a meagre effect on bee bread, but differences among environments were consistent with the changes detected in gut and hive entrance. Bee bread sample composition resembled previous studies8,23,30, and overall natural hives had slight enrichment of acidic and sugar-tolerant bacteria while agricultural hives were enriched for Arsenophonus, previously described in bee bread obtained from multiple habitats23. Herein, Arsenophonus was most likely transmitted from agricultural bees to their food stores or vice versa, as agricultural bee guts possessed slight Arsenophonus enrichment. Contamination of the food reserves by potential pathogens such as Arsenophonus might negatively impact honey bee health at the agricultural environment, as consumption of said food could result in gut microbiome dysbiosis or affect the whole apibiome.
One of the main findings of this study was the intermediated relative abundances observed in semi-natural hives for some bacteria (e.g. Comensalibacter, Lactobacillus, Arsenophonus) and several predicted pathways (e.g. UMP and pyrimidine synthesis in guts, mycolate and LPS synthesis in hive entrance), which signalled the existence of an “intermediate state” in the semi-natural area. The intermediate state was more widespread for pathway recruitment than for community composition, indicating an early functional response of the beehive bacterial community. Importantly, 16 days of exposure to an anthropization gradient were sufficient to shift the bacterial fraction of the apibiome of hives. Agricultural hives stayed anthropized while semi-natural bacterial apibiomes became more balanced, resembling the profiles found in a non-anthropized apiary. As a parallel, quick adaptability of the gut microbiome when under pressure has been reported in humans59,60 and other mammals61. Similarly, honey bee colony-wide analysis of molecular biomarkers also demonstrated an overall increase in the expression of vitellogenin (regulatory protein within bees), antioxidant enzymes and immune proteins in hives situated near areas under wildlife recuperation for the US Conservation Reserve Program62. Our results show that placing beehives in less anthropized areas (more natural with less agricultural pressure) would lead to recruitment of beneficial bacteria (e.g. Lactobacillus and Commensalibacter in honeybee guts) and induce functional reorganisation. Indeed, habitat restoration in agricultural areas by planting native flora can favour recovery of pollinator populations3, including wild bees63.
Conclusion
Decreased anthropization of hives increased the relative abundances of beneficial bacteria in all of the sampled hive niches of the apibiome, albeit at different rates, and induced shifts in predicted functional profiles of guts and hive entrances. These results highlight the quick adaptability of honey bee-associated microbiomes. They offer straightforward management strategies to strengthen bee colonies by reducing the impact of anthropization (by planting of indigenous flora around crops, or relocating hives to more natural areas) whilst maintaining current agricultural production. These results also highlight the relevance of the hive as the unit of study for microbial research, as opposed to bees, in order to understand the contribution of each niche to colony health and resilience, as well as the importance of their interactions. Larger longitudinal and long-term analyses considering seasonal changes would enable the identification of global patterns of anthropization and of core microbes within hive niches, and contribute towards the identification of: (1) beneficial profiles that could be targeted to strengthen honey bee health at any time-period, (2) bacteria that weaken colonies, and (3) biomarkers, as Arsenophonus appears in this work, indicating the risk status of hives under anthropization. Thus, a checklist of safety and hazard markers could be developed as a management tool to employ as a bioindicator of beehive health.
Methods
Hive setup and characterization of study sites
Samples were obtained from 3 apiaries within Croatia. On 20 May 2017, 33 hives were formed in the agricultural region of Marijančaci (45.618139, 18.342667) using 4 capped brood frames with attendant bees, 2 frames of pollen/honey and mated queens. Hives contained one super each. All frames were standard Langstroth, and sister queens and same-origin worker bees were utilised to avoid genetic variation. Hives were moved the next day. Twenty-two hives were relocated 24 aerial km away to the agricultural region of Kozarac (45.717775, 18.680885), henceforth considered the anthropic or agricultural apiary. The remaining 11 colonies were moved to Vardarac (45.621335, 18.775068), which is adjacent to the Kopački Rit Nature Park and 10 aerial km away from Kozarac. These 11 colonies were designated as the semi-natural apiary. Ten additional hives were located in Unije (44.637413, 14.250092), a sparsely populated island in the Adriatic Sea, hereafter referred to as the natural environment. Seven hives were already established on this natural location beforehand. All 7 had (1) lacked management since 2012 (including pesticide treatments), (2) survived Varroa destructor infestations and (3) been previously studied by Muñoz-Colmenero et al.30. Three additional hives were obtained from the 3 strongest natural hives (nnatural = 10). Two supers were added to all 10 hives located in this natural landscape. All three apiaries remained untreated during this experiment.
Apiaries were therefore subjected to an antrophization gradient (agricultural > semi-natural > natural, from most anthropized to least) and surrounded by different flora. Agricultural exploitations and commercial beekeeping practices are regular in Osječko-Baranjska (encompassing both Kozarac and Vardarac apiaries). Grasslands, fruit trees (apple and plum), and intensive commercial crops such as rapeseed, wheat, sunflower, corn, soybeans and barley surrounded the agricultural apiary30,64,65. A similar terrain enclosed the semi-natural apiary, although with greater presence of natural flora due to its proximity (< 3 aerial km) to sedges, reed beds, scrublands and wetlands belonging to the Kopački Rit Nature Park65. In contrast, the natural location of Unije had pastureland, tufted hair grass, maquis (olive groves), coniferous woodland, and mixed broad-leaved trees (holm oaks)66. Arable land was limited to small grassland and shrub plantations around the village65. The semi-natural environment was adjacent (< 1 km) to a Special Protection Area (SPA) and Special Areas of Conservation (SAC), while natural hives were situated inside a SPA.
Measurement of colony strength traits and Varroa destructor load
Colony strength parameters were assessed on the 6th of June 2017 following the Liebefeld method67,69. Portable electronic scales were used to weigh entire hives. Both sides of all frames were checked for in-field measurement of adult bees, brood and pollen areas (dm2), as well as frame walls and bottom board for measurement of adults. Total adult bee, brood and pollen loads were calculated by multiplying the area by 125 (adult bees) or 400 (brood and pollen) as required for standard LR (Langstroth) frames68. Varroa load was measured simultaneously by the Powdered Sugar method69.
Statistical analyses for colony strength differences among environments were performed in R (Rv3.6.6, 2020-02-29). Differences between environmental means were calculated with post-hoc Tukey’s test for factors meeting ANOVA assumptions, and with Kruskal–Wallis test and post hoc Dunn’s test (using Benjamini–Hochberg correction and rstatix package70) for non-normally distributed factors.
Sample collection and processing for 16S rRNA amplification
Samples were collected in June 6, 2017, 17 days after colony formation. When possible, 4 sample types were collected per hive: young worker bees collected from brood frames (most likely nurses) for gut dissection (G), 8 cm2 of bee bread comb randomly selected from a single frame (PB), microorganisms stuck to the hive entrance and collected by swab scrubbing the entire entrance (S), and filters containing vacuum filtered internal air (F) from the hive. Older workers could not be sampled due to the short period elapsed since hive formation. The entrance was swabbed by scrubbing left and right around 6 times per swab tip (3–4 swabs per hive). Sampling of air was done by placing, on top of the honey super, a plastic dome (Lekliško cupola, produced by Dubravko Leskovic) with a perforated side attached to a vacuum hoover (Hf812, J.S.Holdings) with filters. The vacuum was left running for 10 min. Air samples were not collected in the semi-natural environment due to material limitations. Sterilisation of sampling material was undertaken using ethanol 100% and ultraviolet light, and sampling procedures done as established by Muñoz-Colmenero et al.30. Samples were frozen in dry ice until their storage at – 20 °C in the Genetics Laboratory of the University of the Basque Country (UPV/EHU). In total, 158 samples were collected from the 43 hives comprising this study.
For each gut sample (N = 43), 10 bee guts were extracted by dissection, pooled in a 1.5 mL tube with 600 µL of 1xPBS, vortexed, and centrifuged at 8000 rpm for 1 min. The supernatant was collected and placed in a clean 1.5 mL tube. This process was repeated once more by adding 400 µl 1xPBS to recover the maximum liquid sample size. The supernatants were combined and stored at − 20 °C. Afterwards, 200 µL of supernatant were taken and used for DNA extraction with a QIAamp® DNA Mini Kit (Qiagen), following the manufacturer’s protocol.
Regarding bee bread (N = 41, missing 2 agricultural samples), 3–4 pollen cells were randomly collected per sample and placed in a 2 mL tube. Cell lysis and DNA extraction was performed following the protocol established by Muñoz-Colmenero et al.30.
For hive entrance (N = 42, missing 1 agricultural sample), the cotton parts of two swabs were put in a 2 mL tube, and 400 μL of 1xPBS, 20 μL of Proteinase K and 400 μL of AL Buffer were added. Then the tubes were vortexed and incubated at 56 °C and 900 rpm for 90 min. The resulting supernatant was collected and placed in a clean 2 mL tube. This step was performed twice in order to recover as many microorganisms as possible, after which both supernatants were combined and 400 μL of ethanol were added. The mixture was vortexed and 700 μL from this tube was applied to the QIAamp® DNA Mini Kit (Qiagen) columns to perform the DNA extraction, following the manufacturer’s protocol.
DNA extraction of internal air (N = 32) was performed using a PowerSoil DNA Isolation Kit (PowerSoil). A 2 mL PowerBead Tube with 0.1 mm glass beads (Qiagen) was filled with half a filter and 60 μL of pre-heated (up to 60 °C) lysis buffer (0.1 M Tris–HCl, 0.05 M EDTA, 1.25% SDS, 0.002 mg/ml RNase) and pre-heated solution C1. Samples were homogenised using a Precellys 24 tissue homogenizer (Bertin Technologies) for 4 min. Then the tubes were incubated at 65 °C for 15 min and centrifuged at 10,000×g for 30 s. The supernatant was collected and combined with 200 μL of solution C2, after which tubes were again incubated at 4 °C for 5 min and centrifuged at 10,000×g for 1 min. Approximately 700 μL of supernatant were then transferred to a 2 ml collection tube, where 1200 μL of pre-shaken solution C4 were added. Extraction was completed according to the MO BIO Laboratories’ protocol (MO BIO Laboratories).
16S rRNA gene amplification
Characterization of the bacterial community was performed via amplification of the V4 region of the 16S rRNA gene, using the 515F/806R primer set and protocols described in the “Earth Microbiome Project'' (http://www.earthmicrobiome.org/). These primers contained Illumina sequencing adaptors and a 12 bp barcode sequence bound to the forward primers, allowing sample identification. Amplification was performed with the Illumina Amplicon Protocol, following previously described conditions and using the “with blocking primers'' protocol for bee bread samples30. PCR products were examined on a 1.5% agarose gel stained with ethidium bromide. The DNA purification of the PCR products, the preparation of the libraries and the paired-end sequencing were performed at the Sequencing and Genotyping Unit of the University of the Basque Country (SGIKER). Sequencing was performed using an Illumina MiSeq sequencer with a v2 PE 2 × 150 bp kit (300 cycles), and 10% of PhiX was added as the control for the sequencing process.
Quality checking and processing of 16S sequences
Quality of raw sequences was checked with FastQC High Throughput Sequence QC Report v0.11.571. Demultiplexing of the sequences (without Golay error correction), in-depth sequence quality control by the denoise-paired DADA2 method72, and taxonomic assignment were performed in Qiime2 v2.2 (Qiime2-2020.2)73 following the recommended thresholds. Amplicon sequence variants (ASVs) present in a single sample were removed. The original feature table was split by sample type to create sample type specific datasets (gut, bee bread, hive entrance and internal air datasets). Phylogenetic trees were generated from these datasets using mafft and fasttree alignment in Qiime2. A Naive Bayes taxonomic classifier was trained with q2-feature-classifier using our sequence data specifications and the 16S rRNA gene reference sequences from the SILVA 132 database clustered at 99% sequence similarity74. Taxonomic analysis was preceded by mitochondrial and chloroplast sequence removal. Relative abundances of bacteria were represented for phyla and genera via qiime taxa barplot. The bacterial classes presenting relative abundances of ≥ 0.1% in all sample types were identified and their relative frequencies were visualised in percentages via donut charts using the R packages ggplot2 and dplyr. Some ASVs were only classified up to domain level and grouped as “Bacteria” while the additional group “others” was used to group the remaining ASVs. Genera with mean relative frequencies of ≥ 1.0% in each hive niche were represented via bar plots using ggplot2, while genera with mean relative frequencies of ≥ 0.1% in any of the environments for any of the hive niches were represented via tables.
Bacterial community diversity and structure
A common sequencing depth for all sample types was determined through alpha rarefaction curves, and utilised to calculate and compare Alpha diversity values among sample types. Samples presenting lower sequence depths were thus filtered out. For each hive niche, bacterial community phylogenetic diversity was determined in Qiime2 via Faith Phylogenetic Diversity (PD)75, while Pielou’s evenness index76 was used to calculate community evenness. Shannon’s diversity was utilised to account simultaneously for both diversity and evenness. Significant differences were tested based on pairwise Kruskal–Wallis tests and Benjamini & Hochberg False Discovery Rate (BH-FDR) corrected p-values. Visualisation was conducted in R with ggplot2 and dplyr. Sequencing depth for each sample type was then determined through alpha rarefaction curves, and rarefied data sets were obtained per sample type for comparison of environments through Alpha and beta Diversity analyses. Samples presenting lower sequence depths were thus filtered out. For each hive niche, Alpha diversity analyses were performed again and for Beta diversity analyses Bray–Curtis distance (community composition dissimilarity) was computed using Qiime2 and visualised as Principal Coordinate Analysis (PCoA) via Vega editor (v5.22.1). Permutational multivariate analysis of variance (PERMANOVA) was calculated in Qiime2 based on rarefied Bray–Curtis matrices and with pairwise BH-FDR correction, to determine whether the bacterial communities between environments differed significantly. Considering that data dispersion can confound PERMANOVA results, homogeneity of group dispersion (PERMDISP)77 for environments was calculated with betadisper() on the same matrices. Spearman correlation-based circular UPGMA trees (unweighted pair group method with arithmetic mean) were obtained in Qiime2 and displayed via iTOL (Interactive Tree of Life tool, v6.5.8)78. Colors indicating anthropization level within UPGMA trees were added via INKSCAPE (v0.92.3-1).
Identification of taxa driving environmental differences
The feature frequency tables of each sample type were collapsed at genus level and transformed to relative abundances. Tables were uploaded to the Galaxy web application79 where Linear Discrimination Analysis (LDA) size Effect (LEfSe)80 was used to identify the bacteria driving the differences among environments. LEfSe uses a non-parametric factorial Kruskal–Wallis sum rank test81 to identify differentially abundant taxa, followed by a canonical method to calculate which taxa combinations contribute more to environmental differences. Histograms and cladograms of results were plotted within the Galaxy web application, and taxa names within graphs cleaned using INKSCAPE v0.92.3-1. Taxa were considered significant when nonparametric factorial Kruskal–Wallis test p-values ≤ 0.05 and logarithmic LDA scores > 3.0. Possible correlations among bacterial biomarkers (taxa presenting LDA > 3.0 in LEfSe) in gut and hive entrance samples were studied at genus level by performing the Spearman correlation matrix in R via corr.test() from the Hmisc package, applying BH-FDR correction, and visualized using corrplot package . Non-linear sample distribution was checked before Spearman correlation analysis, using shapiro.test normality test in R82 and BH-FDR correction. Mean relative abundances of all significant genera were represented in tables using percentages.
To determine if environmental changes could impact honey bee apibiome functionality, functional prediction of E.C. enzymes83 and MetaCyc pathways84 were performed for gut and hive entrance samples using the PICRUSt2 v2.3.0-b85 qiime2 q2-PICRUSt2 (v2019.10_0) plugin. The resulting E.C. and pathway tables were rarefied for diversity analyses. Functional diversity of environments was determined by Shannon’s diversity index and significance calculated using pairwise Kruskal–Wallis tests. BH-FDR correction was applied to p-values of pairwise analysis. Bray–Curtis distances were visualised via PCoA to determine environmental dissimilarities. Significant predicted function differences among environments were determined by LEfSe, and considered when Kruskal–Wallis p-value ≤ 0.05 and LDA scores > 3.0. Mean relative abundances of significant features were calculated and visualised as histograms via the ggplot2 and dplyr R packages.
Data availability
The 16S rRNA sequences supporting the conclusions of this article were submitted to the Qiita database (https://qiita.ucsd.edu/) with the ID 14084 and are available at the EMBL-EBI with the accession number ERP133378 (https://www.ebi.ac.uk/ena/browser/view/PRJEB48937).
Change history
23 December 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41598-022-25981-x
References
Kulhanek, K. et al. A national survey of managed honey bee 2015–2016 annual colony losses in the USA. J. Apic. Res. 56(4), 328–340. https://doi.org/10.1080/00218839.2017.1344496 (2017).
Potts, S. G. et al. Global pollinator declines: Trends, impacts and drivers. Trends Ecol. Evol. 25(6), 345–353. https://doi.org/10.1016/j.tree.2010.01.007 (2010).
Li, G. et al. The wisdom of honeybee defenses against environmental stresses. Front. Microbiol. 9, 722. https://doi.org/10.3389/fmicb.2018.00722 (2018).
Horak, R. D., Leonard, S. P. & Moran, N. A. Symbionts shape host innate immunity in honeybees. Proc. R. Soc. B. Biol. Sci. 287(1933), 20201184. https://doi.org/10.1098/rspb.2020.1184 (2020).
Dosch, C. et al. The gut microbiota can provide viral tolerance in the honey bee. Microorganisms. 9(4), 871. https://doi.org/10.3390/microorganisms9040871 (2021).
Emery, O., Schmidt, K. & Engel, P. Immune system stimulation by the gut symbiont Frischella perrara in the honey bee (Apis mellifera). Mol. Ecol. 26(9), 2576–2590. https://doi.org/10.1111/mec.14058 (2017).
Forsgren, E., Olofsson, T. C., Vásquez, A. & Fries, I. Novel lactic acid bacteria inhibiting Paenibacillus larvae in honey bee larvae. Apidologie 41(1), 99–108. https://doi.org/10.1051/apido/2009065 (2010).
Anderson, K. E., Sheehan, T. H., Eckholm, B. J., Mott, B. M. & DeGrandi-Hoffman, G. An emerging paradigm of colony health: Microbial balance of the honey bee and hive (Apis mellifera). Insectes Soc. 58(4), 431. https://doi.org/10.1007/s00040-011-0194-6 (2011).
Zheng, H., Powell, J. E., Steele, M. I., Dietrich, C. & Moran, N. A. Honeybee gut microbiota promotes host weight gain via bacterial metabolism and hormonal signaling. Proc. Natl. Acad. Sci. 114(18), 4775–4780. https://doi.org/10.1073/pnas.1701819114 (2017).
Kešnerová, L. et al. Disentangling metabolic functions of bacteria in the honey bee gut. PLoS Biol. 15(12), e2003467. https://doi.org/10.1371/journal.pbio.2003467 (2017).
Bonilla-Rosso, G. & Engel, P. Functional roles and metabolic niches in the honey bee gut microbiota. Curr. Opin. Microbiol. 43, 69–76. https://doi.org/10.1016/j.mib.2017.12.009 (2018).
Ellegaard, K. M. et al. Extensive intra-phylotype diversity in lactobacilli and bifidobacteria from the honeybee gut. BMC Genomics 16(1), 284. https://doi.org/10.1186/s12864-015-1476-6 (2015).
Kwong, W. K. & Moran, N. A. Gut microbial communities of social bees. Nat. Rev. Microbiol. 14(6), 374–384. https://doi.org/10.1038/nrmicro.2016.43 (2016).
Jones, J. C. et al. Gut microbiota composition is associated with environmental landscape in honey bees. Ecol. Evol. 8(1), 441–451. https://doi.org/10.1002/ece3.3597 (2017).
D’Alvise, P. et al. The impact of winter feed type on intestinal microbiota and parasites in honey bees. Apidologie 49(2), 252–264. https://doi.org/10.1007/s13592-017-0551-1 (2018).
Castelli, L. et al. Impact of nutritional stress on honeybee gut microbiota, immunity, and Nosema ceranae infection. Microb. Ecol. 80(4), 908–919. https://doi.org/10.1007/s00248-020-01538-1 (2020).
Campbell, J. B. et al. The fungicide Pristine® inhibits mitochondrial function in vitro but not flight metabolic rates in honey bees. J. Insect. Physiol. 86, 11–16. https://doi.org/10.1016/j.jinsphys.2015.12.003 (2016).
Kakumanu, M. L., Reeves, A. M., Anderson, T. D., Rodrigues, R. R. & Williams, M. A. Honey bee gut microbiome is altered by in-hive pesticide exposures. Front. Microbiol. https://doi.org/10.3389/fmicb.2016.01255 (2016).
Abbo, P. M. et al. Effects of Imidacloprid and Varroa destructor on survival and health of European honey bees, Apis mellifera. Insect Sci. 24(3), 467–477. https://doi.org/10.1111/1744-7917.12335 (2017).
DeGrandi-Hoffman, G., Corby-Harris, V., DeJong, E. W., Chambers, M. & Hidalgo, G. Honey bee gut microbial communities are robust to the fungicide Pristine® consumed in pollen. Apidologie 48(3), 340–352. https://doi.org/10.1007/s13592-016-0478-y (2017).
Motta, E. V. S., Powell, J. E., Leonard, S. P. & Moran, N. A. Prospects for probiotics in social bees. Philos. Trans. R. Soc. B. 377, 20210156. https://doi.org/10.1098/rstb.2021.0156 (2022).
Corby-Harris, V., Maes, P. & Anderson, K. E. The bacterial communities associated with honey bee (Apis mellifera) foragers. PLoS ONE 9(4), e95056. https://doi.org/10.1371/journal.pone.0095056 (2014).
Donkersley, P., Rhodes, G., Pickup, R. W., Jones, K. C. & Wilson, K. Bacterial communities associated with honeybee food stores are correlated with land use. Ecol. Evol. 8(10), 4743–4756. https://doi.org/10.1002/ece3.3999 (2018).
Mullin, C. A. et al. High levels of miticides and agrochemicals in North American apiaries: Implications for honey bee health. PLoS ONE 5(3), e9754. https://doi.org/10.1371/journal.pone.0009754 (2010).
Miller, D. L., Parish, A. J. & Newton, I. L. Transitions and transmission: Behavior and physiology as drivers of honey bee-associated microbial communities. Curr. Opin. Microbiol. 50, 1–7. https://doi.org/10.1016/j.mib.2019.08.001 (2019).
Drew, G. C. et al. Transitions in symbiosis: Evidence for environmental acquisition and social transmission within a clade of heritable symbionts. ISME J. 15, 2956–2968. https://doi.org/10.1038/s41396-021-00977-z (2021).
Forfert, N. et al. Parasites and pathogens of the honeybee (Apis mellifera) and their influence on inter-colonial transmission. PLoS ONE 10(10), e0140337. https://doi.org/10.1371/journal.pone.0140337 (2015).
Erban, T. et al. Bacterial community associated with worker honeybees (Apis mellifera) affected by European foulbrood. PeerJ 5, e3816. https://doi.org/10.7717/peerj.3816 (2017).
Anderson, K. E. et al. Microbial ecology of the hive and pollination landscape: Bacterial associates from floral nectar, the alimentary tract and stored food of honey bees (Apis mellifera). PLoS ONE 8(12), e83125. https://doi.org/10.1371/journal.pone.0083125 (2013).
Muñoz-Colmenero, M. et al. Differences in honey bee bacterial diversity and composition in agricultural and pristine environments—A field study. Apidologie 51(6), 1018–1037. https://doi.org/10.1007/s13592-020-00779-w (2020).
Locke, B. & Fries, I. Characteristics of honey bee colonies (Apis mellifera) in Sweden surviving Varroa destructor infestation. Apidologie 42, 53342. https://doi.org/10.1007/s13592-011-0029-5 (2011).
Kešnerová, L. et al. Gut microbiota structure differs between honeybees in winter and summer. ISME J. 14(3), 801–814. https://doi.org/10.1038/s41396-019-0568-8 (2020).
Raymann, K., Shaffer, Z. & Moran, N. A. Antibiotic exposure perturbs the gut microbiota and elevates mortality in honeybees. PLoS Biol. 15(3), e2001861. https://doi.org/10.1371/journal.pbio.2001861 (2017).
Martinson, V. G., Moy, J. & Moran, N. A. Establishment of characteristic gut bacteria during development of the honeybee worker. Appl. Environ. Microbiol. 78(8), 2830–2840. https://doi.org/10.1128/AEM.07810-11 (2012).
Kwong, W. K., Mancenido, A. L. & Moran, N. A. Immune system stimulation by the native gut microbiota of honey bees. R. Soc. Open. Sci. 4(2), 170003. https://doi.org/10.1098/rsos.170003 (2017).
Koch, H. & Schmid-Hempel, P. Socially transmitted gut microbiota protect bumble bees against an intestinal parasite. Proc. Natl. Acad. Sci. USA. 108(48), 19288–19292. https://doi.org/10.1073/pnas.1110474108 (2011).
Budge, G. E. et al. Identifying bacterial predictors of honey bee health. J. Invertebr. Pathol. 141, 41–44. https://doi.org/10.1016/j.jip.2016.11.003 (2016).
Cornman, R. S. et al. Pathogen webs in collapsing honey bee colonies. PLoS ONE 7(8), e43562. https://doi.org/10.1371/journal.pone.0043562 (2012).
Kwong, W. K., Engel, P., Koch, H. & Moran, N. A. Genomics and host specialization of honey bee and bumble bee gut symbionts. Proc. Natl. Acad. Sci. 111(31), 11509–11514. https://doi.org/10.1073/pnas.1405838111 (2014).
Raza, M. F. et al. Gut microbiota promotes host resistance to low-temperature stress by stimulating its arginine and proline metabolism pathway in adult Bactrocera dorsalis. PLoS Pathog. https://doi.org/10.1371/journal.ppat.1008441 (2020).
Kaivosaari, S., Finel, M. & Koskinen, M. N-glucuronidation of drugs and other xenobiotics by human and animal UDP-glucuronosyltransferases. Xenobiotica 41(8), 652–669. https://doi.org/10.3109/00498254.2011.563327 (2011).
Wang, M., Chen, S., Chen, L. & Wang, D. Responses of soil microbial communities and their network interactions to saline-alkaline stress in Cd-contaminated soils. Environ. Pollut. 252, 1609–1621. https://doi.org/10.1016/j.envpol.2019.06.082 (2019).
Asaf, S., Numan, M., Khan, A. L. & Al-Harrasi, A. Sphingomonas: From diversity and genomics to functional role in environmental remediation and plant growth. Crit. Rev. Biotechnol. 40(2), 138–152. https://doi.org/10.1080/07388551.2019.1709793 (2020).
Zhang, C. et al. Response of soil microbes after direct contact with pyraclostrobin in fluvo-aquic soil. Environ. Pollut. 255, 113164. https://doi.org/10.1016/j.envpol.2019.113164 (2019).
Li, F., Chen, L., Zhang, J., Yin, J. & Huang, S. Bacterial community structure after long-term organic and inorganic fertilization reveals important associations between soil nutrients and specific taxa involved in nutrient transformations. Front. Microbiol. 8, 187. https://doi.org/10.3389/fmicb.2017.00187 (2017).
Gatheru, W. M., Sun, K. & Gao, Y. Sphingomonads in microbe-assisted phytoremediation: Tackling soil pollution. Trends Biotechnol. 35(9), 883–899. https://doi.org/10.1016/j.tibtech.2017.06.014 (2017).
Qin, C. et al. Responses of phyllosphere microbiota and plant health to application of two different biocontrol agents. AMB Express 9(1), 42. https://doi.org/10.1186/s13568-019-0765-x (2019).
Zhou, L. et al. Abundance and diversity of Sphingomonas in Shenfu petroleum-wastewater irrigation zone, China. Environ. Sci. Pollut. Res. 19(1), 282–294. https://doi.org/10.1007/s11356-011-0552-y (2012).
Velić, J., Kišić, K. & Krasić, D. The characteristics of the production and processing of oil and natural gas in Croatia from 2000 to 2014. RGN zbornik. 31(2), 69–112. https://doi.org/10.17794/rgn.2016.2.6 (2016).
De Smet, J., Hendrix, H., Blasdel, B. G., Danis-Wlodarczyk, K. & Lavigne, R. Pseudomonas predators: Understanding and exploiting phage–host interactions. Nat. Rev. Microbiol. 15(9), 517–530. https://doi.org/10.1038/nrmicro.2017.61 (2017).
Haenni, M., Lupo, A. & Madec, J.-Y. Antimicrobial resistance in Streptococcus spp. Microbiol. Spectr. https://doi.org/10.1128/microbiolspec.ARBA-0008-2017 (2018).
Murphy, E. C. & Frick, I.-M. Gram-positive anaerobic cocci–commensals and opportunistic pathogens. FEMS Microbiol. Rev. 37(4), 520–553. https://doi.org/10.1111/1574-6976.12005 (2013).
Genersch, E. American Foulbrood in honeybees and its causative agent, Paenibacillus larvae. J. Invertebr. Pathol. 103(Suppl 1), S10-19. https://doi.org/10.1016/j.jip.2009.06.015 (2010).
Arredondo, D. et al. Lactobacillus kunkeei strains decreased the infection by honey bee pathogens Paenibacillus larvae and Nosema ceranae. Benef. Microbes. 9(2), 279–290. https://doi.org/10.3920/BM2017.0075 (2018).
Dalebroux, Z. D., Svensson, S. L., Gaynor, E. C. & Swanson, M. S. ppGpp conjures bacterial virulence. Microbiol. Mol. Biol. Rev. 74(2), 171–199. https://doi.org/10.1128/MMBR.00046-09 (2010).
Pacios, O. et al. (p)ppGpp and its role in bacterial persistence: New challenges. Antimicrob. Agents. Chemother. https://doi.org/10.1128/AAC.01283-20 (2020).
Cao, C. et al. Inhalable microorganisms in Beijing’s PM2.5 and PM10 pollutants during a severe smog event. Environ. Sci. Technol. 48(3), 1499–1507. https://doi.org/10.1021/es4048472 (2014).
Barberán, A. et al. Continental-scale distributions of dust-associated bacteria and fungi. Proc. Natl. Acad. Sci. USA 112(18), 5756–5761. https://doi.org/10.1073/pnas.1420815112 (2015).
David, L. A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505(7484), 559–563. https://doi.org/10.1038/nature12820 (2014).
Uhr, G. T., Dohnalová, L. & Thaiss, C. A. The dimension of time in host-microbiome interactions. mSystems. https://doi.org/10.1128/mSystems.00216-18 (2019).
Seddik, H., Xu, L., Wang, Y. & Mao, S. Y. A rapid shift to high-grain diet results in dynamic changes in rumen epimural microbiome in sheep. Animal https://doi.org/10.1017/S1751731118003269 (2018).
Ricigliano, V. A. et al. Honey bee colony performance and health are enhanced by apiary proximity to US Conservation Reserve Program (CRP) lands. Sci Rep. 9, 4894. https://doi.org/10.1038/s41598-019-41281-3 (2019).
Tonietto, R. K. & Larkin, D. J. Habitat restoration benefits wild bees: A meta-analysis. J. Appl. Ecol. 55(2), 582–590. https://doi.org/10.1111/1365-2664.13012 (2018).
Glavaš, H., Ivanović, M. & Mandic, N. Resources and possibilities of agro biomass usage for energy purposes in Slavonia region (Croatia). In IEEE International Energy Conference (ENERGYCON), 1150–1155. https://doi.org/10.1109/ENERGYCON.2014.6850568 (2014).
European Environment Agency–Ecosystem types of Europe. Data available at: https://www.eea.europa.eu/data-and-maps/data/ecosystem-types-of-europe-1. (2019) (Accessed 8 April 2021).
Starc, N. Small islands and Large Scale Spatial Development Patterns-story of the Croatian island of Unije. European Regional Science Association, ERSA conference papers. https://www.researchgate.net/publication/23731990_Small_Islands_and_Large_Scale_Spatial_Development_Patterns_-_Story_of_the_Croatian_Island_of_Unije. (2006) (Accessed 4 April 2021).
Delaplane, K. S., van der Steen, J. & Guzman-Novoa, E. Standard methods for estimating strength parameters of Apis mellifera colonies. J. Apic. Res. 52(1), 1–12. https://doi.org/10.3896/IBRA.1.52.1.03 (2013).
Imdorf, A., Buehlmann, G., Gerig, L., Kilchenmann, V. & Wille, H. Überprüfung der Schätzmethode zur Ermittlung der Brutfläche und der Anzahl Arbeiterinnen in freifliegenden Bienenvölkern. Apidologie 18(2), 137–146. https://doi.org/10.1051/apido:19870204 (1987).
Dietemann, V. et al. Standard methods for varroa research. J. Apic. Res. 52(1), 1–54. https://doi.org/10.3896/IBRA.1.52.1.09 (2013).
Kassambara, A. rstatix: Pipe-Friendly Framework for Basic Statistical Tests. R package version 0.7.0. https://CRAN.R-project.org/package=rstatix (2021).
Andrews, S. FastQC: A Quality Control tool for high throughput sequence data. https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2010).
Callahan, B. J. et al. DADA2: High resolution sample inference from Illumina amplicon data. Nat. Methods. 13(7), 581–583. https://doi.org/10.1038/nmeth.3869 (2016).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37(8), 852–857. https://doi.org/10.1038/s41587-019-0209-9 (2019).
Quast, C. et al. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic. Acids. Res. 41(D1), D590–D596. https://doi.org/10.1093/nar/gks1219 (2013).
Faith, D. P. Conservation evaluation and phylogenetic diversity. Biol. Conserv. 61(1), 1–10. https://doi.org/10.1016/0006-3207(92)91201-3 (1992).
Pielou, E. C. The measurement of diversity in different types of biological collections. J. Theor. Biol. 13, 131–144. https://doi.org/10.1016/0022-5193(66)90013-0 (1966).
Anderson, M. J. Distance-based tests for homogeneity of multivariate dispersions. Biometrics 62, 245–253. https://doi.org/10.1111/j.1541-0420.2005.00440.x (2006).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: An online tool for phylogenetic tree display and annotation. Nucleic. Acids. Res. 49(W1), W293–W296. https://doi.org/10.1093/nar/gkab301 (2021).
Afgan, E. et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic. Acids. Res. 46(Web Server issue), W537–W544. https://doi.org/10.1093/nar/gky379 (2018).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12(6), R60. https://doi.org/10.1186/gb-2011-12-6-r60 (2011).
Kruskal, W. H. & Wallis, W. A. Use of ranks in one-criterion variance analysis. J. Am. Stat. Assoc. 47, 583–621. https://doi.org/10.1080/01621459.1952.10483441 (1952).
Royston, J. P. An extension of Shapiro and Wilk’s W test for normality to large samples. J. R. Stat. Soc. Ser. C. Appl. Stat. 31(2), 115–124. https://doi.org/10.2307/2347973 (1982).
Bairoch, A. The ENZYME database in 2000. Nucleic. Acids. Res. 28(1), 304–305. https://doi.org/10.1093/nar/28.1.304 (2000).
Caspi, R. et al. The MetaCyc database of metabolic pathways and enzymes—A 2019 update. Nucleic. Acids. Res. 48(D1), D445–D453. https://doi.org/10.1093/nar/gkz862 (2020).
Douglas, G. M. et al. PICRUSt2 for prediction of metagenome functions. Nat. Biotechnol. 38(6), 685–688. https://doi.org/10.1038/s41587-020-0548-6 (2020).
Acknowledgements
The authors are grateful to the Sequencing and Genotyping Unit—SGIker from UPV/EHU for their technical support. We would also like to thank the funding groups of this work: the Dept. of Economic Development and Competitiveness of the Basque Government (Gobierno Vasco/Eusko Jaurlaritza), R&D&I grants for the agricultural, food and fishing sectors of the Basque Autonomous Community (37-2017-00044) and the Research Group IT1233-19 of the Basque University System. JG was supported by the Department of Agriculture, Fisheries and Food of the Basque Government (Gobierno Vasco/Eusko Jaurlaritza).
Funding
This work was funded by the Dept. of Economic Development and Competitiveness of the Basque Government (Gobierno Vasco/Eusko Jaurlaritza), R&D&I grants for the agricultural, food and fishing sectors of the Basque Autonomous Community (37-2017-00044), and the Research Group IT1233-19 of the Basque University System. JG was supported by the Department of Agriculture, Fisheries and Food of the Basque Government (Gobierno Vasco/Eusko Jaurlaritza) through a subsidy programme of training aid and support. These funding bodies provided the financial support to the research, but did not participate in the design of the study, analysis and interpretation of data, and writing of the manuscript.
Author information
Authors and Affiliations
Contributions
J.G-O. analysed and interpreted the data with inputs from M.M-C., IZ and A.E. M.M-C realised the laboratory experiments. J.G-O. wrote the manuscript and prepared display elements with major contributions of M.M-C., I.Z., A.E. and M.P. M.M-C, I.Z, M.K., J.F., Z.P., N.K. A.E. and R.B. designed the experiment. A.E. and R.B. conceptualised the study. M.K., J.F., Z.P. and N.K. performed field experiments and sampling. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: The original version of this Article contained a typographical error. Figure 5 did not display correctly.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gorrochategui-Ortega, J., Muñoz-Colmenero, M., Kovačić, M. et al. A short exposure to a semi-natural habitat alleviates the honey bee hive microbial imbalance caused by agricultural stress. Sci Rep 12, 18832 (2022). https://doi.org/10.1038/s41598-022-23287-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-23287-6
This article is cited by
-
Colonies under dysbiosis benefit from oxalic acid application: the role of landscape and beekeeping practices in microbiota response to treatment
Journal of Pest Science (2024)
-
Effect of honey bee forage plants in Tunisia on diversity and antibacterial potential of lactic acid bacteria and bifidobacteria from Apis mellifera intermissa and its products
Archives of Microbiology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.