Abstract
The increased frequency of different lifestyles that disrupts circadian rhythms, together with a trend in the accretion of male idiopathic infertility, imposes the necessity to understand the contribution of circadian rhythms disruption to fertility regulation. In this study, the effects of circadian desynchrony (CD) on the steroidogenic capacity of adult Leydig cells were studied. Adult rats were housed under a disturbing light regime (2 days of constant light, 2 days of continual dark, and 3 days of 12:12 h light:dark schedule) designed to mimic shiftwork in humans. CD was characterized by changed and decreased rhythmic locomotor activity and reduced blood testosterone. In the Leydig cells changed transcription of the clock genes (Bmal1, Clock, Cry1 and Reverba/b increased while Per1/2 reversed phase) was detected. This was followed by reduced transcription of genes (Star, Cyp11a1, and Hsd3b1/2) primarily involved in mitosteroidogenesis. In parallel, mitochondrial membrane potential (Δψi) and ATP production declined losing their characteristic oscillatory pattern. Also, the main markers of mitochondrial biogenesis (Ppargc1a, Nrf1, Tfam, Cytc), fusion (Mfn2), and mitophagy (Pink1 and Tfeb) were disturbed. Collectively, CD targets mitochondria in Leydig cells by reducing mitosteroidogenesis, mitoenergetics, and disturbing mitochondrial dynamics. These changes contribute to testosterone decline compromising androgen-dependent functions, including reproduction.
Similar content being viewed by others
Introduction
The essential condition for the survival of the species is successful and efficient reproduction. This condition can be met if the physiological processes are organized to form a unified system synchronized with the external environment. One of the systems which anticipate daily environmental cycles and thereby enable adaptation and maintenance of dynamic organism's equilibrium is the circadian clock. In mammals, the circadian system is hierarchically organized and composed of the master rhythm regulator located in the suprachiasmatic nucleus of the hypothalamus and peripheral clocks present in peripheral cells1. Clock genes orchestrate circadian rhythm in almost every cell operating through transcription/translation negative feedback loop maintaining its rhythm and the rhythmic expression of the clock-dependent genes. The principal genes involved in a clock loops are positive Bmal1, Clock, Npas2, and negative genes Per1/2/3, Cry1/2, and Reverba/b2.
The circadian clock system, together with neuronal, hormonal, and metabolic signals, collectively regulates reproductive physiology on a 24-h scale and drives diurnal oscillation in gene expression1. In males, testosterone needed to complete spermatogenesis and develop and maintain male sexual characteristics is produced by Leydig cells in a daily rhythm. Testosterone is formed by the activity of a battery of steroidogenic elements/enzymes located in mitochondria and endoplasmic reticulum encoded by steroidogenic genes (Star, Cyp11a1, Hsd3b1/2, Cyp17a1, Hsd17b)3. So far, clock genes (Bmal1, Per1/2/3, Cry1/2, Dbp and Reverba/b), as well as genes involved in the process of steroidogenesis (Star, Cyp11a1, Cyp17a1), have rhythmic expressions in Leydig cells4,5,6,7,8,9. Accordingly, blood testosterone fluctuations usually show one or two peaks4,10,11,12,13. The generator of these fluctuations is located both in Leydig cells and outside, and steroidogenic function is tuned to regular episodic gonadotropin input13. Male Bmal1−/− mice are infertile and have low blood testosterone and decreased expression of testicular steroidogenic genes (Star, Hsd3b2, and Hsd17b3)14. Similar results were obtained on BMAL1 knocked out TM3 cells15. In contrast, overexpression of BMAL1 significantly increased StAR and HSD17B3 and improved testosterone production9. All these observations emphasize the importance of the clock system in maintaining rhythmic testosterone synthesis and, consequently, male fertility.
However, modern lifestyles imposes activity that is not in line with the endogenous rhythm dictated by the circadian clock, leading to so-called “circadian desynchrony” (CD)16. This condition can cause changes in hormone secretion, favoring infertility-related changes that may result from reproductive axis dysfunction. As time and synchronization of hormone secretion and tissue sensitivity play a key role in fertility, desynchrony between the master rhythm regulator and the rest of the reproductive axis, especially with peripheral clocks such are in testicular Leydig cells, may become important factors in the development of the conditions associated with reduced testosterone and fertility.
Given the increased frequency of work in shifts and travel that disrupts circadian rhythms, it is crucial to understand the contribution of circadian rhythms to fertility regulation. In particular, understanding the relationship between the circadian clock and testicular function, especially in terms of increased incidence of male infertility17 and that the cause of male infertility often remains unknown in over 70% of cases18.
Therefore, the current study aims to analyze effect of CD, established by changing of photoperiod, on rat androgen-producing Leydig cells. We hypothesized that CD reduces rhythmic function of the Leydig cells by changing activity of clock and steroidogenesis–related genes.
Results
Decreased male fertility is a growing health problem that requires a better understanding of molecular events regulating reproductive competence. Although previously published data suggest an association between circadian rhythm disruption and decreased male fertility19, the knowledge about how CD interrupts male fertility is still incomplete. In this study, the effects of CD on the androgenic capacity of adult Leydig cells were studied. Adult rats were housed under a disturbing light regime (2 days of constant light, 2 days of continual dark, and 3 days of 12:12 h light:dark schedule) designed to mimic shiftwork in humans. After 2 months of life in conditions of the altered light regime, the transcriptional pattern of genes essential for rhythmic Leydig cell's endocrine activity and blood hormones level was monitored at five different time points.
Effects on rhythmic daily activity and blood hormones
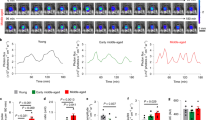
The CD was characterized by decreased and arrhythmic voluntary rat activity (Fig. 1a; Supplemental Fig. 1). It seems that the rats' activity in the experimental group decreased with time (Supplemental Fig. 1) so that in the last week of the experiment, it was about six times lower in the experimental compared to the control group (Fig. 1b). However, the body weight was not changed (Fig. 1c) during 2 months of experiment. In the blood of control rats, 24 h fluctuations of testosterone and corticosterone levels were observed with a peak around ZT11 for both (Fig. 1d,e; Suppl. Table 2). CD disturbed testosterone's diurnal rhythm in blood by decreasing mesor and amplitude (Fig. 1d; Suppl. Table 2) but peak phasing was maintained. To assess stress levels in CD rats' serum corticosterone was measured. No overall increase in diurnal corticosterone blood levels was observed, but the characteristic pattern of blood corticosterone diurnal variation was lost (Fig. 1e; Suppl. Table 2). However relative ratio between testosterone and corticosterone normalized on value ZT3 in control samples (T/C ZT3 = 1) decreased in CD condition (Fig. 1f) suggesting T/C as a possible biomarker for changes in behavior in response to circadian desynchrony.
The CD changed daily activity pattern and reduced testosterone blood level. Adult rats were housed under the controlled light regime of 12 h light–12 h dark (control) or were exposed to a disturbing light regime (2 days of continual light, 2 days of continual dark, and 3 days of 12:12 h light:dark schedule; experimental) for 2 months. The voluntary activity was monitored, and actograms were formed. The representative weekly actogram from control and experimental rats (a) as well as average weekly activity (b) and body mass (c) in the last week of the experiment are shown in bars ± SEM values (n = 25; 5 rats in 5 time points). The experimental conditions reduced diurnal fluctuations of serum testosterone (determined in ZT3, ZT11, ZT17, ZT20, ZT23; ZT0 is time when light switch on) (d), flattened the daily profile of corticosterone levels in serum (e), and reduced testosterone/corticosterone (T/C) ratio (f). For rhythm parameters, please see Suppl. Table 2. ZT is zeitgeber time, i.e., the time when the light switches on. In (d–f) panels of this figure and all others figures data points represent individual values, while the black line represents group means values (n = 5). **Statistical significance p < 0.01 determined by two-way ANOVA; *statistical significance p < 0.05 determined by Mann–Whitney test.
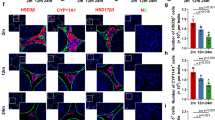
Further, since luteinizing hormone (LH) is the primary regulator of Leydig cell steroidogenesis, the transcription of pituitary genes encoding LH subunits (Lhb, Cga) and genes encoding receptors essential for gonadotroph function (Gnrhr, Nr3c1, Mntr1a which encrypt gonadotropin releasing hormone receptor, glucocorticoid receptor, and melatonin receptor, respectively) were studied. As expected, the result showed the diurnal variation of Cga (Fig. 2a; Suppl. Table 2) and Lhb (Fig. 2b; Suppl. Table 2) but also Gnrhr (Fig. 2c; Suppl. Table 2), Nr3c1 (Fig. 2d; Suppl. Table 2) and Mntr1a (Fig. 2e; Suppl. Table 2) proving circadian activity of pituitary genes involved in the regulation of reproduction. Furthermore, CD disturbed a daily expression profile of Cga and Lhb (Fig. 2a,b; Suppl. Table 2), suggesting a possible disturbance and decrease in LH synthesis that, in addition to changed transcriptional pattern of Gnrhr (Fig. 2c), Nr3c1 (Fig. 2d) and Mntr1a (Fig. 2e) propose disorder in pituitary rhythmic activity, at least the one related to the regulation of reproduction.
The CD changed daily expression patterns of the pituitary genes. Pituitary were collected at different time points during 24 h, RNA isolated for RT followed by RQ-PCR analysis of genes encoded LH subunits (a,b) and genes encoding receptors essential for gonadotroph function (c–e). CD changed daily transcriptional pattern of Cga (a), Lhb (b), Gnrhr (c), Nr3c1 (d) and Mntr1a (e). Statistical significance: *p < 0.05; **p < 0.01; ****p < 0.001 determined by two-way ANOVA.
Expression of the genes related to rhythmic endocrine Leydig cell function
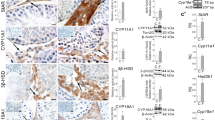
Since CD effectively decreased blood testosterone levels, the transcription of genes encoding the main elements essential for steroidogenesis was analyzed in the Leydig cells. The analysis was performed on the second day of the continual light phase in experimental and at the same time in the control groups. Obtained data in controls confirmed daily fluctuation of the transcription of key steroidogenic genes, Star, Cyp11a1, and Cyp17a1 (Fig. 3b,c,e; Suppl. Table 2) while Lhcgr (Fig. 3a) which determines Leydig cells' sensitivity on LH, and Hsd3b1/2 (Fig. 3d) did not show 24 h variation4,13. CD reduced mesor and amplitude of the Star and Cyp11a1 (Fig. 3b,c; Suppl. Table 2). These genes encode enzymes that are functionally linked to mitochondria, proposing that CD target mitosteroidogenesis. However, overall diurnal rhythmicity and peak phasing of these gene transcripts in the Leydig cells was maintained (Suppl. Table 2). CD also decreased the non-cyclical expression of Hsd3b1/2 and increased the Hsd17b4 (Fig. 3d,f), suggesting that treatments in addition to mitochondrial part also targets the final step of testosterone production in Leydig cells.
CD changed expression of the genes principally related to mitosteroidogenesis. Leydig cells from both groups were individually isolated at different time points for 24 h. RQ-PCR analysis was performed on steroidogenesis-related genes (a–f) and main transcription factors regulating steroidogenic elements expression (g–k). CD affected the transcription pattern of genes involved in steroidogenesis in mitochondria (b–d) but also positive (Nur77, g) and negative regulatory genes (Arr19, j). Statistical significance: *p < 0.05; **p < 0.01; ***p < 0.005; ****p < 0.001 determined by two-way ANOVA.
Next, the gene expression of the most important transcription factors that regulate steroidogenesis was performed (Fig. 3g–k; Suppl. Table 2). All analyzed transcription factors, except Sf1 (Fig. 3h) showed daily fluctuation in its activity: positive regulators, Nur77 (Fig. 3g; Suppl. Table 2) peaked in dark (active) while Creb (Fig. 3i; Suppl. Table 2) reached a maximum at the end of light (inactive) phase; negative regulators Dax1 (Fig. 3k; Suppl. Table 2) peaked at the beginning of dark while Arr19 (Fig. 3j) at the beginning of light period. In CD conditions expression pattern of Sf1, Creb, Dax1 and Arr19 (Fig. 3h–k; Suppl. Table 2) were preserved. However, mRNA for Nur77 was downregulated at ZT17 (Fig. 3g).
In addition, a diurnal clock gene expression profile was measured (Fig. 4). A marked change in diurnal transcription rhythms was observed for: positive clock elements Bmal1 (Fig. 4a) and Clock (Fig. 4b), and negative Cry1 (Fig. 4e), Reverba (Fig. 4f), and Reverbb (Fig. 4g) which have increased without changes in expression times (Suppl. Table 2) while Per1 (Fig. 4c) and Per2 (Fig. 4d) reversed phase. Positive regulators within the secondary loop, Rora (Fig. 4h) and Rorb (Fig. 4i), substantially did not change transcription rhythm.
CD changed clock genes expression. Leydig cells from both groups were individually isolated at different time points for 24 h. RQ-PCR analysis of clock genes were performed in the second day of the light period in the CD protocol. CD changed diurnal transcriptional rhythms: increased positive regulators Bmal1 (a) and Clock (b) and negative Cry1 (e), Reverba (f) and Reverbb (g) but also decreased Per1 (c) and Per2 (d) in ZT11 and ZT17. Transcription of Rora (h) and Rorb (i) were unaltered. Statistical significance: *p < 0.05; **p < 0.01; ***p < 0.005; ****p < 0.001 determined by two-way ANOVA.
CD changed mitochondrial function in Leydig cells
Because the expression of genes involved in mitosteroidogenesis was decreased, it was reasonable to investigate other aspects of mitochondrial physiology, especially energetic function. Since mitochondrial membrane potential (Δψm) contributes to many processes, including mitochondrial energetics and steroidogenesis20, it was interesting to examine whether CD affects the Δψm value in Leydig cells. Changes in the electrochemical gradient Δψm were measured by TMRE fluorescence since the magnitude of TMRE fluorescence is proportional to the Δψm changes. Obtained results revealed daily rhythmic variations of the Δψm in Leydig cells, which reaches a maximum at the end of the light and the beginning of the dark phase (Fig. 5b) and coincides with the time the increase in blood testosterone (Fig. 1d). Also, diurnal rhythmicity of ATP values was observed in Leydig cells from control rats (Fig. 5c; Suppl. Table 2). ATP reached its maximum in the dark phase (Fig. 5c; Suppl. Table 2) close to the peak of the Δψ. However, CD decreased mesor of Δψm fluctuation compared to control groups without altering peak phasing (Fig. 5b) and reduced and flattened the pattern of ATP oscillation in Leydig cells (Fig. 5c; Suppl. Table 2). The observed decrease of Δψm and ATP in CD rats was not associated with the overall decrease of mitochondrial mass estimated through Mt-Nd1/B2m ratio (Fig. 5d) or mitotrack assay (Fig. 5a).
CD changed mitochondrial function. Leydig cells from both groups were individually isolated at different time points during 24 h and followed by measurements to estimate mitochondrial mass and function: mitochondrial mass was estimated through mitotrack assay (a) and Mt-Nd1/B2m ratio (d), mitochondrial membrane potential (ΔΨ) was estimated through TMRE measurement (b) and mitochondrial energetic function was estimate through ATP measurement (c). CD decreased mesor of ΔΨ and ATP production. Mitochondrial biogenesis were estimated by transcriptional activity for several genes (e–j); mitochondrial fusion (k–m); mitochondrial fission (n,o) and mitophagy (p–r). Statistical significance: *p < 0.05; **p < 0.01; ***p < 0.005; ****p < 0.001 determined by two-way ANOVA.
It is well known that mitochondrial function is regulated by mitochondrial fusion/fission closely linked with mitobiogenesis and mitophagy21. To see if CD in Leydig cells is associated with changes in mitochondrial fusion/fission or/and by altered mitochondrial biogenesis or mitophagy the expression of genes involved in these processes were monitored by RQ-PCR. An upregulation of mRNA levels in CD conditions were observed for Ppargc1a (main regulator of mitochondrial biogenesis and function; Fig. 5e), its downstream gene Nrf1 (Fig. 5f) as well as Cytc (Fig. 5h), Cox4 (a subunit of complex IV; Fig. 5i) and Mtnd1 from mitochondrial genome (Fig. 5j). However, transcriptional pattern of Tfam (Fig. 5g) was not significantly altered. Among mitochondrial profusion genes, only Mfn2 (Fig. 5l) was stimulated while Mfn1 (Fig. 5k) and Opa1 (Fig. 5m) were unchanged. Transcription of profission genes Drp1 (Fig. 5n) and Fis1 (Fig. 5o) was without changes in CD group while Pink1 (Fig. 5q) and Tfeb (Fig. 5r) were up regulated but not Prkn (Fig. 5p). Obtained results proposed activation of genes responsible for mitochondrial biogenesis but also activation of genes that promote mitophagy, establishing an equilibrium in the relationship of these processes without changing the mass but with decreased mitochondrial functionality.
Discussion
Infertility or reduced fertility is a global health problem that affects 8–12% of couples worldwide22. Male infertility accounts for about 50% of detected cases, of which for almost 70%, the cause is unknown18. Previous pre-/clinical studies suggested that CD may negatively interfere with blood levels of reproductive hormones and sperm function19,23,24,25. However, the pieces of evidence are still debatable, especially in terms of specific processes that are affected by CD. Therefore, we analyzed the effects of CD, induced by changed photoperiodism, on adult rat Leydig cell testosterone production.
Our data indicate that CD changed rhythmic Leydig cells activity leading to decreased testosterone synthesis. This is associated with altered rhythmic transcription of core clock genes (increased Bmal1, Clock, Cry1, Reverba/b and reversed pattern of Per1/2) and steroidogenic genes (reduced Star, Cyp11a1, Hsd3b1/2 and Hsd17b4). Reduced testosterone production and observed changes in genes expression are not the results of stress since the elevation of blood corticosterone secretion was not detected. Opposite, in CD caused by inverted feeding, high circulating levels of corticosterone were observed26, indicating cooperation of clock and stress systems in response to changes in the environment. On the contrary, our experimental CD model is driven by a mismatch of endogenous circadian rhythms with altered weekly photoperiodic cycles, and adaptation to newly emerging conditions does not include a stress response. Yet, disturbed daily transcriptional profiles of genes encoding LH and GNRHR were observed, suggesting disturbance of pituitary rhythmic activity. Since a temporally organized and synchronized reproductive axis is essential for reproduction27, pituitary and Leydig cells' rhythm disorders could be related to a decrease in testosterone synthesis and weakening of reproductive function.
The obtained results showed that CD alters the endocrine function of Leydig cells by disrupting mitochondrial function. This is supported by three lines of evidences indicating reduced mitosteroidogenesis, reduced mitoenergetics, and altered mitochondrial dynamics. First, CD changed the expression of essential steroidogenic genes functionally connected to mitochondria (Star, Cyp11a1, and Hsd3b1/2). The Leydig cell`s mitosteroidogenesis is triggered by LH-cAMP-PRKA signaling, which activates mitochondria-targeted StAR protein involved in cholesterol transport into the inner mitochondrial membrane28 and its conversion to pregnenolone by CYP11A1 followed by conversion to progesterone by HSD3b1/229. Activation of mitochondrial CYP11A1 is a hormonally regulated and rate-limiting step and is considered as a determinant of the steroidogenic capacity of the cells30. Besides, CD abolishes the characteristic rhythmic transcriptional profile of the Nur77 that regulates expression of steroidogenesis-related genes, including Star and Hsd3b1/231.
Second, CD changed the diurnal profile and dampened ΔΨm fluctuation and ATP production in Leydig cells. It is known that maintenance of ∆ψm is critical for ATP synthesis and steroidogenic mitochondrial activities20,32,33,34. Actually, mitochondrial electron transport chain, which drives formation of the ∆ψm utilized for ATP synthesis, also facilitate cholesterol transport into mitochondria and LH-mediated testosterone production20. In Leydig cells from control rats, these processes are temporally synchronized on 24 h scale because the peak of ∆ψm fluctuations occurs at the end of the light phase close to the height of blood testosterone, followed by a maximum of ATP production. Further, gonadotropin stimulation of Leydig cells provokes increase of ∆ψm, ATP and testosterone production34 suggesting possible synchronization through hormones of reproductive axis.
Third, CD changed the transcription of main markers of mitochondrial biogenesis and mitophagy. It is well known that the effective energetic and steroidogenic cell function is determined by the quantity and dynamics of the mitochondrial network regulated by specific gene activities21,35. In Leydig cells from CD rats, the increased transcription of main markers of mitochondrial biogenesis (Ppargc1a, Nrf1, Cytc, Cox4, and Mtnd1 from the mitochondrial genome) was detected, suggesting a possible effect on mitochondrial mass. However, mitochondrial content, as measured by levels of mtDNA, or Mitotrack assay, was not found to be changed, implying increased mitochondrial removal through mitophagy. Such a prediction is supported by the up-regulation of Pink1 and Tfeb, which are involved in maintaining mitochondrial homeostasis through mitophagy. Still, a lot of data shows circadian clock involvement in regulating mitochondrial biogenesis, fission/fusion, and mitophagy36. For example, the CLOCK-BMAL1 complex stimulates mitochondrial biogenesis and mitophagy by activating SIRT136. A similar scenario may occur in Leydig cells in circadian desynchrony when Bmal1 and Clock are stimulated but also genes regulating mitochondrial biogenesis (Ppargc1a, Nrf1) and mitophagy (Pink1 and Tfeb). Additionally, most of the genes (Ppargc1a, Cytc, Cox4, Tfam, Mfn1/2, Drp, Pink1, and Parkin) are involved in the regulation mitobiogenesis/dynamics/mitophagy in Leydig cells show daily fluctuation in transcriptional activity, but again, without detected effect on mitochondrial mass. This finding is in line with observation in human skeletal muscle37 and in synchronized immortalized human hepatic cells38, supporting the role of the circadian clock orchestrating the functioning of mitochondria which, therefore, may be impaired in circadian desynchrony.
In addition to affecting the Leydig cell's mitochondrial function, the CD also targets the final step in testosterone production by stimulating Hsd17b expression. The Hsd17b encodes the enzyme responsible for the last and critical step in testosterone synthesis, i.e., converting androstenedione to testosterone39. Therefore, an increase in transcription of Hsd17b may be associated with an increase in Bmal1 since it was shown that BMAL1 positively regulates Hsd17b transcription in goat Leydig cells9. However, stimulation of the last step in steroidogenesis is not enough to overcome the adverse effects of CD on testosterone production.
In conclusion, our study reveals the harmful effect of circadian desynchrony established by changing photoperiod on rat Leydig cells physiology. In such conditions, Leydig cells' cost is a decrease in steroidogenic ability, mainly due to a clock disturbance and reduction of mitochondrial function (Fig. 6). Given the importance of testosterone in reproduction, future research is needed to determine the cost of circadian desynchrony on reproductive capacity, including effects on spermatogenesis and sperm functionality.
Methods
Ethical approval
Animal maintenance and conduction of all the experiments were performed in the Laboratory of Chronobiology and Aging and the Laboratory for Reproductive Endocrinology and Signaling, Department of Biology and Ecology, Faculty of Sciences, University of Novi Sad. All the experiments and protocols were approved by the Committee on Animal Care University of Novi Sad (statement no. 2020-01-02), operating under the rules of the National Council for Animal Welfare and following statements of National Law for Animal Welfare (copyright March 2009). Every experiment was performed and conducted followed by “European Convention for the Protection of Vertebrate Animals used for Scientific Purposes” (Council of Europe No 123, Strasbourg, 1985) and NIH Guide for the Use and Care of Laboratory Animals (NIH Publications, No. 80 23, revised, 7th ed., 1996). Experiments were in adherence to the ARRIVE guidelines.
Animals and in vivo experiment
All the experiments were carried out on 9 months old male Wistar rats bred and raised in the animal facility of Faculty of Sciences, University of Novi Sad (Serbia). Rodents were grown in controlled temperature conditions (22 ± 2 °C) with food and water ad libitum. As for the light regime, experimental animals were living under 2 days constant light, 2 days constant darkness and 3 days 12 h light and 12 h dark respectively, for 2 months. Control group of animals were held under LD regime. After 2 months, animals were quickly sacrificed by decapitation in five different time points during 24 h (ZT3, ZT11, ZT17, ZT20, ZT23; ZT0 is time when light switch on; five animals per time point for each group). The method of sacrificing was chosen to avoid the effect of anesthesia on blood hormone levels. The experiment was completed on the second day of the light period in the CD protocol. Rats’ body weight was measured once per week during experiment. Afterwards, trunk blood was collected individually, testes were removed for Leydig cells isolation and purification.
Detection of an animal’s voluntary activity
In order to monitor rat’s voluntary activity, a group of control and experimental rats were placed individually in cages with running wheel system. The detection system was programmed to record turns of the wheel that animal made every 6 min. Activity pattern of rats was monitored for 2 months and data was collected and used for forming actograms as one of the standard ways to represent circadian rhythms. The graphical representation of the animal’s activity was formed using R software.
Hormone level measurement
Hormone levels were measured by radioimmunoassay. According to anti-testosterone No. 250 100% cross-reactivity with DHT40, testosterone level was referred as testosterone + dihydrotestosterone. Samples were measured in duplicate (sensitivity: 6 pg per tube; intraassay coefficient of variation 5–8%). Corticosterone EIA Kit (Caymanchem, Michigan, USA) was used for detecting corticosterone levels, with 30 pg/ml as the lowest standard significantly different from blank. All samples were measured in duplicate.
Leydig cells purification
Testes were decapsulated, placed in 50 ml plastic tubes (per animal) containing 3 ml of collagenase solution (1.25 mg/ml collagenase; 1.5% bovine serum albumin (BSA); 20 mM HEPES, Sigma, St Louise, Missouri) and incubated in shaking-water bath (15 min/34 °C/120 cycles/min). The collagenase reaction was stopped by adding cold M199 medium and seminiferous tubules were removed by filtration. The resulting interstitial cells’ suspension was centrifuged at 160×g/5 min. Afterwards, cell pellet was resuspended (8 ml per animal) using DMEM/F12. Resuspended cells were placed on Percoll gradients (Sigma, St Louis, Missouri) with densities of 1.080, 1.065 and 1.045 g/ml and centrifuged 1100×g/28 min (without brake). Purified Leydig cells were collected from 1.080/1.065 g/ml and 1.065/1.045 g/ml and washed with adequate amount of 0.1% BSA-M199, centrifuged at 200×g/5 min and resuspended in 5 ml DMEM/F12. According to HSD3B staining3 proportion of Leydig cells presence was around 90%, and viability of the cells according to the Trypan blue exclusion test was greater than 90%. The suspension of Leydig cells was further centrifuged and the pellet was stored at − 70 °C until RNA and protein analysis.
Mitochondrial abundance and mitochondrial membrane potential
After Leydig cell isolation, as previously described, cells were plated in two 96 well-plate (1 × 105 and 0.5 × 105 cells/well) and incubated with tetramethylrhodamine (TMRE; Thermo Fisher Scientific, Waltham, MA, USA) and MitoTracker Green, respectively, staining for 20 min/34 °C/5% CO2. Fluorescence was detected on fluorimeter (Fluoroscan, Ascent, FL) using excitation wavelengths 590 and 510 nm whereas emission wavelengths 550 and 485 nm, respectively. Cells were washed using 0.1% BSA-PBS and stored until Bradford method protein quantification.
ATP level measurement
Level of ATP was estimated using the ATP Bioluminescence CLS II kit (Roche Diagnostics, Mannheim, Germany) following the protocol recommended by the manufacturer. Leydig cells (1 × 106/tube) were resuspended in boiling water and Tris–EDTA (1:9), incubated 3 min/100 °C and centrifuged 900×g/1 min. The resulting supernatant was used for ATP level detection whilst the pellet was further utilized for Bradford method analysis. Luciferase reagent was mixed together with standard/sample (1:1) in order to measure luminescence (Biosystems/luminometer, Fluorescan, Ascent, FL).
Genomic DNA isolation, total RNK purification, RT-PCR and RQ-PCR analysis
Genomic DNA from Leydig cells was purified using NucleoSpin Tissue, DNK, RNA and protein purification Macherey-Nagel (Dueren, Germany). Total RNA from Leydig cells and pituitary gland was isolated using GenEluteTM Mammalian Total RNA Miniprep (Qiagen, Hilden, Germany) and EXTRAzol reagent (Birt, Gdańsk, Poland) respectively, following a recommended protocol. DNA and RNA quality was measured and validated using BioSpec-nano (Shimadzu Biotech, Kyoto, Japan). Afterwards, RNA samples were treated with DNase-I treatment (New England Biolabs, Ipswich, Massachusetts, USA). cDNA synthesis was performed using HighCapacity kit (Thermo Fisher Scientific, Waltham, MA, USA) according to manufacturer’s protocol. RQ-PRC SYBR-Green based technology (Applied Biosystems/Thermo Fisher Scientific, Massachusetts, USA) was utilized to determine relative gene expression. The transcription of Gapdh was used to correct the variations in cDNA between the samples. The reaction was performed in presence of 5 µl cDNA and specific primers (Suppl. Table 1).
Statistical analysis
Statistical analysis was performed using GraphPad Prism 8. Experimental results are individually presented as scaterogram or shown as mean value ± SEM variation. Two-way ANOVA was used to analyze results between the groups including all time points. Rhythm parameters (p, MESOR, Amplitude and Acrophase) were obtained using cosinor method online, fitted to 24 h (https://cosinor.online/app/cosinor.php) (Suppl. Table 2).
Data availability
The datasets generated and analyzed during the current study are available in the Cloud of Faculty of Sciences University of Novi Sad, [https://cloud.pmf.uns.ac.rs/s/jDsJS6GwFH6Hrzg]; data from supplemental Fig. 1 are available at [https://cloud.pmf.uns.ac.rs/s/nmcL6idHTDieATy].
References
Albrecht, U. Timing to perfection: The biology of central and peripheral circadian clocks. Neuron 74, 246–260 (2012).
Takahashi, J. S. Transcriptional architecture of the mammalian circadian clock. Nat. Rev. Genet. 18, 164–179 (2017).
Payne, A. H., Downing, J. R. & Wong, K. L. Luteinizing hormone receptors and testosterone synthesis in two distinct populations of Leydig cells. Endocrinology 106, 1424–1429 (1980).
Baburski, A. Z. et al. Melatonin replacement restores the circadian behavior in adult rat Leydig cells after pinealectomy. Mol. Cell. Endocrinol. 413, 26–35 (2015).
Baburski, A. Z. et al. Circadian rhythm of the Leydig cells endocrine function is attenuated during aging. Exp. Gerontol. 73, 5–13 (2016).
Chen, H., Wang, Y., Ge, R. & Zirkin, B. R. Leydig cell stem cells: Identification, proliferation and differentiation. Mol. Cell. Endocrinol. 445, 65–73 (2017).
Li, C. et al. Bisphenol A attenuates testosterone production in Leydig cells via the inhibition of NR1D1 signaling. Chemosphere 263, 128020 (2021).
Zhao, L. et al. Zearalenone perturbs the circadian clock and inhibits testosterone synthesis in mouse Leydig cells. J. Toxicol. Environ. Health A 84, 112–124 (2021).
Xiao, Y. et al. Circadian clock gene BMAL1 controls testosterone production by regulating steroidogenesis-related gene transcription in goat Leydig cells. J. Cell. Physiol. 236, 6706–6725 (2021).
Keating, R. J. & Tcholakian, R. K. In vivo patterns of circulating steroids in adult male rats. I. Variations in testosterone during 24- and 48-hour standard and reverse light/dark cycles. Endocrinology 104, 184–188 (1979).
Thorpe, J. B., Rajabi, N. & Decatanzaro, D. Circadian rhythm and response to an acute stressor of urinary corticosterone, testosterone, and creatinine in adult male mice. Horm. Metab. Res. 44, 429–435 (2012).
Waite, E., Kershaw, Y., Spiga, F. & Lightman, S. L. A glucocorticoid sensitive biphasic rhythm of testosterone secretion. J. Neuroendocrinol. 21, 737–741 (2009).
Baburski, A. Z., Andric, S. A. & Kostic, T. S. Luteinizing hormone signaling is involved in synchronization of Leydig cell’s clock and is crucial for rhythm robustness of testosterone production. Biol. Reprod. 100, 1406–1415 (2019).
Alvarez, J. D. et al. The circadian clock protein BMAL1 is necessary for fertility and proper testosterone production in mice. J. Biol. Rhythms 23, 26–36 (2008).
Ding, H., Zhao, J., Liu, H., Wang, J. & Lu, W. BMAL1 knockdown promoted apoptosis and reduced testosterone secretion in TM3 Leydig cell line. Gene 747, 144672 (2020).
Roenneberg, T., Wirz-Justice, A. & Merrow, M. Life between clocks: Daily temporal patterns of human chronotypes. J. Biol. Rhythms 18, 80–90 (2003).
Barratt, C. L. R. et al. The diagnosis of male infertility: An analysis of the evidence to support the development of global WHO guidance-challenges and future research opportunities. Hum. Reprod. Update 23, 660–680 (2017).
Sciarra, F. et al. Disruption of Circadian rhythms: A crucial factor in the etiology of infertility. Int. J. Mol. Sci. 21, E3943 (2020).
Fusco, F. et al. Impact of Circadian desynchrony on spermatogenesis: A mini review. Front. Endocrinol. (Lausanne) 12, 800693 (2021).
Midzak, A. S., Chen, H., Aon, M. A., Papadopoulos, V. & Zirkin, B. R. ATP synthesis, mitochondrial function, and steroid biosynthesis in rodent primary and tumor Leydig cells. Biol. Reprod. 84, 976–985 (2011).
Sebastián, D., Palacín, M. & Zorzano, A. Mitochondrial dynamics: Coupling mitochondrial fitness with healthy aging. Trends Mol. Med. 23, 201–215 (2017).
Agarwal, A. et al. Male infertility. Lancet 397, 319–333 (2021).
Liu, K. et al. Adverse effects of circadian desynchrony on the male reproductive system: An epidemiological and experimental study. Hum. Reprod. 35, 1515–1528 (2020).
El-Helaly, M., Awadalla, N., Mansour, M. & El-Biomy, Y. Workplace exposures and male infertility—A case-control study. Int. J. Occup. Med. Environ. Health 23, 331–338 (2010).
Kohn, T. P. & Pastuszak, A. W. Shift work is associated with altered semen parameters in infertile men. Fertil. Steril. 108, e323–e324 (2017).
Lassi, M. et al. Disruption of paternal circadian rhythm affects metabolic health in male offspring via nongerm cell factors. Sci. Adv. 7, eabg6424 (2021).
Sen, A. & Hoffmann, H. M. Role of core circadian clock genes in hormone release and target tissue sensitivity in the reproductive axis. Mol. Cell. Endocrinol. 501, 110655 (2020).
Stocco, D. M., Wang, X., Jo, Y. & Manna, P. R. Multiple signaling pathways regulating steroidogenesis and steroidogenic acute regulatory protein expression: More complicated than we thought. Mol. Endocrinol. 19, 2647–2659 (2005).
Payne, A. H. & Hales, D. B. Overview of steroidogenic enzymes in the pathway from cholesterol to active steroid hormones. Endocr. Rev. 25, 947–970 (2004).
Miller, W. L. Steroid hormone synthesis in mitochondria. Mol. Cell. Endocrinol. 379, 62–73 (2013).
Tremblay, J. J. Molecular regulation of steroidogenesis in endocrine Leydig cells. Steroids 103, 3–10 (2015).
Midzak, A. S., Chen, H., Papadopoulos, V. & Zirkin, B. R. Leydig cell aging and the mechanisms of reduced testosterone synthesis. Mol. Cell. Endocrinol. 299, 23–31 (2009).
Allen, J. A. et al. Energized, polarized, and actively respiring mitochondria are required for acute Leydig cell steroidogenesis. Endocrinology 147, 3924–3935 (2006).
Medar, M. L. J. et al. Dependence of Leydig cell’s mitochondrial physiology on luteinizing hormone signaling. Life (Basel) 11, 19 (2020).
Chen, H. & Chan, D. C. Emerging functions of mammalian mitochondrial fusion and fission. Hum. Mol. Genet. 14 Spec No. 2, R283–R289 (2005).
de Goede, P., Wefers, J., Brombacher, E. C., Schrauwen, P. & Kalsbeek, A. Circadian rhythms in mitochondrial respiration. J. Mol. Endocrinol. 60, R115–R130 (2018).
van Moorsel, D. et al. Demonstration of a day-night rhythm in human skeletal muscle oxidative capacity. Mol. Metab. 5, 635–645 (2016).
Cela, O. et al. Clock genes-dependent acetylation of complex I sets rhythmic activity of mitochondrial OxPhos. Biochim. Biophys. Acta 1863, 596–606 (2016).
Zirkin, B. R. & Papadopoulos, V. Leydig cells: Formation, function, and regulation. Biol. Reprod. 99, 101–111 (2018).
Kostic, T. S., Stojkov, N. J., Janjic, M. M. & Andric, S. A. Structural complexity of the testis and PKG I/StAR interaction regulate the Leydig cell adaptive response to repeated immobilization stress. Int. J. Androl. 33, 717–729 (2010).
Acknowledgements
We are grateful to Professor dr Gordon Niswender (Colorado State University) for supplying antibodies for radioimmunoassay analysis. Figure 6 was created with BioRender.com (Agreement No DA23UG6JAH).
Funding
This research was supported by the Ministry of Education, Science and Technological Development of the Republic of Serbia (Grant No. 451-03-68/2022-14/200125) and CeRES grant, and the Autonomic Province of Vojvodina, Serbia, grants no. APV2708.
Author information
Authors and Affiliations
Contributions
M.V.P.—acquisition of the data; analysis and interpretation of the data; revising manuscript critically for important intellectual content; final approval of the version to be submitted. D.Z.M.—acquisition of the data; analysis and interpretation of the data; revising manuscript critically for important intellectual content; final approval of the version to be submitted. S.A.A.—acquisition of the data; analysis and interpretation of the data; revising manuscript critically for important intellectual content; final approval of the version to be submitted. T.S.K.—the conception and design of the research; acquisition of the data; analysis and interpretation of the data; drafting the manuscript; revising manuscript critically for important intellectual content; final approval of the version to be submitted. All authors—approved the submitted version (and any substantially modified version that involves the author’s contribution to the study); agreed both to be personally accountable for all aspect of the work as well as the author’s own contributions and to ensure that questions related to the accuracy or integrity of any part of the work, even ones in which the author was not personally involved, are appropriately investigated, resolved, and the resolution documented in the literature; qualify for authorship, and all those who qualify for authorship are listed.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pavlovic, M.V., Marinkovic, D.Z., Andric, S.A. et al. The cost of the circadian desynchrony on the Leydig cell function. Sci Rep 12, 15520 (2022). https://doi.org/10.1038/s41598-022-19889-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-19889-9
This article is cited by
-
Understanding how space travel affects the female reproductive system to the Moon and beyond
npj Women's Health (2024)
-
An update on the role and potential mechanisms of clock genes regulating spermatogenesis: A systematic review of human and animal experimental studies
Reviews in Endocrine and Metabolic Disorders (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.