Abstract
Amphibians have some of the most variable genome sizes among vertebrates. Genome size variation has been attributed to repetitive and noncoding DNA, including satellite repeats, transposable elements, introns, and nuclear insertions of viral and organelle DNA. In vertebrates, satellite DNAs have been widely described in mammals, but few molecular studies have been carried out in amphibians. Here, we provide a detailed characterization of a new family of satellite DNA, present in all 15 examined species of the family Bufonidae. Southern-blot analysis and PCR reveal that this satellite is formed by monomers of 807 bp, is organized in tandem arrays, and has an AT-content of 57.4%. Phylogenetic analyses show that most clades exhibit species-specific variances, indicating that this satellite DNA has evolved by concerted evolution. The homogenization/fixation process is heterogeneous in Bufonidae, where the genera Bufo and Bufotes do not show species-specific differences, while populations from Rhinella marina exhibit population-specific changes. Additionally, variants of this satellite DNA have been identified in Duttaphrynus melanostictus and R. marina, supporting the ‘library hypothesis’ (a set, ‘library’, of satellite DNAs is shared by a species group). Physical mapping in Bufo bufo, Bufo spinosus, Epidalea calamita and Bufotes viridis provides evidence that this repetitive DNA is not dispersed in the karyotype, but accumulated in pericentromeric regions of some chromosomal pairs. This location, together with its presence in the transcriptomes of bufonids, could indicate a role in centromere function or heterochromatin formation and maintenance.
Similar content being viewed by others
Introduction
Satellite DNA sequences are repeated in tandem arrays, hundreds to thousands of times in the genome. They are predominantly localized in the centromeric, pericentromeric, and telomeric regions, and present the most abundant components of constitutive heterochromatin1. Monomers of satellite DNA accumulate mutations, but the divergence between monomers is usually low as they do not evolve independently but by concerted evolution. This involves a two-step process: (1) changes spread to other sequences (homogenization) by non-reciprocal exchanges (such as uneven crossing-over, gene conversion, rolling-circle amplification, or slippage replication), and (2) they are fixed (or eliminated) in reproductively isolated groups2,3. Furthermore, several sets of variants and satellite DNAs can be shared by a species group (also called a ‘library’), leading to species-specific profiles due to differential amplification and/or contractions of repetitive sequences within such a library2,4. Satellite DNA evolution may also be affected by other factors, such as location within heterochromatin, the reproductive biology of the species, or by its population size2.
Repetitive DNA research has mostly been based on analyses of randomly cloned monomers and short multimers, obtained after digestion of genomic DNA and mapping by fluorescent in situ hybridization (FISH)—techniques, which are still used today5. The application of whole genome sequencing to satellite DNA research has been limited to graph-based clustering analysis and manual annotation, and only recently the availability of long read technology overcomes the problems posed by the assembly of long repetitive motives.
Satellite DNA is no longer considered ‘junk’. These sequences have been implicated in genome regulation6, transcription7, centromere identity8, genome architecture9,10, and other important cell functions11. Applications to evolutionary studies are also interesting, as they can provide information on the divergence between species12 and about the existence of possible mechanisms of reproductive isolation, triggered, for example, by the diversification of centromeric repetitive DNA sequences13. However, even with recognizing the functional importance of satellite DNA, the structural and functional knowledge about these sequences is still limited.
Amphibians are the extant vertebrate class with the largest range of genome sizes14. The differences of DNA-content and chromosome size in this group were not attributed to the visible presence of constitutive heterochromatin15. In the past, numerous studies about the re-association kinetics of satellite DNA and chromosomal organization in amphibians concluded that repetitive sequences play an important role in chromosome size differences in anurans (Supplementary Bibliography, references 1–6). Scarce information about satellite DNA obtained by genome-wide analysis confirms the high amount of satellite DNA present in amphibian genomes (e.g. 15.7% in Proceratophrys boiei16).
Bufonidae presents the fourth largest systematic amphibian family, with 52 genera and 634 known species17 (638 as reported in Amphibian Species of the World18). This group has been subject to many phylogenetic studies, although its systematics is challenging. Along with Pipidae and Ranidae, Bufonidae are among the best-studied amphibian families. However, molecular and genetic data about genome organization in this group remain insufficient. The only information available about repetitive DNA sequences is restricted to a single 420 bp sequence (AY585341) from a satellite DNA (pBv) isolated from a single green toad (Bufotes viridis species group), probably present in several Bufonidae genera (Bufo, Bufotes, and probably Werneria and Wolterstoffina)19.
Repetitive sequences in amphibians are related to/derived from transposons, like the highly repetitive DNA R.e/Tc1 in Ranidae (Pelophylax esculentus)20, from 5S rDNA, like the PcP190 satellite of Hyloidea21, or telomeric sequences, like the ITRs in Pipidae (genus Xenopus)22. Initially, most amphibian satellite DNAs were described in Caudata, primarily in Triturus, and in other genera or families (Supplementary Bibliography, references 7–12). In Anura, satellite DNA has been widely studied in Xenopus, Rana, Pelophylax, and Discoglossidae (Supplementary Bibliography, references 13–28).
Here, we identify a new satellite DNA-family, present in several species of Bufonidae. This satellite DNA has a long monomer size of 807 bp, organized in tandem arrays. Phylogenetic analysis suggests that it has evolved according to the concerted evolution model. Different homogenization/fixation rates are observed in different species. Accordingly, the homogenization of sequences appears unfinished in some species, while we observed differences between populations in other taxa. Furthermore, variants supporting the ‘library hypothesis’ are also present in some species. The accumulation of these repetitive sequences in pericentromeric regions of some chromosomal pairs in Bufo bufo, Bufo spinosus, E. calamita, and Bufotes viridis, together with the evidence of their transcription after the analysis of bufonid transcriptomes, may implicate them in centromere function or heterochromatin formation and maintenance.
Results
Identification, cloning and characterization of a repetitive DNA family in species of the Bufo bufo group
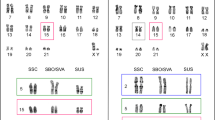
Genomic DNA from one adult Bufo bufo female from Bologna (Italy) was digested with the restriction endonuclease BamHI. Electrophoresis of digested DNA showed several intense bands, indicating the presence of several repetitive DNAs (Fig. 1a). The band of ca. 800 bp was excised, purified, labelled and used as a probe in a Southern blot analysis. Positive bands of about 800 bp, 1600 bp, and 2400 bp revealed a “ladder”-like pattern of bands typical of repetitive DNA, organized in tandem arrays (Fig. 1b). The same band pattern was observed when genomic DNA from Bufo spinosus (Jaén, Spain) was analysed (Fig. 1c). Except for the bands due to star activity of BamHI, no variation in the banding pattern of Southern-blot experiments was found when DNAs from both species or both sexes were analysed (Fig. 1c,d).
(a) Reversed image of the agarose gel electrophoresis of a genomic DNA sample from Bufo bufo (Bologna, Italy) digested with BamHI. Intense bands indicate different repetitive DNA families (*). The white arrowheads point to the position of the analysed band, that was eluted from the gel and cloned into BamHI-digested pCU19 vector. (b) Inverted image of the agarose gel electrophoresis of a DNA sample from Bufo bufo H2 (Jaén, Spain) digested with BamHI and blotted onto a nylon membrane, hybridized with the dig-labelled BamHI-800 bp band. A ladder-like pattern is observed (800, 1600, 2000 and 2400 bp). (c) Southern-Blot of BamHI-digested genomic DNA from Bufo bufo (B. buf: sample from Bologna, Italy) and Bufo spinosus (B. spi: sample from Jaén, Spain). (d) Southern-Blot of BamHI-digested genomic DNA from male (M) and female (F) Bufo spinosus (both from Jaén). (e) PCR using primers Bbu-BamHI-800-F1/R1 (PCR1) or Bbu-BamHI-800-F2/R2 (PCR2) (reversed image) with genomic DNA from Bufo spinosus. Black arrowheads point to 800, 1600, and 2400 bp bands; the bands labelled with (*) in B and C are explained by the star activity of BamHI restriction enzyme, as this band is not always observed. (f) PCR amplification of BamHI-800 sequences in other species of the Bufonidae family. Code used for the name of the species: Bufotes boulengeri (B. bou), E. calamita (E. cal), Bufotes luristanicus (B. lur), Bufotes surdus (B. sur), Bufotes viridis (B. vir), B. bufo (B. buf), R. marina (R. mar), Bufotes latastii (B. lat), S. arabica (S. ara), Bufotes siculus (B. sic), and Bufotes balearicus (B. bal), Ba. brongersmai (B. bro), and D. melanostictus (D. mel). Original gels/blots are presented in Supplementary Fig. S7. For a detailed description of molecular weight markers see Supplementary Table S9.
The band from Bufo bufo of about 800 bp was cloned in a dephosphorylated BamHI-digested pUC19 vector (Fermentas, Vilnius, Lithuania). Recombinant plasmids were purified and sequenced. The sequences from six clones were aligned and compared with each other. Their size ranged between 810 and 811 bp, except for one clone with a small deletion of 22 bp (789 bp) and an incomplete clone of 492 bp (312 bp missing from the 5′-end) that was not used in the analysis. The average A + T content was 57.5%, and no direct or inverted internal repeats were observed, except for a small fragment of 20 bp, repeated 1.9 times (positions 13–50).
Based on these sequences, two primer pairs with opposite orientations were designed (Supplementary Fig. S1). PCRs with both primer pairs gave the same banding pattern: 800 bp, 1600 bp, and 2400 bp (Fig. 1e), indicating that BamHI-800 sequences are monomeric units of a repetitive DNA family, organized in tandem arrays. This was further confirmed as positive clones obtained from the 1600 bp PCR band are dimers of 800 bp monomer units, separated by target sites for BamHI.
Using BamHI-800-specific primers (Supplementary Fig. S1), BamHI-800 sequences were amplified from different DNAs from Bufo spinosus specimens. Cloning and sequencing these amplicons gave five sequences from a Bufo spinosus sample from Morocco and 38 sequences from several Bufo spinosus from Spain (Jaén) (Table 1 and Supplementary Table S1). Incomplete clones (182 and 655 bp) were not analysed (details: Supplementary Table S1).
The alignment of all Bufo bufo group sequences (Supplementary Fig. S2) resulted in a consensus sequence of 819 bp (deletions from 1 to 28 bp were observed in some sequences). This repetitive motif has an average A + T composition of 57.4%, while the average R (transition/transversion ratio) was 0.85. When Bufo bufo-group BamHI-800 sequences are grouped according to species (Bufo bufo and Bufo spinosus), population (Bologna, Spain and Morocco), sex (male and female), or experimental procedure ((i) bands: obtained from BamHI digested genomic DNA fragments, (ii) PCR products with F1/R1 primers, (iii) PCR products with F2/R2 primers), the differences in the average genetic distances within and between groups were not significant (Supplementary Table S2, Supplementary Fig. S3).
BamHI-800 sequences in other amphibian species
To evaluate the presence of this satellite DNA in other amphibian species, we carried out Southern-Blot analysis on BamHI-digested DNAs from several anuran families using BamHI-800 sequences from Bufo bufo as a probe. This analysis comprised six species of Bufonidae (Bufotes viridis, Bufotes boulengeri, Bufotes luristanicus, Bufotes surdus, Epidalea calamita, and Rhinella marina) and three species from other anuran families Discoglossus galganoi (Alytidae), Xenopus tropicalis (Pipidae), and Pelophylax perezi (Ranidae) (Table 1). Interestingly, positive hybridization signals corresponding to BamHI-800 sequences were only observed in Bufonidae (Supplementary Fig. S4a), with all species showing the same ladder-like pattern: ca. 800 bp, 1600 bp, and 2400 bp, corresponding to monomers, dimers, and trimers.
The presence of BamHI-800 repetitive DNA sequences was also examined by PCR in 12 bufonid toad species and in three species of other amphibian families (Table 1 for sample details). Amplification with both primer pairs (Bbu-BamHI-800-F1/R1 and Bbu-BamHI-800-F2/R2) was positive only in Bufonidae. Again, identical patterns corresponding to monomer and sometimes dimer and trimer repeats of 800 bp units were observed in most species (Fig. 1f). Small differences to this common pattern were observed in some cases: (1) less intense bands in Sclerophrys arabica and Duttaphrynus melanosticus, and (2) some extra bands in S. arabica, D. melanosticus and Barbarophryne brongersmai (Fig. 1f and Supplementary Fig. S4b). The monomer band was cloned and sequenced in each bufonid species in Table 1 (Supplementary Table S1 for details about clones/sequences). Of note, some clones from D. melanosticus (5 out of 14) had inserts longer than 800 bp (up to 1100 bp) that were difficult to sequence. The alignment of the sequences obtained (three partial and two complete sequences) revealed the presence of an insertion at the same position in all the clones (Supplementary Fig. S5a). In addition, the inserted sequences had long palindromes (Supplementary Fig. S5a,b) that form long hairpin structures when the secondary structure is predicted (Supplementary Fig. S5c). The existence of this palindromic region produces artefacts in PCRs from these clones when universal primers (M13 (− 20) and M13 Reverse) are used (Supplementary Fig. S5d). These extra bands of smaller size could explain the difficulty experienced to sequence these clones. The inserted region of clone Dmel PCR2-1 23 INS (303 bp) was used in BLASTN searches on DNA databases (NCBI, nr/nt). Hits in several Bufo bufo and Bufo gargarizans genes/sequences were identified, although none was related to transposable elements or repetitive sequences. For further analysis of these sequences, the inserted sequence (red box in Supplementary Fig. S5a) was removed.
BamHI-800 sequences from genetic databases
A BLASTN search on DNA databases (NCBI, nr/nt) produced only three hits: a predicted ncRNA from Bufo bufo (XR_005776185.1, 69% identity) and two predicted transcript variants from Bufo gargarizans (XR_006388564.1, XR_006388563.1, 66% identity). BLASTN searches were also performed against all amphibian genome assemblies available on NCBI (Supplementary Table S3). Exclusively bufonid genome assemblies displayed significant hits: Bufo bufo (GCF_905171765.1), Bufo gargarizans (GCA_014858855.1), and R. marina (GCA_900303285.1). Genomes from two Dendrobatidae and one Leptodactylidae species, the closer outgroups available, did not show BLASTN hits.
BLASTN searches against the Bufo bufo genome produced 5768 hits. Sequence numbers according to their chromosome/unknown localization are shown in Supplementary Table S4. A selection of 1376 sequences without deletions at the 5’- or 3’-ends was further analysed. They have an average monomer size of 809 bp, an A + T content of 57.1%, and identity ranges between 100 and 78% (Table 2).
BLASTN searches against the Bufo gargarizans genome produced 2031 hits. The number of sequences according to their chromosome/unknown localization is shown in Supplementary Table S4. A selection of 953 sequences without deletions at their 5’- or 3’-ends was further analysed. They show an average monomer size of 809 bp, an A + T content of 57.3%, and identity ranges between 100 and 71.7% (Table 2).
Finally, BLASTN searches against the R. marina genome produced 226 hits. No chromosome level assembly is available for this species. A selection of 15 sequences without deletions at their 5’- or 3’-ends was further analysed. Their average monomer size is 797 bp, an A + T content of 57.2%, and identity ranges between 100 and 69% (Table 2).
Analysis of BamHI-800 sequences
In total, 230 BamHI-800 sequences from 15 bufonid species were analysed (200 cloned sequences and 30 sequences retrieved after BLASTN on genome assemblies of Bufo bufo, Bufo gargarizans, and R. marina; first ten hits per species).
Percentages of identity (Supplementary Table S5A–C), pairwise distances between sequences (Supplementary Table S6A,B) and visual observation of the alignment (Supplementary Table S7) revealed two types of monomers (variants) in D. melanosticus, R. marina, and S. arabica. According to this, in these species, we considered as variant 1 (V1) those sequences with higher sequence identity with Bufo bufo group sequences, and as variant 2 (V2) those with lower identity values. V1 and V2 sequences were analysed together and independently.
The comparative analysis of all bufonid BamHI-800 sequences, including the results of the analysis of sequences grouped by species and variants, is shown in Table 2. The average monomer size is 807 bp (814 to 790 bp), and the average A + T% content is 57.4% (56.0% to 60.8%, both in V2 variants). Within species, this satellite DNA shows lower genetic distances when V1 or V2 sequences are analysed independently (between 0.0000 and 0.2143, Supplementary Table S6A and diagonal in Supplementary Table S6B). However, in pairwise comparisons between all the sequences, genetic distances increase to 0.4787 when V2 sequences from D. melanostictus are compared to S. arabica V1 (Supplementary Table S6A,B). Higher genetic distances were observed in pairwise comparisons that include sequences from Ba. brongersmai, D. melanosticus, R. marina, and S. arabica. However, S. arabica V2 sequences are closer to S. arabica V1 sequences than to V2 sequences from R. marina and D. melanosticus. A close look to the alignment of these sequences (Supplementary Table S7) reveal that the increased distances between V1 and V2 variants from S. arabica is mainly due to the presence of long deletions in V2 sequences, and not to a high number of nucleotide differences between them.
Overall genetic distance for bufonid BamHI-800 sequences is 0.2059 ± 0.0081. Island endemic Bufotes siculus from Sicily shows the lowest intra-species distance: 0.0206 ± 0.0038, while the highest distances are observed in species with sequence variants: D. melanosticus (0.1913 ± 0.0139), R. marina (0.1691 ± 0.0091), and S. arabica (0.1313 ± 0.0085) (Supplementary Table S6B). When sequences variants are considered, the lowest intra-species distance is in variant 1 (V1) from D. melanosticus (0.0028 ± 0.0014), while the highest is in variant 2 (V2) from S. arabica (0.1101 ± 0.0076).
The highest inter-specific distance is observed between S. arabica–D. melanosticus (0.3765 ± 0.0238) and S. arabica and Ba. brongersmai (0.3485 ± 0.0221), while the lowest distances are between Bufotes siculus–Bufotes boulengeri (0.0421 ± 0.0037) and Bufotes siculus–Bufotes balearicus (0.0423 ± 0.0046) (Supplementary Table S6C). When net distances were considered, the highest distances are between S. arabica—Ba. brongersmai (0.2462 ± 0.0209) and S. arabica–Bufotes siculus (0.2340 ± 0.0203), while the lowest distances were between Bufotes latastii–Bufotes luristanicus (0.0015 ± 0.0006) and Bufotes latastii–Bufotes surdus (0.0022 ± 0.0007) (Supplementary Table S6D).
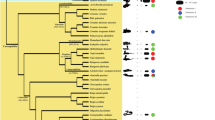
BamHI-800 sequences from 15 bufonid species were used to generate a maximum-likelihoood tree (Fig. 2). The tree contains ten major groups with bootstrap support values of 99% (Fig. 3, detailed subtrees: Supplementary Fig. S6). In each major group, sequences are clustered by species or genera (Table 3).
Circular tree for all BamHI-800 sequences using Maximum Likelihood and the Tamura 3-model of sequence evolution73. A discrete Gamma distribution was used to model evolutionary rate differences among sites (5 categories (+ G, parameter = 2.7852)). The tree with the highest log likelihood (-22,314.97) is shown. The tree is drawn to scale, with branch lengths measured in the number of substitutions per site. The percentage of trees in which the associated taxa clustered together is shown next to the main branches (values lower than 75% have been omitted). This analysis involved 230 nucleotide sequences and 746 positions in the final dataset (all positions with less than 95% site coverage were eliminated, and ambiguous bases were allowed at any position (partial deletion option)). Species names abbreviations and colour codes at branch leafs as in Fig. 1 and Supplementary Table S1.
Sequences from the genus Bufo form two groups: group I includes sequences from Bufo bufo and Bufo spinosus, while group II consists exclusively of Bufo gargarizans. Within Bufotes, the sequences form three distinct groups: subgroup IIIa, consisting of Bufotes viridis, Bufotes boulengeri, Bufotes siculus, and Bufotes balearicus; subgroup III.b, formed by Bufotes latastii and Bufotes luristanicus; and subgroup III.c, formed by Bufotes latastii, Bufotes luristanicus and Bufotes surdus. Finally, sequences from E. calamita, S. arabica, and Ba. brongersmai are included in different groups (IV, V and VI respectively), while sequences from R. marina and D. melanosticus are split in two groups in each species: groups VII and VIII include V1 sequences, and groups IX and X are for V2 sequences.
According to Fig. 3, BamHI-800 in Bufonidae is organized into eight species-specific subfamilies (Bufo gargarizans, E. calamita, S. arabica, Ba. brongersmai, R. marina V1, D. melanosticus V1, R. marina V2, and D. melanosticus V2 from groups II, IV, V, VI, VII, VIII, IX, and X respectively); and two genera-specific subfamilies (Bufo bufo group and Bufotes from groups I and III). The sequences included in the subfamilies IX and X are distant variants (V2) of the V1 sequences in the corresponding species (R. marina and D. melanosticus respectively). In the tree, the same group (V) contains V1 and V2 sequences from S. arabica.
No other BamHI-800 variant has been identified besides V2 sequences from R. marina, D. melanosticus, and S. arabica (BLASTN searches with V2 sequences in genome assemblies from bufonids do not produce hits different from V1 sequences).
Some other interesting features in BamHI-800 tree are: (1) sequences from Bufo gargarizans form a group and cluster with those of the Bufo bufo group; (2) all Bufo bufo species group sequences (from Italy (Bufo bufo), Spain (Bufo spinosus), Morocco (Bufo spinosus), and UK (Bufo bufo sequences retrieved from NCBI)) appear intermixed in their group, without significant differences between them (Supplementary Fig. S3); (3) sequences in group IV (R. marina V1) split into two separate groups (V1 sequences obtained in this work and V1 from NCBI) (Figs. 2, 3; Supplementary Fig. S6).
FISH analysis
Due to limitations in sample availability, the chromosomal distribution of BamHI-800 repetitive DNA could only be analysed in four bufonid species: Bufo bufo (Italy), Bufo spinosus (Spain), E. calamita (Spain) and Bufotes viridis (Greece). The karyotypes of these species have 2n = 22, with six pairs of large and five pairs of small metacentric or submetacentric chromosomes. In situ hybridization using a species-specific BamHI-800 probe produced positive signals located at species-specific positions (Fig. 4).
FISH on metaphase chromosomes from Bufo bufo (a,b); Bufo spinosus (c–f); Epidalea calamita (g) and Bufotes viridis (h) using species specific BamHI-800 sequences as a probe after two (c) or three (a,e,g,h) rounds of amplification. DAPI-stained metaphase chromosomes (b,d,f) corresponding to (a,c,e), respectively. Scale bar equivalent to 2.5 µm.
Bufo bufo samples display intense positive hybridization signals on the centromere and short arm of the smallest pair of the karyotype (chromosome 11). Other faint signals are also observed on pericentromeric regions of all large chromosomal pairs, on the long arm of the 6th chromosomal pair, next to the secondary constriction corresponding to the NOR, and on the long arm of chromosome pair 7, close to the centromere (Fig. 4a,b).
The hybridization pattern in Bufo spinosus is similar to Bufo bufo, except for the absence of a positive signal on chromosome 7 (Fig. 4c,d). The signals are more evident after three rounds of immunological amplification (Figs. 4e,f and 5a). The presence and the intensity of the signal on the long arm of chromosome pairs 2 and 5 (close to the telomere and to the centromere, respectively) showed variability among samples. This variability suggests the existence of polymorphisms in the amount of BamHI-800 repetitive DNA in these chromosomal locations. Analysis of more samples from different geographic locations should be performed to check this hypothesis.
The positive signals in E. calamita sit on the short arm of two of the largest chromosome pairs (chromosomes 1 and 3). In both pairs, BamHI-800 is located close to the centromere, with the signal of pair 1 being more intense than that of pair 3 (Figs. 4g and 5b).
In Bufotes viridis BamHI-800 is located on the 3rd chromosomal pair, in pericentromeric position at both sides of the centromere, with the most intense signal on the short arm. In addition, there is a positive signal in the chromosomal pair that carries the NOR (pair 6), located on the long arm and close to the centromere (Figs. 4h and 5c). As opposed to Bufo bufo, in Bufotes viridis the signal is not located near to the NOR, but in the pericentromeric region of the same chromosome arm.
Information about expression of BamHI-800
To check if this satellite DNA is transcribed, we searched for BamHI-800 sequences in published transcriptomes from several Bufonidae species. We did not detect expression of BamHI-800 sequences in assembled transcriptomes23. However, BamHI-800 sequences produce hits in BLASTN searches against Sequence Read Archive (SRA) data from several Bufonidae RNA sources. The number of hits obtained in several conditions (tissues, stage, spots, etc.) is shown in Supplementary Table S8.
Discussion
We have characterized a new family of repetitive DNA (BamHI-800), isolated from Bufo bufo genomic DNA digested with BamHI. It is organized in tandem arrays of monomeric units of ca. 800 bp. Compared to other satellite DNAs in other amphibians, such as pBv PstI in Bufotes viridis19, Dp-sat1 in Discoglossus pictus24, PcP190 in Hyloidea21, or S1 in Rana ssp.12,25,26,27, BamHI-800 consists of a relatively large repetitive unit. Monomers of such length have not been identified when amphibian repetitive elements are characterized from next-generation sequence reads, e.g. in Proceratophrys boiei16.
Southern-Blot hybridization shows positive signals only in Bufonidae. BamHI-800 sequences were neither identified by Southern-Blot nor by specific PCRs in three other anuran families (e.g. Alytidae (D. galganoi), Pipidae (X. tropicalis), and Ranidae (P. perezi)). Fourteen bufonid toad species (Bufo bufo, Bufo spinosus, Ba. brongersmai, Bufotes boulengeri, Bufotes luristanicus, Bufotes surdus, Bufotes viridis, Bufotes latastii, Bufotes balearicus, Bufotes siculus, D. melanostictus, E. calamita, R. marina, and S. arabica) from seven genera exhibited similar banding patterns by Southern-Blot or PCR for BamHI-800. In addition, BLASTN searches in genome assemblies of Bufonidae confirmed BamHI-800 in Bufo bufo and R. marina, and provided new information in Bufo gargarizans. Genomes of other amphibian species showed no BLASTN hits (Supplementary Table S3), pointing to BamHI-800 as a Bufonidae-specific satellite DNA. Further analysis of closer outgroups (Hyloidea) and early diverging bufonid lineages are needed to confirm the distribution of this satellite DNA family.
Repetitive DNAs have been studied by PCR and sequencing, e.g. the S1 satellite of some species of Rana12,26,27, or the PcP190 satellite in Hyloidea21. In our study, the variability of BamHI-800 sequences obtained by traditional digestion-cloning is similar to that obtained by sequenced PCR products. Therefore, PCR does not select particular sequence groups from BamHI-800 sequences, but amplifies a wide range of variants of this repetitive DNA.
The BamHI-800 satellite-DNA family is conserved in the genomes of all studied bufonids. Genetic distances and phylogenetic groups suggest ten BamHI-800 subfamilies distributed in 2 variants. For some species, like Bufo gargarizans, Ba. brongersmai, D. melanosticus, E. calamita, R. marina, and S. arabica, the BamHI-800 V1-variant shows lower intraspecific variation than interspecific divergence, a sign of concerted evolution. This particular mode of evolution is the consequence of a two-level process called molecular drive, consisting of sequence homogenization and fixation28,29. Thus, high sequence homogeneity within a subfamily of satellite DNA monomers is a result of non-independent evolution of monomers, driven by non-reciprocal sequence transfer, such as unequal crossover29, gene conversion30,31,32, rolling circle replication26,33,34, transposition35 and may be others2. While homogenization depends on mechanisms of genomic turnover, fixation results from random chromosome assortment during sexual reproduction through meiosis and chromosome segregation. As a consequence, fixation depends on population factors. The outcome of this process is higher homogeneity of satellite monomers within lineages than between them. Thus, satellite DNA can evolve by a gradual accumulation of mutations, which results in the divergence of satellite sequences of reproductively isolated organismal groups2. Accordingly, it can be phylogenetically informative, for example, in species36, ecotype-specific variants, or phylogeographic clades37.
We did not find significant sequence divergence between different populations of the Bufo bufo group. This is interesting since we have compared BamHI-800 obtained from Bufo bufo sensu lato. The current taxonomy of the Bufo bufo species group distinguishes four species38,39: Bufo bufo (Linnaeus, 1758) from most of Europe (from northern and eastern France into Russia, including toads from Great Britain, Scandinavia, Italy, the Balkans, and the larger part of Turkey), Bufo eichwaldi40 from the Talysh mountains of Azerbaijan and Iran, Bufo spinosus (Daudin, 1803) from North Africa, Iberia and much of France, and Bufo verrucosissimus (Pallas, 1814) from the Caucasus and the north-eastern part of Turkey. According to their geographic distribution41,42, and their 16S sequence, our samples from Spain and Morocco correspond to Bufo spinosus, while samples from NCBI (London) and Bologna to Bufo bufo. However, in our maximum-likelihood analysis, the BamHI-800 sequences were not clustered according to geographical origin/species. Further studies with additional samples are required to uncover species- or population-specific differences.
Contrary to the situation in Bufo bufo species group, BamHI-800 V1-type sequences in R. marina have diagnostic positions that differentiate two Australian populations (Brisbane, Queensland (our sample) and Oombulgurri, Western Australia (Biosample: SAMEA104558286)). This suggests satellite diversification appears not always indicative of the divergence time. Considering that Australian R. marina can be traced back to 102 individuals introduced in 1935, the rapid BamHI-800 diversification might be explained by an initially very low population size followed by enormous population growth. Future analysis of BamHI-800 in R. marina in its native S-American range could provide information on the effect of biological conditions on the evolution of satellite DNAs.
BamHI-800 sequences in Bufotes could not be clearly classified into species-specific subfamilies. In this genus, intraspecific variability is greater than interspecific divergence, and consequently, BamHI-800 sequences appear scattered in the phylogenetic tree. However, two differentiated groups were identified in Bufotes. These groups correspond with the two ancient clades of diploid toads: taxa diversified around the Mediterranean Basin and the Middle East and diploid taxa occurring between Iran, Iraq, Pakistan, and the Indian Himalayas43,44,45.
Barbarophyne brongersmai previously occupied a controversial taxonomic position. It was initially placed in the “Bufotes viridis–E. calamita group” based on morphology46, and then into the “Bufotes viridis group”47. Subsequent research on larval morphology, karyology, osteology, and bioacoustics revealed that Ba. brongersmai is well-diverged from all other Western Palearctic bufonid taxa48,49,50. Later mitochondrial and nuclear DNA-based phylogenies43,51,52,53 distinguished and clearly separated Ba. brongersmai from green toads (Bufotes), which was confirmed by54. According to this phylogenetic position, it is not surprising that the BamHI-800 sequences of Ba. brongersmai appear in an independent group in the phylogenetic tree.
The BamHI-800 sequences found in two other bufonid genera, R. marina and D. melanostictus, are of two types: those clustering in separate subfamilies as V2 variants and those phylogenetically related to the V1-variant of BamHI-800. The ‘library hypothesis’, not mutually exclusive with concerted evolution, assumes the existence of a collection (‘library’) of satellite DNAs shared by a species group. Accordingly, these taxa share a library of distinct conserved satellite DNA monomers4,55,56 or subfamilies, differentially amplified in each taxon. Variability between sequences reduces the homogenization effects of molecular mechanisms of non-reciprocal exchange, and sequence variants persist as a library57,58,59. Any of these variants may be differentially amplified in each taxon. Thus, in different lineages (species or genera), the subsequent replacement or amplification of one sequence variant can take place.
Replacement of a sequence variant by another in different species is a common feature of satellite DNAs2,60. When it occurs, unrelated species-specific dominant repeats reveal the presence of low-copy counterparts of each in others. This has been suggested in insects55,56,57 and plants61,62, and could explain our observations in Bufonidae, in which both hypotheses (concerted evolution and differential amplification of satellite variants) appear applicable.
FISH-localization of BamHI-800 in four species (Bufo bufo, Bufo spinosus, Bufotes viridis, and E. calamita) is not conserved but shows species-specificity in different chromosomes. BamHI-800 is mostly a component of the repetitive DNA in most pericentromeric regions of Bufo bufo species group (mainly in the large chromosomes and in the smallest pair). However, our comparison of FISH and C-banded karyotypes shows this is not the case in the other two species. Some C-positive bands could present BamHI-800, e.g. two pericentromeric C-bands on the short arm of pairs 1 and 3 in E. calamita15, occurring in the same position as BamHI-800. Similarly, in Bufotes viridis, the pericentromeric C-band on the short arm of chromosome 3 and on the long arm of chromosome 6 may probably show BamHI-800 accumulation. The short arms of chromosome 3 in Bufotes viridis also exhibit pBv satellites19, apparently at the same location as BamHI-800. Our analyses show no relation between these two repetitive DNA families.
Of note is the association between the NOR and the accumulation of BamHI-800 in Bufo bufo and Bufo spinosus. Considering that the NOR in Bufo bufo is flanked by constitutive heterochromatin15, the telomeric BamHI-800 FISH-signal of pair 6 may correspond to one of the C-positive bands flanking the NOR. BamHI-800 sequences in Bufo viridis, and E. calamita are not associated with NORs, although this repetitive DNA in Bufotes viridis sits on the NOR-bearing chromosome 663. The NOR regions of Bufo bufo/Bufo spinosus and Bufo viridis are located in subtelomeric and telomeric positions of the long arm of the sixth chromosome pair, while they are located at terminal position of the long arm of the eleventh chromosomal pair in E. calamita15.
BamHI-800 is associated with constitutive heterochromatin of centromeric/pericentromeric regions. Considering its location, a role of these sequences in centromere function/heterochromatin maintenance could be proposed. The existence of hits for this repetitive DNA in adult transcriptomes from several Bufonidae species is of interest. More studies about its expression in more species and earlier developmental stages could add more information to this interesting question regarding a monomer unit with a long size.
Conclusions
This work provides new data on evolutionary dynamics of satellite DNA in amphibians. This is the first study to reveal the occurrence of a specific satellite DNA family (BamHI-800) and its evolution in bufonid toads. Comparative analysis of BamHI-800 in seven bufonid toad genera shows that the structure, organization, and chromosomal distribution of this satellite DNAs is a relevant feature to distinguish the genomes of related species. The consequences of concerted evolution on intra-species homogenization depend on the species analysed. Most species clearly show specific differences (in R. marina even differentiated between populations), while in others the homogenization process is still ongoing. Moreover, the existence of variants in some species supports the ‘library hypothesis’. The results indicate that this repetitive DNA has a high degree of evolutionary conservation in one of the largest anuran families.
Methods
Samples
Animals were euthanized by immersion in buffered 2% Tricaine methanesulfonate (MS 222; Sigma-Aldrich, Germany) and then decapitated and dissected. Samples from 14 amphibian species belonging to seven genera of Bufonidae have been analysed. Three other anurans (Discoglossus galganoi, Xenopus tropicalis, and Pelophylax perezi) were used as outgroups. The examined species, the number of individual, their origin, chromosome number and the number of sequences considered are listed in Table 1. Individuals from Bufo bufo species group analysed in this work were assigned to Bufo bufo or Bufo spinosus species according to the 16S sequence64.
This study was approved by the relevant Institutional Animal Care and Use Committee (Bioethics Committee of the University of Jaén, 2009). Samples were collected in accordance with regulations for the protection of terrestrial wild animals. Capture permits for Bufo bufo, Bufo spinosus E. calamita, D. galganoi, and P. perezi were provided by the Junta de Andalucía, Dirección General de Gestión del Medio Natural (2010, 2012). Sampling green toad clutches in Crete was done under permit 115790/229. Care and sacrifice of animals used in this research was conducted in accordance with policies on experimental animals provided by Spanish and EU regulations. When applicable, this study is reported according to the ARRIVE guidelines.
DNA extraction, repeat DNA isolation and cloning
Genomic DNA was extracted from skin or liver following the standard phenol–chloroform protocol65. Repetitive sequences from Bufo bufo and E. calamita were revealed by digestion of genomic DNA with several restriction endonucleases (EcoRI, HindIII, PstI, BamHI, AluI, SalI, KpnI y SmaI) according to the manufacturers’ protocols and subsequent agarose gel electrophoresis. After BamHI digestion, the intense bands of about 800 bp and 1600 bp were eluted (GeneJET Gel Extraction Kit, Thermo Fisher Scientific (Waltham, Massachusetts)) and cloned into BamHI-digested pUC19 vector (Fermentas (Vilnius, Lithuania)). Competent JM109 bacteria were transformed by thermal shock. Recombinant clones were denaturalized, dot-blotted onto nylon membranes and hybridized using the BamHI-800 digoxigenin-labelled band as a probe. Positive clones were sequenced using ABI BigDye Terminator kit (Applied Biosystems (Massachusetts, USA)) and T7 or SP6 universal primers. Nucleotide sequences were obtained in an ABI PRISM 3730xl capillary DNA analyser at the DNA sequencing Service from the Centro Nacional de Investigaciones Oncológicas (CNIO (Madrid, Spain)).
Design of specific primers and PCR cloning
The sequences obtained from cloned BamHI-800 bands from the species Bufo bufo and E. calamita were used to design specific primers to PCR-amplify the repetitive sequences in other amphibian species (Supplementary Fig. S1). BamHI-800 sequences were PCR-amplified in other amphibian species in volumes of 50 μl, using 100 ng of genomic DNA, 0.7 U of Taq DNA polymerase (Bioline GmbH, Luckenwalde, Germany), 0.4 μM of each primer, 100 μM of each dNTP and a final MgCl2 concentration of 2 mM in 1× PCR buffer. The thermal conditions used were as follows: 1 cycle of 95 °C for 4 min; 33 cycles of 92 °C for 30 s, 52 °C for 45 s, 72 °C for 45 s; and a final extension step of 5 min at 72 °C.
The PCR products from several species were separated by agarose electrophoresis purified with the GeneJET Gel Extraction Kit (Thermo Fisher Scientific (Waltham, Massachusetts)) and cloned into pGEM-T Easy Vector (Promega (Wisconsin, USA)) as described by the supplier. Positive recombinant colonies were selected and sequenced as indicated. All sequences used in this work were deposited on GenBank under accession numbers ON491183–ON491407.
Southern-blot and probe hybridization
Samples of genomic DNA from different species were digested with BamHI. The resulting fragments were separated by electrophoresis on a 1% agarose gel (5 V/cm) and blotted onto Hybond N + nylon membranes (GE Healthcare (Chalfont St. Giles, UK)) as previously described65,66. Pre-hybridization was done at 68 °C for 3 h in absence of formamide; hybridization was performed overnight at 60–68 °C, using the BamHI-800 band, digoxigenin-labelled by random primers (the optimal hybridization temperatures in each case were obtained experimentally). Additionally, species-specific PCR-labelled probes with digoxigenin-11-dUTP (Roche, Mannheim, Germany) were obtained from positive clones for BamHI-800 sequences, using BamHI-800-F1/R1 as primers. The post-hybridization washes were done in agitation. The wash I (2 × 15 min each) was performed at room temperature in 2 × SSC/0.1% SDS, while the wash II (2 × 30 min each) was done at the hybridization temperature in 0.5 × SSC/0.1% SDS. Hybridization signals were detected with an anti-digoxigenin-11-dUTP antibody conjugated with alkaline phosphatase (Roche, Mannheim, Germany) and revealed using NBT-BCIP as substrate (Roche, Mannheim, Germany) according to the supplier’s recommendations.
Sequence analysis
Multiple sequence alignments were performed with MUSCLE67 and edited manually in MEGA68. Sequence similarity searches were implemented at NCBI (http://Blatst.ncbi.nlm.nih.gov/Blatst.cgi), using blastn suite (megablast and discontiguous megablast) with default parameters69. Similarities with known repetitive elements were queried in Repbase70 and NCBI GenBank.
BamHI-800 sequences were searched in NCBI using BLASTN in standard databases (nr/nt) or in amphibian genome databases (Supplementary Table S3). The BLAST algorithm was optimized for more dissimilar sequences (discontiguous megablast), removing filtering and mask from algorithm parameters. Sequence from clone Bbuf.BH2_B-86 was used as entry query sequence in all cases except for R. marina genome, where sequence from clone Rmar_PCR1.1-5 was used. The hits obtained after BLASTN searches in Bufo bufo, Bufo gargarizans and R. marina genome databases were retrieved and complete sequences used in this analysis.
Dot Plot analysis was performed with YASS71 (https://bioinfo.lifl.fr/yass/yass.php), and DNA secondary structure prediction was modelled with RNA structure72 (V 6.4) with command line: Fold Predict1.fasta Fold.ct -loop 30 -maximum 20 -percent 10 -temperature 310.15 -window 3.
Evolutionary analyses were conducted in MEGA X68. The nucleotide substitution model with the lowest BIC (Bayesian Information Criterion) score was chosen (T92 + G): evolutionary divergence between sequences was estimated using the Tamura 3-parameter model73, and the rate of variation among sites was modelled with a gamma distribution. All ambiguous positions were removed for each sequence pair (pairwise deletion option). Standard error estimates were obtained by a bootstrap procedure (1000 replicates). Maximum likelihood (ML) analysis using the Tamura 3-model of sequence evolution73 was employed to generate a sequence tree. In our analysis, all positions with less than 95% site coverage were eliminated, and ambiguous bases were allowed at any position (partial deletion option). No big differences were observed when all positions with gaps are retained (all positions) or removed (complete deletion). The initial tree(s) for the heuristic search were obtained automatically by applying Neighbour-Joining and BioNJ algorithms to a matrix of pairwise distances estimated using the Maximum Composite Likelihood (MCL) approach, and then selecting the topology with the superior log likelihood. The significance of the phylogenetic lineages was assessed by bootstrap support values, calculated for 1000 replicates74.
Information about expression of BamHI-800 sequences was obtained using BLASTN optimized for more dissimilar sequences (discontiguous megablast) on NCBI Sequence Read Archive (SRA) data from Bufonidae RNA sequences. When available, the query sequence was from the same species/genus. In any other case, the sequence used was from clone Bbuf.BH2_B-86. Default algorithm parameters were used except for the lack of filtering and masking.
Chromosome preparation and fluorescent in-situ hybridization (FISH)
Chromosome preparations were obtained from bone marrow cells of adult individuals or from cell culture from tadpoles according to the protocols described in75.
FISH was performed as previously described in 76. The probe used was a BamHI-800 positive clone from the same species as the chromosome preparations (Bbu-BamHI-800-86 in Bufo bufo, Bca-BamHI-800-9 and Bca-BamHI-800-10 in E. calamita and Bvi-BamHI-800-5 in Bufotes viridis), PCR-labelled with biotin-16-dUTP (Roche (Mannheim, Germany)). When indicated, three rounds of immunological amplification with avidin-FITC/anti-avidin–biotin were used. For each species and sample, more than 20 hybridized metaphase plates were photographed and analysed. The images were taken with FITC and DAPI filters in an Olympus BX-51 fluorescence microscope. Images were processed with Adobe Photoshop. Processing was applied equally across the entire image.
Data availability
All BamHI-800 sequences obtained in this study were submitted to Genbank (Accession numbers: ON491183–ON491407). All data generated and analysed during this study are included in this published article and its Supplementary Information files.
References
John, B. & Gabor Miklos, G. L. Functional aspects of satellite DNA and heterochromatin. Int. Rev. Cytol. 58, 1–114 (1979).
Plohl, M., Meštrović, N. & Mravinac, B. Satellite DNA evolution. Genome Dyn. 7, 126–152 (2012).
Dover, G. Molecular drive: A cohesive mode of species evolution. Nature 299, 111–117 (1982).
Fry, K. & Salser, W. Nucleotide sequences of HS-α satellite DNA from kangaroo rat Dipodomys ordii and characterization of similar sequences in other rodents. Cell 12, 1069–1084 (1977).
Garrido-Ramos, M. A. Satellite DNA: An evolving topic. Genes 8, 230 (2017).
Pezer, Ž, Brajković, J., Feliciello, I. & Ugarkovć, D. Satellite DNA-mediated effects on genome regulation. Genome Dyn. 7, 153–169 (2012).
Biscotti, M. A., Canapa, A., Forconi, M., Olmo, E. & Barucca, M. Transcription of tandemly repetitive DNA: Functional roles. Chromosom. Res. 23, 463–477 (2015).
Plohl, M., Meštrović, N. & Mravinac, B. Centromere identity from the DNA point of view. Chromosoma 123, 313–325 (2014).
Louzada, S. et al. Decoding the role of satellite DNA in genome architecture and plasticity—An evolutionary and clinical affair. Genes 11, 72 (2020).
Jagannathan, M., Cummings, R. & Yamashita, Y. M. A conserved function for pericentromeric satellite DNA. Elife 7, 1–19 (2018).
Shatskikh, A. S., Kotov, A. A., Adashev, V. E., Bazylev, S. S. & Olenina, L. V. Functional significance of satellite DNAs: Insights from Drosophila. Front. Cell Dev. Biol. 8, 1–19 (2020).
Picariello, O., Feliciello, I., Bellinello, R. & Chinali, G. S1 satellite DNA as a taxonomic marker in brown frogs: Molecular evidence that Rana graeca graeca and Rana graeca italica are different species. Genome 45, 63–70 (2002).
Ferree, P. M. & Prasad, S. How can satellite DNA divergence cause reproductive isolation? Let us count the chromosomal ways. Genet. Res. Int. 2012, 1–11 (2012).
Liedtke, H. C., Gower, D. J., Wilkinson, M. & Gomez-Mestre, I. Macroevolutionary shift in the size of amphibian genomes and the role of life history and climate. Nat. Ecol. Evol. 2, 1792–1799 (2018).
Schmid, M. Chromosome banding in amphibia I. Constitutive heterochromatin and nucleolus organizer regions in bufo and hyla. Chromosoma 66, 361–388 (1978).
Da Silva, M. J. et al. Great abundance of satellite DNA Proceratophrys (Anura, Odontophrynidae) revealed by genome sequencing. Cytogenet. Genome Res. 160, 141–147 (2020).
AmphibiaWeb. University of California, Berkeley, CA, USA. https://amphibiaweb.org (Accessed 14 June 2022)
Frost, D. R. Amphibian Species of the World: An Online Reference. Version 6.1. American Museum of Natural History, New York, USA. (2021). https://amphibiansoftheworld.amnh.org/index.php. (Accessed 14 June 2022)
Odierna, G., Aprea, G., Capriglione, T., Castellano, S. & Balletto, E. Evidence for chromosome and PstI satellite DNA family evolutionary stasis in the Bufo viridis group (Amphibia, Anura). Chromosom. Res. 12, 671–681 (2004).
Pontecorvo, G., De Felice, B. & Carfagna, M. A novel repeated sequence DNA originated from a Tc1-like transposon in water green frog Rana esculenta. Gene 261, 205–210 (2000).
Vittorazzi, S., Lourenço, L. B. & Recco-Pimentel, S. M. Long-time evolution and highly dynamic satellite DNA in leptodactylid and hylodid frogs. BMC Genet. 15, 111 (2014).
Nanda, I., Fugate, M., Steinlein, C. & Schmid, M. Distribution of (TTAGGG)n telomeric sequences in karyotypes of the Xenopus species complex. Cytogenet. Genome Res. 122, 396–400 (2008).
Gerchen, J. F. et al. A single transcriptome of a green toad (Bufo viridis) yields candidate genes for sex determination and -differentiation and non-anonymous population genetic markers. PLoS ONE 11, e0156419 (2016).
Amor, N., Odierna, G., Chinali, G., Said, K. & Picariello, O. Unusual chromosomal distribution of a major satellite DNA from Discoglossus pictus (Amphibia, Anura). Cytogenet. Genome Res. 127, 33–42 (2009).
Feliciello, I., Picariello, O. & Chinali, G. Intra-specific variability and unusual organization of the repetitive units in a satellite DNA from Rana dalmatina: Molecular evidence of a new mechanism of DNA repair acting on satellite DNA. Gene 383, 81–92 (2006).
Feliciello, I., Picariello, O. & Chinali, G. The first characterisation of the overall variability of repetitive units in a species reveals unexpected features of satellite DNA. Gene 349, 153–164 (2005).
Picariello, O., Feliciello, I. & Chinali, G. S1 satellite DNA repetitive units display identical structure and overall variability in all Anatolian brown frog taxa. Genetica 144, 47–57 (2016).
Dover, G. Molecular drive. Trends Genet. 18, 587–589 (2002).
Dover, G. A. & Tautz, D. Conservation and divergence in multigene families: Alternatives to selection and drift. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 312, 275–289 (1986).
Smith, G. P. Evolution of repeated DNA sequences by unequal crossover. Science 191, 528–535 (1976).
Mravinac, B., Ugarković, D., Franjević, D. & Plohl, M. Long inversely oriented subunits form a complex monomer of Tribolium brevicornis satellite DNA. J. Mol. Evol. 60, 513–525 (2005).
Glinka, S., De Lorenzo, D. & Stephan, W. Evidence of gene conversion associated with a selective sweep in Drosophila melanogaster. Mol. Biol. Evol. 23, 1869–1878 (2006).
Walsh, J. B. Persistence of tandem arrays: Implications for satellite and simple-sequence DNAs. Genetics 115, 553–567 (1987).
Cohen, S. & Segal, D. Extrachromosomal circular DNA in eukaryotes: Possible involvement in the plasticity of tandem repeats. Cytogenet. Genome Res. 124, 327–338 (2009).
Okumura, K., Kiyama, R. & Oishi, M. Sequence analyses of extrachromosomal Sau3A and related family DNA: Analysis of recombination in the excision event. Nucleic Acids Res. 15, 7477–7489 (1987).
Garrido-Ramos, M. A. et al. Evolution of centromeric satellite DNA and its use in phylogenetic studies of the Sparidae family (Pisces, Perciformes). Mol. Phylogenet. Evol. 12, 200–204 (1999).
Robles, F. et al. Evolution of ancient satellite DNAs in sturgeon genomes. Gene 338, 133–142 (2004).
Arntzen, J. W., Recuero, E., Canestrelli, D. & Martínez-Solano, Í. How complex is the Bufo bufo species group?. Mol. Phylogenet. Evol. 69, 1203–1208 (2013).
Arntzen, J. W., Wilkinson, J. W., Butôt, R. & Martínez-Solano, Í. A new vertebrate species native to the British Isles: Bufo spinosus Daudin, 1803 in Jersey. Herpetol. J. 24, 209–216 (2014).
Litvinchuk, S. N., Borkin, L. J., Skorinov, D. V. & Rosanov, J. M. A new species of common toads from the Talysh Mountains of south-eastern Caucasus: Genome size, allozyme, and morphological evidences. Russ. J. Herpetol. 15, 19–43 (2008).
Arntzen, J. W. et al. Morphological and genetic differentiation of Bufo toads: Two cryptic species in Western Europe (Anura, Bufonidae). Contrib. Zool. 82, 147–169 (2013).
Recuero, E. et al. Multilocus species tree analyses resolve the radiation of the widespread Bufo bufo species group (Anura, Bufonidae). Mol. Phylogenet. Evol. 62, 71–86 (2012).
Stöck, M. et al. Evolution of mitochondrial relationships and biogeography of Palearctic green toads (Bufo viridis subgroup) with insights in their genomic plasticity. Mol. Phylogenet. Evol. 41, 663–689 (2006).
Dufresnes, C. et al. Fifteen shades of green: The evolution of Bufotes toads revisited. Mol. Phylogenet. Evol. 141, 106615 (2019).
Betto-Colliard, C., Hofmann, S., Sermier, R., Perrin, N. & Stöck, M. Profound genetic divergence and asymmetric parental genome contributions as hallmarks of hybrid speciation in polyploid toads. Proc. R. Soc. B Biol. Sci. 285, 20172667 (2018).
Hoogmoed, M. S. On a new species of toad from southern Morocco. Zool. Meded. 47, 49–64 (1972).
Amphibian species of the world. A taxonomic and geographical reference. (Allen Press and Ass. of Systematics Collections, 1985).
Grillitsch, B., Grillitsch, H. & Splechtna, H. The tadpole of Bufo brongersmai Hoogmoed 1972. Amphib. Reptil. 10, 215–229 (1989).
Herrero, P., López-Jurado, L. F., Arano, B. & García-Paris, M. Analyses and nuclear DNA content of Bufo brongersmai Hoogmoed. J. Herpetol. 27, 463–465 (1993).
Delfino, M., Doglio, S., Roček, Z., Seglie, D. & Kabiri, L. Osteological Peculiarities of Bufo brongersmai (Anura: Bufonidae) and their possible relation to life in an arid environment. Zool. Stud. 48, 108–119 (2009).
Van Bocxlaer, I., Biju, S., Loader, S. P. & Bossuyt, F. Toad radiation reveals into-India dispersal as a source of endemism in the Western Ghats-Sri Lanka biodiversity hotspot. BMC Evol. Biol. 9, 1–10 (2009).
Pyron, R. A. & Wiens, J. J. A large-scale phylogeny of Amphibia including over 2800 species, and a revised classification of extant frogs, salamanders, and caecilians. Mol. Phylogenet. Evol. 61, 543–583 (2011).
Garcia-Porta, J. et al. Molecular phylogenetics and historical biogeography of the west-palearctic common toads (Bufo bufo species complex). Mol. Phylogenet. Evol. 63, 113–130 (2012).
Beukema, W. et al. Review of the systematics, distribution, biogeography and natural history of Moroccan amphibians. Zootaxa 3661, 1–60 (2013).
Cesari, M., Luchetti, A., Passamonti, M., Scali, V. & Mantovani, B. Polymerase chain reaction amplification of the Bag320 satellite family reveals the ancestral library and past gene conversion events in Bacillus rossius (Insecta Phasmatodea). Gene 312, 289–295 (2003).
Mestrović, N., Plohl, M., Mravinac, B. & Ugarković, D. Evolution of satellite DNAs from the genus Palorus–experimental evidence for the ‘library’ hypothesis. Mol. Biol. Evol. 15, 1062–1068 (1998).
Mravinac, B., Plohl, M., Mestrović, N. & Ugarković, D. Sequence of PRAT satellite DNA ‘frozen’ in some coleopteran species. J. Mol. Evol. 54, 774–783 (2002).
Ugakovic, D. & Plohl, M. Variation in satellite DNA profiles-causes and effects. EMBO J. 21, 5955–5959 (2002).
Meštrović, N., Castagnone-Sereno, P. & Plohl, M. Interplay of selective pressure and stochastic events directs evolution of the MEL172 satellite DNA library in root-knot nematodes. Mol. Biol. Evol. 23, 2316–2325 (2006).
Plohl, M. Those mysterious sequences of satellite DNAs. Period. Biol. 112, 403–410 (2010).
Quesada Del Bosque, M. E. et al. A satellite DNA evolutionary analysis in the North American endemic dioecious plant Rumex hastatulus (Polygonaceae). Genome 54, 253–260 (2011).
Navajas-Pérez, R. & Paterson, A. H. Patterns of tandem repetition in plant whole genome assemblies. Mol. Genet. Genomics 281, 579–590 (2009).
Stöck, M., Steinlein, C., Lamatsch, D. K., Schartl, M. & Schmid, M. Multiple origins of tetraploid taxa in the Eurasian Bufo viridis subgroup. Genetica 124, 255–272 (2005).
Stöck, M. et al. Post-Messinian evolutionary relationships across the Sicilian channel: Mitochondrial and nuclear markers link a new green toad from Sicily to African relatives. BMC Evol. Biol. 8, 1–19 (2008).
Sambrook, J. & Russel, D. W. Molecular Cloning: A Laboratory Manual Vol. 3 (Cold Spring Harbor Laboratory Press, 2001).
Bullejos, M. et al. Multiple, polymorphic copies of SRY in both males and females of the vole Microtus cabrerae. Cytogenet. Cell Genet. 79, 167–171 (1997).
Edgar, R. C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Kohany, O., Gentles, A. J., Hankus, L. & Jurka, J. Annotation, submission and screening of repetitive elements in Repbase: RepbaseSubmitter and Censor. BMC Bioinform. 7, 1–7 (2006).
Noé, L. & Kucherov, G. YASS: Enhancing the sensitivity of DNA similarity search. Nucleic Acids Res. 33, 540–543 (2005).
Bellaousov, S., Reuter, J. S., Seetin, M. G. & Mathews, D. H. RNAstructure: Web servers for RNA secondary structure prediction and analysis. Nucleic Acids Res. 41, W471–W474 (2013).
Tamura, K. Estimation of the number of nucleotide substitutions when there are strong transition-transversion and G+C-content biases. Mol. Biol. Evol. 9, 678–687 (1992).
Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 39, 783–791 (1985).
Roco, Á. S., Liehr, T., Ruiz-García, A., Guzmán, K. & Bullejos, M. Comparative distribution of repetitive sequences in the karyotypes of Xenopus tropicalis and Xenopus laevis (Anura, pipidae). Genes 12, 617 (2021).
Fernández, R. et al. Molecular and cytogenetic characterization of highly repeated DNA sequences in the vole Microtus cabrerae. Heredity 87, 637–646 (2001).
Acknowledgements
This research was funded by The Spanish Ministry of Science and Innovation, grant number CGL2009-08894. The APC was funded by the University of Jaén throughout the program Plan de Apoyo a la Investigación 2019–2020 (Acción 4) to M.B.
The authors want to thank Rosanna Falconi and Francesco Zaccanti for DNA samples from Bufo bufo (Bologna, Italy), Dagmar Wilhelm for genomic DNA from Rhinella marina and the Functional Ecology Group (Centro de Investigação em Biodiversidade e Recursos Genéticos, CIBIO) for tissue samples from B. bufo and Bufotes boulengeri. We thank Petros Lymberakis (Natural History Museum of Crete, University of Crete) for help with the permit 115790/229 to sample green toad clutches. The authors also thank Pedro Lorite for helpful comments on the manuscript.
Author information
Authors and Affiliations
Contributions
Conceptualization, M.B.; methodology, K.G., Á.S.R; formal analysis, K.G., A.S.R. and M.B.; investigation, K.B., Á.S.R, A.R.-G.; resources, M.S., E.G-M.; writing—original draft preparation, M.B. and K.G.; writing—review and editing, M.B., K.G., Á.S.R., and M.S.; supervision, M.B.; project administration, M.B.; funding acquisition, M.B. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Guzmán, K., Roco, Á.S., Stöck, M. et al. Identification and characterization of a new family of long satellite DNA, specific of true toads (Anura, Amphibia, Bufonidae). Sci Rep 12, 13960 (2022). https://doi.org/10.1038/s41598-022-18051-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-18051-9
This article is cited by
-
A candidate sex determination locus in amphibians which evolved by structural variation between X- and Y-chromosomes
Nature Communications (2024)
-
Interspecific cytogenomic comparison reveals a potential chromosomal centromeric marker in Proceratophrys frog species
Chromosoma (2024)
-
Consequences of polyploidy and divergence as revealed by cytogenetic mapping of tandem repeats in African clawed frogs (Xenopus, Pipidae)
European Journal of Wildlife Research (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.