Abstract
Hepatocellular carcinoma (HCC) is a highly heterogeneous cancer, and resistant to both conventional and targeted chemotherapy. Recently, nonsteroidal anti-inflammatory drugs (NSAIDs) have been shown to decrease the incidence and mortality of different types of cancers. Here, we investigated the cellular bioactivities of a series of triazolothiadiazine derivatives on HCC, which have been previously reported as potent analgesic/anti-inflammatory compounds. From the initially tested 32 triazolothiadiazine NSAID derivatives, 3 compounds were selected based on their IC50 values for further molecular assays on 9 different HCC cell lines. 7b, which was the most potent compound, induced G2/M phase cell cycle arrest and apoptosis in HCC cells. Cell death was due to oxidative stress-induced JNK protein activation, which involved the dynamic involvement of ASK1, MKK7, and c-Jun proteins. Moreover, 7b treated nude mice had a significantly decreased tumor volume and prolonged disease-free survival. 7b also inhibited the migration of HCC cells and enrichment of liver cancer stem cells (LCSCs) alone or in combination with sorafenib. With its ability to act on proliferation, stemness and the migration of HCC cells, 7b can be considered for the therapeutics of HCC, which has an increased incidence rate of ~ 3% annually.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma (HCC) is one of the most common cancer types and second in terms of cancer-related mortality worldwide1. HCC has a very heterogeneous structure and develops over many years as a multi-step process2. Patients with advanced stage HCC have very limited and mostly palliative treatment options due to the chemo-resistant nature of the disease3,4. Hepatic injury initiated by any of the etiologies such as chronic hepatitis, alcohol abuse or aflatoxin-B1-intoxication, and recently obesity, results in continuous destructive-regenerative cycles and causes cirrhosis, which further induces carcinogenesis leading to HCC5. Recently, sorafenib, lenvatinib, regorafenib, atezolizumab plus bevacizumab, cabozantinib and ramucirumab are the FDA-approved drugs for the systemic therapy of HCC6. Although sorafenib and Lenvatinib are regarded as the best treatment options for advanced HCC patients, capable of inducing apoptosis and inhibiting angiogenesis and proliferation of tumor cells, they can extend patient survival for about 3 months7,8. Combinatorial approaches involving the use of chemotherapeutic agents together with sorafenib in HCC have been reported to show limited usage due to the high toxicity and unconvincing efficacy in prolonging the survival of HCC patients9,10. In addition, recent studies report that acquired drug resistance to sorafenib is very common. Multiple factors and mechanisms are identified to be involved in the development of sorafenib resistance followed by the progression of the disease. Tumor microenvironment, EGFR activation, compensatory pathways such as PI3K/AKT and JAK/STAT, and the presence of cancer stem cells (CSCs) are only some of these mechanisms11. Therefore, it is essential to discover reliable drug candidates that could provide novel therapeutic options for advanced HCC patients.
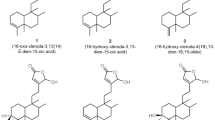
Nonsteroidal anti-inflammatory drugs (NSAIDs) have been used to treat acute and chronic conditions with pain and inflammation for many years. Recently, clinical studies have emphasized the role of NSAIDs in cancer treatment, such that NSAIDs are effective in decreasing the incidence and mortality of many cancer types12,13,14,15. 1,2,4-Triazole is an important scaffold in the field of medicinal chemistry. Currently, a group of compounds carrying the 1,2,4-triazole ring, named anastrozole (1), letrozole (2) vorozole (3), are explicitly used in the treatment of estrogen receptor-positive breast cancer as nonsteroidal aromatase inhibitors (Fig. 1)16,17. For this reason, the five-membered 1,2,4-triazole ring system is a suitable heterocyclic core structure for designing new anticancer compounds. During the last two decades, it has been determined that triazolopyridazine, triazolotriazine, triazolothiadiazine, and triazolothiadiazole derivatives (condensed 1,2,4-triazole derivatives) have cytotoxic activities in many cancer cells by inducing various molecular mechanisms18,19,20,21,22,23,24,25.
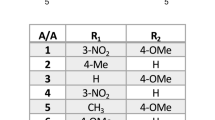
Since more than 90% of the liver cancer cases are associated with chronic inflammation26, NSAIDs could be effective against HCC. Indeed, in one of our previous studies, the anticancer effects of triazolothiadiazines on epithelial cancers, especially liver cancer, were demonstrated, where compounds could induce apoptotic cell death due to oxidative stress, acting on Akt protein13. Furthermore, a compound with triazolothiadiazine scaffold, displaying antiproliferative bioactivities against human hepatoma cell line (HepG2), was reported recently27. In light of this information and as a part of our ongoing interest in liver cancer therapeutics28, we investigated the anticancer effects of our newly synthesized compounds bearing triazolethiones (1–8) or triazolothiadiazine (1a-8c) cores, which we reported as effective analgesic/anti-inflammatory compounds previously (Fig. 2)24,25.
This study not only evaluates the in vitro and in vivo anti-tumor activity of the compounds against liver cancer, but also characterizes the form of cell death induced by the compounds and identifies the underlying molecular mechanism of action with the most potent compound.
Materials and methods
Synthesis of NSAID triazolothiadiazine derivatives
Synthesis of 4-amino-3-substituted-1,2,4-triazole-5-thiones (Compounds 1–8), was carried out by melting some aralkyl carboxylic acid derivatives with thiocarbohydrazide according to the procedure described previously25,29. Synthesis of 3,6-disubstituted-7H-1,2,4-triazolo[3,4-b]-1,3,4-thiadiazines (1a-8c) was carried out by reacting relevant 4-amino-3-substituted-1,2,4-triazole-5-thiones (1–8) with appropriate phenacyl halides in anhydrous ethanol under reflux, 3,6-disubstituted-7H-1,2,4-triazolo[3,4-b]-1,3,4-thiadiazines (1a-8c)25,29. Physical properties and spectral data characterizing the structure of the synthesized compounds were reported previously24,25.
Drugs and Chemicals
JNK inhibitor V (cat. no. 420129) and DMSO (cat. no. D2650) were provided from Calbiochem;Sigma-Aldrich; Merck KGaA (Burlington, MA, United States), sorafenib (cat. no. S7397), and DAPT (cat. no. S2215) were provided from Selleck chemicals (Houston, TX, USA), Taxol was provided from Bristol Myers Squibb (New York City, New York, USA). Camptothecin (cat. no. C9911) was purchased from Sigma-Aldrich; Merck KGaA. N-acetyl-cysteine (NAC) was provided from Birsen Tozkoparan (Department of Pharmaceutical Chemistry, Hacettepe University, Ankara, Turkey).
Cell culture
Nine hepatocellular carcinoma, and one colon carcinoma cell lines were obtained from the following sources: Huh7 (JCRB0403), HepG2 (ATCC HB-8065), Hep3B (ATCC HB-8064), PLC (ATCC CRL-8024), SK-Hep1 (ATCC HTB52) Mahlavu30, FOCUS31, SNU182 (ATCC CRL-2235), SNU387 (ATCC CRL-2237), SNU475 (ATCC CRL-2236) and, HCT116 (ATCC CCL-247). Cells were grown in DMEM (cat. no. 31885), (Dulbecco's Modified Eagle Medium) or RPMI (Roswell Park Memorial Institute)-1640 growth medium (cat. no. 31870) supplemented with 10% fetal bovine serum (FBS) (cat. no. 10270), 1% non-essential amino acids (cat. no. 11140050), 1% L-glutamine (cat. no. 25030), 1% penicillin and streptomycin (cat. no. 15140–122) from Invitrogen; Thermo Fisher Scientific, Inc. (Waltham, MA, USA) at 37 °C under 5% CO2. All cell lines used in this study are STR (short tandem repeat) authenticated. Cells are regularly tested for mycoplasma contamination using a mycoplasma detection kit (cat. no. LT07-318) (MycoAlert™, Lonza, Basel, Switzerland). The passaging of the cells did not exceed 8–10 passages (2 times a week) throughout the experiments.
Sulforhodamine B (SRB) assay
Cells were inoculated into 96-well plates (1000–5000 cell/well in 150 μL/well) for 24 h and treated with the increasing concentrations of compounds (40–0.01 µM) for 24, 48, or 72 h. Cells were washed with 1xPBS (CaCl2-, MgCl2-free) (Invitrogen, cat. no. 14190144) and fixed with ice-cold 10% trichloroacetic acid (cat. no. 27242, Sigma Aldrich; Merck KGaA,). Microplates were kept at + 4 °C, in the dark for 1 h, then washed with ddH2O. Air-dried plates were stained with SRB (cat. no. 2S1403 Sigma Aldrich; Merck KGaA) (w/v 0.4%SRB in 1% acetic acid solution) for 10 min in the dark. The plates were rinsed with 1% acetic acid solution to wash away the unbound SRB. The protein-bound SRB was then solubilized in a 10 mM Tris-base solution. The absorbance values were obtained at 515 nm using a ELx800 plate reader (BioTek Instruments, Inc., Winooski, VT, USA).
Real-time cell growth analysis (RT-CES)
HCC cells were plated into E-96 plates (1000–5000 cells/well). The cell index (CI) values were recorded to obtain the proliferation curve of the cells through real-time cell electronic sensing (RT-CES) (ACEA Biosciences; Agilent Technologies, Santa Clara, CA, USA). On the next day, the cells were treated with the compounds. The CI values, representing the cell impedance, were recorded every 10 min for the first 4 h to monitor the fast drug response and then every 30 min to monitor the long-term drug response. Cell growth curves were generated using the time-zero normalized CI values.
Annexin V/PI staining analysis of apoptosis
Cells were plated into 6-well plates (30,000–60,000 cells/well). The next day cells were treated in increasing concentrations of 7b or DMSO. For experiments with NAC and JNKinhV, cells were treated with either NAC (5 mM), JNKinhV (1 µM) only, or pre-treated with NAC for 1 h followed by 7b treatment, or with JNKinhV and 7b (simultaneously) for 24 h. Cells were harvested in falcon tubes by trypsinization and centrifuged at 1500 rpm for 5 min. Cell media was removed, and pellets were washed twice in ice-cold 1xPBS and centrifuged again. Then, cells were resuspended in Annexin-V-FLUOS labeling solution which consisted of 2 µl Annexin-V-FLUOS labeling reagent and 2 µL PI solution in 100 µl incubation buffer per sample and incubated for 15 min at room temperature in dark. 100–200 µl of incubation buffer was added on top of each sample and analyzed using Novocyte (ACEA Biosciences; Agilent Technologies) flow cytometry. The percentage of apoptotic cells was determined using NovoExpress software (ACEA Biosciences; Agilent Technologies).
Cell cycle analysis
HCC cells were inoculated in 100 mm culture dishes. After 24 h, cells were treated with compound 7b and its DMSO control for 24 h. For experiments with NAC, cells were treated with either NAC (5 mM) or pre-treated with NAC for 1 h followed by 7b treatment for 24 h. Cell pellets were harvested and re-suspended in 1 ml, ice-cold 1xPBS, and fixed by adding 2.5 mL, 70% ice-cold ethanol. Next, the cell pellets were re-suspended in Propidium iodide (PI) solution (50 μg/mL PI (cat. no. P4854, Sigma Aldrich; Merck KGaA), 0.1 μg/mL RNaseA (cat. no. EN0531Fermentas;Thermo Fisher Scientific Inc.), 0.05% Triton-X-100, 1xPBS) and incubated for 40 min at 37 °C at dark. Cell cycle analysis was performed using FACSCalibur and CellQuest Software (BD Biosciences, Franklin Lakes, NJ, USA) or Novocyte Flow Cytometry (ACEA Biosciences; Agilent Technologies).
Detection of ER-stress
Total RNA was isolated from cells via the Nucleospin RNA II kit (cat. no. 740955, Macherey–Nagel, Düren, Germany) according to the manufacturer’s protocol. First strand cDNA synthesis was performed using the RevertAid First Strand cDNA synthesis kit (cat. no. K1622, Thermo Scientific Inc.). Semi-quantitative reverse transcriptase PCR (RT-PCR) assay was performed using XBP1 specific primers. The primer sequences; GAPDH forward primer: GGCTGAGAACGGGAAGCTTGTCAT, GAPDH reverse primer: CAGCCTTCTCCATGGTGGTGAAGA, XBP1 forward primer: TTACGAGAGAAAACTCATGGCC, XBP1 reverse primer: GGGTCCAAGTTGTCCAGAATGC.
Reactive Oxygen Species (ROS) detection
Huh7 and Mahlavu cells were seeded into 10 cm culture dishes (100,000–300,000 cells/dish). Next day, cells were treated with compound 7b, DMSO or Selenium (Se) deficient serum-free media for 8, 12 or 24 h. For experiments with NAC, cells were treated with compound 7b only or pretreated with NAC for 1 h following 7b treatment for 12 h. Flow cytometric analysis of ROS accumulation in cells at 12 h was done with the Oxidative Stress kit (cat. no. MCH100111, Merck Millipore) using the MUSE™ Cell Analyzer. In parallel, cells were incubated with ROS assay solution (10 mM HEPES buffer (cat. no. H-1016, Sigma Aldrich; Merck KGaA) 10 mM glucose (cat. no 0.0188, Amresco, OH, USA), 1 µM DCFH-DA (Dichloro-dihydro-fluorescein diacetate) (cat. no. D6883, Sigma Aldrich; Merck KGaA) in 1 × PBS) for 15 min at 37 °C and then visualized under a fluorescence microscope (Nikon Eclipse Ti-E, Minato City, Tokyo, Japan). Se-deficient serum-free medium was used as the positive control 34.
Western Blot analysis
HCC cells were treated with the increasing concentrations of compound 7b (+ : IC50, + + : 2 × IC50) or with DMSO control. For experiments with NAC, cells were treated with either NAC (5 mM) or pre-treated with NAC for 1 h followed by 7b treatment for 24 h. Cells were harvested by scraping, washed with PBS, lysed with RIPA lysis buffer on ice and centrifuged at 13,000 rpm for 20 min. The supernatants were collected, and the protein concentration was measured using the Bradford assay. For SDS-PAGE, 20–50 µg of protein was prepared, and samples were run using the Novex® NuPAGE® Bis–Tris Electrophoresis system (Thermo Fisher Scientific Inc.) according to the manufacturer’s protocol. Transfer of proteins to nitrocellulose membrane was done via XCell IITM Blot Module (Thermo Fisher Scientific Inc.). The blots were incubated with primary antibodies against PARP-1 (Santa Cruz Biotechnology, Dallas, TX, USA, cat. no. sc‐8007, 1:1,000), caspase-3 (Cell Signaling Technology, Danvers, MA, USA, cat no. 9661S, 1:200), caspase-9 (Santa Cruz Biotechnology, cat no. sc-22182, 1:200), p53 (Cell Signaling Technology cat. no. 9286S, 1:1000), p21 (Cell Signaling Technology cat. no. 2946S), cyclin B1 (Cell Signaling Technology cat. no. 4135S), CDK1, SAPK/JNK (Cell Signaling Technology cat no. 9252, 1:300), Phospho-SAPK/JNK (Thr183/Tyr185) (Cell Signaling Technology cat. no. 9251, 1:200), phospho-p38 (Thr180/Tyr182) (Cell Signaling Technology cat. no. 9211S, 1:500), phospho-c-Jun (Santa Cruz Biotechnology cat. no. sc-822, 1:200), phospho-ASK1 (Thr845) (Cell Signaling Technology cat. no. 3765, 1:500), phospho-ASK1 (Ser966) (Genscript, Piscataway, NJ, USA, cat. no. A00340, 1:500), phospho-ASK1 (Ser83) (Abcam, Cambridge, UK, cat. no. ab47304, 1:500), phospho-MKK7 (Ser271/Thr275) (Cell Signaling Technology cat. no. 4171, 1:500), phospho-MKK4 (S257 + T261) (Abcam cat. no. ab4760, 1:300), calnexin (Sigma-Aldrich; Merck KGaA cat. no. C4731, 1:5000), and actin (Santa Cruz Biotechnology cat. no. sc-1616, 1:5000) in 0.1% TBST at 4 °C overnight, followed by secondary antibody incubations with HRP-conjugated goat anti-mouse IgG (Sigma-Aldrich; Merck KGaA cat. no. A0168, 1:5000), rabbit anti-goat IgG (Sigma-Aldrich; Merck KGaA cat. no. A8919, 1:5000), or goat anti-rabbit IgG (Sigma-Aldrich; Merck KGaA cat. no. A6154, 1:5000) for 1 h at room temperature. Proteins were visualized by using the enhanced chemiluminescence (ECL) system or C-DiGit Blot Scanner (LI-COR, Lincoln, NE, USA). All original western blot images are provided in Supplementary Fig. S8.
In vivo mouse xenograft experiments
All animals received human care, and study protocols comply with the institution's guidelines. The animal ethics committee of Bilkent University approved the study protocol. In addition, all studies were reported in accordance with the ARRIVE (Animal Research: Reporting of In Vivo Experiments) guidelines. Mahlavu cells prepared in DMEM (10,000,000 cells/mouse) were injected subcutaneously (SC) to the flank of 8–16 weeks old male nude mice as described previously35. Drug treatment was initiated once the tumor volume reached 150mm3. The subjects received compound 7b (100 mg/kg) suspended in simple syrup (16 g glucose in 9 g ddH2O) by oral feeding, and the control group of mice received only 100 µl simple syrup twice a week for 21 days. Nude mice were not treated for the following 3 weeks and imaged with Magnetic Resonance Imaging (acquired with 3-TESLA Siemens MAGNETOM Trio, UMRAM Center, Bilkent University) following intraperitoneal (ip) injection of anesthesia regimen consisting of 10 mg/kg xylaxin and 90 mg/kg ketamine.
Detection of liver cancer stem cell (LCSC) enrichment by flow cytometry
Huh7 and Mahlavu cells were seeded onto 100 mm culture dishes. The next day cells were treated with compound 7b, sorafenib at their IC50 concentrations or their combinations, and corresponding DAPT (positive), or DMSO (vehicle) controls. Fluorescence labeling of LCSCs was done using primary antibodies against CD133 (Miltenyi Biotec, Bergisch Gladbach, North Rhine-Westphalia, Germany cat. no. 130-090-664), and anti-biotin-PE (Miltenyi Biotec, cat. no. 130-090-756), EpCAM (Miltenyi Biotec, cat. no. 130–080-301), or CD90 (Miltenyi Biotec, cat. no. 130-095-403) for flow cytometry analysis as described previously36. Mouse-IgG-FITC (Miltenyi Biotec, cat. no. 130–092-213), and mouse-IgG-biotin antibodies (Miltenyi Biotec, cat. no. 130-093-018) were used as isotype controls. DAPT (Notch pathway inhibitor) was used as a positive control for cancer stem cell inhibition. Results for each treatment group were compared to that of the DMSO control. Changes in the positivity of CD133 + /EpCAM + cells or CD90 + cells were indicative of enrichment or reduction of the LCSC population. BD Accuri C6 and Novoctye flow cytometer (ACEA Biosciences; Agilent Technologies) were used for flow cytometric analysis.
Sphere formation assay
Sphere formation was triggered using DMEM/F12 serum-free medium (Invitrogen) supplemented with epidermal growth factor (20 ng ml−1; cat. no. PHG0311, Thermo Fisher Scientific Inc.), basic fibroblast growth factor (10 ng ml−1; cat. no. F0291, Sigma Aldrich; Merck KGaA), B27 supplement (1:50; cat. no. 10889038, Invitrogen; Thermo Fisher Scientific Inc.), Heparin (2 µg ml−1; cat. no. H3149, Sigma Aldrich; Merck KGaA), insulin (5 µg ml−1; cat. no. I-6634, Sigma Aldrich; Merck KGaA), hydrocortisone (0.5 µg ml−1; cat. no. H0888, Sigma Aldrich; Merck KGaA) and 100 units mL-1 penicillin and streptomycin (Invitrogen; Thermo Fisher Scientific Inc). The cells were cultured in ultra-low attachment 96-well plates (cat. no. CLS3474-24EA, Corning, Millipore; Merck KGaA) for 6–12 days. Images of spheres and measurements of sphere size and number were assessed using light microscopy (Zeiss, Oberkochen, Germany).
Cell migration assay
RT-CES DP system (ACEA Biosciences; Agilent Technologies) was used to test the effect of compound 7b on the migration capacity of Huh7 and Mahlavu cells. The lower chamber of the 16-well CIM-plate (cat. no. 5665817001, ACEA Biosciences; Agilent Technologies) was filled with 160 µl of 10% FBS containing complete DMEM, and the upper chamber was placed on top of the lower chamber. After 1 h at 37 °C, cells that were prepared in the presence of different concentrations of compound 7b (2 µM for Huh7 and 3 µM for Mahlavu), Taxol (20 ng/ml) or DMSO inside serum-free DMEM, to be seeded into the upper chamber (30,000–50,000 cell/well). CIM-plates were placed into the system after 30 min of incubation at room temperature (RT), and CI values were obtained every 15 min for 24 h. Time-zero normalized CI values were used to generate time-dependent migration curves for each experimental group.
Statistical analysis
Data were obtained from three independent experiments, and all experiments were carried out with n ≥ 3 biological replicates. Statistical analysis for in vitro data was done using a Student's t-test. All in vivo experimental data were analyzed using ANOVA, n = 5–6 mice/group (Graphpad Prism version 7.0, or Microsoft Excel). * p < 0.05, ** p < 0.01, *** p < 0.001.
Results
Cytotoxic bioactivities of 4-amino-3-substituted-1,2,4-triazole-5-thiones (1–8) and 3,6-disubstituted-7H-1,2,4-triazolo[3,4-b]-1,3,4-thiadiazines (1a-8c) on cancer cells
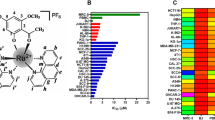
Cytotoxic bioactivities of 8 compounds with 4-amino-3-substituted-1,2,4-triazole-5-thiones (1–8) and 24 compounds with 3,6-disubstituted-7H-1,2,4-triazolo[3,4-b]-1,3,4-thiadiazines (1a-8c) were tested on primary liver and colon cancer cell lines by SRB assay (Table 1). While 4-amino-3-substituted-1,2,4-triazole-5-thiones (1–8) did not exhibit cytotoxic activities on two epithelial cancer cell lines (Huh7 and HCT116), compounds with triazolothiadiazine core had significant cytotoxicity, comparable to the effect of a well-known chemotherapeutic drug camptothecin (CPT). Among all the compounds, three of them (7a, 7b, and 7c) had prominent cytotoxic activities at concentrations ranging from 0.3–5.3 µM (Table 1). These compounds were then screened against a panel of HCC cell lines (Huh7, HepG2, Hep3B, PLC, SK-Hep1, Mahlavu, FOCUS, SNU182, and SNU475). IC50 values were ~ 7–23 µM for 7a, ~ 0.2–1 µM for 7b ~ 12–50 µM for 7c (Supplementary Table S1). With cytotoxic doses smaller than 5 µM, compound 7b was identified as a potent anticancer agent against liver cancer cells. Interestingly, this compound was previously reported to have the most promising and reliable anti-inflammatory activity among newly synthesized 3,6-disubstituted 7H-1,2,4-triazolo[3,4-b]-1,3,4-thiadiazine derivatives25,29. Next, we evaluated the anticancer activities of compound 7b against a panel of HCC cell lines consisting of Huh7, HepG2, Mahlavu, FOCUS, SNU475, and SNU387 to determine the time- and dose-cell growth inhibitory effect. Time-dependent IC50 values of compound 7b for each HCC cell line were calculated accordingly. (Table 2).
Cell growth inhibition, apoptosis, and cell cycle arrest induction by compound 7b
A label-free real-time cell monitoring system (RT-CES) that can measure cell growth by electrical impedance detection was used to determine the real-time anticancer activity of compound 7b. It was demonstrated that compound 7b inhibits cell growth in a dose- and cell line-dependent manner, similar to our initial results with the SRB assay (Fig. 3a). Further experiments were performed to decipher the mechanism underlying this activity. In the presence of compound 7b, cleaved Poly-ADP-ribosyl-polymerase (PARP) fragments indicative of activated apoptotic pathway activation, were visible in most of the HCC cells after 24 h (Fig. 3b). The percentage of apoptotic cells increased significantly, as shown by Annexin-V/PI staining (Supplementary Fig. S1a). Furthermore, caspase-9 and caspase-3 cleavage was also observed, indicating activation of intrinsic apoptotic pathway upon 7b treatment in HCC cells (Supplementary Fig. S1b). To further enlighten the mechanism beneath the apoptotic cell death in these cells, cell cycle distribution in the presence of compound 7b was examined by flow cytometry analysis of propidium iodide-stained cells. Compound 7b induced dose-dependent G2/M arrest in HCC cell lines after 24 h (Fig. 3c). In addition, 7b caused a significant increase in p53 and p21 expression levels and a minor decrease in the levels of CDK1 in HCC cells (Supplementary Fig. S2).
7b induces growth inhibition, apoptosis, and cell cycle arrest in HCC cell lines. (a) Real-time cell growth in the presence of compound 7b and its DMSO control. The cell growth index of 7b treated cells was recorded every 30 min. All compounds were administered in triplicates for RT-CES analysis. (b) Western blot analysis displaying the cleaved PARP upon compound 7b treatment. All HCC cell lines were treated with 2.5 μM 7b or its corresponding DMSO for 24 h. Calnexin protein was used as an equal loading control. (c) Histograms and the bar graph indicating cell cycle distribution of Huh7, HepG2, Mahlavu, FOCUS, SNU475 and SNU387 cells treated with increasing concentrations of compound 7b and their DMSO controls for 24 h.
Induction of ROS accumulation and activation of JNK pathway upon treatment with 7b
The most common underlying mechanisms inducing G2/M arrest followed by apoptosis in cells are ER-stress and the accumulation of reactive oxygen species (ROS)38,39. Since XBP-1 splicing via semi-quantitative PCR is indicative of ER-stress induction40, we evaluated the spliced XBP-1 in Huh7 cells upon treatment with compound 7b, DMSO or 500 ng/mL tunicamycin (TN) for 24 h. Tunicamycin was used as a positive control at an ER stress-inducing dose41. Compound 7b did not induce ER-stress-specific XBP-1 splicing in these cells, unlike TN (Supplementary Figure S3). Then, we tested whether oxidative stress upon ROS accumulation was triggered by compound 7b in Huh7 and Mahlavu cells, which were treated with increasing concentrations of compound 7b (1.5 and 3 µM) for 8, 12, and 24 h. In the presence of compound 7b, the accumulation of ROS was detected significantly in both Huh7 and Mahlavu cells by DCFH-DA staining, which was in parallel quantified and statistically analyzed by flow cytometric analysis (Fig. 4a and b). Additionally, ROS induction was significantly decreased once the cells were pretreated with NAC prior to 7b treatment (Fig. 4c). NAC pretreatment decreased apoptosis caused by 7b and resulted in lower levels of cell cycle arrest at the G2/M phase (Suppl. Fig. S4), which further supported the compound's mechanism of action as oxidative stress-induced apoptosis.
7b induces oxidative stress in HCC cell lines. (a) Representative images of DCFH-DA staining of Huh7 and Mahlavu cells treated with Se-deficient media or increasing concentrations of 7b for 8, 12, and 24 h. All images are obtained under a fluorescent microscope with 20 × magnification. (b) Flow cytometric analysis of ROS accumulation in HCC cells. Bar graphs indicate the fold change and quantitative analysis of ROS-positive cells relative to DMSO after 12 h for each treatment condition. Se-deficient medium was used as a positive control for ROS induction. (c) Quantitative analysis of reversal of ROS induction upon NAC pretreatment in HCC cells is represented in bar graphs. Cells were either treated with 3 µM 7b only or 5 mM NAC for 1 h and then with 7b for 12 h. ROS accumulation in HCC cells was analyzed using flow cytometry.
It is well described in the literature that activation of c-Jun NH2-terminal kinases (JNKs), also known as stress-activated protein kinases (SAPKs), is a typical response to many forms of stress42,43. Therefore, we checked the levels of activated JNK and its downstream element c-Jun, a transcription factor phosphorylated by JNK1 and JNK2, in HCC cells treated with 7b, or JNKInhV relative to DMSO treated cells for 24 h. We observed a significant increase in the phosphorylated-JNK and phosphorylated-c-Jun protein levels upon compound 7b treatment (Fig. 5a). We tested the effect of 7b on the regulation of p38, another ROS-mediator-activated protein 44, and found that phospho-p38 levels also increase significantly upon 7b treatment in HCC cell lines (Fig. 5a). To further investigate the effects of ROS accumulation on the upstream components of the JNK signaling pathway, levels of MKK-4, MKK745 and apoptosis signal-regulating kinase 1 (ASK1) proteins46,47 were analyzed. We found that compound 7b increased the phospho-MKK7 protein levels (Fig. 5b) but did not change the phospho-MKK4 levels in HCC cells (Supplementary Fig. S5). Furthermore, 7b also altered the phosphorylation status of its upstream activator ASK1 (Fig. 5b). In contrast, NAC pretreatment reduced levels of JNK phosphorylation in HCC cells, further supporting our findings that JNK pathway activation is caused by ROS accumulation (Supplementary Fig. S6). To further analyze the role of JNK pathway activation and apoptosis induction by 7b, HCC cells were treated with JNK inhibitor together with 7b for 24 h and apoptotic cells were measured by Annexin-V/PI staining. Cells were partially resistant to apoptosis in the presence of JNK inhibitor compared to cells treated with 7b alone (Supplementary Fig. S4).
7b activates the JNK pathway in HCC cell lines. (a) Representative western blot images of total or active (phosphorylated) JNK1 and JNK2 protein levels in HCC cell lines treated with increasing concentrations of compound 7b, cell line-specific IC50 concentrations of JNKInhV (Huh7: 2 μM, HepG2: 2 μM, Mahlavu: 5 μM, FOCUS: 0.5 μM), and DMSO control (top left panel); phospho-p38 and phospho-c-Jun levels in HCC cell lines treated with increasing concentrations of 7b (+ : IC50, + + : 2 × IC50) and DMSO control (bottom left) for 24 h. Bar graphs represent quantitative analysis of the relative band intensity values obtained from 3 different experiments of each treatment group for phosphorylated JNK, phosphorylated c-Jun proteins compared to DMSO using the ImageJ analysis tool (right panel). (b) Representative images indicate the changes in the levels of P-MKK7, P-ASK1Thr845, P-ASK1Ser966 and P-ASK1Ser83 proteins by western blot analysis. Huh7, HepG2, Mahlavu, and FOCUS cells were treated with increasing concentrations of compound 7b and corresponding DMSO for 24 h (bottom left panel). Bar graphs represent quantitative analysis of the relative band intensity values obtained from 3 different experiments of each treatment group for MKK-7, and phosphorylated ASK1 proteins compared to DMSO using ImageJ analysis tool (right panel). Calnexin or β-actin were used as equal loading controls for all western blot experiments.
In vivo anti-tumor activity of compound 7b in mouse xenografts
Mahlavu cell xenografts in nude mice were used to determine the in vivo anti-tumor effect of compound 7b. Mahlavu cells were selected due to their significant response to compound 7b and their poorly differentiated and highly metastatic nature48. We observed that compound 7b increased the overall survival of nude mice (Fig. 6a and Supplementary. Fig. S7). Moreover, the tumor volumes were significantly decreased in the compound 7b treated group, as demonstrated by the representative MRI images (Fig. 6b). Altogether, our in vivo data revealed that compound 7b is orally tolerable in nude mice and is a novel and potentially effective drug candidate against HCC.
7b reduces tumor growth and increases the overall survival of nude mice. (a) Mahlavu xenografted nude mice were treated with either (100 mg/kg) compound 7b suspended in simple syrup or only simple syrup twice a week once the tumor size reached a volume of 150 mm3. Tumor sizes are recorded twice a week throughout the experiment. Time-zero normalized data are shown as mean ± SD for each group of animals (n = 5); ***P < 0.001, as calculated using one-way ANOVA (b) Representative MRI images taken at the 135th day for compound 7b treated group and 105th day for control.
Effect of compound 7b on liver cancer stem cell enrichment
Tumor initiating cells of liver, so-called (liver cancer stem cells-LCSCs), are known to be capable of reconstituting the tumor by themselves, triggering metastatic events, and enhancing drug resistance in cancer cells49. Therefore, we tested compound 7b for its effect on enrichment LCSCs, which can be quantified by detecting CSC markers found on the surface of these cells. For this purpose, Huh7 and Mahlavu cells were treated with compound 7b and other inhibitors such as sorafenib, DAPT, or DMSO control at IC50 concentrations for 72 h. Cells that remained viable after treatment were collected, and the expression of LCSC markers CD133 and EpCAM (for Huh7) and CD90 (for Mahlavu) were evaluated by flow cytometry. Results have revealed that compound 7b was able to decrease the CD133 + /EpCAM + population in Huh7 cells and the CD90 + population in Mahlavu cells significantly (Fig. 7a). Besides, sorafenib was known to enrich the LCSC population, as demonstrated previously36. To test the combinatory effect of compound 7b with sorafenib, Huh7 cells were treated with the increasing concentrations of both compounds simultaneously for 72 h. Flow cytometry analysis and the sphere formation assay have shown that compound 7b alone and, in combination with sorafenib, reduced the LCSC ratio and the sphere formation capacity of Huh7 cells. (Fig. 7b).
7b reduces enrichment of LCSCs and the migration capacity of HCC cells. Huh7 and Mahlavu cells were treated with sorafenib (5 µM), DAPT (5 µM), compound 7b (2 µM), or DMSO control for 72 h. (a) Images represent flow cytometric analysis of CD133/EpCAM positivity in Huh7 cells, and CD90 positivity in Mahlavu cells upon treatment with indicated compounds/inhibitors. Upper-right quadrant (UR): double-positive cells for CD133 and EpCAM for Huh7 cells; upper-window: positivity for CD90 for Mahlavu cells. (b) Flow cytometric analysis of CD133/EpCAM positivity in Huh7 cells treated with compound 7b or sorafenib alone and with combinations of compound 7b and sorafenib and representative images of spheres formed by Huh7 cells. Bar graphs indicate quantitative analysis of sphere count and diameter in different treatment groups. (c) Cell migration index of Huh7 and Mahlavu cells treated with the positive control, taxol (20 ng/ml) or compound 7b (2 µM for Huh7 and 3 µM for Mahlavu) for 24 h.
Effect of compound 7b on cell migration
To identify the effect of compound 7b on the migration capacity of HCC cells, a real-time cell migration system (RT-CES, DP system, xCELLigence) was used. Huh7 and Mahlavu cells were treated with taxol as a positive control for cell migration inhibition or with compound 7b (Huh7: 2 µM, Mahlavu: 3 µM) for 24 h. Comparable with the effect of taxol, compound 7b inhibited the migration of both cell lines compared to the DMSO treated cells (Fig. 7c). Altogether, 7b could effectively interfere with the drug resistance-related cellular mechanisms such as stemness and migration.
Discussion
HCC is one of the most common and deadly cancers in the world. Sorafenib, which is a multi-kinase inhibitor approved by FDA was reported to prolong survival of advanced stage HCC patients by only 3 months50. Although second-line treatment options such as Regorafenib were also studied for patients who are intolerant to sorafenib, drug resistance has become a critical problem for these patients due to the highly heterogeneous molecular nature of HCC.
Recent studies have reported that NSAIDs effectively decrease the incidence and mortality of various types of cancers12,51,52. In this study, we demonstrated that compound 7b, a synthetic 1,2,4-triazolo[3,4-b]-1,3,4-thiadiazine NSAID derivative, is a potent anticancer agent for HCC cells in vitro and in vivo, which was previously reported to have promising analgesic and anti-inflammatory activities25,29. Among the reported numerous biological effects, accumulation of reactive oxygen species (ROS) is considered as the underlying mechanism of the anticancer potential of NSAIDs53,54. Compatible with the literature, we have demonstrated that ROS accumulation induces growth inhibition, apoptosis, and G2/M cell cycle arrest in the presence of 7b (Figs. 3 and 4). We also examined signaling pathways activated by oxidative stress and found that activation of JNK and p38 as well as the downstream protein c-Jun was induced upon 7b treatment in HCC cells. (Fig. 5a). Further analysis of the upstream components of the JNK pathway revealed that compound 7b treatment causes cell line-specific activation of MKK7 protein (Fig. 5b) as well as dynamic regulation of phosphorylation sites on the ASK1 protein (Fig. 5b). Our in vivo data supported our findings on anticancer activity of 7b, where orally administered compound increased the overall disease-free survival for more than 4 months and reduced tumor size significantly in Mahlavu xenografts in nude mice (Fig. 6).
One of the most important factors associated with drug resistance in HCC is the presence of cancer stem cells. Conventional therapies fail to affect slowly dividing stem cell-like cancer cells and mainly target rapidly dividing tumor cells55. CSCs can reconstitute the tumor by themselves and manage to acquire metastatic features to migrate to distant organs56,57. Hence, we have also tested the compounds against LCSC and revealed that compound 7b was also effective in inhibiting the enrichment of LCSCs (Fig. 7), meaning that compound 7b is not solely active on tumor cells (non-stem), but also tumor-initiating cells in HCC. We also tested the effect of compound 7b on the migration capacity of HCC cells, since CSC activity is associated with the migration and metastatic capacity of cancer cells58. 7b could effectively inhibit the migration of Huh7 and Mahlavu cells, comparable to the effect of Taxol, which is known as a microtubule stabilizing agent59 that is widely used as a positive control for inhibition of migration in cancer cells60,61.
In the last decade, the anticancer and anti-metastatic effects of a well-defined NSAID, aspirin, has been described in cancer including HCC62. It is reported that by the induction of metabolic and oxidative stress in HepG2 cells, aspirin causes apoptosis through mitochondrial dysfunction63. In another study, aspirin was shown to attenuate pro-metastasis caused by sorafenib by upregulating the tumor suppressor HTATIP2 in nude mice xenografts64. Yet, the use of aspirin in treatment of cancer and its prevention remains uncertain due to the problems with the optimal dosage of aspirin and its serious side-effects such as gastrointestinal bleeding, increased uric acid, and coagulation inhibition65,66. Since these drugs are not designed to treat or prevent cancer progression, designing and developing novel NSAIDs that target cancer-related mechanisms is very crucial and can be considered as a new class of anticancer pharmaceutical agents such as non-steroid anti-inflammatory chemotherapeutic drugs (NSAICD).
In conclusion, our results revealed that 7b, a synthetic 1,2,4-triazolo[3,4-b]-1,3,4-thiadiazine NSAID derivative, is a promising compound with anticancer and anti-stem cell activities against HCC cells in vitro and in vivo. The anticancer effect was attributed to the induction of oxidative stress, cell cycle arrest, and eventually apoptosis through the JNK pathway regulation (Fig. 8). These findings highlight the potency of 7b as a new NSAICD, bearing a thiadiazine core, to be considered a promising anticancer drug for HCC patients and deserves further analysis.
Illustration of molecular mechanisms involved in response to compound 7b treatment in HCC cell lines. 7b blocks cell proliferation through induction of ROS accumulation and JNK pathway activation which further stimulates transcription of genes related to apoptosis and cell cycle arrest. As a result, G2/M arrest (through the regulatory effect of p53 and p21), and apoptosis (through intrinsic pathway) are observed. 7b is also a potent compound acting on cancer stemness and migration capacity of HCC cell lines. This figure was created with BioRender.com.
Abbreviations
- HCC:
-
Hepatocellular carcinoma
- NSAID:
-
Nonsteroidal anti-inflammatory drug
- ROS:
-
Reactive oxygen species
- SRB:
-
Sulforhodamine B
- DMSO:
-
Dimethyl sulfoxide
- LCSC:
-
Liver cancer stem cell
- RT:
-
Room temperature
- IC50:
-
50% Inhibitory concentration
- DMEM:
-
Dulbecco's Modified Eagle Medium
- FBS:
-
Fetal bovine serum
- PBS:
-
Phosphate-buffered saline
- CI:
-
Cell index
- RT-PCR:
-
Real-time polymerase chain reaction
- Se:
-
Selenium
- DCFH-DA:
-
Dichloro-dihydro-fluorescein diacetate
References
Villanueva, A. Hepatocellular Carcinoma. N. Engl. J. Med. 380, 1450–1462. https://doi.org/10.1056/NEJMra1713263 (2019).
Jeng, K. S. et al. Heterogeneity of hepatocellular carcinoma contributes to cancer progression. Crit. Rev. Oncol. Hematol. 94, 337–347. https://doi.org/10.1016/j.critrevonc.2015.01.009 (2015).
Finn, R. S. Emerging targeted strategies in advanced hepatocellular carcinoma. Semin. Liver Dis. 33(Suppl 1), S11-19. https://doi.org/10.1055/s-0033-1333632 (2013).
Gong, X. L. & Qin, S. K. Progress in systemic therapy of advanced hepatocellular carcinoma. World J. Gastroenterol. 22, 6582–6594. https://doi.org/10.3748/wjg.v22.i29.6582 (2016).
Frager, S. Z. & Schwartz, J. M. Hepatocellular Carcinoma: Epidemiology, Screening, and Assessment of Hepatic Reserve. Curr. Oncol. 27, 138–143. https://doi.org/10.3747/co.27.7181 (2020).
Llovet, J. M. et al. Hepatocellular carcinoma. Nat. Rev. Dis. Primers 7, 6. https://doi.org/10.1038/s41572-020-00240-3 (2021).
Llovet, J. M. et al. Sorafenib in advanced hepatocellular carcinoma. N. Engl. J. Med. 359, 378–390. https://doi.org/10.1056/NEJMoa0708857 (2008).
Kim, K. et al. Regorafenib in advanced hepatocellular carcinoma (HCC): Considerations for treatment. Cancer Chemother. Pharmacol. 80, 945–954. https://doi.org/10.1007/s00280-017-3431-5 (2017).
Deng, G. L., Zeng, S. & Shen, H. Chemotherapy and target therapy for hepatocellular carcinoma: New advances and challenges. World J. Hepatol. 7, 787–798. https://doi.org/10.4254/wjh.v7.i5.787 (2015).
Yeo, W. et al. A randomized phase III study of doxorubicin versus cisplatin/interferon alpha-2b/doxorubicin/fluorouracil (PIAF) combination chemotherapy for unresectable hepatocellular carcinoma. J. Natl. Cancer Inst. 97, 1532–1538. https://doi.org/10.1093/jnci/dji315 (2005).
Tang, W. et al. The mechanisms of sorafenib resistance in hepatocellular carcinoma: Theoretical basis and therapeutic aspects. Signal Transduct. Target Ther. 5, 87. https://doi.org/10.1038/s41392-020-0187-x (2020).
Rayburn, E. R., Ezell, S. J. & Zhang, R. Anti-inflammatory agents for cancer therapy. Mol. Cell Pharmacol. 1, 29–43. https://doi.org/10.4255/mcpharmacol.09.05 (2009).
Aytac, P. S. et al. Novel triazolothiadiazines act as potent anticancer agents in liver cancer cells through Akt and ASK-1 proteins. Bioorg. Med. Chem. 24, 858–872. https://doi.org/10.1016/j.bmc.2016.01.013 (2016).
Greenspan, E. J., Madigan, J. P., Boardman, L. A. & Rosenberg, D. W. Ibuprofen inhibits activation of nuclear {beta}-catenin in human colon adenomas and induces the phosphorylation of GSK-3{beta}. Cancer Prev. Res. (Phila) 4, 161–171. https://doi.org/10.1158/1940-6207.CAPR-10-0021 (2011).
Kim, M. S. et al. Naproxen induces cell-cycle arrest and apoptosis in human urinary bladder cancer cell lines and chemically induced cancers by targeting PI3K. Cancer Prev. Res. (Phila) 7, 236–245. https://doi.org/10.1158/1940-6207.CAPR-13-0288 (2014).
Santen, R. J. Inhibition of aromatase: Insights from recent studies. Steroids 68, 559–567. https://doi.org/10.1016/S0039-128x(03)00096-5 (2003).
Goss, P. E. & Strasser-Weippl, K. Aromatase inhibitors for chemoprevention. Best Pract. Res. Cl. En. 18, 113–130. https://doi.org/10.1016/S1521-690x(03)00070-8 (2004).
Albrecht, B. K. et al. Discovery and optimization of triazolopyridazines as potent and selective inhibitors of the c-Met kinase. J. Med. Chem. 51, 2879–2882. https://doi.org/10.1021/jm800043g (2008).
Boezio, A. A. et al. Discovery and optimization of potent and selective triazolopyridazine series of c-Met inhibitors. Bioorg. Med. Chem. Lett. 19, 6307–6312. https://doi.org/10.1016/j.bmcl.2009.09.096 (2009).
Bhat, K. S. et al. Synthesis and anti-tumor activity studies of some new fused 1,2,4-triazole derivatives carrying 2,4-dichloro-5-fluorophenyl moiety. Eur. J. Med. Chem. 44, 5066–5070. https://doi.org/10.1016/j.ejmech.2009.09.010 (2009).
Ibrahim, D. A. Synthesis and biological evaluation of 3,6-disubstituted [1,2,4]triazolo[3,4-b][1,3,4]thiadiazole derivatives as a novel class of potential anti-tumor agents. Eur. J. Med. Chem. 44, 2776–2781. https://doi.org/10.1016/j.ejmech.2009.01.003 (2009).
Sunil, D. et al. Synthesis, characterization, antioxidant, and anticancer studies of 6-[3-(4-chlorophenyl)-1H-pyrazol-4-yl]-3-[(2-naphthyloxy)methyl][1,2,4]triazolo[3,4-b][1,3,4]thiadiazole in HepG2 cell lines. Med. Chem. Res. 20, 1074–1080. https://doi.org/10.1007/s00044-010-9436-9 (2011).
Khan, I. et al. Synthesis, crystal structure and biological evaluation of some novel 1,2,4-triazolo[3,4-b]-1,3,4-thiadiazoles and 1,2,4-triazolo[3,4-b]-1,3,4-thiadiazines. Eur. J. Med. Chem. 78, 167–177. https://doi.org/10.1016/j.ejmech.2014.03.046 (2014).
Tozkoparan, B. et al. Novel 1,2,4-Triazolo[3,4-b]-1,3,4-thiadiazole derivatives as dual analgesic/anti-inflammatory and antimicrobial agents. Lett. Drug Des. Discov. 9, 204–212. https://doi.org/10.2174/157018012799079626 (2012).
Tozkoparan, B., Aytac, S. P. & Aktay, G. Novel 3,6-disubstituted 7H–1,2,4-triazolo[3,4-b][1,3,4]thiadiazines: Synthesis, characterization, and evaluation of analgesic/anti-inflammatory, antioxidant activities. Arch. Pharm. 342, 291–298. https://doi.org/10.1002/ardp.200800188 (2009).
Bishayee, A. The role of inflammation and liver cancer. Adv. Exp. Med. Biol. 816, 401–435. https://doi.org/10.1007/978-3-0348-0837-8_16 (2014).
Radwan, R. R., Zaher, N. H. & El-Gazzar, M. G. Novel 1,2,4-triazole derivatives as anti-tumor agents against hepatocellular carcinoma. Chem.-Biol. Interact 274, 68–79. https://doi.org/10.1016/j.cbi.2017.07.008 (2017).
Tuncbilek, M., Guven, E. B., Onder, T. & Cetin Atalay, R. Synthesis of novel 6-(4-substituted piperazine-1-yl)-9-(beta-D-ribofuranosyl)purine derivatives, which lead to senescence-induced cell death in liver cancer cells. J. Med. Chem. 55, 3058–3065 (2012).
Aytac, S. P. et al. Synthesis of 3,6-disubstituted 7H–1,2,4-triazolo[3,4-b]-1,3,4-thiadiazines as novel analgesic/anti-inflammatory compounds. Eur. J. Med. Chem. 44, 4528–4538. https://doi.org/10.1016/j.ejmech.2009.06.026 (2009).
Oefinger, P. E., Bronson, D. L. & Dreesman, G. R. Induction of hepatitis B surface antigen in human hepatoma-derived cell lines. J. Gen. Virol. 53, 105–113. https://doi.org/10.1099/0022-1317-53-1-105 (1981).
He, L. et al. Establishment and characterization of a new human hepatocellular carcinoma cell line. In Vitro 20, 493–504. https://doi.org/10.1007/BF02619623 (1984).
Onen-Bayram, F. E. et al. A novel thiazolidine compound induces caspase-9 dependent apoptosis in cancer cells. Bioorg. Med. Chem. 20, 5094–5102. https://doi.org/10.1016/j.bmc.2012.07.016 (2012).
Hawash, M. M. et al. Synthesis and biological evaluation of novel pyrazolic chalcone derivatives as novel hepatocellular carcinoma therapeutics. Eur. J. Med. Chem. 129, 12–26. https://doi.org/10.1016/j.ejmech.2017.02.002 (2017).
Irmak, M. B., Ince, G., Ozturk, M. & Cetin-Atalay, R. Acquired tolerance of hepatocellular carcinoma cells to selenium deficiency: A selective survival mechanism?. Cancer Res. 63, 6707–6715 (2003).
Durmaz, I. et al. Liver cancer cells are sensitive to Lanatoside C induced cell death independent of their PTEN status. Phytomedicine 23, 42–51. https://doi.org/10.1016/j.phymed.2015.11.012 (2016).
Kahraman, D. C. et al. Quinoides and VEGFR2 TKIs influence the fate of hepatocellular carcinoma and its cancer stem cells. Med. Chem. Comm. 8, 81–87. https://doi.org/10.1039/c6md00392c (2017).
Kahraman, D. C., Kahraman, T. & Cetin-Atalay, R. Targeting PI3K/Akt/mTOR pathway identifies differential expression and functional role of IL8 in liver cancer stem cell enrichment. Mol. Cancer Ther. 18, 2146–2157. https://doi.org/10.1158/1535-7163.MCT-19-0004 (2019).
Shi, Y. et al. ROS-dependent activation of JNK converts p53 into an efficient inhibitor of oncogenes leading to robust apoptosis. Cell Death Differ. 21, 612–623. https://doi.org/10.1038/cdd.2013.186 (2014).
Redza-Dutordoir, M. & Averill-Bates, D. A. Activation of apoptosis signalling pathways by reactive oxygen species. Biochim. Biophys. Acta 2977–2992, 2016. https://doi.org/10.1016/j.bbamcr.2016.09.012 (1863).
van Schadewijk, A., van’t Wout, E. F., Stolk, J. & Hiemstra, P. S. A quantitative method for detection of spliced X-box binding protein-1 (XBP1) mRNA as a measure of endoplasmic reticulum (ER) stress. Cell Stress Chaperones 17, 275–279 (2012).
Oslowski, C. M. & Urano, F. Measuring ER stress and the unfolded protein response using mammalian tissue culture system. Methods Enzymol 490, 71–92. https://doi.org/10.1016/B978-0-12-385114-7.00004-0 (2011).
Gutierrez, G. J. et al. JNK-mediated phosphorylation of Cdc25C regulates cell cycle entry and G(2)/M DNA damage checkpoint. J. Biol. Chem. 285, 14217–14228. https://doi.org/10.1074/jbc.M110.121848 (2010).
MacCorkle, R. A. & Tan, T. H. Mitogen-activated protein kinases in cell-cycle control. Cell Biochem. Biophys. 43, 451–461. https://doi.org/10.1385/CBB:43:3:451 (2005).
Min, L., He, B. & Hui, L. Mitogen-activated protein kinases in hepatocellular carcinoma development. Semin. Cancer Biol. 21, 10–20. https://doi.org/10.1016/j.semcancer.2010.10.011 (2011).
Haeusgen, W., Herdegen, T. & Waetzig, V. The bottleneck of JNK signaling: Molecular and functional characteristics of MKK4 and MKK7. Eur. J. Cell Biol. 90, 536–544. https://doi.org/10.1016/j.ejcb.2010.11.008 (2011).
Shiizaki, S., Naguro, I. & Ichijo, H. Activation mechanisms of ASK1 in response to various stresses and its significance in intracellular signaling. Adv. Biol. Regul. 53, 135–144. https://doi.org/10.1016/j.jbior.2012.09.006 (2013).
Hattori, K., Naguro, I., Runchel, C. & Ichijo, H. The roles of ASK family proteins in stress responses and diseases. Cell Commun. Signal 7, 9. https://doi.org/10.1186/1478-811X-7-9 (2009).
Buontempo, F. et al. Inhibition of Akt signaling in hepatoma cells induces apoptotic cell death independent of Akt activation status. Invest. New Drugs 29, 1303–1313. https://doi.org/10.1007/s10637-010-9486-3 (2011).
Cheng, Z., Li, X. & Ding, J. Characteristics of liver cancer stem cells and clinical correlations. Cancer Lett. 379, 230–238. https://doi.org/10.1016/j.canlet.2015.07.041 (2016).
Bruix, J., da Fonseca, L. G. & Reig, M. Insights into the success and failure of systemic therapy for hepatocellular carcinoma. Nat. Rev. Gastroenterol. Hepatol. 16, 617–630. https://doi.org/10.1038/s41575-019-0179-x (2019).
Roxburgh, C. S. & McMillan, D. C. Cancer and systemic inflammation: Treat the tumour and treat the host. Br. J. Cancer 110, 1409–1412. https://doi.org/10.1038/bjc.2014.90 (2014).
Zappavigna, S. et al. Anti-inflammatory drugs as anticancer agents. Int. J. Mol. Sci. https://doi.org/10.3390/ijms21072605 (2020).
Adachi, M. et al. Nonsteroidal anti-inflammatory drugs and oxidative stress in cancer cells. Histol. Histopathol. 22, 437–442. https://doi.org/10.14670/HH-22.437 (2007).
Tinsley, H. N. et al. New NSAID targets and derivatives for colorectal cancer chemoprevention. Recent Results Cancer Res. 191, 105–120. https://doi.org/10.1007/978-3-642-30331-9_6 (2013).
Senthebane, D. A. et al. The role of tumor microenvironment in chemoresistance: To survive, keep your enemies closer. Int. J. Mol. Sci. https://doi.org/10.3390/ijms18071586 (2017).
Oishi, N. & Wang, X. W. Novel therapeutic strategies for targeting liver cancer stem cells. Int. J. Biol. Sci. 7, 517–535 (2011).
Yamashita, T. & Kaneko, S. Liver cancer stem cells: Recent progress in basic and clinical research. Regen. Ther. 17, 34–37. https://doi.org/10.1016/j.reth.2021.03.002 (2021).
Schulte, L. A., Lopez-Gil, J. C., Sainz, B. Jr. & Hermann, P. C. The cancer stem cell in hepatocellular carcinoma. Cancers (Basel) https://doi.org/10.3390/cancers12030684 (2020).
Belotti, D. et al. The microtubule-affecting drug paclitaxel has antiangiogenic activity. Clin. Cancer Res. 2, 1843–1849 (1996).
Kuljittichanok, D. et al. Effect of Derris scandens extract on a human hepatocellular carcinoma cell line. Oncol. Lett. 16, 1943–1952. https://doi.org/10.3892/ol.2018.8824 (2018).
Wang, X. et al. In vitro wound healing of tumor cells: inhibition of cell migration by selected cytotoxic alkaloids. BMC Pharmacol. Toxicol. 20, 4. https://doi.org/10.1186/s40360-018-0284-4 (2019).
Zhang, Z., Chen, F. & Shang, L. Advances in anti-tumor effects of NSAIDs. Cancer Manag. Res. 10, 4631–4640. https://doi.org/10.2147/CMAR.S175212 (2018).
Raza, H., John, A. & Benedict, S. Acetylsalicylic acid-induced oxidative stress, cell cycle arrest, apoptosis and mitochondrial dysfunction in human hepatoma HepG2 cells. Eur. J. Pharmacol. 668, 15–24. https://doi.org/10.1016/j.ejphar.2011.06.016 (2011).
Lu, L. et al. Aspirin minimized the pro-metastasis effect of sorafenib and improved survival by up-regulating HTATIP2 in hepatocellular carcinoma. PLoS ONE 8, e65023. https://doi.org/10.1371/journal.pone.0065023 (2013).
Huang, E. S. et al. Long-term use of aspirin and the risk of gastrointestinal bleeding. Am. J. Med. 124, 426–433. https://doi.org/10.1016/j.amjmed.2010.12.022 (2011).
Li, Z. et al. Effects of aspirin on the gastrointestinal tract: Pros. vs cons.. Oncol. Lett. 20, 2567–2578. https://doi.org/10.3892/ol.2020.11817 (2020).
Acknowledgements
This work was supported by The Scientific and Technological Research Council of Turkey Grant no.112S030 and The Turkish Ministry of Development project no.2016K121540. We would like to thank Dr. Aybar Can Acar for his valuable contribution to KanSiL infrastructure grant, and Tugce Oksakli for laboratory technical assistance.
Author information
Authors and Affiliations
Contributions
B.T. and R.C.A. participated in research design and supervised the study. D.C.K., E.B.G., and P.A. performed the experiments. G.A. guided and contributed to the in vivo experiments. D.C.K., E.B.G. and R.C.A. analyzed the data. D.C.K., E.B.G., B.T., and R.C.A. prepared the figures and contributed to the writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kahraman, D.C., Bilget Guven, E., Aytac, P.S. et al. A new triazolothiadiazine derivative inhibits stemness and induces cell death in HCC by oxidative stress dependent JNK pathway activation. Sci Rep 12, 15139 (2022). https://doi.org/10.1038/s41598-022-17444-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-17444-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.