Abstract
Knowledge about contagiousness is key to accurate management of hospitalized COVID-19 patients. Epidemiological studies suggest that in addition to transmission through droplets, aerogenic SARS-CoV-2 transmission contributes to the spread of infection. However, the presence of virus in exhaled air has not yet been sufficiently demonstrated. In pandemic situations low tech disposable and user-friendly bedside devices are required, while commercially available samplers are unsuitable for application in patients with respiratory distress. We included 49 hospitalized COVID-19 patients and used a disposable modular breath sampler to measure SARS-CoV-2 RNA load in exhaled air samples and compared these to SARS-CoV-2 RNA load of combined nasopharyngeal throat swabs and saliva. Exhaled air sampling using the modular breath sampler has proven feasible in a clinical COVID-19 setting and demonstrated viral detection in 25% of the patients.
Similar content being viewed by others
Introduction
SARS-CoV-2 transmission is thought to largely depend on droplets arising from the upper respiratory tract, expelled through talking, coughing, and sneezing, which settle down quickly and relatively close to its source1. However, aerosols, in which the virus remains replicative for at least three hours, remain suspended in the air drifting long distances2,3,4,5, suggesting a role for transmission via aerosols. This was first confirmed by Richard et al., showing that SARS-CoV-2 can be transmitted via air between ferrets6. The role of superspreading events further suggests that aerosol transmission contributes to the pandemic7. It was recently found that the newly emerged variants of concern show increased infectivity and further confirming the importance of aerosol-mediated spread8,9,10.
Nasopharyngeal throat swabs are a common diagnostic sample, and it is challenging to effectively sample exhaled virions11. Given the multiple modes of transmission, it remains questionable whether the SARS-CoV-2 RNA load in the upper respiratory tract is the best proxy for contagiousness. Therefore, we assessed the feasibility of sampling exhaled air from hospitalized COVID-19 patients to measure SARS-CoV-2 RNA using a modular breath sampler. This enabled the collection of a liquid sample, compatible with the conventional molecular diagnostic infrastructure, using a disposable device compatible with application in a highly infectious surrounding. Additionally, we aimed to determine the influence of patient characteristics on RNA detection and load.
Materials and methods
Ethical considerations
This cohort study was conducted between October 2020 and February 2021, according to the principles of Helsinki and the Medical Research Involving Human Subjects Act (WMO) at Radboud University Medical Centre (Radboudumc, Nijmegen, the Netherlands). All study protocols were reviewed and approved by the local ethics board, the Committee on Research Involving Human Subjects (CMO) Arnhem-Nijmegen (CMO 2020–6517), which deemed verbal informed consent sufficient. All participants provided verbal informed consent at inclusion.
Participants
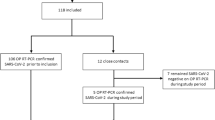
Patients ≥ 18 years of age were included prospectively within 7 days after admission to the Radboudumc. COVID-19 was confirmed by RT-qPCR on a combined nasopharyngeal throat swab.
Sample collection
Patient characteristics, clinical features, and routine hemocytometric and inflammatory laboratory measurements were collected on standardized patient charts. Routine diagnostic laboratory measurements were registered on the day of sampling (+ /− 1 day).
From each patient, samples were simultaneously collected using three different methodologies. A combined nasopharyngeal throat swab was taken using the hospital’s protocol according to national guidelines12. After collection, the sample was placed in 5.0 mL virus transport medium consisting of Hank’s balanced salt solution (Gibco) containing 2% FCS (Sigma-Aldrich), 100 µg/mL gentamicin (Gibco), and 0.5 µg/mL amphotericin-B (Gibco) and stored at -20 °C until further processing.
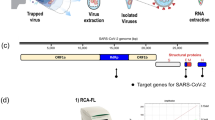
Exhaled air was assessed using a modular breath sampler (MBS, Xheal Diagnostics, Fig. 1a). Patients were instructed to inhale and exhale normally through the mouthpiece for one minute, after which they inhaled and exhaled deeply three times before continuing to breath normally to complete two minutes in total. The sample, collected in capture buffer, was stored at − 20 °C until further processing.
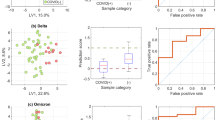
(a) Modular breath sampler. During sampling, the patient breathes through the mouthpiece; inhaling (I) and exhaling (II). The exhaled air is guided through the sampler to the diffuser into the capture buffer. The exhaled air leaves the sampler on the back end. After sampling, the collection tube can be disconnected and stored for analysis. (b) Correlation plot depicting SARS-CoV-2 viral RNA load measured in nasopharyngeal throat swab and saliva samples. (c) SARS-CoV-2 RNA detection in exhaled air samples and nasopharyngeal throat swab samples. Filled dots ( ) represent cases in which SARS-CoV-2 RNA was detected in exhaled air samples. Open dots (○) represent cases in which SARS-CoV-2 RNA was not detected.
) represent cases in which SARS-CoV-2 RNA was detected in exhaled air samples. Open dots (○) represent cases in which SARS-CoV-2 RNA was not detected.
Saliva was collected by instructing patients to spit in a sterile 15 or 50 mL container (Greiner) and stored at − 20 °C.
RT-qPCR
The presence and viral load of SARS-CoV-2 were determined using RT-qPCR adapted from that of the Dutch National Institute for Public Health and the Environment (RIVM). Briefly, 500 µl material was lysed in 450 µl MagNAPure lysis/binding buffer (Roche). RNA internal extraction control (Plasmodium falciparum PfMGET ivRNA) was added prior to extraction using the MagNAPure LC Total Nucleic Acid—High Performance kits (Roche). RT-qPCR was performed using the Luna Universal Probe One-Step RTqPCR kit (NEB) with 400 nM E-gene primers (FW: 5’- ACAGGTACGTTAATAGTTAATAGCGT-3’ RV: 5’- ATATTGCAGCAGTACGCACACA-3’) and 200 nM E-gene probe (5’-FAM ACACTAGCCATCCTTACTGCGCTTCG-BHQ1-3’ (Biolegio)) on a CFX96 C1000 Real-Time PCR etection System (BioRad). Transcript quantities were calculated using a tenfold dilution series of E gene ivRNA. The extraction efficiency was checked in a separate RT-qPCR using the Luna Universal Probe One-Step RT-qPCR kit (NEB) with primers targeting PfMGET ivRNA.
Statistical analyses
Analyses were conducted using SPSS software, 27th version (IBM Corp., 2021) and GraphPad Prism, version 8.0.2 (GraphPad Software, 2019). Categorical data are presented as numbers with percentages and continuous data are presented as medians with interquartile ranges. Mann–Whitney U tests were used to assess differences between groups. Spearman rank correlation was used to assess the correlation between SARS-CoV-2 RNA loads in saliva and nasopharyngeal throat swabs.
Results
Patient characteristics
Forty-nine patients (64.4% male) with a median age of 68 years (IQR 52–75) (Table 1) were included. Patients were admitted up to 21 days after symptom onset (median 8, IQR 5–10) and included at a median of 2 days (IQR 2–3.5 days) after admission. Forty-four patients (89.9%) had one or more comorbidities, with cardiovascular diseases including hypertension in 28 (63.3%) and pulmonary diseases including COPD in 16 (36.7%) being the most common. Six patients (12.2%) used immunosuppressive medication prior to SARS-CoV-2 infection.
The majority of patients (87.8%, 43/49) received immunomodulatory treatment for COVID-19 in the form of corticosteroid treatment. Thirty-six patients (73.5%) showed clinical improvement and were discharged, 8 patients (16.3%) were transferred to a rehabilitation center, and 5 patients (10.2%) died within the hospital. Median length of admission was 7 days (ICR 4–12.5).
Correlation of viral RNA load in saliva and nasopharyngeal throat swabs
Out of 30 patients from whom we collected saliva samples, 27 (90%) had detectable SARS-CoV-2 RNA in both saliva and the combined nasopharyngeal-throat swab. SARS-CoV-2 RNA loads in saliva and the combined nasopharyngeal throat swab were correlated (rs = 0.566, p = 0.002) (Fig. 1b).
Detection of SARS-CoV-2 RNA in exhaled air
SARS-CoV-2 RNA was detected in exhaled air in 12 (24.5%) patients (Fig. 1c) up to 23 days after symptom onset. Patients with and without detectable SARS-CoV-2 RNA in exhaled air did not differ in age, BMI, time from symptom onset to admission, length of admission, or laboratory measures (Table 1). The median time from symptom onset to sampling was 7.5 days (IQR 6–14.3) in patients with detectable SARS-CoV-2 RNA versus 11 days (IQR 8.5–13) in patients without detectable SARS-CoV-2 RNA (p = 0.295). Interestingly, the presence of SARS-CoV-2 RNA in exhaled air was not limited to patients with high SARS-CoV-2 viral load in the combined nasopharyngeal throat swab. Furthermore, patients with and without detectable SARS-CoV-2 RNA in exhaled air showed no differences in SARS-CoV-2 RNA load in the combined nasopharyngeal throat swab (p = 0.335) or saliva (p = 0.938, Supplementary Fig. 1).
Discussion
In this study, we showed for the first time that SARS-CoV-2 RNA can be detected by RT-qPCR in exhaled air of hospitalized COVID-19 patients. SARS-CoV-2 RNA has previously been detected in air collected from COVID-19 wards13 and in exhaled air from ambulant COVID-19 patients14, but not yet from hospitalized patients. In our study, sampling of exhaled air was feasible using a handheld modular breath sampler and viral RNA was detected in almost 25% of the patients.
We demonstrated that viral RNA is still detectable 7 days after disease onset in a clinical setting, later than Coleman et al.14 who detected SARS-CoV-2 RNA after a median of 3 days of symptoms in an ambulant setting. Their positivity rate was higher, which can probably also be attributed to a longer sampling time of 30 min. It must be noted that the two-minute breathing exercise was well tolerated by all patients in our study, including patients receiving oxygen supplementation via nasal cannula as well as patients using high-flow oxygen therapy. The sampling could be prolonged to increase sensitivity, however, increased sampling time will reduce the usability of the modular breath sampler and hamper its clinical implication.
Viral load in nasopharyngeal throat swab and saliva samples was positively correlated, confirming previous observations both in inpatients with confirmed COVID-1915 and in routine sampling16. Sampling saliva can be a patient-friendly alternative to nasopharyngeal throat swab sampling, but sensitivity can be an issue, as shown by the lower viral loads in saliva.
The fact that SARS-CoV-2 RNA was measured up to 23 days after symptom onset suggests long-term persistence of viral RNA. Moreover, the absence of any association between SARS-CoV-2 positivity in exhaled air samples and viral load in nasopharyngeal throat swab and saliva samples makes contamination from the upper airway unlikely.
This study shows the feasibility of sampling exhaled air from hospitalized COVID-19 patients for the presence of SARS-CoV-2 RNA. The modular breath sampler could be used in epidemiological studies to determine the best proxy of SARS-CoV-2 contagiousness.
Data availability
The data that support the findings of this study are available from the corresponding author, upon reasonable request.
References
Wiersinga, W. J., Rhodes, A., Cheng, A. C., Peacock, S. J. & Prescott, H. C. Pathophysiology, transmission, diagnosis, and treatment of coronavirus disease 2019 (COVID-19): A review. JAMA 324, 782–793. https://doi.org/10.1001/jama.2020.12839 (2020).
Knight, V. Viruses as agents of airborne contagion. Ann. N. Y. Acad. Sci. 353, 147–156. https://doi.org/10.1111/j.1749-6632.1980.tb18917.x (1980).
Tang, S. et al. Aerosol transmission of SARS-CoV-2? Evidence, prevention and control. Environ. Int. 144, 106039. https://doi.org/10.1016/j.envint.2020.106039 (2020).
Somsen, G. A., van Rijn, C., Kooij, S., Bem, R. A. & Bonn, D. Small droplet aerosols in poorly ventilated spaces and SARS-CoV-2 transmission. Lancet Respir. Med. 8, 658–659. https://doi.org/10.1016/S2213-2600(20)30245-9 (2020).
van Doremalen, N. et al. Aerosol and surface stability of SARS-CoV-2 as compared with SARS-CoV-1. N. Engl. J. Med. 382, 1564–1567. https://doi.org/10.1056/NEJMc2004973 (2020).
Richard, M. et al. SARS-CoV-2 is transmitted via contact and via the air between ferrets. Nat Commun. 11, 3496. https://doi.org/10.1038/s41467-020-17367-2 (2020).
Lakdawala, S. S. & Menachery, V. D. Catch Me if You Can: Superspreading of COVID-19. Trends Microbiol. 29, 919–929. https://doi.org/10.1016/j.tim.2021.05.002 (2021).
Port, J. R. et al. Increased small particle aerosol transmission of compared with SARS-CoV-2 lineage A in vivo. Nat. Microbiol. 7, 213–223. https://doi.org/10.1101/2021.07.26.453518 (2022).
Riediker, M. et al. Higher viral load and infectivity increase risk of aerosol transmission for Delta and Omicron variants of SARS-CoV-2. Swiss Med. Wkly. 152, 30133. https://doi.org/10.4414/smw.2022.w30133 (2022).
Zheng, J. et al. High amounts of SARS-CoV-2 in aerosols exhaled by patients with Omicron variant infection. J. Infect. https://doi.org/10.1016/j.jinf.2022.02.015 (2022).
Tellier, R. et al. Recognition of aerosol transmission of infectious agents: a commentary. BMC Infect. Dis. 19, 101. https://doi.org/10.1186/s12879-019-3707-y (2019).
Rijksinstituut voor Volksgezondheid en Milieu. Afnametechniek specifieke virale diagnostiek (COVID-19, influenza). https://lci.rivm.nl/bijlage/afnametechniek (2020).
Liu, Y. et al. Aerodynamic analysis of SARS-CoV-2 in two Wuhan hospitals. Nature 582, 557–560. https://doi.org/10.1038/s41586-020-2271-3 (2020).
Coleman, K. K. et al. Viral load of SARS-CoV-2 in respiratory aerosols emitted by COVID-19 patients while breathing, talking, and singing. Clin. Infect Dis. https://doi.org/10.1093/cid/ciab691 (2021).
Wyllie, A. L. et al. Saliva or nasopharyngeal swab specimens for detection of SARS-CoV-2. N. Engl. J. Med. 383, 1283–1286. https://doi.org/10.1056/NEJMc2016359 (2020).
de Oliveira, C. M., Brochi, L., Scarpelli, L. C., Lopes, A. C. W. & Levi, J. E. SARS-CoV-2 saliva testing is a useful tool for Covid-19 diagnosis. J. Virol. Methods. 296, 114241. https://doi.org/10.1016/j.jviromet.2021.114241 (2021).
Funding
AvL and RvC are supported by the National Institute of Health for the project “Using Tryptophan Metabolism and Response to Corticosteroids to Define New Therapeutic Targets for Tuberculosis Meningitis: Integration of Large Scale Clinical, Metabolomic, and Genomic Data” [R01AI145781]. GJO and RPvR are supported by a VICI grant [16.VICI.170.090] from the Dutch Research Council (NWO). MGN is supported by an ERC Advanced Grant [#833247] and a Spinoza Grant of the Netherlands Organization for Scientific Research. MIdJ is supported by the TARGET grant, funded by ZonMW, the funding organization for medical sciences within NWO [549009004].
Author information
Authors and Affiliations
Contributions
All authors contributed to the conception of the work and interpretation of the data. Clinical data were acquired by L.K. RT-qPCR was performed by K.L. under supervision of T.B. L.K. conducted statistical analyses and prepared graphical visualisation of the data. L.K., A.v.L., and M.I.d.J. wrote the main manuscript text. All authors have read and revised the paper and approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
LK, KL, DAD, GJO, MGN, JtO, RvC, KMM, FLvdV, HW, JD, RJL, RPvR, TB, and AvL declare no competing interests. CHvdK and MIdJ are involved in Xheal Diagnostics that aims to exploit the presented modular breath sampler.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kurver, L., van den Kieboom, C.H., Lanke, K. et al. SARS-CoV-2 RNA in exhaled air of hospitalized COVID-19 patients. Sci Rep 12, 8991 (2022). https://doi.org/10.1038/s41598-022-13008-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-13008-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.