Abstract
Phosphatidylinositol 4-kinases (PI4Ks) are the first enzymes that commit phosphatidylinositol into the phosphoinositide pathway. Here, we show that Arabidopsis thaliana seedlings deficient in PI4Kβ1 and β2 have several developmental defects including shorter roots and unfinished cytokinesis. The pi4kβ1β2 double mutant was insensitive to exogenous auxin concerning inhibition of root length and cell elongation; it also responded more slowly to gravistimulation. The pi4kß1ß2 root transcriptome displayed some similarities to a wild type plant response to auxin. Yet, not all the genes displayed such a constitutive auxin-like response. Besides, most assessed genes did not respond to exogenous auxin. This is consistent with data with the transcriptional reporter DR5-GUS. The content of bioactive auxin in the pi4kß1ß2 roots was similar to that in wild-type ones. Yet, an enhanced auxin-conjugating activity was detected and the auxin level reporter DII-VENUS did not respond to exogenous auxin in pi4kß1ß2 mutant. The mutant exhibited altered subcellular trafficking behavior including the trapping of PIN-FORMED 2 protein in rapidly moving vesicles. Bigger and less fragmented vacuoles were observed in pi4kß1ß2 roots when compared to the wild type. Furthermore, the actin filament web of the pi4kß1ß2 double mutant was less dense than in wild-type seedling roots, and less prone to rebuilding after treatment with latrunculin B. A mechanistic model is proposed in which an altered PI4K activity leads to actin filament disorganization, changes in vesicle trafficking, and altered auxin homeostasis and response resulting in a pleiotropic root phenotypes.
Similar content being viewed by others
Introduction
Plant health and productivity depends on root outgrowth, which allows water and nutrient uptake, and is equally crucial for an efficient photosynthesis rate1,2. Root morphogenesis is a complex process, orchestrated by a complex signaling crosstalk at different levels, from single-cell metabolism to hormone transport within plant organs. On-point spatial and temporal organization of cell organelles, polar establishment of cell architecture and directed shoot ward auxin transport are fundamental for correct root cell differentiation. Root hair cell priming and plasticity require fine-tuned, interconnected cellular processes driven by a properly established cytoskeleton that controls the polar delivery of membranes to the root apex in order to enlarge the cell unidirectionally, and by the transport of auxin through the root tip1,2. Auxin regulates cell polarity through the activation of ROPs (Rho-like GTPase) that participate in the polar localization of PIN-FORMED (PIN) family proteins. Carriers of the PIN family are plasma membrane-integrated auxin efflux carriers responsible for the direction and intensity of auxin flow through the plant body. Their cellular localization and activity are regulated at many levels3,4,5, and rely on the lipid composition of the membrane they are in.
Phosphoinositides, minor components of plasma membrane, are phosphorylated derivatives of phosphatidylinositol (PI), such as phosphatidylinositol-4-phosphate (PI4P) and phosphatidylinositol-4,5-bisphosphate (PI4,5P2). Phosphoinositides are important signaling molecules as they are substrates or cofactors of important signaling enzymes. In plants, both PI4P and PI4,5P2 can be substrates to phospholipases C (PLCs) leading to diacylglycerol and the corresponding phosphorylated inositol. PI4,5P2 is a cofactor of some phospholipases D (PLDs), that catalyze the production of phosphatidic acid, a major plant signaling lipid6. More generally, phosphoinositides can directly interact with membrane proteins (such as ion channels or G protein-coupled receptors) or cytosolic proteins that they can recruit to membranes7,8. Interestingly, specific relative levels of phosphoinositides are a characteristic feature of different membranes: plasma membrane, endoplasmic reticulum and Golgi membranes do not have the same relative composition in phosphoinositides7,8,9. Besides, membrane nanoclusters enriched in certain proteins crucial for signal transduction and transport proteins also have a specific composition in phosphoinositides8,10. Formation of membrane domains enriched in PI4P and PI4,5P2 is a crucial component of plasma membrane dynamics. Such phosphoinositide-enriched domains are important for the localization of REMORINs, scaffold proteins governing PM–bound signaling11, and FLS2, a pattern-recognition receptor that determines the specific perception of the bacterial protein flagellin12.
Composition in phosphoinositides is modified by the activities of lipid kinases. PI4Ks phosphorylate the 4th hydroxyl position in the inositol head group of PI to generate PI4P. PI4P can be further phosphorylated by phosphatidylinositol-4,5-kinases (PI4,5 K) into PI4,5P2. There are two types of PI4Ks according to their primary sequences and pharmacological sensitivities. Type-II PI4Ks are inhibited by adenosine while type III PI4Ks are inhibited by micromolar concentrations of wortmannin, a steroid produced by the fungi Penicillium funiculosum. In the A. thaliana genome, twelve putative PI4K isoforms have been identified. Eight belong to type-II (AtPI4Kγ1-8), and four belong to type-III (AtPI4Kα1 and α2 and AtPI4Kβ1 and β2)13. Not much is known about type-II PI4Ks and they could actually be protein kinases and not lipid kinases13,14. We have previously shown that type-III PI4Ks are upstream of the PLC activity that controls the responses of tobacco BY2 cells to cryptogein, a fungi elicitor15. Type-III PI4Ks are also upstream of plant cold response PLC activity16 but also of the PLC activity that controls gene expression, in basal, non-stimulated conditions17. Type-III PI4Ks have been shown also to be activated in response to salicylic acid (SA), and the consequent increase in phosphoinositides is an important part of the specific response of Arabidopsis to this phytohormone18,19,20. AtPI4Kα2 is a pseudogene and viable homozygous PI4Kα1 mutants have never been obtained. We have worked previously on a double mutant defective in both PI4Kβ genes. In 4-week-old plants, pi4kβ1β2 exhibited a constitutively high SA level that led to a stunted phenotype21. However, SA accumulation did not occur in young pi4kβ1β2 seedlings21,22 and therefore, they appeared to be the material of choice to study the roles of PI4Ks and phosphoinositides in root development. Several aspects of the role of PI4Ks in plant cell biology have been discovered using pi4kß1ß2 double mutant, such as the involvement of PI4Kβ1 in cell plate formation during cytokinesis23, in the formation of secretory vesicles24, root hair shaping and polar growth25. Here, we show that the pi4kß1ß2 double mutant exhibits several root phenotypes: impaired primary root growth, lower sensitivity to exogenous indole-3-acetic acid (IAA), impaired elongation and bending in response to gravistimulation, and misshapen root hair growth. These changes appeared to coincide with an altered subcellular distribution and turnover of PIN2, a less stable actin cytoskeleton and generally altered intracellular trafficking dynamics. Moreover, expression of several auxin-associated genes in roots was not responsive to exogenous auxin. However, some transcript levels in non-treated mutant roots were already at the auxin-response levels of wild type (WT) roots, thus displaying an apparent auxin-like response. Remarkably, the measured content of bioactive IAA in double mutant roots did not differ from that of WT, but a considerable increase of glutamate-conjugated form of auxin was monitored. Besides, the auxin level reporter DII-VENUS did not respond to exogenous auxin in pi4kß1ß2 mutant. Our data, therefore, link altered PI4K activity to the modification of vesicular trafficking and actin filaments organization one the one hand to altered auxin response likely due to alteration in auxin homeostasis one the other hand.
Results
The pi4kß1ß2 mutant is impaired in root growth
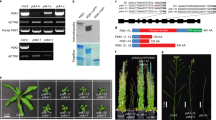
The PI4Kß1ß2 deficiency in pi4kß1ß2 seedlings led to a decreased primary root length of up to fourfold compared to the WT control (Fig. 1a,b). The shorter primary root of the mutant appeared to be due to shorter meristem and elongation zones (Fig. 1c). The shorter meristem of pi4kß1ß2 was due to fewer cells (Fig. 1d), some of which showed unfinished cytokinesis. Interestingly, the CycB1::GUS associated signal occupied a smaller percentage area of the meristem in pi4kß1ß2 roots when compared to the WT (Fig. 1e,f). The elongation zone was almost missing. In the differentiation zone, the pi4kß1ß2 mutant had smaller cortical cells (Supplementary Fig. S1) and either similar or very small root hair lengths when compared to the WT. This created apparent “bald-like” zones (Fig. 1g,h), while the overall total root hair density in pi4kß1ß2 plants did not differ to that of the WT (Fig. 1i). An analysis of the epidermal cell lines26 showed that the regularity of trichoblasts/atrichoblasts formation was not affected in the mutant (Supplementary Fig. S2). This confirmed that the apparent "bald-like zones'' were not due to an absence of hairs but to shorter root hairs.
Impaired root growth and morphological characteristics of the pi4kß1ß2 mutant. (a) representative pictures of the apical root parts of 11-day-old seedlings of Arabidopsis thaliana Col-0 (WT) and the pi4kß1ß2 mutant: meristem (M), elongation zone (EZ) and differentiation zone (DZ) are marked, scale bar = 100 μm; (b) primary root length, n = 40; (c) length of the meristematic and elongation zones, n = 12, error bars represent mean ± SEM; different letters indicate statistically significant groups, one-way ANOVA with Tukey-HSD post-hoc test (p > 0.05); (d) number of separated cells in the meristem, n = 36; (e) representative images of GUS staining in the root meristem of 4-day-old plants expressing CycB1::GUS, scale bar = 100 μm; (f) relative area of CycB1::GUS expression, % of the meristematic zone; n = 72; (g) representative images of root hair distribution in the DZ of roots, scale bar = 100 μm; (h) root hair length, n = 180; (i) root hair density, n = 90. Central line of the boxplots represents the median, plus represents the mean, circles represent individual values from three biological repeats. p-value was calculated by Student t-test.
Responses to IAA and to gravistimulation are impaired in pi4kß1ß2
Mutant and WT 5-day-old seedlings were transferred to a cultivation medium containing various phytohormones. Seven days later, the lengths of the primary root, of the meristem and of the cortical cells within the differentiation zone were measured. The presence of IAA led to a decrease in the root length of WT plants; the decrease was more than 60% at 100 nM IAA. The pi4kß1ß2 mutant was less sensitive to the auxin treatment, the decrease being only 20% at the concentrations tested (Fig. 2a, Supplementary Fig. S3). A lower sensitivity to exogenous auxin was also detected at the cellular and/or tissue levels. At 50 nM IAA, the length of WT cortical cells showed a 30% decrease compared to the control, while the mutant was insensitive. At 1 µM IAA, the decrease in length of WT cortical cells was 50%, compared to the control, while the mutant remained insensitive (Supplementary Fig. S4a). Concerning meristem size, 100 nM IAA caused a 20% shortening of its length in WT seedlings but no response was observed for the pi4kß1ß2 mutant; this difference in IAA sensitivity was still apparent even at 1 µM (Supplementary Fig. S4b). Interestingly, the sensitivity of primary root length to a cytokinin (BAP) or to salicylic acid (SA) did not differ between pi4kß1ß2 and WT seedlings (Supplementary Fig. S4c,d), thus indicating a specific response to auxins.
Auxin-related phenotypes of the pi4kß1ß2 mutant. (a) primary root length of 11-day-old seedlings in response to different IAA concentrations, n = 22. Central line of the boxplots represents the median, circles represent individual values; p-value is indicated for significantly different groups; t-test with correction for multiple comparisons; (b) Elongation rate of primary root under gravistimulation, n = 10; (c) root tip orientation angle, n = 10; (b,c) gravitropic assay, 5-day-old seedlings were rotated to 90° on a horizontal microscope, images were taken every hour. Asterisks indicate statistically significant differences between genotypes, p < 0.05, paired t-test with correction for unequal variances. Experiments were repeated three times; data from a representative repeat are shown.
We then focused on another auxin-related process, the response to gravistimulation. Interestingly, both root elongation (i.e. the distance that the root tip grew since the 0’ time-point) and root orientation (i.e. the angle between the root tip at current and 0’ time-point) were affected in the double mutant in due course of 12 h experiment (Fig. 2b,c Supplementary Fig. S5, Supplementary movie SM1 for WT, Supplementary movie SM2 for pi4kß1ß2).
The transcriptome of pi4kß1ß2 roots shows partial similarities to IAA-treated WT roots
In order to better detail the pi4kß1ß2 root phenotypes, an RNAseq transcriptomic analysis of roots was performed (Supplementary table S2). It was found that 2517 and 3418 genes were either up- or down-regulated, respectively, in pi4kß1ß2 roots compared to WT roots. To be more stringent, we then only considered the genes passing a threshold of log2-fold change of 1.5. On these genes we performed a Gene Ontology classification (Supplementary Fig. S6). Among the genes induced in pi4kß1ß2 roots compared to WT, we found enrichment in genes encoding extracellular, plasma membrane, or cell wall localized proteins, and underrepresentation of genes encoding cytoskeleton or mitochondria-associated proteins. Interestingly, among the repressed genes, the cell wall-associated proteins were also enriched, while cytoskeleton-localized proteins were overrepresented. As for biological processes, we found enrichment in the categories of “response to stress”, “signal transduction”and “development” for both groups of genes. Results of the RNAseq analysis were confirmed by qPCR on a selection of genes (Supplementary Fig. S7). Among the genes most induced in pi4kß1ß2 roots, we found several that were involved in response to hypoxia, oxidative stress and induced systemic resistance (Supplementary Fig. S8). A focus on genes involved in auxin transport or metabolism is displayed (Supplementary table S4). GH3.12 (AT5G13320) and GH3.3 (AT2G23170) are markedly up-regulated in pi4kß1ß2 roots compared to WT ones.
Next, the list of the 200 most up-regulated and 200 most down-regulated genes in pi4kß1ß2 mutant roots versus WT roots was used as a signature to interrogate public transcriptomic data using the Genevestigator similarity search program25. This was performed against curated root experiments dealing with root samples and classified as “Hormone”, “Temperature” or “Stress”. Out of the 10 most similar experiments, 7 concerned treatments with auxin (Fig. 3a). Within this set of curated root experiments (Fig. 3a), we then only selected the experiments dealing with response to auxins. According to the responses in these experiments of the 200 most repressed genes in our pi4kß1ß2 versus WT root comparison, the experiments and the genes were clustered (Fig. 3b). This allowed the identification of clusters of genes, down-regulated in pi4kß1ß2 mutant roots compared to WT ones and down-regulated in some experiments dealing with the response to auxin (Fig. 3b, clusters A,B,C; list of genes of these clusters in supplementary table 3). We did the same with the 200 most up-regulated genes in the pi4kß1ß2 double mutant compared to WT roots and thus identified genes upregulated both in pi4kß1ß2 mutant roots versus WT and up-regulated in curated experiments dealing with response to auxin in roots (Fig. 3c, cluster E). These clusters represent genes for which the effect of the pi4kß1ß2 double mutation in the root compared to WT is similar to a treatment with auxin. Yet other clusters exist, consisting of genes that are down-regulated in pi4kß1ß2 roots, but were shown to be upregulated by auxins in public transcriptomics data (Fig. 3b, cluster D; supplementary table S3), or genes that are up-regulated in pi4kß1ß2 roots, but were shown to be upregulated by auxins in public transcriptomics data (Fig. 3c, cluster F; supplementary table S3). The transcript levels of selected auxin responsive genes representing different clusters were monitored by qPCR in mutant and WT plants, treated or not with 10 nM IAA for 24 h (Fig. 3d). The transcript level of AT1G64590, CSLB5, SAUR9, NPF2.4 and BRU6 in the untreated roots of pi4kß1ß2 mutant was similar to that in WT roots treated with auxins. On the other hand, the transcription of CSLB5, FLA13 and BRU6 did not change in response to auxin in the pi4kß1ß2 mutant, showing another evidence of affected auxin response.
Transcriptomic analysis of pi4kß1ß2 roots. (a) Similarity between the pi4kß1ß2 roots transcriptome (compared to WT) and the stress-, hormone- or temperature- responsive transcriptomes. The 200 genes most up-regulated in pi4kß1ß2 roots compared to the WT and the 200 genes most down-regulated in pi4kß1ß2 roots compared to the WT were used as a signature to search for transcriptome experiments with the highest similarity. The similarity search was performed against the 56 root experiments classified as “stress”, “temperature” or “hormone” by Genevestigator (Hruz et al., 2008). Experiments were sorted according to Euclidean distance. Expression of the signature genes in the 10 most similar experiments are shown in color-scale; (b,c) Hierarchical clustering of curated root experiments dealing with the response to auxins. The 9 curated root experiments dealing with auxins in Genevestigator were retrieved. According to the expression in these experiments of the 200 most down-regulated (b) genes in our pi4kß1ß2 vs. WT root comparison, the genes and experiments were clustered with the Biclustering tool in Genvestigator. The same was done using the 200 most up-regulated (c) genes in our pi4kß1ß2 vs. WT root comparison. Similarities between expression profiles were determined using Pearson correlation. For each experiment, the duration of hormone treatment is indicated. Separated gene clusters with highest levels of induction/repression are labeled and genes are specified on the right panel; (d) Response of selected genes to auxin. Five-day-old seedlings were transferred to a medium containing 10 nM IAA, and roots for RNA extraction were harvested after 24 h. The data are presented in means ± SE, n = 9, with a Tukey honestly significant difference (HSD) multiple mean comparison post hoc test. Different letters indicate a significant difference (one-way ANOVA, Tukey HSD, p-value < 0.05).
Assessing auxin sensitivity of pi4kß1ß2 roots
We next checked auxin transcriptional response by a reporter system, introducing by crossing the auxin sensitive synthetic promoter DR527 fused to a GUS reporter gene into pi4kß1ß2 background. Surprisingly, the basal level of DR5 promoter activity was lower in root and leaf meristem of the pi4kß1ß2 plants (Fig. 4a,b). After exposure to 10 nM IAA, an important increase of DR5-GUS signal was detected in WT meristems, but not in the pi4kß1ß2 mutant (Fig. 4a,b), confirming that the sensitivity to IAA is impaired in the mutant line.
Auxin sensitivity of the pi4kß1ß2 mutant. (a) representative images of DR5-GUS activity in 5-day-old root and cotyledons in the presence or not of 0.01 µM IAA for 12 h, scale bar = 100 µm; (b) DR5-GUS quantification, % of GUS-stained area in root meristem, n = 10; (c) representative images of DII-VENUS fluorescence in root tip of 7-day-old seedlings in the presence or not of 0.01 µM IAA for 1 h, maximum intensity Z-projections of 10 nm stacks, scale bar = 50 µm; (d) DII-VENUS fluorescence quantification, % of meristematic zone; n = 10; (e) quantitation of IAA metabolites and conjugates in 7-day-old roots, n = 6; Central line of the boxplots represents the median, circles represent individual values; p-value is indicated for significantly different groups, ns – non significant; unpaired t-test (d,e); The data are presented in means ± SD, n = 10, with a Tukey honestly significant difference (HSD) multiple mean comparison post hoc test. Different letters indicate a significant difference (one-way ANOVA, Tukey HSD, P < 0.05) (b); Experiments were repeated three times; data from a representative repeat are shown. IAA = indole-3-acetic acid, IAA-Asp = IAA-aspartate, IAA-Glu = IAA-glutamate, CamX = camalexin, IAA-GE = IAA-glucose ester, OxIAA = oxo-IAA, IAM = Indole-3-acetamide (IAA precursor), OxIAA-GE = oxo-IAA-glucose ester, I3A = indole-3-aldehyde, IAN = Indole-3-acetonitrile (IAA precursor), OxIAA-Asp = oxo-IAA-aspartate.
The DII-VENUS28 construct was introduced into the pi4kß1ß2 mutant by floral-dip agrobacterium transformation. DII-VENUS is a fast maturing form of a yellow fluorescent protein fused in-frame to the Aux/IAA-interaction domain (termed domain II;) and it is rapidly degraded in response to auxin28. It is used as a reporter of auxin level. As the DII-VENUS reporter was introduced by agrobacterium transformation, the potential positional effect of the insert cannot be excluded, so the basal fluorescent signal cannot be compared between the lines but signals can be compared within one line. After exposure to 10 nM IAA, a significant decrease of DII-VENUS fluorescence signal was detected in WT plants, but not in the pi4kß1ß2 mutant (Fig. 4c,d). To check whether the mutant insensitivity to IAA might be a consequence of an elevated IAA level in control conditions, we extracted hormones from the total root system and measured the content of IAA metabolites and conjugates. No difference in the measured free IAA content was detected between genotypes, while IAA-Glu, CamX, I3A and IAN concentrations were higher in the pi4kß1ß2 mutant than in the wild type (Fig. 4e).
Localization of auxin efflux transporter PIN2 is altered in the pi4kß1ß2 mutant
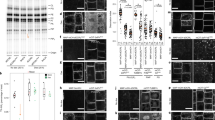
As auxin signaling is relying on the correct auxin transport between and within the cells, we investigated the localization and dynamics of auxin transporter PIN2. We analyzed plants expressing PIN2::PIN2-GFP by immunostaining (Fig. 5a–e) and confocal microscopy of PIN2-GFP in both WT and pi4kß1ß2 backgrounds (Fig. 5f–k). Overall, PIN2-GFP was distributed on the plasma membrane in the same cell types and with a similar polar distribution in mutant roots compared to WT roots. However, in the pi4kß1ß2 mutant, several “black holes” in the signal were detected along the plasma membrane (Fig. 5b–e, Supplementary Fig. S9). When counterstained with FM 4–64, a dye that labels the plasma membrane, it was seen that the unstained parts in the pi4kß1ß2 roots corresponded to tunnels between adjacent cells (Fig. 5c–e). Confocal microscopy color-coded projections of pictures were taken over time to track PIN2-GFP intracellular movement in the meristematic zone. The chaotic distribution of vesicles in pi4kß1ß2 compared to the vesicles aligned in WT showed not only differences in the amount of GFP-marked intracellular vesicles, but also that their movement was less rectilinear and very fast in pi4kß1ß2 (compare Fig. 5f–h, where vesicles are indicated by white arrows, and the corresponding Supplementary movies SM3 for WT and SM4 for pi4kß1ß2). Differences in vacuolar morphology were also observed in pi4kß1ß2 (Fig. 5i,j; Supplementary Fig. S10), with bigger and less fragmented vacuoles than the WT. When focused on growing root hair cells, altered movement of fluorescent marked vesicles in mature root hair cells and elongating root hairs in pi4kß1ß2 PIN2::PIN2-GFP was observed (Supplementary movies SM5 for WT and SM6 for pi4kß1ß2). Bright field imaging also revealed differences in the flow of cytoplasmic streaming. Circulation of the cytoplasmic stream occurred close to the plasma membrane and in a straight path in the WT, whereas in the mutant stream the stream flowed in less coordinated lanes. We then studied the response to a dark shift of whole seedlings, a treatment known to enhance PIN2-GFP delivery to the lytic vacuole26. A 1 h dark shift caused the translocation of PIN2-GFP to lytic vacuoles in WT roots but not in the double mutant (Fig. 5k,l). All these results point to altered intracellular trafficking dynamics in the roots of pi4kß1ß2 seedlings.
Visualization of PIN2-GFP subcellular distribution by confocal microscopy. (a) distribution of PIN2 along the PM in WT roots, immunostaining; (b) distribution of PIN2-GFP along the plasma membrane in pi4kß1ß2 roots, immunostaining; (c,d,e) show PIN2-GFP signal overlapping with FM4-64 dye, (c, FM4-64; d, PIN2-GFP; e, merged signals); (f,g) color-coded projection of PIN2-GFP distribution and intracellular movement over time in (f) WT and (g) pi4kß1ß2 backgrounds; arrows point to vesicles moving in time; (h), zoomed part of f and g, scale bars 5 μm, arrows point to vesicles moving in time; (i,j) merged 3D reconstruction of pictures taken along the z-axis of the bright field and fluorescent channel of PIN2 distribution along the plasma membrane and vacuole morphology in (i) WT and (j) pi4kß1ß2 backgrounds; arrows point to enlarged vacuoles in pi4kß1ß2; (k) visualization of PIN2-GFP movement towards the lytic vacuole upon a dark shift of whole seedlings. After 1 h, the GFP signal was visible in the WT background, but not in pi4kß1ß2; (l) quantification of the GFP signal intensity in the lytic vacuole, each circle represents the plasma membrane/intracellular ratio for a single cell; p-value is indicated for significantly different groups, ns—non significant; unpaired t-test with correction for multiple comparisons; n = 25; scale bars 10 μm.
Actin stability and remodeling are affected in the pi4kß1ß2 mutant
Five-day-old pi4kß1ß2 seedlings expressing 35S::LifeAct-GFP were sprayed with 10 µM latB, a drug that inhibits actin polymerization. Treated seedlings were then observed under a confocal microscope (Fig. 6a). Without a latB treatment, the fluorescence signal occupancy was lower in pi4kß1ß2 compared to WT seedlings. After a 90 min exposure to latB, the fluorescence signal occupancy in pi4kß1ß2 decreased 40%, while no change was detected in WT plants (Fig. 6b). After a 150 min of exposure to latB, the signal occupancy in WT showed a 35% decrease compared to the control, while the occupancy decreased to 54% for the pi4kß1ß2 mutant compared to control conditions. Interestingly, while WT roots showed a gradual decrease in actin filament bundling (Fig. 6c) in due course of latB treatment, no significant changes were observed in the pi4kß1ß2 double mutant.
Actin reorganization in the pi4kß1ß2 mutant in response to latrunculin B. Five-day-old seedlings expressing 35S::LifeAct-GFP were sprayed with 10 µM latB. (a) representative maximum intensity projections of root epidermis of WT and pi4kß1ß2 plants; confocal microscopy, scale bar = 10 μm; (b) quantitative analysis of the density (expressed as percentage of occupancy) of actin filament arrays in epidermal cells; (c) quantitative analysis of the extent of filament bundling (expressed as skewness) in epidermal cells. Central line of the boxplots represents the median, plus represents the mean; circles represent individual values; p-value is indicated for significantly different time points within each genotype and for the comparison of genotypes immediately after treatment; one-way ANOVA with Tukey HSD post-hoc test; n = 10.
Discussion
Here, we show that PI4Kß1ß2 deficiency led to up to a fourfold decrease of primary root length compared to WT seedlings. A dwarf phenotype, both in the roots and aerial parts, has already been reported for the pi4kß1ß2 mutant21. Notably, the small rosette size of 4-week-old pi4kß1ß2 mutant plants has been linked to an increased constitutive SA level21. Indeed, a pi4kß1ß2sid2 triple mutant did not accumulate SA and it did not display the stunted rosette phenotype. However, pi4kß1ß2sid2 seedlings still exhibited shorter roots than WT plants, thus showing that this root phenotype was a SA-independent process21,22. Furthermore, SA accumulation did not occur in young pi4kß1ß2 seedlings22, thereby confirming that the root length phenotype was not due to high SA levels. Similar SA levels in pi4kß1ß2 and WT roots were found in this work (Supplementary Fig. S11), thus confirming that the observed root phenotype was not related to altered SA levels and therefore it was an SA-independent process.
So what causes the short root phenotype of pi4kß1ß2 seedlings? To answer this question, a detailed analysis of root morphology was undertaken (Fig. 1). The shorter primary roots of the double mutant appeared to be due to a reduced meristematic zone due to a lower number of cells. The CycB1::GUS associated signal occupied a significantly smaller (about 10%) area of the meristematic zone in pi4kß1ß2 seedling roots when compared to the WT. This might explain in part why there were fewer cells in the meristematic zone of the mutant. An absent or a very short transition zone might also result from elevated auxin levels or an enhanced response to auxin. Indeed, the transition zone in a root begins where auxin levels attain a minimum28. The shorter primary root length in the pi4kß1ß2 double mutant was also associated with smaller cortical cells measured in the differentiation zone. A reduced meristematic zone in the pi4kß1ß2 mutant has been documented previously29, but we have supplemented these data by measuring meristem cell number and cell length in the differentiation zone. Concerning root hair length, we observed a bimodal distribution in the pi4kß1ß2 plants, with very short hairs that gave the impression of bald zones. Interestingly, the regularity of trichoblasts/atrichoblasts formation was not affected in the mutant.
Due to the observed root phenotypes, an obvious next step was to assess the sensitivity of the double mutant to different hormones known to alter root growth. Root sensitivity to BAP or SA did not differ between pi4kß1ß2 and WT seedlings. On the contrary, a loss of sensitivity in the double mutant to exogenous IAA was observed with respect to inhibition of primary root length, inhibition of cortical cell elongation, and elongation of the meristematic zone (Fig. 2, Supplementary Fig. S4a,b). This was in agreement with the experiments of Löfke et al., 2015, showing that altered vesicular trafficking due to inhibited PI4Kß1ß2 activity resulted in lower sensitivity to auxin NAA, altered vacuolar morphology and cell elongation30. Interestingly, pi4kß1ß2 double mutant was less efficient in response to gravistimulation, another auxin-related process. Notably, not only the root elongation, but also the root tip orientation towards gravity vector were impaired in the mutant, suggesting gravity sensing defects.
To detail the cellular processes that were interfering with the response to stimulation in mutant roots, we performed full transcriptomic analysis. We found that differences in gene expression between pi4kß1ß2 and WT roots were in part similar to those observed between auxin treated and non-treated WT roots. Yet, not all genes followed this trend and thus this similarity is only apparent. Besides, we tested by qPCR the response to auxin on a selection of genes, previously described as auxin responsive. The addition of exogenous IAA had no or only a small effect on the expression of those genes in pi4kß1ß2 compared to the WT. Similarly, DR5-GUS was not induced in the pi4kß1ß2 mutant after exposure to exogenous IAA. Besides, we also monitored a lower activity of the transcriptional reporter DR5-GUS in non-treated pi4kß1ß2 mutant meristems (in root tip or cotyledon tip) compared to wild type ones. Therefore, there seems to be a lower sensitivity to auxin, either exogenous or endogenous, as far as gene expression is concerned.
Why is the double mutant less sensitive to exogenous IAA? A possibility could be that the mutant is no longer responsive because of a constitutive high auxin level. As mentioned above, some of the observed root traits of the double mutant (such as a very short transition zone) already resemble an auxin response even in non-treated conditions. Yet, the DR5-GUS in control conditions does not plead for higher IAA content in the mutant. Besides, the level of free bioactive IAA was comparable between mutant and WT roots. The use of the DII-VENUS, a reporter directly related to the bioactive signal itself31, gives another block of valuable information. DII corresponds to the auxin binding domain of AUX/IAA ; when IAA binds to this domain, AUX/IAA proteins are released from ARF factors and they can interact with SCFTIR1 that ubiquitinylates them for degradation by the proteasome. Because the reporter was introduced by transformation in the pi4kß1ß2 mutant, we cannot directly compare data obtained in wild type to mutant background, but we can compare data within one line. In the mutant background, the DII-VENUS fluorescence was not significantly reduced by addition of exogenous IAA, as it was in WT seedlings. This is coherent with a non- or low- sensitivity to auxin we already documented based on gene expression data. The point is therefore to understand why DII-VENUS fluorescence is not reduced by addition of exogenous IAA. An explanation might be related to auxin homeostasis. In the pi4kß1ß2 mutant, we detected elevated levels of several inactive auxin metabolites including the glutamate-conjugate form. IAA-Glu is an early metabolite and storage form of IAA and is synthesized upon incubation of plants with high concentrations of IAA, and is considered precursors for auxin degradation32. High level of IAA-Glu might result from a constitutive conjugation activity in the double mutant. Transcripts of different GH3 enzymes involved in aminoacid conjugation are indeed markedly up-regulated in the pi4kß1ß2 roots. The affected balance between precursors or conjugates or IAA might then explain partial auxin-like response in roots. Conversely, this affected balance also probably reflects the affected sensitivity to active auxin. It is likely that in pi4kß1ß2 mutant, exogenous IAA is conjugated, resulting in the IAA level not changing, as seen by DII-Venus monitoring.
The pi4kß1ß2 mutant also showed an altered subcellular trafficking behaviour of PIN2, including trapping of the PIN2-GFP fusion protein in rapidly moving vesicles and a reduced transport towards the lytic vacuole upon a dark shift of pi4kß1ß2 seedlings. Differences in pi4kß1ß2 vacuolar morphology were also observed, with bigger and less fragmented vacuoles compared to the WT. This phenotype corresponds to that observed when WT Arabidopsis were treated with wortmannin, an inhibitor of PI4K activity33. In pi4kß1ß2 roots, PIN2 localization by immunostaining and staining with FM64 evidenced “black holes” or stubs corresponding to tunnels between adjacent cells, also referred to as “cell wall stubs”. This can be linked with unfinished cytokinesis23,34. Caillaud et al. (2008)35 demonstrated that map65-3/ple mutants displayed cell wall stubs and multiple nuclei in the root meristem, both features of cytokinesis-defective mutants. Interestingly, MAP65-3 is a downstream target for inhibition by MAP kinase MPK4, and also a physical interaction between PI4Kβ1 and MPK4 has been reported23. Lin et al., (2019)23 proposed that PI4Kβ1 and MPK4 influence localization and activity of MAP65-3, acting synergistically to control phragmoplast dynamics. The altered cytoskeleton organization in our mutant could explain some of the trafficking issues, as the movement of membrane vesicles depends on the cytoskeleton36. For example, it was shown that PIN1 cycling is actin-dependent37. Proper assembly of the cytoskeleton in concert with the molecular motors, myosins, is essential for active internal transport, and therefore proper distribution of cargos, like PIN21,38. Rho proteins mediate signals for cytoskeletal reorganization and cell polarity and they are also implicated in the regulation of endo- and exocytosis, and correct localization of PIN139. Moreover, a direct interaction between both PI4Kβ1 and PI4Kβ2 and another GTPase protein involved in membrane trafficking, RabA4B, has been reported40. Membrane recruitment of ROP-GTPase ROP6 (and possibly also other ROPs) is also dependent on anionic phospholipids41,42. We therefore suggest that an altered regulation of Rho proteins by phosphoinositides could help explain, in part, the observed problems of pi4kß1ß2 roots.
Can the alteration in trafficking and cytoskeleton explain the insensitivity to auxin? Proper spatial and temporal distribution of auxin in the root tip ensures differentiation of cell files and therefore conditions normal root hair outgrowth. This relies on an auxin efflux carrier modulating auxin circulation within root tip1. Therefore, altered trafficking can have consequences on root response to endogenous auxin. Concerning exogenously added auxin, the temporal distribution might not be the key factor and as mentioned above, auxin homeostasis probably explain the phenotypes.
Similar root phenotypes have been observed already in mutants affected in other steps of phosphoinositide turn-over. The pip5k1pi5k2 mutant lacking two isoforms of PI4,5 K also showed shorter roots, a reduced meristematic zone and a lower sensitivity to exogenous auxins43. The pip5k2 mutant has less lateral root formation and impaired gravitropism43,44. On the contrary, our pi4kß1ß2 mutant has more lateral roots45. We can thus speculate that some of the observed phenotypes of pi4kß1ß2 seedlings are in part due to altered production of PI4,5P2. As mentioned earlier, pi4kß1ß2 roots did not respond to a gravitropic stimulus, which is dependent on fine-tuned spatial and temporal modulation of PIN2 distribution as well as auxin gradient regulation46,47,48,49. On the other hand, root bending requires a functional cytoskeleton network49. Proper actin cytoskeleton assembly is also required to trigger and maintain root hair integrity50,51, which is compromised in the pi4kß1ß2 mutant. Furthermore, phosphoinositides can regulate actin dynamics by direct interaction with actin-binding proteins (ABPs)52,53, or affect actin polymerization, dynamics, and association with membranes indirectly through regulation of the activity and localization of Rho GTPases54 or via recruiting scaffolding proteins to the PM52,55. Indeed an altered actin cytoskeleton was observed in our mutant and phosphoinositides are well known to regulate actin organization56. A pi4p5k10/11 double mutant also displayed an increased sensitivity to latB, whereas PI4P5K10 overexpression resulted in aggregation of the apical actin fringe in tobacco pollen tubes43.
Nevertheless, the intermediate signaling links connecting PI4Kbeta 1/2 deficiency and the resulting misregulation of PI4P on endomembranes with altered ectopic auxin signaling activities remain to be clarified. For instance, root gravitropic growth requires establishing of PIN2 polarity, that also involves MEMBRANE ASSOCIATED KINASE REGULATOR 4, MAKR4, acting downstream of auxin receptors TRANSMEMBRANE KINASE1 (TMK1). At the same time, PIN2 and MAKR4 plasma membrane localisation is dependent on anionic phospholipids turnover57. On the other hand, root bending requires cell wall modification that is under control of AHA H+-ATPases in cooperation with TMKs58. Interestingly, a mutant cngc2, deficient in plasma membrane-localized CYCLIC NUCLEOTIDE-GATED ION CHANNEL 2 exhibits a phenotype, partially similar to pi4kß1ß2. This concerns stunted root and rosette growth, SA accumulation in the leaves, lower gravitropic bending and impaired sensitivity to exogenous auxin, but, unlike pi4kß1ß2, cngc2 mutant accumulates higher endogenous IAA59.
Based on our observations, a working model is proposed that assembles multiple causes leading to the short root phenotype of the pi4kß1ß2 mutant that arises from several root developmental defects, including reduced cell number and length (Fig. 7). Many correlate with altered dynamics of intracellular delivery processes. Plasma membrane establishment remains incomplete, cell architecture is misshaped, and PIN2 turnover is altered in the root elongation zone. This can be associated with a lower stability of the actin filaments network. Based on DII-VENUS degradation and gene expression, there appears to be a lack of response to auxin, endogenous or exogenous, in the pi4kß1ß2 mutant. A link between altered trafficking/cytoskeleton integrity and this lack of gene expression response will require further investigations.
Working model for the impact of pi4kß1ß2 mutations on root length. The pi4kß1ß2 mutations lead to an altered actin cytoskeleton, an altered vesicle trafficking and an altered sensitivity to auxin including at the gene expression level. Altered trafficking can be linked to PI4K interacting with small G proteins like Rab or Rho proteins; it could also be a consequence of the weakened cytoskeleton. It is hypothesized that both altered cytoskeleton and trafficking prevent a correct cytokinesis. Finally, we propose that the short root phenotype results from multiple causes: altered actin cytoskeleton, altered cytokinesis, altered trafficking, and altered auxin responses.
Materials and methods
Plant material
Experiments were performed using A. thaliana Col-0 as the WT control and the following mutant lines: pi4kβ1β2 (SALK_040479/SALK_0906930), 60, CycB1::GUS60, DR5::GUS27, PIN2::PIN2-GFP43, PIN2::PIN2-GFP in a pi4kβ1β2 background23, DII-VENUS28, DII-VENUS in pi4kβ1β2 (this study), 35S::LifeAct-GFP, 35S::LifeAct-GFP in a pi4kβ1β2 background61. The DII-VENUS construct was introduced into pi4kβ1β2 by floral dip transformation; three independent lines were selected and the T4 generation was studied. CycB1::GUS60 and DR5::GUS27 constructs were introduced into the pi4kβ1β2 background by crossing, and homozygous F3 seeds were used. Genotyping primers are listed in Supplementary Table 1. This study complies with relevant institutional, national, and international guidelines and legislation for using plant material. Analytical grade chemicals used in this study were purchased from Sigma Aldrich (St. Louis, Missouri, USA).
Plant cultivation
Seeds were surface sterilized with 1.6% sodium hypochlorite (30% of SAVO®, Unilever) solution containing 0.02% (v/v) TWEEN20 (Sigma Aldrich, St. Louis, Missouri, USA). Seeds were stratified for 2 days at 4 °C in the dark. Seeds were germinated for 3 days in Petri dishes containing half-strength Murashige–Skoog basal salt medium (Duchefa, Haarlem, Netherlands), pH 5.7, supplemented with 1% (w/v) sucrose and 0.8% (w/v) plant agar (Duchefa, Haarlem, Netherlands) at 22 °C under a 16 h light/8 h dark regime in a vertical position.
For the primary root length analysis, 4 days after germination, seedlings were transferred to square Petri plates containing the same medium supplemented or not with hormones (IAA at 0.05, 0.1 or 1 µM final concentration; BAP, at 0.1, 0.5, 1 or 5 µM; SA at 2, 10 or 20 µM). Stock solutions at 200 mM were prepared in distilled water and few drops of 1 N NaOH. After 7 days of cultivation in vertical position Petri dishes were scanned for the primary root length measurement (Epson Perfection V700 Photo, Suwa, Japan, at 600 dpi resolution). For the measurement of the lengths of meristem, elongation zone and cortical cells, roots were observed under an ApoTome Zeiss microscope with a 5 × objective at bright field settings. Images were analyzed with FiJi software62. At least 12 seedlings were analyzed for each variant. For the measurement of root hair length and density, 5-day-old seedlings were photographed under a stereo microscope (SteREO Discovery V8, Carl Zeiss GmbH, Jena, Germany) equipped with an AxioCam HRc camera. Images were imported into FiJi software and root hair length was measured manually using a segmented line tool. At least 60 root hairs from 10 seedlings were analyzed for each variant. For PIN2 localization and dynamics analysis, DII-Venus assay or actin structure evaluation, 7-days old seedlings were used. For actin structure evaluation, seedlings expressing 35S::LifeAct-GFP were sprayed with 10 μM latB (latrunculin B) for different time incubations (30 min, 90 min and 150 min) and were used for confocal microscopy. For DII-VENUS assay, seedlings were transferred to the media supplemented 0.01 μM IAA for 1 h and subjected to microscopy. The fluorescence intensity of nuclei was extracted using FiJi software.
Gravitropic test
Gravitropic response test was performed as previously described46. Five-day-old seedlings were transferred onto fresh Petri Dishes containing half-strength Murashige–Skoog basal salt medium (Duchefa, Haarlem, Netherlands), pH 5.7, supplemented with 1% (w/v) sucrose and 0.8% (w/v) plant agar (Duchefa, Haarlem, Netherlands) and aligned in a horizontal orientation. Plants were scanned at indicated time points using a Horizontal LSM880 with Airyscan module for 12 h and images were used to determine root reorientation. The root turning angle and length were calculated for each time point. Ten roots were imaged for each genotype.
GUS staining
GUS staining was performed as previously described63. Briefly, 4- or 8-day-old seedlings were incubated in 2 mM X-Gluc, 50 mM NaH2PO4, pH 7, 0.5% (v/v) Triton-X, 0.5 mM K-ferricyanide, for 16 h at 37 °C. Chlorophyll was removed by repeated washing with 80% (v/v) ethanol. Imaging was performed using an ApoTome Zeiss microscope with a 5 × objective at bright field settings.
Confocal microscopy
A Zeiss LSM 880 inverted confocal laser scanning microscope (Carl Zeiss AG, Germany) was used with a 40 × C-Apochromat objective (NA = 1.2 W). Fluorescence signals were processed with Zen Blue software (Zeiss), where PIN2 distribution was evaluated as a ratio of mean fluorescence intensity at the apical PM to mean intracellular fluorescence intensity of individual cells. Fluorescence associated with actin filaments (LifeAct-GFP) or DII-VENUS was acquired by excitation at 488 nm and emission at 490–540 nm for GFP. Images were acquired in z-stacks (step size 0.43 μm, 40–50 sections per stack). Actin filaments density and DII-VENUS signal intensity was calculated by FiJi software as the percent occupancy of GFP signal in each Maximum intensity projection. For each variant, fluorescent intensity of at least 5 roots was analyzed with 1–5 ROI (region of interest) per 1 root (ROI corresponds to one entire cell for actin; ROI corresponds to meristematic zone for DII-VENUS). For analyzing the skewness, all z-stack images were skeletonized and projected using a plugin moment calculator.
For tracking PIN2-GFP distribution in WT and pi4kß1ß2 over time (supplementary movies SM3 and SM4), ten frames were continuously obtained by confocal microscopy to track the movement of PIN2:GFP in root epidermis cells in the transition zone and compiled to a movie. PIN2-GFP subcellular distribution and cell properties were monitored on a Zeiss LSM880 microscope (AxioObserver, objective C-Apochromat 40x/1.2 W Korr FCS M27, Filter 493–598, Laser 488 nM, using zoom factor 6. Original picture size is 35,42 μm × 35,42 μm, scale bar is 10 μm.
For root hair video showing cytoplasmic streaming (supplementary movies SM5 and SM6), maximum intensity projections of a Z-stack of a root hair were taken over time. Fluorescent channel and bright field are presented together. Fluorescent channel: visualization of cytoplasmic streaming in root hair cell outgrowing a root hair, based on differential movement of fluorescent intracellular structures in the line PIN2::PIN2-GFP compared to the mutant expressing PIN2-GFP. The movie was reconstructed from confocal pictures captured in 20 frames (time-lapse) and in 18 (WT background)/19 slices (mutant background) through the root hair along the z-axis. Original picture size is 106.27 μm × 106.27 μm, pictures were captures with EC Plan-Neofluar 20x/0.50 (WD = 2.0 mm) objective, using zoom factor 4. Scale bar is 10 μm. Brightfield channel: visualization of cytoplasmic streaming in a movie reconstructed from confocal pictures captured in 20 frames (time-lapse) and in 18 (WT background)/19 slices (mutant background) along the root hair in the z-axis. Original picture size is 106.27 μm × 106.27 μm, pictures were captures with EC Plan-Neofluar 20x/0.50 (WD = 2.0 mm) objective, using zoom factor 4. Scale bar is 10 μm.
FM 4–64 staining of the plasma membrane
Five-day-old A. thaliana seedlings expressing PIN2::PIN2-GFP47 were incubated with 2 μM FM 4–64 (Molecular Probes, catalogue number T13320) in half-strength Murashige and Skoog (Sigma Aldrich, St. Louis, Missouri, USA) liquid medium in multi-well plates for 5 min and then rinsed 3 times in liquid medium64. The seedlings were observed using a confocal scanning microscope Zeiss LSM 880 with Airyscan module.
PIN2 immunolocalization
Whole mount immunolocalization of 5–day–old seedlings was performed as described previously65 with minor changes. The protocol was adapted to the InSituPro VS liquid-handling robot (Intavis AG, Germany). Prior to immunolocalization, seedlings were fixed 1 h with 4% paraformaldehyde dissolved in MTSB (50 mM PIPES, 5 mM EGTA, 5 mM MgSO4·7H2O pH 7, adjusted with KOH), at room temperature, with no vacuum. In the robot, the procedure started with several washes with MTSB-T (MTSB + 0.01% TritonX-100) then cell walls were digested with 0.05% Pectolyase Y-23 in MTSB-T and membranes were permeated with DMSO/Igepal in MTSB-T. Samples were blocked with BSA (blocking solution: 2% BSA in MTSB-T) and incubated first with anti-PIN2 rabbit antibody (kindly provided by Prof. C. Luschnig, dilution 1:500) and then a secondary anti-rabbit Alexa Fluor 546 antibody (Thermo Fisher Scientific, dilution 1:1000). Both antibodies were diluted in BSA. Between the described steps, washes with MTSB-T were provided and at the end MTSB-T was exchanged for deionized water. Seedlings were then transferred from the robot to 50% glycerol in deionized water and the fluorescence signal was measured using a confocal scanning microscope Zeiss LSM 880 with Airyscan module.
RNA extraction and RNA-seq
For each of the 3 biological repetitions, RNA samples were obtained by pooling RNAs from more than 70 plants. Seven-days-old seedlings roots (100–200 mg fresh weight) were frozen in liquid N2. Roots were homogenized in tubes with 1 g of 1.3 mm silica beads using a FastPrep-24 instrument (MP Biomedicals, USA). Total RNA was isolated using a Spectrum Plant Total RNA kit (Sigma-Aldrich, USA) and treated with a DNA-free kit (Ambion, USA). The quantity of extracted RNA was measured using NanoDrop.
Sequencing was carried out using an Illumina NexSeq500 (IPS2 POPS platform). RNA-seq libraries were made using the TruSeq Stranded mRNA kit (Illumina®, California, USA). The RNA-seq samples were Single End (SE) sequenced, stranded with a sizing of 260 bp and a read length of 75 bases, lane repartition and barcoding gave approximately 45 million SE reads per sample.
Gene transcription measurement was conducted as described previously61. In general, 1 μg of RNA was converted into cDNA with M-MLV RNase H−Point Mutant reverse transcriptase (Promega Corp., USA) and an anchored oligo dT21 primer (Metabion, Germany). Gene expression was quantified by qRT-PCR using a LightCycler 480 SYBR Green I Master kit and LightCycler 480 (Roche, Switzerland). The PCR conditions were 95 °C for 10 min followed by 45 cycles of 95 °C for 10 s, 55 °C for 20 s, and 72 °C for 20 s. Melting curve analysis was then conducted. CT values of target genes were normalized to the housekeeping gene TIP41. A list of the analyzed genes and primers is available in Supplementary table 1.
RNA-seq bioinformatic treatments and analyses
To facilitate comparisons, each sample followed the same steps from trimming to counts. RNA-Seq preprocessing included trimming library adapters and performing quality controls. The raw data (fastq) were trimmed using the Trimmomatic66 tool for a Phred Quality Score Qscore > 20, read length > 30 bases, and ribosome sequences were removed with the sortMeRNA tool67.The genomic mapper STAR (version 2.7. 3a68) was used to align reads against the A. thaliana genome (from TAIRv10), with options–outSAMprimaryFlag AllBestScore–outFilterMultimapScoreRange 0 to keep the bests results. Transcript abundance of each gene was calculated with STAR and counts only single reads for which reads map unambiguously one gene, thus removing multi-hits. According to these rules, around 97% of SE reads were associated with a gene, 1–2% of SE reads were unmapped and 1.22–1.66% of SE reads with multi-hits were removed. Differential analyses followed the procedure previously described69. Briefly, genes with less than 1 read after a counts-per-million (CPM) normalization in at least one half of the samples were discarded. Library size was normalized using the trimmed mean of M-value (TMM) method and count distribution was modeled with a negative binomial generalized linear model. Dispersion was estimated by the edgeR method70 in the statistical software ‘R’71 (Version 3.2.5 R Development Core Team (2005). Expression differences compared 2 samples using likelihood ratio tests and p-values were adjusted with the Benjamini–Hochberg procedure to control False Discovery Rate (FDR). A gene was declared differentially expressed if the adjusted p-value < 0.05.
Genes were classified using the Classification SuperViewer Tool developed by72 as described previously20. The classification source was set to Gene Ontology categories as defined by73. The frequency of each category was normalized to the whole Arabidopsis set. The mean and standard deviation for 100 boot-straps of our input set were calculated to provide some idea as to over- or under-representation reliability. Similarity analysis were performed using tools developed by Genevestigator29. The “Hierarchical clustering” tool works on the expression matrix defined by a microarray experiment selection and a gene selection. The “Biclustering” tool identifies groups of genes that are expressed above or under a set threshold ratio in a subset of conditions rather than in all conditions.
Hormone measurements
Whole roots (50–100 mg) were harvested from 7-day-old vertical grown seedlings. At least 6 samples were analyzed for WT and pi4kβ1β2. Hormone analysis was performed with a LC/MS system consisting of UHPLC 1290 Infinity II (Agilent, Santa Clara, CA, USA) coupled to 6495 Triple Quadrupole Mass Spectrometer (Agilent, Santa Clara, CA, USA), operating in MRM mode, with quantification by the isotope dilution method. Detailed methodology was described previously74.
Data deposition
Experimental steps, from growth conditions to bioinformatic analyses, have been deposited in the CATdb database75 as ProjectID NGS2020_14_pi4kb1b2 and further submitted to the international repository GEO76 as ProjetID = GSE179635.
Statistical analysis
At least three biological repetitions were carried out for all experiments, and at least 10 seedlings were analyzed for each treatment. Student’s t-test with correction for multiple comparisons and one-way ANOVA with Tukey’s HSD post-hoc test were applied; the exact number of values and statistical procedures are stated in the figure legends.
References
Retzer, K. & Weckwerth, W. The TOR–auxin connection upstream of root hair growth. Plants 10, 150 (2021).
Waldie, T. & Leyser, O. Cytokinin targets auxin transport to promote shoot branching. Plant Physiol. 177, 803–818 (2018).
Luschnig, C. & Vert, G. The dynamics of plant plasma membrane proteins: PINs and beyond. Development (Cambridge, England) 141, 2924–2938 (2014).
Habets, M. & Offringa, R. PIN-driven polar auxin transport in plant developmental plasticity: A key target for environmental and endogenous signals. The New Phytol. 203, 1 (2014).
Semeradova, H., Montesinos, J. C. & Benkova, E. All roads lead to Auxin: Post-translational regulation of Auxin transport by multiple hormonal pathways. Plant Commun. 1, 100048 (2020).
Pokotylo, I., Kravets, V., Martinec, J. & Ruelland, E. The phosphatidic acid paradox: Too many actions for one molecule class? Lessons from plants. Prog. Lipid Res. 71, 43–53 (2018).
Noack, L. C. & Jaillais, Y. Functions of anionic lipids in plants. Annu. Rev. Plant. Biol. 71, 71–102 (2020).
Jaillais, Y. & Ott, T. The nanoscale organization of the plasma membrane and its importance in signaling: A proteolipid perspective1. Plant Physiol. 182, 1682–1696 (2020).
Gronnier, J. et al. Structural basis for plant plasma membrane protein dynamics and organization into functional nanodomains. eLife 6, e26404 (2017).
Galvan-Ampudia, C. S. et al. Temporal integration of auxin information for the regulation of patterning. eLife 9, e55832 (2020).
Ke, M. et al. Salicylic acid regulates PIN2 auxin transporter hyperclustering and root gravitropic growth via Remorin-dependent lipid nanodomain organisation in Arabidopsis thaliana. New Phytol. 229, 963–978 (2021).
McKenna, J. F. et al. The cell wall regulates dynamics and size of plasma-membrane nanodomains in Arabidopsis. PNAS 116, 12857–12862 (2019).
Akhter, S. et al. Role of Arabidopsis AtPI4Kγ3, a type II phosphoinositide 4-kinase, in abiotic stress responses and floral transition. Plant Biotechnol. J. 14, 215–230 (2016).
Galvão, R. M., Kota, U., Soderblom, E. J., Goshe, M. B. & Boss, W. F. Characterization of a new family of protein kinases from Arabidopsis containing phosphoinositide 3/4-kinase and ubiquitin-like domains. Biochem. J. 409, 117–127 (2008).
Cacas, J.-L. et al. Revisiting plant plasma membrane lipids in tobacco: A focus on sphingolipids. Plant Physiol 170, 367–384 (2016).
Delage, E., Ruelland, E., Guillas, I., Zachowski, A. & Puyaubert, J. Arabidopsis type-III phosphatidylinositol 4-kinases β1 and β2 are upstream of the phospholipase C pathway triggered by cold exposure. Plant Cell Physiol. 53, 565–576 (2012).
Djafi, N. et al. The Arabidopsis DREB2 genetic pathway is constitutively repressed by basal phosphoinositide-dependent phospholipase C coupled to diacylglycerol kinase. Front. Plant Sci. 4, 307 (2013).
Krinke, O., Novotná, Z., Valentová, O. & Martinec, J. Inositol trisphosphate receptor in higher plants: Is it real?. J. Exp. Bot. 58, 361–376 (2007).
Ruelland, E. et al. Salicylic acid modulates levels of phosphoinositide dependent-phospholipase C substrates and products to remodel the Arabidopsis suspension cell transcriptome. Front. Plant Sci. 5, 608 (2014).
Kalachova, T. et al. The inhibition of basal phosphoinositide-dependent phospholipase C activity in Arabidopsis suspension cells by abscisic or salicylic acid acts as a signalling hub accounting for an important overlap in transcriptome remodelling induced by these hormones. Environ. Exp. Bot. 123, 37–49 (2016).
Šašek, V. et al. Constitutive salicylic acid accumulation in pi4kIIIβ1β2 Arabidopsis plants stunts rosette but not root growth. New Phytol. 203, 805–816 (2014).
Pluhařová, K. et al. “Salicylic Acid Mutant Collection” as a Tool to Explore the Role of Salicylic Acid in Regulation of Plant Growth under a Changing Environment. Int. J. Mol. Sci. 20, 1 (2019).
Lin, F. et al. A dual role for cell plate-associated PI4Kβ in endocytosis and phragmoplast dynamics during plant somatic cytokinesis. The EMBO Journal 38, e100303 (2019).
Kang, B.-H., Nielsen, E., Preuss, M. L., Mastronarde, D. & Staehelin, L. A. Electron tomography of RabA4b- and PI-4Kβ1-labeled trans Golgi network compartments in Arabidopsis. Traffic 12, 313–329 (2011).
Preuss, M. L. et al. A role for the RabA4b effector protein PI-4Kbeta1 in polarized expansion of root hair cells in Arabidopsis thaliana. J. Cell Biol. 172, 991–998 (2006).
Singh, S. K., Fischer, U., Singh, M., Grebe, M. & Marchant, A. Insight into the early steps of root hair formation revealed by the procuste1 cellulose synthase mutant of Arabidopsis thaliana. BMC Plant Biol. 8, 57 (2008).
Ulmasov, T., Liu, Z. B., Hagen, G. & Guilfoyle, T. J. Composite structure of auxin response elements. Plant Cell 7, 1611–1623 (1995).
Brunoud, G. et al. A novel sensor to map auxin response and distribution at high spatio-temporal resolution. Nature 482, 103–106 (2012).
Hruz, T. et al. Genevestigator V3: A Reference Expression Database for the Meta-Analysis of Transcriptomes. Adv. Bioinf. 2008, e420747 (2008).
Preuss, M. L., Serna, J., Falbel, T. G., Bednarek, S. Y. & Nielsen, E. The arabidopsis rab GTPase RabA4b localizes to the tips of growing root hair cells. Plant Cell 16, 1589–1603 (2004).
Sassi, M. et al. COP1 mediates the coordination of root and shoot growth by light through modulation of PIN1- and PIN2-dependent auxin transport in Arabidopsis. Development 139, 3402–3412 (2012).
Staswick, P. E. The tryptophan conjugates of jasmonic and indole-3-acetic acids are endogenous Auxin inhibitors. Plant Physiol. 150, 1310–1321 (2009).
Löfke, C., Dünser, K., Scheuring, D. & Kleine-Vehn, J. Auxin regulates SNARE-dependent vacuolar morphology restricting cell size. eLife 4, e05868 (2015).
Kang, B.-H., Busse, J. S. & Bednarek, S. Y. Members of the arabidopsis dynamin-like gene family, ADL1, are essential for plant cytokinesis and polarized cell growth[W]. Plant Cell 15, 899–913 (2003).
Caillaud, M.-C. et al. MAP65-3 microtubule-associated protein is essential for nematode-induced giant cell ontogenesis in arabidopsis. Plant Cell 20, 423–437 (2008).
Glanc, M., Fendrych, M. & Friml, J. PIN2 polarity establishment in arabidopsis in the absence of an intact cytoskeleton. Biomolecules 9, 222 (2019).
Geldner, N., Friml, J., Stierhof, Y.-D., Jürgens, G. & Palme, K. Auxin transport inhibitors block PIN1 cycling and vesicle trafficking. Nature 413, 425–428 (2001).
Goldstein, R. E. & van de Meent, J.-W. A physical perspective on cytoplasmic streaming. Interface Focus 5, 20150030 (2015).
Nagawa, S. et al. ROP GTPase-dependent actin microfilaments promote PIN1 polarization by localized inhibition of clathrin-dependent endocytosis. PLOS Biol. 10, e1001299 (2012).
Antignani, V. et al. Recruitment of PLANT U-BOX13 and the PI4Kβ1/β2 phosphatidylinositol-4 kinases by the small GTPase RabA4B plays important roles during salicylic acid-mediated plant defense signaling in arabidopsis. Plant Cell 27, 243–261 (2015).
Smokvarska, M., Jaillais, Y. & Martinière, A. Function of membrane domains in rho-of-plant signaling. Plant Physiol. 185, 663–681 (2021).
Smokvarska, M. et al. A plasma membrane nanodomain ensures signal specificity during osmotic signaling in plants. Curr. Biol. 30, 4654-4664.e4 (2020).
Ischebeck, T. et al. Phosphatidylinositol 4,5-bisphosphate influences PIN polarization by controlling clathrin-mediated membrane trafficking in Arabidopsis. Plant Cell 25, 4894–4911 (2013).
Mei, Y., Jia, W.-J., Chu, Y.-J. & Xue, H.-W. Arabidopsis phosphatidylinositol monophosphate 5-kinase 2 is involved in root gravitropism through regulation of polar auxin transport by affecting the cycling of PIN proteins. Cell Res. 22, 581–597 (2012).
Reyes-Hernández, B. J. et al. Root stem cell niche maintenance and apical meristem activity critically depend on threonine synthase1. J. Exp. Bot. 70, 3835–3849 (2019).
Retzer, K. et al. Brassinosteroid signaling delimits root gravitropism via sorting of the Arabidopsis PIN2 auxin transporter. Nat. Commun. 10, 5516 (2019).
Abas, L. et al. Intracellular trafficking and proteolysis of the Arabidopsis auxin-efflux facilitator PIN2 are involved in root gravitropism. Nat Cell Biol 8, 249–256 (2006).
Luschnig, C., Gaxiola, R. A., Grisafi, P. & Fink, G. R. EIR1, a root-specific protein involved in auxin transport, is required for gravitropism in Arabidopsis thaliana. Genes Dev. 12, 2175–2187 (1998).
Kleine-Vehn, J. et al. Differential degradation of PIN2 auxin efflux carrier by retromer-dependent vacuolar targeting. PNAS 105, 17812–17817 (2008).
Ketelaar, T. The actin cytoskeleton in root hairs: all is fine at the tip. Curr Opin Plant Biol 16, 749–756 (2013).
Grierson, C., Nielsen, E., Ketelaarc, T. & Schiefelbein, J. Root Hairs. Arabidopsis Book 12, e0172 (2014).
Wu, C.-Y. et al. The role of phosphoinositide-regulated actin reorganization in chemotaxis and cell migration. Br. J. Pharmacol. 171, 5541–5554 (2014).
Sun, T., Li, S. & Ren, H. Profilin as a regulator of the membrane-actin cytoskeleton interface in plant cells. Front. Plant Sci. 4, 512 (2013).
Molendijk, A. et al. Arabidopsis thaliana Rop GTPases are localized to tips of root hairs and control polar growth. EMBO J. 20, 2779–2788 (2001).
Pleskot, R., Pejchar, P., Staiger, C. & Potocký, M. When fat is not bad: the regulation of actin dynamics by phospholipid signaling molecules. Front. Plant Sci. 5, 5 (2014).
Janmey, P. A., Bucki, R. & Radhakrishnan, R. Regulation of actin assembly by PI(4,5)P2 and other inositol phospholipids: An update on possible mechanisms. Biochem. Biophys. Res. Commun. 506, 307–314 (2018).
Marquès-Bueno, M. M. et al. Auxin-regulated reversible inhibition of TMK1 signaling by MAKR2 modulates the dynamics of root gravitropism. Curr. Biol. 31, 228-237.e10 (2021).
Lin, W. et al. TMK-based cell-surface auxin signalling activates cell-wall acidification. Nature 599, 278–282 (2021).
Chakraborty, S. et al. Cyclic nucleotide-gated ion channel 2 modulates Auxin homeostasis and signaling. Plant Physiol 187, 1690–1703 (2021).
Colón-Carmona, A., You, R., Haimovitch-Gal, T. & Doerner, P. Spatio-temporal analysis of mitotic activity with a labile cyclin–GUS fusion protein. Plant J. 20, 503–508 (1999).
Kalachova, T. et al. Interplay between phosphoinositides and actin cytoskeleton in the regulation of immunity related responses in Arabidopsis thaliana seedlings. Environ. Experim. Bot. 167, 103867 (2019).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Figueroa-Balderas, R. E., García-Ponce, B. & Rocha-Sosa, M. Hormonal and stress induction of the gene encoding common bean acetyl-coenzyme A carboxylase. Plant Physiol. 142, 609–619 (2006).
Marhava, P. et al. Plasma membrane domain patterning and self-reinforcing polarity in Arabidopsis. Dev. Cell 52, 223-235.e5 (2020).
Sauer, M. et al. Canalization of auxin flow by Aux/IAA-ARF-dependent feedback regulation of PIN polarity. Genes Dev 20, 2902–2911 (2006).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Kopylova, E., Noé, L. & Touzet, H. SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data. Bioinformatics 28, 3211–3217 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Rigaill, G. et al. Synthetic data sets for the identification of key ingredients for RNA-seq differential analysis. Brief Bioinform. 19, 65–76 (2018).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucl. Acids Res. 40, 4288–4297 (2012).
R Core Team. R: A Language and Environment for Statistical Computing. (2018).
Zhu, T. A browser-based functional classification SuperViewer for Arabidopsis genomics. (2003).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Prerostova, S. et al. Light quality and intensity modulate cold acclimation in Arabidopsis. Int. J. Mol. Sci. 22, 2736 (2021).
Gagnot, S. et al. CATdb: A public access to Arabidopsis transcriptome data from the URGV-CATMA platform. Nucl. Acids Res. 36, D986-990 (2008).
Edgar, R., Domrachev, M. & Lash, A. E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucl. Acids Res. 30, 207–210 (2002).
Acknowledgements
This work was supported by the European Regional Development Fund, Project “Centre for Experimental Plant Biology” [Grant No. CZ.02.1.01/0.0/0.0/16_019/0000738] and Czech Science Foundation (Grant No. 19-13375Y). IEB Imaging Facility is supported by a project of the Ministry of Education, Youth and Sports “National Infrastructure for Biological and Medical Imaging (Czech-BioImaging – LM2018129)". The POPS platform benefits from the support of the LabEx Saclay Plant Sciences-SPS (ANR-10-LABX-0040-SPS). AS benefited from the French government grant for PhD studies in co-tutelle.
Author information
Authors and Affiliations
Contributions
A.S., T.K. and E.R. designed the study. A.S., T.K., K.R., A.J., P.D., J.L., R.P., J.A., A.G., S.P., L.S.-T. performed the experiments. A.S.., T.K., L.B. and E.R. analyzed data. A.S., T.K., and E.R. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Video 1.
Supplementary Video 2.
Supplementary Video 3.
Supplementary Video 4.
Supplementary Video 5.
Supplementary Video 6.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Starodubtseva, A., Kalachova, T., Retzer, K. et al. An Arabidopsis mutant deficient in phosphatidylinositol-4-phosphate kinases ß1 and ß2 displays altered auxin-related responses in roots. Sci Rep 12, 6947 (2022). https://doi.org/10.1038/s41598-022-10458-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-10458-8
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.