Abstract
We aim to evaluate the performance of a deep convolutional neural network (DCNN) in predicting the presence or absence of sarcopenia using shear-wave elastography (SWE) and gray-scale ultrasonography (GSU) of rectus femoris muscle as an imaging biomarker. This retrospective study included 160 pair sets of GSU and SWE images (n = 160) from December 2018 and July 2019. Two radiologists scored the echogenicity of muscle on GSU (4-point score). Among them, 141 patients underwent CT and their L3 skeletal muscle index (SMI) were measured to categorize the presence or absence of sarcopenia. For DCNN, we used three CNN architectures (VGG19, ResNet-50, DenseNet 121). The accuracies of DCNNs for sarcopenia classification were 70.0–80.0% (based on SWE) and 65.0–75.0% (based on GSU). The DCNN application to SWE images highlights the utility of deep-learning base SWE for sarcopenia prediction. DCNN application to SWE images might be a potentially useful biomarker to predict sarcopenic status.
Similar content being viewed by others
Introduction
The term “sarcopenia” was coined in 1988 by Rosenberg and was originally defined as a muscle loss in the appendicular muscle mass in older people1. In 2010, the definition was modified to also refer to low muscle function2. Muscle strength differs among individuals and decreases with age3. Muscle mass measurement for sarcopenia evaluation is usually assessed by medical imaging modalities, such as dual-energy X-ray absorptiometry, computed tomography (CT), magnetic resonance imaging (MRI), and ultrasonography (USG).
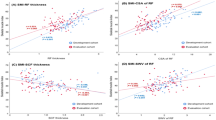
USG can measure targeted muscle size and echogenicity in real-time with several advantages (e.g., relatively low cost, portability, lack of radiation exposure, provision of muscle components such as fibrosis, adipose tissue infiltration). The thickness of the quadriceps is known to be highly correlated with voluntary contraction force, and the echogenicity of the rectus femoris muscle is known to be associated with muscle strength4,5. Intramuscular fat and fibrosis affect the echogenicity of gray-scale USG (GSU), and the qualitative information is essential for evaluating the functional information; it is not always correlated with quantitative information6. Shear-wave elastography (SWE) USG is a relatively new non-invasive functional imaging method for measuring soft tissue elasticity (i.e., tissue stiffness) with high reproducibility and objective quantitative imaging capability, which might contain more objective qualitative information7. In the evaluation of idiopathic inflammatory myopathy, the SWE measurements in Young’s modulus of the muscle demonstrated significant associations with disease activity, suggesting that it could be considered as a new modality for monitoring disease activity8.
Imaging analysis and interpretation of radiologic imaging are basic tasks performed by radiologists in providing qualitative radiologic reading, depending on their experience and medical knowledge. However, in this era of big data and artificial intelligence, radiologic imaging has been enhanced with a capability to provide quantitative imaging biomarkers for early detection, further characterization, activity monitoring, and response to treatment. Recently, deep learning and radiomics have been introduced in radiology for analyzing and interpreting images. Deep learning is a subset of machine learning where multiple hidden neural networks learn representations of data through simple and complex feature abstraction to perform tasks9,10. Convolutional neural networks (CNNs) are a type of artificial neural network designed to use pixel data in images to learn abstract representations with high levels of semantics, where deep convolutional neural network (DCNN) has been shown to exhibit high performance in medical classification, detection, and segmentation tasks [8–12]9,11,12,13,14. Radiomics is a method of high throughput data mining that extracts numerous image features from routine clinical images to assess tumor characteristics on radiologic images, such as spatial heterogeneity, texture, or shape for precision medicine and decision support15,16. However, to date, there has been no study that predicts the sarcopenia on muscle USG using either DCNNs or radiomics.
Therefore, the aim of this study is to evaluate the performance of a deep convolutional neural network (DCNN) in predicting the presence or absence of sarcopenia using shear-wave elastography (SWE) and gray-scale ultrasonography (GSU) of rectus femoris muscle as an imaging biomarker.
Results
Demographics
Among 160 subjects who underwent USG, 69.4% (111/160) were classified as low-grade muscle echogenicity (grade 0, n = 27; grade 1, n = 84), and the other 30.6% (49/160) were classified as high-grade muscle echogenicity (grade 2, n = 39; grade 3, n = 10). Among 141 subjects who underwent USG and CT, 20.6% (29/141) were categorized as “a sarcopenia” (men, n = 22; women, n = 7) and 79.4% (112/141) were categorized as “not a sarcopenia” (men, n = 34; women, n = 78%).
Performance of the three DCNNs
The diagnostic performance of the three DCNNs in classifying the echogenicity of the muscle grades is shown in Table 1. The VGG19 and DenseNet121 pre-trained model outperformed ResNet model (accuracy, 85.0%) (Fig. 1). The best performing architecture (DenseNet121) showed a sensitivity of 83.3% and specificity of 85.7% for grading muscle echogenicity. The comparison of the diagnostic performance in sarcopenia classification on GSU and SWE USG by three DCNNs are shown in Table 1 and Fig. 1. Using data augmentation and image resizing, the accuracy of sarcopenia classification based on GSU images increased considerably from 65.0 to 75.0% on VGG19 pre-trained model, yielding performance of 77.8% sensitivity, 72.7% specificity, and 0.77 AUC. In predicting sarcopenia with GSU images, Grad-CAM was applied and showed high activations in hyperechoic areas due to muscle fascia/fibrosis and hypoechoic areas considered as intramuscular fat area (Fig. 2). The VGG19 pre-trained model with SWE USG images yielded the best performance of 80.0% accuracy, 88.9% sensitivity, and 72.7% specificity (Fig. 1b).
Discussion
Muscle mass is positively associated with muscle strength and is supported by epidemiologic cohorts17,18. However, some researchers found that muscle quality rather than quantity determines muscle function6,19. The definition for sarcopenia has been modified to include both muscle quantity and function. Skeletal muscle is composed of a group of muscle fibers, and muscle fiber is broadly classified into “slow twitch” (type I) and “fast switch” (type II)20. A selective reduced size and the atrophy of type 2 fiber are associated with sarcopenia21. Additionally, intramyocellular and intermuscular fat may increase with aging (myosteatosis)22. The suggested mechanism for these histologic alterations of aging muscles involves age-related change in activation, proliferation, and differentiation of quiescent skeletal muscle precursor “stem cells” into adipocytes.
Although MRI or CT is considered as gold-standard modality for evaluating the whole body’s composition, ultrasound is a simple, reliable, and non-invasive imaging modality for muscle imaging that provides muscle quality (i.e., histologic changes) and quantity (i.e., muscle mass or volumes) in a relatively short time without exposure to ionizing radiation. Investigators have reported inter-rater consistency in measuring the cross-sectional area of the muscle and echogenicity evaluation using USG, and it was less affected by the operator’s skill23,24,25. These results might support the role of USG as an easily assessable screening modality for evaluating sarcopenia.
In this current study, we applied DCNNs to classify the echogenicity of the muscle on GSU and to predict the presence of sarcopenia on both of GSU and SWE USG images. The results showed a good diagnostic performance (80.0% to 85.0% accuracy) in classifying muscle echogenicity on GSU images. This result is similar to a previous study that used DCNN to classify the liver cirrhosis on US images (85.7% accuracy), and it outperformed all five radiologists13. For predicting sarcopenia on both GSU and SWE USG, the DCNNs slightly performed better on SWE USG (70.0% to 80.0% accuracy) than GSU USG (65.0% to 75.0% accuracy). The prediction of sarcopenia on GSU showed an enhanced performance (65.0% to 75.0% accuracy) with an end-to-end DCNN approach compared to MLP using radiomic features. The significant feature attributes of VGG19 model predictions can be observed through Grad-CAM (Fig. 2).
We have shown that DCNN plays an important role in the proposed system in automatically extracting useful features from a limited dataset. DCNNs compared to radiomics-based networks avoid human hand-crafted feature extraction, which is time consuming and have inter-subject variations in image acquisition. Additionally, in the case of MLP with radiomic features, it is difficult to determine precisely how our model arrived at its decision due to the “black box” of the neural network26.Our results reveal that a combination of appropriate image pre-processing pipelines and pre-trained DCNN architecture selection serves as an automatic high-performing model for sarcopenia/muscle grade to overcome classification tasks with small datasets. Although MLP classifier combined with radiomic features results in a relatively low performance compared to end-to-end deep learning approach, they give insights into the possible approaches to improve performance and reproducibility of the learning systems to become fully quantitative imaging biomarkers.
There are some limitations to this study. First, the number of subjects was relatively small for deep learning, and the pre-trained networks were not directly optimized from ultrasound images. Nevertheless, this is the first study to determine the performance of DCNNs in evaluating muscle echogenicity grades and in diagnosing sarcopenia, yet it demonstrated a good performance. The performance is anticipated to improve with more suitable pre-trained network and a larger dataset. Second, we obtained data from a Korean cohort. Therefore, we used the definition of sarcopenia based on a one–population skeletal muscle index. Hence, the results may not be generalized to other groups. Further studies with various cohorts, including functional performance evaluation, are needed to evaluate the robustness of the network. Third, we did not evaluate the added value of SWE on conventional GSU image. The image features of GSU and SWE are complementary in a clinical setting. In current study, we cannot extract elastography images from fusion images of underlying GSU. As a future study, feeding the DCNN models with both GSU and SWE images to predict the presence or absence of sarcopenia is needed to image the accuracy of the model since the might provide complementary information.
In conclusion, DCNNs exhibited a high performance in sarcopenia in based on SWE images. The DCNN application to SWE images highlights the utility of deep-learning base SWE for sarcopenia prediction. DCNN application to SWE images might be a potentially useful biomarker to predict sarcopenic status.
Methods
Datasets
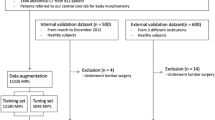
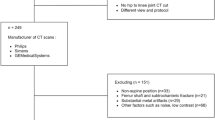
This study was approved by the institutional review board of Inje University Haeundae Paik Hospital (Approval No. 2020-02-013). Informed consent was waived by the ethics committee/institutional review board of Inje University Haeundae Paik Hospital due to the retrospective design of the study. All research was carried out in accordance with relevant guidelines and regulations. Between December 2018 and July 2019, a total of 160 consecutive adult patients underwent lower extremity ultrasonography including GSU and SWE evaluation of the mid-third portion of the right rectus femoris muscle. The patients were included in the muscle echogenicity grading evaluation, and they comprised 62 men (aged 21–87 years; mean age, 57.4 years) and 98 women (aged 24–85 years; mean age, 56.4 years) (Fig. 3). For the muscle grade and sarcopenia prediction, we randomly selected 17 cases as the validation dataset for model selection and hyperparameter tuning, and 20 cases as test dataset to evaluate the performance of the model. For the prediction of sarcopenia, 19 subjects were excluded for the following reasons: (1) unavailable CT date within 1 month of USG (n = 16), (2) unavailable body weight information (n = 1), and (3) surgical hardware in L3 level on CT image (n = 2). Finally, 141 patients enrolled for sarcopenia classification, and it comprised 56 men (aged 21–82 years; mean age, 56.1 years) and 85 women (aged 24–85 years; mean age, 56.5 years). From the 141 cases, we randomly selected 17, 20, and 104 cases as the validation, test, and training sets, respectively. Collectively, the data sets were divided into training set, validation set, and test set (training:validation:test = 115:15:30 for muscle echogenicity and training:validation: test = 96:15:30 for sarcopenia prediction) (Fig. 3).
Imaging study and analysis
Ultrasonography for muscle echogenicity grading and shear-wave elastography
All subjects underwent USG evaluation at the mid-portion of the right rectus femoris muscle with the same protocol and US scanner (LOGIQ E9; GE Healthcare, Wauwatosa, WI, USA) using a linear 9- to 5-MHz probe. This was performed by a musculoskeletal radiologist with 5 years of experience. All subjects were examined in the supine position with a neutral foot position and were asked not to move or strain the lower extremity. The GSU and SWE images of the right mid rectus femoris muscle in a transverse plane were acquired simultaneously. Color box was placed on rectus femoris muscle and the visual color scale bar is in the left aspect of the screen (red means hard and blue means soft) (Fig. 4). During USG, copious gel was applied and without flattening or deforming the superficial epimysium layer to minimize the external pressure which might affect the measurement27. To obtain the ground truth muscle grade, two musculoskeletal radiologists (13 years and 5 years of musculoskeletal radiology experience) classified the muscle echogenicity using a four-grade scale by consensus on the GSU image: (1) grade 0: low echoic muscle with inner speckled appearance of the perimysial connective tissue, (2) grade 1: area of increased echo relative to background muscle, (3) grade 2: nearly as echogenic as perimysial fat, and (4) grade 3: isoechoic to fat (Fig. 5)28. The muscle echogenicity grades were dichotomized as low grade (grade 0 and grade 1) and high grade (grade 2 and grade 3). This system was used as the muscle echogenicity grading ground truth system.
Assessment of sarcopenia: muscle quantification on computed tomography
All examinations were performed in a center using two multidetector-row computed tomography on an axial plane including the L3 level: a 128-slice system (Definition AS+, Siemens Healthineers, Forchheim, Germany) and a 64-slice system (Discovery CT 750 HD, GE Healthcare, Waukesha, WI, USA). The scan parameters were as follows: fixed tube potential = 120 kVp; beam collimation = 128 × 0.6 mm and 64 × 0.625 mm, respectively; slice thickening of 5 mm without interval. To obtain the ground truth for sarcopenia diagnosis, the inferior end plate level of the L3 image was evaluated to measure the total abdominal muscle area on the axial image. The Asan-J software, developed based on ImageJ (NIH, Bethesda, MD, USA), was used to measure the skeletal muscle mass on the CT image (available at http://datasharing.aim-aicro.com/morphometry). Sarcopenia was defined using the L3 skeletal muscle index (cm2/m2), which is based on the Korean National Health and Nutrition Examination Study (KNHANES): \(\le\) 49 cm2/m2 for men and \(\le\) 31 cm2/m2 for women29.
Pre-processing
Preprocessing techniques were applied for preparing the images to DCNN submission: (1) intensity normalization was performed in a nonlinear way to convert GSU and SWE images into standardized intensity ranges for all subjects, (2) image crop was applied for region-of-interest : elastographic areas in SWE images and ultrasonographic area in GSU images (except for text labels) (2) The dataset was augmented by applying a rotation (− 5° to 5°), shifting (shift limit of 0.0625), scaling (scale limit of 0.1), horizontal/vertical flipping, and additional random contrast, brightness, sharpness, blurring, and Gaussian noise to increase the generalization of our networks, (3) Finally, all images were resampled ass the retrieved ultrasonography images had different heights and widths. We resized the input images with 84 × 84 input size for SWE images and 112 × 112 input size for GSU images (Fig. 6). The pre-processing was performed using an in-house code written in MATLAB (Version 2018b, Math Works, Natick, MA, USA).
DCNN architectures
Three different DCNNs (VGG19, ResNet-50, and DenseNet121) were used to classify sarcopenia muscle images to measure the efficacy of the DCNNs (Fig. 6). The pre-trained weights of all models were obtained from training a subset of ImageNet dataset—a large scale benchmark dataset that contains 1.28 million natural images with 1000 categories30. The saved DCNN models were then fine-tuned with the USG training dataset after replacing the final fully connected and softmax layer with a new fully connected layer and a random initialized sigmoid layer. The training for all the models was performed using a mini-batch gradient descent with Adam optimizer and a base learning rate of 0.0001 annealed by a factor of 0.1 when the gradient was stuck on a plateau. During the classification of the sarcopenia on GSU images, we utilized a gradient-weighted class activation map (Grad-CAM) technique to produce “visual explanations” for the decisions from our classification model31.
The pre-trained DCNN models were imported from the official repository in Keras, and all deep learning DCNN models were implemented using Keras (version.2.2.4) with a Tensorflow backend (version.1.10.0).
Statistical analysis
Three DCNNs in each clinical situation were compared in accuracy. This was followed by a diagnostic performance evaluation of the DCNN architecture in sensitivity, specificity, positive and negative likelihood ratio, and area under the receiver operating characteristic curve (AUC). The best performing DCNN architecture for different clinical situations were selected based on accuracy and AUC. The considered clinical situations were (a) classification of normal vs. sarcopenia with GSU images; (b) classification of normal vs. sarcopenia with SWE images, and (c) muscle echogenicity grading (low vs. high) with GSU images. All statistical analyses were performed using R (version 3.6.2) (R Foundation for Statistical Computing, Vienna, Austria, https://www.R-project.org).
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Rosenberg, I. Epidemiologic and methodologic problems in determining nutritional status of older persons (summary comments). Am. J. Clin. Nutr. 50, 1231–1233 (1989).
Bauer, J. et al. Sarcopenia: A time for action. An SCWD position paper. J. Cachexia Sarcopenia Muscle 10, 956–961. https://doi.org/10.1002/jcsm.12483 (2019).
Cruz-Jentoft, A. J. et al. Sarcopenia: Revised European consensus on definition and diagnosis. Age Ageing 48, 16–31 (2019).
Strasser, E. M., Draskovits, T., Praschak, M., Quittan, M. & Graf, A. Association between ultrasound measurements of muscle thickness, pennation angle, echogenicity and skeletal muscle strength in the elderly. Age 35, 2377–2388. https://doi.org/10.1007/s11357-013-9517-z (2013).
Watanabe, Y. et al. Echo intensity obtained from ultrasonography images reflecting muscle strength in elderly men. Clin. Interv. Aging 8, 993–998. https://doi.org/10.2147/cia.S47263 (2013).
Barbat-Artigas, S., Rolland, Y., Vellas, B. & Aubertin-Leheudre, M. Muscle quantity is not synonymous with muscle quality. J. Am. Med. Dir. Assoc. 14, 852.e851-857. https://doi.org/10.1016/j.jamda.2013.06.003 (2013).
Arda, K., Ciledag, N., Aktas, E., Aribas, B. K. & Kose, K. Quantitative assessment of normal soft-tissue elasticity using shear-wave ultrasound elastography. AJR Am. J. Roentgenol. 197, 532–536. https://doi.org/10.2214/ajr.10.5449 (2011).
Alfuraih, A. M. et al. Muscle shear wave elastography in idiopathic inflammatory myopathies: A case-control study with MRI correlation. Skeletal Radiol. 48, 1209–1219. https://doi.org/10.1007/s00256-019-03175-3 (2019).
Chartrand, G. et al. Deep learning: A primer for radiologists. Radiographics 37, 2113–2131. https://doi.org/10.1148/rg.2017170077 (2017).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444. https://doi.org/10.1038/nature14539 (2015).
Kooi, T. et al. Large scale deep learning for computer aided detection of mammographic lesions. Med. Image Anal. 35, 303–312. https://doi.org/10.1016/j.media.2016.07.007 (2017).
Anthimopoulos, M., Christodoulidis, S., Ebner, L., Christe, A. & Mougiakakou, S. Lung pattern classification for interstitial lung diseases using a deep convolutional neural network. IEEE Trans. Med. Imaging 35, 1207–1216. https://doi.org/10.1109/tmi.2016.2535865 (2016).
Lee, J. H. et al. Deep learning with ultrasonography: Automated classification of liver fibrosis using a deep convolutional neural network. Eur. Radiol. https://doi.org/10.1007/s00330-019-06407-1 (2019).
Narayana, P. A. et al. Deep learning for predicting enhancing lesions in multiple sclerosis from noncontrast MRI. Radiology 294, 398–404. https://doi.org/10.1148/radiol.2019191061 (2019).
Aerts, H. J. The potential of radiomic-based phenotyping in precision medicine: A review. JAMA Oncol. 2, 1636–1642. https://doi.org/10.1001/jamaoncol.2016.2631 (2016).
Gillies, R. J., Kinahan, P. E. & Hricak, H. Radiomics: Images are more than pictures, they are data. Radiology 278, 563–577. https://doi.org/10.1148/radiol.2015151169 (2016).
Beliaeff, S., Bouchard, D. R., Hautier, C., Brochu, M. & Dionne, I. J. Association between muscle mass and isometric muscle strength in well-functioning older men and women. J. Aging Phys. Act. 16, 484–493. https://doi.org/10.1123/japa.16.4.484 (2008).
Reed, R. L., Pearlmutter, L., Yochum, K., Meredith, K. E. & Mooradian, A. D. The relationship between muscle mass and muscle strength in the elderly. J. Am. Geriatr. Soc. 39, 555–561. https://doi.org/10.1111/j.1532-5415.1991.tb03592.x (1991).
Manini, T. M. & Clark, B. C. Dynapenia and aging: An update. J. Gerontol. A Biol. Sci. Med. Sci. 67, 28–40. https://doi.org/10.1093/gerona/glr010 (2012).
Talbot, J. & Maves, L. Skeletal muscle fiber type: Using insights from muscle developmental biology to dissect targets for susceptibility and resistance to muscle disease. Wiley Interdiscip. Rev. Dev. Biol. 5, 518–534 (2016).
Nilwik, R. et al. The decline in skeletal muscle mass with aging is mainly attributed to a reduction in type II muscle fiber size. Exp. Gerontol. 48, 492–498. https://doi.org/10.1016/j.exger.2013.02.012 (2013).
Kuk, J. L., Saunders, T. J., Davidson, L. E. & Ross, R. Age-related changes in total and regional fat distribution. Ageing Res. Rev. 8, 339–348. https://doi.org/10.1016/j.arr.2009.06.001 (2009).
Thomaes, T. et al. Reliability and validity of the ultrasound technique to measure the rectus femoris muscle diameter in older CAD-patients. BMC Med. Imaging 12, 7 (2012).
Zaidman, C. M., Wu, J. S., Wilder, S., Darras, B. T. & Rutkove, S. B. Minimal training is required to reliably perform quantitative ultrasound of muscle. Muscle Nerve 50, 124–128 (2014).
Sarwal, A. et al. Interobserver reliability of quantitative muscle sonographic analysis in the critically Ill population. J. Ultrasound Med. 34, 1191–1200. https://doi.org/10.7863/ultra.34.7.1191 (2015).
Castelvecchi, D. Can we open the black box of AI?. Nat. News 538, 20 (2016).
Alfuraih, A. M. et al. The effect of unit, depth, and probe load on the reliability of muscle shear wave elastography: Variables affecting reliability of SWE. J. Clin. Ultrasound 46, 108–115 (2018).
Heckmatt, J. Z., Leeman, S. & Dubowitz, V. Ultrasound imaging in the diagnosis of muscle disease. J. Pediatr. 101, 656–660 (1982).
Lee, J. S., Kim, Y. S., Kim, E. Y. & Jin, W. Prognostic significance of CT-determined sarcopenia in patients with advanced gastric cancer. PLoS ONE 13, e0202700. https://doi.org/10.1371/journal.pone.0202700 (2018).
Deng, J. et al. In 2009 IEEE conference on computer vision and pattern recognition. 248–255 (IEEE).
Selvaraju, R. R. et al. Grad-CAM: Why did you say that? arXiv preprint arXiv: arXiv:1611.07450 (2016).
Acknowledgements
This work was supported by a National Research Foundation (NRF) grant funded by the Korea government, Ministry of Science and ICT (MSIP, 2018R1A2B6009076). This work was also supported by an NRF grant funded by the Korea government (NRF-2019R1G1A1008440). This study was supported by a new faculty research seed money grant of Yonsei University College of Medicine for 2021 (2021-32-0046).
Author information
Authors and Affiliations
Contributions
J.Y., Y.S., S.H., and Y.H.L. wrote the main manuscript. J.Y. completed the data collection. Y.S. and Y.H.L. built the imaging analysis software and DCNN architecture. J.Y. and Y.H.L. analyzed and interpreted the data. All authors reviewed the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yi, J., Shin, Y., Hahn, S. et al. Deep learning based sarcopenia prediction from shear-wave ultrasonographic elastography and gray scale ultrasonography of rectus femoris muscle. Sci Rep 12, 3596 (2022). https://doi.org/10.1038/s41598-022-07683-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-07683-6
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.