Abstract
Serratia marcescens is an opportunistic bacterium that infects a wide range of hosts including humans. It is a potent pathogen in a septic injury model of Drosophila melanogaster since a few bacteria directly injected in the body cavity kill the insect within a day. In contrast, flies do not succumb to ingested bacteria for days even though some bacteria cross the intestinal barrier into the hemolymph within hours. The mechanisms by which S. marcescens attacks enterocytes and damages the intestinal epithelium remain uncharacterized. To better understand intestinal infections, we performed a genetic screen for loss of virulence of ingested S. marcescens and identified FliR, a structural component of the flagellum, as a virulence factor. Next, we compared the virulence of two flagellum mutants fliR and flhD in two distinct S. marcescens strains. Both genes are required for S. marcescens to escape the gut lumen into the hemocoel, indicating that the flagellum plays an important role for the passage of bacteria through the intestinal barrier. Unexpectedly, fliR but not flhD is involved in S. marcescens-mediated damages of the intestinal epithelium that ultimately contribute to the demise of the host. Our results therefore suggest a flagellum-independent role for fliR in bacterial virulence.
Similar content being viewed by others
Introduction
Infectious diseases remain one of the leading causes of death in the world due to new emerging strains of pathogens, multidrug resistance of microorganisms, and persistent infections. The understanding of these diseases requires deep knowledge about the interactions between host and pathogen, which can be studied from two different perspectives: that from the host and that from the pathogen. Host defense against infections encompasses two distinct but complementary facets: resistance and resilience1, also referred to as disease tolerance2,3,4. Resistance, which relies on the immune system, is the ability of the host to directly attack the pathogen to lower the microbial burden and ultimately clear the infection. In contrast, resilience, is the ability of the host to withstand and repair damages provoked directly by the pathogen or indirectly by the host’s own immune response. Viewed from the pathogen perspective, the invading microorganism needs to survive inside the host, that is, to withstand, elude or neutralize host defenses, and to gather nutrients to sustain its growth, proliferation, and ultimately its dissemination.
The digestive tract is in constant contact with various pathogens that may occasionally contaminate the food. Therefore, the intestine has developed robust resistance and resilience mechanisms to confront and to endure such infections. Numerous reports have documented the use of Drosophila as a model to study intestinal infections5,6,7. It is thought that the fly midgut prevents the passage of bacteria to the hemolymph via three major arms8: the peritrophic matrix barrier, which is a passive physical defense mechanism, and two active chemical defense mechanisms, the local secretion in the lumen of Anti-Microbial Peptides (AMP) and the local release of Reactive Oxygen Species (ROS). The peritrophic matrix lines the gut epithelium and confines microorganisms to the lumen of the digestive tract9,10. The secretion of AMPs in the midgut is regulated by the IMD and not the Toll signaling pathway10,11,12,13. The detection of microbial uracil has been proposed to trigger the production of ROS in the lumen through the Dual Oxidase (DuOx) enzyme14,15. Additionally, at least three resilience mechanisms may contribute to maintain the homeostasis of the intestinal epithelium: (i) the secretion of Immune Response Catalase (IRC) limits the detrimental effect of ROS on intestinal epithelial cells (Enterocytes = ECs)16; (ii) the proliferation of Intestinal Stem Cells (ISCs) compensates EC cell death17,18,19,20; (iii) the extrusion of EC cytoplasm within hours of ingestion eliminates intracellular toxins and damaged organelles21. Even though the fly intestine harbors sophisticated defense mechanisms, some microorganisms, such as Serratia marcescens or Pseudomonas aeruginosa, are able to resist, survive, damage and cross the intestinal barrier10,22,23.
S. marcescens is a Gram-negative entomopathogen and also a human opportunistic bacterium associated with nosocomial infections24,25,26. The pathogenicity of S. marcescens relies on multiple virulence factors such as the pore-forming toxin hemolysin27,28,29, the serralysin protease30,31,32 or a phospholipase33. S. marcescens is a potent pathogen in the septic injury model of Drosophila. When introduced directly in the hemocoel, a few bacterial cells are sufficient to kill the fly within a day. The bacteria proliferate rapidly in the hemolymph causing bacteremia followed by death. Upon detection of the bacteria, the Immune deficiency (IMD) signaling pathway stimulates the secretion of AMPs by fat body cells. However, this systemic immune response does not affect S. marcescens since IMD-deficient flies are as susceptible as wild-type flies to septic injury10. However, in the oral infection model, S. marcescens invades and damages the intestinal epithelium and causes EC cell death; yet, the flies do not succumb to the infection for days. This delay is likely accounted by resilience mechanisms such as ISC compensatory proliferation20. In the midgut, the bacteria trigger the local release of AMPs by the IMD signaling pathway10,12,13 and are thought to induce the local secretion of ROS through the DuOx enzyme15. Interestingly, a low but significant number of bacteria can cross the intestinal barrier and manage to reach the hemolymph. In contrast to the septic injury model, S. marcescens is not able to proliferate in the hemocoel as it is controlled by phagocytosis. Indeed, phagocytosis-impaired flies are highly susceptible to the oral infection as ingested bacteria proliferate in the hemolymph10. In keeping with this cellular control of bacteria that have escaped in the hemocoel, the bacteria do not trigger the systemic immune response, which monitors short peptidoglycan fragments released by bacteria during their divisions34. Thus, under normal conditions, bacteria that have crossed the intestinal barrier do not appear to contribute to the virulence of this pathogen in the oral infection model.
The difference in the virulence of the bacteria between the septic injury and the oral infection model indicates that the virulence program of S. marcescens is downregulated after its passage from the gut lumen through the midgut epithelium to the hemolymph10. How S. marcescens modulates its virulence program according to its infection route remains unknown. Additionally, the virulence factors that the bacteria employ to damage the intestinal epithelium and to cross the fly intestinal barrier are still uncharacterized. To better understand intestinal infection by S. marcescens, we performed a small-scale genetic screen to isolate bacterial mutants displaying an impaired virulence in the Drosophila oral infection model. This screen identified a novel virulence factor, fliR, that is needed for S. marcescens to severely damage the intestinal epithelium and to efficiently kill the flies. Furthermore, this study sheds light on the importance of the flagellum for the dissemination of gut bacteria through the intestinal epithelium into the internal milieu potentially causing systemic infections.
Results
The fliR gene as a novel virulence factor in S. marcescens
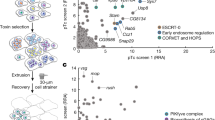
To identify new virulence factors for S. marcescens, we partially screened a transposon (mini-Tn5) insertion mutant library generated in the Db10 strain35. We examined the survival of eater−/− flies following the oral infection with individual mutant clones. Of note, the eater−/− mutants are phagocytosis-impaired flies proven useful for the screen because of their susceptibility to wild-type Serratia intestinal infection: the bacteria proliferate in the hemolymph and rapidly kill the flies, making easier the selection for less virulent bacterial strains.
We have tested 1348 bacterial mutants and identified a strain (19H12) that exhibited reduced virulence in the intestinal infection (Supplementary Fig. 1). Sequencing analysis of the 19H12 clone revealed an insertion mutation in the gene fliR, which encodes a structural component of the flagellum and is required for its biosynthesis by participating in the export machinery of its components as well as some virulence factors36,37,38. FliR is a protein that forms part of the export gate of the flagellum, a structure embedded within the MS-ring, the basal body that anchors the flagellum to the cytoplasmic membrane and the cell wall.
fliR, like flhD, is required for the formation of flagella in S. marcescens, as determined in in vivo studies
The function of fliR in the virulence of Serratia might be dependent on its role in the assembly of the flagellum. The latter is a complex process initiated by the major (class I) regulator FlhDC that controls the expression of several flagellar genes, including fliR36.
To validate the implication of fliR in the virulence of the bacteria, and to assess whether it is related to its function in the flagellum apparatus, we designed pKNOCK insertion mutants39 for the fliR gene as well as for the flhD regulatory gene. These insertion mutants, in addition to a fliR plasmidic rescue (fliR was cloned in the pBB1:lacI:MCS expression plasmid resulting in pBB1:lacI:fliR)40, were generated in two different S. marcescens wild-type strains of distinct origins: Db10 (a derivative of a Drosophila isolate from Stockholm, Sweden)41 and RM66262 (a clinical isolate from Rosario, Argentina)42.
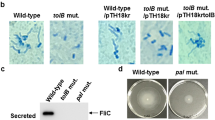
After selecting mutants in both the Db10 and the RM66262 backgrounds, we first confirmed that the mutations in fliR or flhD do not alter the growth of the bacteria in the LB medium and in the infection solution (50 mM sucrose + 10% LB) (Supplementary Fig. 2). We then determined the loss of flagellum-dependent activities in all mutants (Supplementary Fig. 3): the flagellin expression is lost and the motility is impaired in the flhD and the fliR mutants as shown by western blot, swimming, and swarming assays. Also, the phospholipase of S. marcescens is secreted through the flagellum export system, which is a type 3 secretion system (T3SS)38. As expected, we did not detect phospholipase activity for either flhD or fliR mutants as compared to the wild-type strains (Supplementary Fig. 3). The flagellum-dependent functions of the bacteria are restored in the fliR rescue indicating that the observed phenotypes are due to the lack of fliR expression and that the two independent fliR mutations do not induce polar effects in the operon.
fliR has a flagellum-independent role in the virulence of S. marcescens in the intestinal infection model
Our first attempt in the in vivo study was to validate the result of the screen with fliR insertion mutant bacteria and to compare its virulence to the flhD bacteria in the oral infection model of phagocytosis-impaired flies. Indeed, eater−/− flies fed with fliR mutant bacteria were less susceptible to the infection when compared to flies fed with either otherwise isogenic flhD mutant or wild-type bacteria of the Db10 (Fig. 1A) or the RM66262 (Fig. 1B) strains. Similar results were found when phagocytosis was impaired by the prior injection of latex beads that ultimately saturate the phagocytes after their engulfment (Supplementary Fig. 4A). As expected, the fliR strain harboring the plasmid rescue of fliR was as virulent as the wild-type strain, further demonstrating in vivo that the mutation in fliR is solely responsible for the observed phenotype (Fig. 1A,B). Additionally, we verified the virulence of the mutants in wild-type wA5001 flies. Likewise, fliR mutant bacteria in the two bacterial strain backgrounds were less virulent in the oral infection model when compared to the flhD mutant or wild-type bacteria (Fig. 1C,D). Moreover, we tested a possible role for fliR gene in the virulence of the bacteria in septic injury. We found that fliR mutant bacteria were as virulent as flhD mutant and wild-type S. marcescens when introduced directly in the hemolymph (Fig. 1E). In conclusion, these results reveal a flhD-independent role for fliR in the virulence of S. marcescens in intestinal infection, but not in the septic injury model. Similar results were obtained when monitoring the survival of IMD-deficient (kenny) and DuOx-deficient (silenced in ECs) flies following an oral infection (Supplementary Fig. 4B,C). These experiments indicate that the contribution of fliR in the virulence of S. marcescens is not related to its interactions with the fly immune system such as eliciting or evading the immune response during intestinal infections. As we had confirmed the role of fliR in the virulence of S. marcescens in two different bacterial strains, we focused only on the RM66262 strain for further investigations.
fliR and not flhD is required for S. marcescens full virulence in the D. melanogaster oral infection model. Flies were fed on filter pads with sucrose solution and 10% LB containing the bacteria at OD600nm = 0.1. IPTG (0.5 mM) was added for the fliR rescue strain (A–D). (A, B) Survival test of eater−/− mutant flies upon oral infection by bacterial strains in either the Db10 (A) or the RM66262 (B) genetic background. (C–D) Survival test of wA5001 flies upon oral infection by bacterial strains in either the Db10 (C) or the RM66262 (D) genetic background. (E) Survival test of wA5001 after injection of bacteria at OD600nm = 0.1. The slightly faster lethality observed after a challenge with the wild-type strain was not confirmed in subsequent experiments. Each graph represents one out of three independent experiments that yielded similar results. Error bars represent the standard error. Statistical tests were performed using Log-rank.
The flagellum is essential for S. marcescens to traverse the epithelial barrier
The bacteria in the gut lumen of flies are subjected to various stressors such as immune effectors and digestive enzymes. To monitor the survival of the fliR mutant in the digestive tract, we applied an assay consisting in the ingestion of bacteria that constitutively express GFP from a plasmid together with the propidium iodide stain: the GFP label indicates the presence of live bacteria, whereas the propidium iodide penetrates and stains only dead bacteria. We found that the intestinal lumen of wA5001 flies that ingested RM66262 wild-type, flhD or fliR mutants contained only live bacteria marked with GFP (Fig. 2A,B) as compared to the lumen of flies that have ingested the E. coli control. The absence of propidium iodide staining for the tested S. marcescens strains suggests that the mutants are not killed in the midgut at least at 4 h post-infection, whereas E. coli was killed in the posterior midgut after having passed through the acidic region (Fig. 2A,B). In addition, we measured the bacterial titer in the midgut of eater flies at 24 h post-infection. We observed that the CFU count of fliR mutants in the intestine is comparable to the values determined for either the flhD mutant or the wild-type bacteria (Fig. 2C). Taken together, these results indicate that flhD and fliR mutants are able to resist to the stressful environment of the midgut as well as wild-type bacteria.
flhD and fliR mutants survive in the Drosophila midgut like wild-type S. marcescens but have decreased ability to cross the epithelial barrier. (A) Confocal pictures of wA5001 midguts after ingestion of GFP-expressing bacteria (green) and propidium iodide (PI) (red). Flies were fed on filter pads containing a mix of bacteria (OD600nm = 10) and PI (50 µg/mL) for 4 h at 25 °C. (B) Quantification of wA5001 midguts with live or live and dead bacteria in the RM66262 background. Number of midguts per column = 10–12. (C) CFU count of bacteria in the midgut 24 h post-infection. eater−/− mutant flies were fed with bacteria in the RM66262 background (OD600nm = 0.1) at 25 °C. Each dot in the graph represent one infected midgut. Number of midguts per column = 30. (D) CFU count of bacteria in the hemolymph 4 h post-infection. eater−/− mutant flies were orally infected by bacteria in the RM66262 background (OD600nm = 0.1). Number of dots per column = 9–12. Statistical test was performed using Kruskal–Wallis and Dunn’s post-hoc tests (C, D). IPTG (0.5 mM) was added to the fliR rescue strain. Each graph represents one out of three independent experiments that yielded similar results, except for graph (D) that represent the pooled data of three independent experiments.
Besides motility, adherence to and invasion of host cells are two other important functions of the flagellum43,44. Since flagellum mutants are not motile, we tested the ability of both flhD and fliR mutant bacteria to adhere to and invade CHO cells by forcing the contact between bacteria and host cells by centrifugation. As expected, we observed a decreased adhesion and invasion for both mutants in comparison to wild-type bacteria (Supplementary Fig. 5A,B). We also found that the invasion of Drosophila S2 cells by fliR bacteria is highly diminished when compared to wild-type bacteria (Db10) (Supplementary Fig. 5C). Therefore, these impaired functions of the flagellum may affect the ability of S. marcescens to traverse the intestinal epithelium and to cause septicemia. Therefore, to examine the ability of the bacterial mutants to cross the epithelial barrier, we quantified the amount of bacteria present in the hemolymph of phagocytosis-impaired flies 4 h after the beginning of RM66262 ingestion. We showed that both flhD and fliR bacteria were less abundant in the hemolymph as compared to wild-type bacteria (Fig. 2D). However, the ability of the fliR mutant to cross the intestinal barrier was not rescued by the complementation (Fig. 2D). Of note, the latter is carried out under the control of an inducible promoter that requires IPTG. The IPTG used to activate the expression of the fliR gene may have not been able to pass the intestinal barrier.
These results were confirmed upon dissection of wild-type midguts after gentamicin solution feeding to clear previously ingested bacteria remaining in the lumen. As gentamicin is not able to cross eukaryotic membranes, the microbial titer measured in the treated midguts corresponded to bacteria within ECs or adhering to the basal part of the epithelium, which is in contact with hemolymph10. We observed less fliR bacterial loads than wild-type ones (Supplementary Fig. 5D). In conclusion, both flagellum mutants exhibit difficulties to traverse the intestinal barrier. These findings pinpoint a requirement for the flagellum in the passage of the bacteria from the gut lumen to the body cavity.
The fliR gene is required for S. marcescens to impact the homeostasis of the intestinal epithelium
Following the ingestion of S. marcescens, two distinct resilience mechanisms are activated in the intestinal epithelium: the cytoplasmic purge and the compensatory proliferation of ISCs. In the early phase of infection, pore-forming toxins such as hemolysin elicit the extrusion of EC cytoplasm21. This short-term cytoplasmic purge prevents the toxic effect of the hemolysin on the ECs and results in a drastic thinning of the intestinal epithelium 3 h post-infection. We examined the induction of the cytoplasmic purge by measuring the thickness of the intestinal epithelium 3 h post-infection. The cytoplasmic purge was triggered in midguts infected with fliR, flhD mutants or the wild-type control, as the thinning of the epithelium (~ 10 µm) occurred in midguts infected with either mutant or control strains (Fig. 3A and Supplementary Fig. 6A). Thus, both mutants are toxic enough to trigger the cytoplasmic purge in ECs possibly because they secrete equivalent levels of hemolysin.
In comparison to flhD, fliR mutant bacteria trigger a lessened compensatory proliferation of Drosophila intestinal stem cells. (A) Epithelium thickness measured 3 h post-infection at 25 °C on wA5001 midguts stained with phalloidin (actin). The thickness was measured using the FIJI software, each dot represents the mean of 10 different measurements along the anterior midgut (R2 region). (B) Number of mitoses measured using a PH3 staining in whole midguts of eater−/− mutant flies at 25 °C 24 h post-infection. Flies were orally infected with bacteria in the RM66262 background using OD600nm = 10. IPTG (0.5 mM) was added in the infection solution containing the fliR rescue strain. Each graph represents three independent experiments. Statistical tests were performed using one-way ANOVA.
Despite several midgut defense mechanisms, the bacteria manage to inflict damages to the epithelium, to stress and to kill ECs via unknown virulence factors10,20. Subsequently, ISCs proliferate at 24 h in response to EC stress or death. A phosphohistone H3 (PH3) staining, which marks dividing ISCs in the gut, allows to indirectly monitor the extent of gut damages: an increase in the PH3 level results from an enhanced proliferation of ISC, which may reflect the extent of epithelial damage. Of note, ISC compensatory proliferation in response to EC cell death was previously detected throughout the midgut epithelium of flies that have ingested S. marcescens20. To examine the ability of both mutants to damage the intestinal epithelium, we performed a PH3 staining on eater−/− infected midguts. We detected a significant decrease in the PH3-positive cell count in the midguts infected with fliR mutants as compared to the ones infected with either flhD mutants or wild-type RM66262 bacteria (Fig. 3B). Similar results were obtained following the infection of wA5001 flies with flagellar mutants in the Db10 genetic background (Supplementary Fig. 6B,C). This finding suggests a diminished efficiency for the fliR bacteria to attack the intestinal cells, a process which is at least partially independent from flhD.
Discussion
Intestinal infection with S. marcescens shares similar features with P. aeruginosa oral infection including the passage through the epithelial barrier and the damages to ECs10,22,23. However, the mechanisms used by these two bacterial species to exert these two features has not yet been characterized. Here we have presented evidence that the flagellum of S. marcescens is required for its passage from the gut to the body cavity of the flies. Importantly, we have identified FliR as a novel virulence factor that is needed for the bacteria to severely damage the intestinal epithelium, apparently independently from its major function in building up flagella.
Bacteria can cross the intestinal barrier via two distinct strategies: paracellular/extracellular passage by swimming in between the closely apposed enterocytes through the septate junctions or intracellular passage through the intestinal cells. In this study, we showed that the flagellum of S. marcescens plays a crucial role in the passage of bacteria from the gut lumen to the hemolymph as both flagellar mutants flhD and fliR displayed decreased bacterial loads in the body cavity of the fly (Fig. 2D, Supplementary Fig. 5D). Most S. marcescens bacteria remain confined to the gut endoperitrophic compartment as the peritrophic matrix forms an efficient barrier10. It remains to be determined whether the flagellum is required for the passage through the peritrophic matrix of the few bacteria that manage to cross it. It has been previously shown that some bacteria were attempting to traverse the epithelium in between ECs at late stages of infection10. An open possibility is that for the early passage that occurs within 2 h of feeding, bacteria may cross at the proximal part of the midgut, in the proventriculus region where the peritrophic matrix is synthesized before being reinforced by ECs along the midgut9,10. In both cases, the role of the flagellum may be restricted to its motility function. S. marcescens appears to traverse the intestinal barrier more efficiently when they do not express hemolysin and therefore do not trigger the cytoplasmic purge enterocyte defense21. This observation suggests the possibility that S. marcescens crosses the epithelial barrier by invading intestinal cells, in keeping with a study that also showed that S. marcescens requires the flagellum to adhere to and invade CHO cells44 (Supplementary Fig. 5A,B). The lack of adherence and invasion observed for flagellar mutants can be related to the motility function of the flagellum or to the secretion, through the T3SS, of several virulence factors such as the phospholipase or the S. marcescens nuclease38,45. We note that in Caulobacter crescentus, the synthesis of the type IV pilus, which plays a primordial role in adherence, depends on flagellar genes for the production of pilin46.
Altogether, our results suggest that the escape of a few bacteria into the hemocoel does not contribute to the fatal outcome of the infection as the flhD mutant is as lethal as wild-type bacteria. This result is in keeping with the low bacterial burden detected in the hemolymph throughout the infection, which is limited by hemocytes that phagocytose S. marcescens10.
Here we showed that fliR mutants are less virulent in the intestinal infection model when compared to flhD mutant bacteria in two distinct S. marcescens strains (Fig. 1A–D). This difference in the virulence observed between flhD and fliR mutants implies a possible role for fliR in the virulence of the bacteria independently from its role in the flagellum or T3SS formation, which are both impaired in fliR and flhD mutants. A function of some, including a putative fliR homologue, but not all, flagellar genes in type IV pilus-dependent twitching motility has been reported in the nonflagellated bacterium Lysobacter enzymogenes, highlighting the possibility that some flagellar components have functions in bacterial physiology beyond the synthesis of the flagella47. Also, the fliK flagellar gene in Bacillus thurengiensis is required to counteract Drosophila melanogaster antimicrobial host defenses and is required for virulence independently of its role in the biosynthesis of the flagellum48.
On the one hand, the analysis of the epithelial thickness revealed a normal induction of the cytoplasmic purge following the infection with the fliR mutant bacteria (Fig. 3A and Supplementary Fig. 6A). This purge is triggered in response to the release of the hemolysin pore-forming toxin by S. marcescens. Therefore, the role of fliR in the virulence of the bacteria is not related to the secretion of hemolysin. Indeed, hemolysin is secreted by a T5SS27,49,50. On the other hand, we showed that the fliR strain is likely to induce less damage to the intestinal epithelium as the proliferation rate of ISCs is diminished in the midguts infected with fliR as compared to the ones infected with flhD bacteria (Fig. 3B and Supplementary Fig. 6B,C). This finding strongly suggests that FliR is implicated in the attack and the death of the ECs independently from its function in the flagellum. An attractive hypothesis is that FliR may be needed for the formation of a distinct secretory apparatus required for the secretion of unknown virulence factors that may directly attack and kill the fly intestinal cells (Fig. 4).
An flhD-independent function of fliR in the Drosophila intestinal infection model. (Left) The flhD-dependent function of fliR is restricted to its role in the flagellum formation. The flagellum is required for S. marcescens to traverse the epithelial barrier to reach the hemolymph. (Right) fliR acquired a function in the virulence of the bacteria independently from its role in the flagellum. fliR may be needed for the assembly of an flhD-independent secretory apparatus that would release virulence factors that may damage the intestinal cells.
Methods
Fly strains
Flies were raised at 25 °C with 60% humidity on a semi-solid standard medium composed of 50 L of sterile water containing 3.2 kg of cornmeal, 2.4 kg of sugar, 580 g of yeast brewer’s dry powder, 240 g of agar and 260 g of 4-hydroxybenzoate sodium salt (Merck). The different fly strains used in the experiments were: wA5001 and eater−/−51.
Bacterial strains and culture
Two strains of S. marcescens were used: Db1041 and RM6626242. The different mutants were generated by the pKNOCK plasmid insertion technique39. This plasmid carries antibiotic resistance to chloramphenicol (20 µg/mL) or to gentamicin (15 µg/mL). The fliR; pBB1:lacI:fliR (fliR; pBB1::fliR) strain expresses a wild-type copy of fliR under the control of an Isopropyl ß-D-1-thiogalactopyranoside (IPTG)-inducible promoter. The bacteria were cultured overnight on LB agar plates or in liquid medium at 37 °C with the corresponding antibiotics.
Infection experiments and survival
The oral infection and the septic injury were performed at 25 °C essentially as described in Nehme et al.10. Bacterial pellet was diluted in 50 mM sucrose solution and 10% LB for the oral infection or in PBS for septic injury to a final OD600 of 0.1, 1 or 10 as needed. The survival of oral infected flies was monitored every day and 200 µL of 100 mM sucrose was added daily to the tubes.
Bacterial loads
The bacterial titer of the intestine was measured 24 h post-infection. A single midgut was dissected and homogenized in 100 µL of PBS. The bacterial titer in the hemolymph was determined 4 h post-infection. The hemolymph was retrieved from five flies using a Nanoject II microinjector (Drummond) and collected in 10 µL of PBS. A serial dilution was applied on the samples, then each dilution was plated on LB-agar plates with ampicillin for the RM66262 wild-type strain and mutants thereof.
Staining and imaging
To perform a propidium iodide staining, the flies were fed for 4 h with a solution containing 50 mM sucrose, 10% LB, bacteria that constitutively express GFP from a plasmid (OD600 of 10) and 50 µg/mL of propidium iodide. The midguts were dissected in PBS, fixed with 8% PFA then washed three times with PBS.
To measure epithelial thinning, midguts were dissected and fixed as described above. Actin staining was performed by incubating the samples for 1h30 in 10 µM of FITC-labeled phalloidin (Sigma-Aldrich #P5282). The epithelium thickness was measured using the FIJI software. The PH3 staining was performed at 24 h post-infection (OD600 of 10). The midguts were dissected in PBS, fixed with 8% PFA, incubated with the PH3 antibody (Millipore, ref 09-797) overnight at 4 °C, then stained with an anti-rabbit FITC-labeled antibody (Abcam #6717) overnight at 4 °C or 2 h at room temperature. All stained midguts were mounted in the Vectashield mounting medium (Vector Laboratories). The samples were observed and imaged using a LSM780 confocal microscope (Zeiss).
Statistical analysis
All graphs and statistical tests were performed using GraphPad Prism. The statistical test used for the survival curves was Log-rank. Mann–Whitney, one-way ANOVA or Kruskal Wallis tests were performed for all other experiments (as specified in figure legends). The number of stars (*) represents the P values P ≥ 0.05 (ns), P < 0.05 (*), P < 0.01 (**), P < 0.001 (***) and P < 0.0001 (****).
Data availability
All data and materials are available upon request.
References
Ferrandon, D. The complementary facets of epithelial host defenses in the genetic model organism Drosophila melanogaster: From resistance to resilience. Curr. Opin. Immunol. 25, 59–70 (2013).
Medzhitov, R., Schneider, D. S. & Soares, M. P. Disease tolerance as a defense strategy. Science 335, 936–941 (2012).
Soares, M. P., Gozzelino, R. & Weis, S. Tissue damage control in disease tolerance. Trends Immunol. 35, 483–494 (2014).
Soares, M. P., Teixeira, L. & Moita, L. F. Disease tolerance and immunity in host protection against infection. Nat. Rev. Immunol. 17, 83–96 (2017).
Limmer, S., Quintin, J., Hetru, C. & Ferrandon, D. Virulence on the fly: Drosophila melanogaster as a model genetic organism to decipher host-pathogen interactions. Curr. Drug Targets 7, 978–999 (2011).
Kuraishi, T., Hori, A. & Kurata, S. Host-microbe interactions in the gut of Drosophila melanogaster. Front. Physiol. 4, 1–8 (2013).
Liu, X., Hodgson, J. J. & Buchon, N. Drosophila as a model for homeostatic, antibacterial, and antiviral mechanisms in the gut. PLoS Pathog. 13, 10–17 (2017).
Lemaitre, B. & Miguel-Aliaga, I. The digestive tract of Drosophila melanogaster. Annu. Rev. Genet. 47, 377–404 (2013).
Kuraishi, T., Binggeli, O., Opota, O., Buchon, N. & Lemaitre, B. Genetic evidence for a protective role of the peritrophic matrix against intestinal bacterial infection in Drosophila melanogaster. Proc. Natl. Acad. Sci. 108, 15966–15971 (2011).
Nehme, N. T. et al. A model of bacterial intestinal infections in Drosophila melanogaster. PLoS Pathog. 3, 1694–1709 (2007).
Tzou, P. et al. Tissue-specific inducible expression of antimicrobial peptide genes in Drosophila surface epithelia. Immunity 13, 737–748 (2000).
Ryu, J. H. et al. An essential complementary role of NF-κB pathway to microbicidal oxidants in Drosophila gut immunity. EMBO J. 25, 3693–3701 (2006).
Liehl, P., Blight, M., Vodovar, N., Boccard, F. & Lemaitre, B. Prevalence of local immune response against oral infection in a Drosophila/Pseudomonas infection model. PLoS Pathog. 2, 0551–0561 (2006).
Ha, E.-M., Oh, C.-T., Bae, Y. S. & Lee, W. J. A direct role for dual oxidase in drosophila gut immunity. Science 310, 847–850 (2005).
Lee, K. A. et al. Bacterial-derived uracil as a modulator of mucosal immunity and gut-microbe homeostasis in Drosophila. Cell 153, 797–811 (2013).
Ha, E. M. et al. An antioxidant system required for host protection against gut infection in Drosophila. Dev. Cell 8, 125–132 (2005).
Biteau, B., Hochmuth, C. E. & Jasper, H. JNK activity in somatic stem cells causes loss of tissue homeostasis in the aging drosophila gut. Cell Stem Cell 3, 442–455 (2008).
Buchon, N. et al. Invasive and indigenous microbiota impact intestinal stem cell activity through multiple pathways in Drosophila. GENES Dev. 23, 2333–2344 (2009).
Jiang, H. et al. Cytokine/Jak/Stat signaling mediates regeneration and homeostasis in the Drosophila midgut. Cell 137, 1343–1355 (2009).
Cronin, S. J. et al. Genome-wide RNAi screen identifies genes involved in intestinal pathogenic bacterial infection. Science 325, 340–343 (2009).
Lee, K. Z. et al. Enterocyte purge and rapid recovery is a resilience reaction of the gut epithelium to pore-forming toxin attack. Cell Host Microbe 20, 716–730 (2016).
Apidianakis, Y., Pitsouli, C., Perrimon, N. & Rahme, L. Synergy between bacterial infection and genetic predisposition in intestinal dysplasia. Proc. Natl. Acad. Sci. 106, 20883–20888 (2009).
Limmer, S. et al. Pseudomonas aeruginosa RhlR is required to neutralize the cellular immune response in a Drosophila melanogaster oral infection model. Proc. Natl. Acad. Sci. 108, 17378–17383 (2011).
Hejazi, A. & Falkiner, F. R. Serratia marcescens. J. Med. Microbiol. 46, 903–912 (1997).
Iosifidis, E. et al. Outbreak of bloodstream infections because of Serratia marcescens in a pediatric department. Am. J. Infect. Control 40, 11–15 (2012).
Maltezou, H. C. et al. Consecutive Serratia marcescens multiclone outbreaks in a neonatal intensive care unit. Am. J. Infect. Control 40, 637–642 (2012).
Hertle, R. Serratia type pore forming toxins. Curr. Protein Pept. Sci. 1, 75–89 (2000).
Kurz, C. L. et al. Virulence factors of the human opportunistic pathogen Serratia marcescens identified by in vivo screening. EMBO J. 22, 1451–1460 (2003).
Di Venanzio, G., Lazzaro, M., Morales, E. S., Krapf, D. & García Véscovi, E. A pore-forming toxin enables Serratia a nonlytic egress from host cells. Cell Microbiol. 19, e12656 (2017).
Marty, K. B., Williams, C. L., Guynn, L. J., Benedik, M. J. & Blanke, S. R. Characterization of a cytotoxic factor in culture filtrates of Serratia marcescens characterization of a cytotoxic factor in culture filtrates of Serratia marcescens. Infect. Immun. 70, 1121–1128 (2002).
Shanks, R. M. Q. et al. Identification of slpB, a cytotoxic protease from Serratia marcescens. Infect. Immun. 83, 2907–2916 (2015).
Bruna, R. E., Molina, M. V., Lazzaro, M., Mariscotti, J. F. & García Véscovi, E. CpxR-dependent thermoregulation of Serratia marcescens PrtA metalloprotease expression and its contribution to bacterial biofilm formation. J. Bacteriol. 200, 1–18 (2018).
Shimuta, K. et al. The hemolytic and cytolytic activities of Serratia marcescens phospholipase A (PhlA) depend on lysophospholipid production by PhlA. BMC Microbiol. 9, 1–10 (2009).
Ferrandon, D., Imler, J. L., Hetru, C. & Hoffmann, J. A. The Drosophila systemic immune response: Sensing and signalling during bacterial and fungal infections. Nat. Rev. Immunol. 7, 862–874 (2007).
Pradel, E. et al. Detection and avoidance of a natural product from the pathogenic bacterium Serratia marcescens by Caenorhabditis elegans. Proc. Natl. Acad. Sci. 104, 2295–2300 (2007).
Apel, D. & Surette, M. G. Bringing order to a complex molecular machine: The assembly of the bacterial flagella. Biochim. Biophys. Acta - Biomembr. 1778, 1851–1858 (2008).
Zhuang, W. Y. & Shapiro, L. Caulobacter fliQ and fliR membrane proteins, required for flagellar biogenesis and cell division, belong to a family of virulence factor export proteins. J. Bacteriol. 177, 343–356 (1995).
Givskov, M., Eberl, L., Christiansen, G., Benedik, M. J. & Molin, S. Induction of phospholipase- and flagellar synthesis in Serratia liquefaciens is controlled by expression of the flagellar master operon flhD. Mol. Microbiol. 15, 445–454 (1995).
Alexeyev, M. F. The pKNOCK series of broad-host-range mobilizable suicide vectors for gene knockout and targeted DNA insertion into the chromosome of gram-negative bacteria. Biotechniques 26, 824–828 (1999).
Kovach, M. E. et al. Four new derivatives of the broad-host-range cloning vector pBBR1MCS, carrying different antibiotic-resistance cassettes. Gene 166, 175–176 (1995).
Flyg, C., Kenne, K. & Boman, H. G. Insect pathogenic properties of Serratia marcescens: Phage-resistant mutants with a decreased resistance to cecropia immunity and a decreased virulence to Drosophila. J. Gen. Microbiol. 120, 173–181 (1980).
Bruna, R. E. et al. Draft whole-genome sequence of Serratia marcescens strain RM66262, isolated from a patient with a urinary tract infection. Genome Announc. 3, e01423-e1515 (2015).
Chaban, B., Hughes, H. V. & Beeby, M. The flagellum in bacterial pathogens: For motility and a whole lot more. Semin. Cell Dev. Biol. 46, 91–103 (2015).
Fedrigo, G. V., Campoy, E. M., Di Venanzio, G., Colombo, M. I. & Véscovi, E. G. Serratia marcescens is able to survive and proliferate in autophagic-like vacuoles inside non-phagocytic cells. PLoS ONE 6, e24054 (2011).
Liu, J. H. et al. Role of flhDC in the expression of the nuclease gene nucA, cell division and flagellar synthesis in Serratia marcescens. J. Biomed. Sci. 7, 475–483 (2000).
Ellison, C. K., Rusch, D. B. & Brun, Y. V. Flagellar mutants have reduced pilus synthesis in Caulobacter crescentus. J. Bacteriol. 201, e00031-e119 (2019).
Fulano, A. M., Shen, D., Kinoshita, M., Chou, S. H. & Qian, G. The homologous components of flagellar type III protein apparatus have acquired a novel function to control twitching motility in a non-flagellated biocontrol bacterium. Biomolecules 10, 733 (2020).
Attieh, Z. et al. The fliK gene is required for the resistance of Bacillus thuringiensis to antimicrobial peptides and virulence in Drosophila melanogaster. Front. Microbiol. 11, 1–16 (2020).
Petersen, L. M. & Tisa, L. S. Friend or foe? A review of the mechanisms that drive Serratia towards diverse lifestyles. Can. J. Microbiol. 59, 627–640 (2013).
Schiebel, E., Schwarz, H. & Braun, V. Subcellular location and unique secretion of the hemolysin of Serratia marcescens. J. Biol. Chem. 264, 16311–16320 (1989).
Kocks, C. et al. Eater, a transmembrane protein mediating phagocytosis of bacterial pathogens in Drosophila. Cell 123, 335–346 (2005).
Acknowledgements
We are grateful to Drs. P. Giammarinaro, S. Niehus, J. Mariscotti, R. Bruna, and J. Nguyen for help in some experiments; W.J. Lee and the Bloomington Stock Center (NIH P40OD018537) for fly stocks. A.A has been supported by the Higher Education Commission Pakistan, G.D.V and M.La by ANPCyT (Agencia Nacional de Promoción Científica y Tecnológica, Argentina) and CONICET (Consejo Nacional de Investigaciones Cientifícas y Técnicas, Argentina), and B.S.R by an IdEx fellowship (Université de Strasbourg-France). The work in E.G.V laboratory has been funded by CONICET (Argentina) and the work in D.F laboratory has been funded by CNRS (Centre National De La Recherche Scientifique), Fondation pour la Recherche Médicale (Equipe FRM to DF), and ANR (Agence Nationale de la Recherche, France) (DROSOGUT, ENTEROCYTE_PURGE_RECOVERY). The D.F and E.G.V laboratories have developed a collaboration within the framework of the ECOS-Sud (France)-MINCT (Ministerio deCiencia y Tecnología e Innovación) exchange program (#A12B04).
Author information
Authors and Affiliations
Contributions
A.A. performed the screen; G.D.V., M.La and E.G.V. designed and analyzed the in vitro experiments; M.La, G.D.V and B.S.R performed the in vitro experiments; M.Le, B.S.R. and D.F. designed and analyzed the in vivo experiments; M.W.Y., M.Le, B.S.R., and S.L. performed the in vivo experiments. B.S.R. and D.F. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sina Rahme, B., Lestradet, M., Di Venanzio, G. et al. The fliR gene contributes to the virulence of S. marcescens in a Drosophila intestinal infection model. Sci Rep 12, 3068 (2022). https://doi.org/10.1038/s41598-022-06780-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-06780-w
This article is cited by
-
Shaping the Microbial Landscape: Parasitoid-Driven Modifications of Bactrocera dorsalis Microbiota
Microbial Ecology (2024)
-
The pivotal roles of gut microbiota in insect plant interactions for sustainable pest management
npj Biofilms and Microbiomes (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.