Abstract
Recently it was proposed that the redox status of cysteines acts as a redox switch to regulate both the oligomeric status and the activity of human dUTPase. In a separate report, a human dUTPase point mutation, resulting in a tyrosine to cysteine substitution (Y54C) was identified as the monogenic cause of a rare syndrome associated with diabetes and bone marrow failure. These issues prompt a critical investigation about the potential regulatory role of cysteines in the enzyme. Here we show on the one hand that independently of the redox status of wild-type cysteines, human dUTPase retains its characteristic trimeric assembly and its catalytic activity. On the other hand, the Y54C mutation did not compromise the substrate binding and the catalytic properties of the enzyme at room temperature. The thermal stability of the mutant protein was found to be decreased, which resulted in the loss of 67% of its activity after 90 min incubation at the physiological temperature in contrast to the wild-type enzyme. In addition, the presence or absence of reducing agents had no effect on hDUTY54C activity and stability, although it was confirmed that the introduced cysteine contains a solvent accessible thiol group.
Similar content being viewed by others
Introduction
Preservation of genome integrity is essential for viability and is accomplished by efficient interlinked regulatory pathways of DNA damage recognition / repair and nucleotide biosynthesis. In eukaryotes, these processes are generally compartmentalized in the nucleus and in the mitochondrion. These two organelles represent altered redox milieu, as shown by several independent experimental approaches (e.g. refs1,2,3, cf also for a review4). It is also of importance to provide functionality of organelle-specific genomic repair and nucleotide biosynthesis in line with the actual redox status (cf e.g. refs5,6,7,8,9, cf also for a review10).
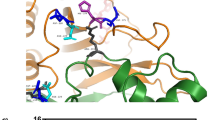
The dUTPase enzymes play a dual role in sanitizing the nucleotide pool and in providing precursor for thymidylate biosynthesis11,12. It has been established that in humans and in several other eukaryotes, two dUTPase isoforms are present, one nuclear and one mitochondrial and their expression is under control of alternative promoters13,14. The human dUTPase isoforms differ only in their respective intracellular trafficking signals: a well characterized nuclear localization signal13,15 for the nuclear isoform and a mitochondrial leader sequence for the mitochondrial isoform13 (Fig. 1A). With regard to the different redox status of the nucleus and the mitochondrion, it is important to discuss the location of cysteine residues in the structure. There are two cysteine residues (Cys78 and Cys134) within the conserved major part of the human dUTPase isoforms, which are buried within the well folded structure, more than 5 Å distance from the active site. There is one additional cysteine on the flexible N-terminus of the nuclear isoform (Cys3), which could not be included in the 3D structure based on X-ray crystallographic data as the electron density map is poorly defined around this mobile section of the protein16 (Fig. 1B). The mature mitochondrial isoform that is produced after the mitochondrial import by cleaving the organelle-specific leader segment, does not contain this latter, potentially solvent exposed cysteine.
(A) Sequences of the nuclear (nDut) and mitochondrial (mDut) isoforms of the human dUTPase (Uniprot ID: P33316-2 and -3). Numbering of nuclear isoform of the human dUTPase residues is shown above the sequences. Stars denote the identical residues. In case of mDut the functional protein sequence remaining after the removal of mitochondrial localization sequence is shown in bold. Residues included in the crystal structure shown on Panel A are underlined. Cysteines are with yellow outline, position of Y54C mutation is coloured by red. Residues of the five conserved motif are coloured blue. (B) Crystal structure of the human dUTPase (PDB ID: 2HQU). Protein is shown as cartoon, chains are coloured cyan, salmon and green. Substrate analogue 2ʹ-deoxyuridine 5ʹ-alpha,beta-imido-triphosphate (dUPNPP) is shown as spheres with atomic coloring (carbon: black, oxygen: red, nitrogen:blue, phosphorus:orange). Residues within 5 Å distance from the substrate analogue are shown as white partially transparent surface. Cysteine side chains are shown as sticks with atomic coloring (carbon: magenta, sulfur: yellow), labels of cysteines are shown. Both Cys-78 and Cys-134 residues are in the globular core of the protein more than 5 Å distance from the active site of the protein.
dUTPases belong to two enzyme families, the all-alpha and the all-beta dUTPase family11,17. Most dUTPases, including mammalian dUTPases are all-beta dUTPases18. All-beta dUTPases are homotrimers, constituted by intrinsically folded beta-pleated subunits13,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37. The three active sites are found within the clefts formed by subunit interfaces. In most cases, the C-terminal part of a given subunit folds back to the remote active site in such a manner that each active site involves conserved residues from all the three subunits (Fig. 1B). This fold is highly specific and is strictly conserved in the trimeric dUTPase family. The well-folded core is frequently complemented by lineage-specific N-terminal and C-terminal flexible extensions38,39, or by intrastrand extensions that fold out from the core24,29,32,35,40. In eukaryotes the flexible N-termini usually encode organelle specific signal sequences for intracellular trafficking11,13,38.
It has been proposed that the N-terminal cysteine of the nuclear isoform leads to formation of a covalently linked dimer through a disulfide bridge between Cys3 residues with regulatory consequences41. Rotoli et al. also claimed that this disulfide bridge is essential for activity based on experiments indicating inactivation of the enzyme by the Cys3Ala mutation or by the addition of excess reducing agents41. The proposal of an enzymatically active covalently linked dimeric state of human dUTPase is highly intriguing. However, it cannot be reconciled in the current structure–function knowledge on this enzyme, which clearly indicated an indispensable condition for the homotrimeric assembly to form functional active sites based on all available 3D crystal structure data and solution studies13,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,42,43. Although an artificial covalent dimer construct of Drosophila virilis dUTPase could have been created, it showed decreased enzymatic activity as compared to the respective trimeric enzyme21.
These results of Rotoli et al. are also in contradiction with data reported for full enzymatic activity preserved in an N-terminally truncated human dUTPase construct (M24-N164) lacking the flexible N-terminus and the Cys3 residue located within. The kcat for this enzyme construct was reported as 8.2 ± 0.6 s−1 (37 °C)19 to be compared with kcat = 6.8 ± 2.0 s−1 (20 °C) for the full-length and His-tagged protein44. Also, limited trypsinolysis leading to cleavage of the flexible N-terminus of human dUTPase did not lead to any loss of enzymatic activity16.
dUTPase is an enzyme essential for viability and its regulation is of key importance, therefore in the present study we set out to shed light on this contradiction. The essential character of dUTPase has been well established in most free-living species40,45,46,47,48,49,50 investigated so far, with some potential exceptions to this general rule in some prokaryotes51. Potentially in line with the importance of dUTPase in cell viability, no human dUTPase knock-outs could have been obtained to date. However, silencing studies reinforced the essentiality of human dUTPase and its key role in regulating thymidylate metabolism31,52,53,54,55. It has also been shown that dUTPase overexpression in cancers decrease the effectivity of thymidylate-synthase targeted chemotherapy (e. g. 5-fluorouracil, methotrexate)54,55,56,57 and is involved in development of resistance against the treatment58,59. Interestingly, a human dUTPase mutation (Tyr54Cys) located outside the conserved region was suggested to be responsible for monogenic syndrome associated with diabetes type 2 and bone marrow failure53. This report was the first and to date the only case where a dUTPase mutation has been associated with human disease. It was suggested that the mutation affects enzymatic activity, however, no kinetic data was reported. It was therefore of immediate interest to characterize the structural and functional consequences of this mutation.
Results
Redox status of cysteines does not alter functional properties of human dUTPase
First, we set out to explore whether the function of human dUTPase is changed depending on the reducing or non-reducing experimental conditions. For these investigations, we chose the hDUTF158W quasi-wild-type human dUTPase variant, which has the same sequence as the nuclear isoform (Fig. 1A, nDut), but a tryptophan sensor replaces Phe-158 at its active site (Fig. 2A). The effect of this tryptophan has been characterized in the literature16,44,60. The F/W substitution in human dUTPase was found not to affect the activity and substrate binding properties of the enzyme, hence this construct is considered to be a quasi-wild-type human dUTPase. Besides, the tryptophan substitution allowed the exploration of the kinetic mechanism of this enzyme, since the aromatic residue serves as a sensitive sensor to follow substrate/product binding and release and thus the enzymatic cycle of human dUTPase, even on the millisecond scale (Fig. 2B,C).
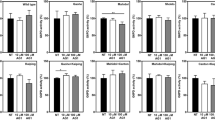
(A) Model structure of the F158W mutant human dUTPase. The model was created with PyMOL Mutagenesis tool from the crystal structure of the human dUTPase (PDB ID: 2HQU). Protein is shown as cartoon, chains are coloured cyan, salmon and green. Substrate analogue dUPNPP is shown as sticks with atomic coloring (carbon: black, oxygen: red, nitrogen: blue, phosphorus: orange). The sidechain of the introduced tryptophan sensor (F158W) is shown as sticks with atomic coloring (carbon: grey, nitrogen: blue). Magnesium ion is shown as green sphere, black dashed lines represent the polar contacts of the magnesium ion with the phosphate chain of the substrate analogue dUPNPP. The built in F158W tryptophan sensor can detect the presence or the absence of the substrate or the substrate analogue, thus the reaction can be followed by tryptophan fluorescence as shown on Panel B. (B) Representative transient kinetic reaction curves of hDUTF158W followed by tryptophan fluorescence. The bright green reaction curve was recorded under single turnover (substrate concentration < active site concentration), while the dark green reaction curve under multiturnover conditions. The fluorescence decreases upon substrate binding and increases upon substrate/product release. The product release is rate limited by the hydrolysis event (see Panel C also). Under multiturnover conditions, a longer low fluorescent phase equivalent to steady-state hydrolysis can be observed. Single turnover curves can, while the multiturnover reaction curves can not be fitted with triple exponential function. (C) Schematic representation of dUTPase kinetic mechanism. The “†” marks represent fluorescent quenching. The rate constants of the steps which can be detected by tryptophan fluorescence in stopped flow under single turnover conditions are indicated on the reaction scheme. However, under single turnover conditions the rate constants of the collision complex formation (k1, k−1) and the isomerization (k2, k−2) can not be directly determined. Under these conditions only “observed binding rate constants” can be detected (k1obs, k2obs) that are originated from the net result of the forward and backward reaction of the given step, and are characteristic for the used enzyme and substrate concentration. (D) SDS-PAGE gel of the hDUTF158W protein in the absence and presence of 10 mM β-ME. (Full-length gel image is included in a Supplementary Information.) (E–F) Representative single turnover fluorescence reaction curves of hDUTF158W in the presence (E) and in the absence (F) of 10 mM β-ME. The reaction curves were fitted with triple exponential function (Eq. 1) which yielded the following parameters A1 = 3.95 ± 0.11, A2 = 1.15 ± 0.01, A3 = − 5.02 ± 0.004, k1 = 730.2 ± 15.3, k2 = 96.6 ± 1.5, k3 = 6.58 ± 0.01, y0 = 7.89 ± 0.0009 for Panel E and A1 = 4.12 ± 0.36, A2 = 0.39 ± 0.04, A3 = − 3.14 ± 0.003, k1 = 1024.3 ± 55.0, k2 = 193.9 ± 14.4, k3 = 7.0 ± 0.01, y0 = 7.70 ± 0.0009 for Panel F (the errors represent the error of the fitted parameters). Several (3–5) single turnover reaction curves were recorded, and the average and standard deviation of the resulted rate constants are shown in Table 1.
We purified this protein in the absence of any reducing agent. The SDS-PAGE gel of this enzyme preparation (Fig. 2D) indicated that in addition to the expected protein band at the ~ 19 kDa position (corresponding to a protein monomer), an extra band was also observed at around 38 kDa. This position matches with the molecular mass of a human dUTPase dimer.
As it was suggested by Rotoli et al. human dUTPase dimers may be formed by cysteine disulfide bonds in the absence of reducing agent. To see whether the second band is due to covalent intersubunit cysteine bridges, we added 10 mM 2-mercaptoethanol (β-ME) to the solution. Elimination of the band at 38 kDa position by addition of reducing agent (Fig. 2D) suggested that this band indeed resulted from covalent intersubunit link formation through cysteines. Thus, we observed that intersubunit disulfide bond between two human dUTPase subunits can be detected in the absence of reducing agent, although this observation was reached under denaturing conditions, i.e. by SDS-PAGE. It has to be noted that under these conditions the protein chain is unfolded, hence the covalent dimer may not be relevant for the native state (i.e. it may be an artefact). In our former SAXS experiments the nuclear isoform of the human dUTPase could only be detected as trimers in solution42,43. This result does not exclude the possibility of an intersubunit disulfide bond through Cys3 residues between two subunits within one trimer. Such an intersubunit disulfide bond would result in a dimer band on a non-reducing SDS-PAGE gel.
To gain relevant data about oligomerization of hDUT under native conditions we also investigated the oligomeric state of hDUT with native mass spectrometry (MS) and chemical crosslinking experiments (Supplementary Fig. 1). Trimers were the most abundant form of hDUTF158W (61%) in the mass spectra; monomers were also detected in considerable amount (36%), while only few dimers were present (3%) (Supplementary Fig. 1A). The SDS-PAGE analysis of purified hDUTF158W samples crosslinked with disuccinimidyl suberate, an agent of 11 Å-spacer length, indicated crosslinked trimers, dimers and monomers as well (Supplementary Fig. 1B). In contrast to our present crosslinking results, when formaldehyde crosslinking experiment was performed on human cells, no trimers from the human dUTPase nuclear isoform were observed41. We propose that this contradiction is due to the small size of formaldehyde (zero-length crosslinker), which limits its ability to crosslink larger assemblies and the low amount of dUTPase protein within the cell. Based on our results acquired with three independent experimental methods (previous SAXS results43, together with the present native mass spectrometry and crosslinking experiments), we suggest that the trimeric form of the nuclear isoform of human dUTPase is abundant in solution.
To test whether the presence of the reducing agent influences the activity of human dUTPase we performed transient kinetic experiments. If the active site concentration does not exceed the substrate concentration, then only one substrate molecule will be processed by one active site (single turnover condition). If the protein and substrate concentration is high enough to prevent the rate limitation of complex formation then under single turnover conditions the intrinsic catalytic rate constant of human dUTPase can be determined by triple exponential fitting (Eq. 1)44.
According to our results (Fig. 2E,F; Table 1) the triple exponential function could be fitted to the reaction curve where the enzyme:substrate ratio is 1:1, indicating that (i) the enzymatic mechanism of hDUTF158W remains the same under both reducing and non-reducing conditions, and (ii) 100% of the protein is active under both conditions. We presume that the trimer state is prevalent, as the applied assay buffer is highly similar to the one previously used in non-reducing SAXS experiments43, where only the trimeric form of the enzyme could be detected. Moreover, based on the current structure function knowledge the homotrimeric assembly is necessary for active site formation (cf. for example refs19,44). The 100% activity argues for negligible amount of inactive monomers or dimers of hDUTF158W under both conditions.
Regarding rate constants involved in formation of enzyme:substrate complex, under single turnover conditions two “observed binding rate constants” can be determined (k1obs, k2obs). These observed rate constants (k1obs and k2obs) are related to the intrinsic rate constants (Fig. 2B,E,F)44, however these are concentration dependent. Therefore, we have used the same enzyme and substrate concentrations for reducing and non-reducing conditions to allow correct comparisons. We could not observe any remarkable difference, as presented in Table 1.
We conclude, that even if a covalent link between two subunits of a homotrimer is present in the solution through cysteines under non-reducing conditions, the measured kinetic properties do not differ. Thus, it does not seem likely that the cysteines of human dUTPase would work as a redox switch.
Y54C mutation involved in monogenic diabetes decreases protein stability
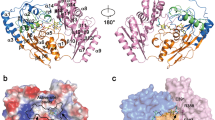
To investigate the effect of Y54C mutation in vitro, we introduced this substitution in the hDUTF158W protein. Based on the crystal structures of the human dUTPase Tyr-54 is situated on the surface of the protein (Fig. 3A–C), thus it was straightforward to assume that the mutant cysteine is also solvent accessible. To verify this hypothesis, we performed a thiol quantitation assay (Fig. 3D). Normalized fluorescence of hDUTF158W,Y54C was found to be significantly higher than that of hDUTF158W (cf. Figure 3E), this difference was consistently observable in a wide concentration range. Thus, we concluded that Cys54 is on the surface of the protein.
(A) Crystal structure of the human dUTPase (PDB ID: 2HQU). Protein is shown as cartoon, chains are coloured cyan, salmon and green. Substrate analogue dUPNPP is shown as spheres with atomic coloring (carbon: black, oxygen: red, nitrogen: blue, phosphorus: orange). Residues within 5 Å distance from the substrate analogue are shown as white partially transparent surface. In case of one of the dUTPase protomers side chains of Tyr-54 and its interaction partner Asp-69 are shown as sticks with atomic coloring (carbon: cyan, oxygen: red). Tyr-54 and Asp-69 residues are more than 5 Å distance from the active site of the protein. Black frame shows the position of close-ups on Panels B and C. (B) Close up of Tyr-54 in one of the dUTPase protomers. Coloring and representation is as on Panel A. Dashed black lines illustrate the hydrogen bond formed between Tyr-54 and Asp-69 (distance of the two residues in Å is indicated on the figure). (C) Close up of the introduced Cys-54 mutation to the protein with PyMOL in most abundant conformation (> 80%) of the residue (cysteine side chain is shown as sticks with atomic coloring: carbon: magenta, sulfur: yellow). Cys-54 and Asp-69 can not form a hydrogen bond as the distance of the sulfur from the oxygen atoms is 5.2 Å and 5.9 Å. (D) Scheme of the thiol quantitation assay. Solvent accessible, free cysteines react with the thiol assay reagent and a fluorophore is formed. The observed fluorescent signal is proportional to the amount of free cysteines in the sample. (E) Results of thiol quantitation assay of hDUTF158W and the hDUTF158W,Y54C. (F) SDS-PAGE gel of the hDUTF158W,Y54C protein in the absence and presence of 10 mM β-ME. (Full-length gel image is included in a Supplementary Information).
The reactivity of this residue in the thiol quantitation assay also proposes that probably these cysteines do not participate in extra disulfide bonds. This was confirmed on SDS-PAGE, where we observed similar pattern under non-reducing, non-native conditions as in the case of hDUTF158W (Fig. 3F to be compared with Fig. 2D and its analysis).
We also performed active site titration and transient kinetic analysis on the mutant enzyme under reducing and non-reducing conditions (Fig. 4A,B, Table 1). The determined kcat value was in good agreement with both the previously published kcat of the hDUTF158W enzyme (6.8 ± 2.0 s−1) determined from transient kinetic measurements44, and with our measurements with the hDUTF158W variant. There was no difference between the reducing and the non-reducing conditions, and again 100% of the protein preparation was active under both conditions.
(A–B) Representative single turnover fluorescence reaction curves of hDUTF158W,Y54C in the presence (A) and in the absence (B) of 10 mM β-ME. The reaction curves were fitted with triple exponential function (Eq. 1) which yielded the following parameters A1 = 3.20 ± 0.14, A2 = 1.2 ± 0.01, A3 = -4.02 ± 0.003, k1 = 877.8 ± 23.9, k2 = 118.8 ± 1.4, k3 = 7.0 ± 0.01, y0 = 6.40 ± 0.0006 for Panel A and A1 = 4.61 ± 0.12, A2 = 1.74 ± 0.01, A3 = − 5.70 ± 0.52, k1 = 728.9 ± 13.1, k2 = 61.4 ± 0.01, k3 = 6.9 ± 0.01, y0 = 8.18 ± 0.001 for Panel B (the errors represent the error of the fitted parameters). Several (3–5) single turnover reaction curves were recorded, and the average and standard deviation of the resulted parameters are shown in Table 1. (C) Differential Scanning Fluorimetry measurements of the hDUTF158W and hDUTF158W,Y54C proteins in the presence and absence of 10 mM β-ME. Melting temperatures are given in Table 1. (D) Stability measurement of the hDUTF158W and hDUTF158W,Y54C proteins. Results of steady-state initial velocity measurements with fixed concentration of dUTP after incubation at 37 °C for various time intervals.
The observed rate constants of substrate binding were also comparable under the two conditions and they did not differ from the observed rate constants determined for hDUTF158W (Table 1).
As previously the mutant enzyme was not characterized, we also performed steady-state activity measurements to determine the Michaelis–Menten parameter of the mutant (Supplementary Fig. 2, Table 1). The determined kcat (6.7 ± 0.7 s−1) and KM (2.31 ± 0.98 µM) values did not differ from that of the wild-type and the hDUTF158W enzyme44. We concluded that the Y54C mutation does not alter the activity and substrate binding properties of the human dUTPase.
To further test the properties of the mutant enzyme we performed differential scanning fluorimetry (DSF) measurements which is applicable for the determination of the melting temperature (TM) of the protein referring to its stability. We have found (Fig. 4C) that the mutant enzyme has about 6 °C lower TM (52.5 ± 0.5 °C) than the hDUTF158W (58.5 ± 0.5 °C). Addition of β-ME did not influence the thermal unfolding of either protein (Fig. 4C). As TM = 52.5 °C is still relatively high compared to the physiological temperature where the enzyme has to function, we performed stability measurements at 37 °C. For this purpose, we incubated the mutant and the hDUTF158W enzyme at 37 °C for different time intervals and then measured activity (Fig. 4D). We found that the mutant enzyme lost its activity faster than hDUTF158W. After 90 min incubation the hDUTF158W variant preserved 75% while the hDUTF158W,Y54C variant only 33% of its activity.
As a structural background, based on the analysis of the interaction network around residue 54 of the wild-type and the Y54C mutant proteins we found only one major change as a consequence of the mutation61. In case of the wild-type protein Tyr-54 forms a hydrogen bond with Asp-69, while the cysteine residue cannot establish the same interaction (Fig. 3A–C). Lack of this hydrogen bond in hDUTF158W,Y54C may be involved in the observed decrease of thermal stability of the mutant protein.
We concluded based on our in vitro investigation that the hDUTF158W,Y54C protein is less stable than the wild-type dUTPase at physiological temperature. This may result in shorter half-life and/or lower activity in vivo.
Discussion
In the present work we focused on the question whether cysteine oxidation may have any effect on human dUTPase structure and activity. Using a combination of independent methods of structural biochemistry and enzyme kinetics, we report that both under reducing and non-reducing conditions the homotrimeric organization is readily visualized and the enzyme activity is fully preserved.
We observed no significant difference of the catalytic rate constant of the quasi-wild-type human dUTPase, hDUTF158W in the presence and absence of 250-times excess reducing agent (ie. 40 μM dUTPase and 10 mM β-ME) (cf. Table 1). In an earlier study steady state activity of a human dUTPase construct (M24-N164) lacking Cys3 was reported to be 8 ± 3 s-1 16, which is in good agreement of the data obtained for the hDUTF158W enzyme with transient kinetics experiments (kcat = 6.8 ± 2.0 s−1)44.
Thus, based on these results we can state that the enzymatic activity of human dUTPase is not dependent on the oxidative state of cysteines. Consequently, regulation of dUTPase activity through redox potential in vivo is unlikely. We therefore respectfully disagree with the previous conclusions of Rotoli et al., who observed marked decrease of enzymatic activity upon addition of 2% β-ME (ca. 289 mM) to the assay buffer. We propose that the observed change is likely to be an artefact caused by the rather high concentration of reducing agent. In the same work Rotoli et al. also observed reduced enzymatic activity in the C3A single point mutant human dUTPase protein, which based on our current results, may have no relation to the redox nature of the protein.
With respect to the proposed presence of dimeric forms of human dUTPase, we note that the method of choice in native mass spectrometry is the Electrospray Ionization (ESI)-MS for detection of protein oligomers62. In the present study we have used ESI–MS and found that the majority of the protein sample corresponds to the trimeric assembly, whereas in the previous report MALDI-TOF was used41, that is much less applicable for studying native protein complexes.
Here we also report, that the intriguing Y54C mutation identified in patients of a monogenic syndrome associated with diabetes and bone marrow failure, also did not change the enzyme activity at room temperature but led to decreased thermal stability. There might be two different explanations for the observed pathological effects in patients carrying this mutation. On the one hand the decreased thermal stability may result in shorter half-life, and in insufficient in vivo activity. In some other systems as well, different enzyme mutations leading to impaired cellular functions may similarly involve temperature sensitivity63,64.
On the other hand, the mutation may perturb protein–protein interactions. This hypothesis is based on the premise that Cys54 is located on the surface of the protein. The significantly increased amount of reactive cysteines observed in the thiol reactivity assay of hDUTF158W,Y54C compared to that of hDUTF158W indicate that the mutant cysteine is indeed on the surface of the protein, thus could potentially interfere with binding of the human dUTPase to other proteins in vivo.
Finally based on our results we conclude that cysteines do not act as redox switches in the human dUTPase. Introduction of cysteine mutation at position 54 of the nuclear isoform of the human dUTPase alters the stability of the protein, which could be the reason why this mutation leads to a monogenic syndrome associated with diabetes and bone marrow failure.
Methods
Cloning and mutagenesis
Site-directed mutagenesis was performed by the QuikChange method (Stratagene) and was verified by sequencing. The enzyme conferring a tryptophan sensor in the active site (hDUTF158W) was used as wild-type16,43,44,60,65,66. The Tyr54 to Cys mutant was created within this construct using the following forward (FW) and reverse (REV) primers: Y54C_FW: 5ʹ CGACCTGTACAGTGCCTGTGATTACACAATACCACCTATGG 3ʹ and Y54C_REV: 5ʹ GCTGGACATGTCACGGTCTCTAATGTGTTATGGTGGATACC 3ʹ. Thus the hDUTF158W and the hDUTF158W,Y54C constructs have the same sequence as the nuclear isoform of the human dUTPase (Fig. 1A, nDut) except the indicated mutations and an N-terminal polyhistidine tag (MAHHHHHHVGT).
Protein expression and purification
Expression and purification of the human dUTPase proteins were done as described previously in 66. Briefly, the hDUTF158W and the hDUTF158W,Y54C enzymes were expressed in BL21 Rosetta (pLysS) cells (Novagen). The protein expression was induced by the addition of 500 µM isopropyl β-D-1-thiogalactopyranoside at OD = 0.5 and was conducted for 4 h at 37 °C. The proteins were purified on His-NTA column as described in 66, except that before the elution (500 mM imidazole) a two-step imidazole washing was done, by 75 mM and 100 mM imidazole. The proteins were purified either in the presence or in the absence of reducing agent. As a reducing agent 10 mM 2–mercaptoethanol (β–ME) was used. The protein concentration was measured by UV absorbance. Extinction coefficients were calculated based on the amino acid sequence using the ProtParam tool (http://web.expasy.org/protparam/). Extinction coefficients for the proteins were: λ280 = 15,930 M−1 cm−1 and λ280 = 16,055 M−1 cm−1 for the hDUTF158W (reduced/non-reduced), λ280 = 14,440 M−1∙cm−1 and λ280 = 14,690 M−1∙cm−1 for the hDUTF158W,Y54C (reduced/non-reduced). Protein concentration is given in monomer/subunit concentration in every case. All measurements were carried out in a buffer comprising 20 mM HEPES pH 7.5, 100 mM NaCl, 2 mM MgCl2 and 10 mM β-ME (“assay buffer”) if not stated otherwise.
Enzyme activity assay
Michaelis–Menten enzyme kinetics
Proton release during the transformation of dUTP into dUMP and PPi was followed continuously at 559 nm at 20 °C using a JASCO-V550 spectrophotometer67,68. Reaction mixtures contained 10 nM enzyme and varying concentrations of dUTP. The reaction was started with the addition of dUTP. Initial velocity was determined from the slope of the first 10% of the progress curve. Initial velocities were plotted against substrate concentration and the results were fitted with the Michaelis–Menten equation.
Investigation of enzyme stability by activity measurement
This measurement was done with the same assay, except that fixed concentration of dUTP (30 µM) was used, and the enzyme was pre-incubated at 37 °C for different time intervals (0–90 min).
Transient kinetics experiments
Fluorescence stopped-flow measurements were carried out using an SX20 (Applied Photophysics, UK) stopped-flow instrument, following tryptophan fluorescence at 20 °C, as described previously44,60,65,66. Equal volumes (50 μl) of 40 μM dUTPase enzyme and 40 μM dUTP solutions (both diluted in assay buffer) were mixed and typically 3–5 traces were collected and averaged. A triple exponential function (Eq. 1) was fitted to the averaged traces to determine the catalytic constants, and the observed binding rates at the given substrate concentration based on Tóth et al. 44.
where F is the observed fluorescence, x is the variable (time), A1,2,3 are the amplitudes, k1,2,3obs are the rate constants of the observable fluorescence phases, while y0 is the y offset.
Differential Scanning Fluorimetry (DSF)
DSF assays were carried out on a Bio-Rad CFX96 qPCR in FrameStar® 96 Well Skirted PCR Plates with black wire and white wells sealed with Eppendorf adhesive PCR films. Reactions were performed in a total volume of 25 μl containing 500 × diluted Sypro® Orange dye. Samples were heated from 25 to 90 °C. The speed of heating was 2 °C/minute. The protein concentration was 0.8 mg/ml in the measurements. The melting point was determined by taking the negative derivate of the curves. β-ME was added as indicated on Fig. 4C.
Thiol quantitation assay
Reactive thiols were measured by MAK151 Sigma Fluorometric Thiol Quantitation Kit according to manufacturer instructions. Briefly 10 μl of the protein samples was added to 10 μl of thiol reagent in a well of a low volume black bottom plate (CLS4514 Sigma). Samples were mixed and then kept in dark for 30 min. Fluorescence intensity was measured at λex = 490 nm/ λem = 535 nm by Spectramax M5 Plate reader (Molecular Devices). Fluorescence values were normalized by subtraction of the fluorescence measured for the buffer. Each sample were measured in triplicates.
Electrospray ionization mass spectrometry
The oligomerization state of the human dUTPase was studied by a Waters QTOF Premier mass spectrometer (Waters, Milford, MA, USA) equipped with electrospray ionization source (Waters, Milford, MA, USA) operated in positive ion mode. Purified hDUT samples of 20 µM were subjected to buffer exchange to 200 mM NH4HCO3 buffer performed with Vivaspin® 500 Polyethersulfone centrifugal concentrators of 10 kDa weight cutoff. Mass spectra were measured under native conditions: namely, the ions were generated from aqueous 5 mM NH4HCO3 buffer solution (pH = 7.5) containing hDUT at ca. 0.5 µM monomer concentration, which conditions favor the transfer of the protein complexes from solution into the gas phase. The capillary voltage was 2600–2800 V, the sampling cone voltage was 125 V and the temperature of the source was set at 80 °C, collision cell pressure was 3.43 × 10–3 mbar and ion guide gas flow was 35 ml/min. Mass spectra were recorded applying the software MassLynx 4.1 (Waters, Milford, MA, USA) in the 1000–8000 m/z mass range43.
Change history
29 October 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41598-021-01241-2
References
Hurd, T. R., Prime, T. A., Harbour, M. E., Lilley, K. S. & Murphy, M. P. Detection of reactive oxygen species-sensitive thiol proteins by redox difference gel electrophoresis: Implications for mitochondrial redox signaling. J. Biol. Chem. 282, 22040–22051 (2007).
Hanson, G. T. et al. Investigating mitochondrial redox potential with redox-sensitive green fluorescent protein indicators. J. Biol. Chem. 279, 13044–13053 (2004).
Dooley, C. T. et al. Imaging dynamic redox changes in mammalian cells with green fluorescent protein indicators. J. Biol. Chem. 279, 22284–22293 (2004).
Go, Y.-M. & Jones, D. P. Redox compartmentalization in eukaryotic cells. Biochim. Biophys. Acta 1780, 1273–1290 (2008).
Yasuhira, S. & Yasui, A. Alternative excision repair pathway of UV-damaged DNA in Schizosaccharomyces pombe operates both in nucleus and in mitochondria. J. Biol. Chem. 275, 11824–11828 (2000).
Bacman, S. R., Williams, S. L. & Moraes, C. T. Intra- and inter-molecular recombination of mitochondrial DNA after in vivo induction of multiple double-strand breaks. Nucleic Acids Res. 37, 4218–4226 (2009).
de Souza-Pinto, N. C. et al. Novel DNA mismatch-repair activity involving YB-1 in human mitochondria. DNA Repair (Amst). 8, 704–719 (2009).
Boesch, P. et al. Plant mitochondria possess a short-patch base excision DNA repair pathway. Nucleic Acids Res. 37, 5690–5700 (2009).
Szczesny, B., Tann, A. W., Longley, M. J., Copeland, W. C. & Mitra, S. Long patch base excision repair in mammalian mitochondrial genomes. J. Biol. Chem. 283, 26349–26356 (2008).
Boesch, P. et al. DNA repair in organelles: pathways, organization, regulation, relevance in disease and aging. Biochim. Biophys. Acta 1813, 186–200 (2011).
Vértessy, B. G. & Tóth, J. Keeping uracil out of DNA: physiological role, structure and catalytic mechanism of dUTPases. Acc. Chem. Res. 42, 97–106 (2009).
Hirmondo, R., Lopata, A., Suranyi, E. V., Vertessy, B. G. & Toth, J. Differential control of dNTP biosynthesis and genome integrity maintenance by dUTPases. Sci. Rep. 7, 6043 (2017).
Ladner, R. D., McNulty, D. E., Carr, S. A., Roberts, G. D. & Caradonna, S. J. Characterization of distinct nuclear and mitochondrial forms of human deoxyuridine triphosphate nucleotidohydrolase. J. Biol. Chem. 271, 7745–7751 (1996).
Rácz, G. A. et al. Evaluation of critical design parameters for RT-qPCR-based analysis of multiple dUTPase isoform genes in mice. FEBS Open Bio 9, 1153–1170 (2019).
Róna, G. et al. Dynamics of re-constitution of the human nuclear proteome after cell division is regulated by NLS-adjacent phosphorylation. Cell Cycle 13, 3551–3564 (2014).
Varga, B. et al. Active site closure facilitates juxtaposition of reactant atoms for initiation of catalysis by human dUTPase. FEBS Lett. 581, 4783–4788 (2007).
Nagy, G. N. & Leveles, I. Preventive DNA repair by sanitizing the cellular (deoxy)nucleoside triphosphate pool. Sci. Rep. 281, 4207–4223 (2014).
Takács, E., Grolmusz, V. K. & Vértessy, B. G. A tradeoff between protein stability and conformational mobility in homotrimeric dUTPases. FEBS Lett. 566, 48–54 (2004).
Mol, C. D., Harris, J. M., McIntosh, E. M. & Tainer, J. A. Human dUTP pyrophosphatase: uracil recognition by a beta hairpin and active sites formed by three separate subunits. Structure 4, 1077–1092 (1996).
Chu, R., Lin, Y., Rao, M. S. & Reddy, J. K. Cloning and identification of rat deoxyuridine triphosphatase as an inhibitor of peroxisome proliferator-activated receptor α. J. Biol. Chem. 271, 27670–27676 (1996).
Benedek, A. et al. Potential steps in the evolution of a fused trimeric all-β dUTPase involve a catalytically competent fused dimeric intermediate. FEBS J. 283, 3268–3286 (2016).
Badalucco, L., Poudel, I., Yamanishi, M., Natarajan, C. & Moriyama, H. Crystallization of Chlorella deoxyuridine triphosphatase. Acta Crystallogr. Sect. F. Struct. Biol. Cryst. Commun. 67, 1599–1602 (2011).
Rafael Ciges-Tomas, J. et al. The structure of a polygamous repressor reveals how phage-inducible chromosomal islands spread in nature. Nat. Commun. 10, 3676 (2019).
Whittingham, J. L. et al. dUTPase as a platform for antimalarial drug design: structural basis for the selectivity of a class of nucleoside inhibitors. Structure 13, 329–338 (2005).
Samal, A., Schormann, N., Cook, W. J., DeLucas, L. J. & Chattopadhyay, D. Structures of vaccinia virus dUTPase and its nucleotide complexes. Acta Crystallogr. D. Biol. Crystallogr. 63, 571–580 (2007).
Tchigvintsev, A. et al. Structure and activity of the Saccharomyces cerevisiae dUTP pyrophosphatase DUT1, an essential housekeeping enzyme. Biochem. J. 437, 243–253 (2011).
Li, C. et al. Crystal structure of African Swine fever virus dUTPase reveals a potential drug target. MBio 10, 19 (2019).
Prasad, G. S., Enrico, A., Elder, J. H. & Stout, C. D. Structures of feline immunodeficiency virus dUTP pyrophosphatase and its nucleotide complexes in three crystal forms research papers. Acta Crystallogr. D Biol. Crystallogr. 56, 1100–1109 (2000).
Maiques, E. et al. Another look at the mechanism involving trimeric dUTPases in Staphylococcus aureus pathogenicity island induction involves novel players in the party. Nucleic Acids Res. 44, 5457–5469 (2016).
Németh-Pongrácz, V. et al. Flexible segments modulate co-folding of dUTPase and nucleocapsid proteins. Nucleic Acids Res. 35, 495–505 (2007).
Cedergren-Zeppezauer, E. S., Larsson, G., Nyman, P. O., Dauter, Z. & Wilson, K. S. Crystal structure of a dUTPase. Nature 355, 740–743 (1992).
Zang, K., Li, F. & Ma, Q. The dUTPase of white spot syndrome virus assembles its active sites in a noncanonical manner. J. Biol. Chem. 293, 1088–1099 (2018).
Chan, S. et al. Crystal structure of the Mycobacterium tuberculosis dUTPase: insights into the catalytic mechanism. J. Mol. Biol. 341, 503–517 (2004).
García-Nafría, J. et al. The structure of the genomic Bacillus subtilis dUTPase: novel features in the Phe-lid. Acta Crystallogr. Sect. D Biol. Crystallogr. 66, 953–961 (2010).
Leveles, I. et al. Structure and enzymatic mechanism of a moonlighting dUTPase. Acta Crystallogr. Sect. D Biol. Crystallogr. 69, 2298–2308 (2013).
Freeman, L. et al. The flexible motif V of Epstein-Barr virus deoxyuridine 5’-triphosphate pyrophosphatase is essential for catalysis. J. Biol. Chem. 284, 25280–25289 (2009).
Dauter, Z. et al. Crystal structure of dUTPase from equine infectious anaemia virus; active site metal binding in a substrate analogue complex. J. Mol. Biol. 285, 655–673 (1999).
Kan, L. et al. Cloning and expression of the mouse deoxyuridine triphosphate nucleotidohydrolase gene: differs from the rat enzyme in that it lacks nuclear receptor interacting LXXLL motif. Gene Expr. 8, 231–246 (1999).
Kovári, J. et al. Altered active site flexibility and a structural metal-binding site in eukaryotic dUTPase: kinetic characterization, folding, and crystallographic studies of the homotrimeric Drosophila enzyme. J. Biol. Chem. 279, 17932–17944 (2004).
Pecsi, I. et al. The dUTPase enzyme is essential in Mycobacterium smegmatis. PLoS ONE 7, e37461 (2012).
Rotoli, S. M., Jones, J. L. & Caradonna, S. J. Cysteine residues contribute to the dimerization and enzymatic activity of human nuclear dUTP nucleotidohydrolase (nDut). Protein Sci. 27, 1797–1809 (2018).
Takács, E., Barabás, O., Petoukhov, M. V., Svergun, D. I. & Vértessy, B. G. Molecular shape and prominent role of beta-strand swapping in organization of dUTPase oligomers. FEBS Lett. 583, 865–871 (2009).
Nyíri, K. et al. Structural model of human dUTPase in complex with a novel proteinaceous inhibitor. Sci. Rep. 8, 4326 (2018).
Tóth, J., Varga, B., Kovács, M., Málnási-Csizmadia, A. & Vértessy, B. G. Kinetic mechanism of human dUTPase, an essential nucleotide pyrophosphatase enzyme. J. Biol. Chem. 282, 33572–33582 (2007).
Gadsden, M. H., McIntosh, E. M., Game, J. C., Wilson, P. J. & Haynes, R. H. dUTP pyrophosphatase is an essential enzyme in Saccharomyces cerevisiae. EMBO J. 12, 4425–4431 (1993).
Pálinkás, H. L. et al. Crispr/cas9-mediated knock-out of dutpase in mice leads to early embryonic lethality. Biomolecules 9, 18 (2019).
Muha, V. et al. Uracil-containing DNA in drosophila: stability, stage-specific accumulation, and developmental involvement. PLoS Genet. 8, e1002738 (2012).
El-hajj, H. H., Zhang, H. U. I. & Weiss, B. Lethality of a dut ( Deoxyuridine Triphosphatase ) Mutation in E. coli. 170, 1069–1075 (1988).
Dubois, E. et al. Homologous recombination is stimulated by a decrease in dUTPase in Arabidopsis. PLoS One 6, e18658 (2011).
Castillo-Acosta, V. M. et al. Pyrimidine requirements in deoxyuridine triphosphate nucleotidohydrolase deficient Trypanosoma brucei mutants. Mol. Biochem. Parasitol. 187, 9–13 (2013).
Kerepesi, C. et al. Life without dUTPase. Front. Microbiol. 7, 1–10 (2016).
Merényi, G. et al. Cellular response to efficient dUTPase RNAi silencing in stable HeLa cell lines perturbs expression levels of genes involved in thymidylate metabolism. Nucleosides. Nucleotides Nucleic Acids 30, 369–390 (2011).
Dos Santos, R. S. et al. dUTPase (DUT) Is mutated in a novel monogenic syndrome with diabetes and bone marrow failure. Diabetes 66, 1086–1096 (2017).
Wilson, P. M., LaBonte, M. J., Lenz, H.-J., Mack, P. C. & Ladner, R. D. Inhibition of dUTPase induces synthetic lethality with thymidylate synthase-targeted therapies in non-small cell lung cancer. Mol. Cancer Ther. 11, 616–628 (2012).
Koehler, S. E. & Ladner, R. D. Small interfering RNA-mediated suppression of dUTPase sensitizes cancer cell lines to thymidylate synthase inhibition. Mol. Pharmacol. 66, 620–626 (2004).
Kiyonari, S. et al. The 1,2-diaminocyclohexane carrier ligand in oxaliplatin induces p53-dependent transcriptional repression of factors involved in thymidylate biosynthesis. Mol. Cancer Ther. 1535-7163.MCT-14-0748- (2015). https://doi.org/10.1158/1535-7163.MCT-14-0748
Wilson, P. M., Danenberg, P. V., Johnston, P. G., Lenz, H.-J. & Ladner, R. D. Standing the test of time: targeting thymidylate biosynthesis in cancer therapy. Nat. Rev. Clin. Oncol. 11, 282–298 (2014).
Ladner, R. D. et al. dUTP nucleotidohydrolase isoform expression in normal and neoplastic tissues: association with survival and response to 5-fluorouracil in colorectal cancer. Cancer Res. 60, 3493–3503 (2000).
Wilson, P. M. et al. Novel opportunities for thymidylate metabolism as a therapeutic target. Mol. Cancer Ther. 7, 3029–3037 (2008).
Pécsi, I., Leveles, I., Harmat, V., Vértessy, B. G. & Tóth, J. Aromatic stacking between nucleobase and enzyme promotes phosphate ester hydrolysis in dUTPase. Nucleic Acids Res. 38, 7179–7186 (2010).
Piovesan, D., Minervini, G. & Tosatto, S. C. E. The RING 2.0 web server for high quality residue interaction networks. Nucleic Acids Res. 44, W367–74 (2016).
Donnelly, D. P. et al. Best practices and benchmarks for intact protein analysis for top-down mass spectrometry. Nat. Methods 16, 587–594 (2019).
Hochhauser, S. J. & Weiss, B. Escherichia coli mutants deficient in deoxyuridine triphosphatase. J. Bacteriol. 134, 157–166 (1978).
Marton, L. et al. Molecular mechanism for the thermo-sensitive phenotype of CHO-MT58 cell line harbouring a mutant CTP: Phosphocholine Cytidylyltransferase. PLoS One 10, e0129632 (2015).
Pécsi, I. et al. Nucleotide pyrophosphatase employs a P-loop-like motif to enhance catalytic power and NDP/NTP discrimination. Proc. Natl. Acad. Sci. U. S. A. 108, 14437–14442 (2011).
Szabó, J. E., Takács, E., Merényi, G., Vértessy, B. G. & Tóth, J. Trading in cooperativity for specificity to maintain uracil-free DNA. Sci. Rep. 6, 24219 (2016).
Vertessy, B. G., Zalud, P., Nyman, P. O. & Zeppezauer, M. Identification of tyrosine as a functional residue in the active site of E. coli dUTPase. Biochim. Biophys. Acta 1205, 146–150 (1994).
Varga, B. et al. Active site of mycobacterial dUTPase: structural characteristics and a built-in sensor. Biochem. Biophys. Res. Commun. 373, 8–13 (2008).
Funding
This research was funded by National Research, Development and Innovation Office of Hungary, Grant numbers PD124330 (JES), PD134324 (KN), K135231 (BGV) VEKOP-2.3.2-16-2017-00013, NKP-2018-1.2.1-NKP-2018-00005, and the BME-Biotechnology FIKP grant of EMMI (BME FIKP-BIO) . J.E.S. is the recipient of the János Bolyai Research Scholarship of the Hungarian Academy of Sciences; J.E.S. is the recipient of ÚNKP-18-4-BME-391 and ÚNKP-19-4-BME-420 New National Excellence Program of the Ministry of Human Capacities. The work of KN was supported through the New National Excellence Program of the Ministry of Human Capacities (ÚNKP-20-4-II-BME-311).
Author information
Authors and Affiliations
Contributions
All authors have read and agree to the published version of the manuscript. Conceptualization, J.E.S., K.N. and B.G.V.; methodology, J.E.S, K.N. and O.O.; investigation, J.E.S., D.A., K.N., M. J. and O.O.; resources, J.E.S., K.N. and B.G.V.; data curation, J.E.S., D.A., K.N. and O.O.; writing—original draft preparation, J.E.S., K.N. and B.G.V.; writing—review and editing, J.E.S., K.N. and B.G.V.; visualization, J.E.S., K.N. and B.G.V.; supervision, J.E.S., B.G.V.; funding acquisition, J.E.S., K.N. and B.G.V.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: The original version of this Article contained an error in the spelling of the author Olivér Ozohanics which was incorrectly given as Olivér Ozahonics.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Szabó, J.E., Nyíri, K., Andrási, D. et al. Redox status of cysteines does not alter functional properties of human dUTPase but the Y54C mutation involved in monogenic diabetes decreases protein stability. Sci Rep 11, 19197 (2021). https://doi.org/10.1038/s41598-021-98790-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-98790-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.