Abstract
Macropod progressive periodontal disease (MPPD) is a necrotizing, polymicrobial, inflammatory disease commonly diagnosed in captive macropods. MPPD is characterized by gingivitis associated with dental plaque formation, which progresses to periodontitis and then to osteomyelitis of the mandible or maxilla. However, the underlying microbial causes of this disease remain poorly understood. In this study, we collected 27 oral plaque samples and associated clinical records from 22 captive Macropodidae and Potoroidae individuals that were undergoing clinical examination at Adelaide and Monarto Zoos in South Australia (15 healthy, 7 gingivitis and 5 periodontitis-osteomyelitis samples). The V3-V4 region of the 16S ribosomal RNA gene was sequenced using an Illumina Miseq to explore links between MPPD and oral bacteria in these animals. Compositional differences were detected between the microbiota of periodontitis-osteomyelitis cases compared to healthy samples (p-value with Bonferroni correction < 0.01), as well as gingivitis cases compared to healthy samples (p-value with Bonferroni correction < 0.05) using Permutational Multivariate Analysis of Variance (PERMANOVA). An overabundance of Porphyromonas, Fusobacterium, and Bacteroides taxa was also identified in animals with MPPD compared to healthy individuals using linear discriminant analysis effect size (LEfSe; p = < 0.05). An increased abundance of Desulfomicrobium also was detected in MPPD samples (LEfSe; p < 0.05), which could potentially reflect differences in disease progression. This is the first microbiota analysis of MPPD in captive macropods, and these results support a polymicrobial pathogenesis of MPPD, suggesting that the microbial interactions underpinning MPPD may be more complex than previously documented.
Similar content being viewed by others
Introduction
Macropods, which include kangaroos and wallabies, are herbivorous marsupials in the superfamily Macropodoidea within the order Diprotodontia. The regular macropod dentition varies across species, and the dental formula is I 3/1, C 0–1/0, PM 1–2/1–2, M 4/4, reflecting adaptation to a range of foraging strategies. Incisors are used for prehension, and pre-molars and molars are used for mastication. Molar progression occurs normally in grazing and some intermediate grazer-browser species, where premolar and molar teeth progressively wear down and are shed at the rostral end of the quadrant when they cease to be functional1,2.
Macropod progressive periodontal disease (MPPD), commonly termed ‘lumpy jaw’, is a necrotising polymicrobial dental disease characterised by the proliferation of anaerobic bacteria. MPPD is one of the most common diseases identified in macropods, especially in captive environments. Appropriate treatment is often successful with early intervention, though recurrence is common. Euthanasia is often required in advanced cases3,4,5,6.
The pathogenesis of MPPD is not fully understood. Commonly reported risk factors for MPPD include stressors (e.g. cold, wet weather and overcrowding), molar progression, food impaction and plaque accumulation, faecal contamination of feed areas, age, and inappropriate diet7,8,9. Coarse feed may cause gingival injury that could allow bacterial invasion. Soft foods with insufficient abrasiveness may also lead to a lack of natural ‘toughening’ of the mucosa and affect toothwear1,2,3,7.
Periodontal disease (PD) is a common, inflammatory, oral disease recognised in humans and animals. PD is generally initiated by the accumulation of dental plaque—a diverse biofilm of commensal microorganisms—that adhere to teeth. In humans, primary colonizers, such as Streptococci spp. and Corynebacterium spp. adhere to tooth enamal, followed by secondary colonisers, such as Fusobacterium spp., which provide a foundation that can allow late-colonising bacteria, including anaerobic gram-negative bacteria, to attach10,11. Gingivitis, a reversible form of PD, presents as inflammation of gingivae resulting from the host’s innate and adaptive immune response to bacterial toxins, including enzymes, structural components and leukotoxins. Periodontitis, an irreversible form of PD, progresses to loss of integrity of the gingival epithelium and inflammation of the periodontal ligament, absorption of alveolar bone, tooth mobility and eventual tooth loss4,5,6.
Unlike PD in humans, MPPD commonly progresses to necrotising osteomyelitis of the mandible or maxilla, with formation of sequestra and proliferation of subperiosteal bone subsequently leading to bone deformity in the jaw. Although the pathogenesis of PD in macropods is considered to be similar to that for humans8, the progression to osteomyelitis, suppurative inflammation and necrosis of adjacent soft tissues observed in macropods is rare in humans3.
Various hypotheses have been offered to describe the pathogenesis of PD, mostly in humans. The ‘non-specific plaque’ hypothesis states that PD progression correlates with the total quantity of bacterial plaque present, while the ‘specific plaque’ hypothesis proposes that disease correlates with the presence of specific bacterial species12. Polymicrobial synergy and dysbiosis working together has also been proposed as a model for PD development13. The models assumes that bacterial interactions can be antagonistic or synergistic and that a change in the local environment initiates disease that leads to an imbalance or dysbiosis within the microbial community.
A shift in the microbial community that favours survival and toxin production by some bacteria is likely to result in polymicrobial disease. A further model, the ‘keystone-pathogen hypothesis,’13 contends that disease depends on the presence of a single species or group of bacteria that allows environmental change and dysbiosis to occur. This hypothesis suggests that the immune response is subverted by specific keystone-bacteria to allow progression of disease14. Poryphomonas gingivialis is considered to be the keystone pathogen in human PD12,13,15. In animals, a closely related pathogen Porphyromonas gulae, and in marsupials specifically, the recently-described Porphyromonas loveana, may act as similar keystone pathogens16.
In macropods, culture-dependent methods have been used to isolate bacterial species from the oral cavity, including those involved in MPPD. It has been reported that the major causative agents of MPPD are bacteria within the Norcardia, Actinomyces, Bacteroides, and Fusobacterium genera. Fusobacterium necrophorum has been the most commonly isolated bacteria from MPPD lesions8,17,18. However, there have been cases of MPPD presumed to be attributable to Bacteroides, with no Fusobacterium species isolated, as well as cases where neither species were identified19,20. In the healthy oral cavity, aerobic Gram-positive organisms are predominant, although Fusobacterium species, routinely identified in human oral microbiota and PD, have typically not been isolated in macropods8,21,22,23. This striking difference between healthy and diseased groups in the study of Samuel, questioned whether F. necrophorum escaped detection, and is part of the endogenous micriobiota or introduced from an external source18.
Culture-independent studies using PCR-Denatured Gradient Gel Electrophoresis (PCR-DGGE) have detected F. necrophorum at different stages of MPPD, along with other Gram-negative anaerobic bacteria belonging to Bacteroides and Porphyromonas genera3. This finding was supported by quantitative PCR for a leucotoxin gene of F. necrophorum, but not all animals with disease were PCR-positive. F. necrophorum and its leukotoxin gene were also detected at low levels in some healthy animals24. Bacterial families found only in the healthy group were Pasteurellaceae and Moraxellaceae. Bacterial diversity was greater in healthy mouths than in periodontitis-osteomyelitis cases, but was highest in gingivitis cases. Analysis of bacterial community structure in diseased mouths showed that a small proportion of organisms were responsible for a large amount of functional interaction, supporting the ecological plaque hypothesis3. P. loveana and P. gulae were the predominant porphyromonads isolated from the macropod oral cavity16.
Next generation sequencing (NGS) methods can be applied to profile bacterial communities by sequencing the hypervariable regions of amplified 16S rRNA-encoding genes. This region of the rRNA gene is able to identify prokaryotes at the individual species level in some cases, and it has been used to study oral, gut and other microbiotas of animals, including macropods25,26. Sequencing methods have also defined the oral microbiota of Tasmanian devils (Sarcophilus harrisii), identifying over 1000 unique microbial sequences, or operational taxonomic units (OTUs), which would have been impossible to determine using culture based methods27. The oral microbiota of macropods, in either health or disease, has not been studied using High Throughput Seuqencing (HTS) methods. This study uses HTS to explore the oral microbiota of healthy captive macropods compared to those diagnosed with MPPD. We hypothesise that specific changes in microbiota profile contribute to the induction and progression of MPPD. The aim of this study was to identify microbiota changes in two stages of disease (gingivitis and periodontitis-osteomyelitis) compared to the healthy condition. This will add to our understanding of the microbiology of MPPD in macropods and may lead to improved preventative measures or treatment of the disease.
Results
Clinical descriptions
After examination, 27 samples from 22 animals were classified as healthy (n = 15), gingivitis (n = 7) or periodontitis-osteomyelitis cases (n = 5). Supplementary 1 provides further clinical details on the two most severe cases of MPPD sampled.
Microbiota profiling
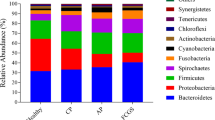
The relative abundances of bacteria at the phylum, genus, and species taxonomic levels were explored in 27 animals belonging to healthy, gingivitis and periodontitis-osteomyelitis categories. In total, 1178 OTUs were detected, assigned to 28 phyla, 186 families, 438 genera and 181 species (Fig. 1).
Proteobacteria and Actinobacteria were the most abundant phyla in healthy samples, forming 80.8% of the entire microbiota (Supplementary 2). We also explored microbiota identified at the class and genus levels in healthy, gingivitis, and periodontitis-osteomyelitis samples (Fig. 2 and Table 1). At the genus level, Moraxella, Fusobacterium and Lautropia were the most abundant genera across all samples (Supplementary 3).
MPPD is associated with compositional shifts in oral microbiota, but not diversity.
We examined if shifts in alpha diversity were apparent between health and diseased animals using the Shannon’s Diversity Index (Fig. 3). There were no significant shift in alpha diversity between healthy, gingivitis, and periodontitis-osteomyelitis sample groups (Fig. 3).
In contrast, the composition (beta diversity) of oral microbiota in periodontitis-osteomyelitis samples were distinguishable from the healthy and gingivitis samples using Principal Coordinate Analysis (PCoA) plot of Bray Curtis Values (Fig. 4). In a PCoA, periodontitis-osteomyelitis samples grouped to the exclusion of healthy samples on the first PCoA component of a 3D PCoA plot (Fig. 4). Supplementary 7 presents the formula PCoA components. The first component explains 31% of variance in the microbiome data (Fig. 4). The first component separated periodontitis-osteomyelitis samples from healthy and gingivitis samples, as periodontitis-osteomyelitis samples had the lowest negative coefficient values for this component (Supplementary 7). Further, the microbial composition of healthy animals was significantly different from those suffering from both periodontitis-osteomyelitis (PERMANOVA with Bonferoni correction; 0.0002) and gingivitis (p = 0.0323) (Table 2). Further, the composition of oral micorbiota in animals with periodontitis-osteomyelitis was not significantly different from those with gingivitis (p = 0.0833) (Table 2), suggesting that there may be compositional similarities between these two stages of disease. Together, these results suggest that a distinct oral microbial communities are associated with PD and gingivitis in macropods compared to healthy animals.
Porphyromonas, Fusobacterium and Bacteroides dominate microbiota in both gingivitis and MPPD
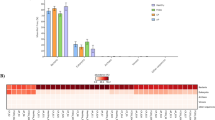
A LEfSe test was utilized to identify specific taxa associated with each disease state. We observed significant differences in taxa abundance in the course of disease. The abudance of 27 genera were significantly different in healthy animals versus those suffering from gingivitis, while 66 genera were differnet between healthy and periodontitis-osteomyelitis animals (Fig. 5; Table 2; Supplementary 4 and Supplementary 5).
Biomarker discovery in macropod progressive periodontal disease using LEfSe test. (A) The main genera that differentiate gingivitis from healthy samples. (B) The main genera that differentiate periodontitis-osteomyelitis from healthy samples. LEfSe test with alpha value = 0.05 and threshold of absolute logarithmic linear discriminant analysis (LDA) score > 2. Higher absolute values of the LDA score represent higher enrichment and discriminative performance. Compared to healthy, Porphyromonas, Fusobacterium are overabundant in gingivitis and periodontitis-osteomyelitis samples. In contrast, Neisseria is underabundant in gingivitis and periodontitis-osteomyelitis samples.
In the comparison of gingivitis and healthy samples, Porphyromonas (logarithmic LDA score = 4.743 and p-value = 0.001), Fusobacterium (logarithmic LDA score = 4.648 and p-value = 0.031), and Bacteroides (logarithmic LDA score = 4.666 and p-value = 0.044) were the dominant taxa that were overrepresented in gingivitis samples. In contrast, Neisseria, a genera within the Proteobacteria (the most abundant phyla in healthy samples) (logarithmic LDA score = 4.528 and p-value = 0.037) and Akkermansia (logarithmic LDA score = 3.297 and p-value = 0.077) were overrepresented in healthy samples (Fig. 5, Table 1, Supplementary 4).
In the comparison of periodontitis-osteomyelitis and healthy samples, Porphyromonas (logarithmic LDA score = 4.743 and p-value = 0.001), Fusobacterium (logarithmic LDA score = 4.648 and p-value = 0.031), and Bacteroides (logarithmic LDA score = 4.666 and p-value = 0.044) were the dominante species that were overrepresented in periodontitis-osteomyelitis, similar to the gingivitis. In contrast, Neisseria was again overabundant in healthy animals compared to those suffering from periodontitis-osteomyelitis (Fig. 5, Table 1, Supplementary 5).
We also investigated differences in taxa abunance in animals suffering from gingivitis or periodontitis-osteomyelitis. Eighteen genera were significantly different in abundance in both the gingivitis versus healthy and periodontitis-osteomyelitis versus healthy sample comparisons (Table 1, Supplementary 6). Of particular note, the abundance of Desulfomicrobium increased in both gingivitis and periodontitis-osteomyelitis in comparison to healthy samples (Table 1), with the marked increase in periodontitis-osteomyelitis samples potentially having additive effects on disease progress (logarithmic LDA score = 3.564 and p-value = 0.002) (Supplementary 5).
We confirmed these resulst by additional performing Pearson correlations on species abundance and disease. Pearson correlations revealed highly significant, positive correlations (> 72%, p < 0.01) between the relative abundance of Porphyromonas, Fusobacterium, Bacteroides, and Desulfomicrobium in oral macropod samples (Table 3). In contrast, there was negative correlation between the abundance of Neisseria genus with Porphyromonas, Fusobacterium, Bacteroides, and Desulfomicrobium genera (Table 3). Signature of Porphyromonas, Fusobacterium, and Bacteroides in MPPD was robust and consistent in different sexes, species, and zoos (Supplementary 8).
Discussion
Periodontal diseases are commonly reported clinical disorders in animals. Here, for the first time, the oral microbiota of healthy and MPPD-affected captive macropods were explored using 16S rRNA gene seuqencing. This study characterised the healthy macropod oral microbiota and identified differences in bacterial composition and taxa abundances between healthly samples and the different stages of MPPD, thus providing novel information about this polymicrobial disease and its prospective pathogenesis.
There are similarities and differences between PD in human and MPPD in macropods. In humans, PD is initiated by dental plaque and exhibits progression from reversible gingivitis to irreversible periodontitis. However, unlike PD in humans, MPPD commonly progresses to necrotising osteomyelitis of the mandible or maxilla, with the formation of sequestra and proliferation of subperiosteal bone subsequently leading to bone deformity in the jaw (the characteristic ‘lumpy jaw’). The progression to osteomyelitis, suppurative inflammation and necrosis of adjacent soft tissues observed in macropods is rare in humans3,28. In humans, early colonisers, such as Streptococcus spp. and Fusobacterium spp. provide a foundation that can allow late colonising bacteria to attach, which includes many anaerobic Gram-negative bacteria4,5,6,29. In the present study, we found that Porphyromonas, Fusobacterium, and Bacteroides are the most abundant genera in gingivitis samples. These genera also dominated the microbiome in periodontitis-osteomyelitis cases (the advanced stages of disease). Also, we observed a significant and positive correlation between the relative abundances of Porphyromonas and Fusobacterium genera and disease, similar to human PD. In contrast to humans, Streptococcus did not appear to be a major component of plaque microbiota in macropods.
It has been well-documented that Fusobacterium nucleatum can enhance the attachment and invasion of Porphyromonas gingivalis or Aggregatibacter actinomycetemcomitans to human gingival epithelial cells29. In another study, Porphyromonas gingivalis entry into gingival epithelial cells was modulated by Fusobacterium nucleatum30. Additionally, it has been proposed that Fusobacterium spp. can inhibit the initial host innate immune response29. Scanning electron microscopy has shown that P. gingivalis and F. nucleatum can form consortia and penetrate Ca9-22 cells within 30–60 min after infection (early colonisation)30. Altogether, we suggest that Fusobacterium may be involved in early colonisation in MPPD, enhancing adhesion and invasion of species within the Porphyromonas genus that are likely to be either P. gulae or loveana. This interplay has been observed in murine alveolar bone loss and arthritis onset31.
Microbiota profiling using 16S rRNA gene pyrosequencing identified the Gram-negative bacterial genera Fusobacterium, Bacteroides, and Porphyromonas as the dominant taxa in MPPD, as noted in previous culture and DGGE-based studies3,8,17,18,20. These findings support the hypothesis that Porphyromonas and Bacteroides may have a bigger role in disease pathogenesis than has historically been proposed. The importance of pathogens other than F. necrophorum in MPPD was first noted in 1977, and recent DGGE-based molecular studies, confirmed that Porphyromonas and Bacteroides are important genera in MPPD pathogenesis3,20,24. However, caution should be applied, as bacterial abundance may not necessarily correlate with pathogenicity.
Abundance of Desulfomicrobium also increased in MPPD samples (LEfSe test, p < 0.05), which could potentially have additive effects on disease progress. Desulfomicrobium orale has also been isolated from subgingival plaque of human patients with PD32, identified as a human oral pathogen33. This is the first time that another significant genera associated with PD in humans has been reported in MPPD, albeit at low abundance when compared to Porphyromonas, Bacteroides and Fusobacterium.
The dominance of the Proteobacteria phylum found in the oral microbiota of healthy macropods was similar to another study of the salivary microbiota in Tammar wallabies25. This contrasts with previous reports on the oral microbiota of other marsupials, such as Tasmanian Devils, which have a similar proportion of Proteobacteria, Bacteroidetes, Fusobacteria, and Firmicutes phyla (at around 20% of each) and koalas, where Bacteroidetes and Firmicutes were in the top three phyla27,34. In macropods, the highly abundant Proteobacteria phylum is composed of many species associated with gingival health in comparison to either gingivitis or periodontitis-osteomyelitis3. In particular, Pasteurellaceae and Moraxellaceae were the two major families found to be abundant in healthly samples3. An increased abundance of Gram-positive non-sporulating rods such as Corynebacterium sp. and Actinomyces sp. in healthy compared to diseased samples has also been reported in culture-based studies18,23. In line with those studies, species in the genera Corynebactrium, Actinomyces, Streptococcus, Lautropia, Leptotrichia and Capnocytophaga are associated with the healthy oral cavity in human microbiota studies35. It has also been noted that some disease-associated genera have overlapping species that are also present in healthy samples35.
Despite being the first study to profile the microbiota of MPPD using 16S rRNA gene pyrosequencing, this study had some limitations. In microbiota studies, sequencing depth normally includes the family, genus and sometimes species level, but there can be many OTUs that cannot be identified at the lowest level. The current study was based on a single zoological collection of animals at two sites, and the availability of MPPD cases was a limiting factor on sample size. Subsequently, there was an unequal number of healthy, gingivitis and periodontitis-osteomyelitis cases available for study, with only a limited number of gingivitis and periodontitis-osteomyelitis samples. Additionally, the low sample size meant that different species of macropods were included together in the study and the yellow-footed rock wallaby (YFRW) was overrepresented. Larger sample sizes, and comparison between macropod species and zoological collections, would be beneficial in future studies.
Conclusion
Despite individual-to-individual variation, bacterial communities likely undergo largely conserved changes during PD36. For the first time, we have profiled the shift in oral microbiota of captive macropods at different stages of MPPD, as well as characterised the healthy gingival microbiome. Porphyromonas, Fusobacterium, Bacteroides, and Desulfomicrobium may play key roles in this disease, as they appear at higher prevalence in MPPD cases compared with healthy animals. Overall, these results support a polymicrobial pathogenesis of MPPD and suggest that the diversity of bacteria involved, and the interactions between them, may be more complex than has been documented previously. Improving our understanding of the pathogenesis of MPPD is key to the development of more effective preventative and therapeutic measures.
Materials and methods
Ethics
All experiments and experimental protocols were approved by Zoos South Australia Animal Welfare and Ethics Committee, Australia. All experiments were performed in accordance with relevant guidelines and regulations of Zoos South Australia as well as School of Animal and Veterinary Sciences, The University of Adelaide. The study was carried out in compliance with the ARRIVE guidelines (https://arriveguidelines.org).
Selected species, clinical assessment and sample collection
Periodontal plaque samples were collected opportunistically by veterinarians during routine preventative medicine health checks or investigation of clinical oral disease from 22 captive individuals of the Macropodidae and Potoroidae families at Adelaide Zoo (AZ; n = 7) and Monarto Zoo (MZ; n = 15) in South Australia. Species sampled included Yellow-footed rock wallaby (YFRW, Petrogale xanthopus n = 19), Black-flanked rock wallaby (BFRW, P. lateralis, n = 1), Long-nosed potoroo (LNP, Potorous tridactylus, n = 1) and Tammar wallaby (TW, Notamacropus eugenii, n = 6). Most were captive born at AZ or elsewhere in South Australia and had been resident at their respective zoo from < 1 to 11 years. At AZ, wallabies were maintained on a grassed enclosure, supplemented with kangaroo pellets (Wombaroo Food Products, Glen Osmond, South Australia), lucerne chaff, browse, carrots and spinach. The LNP was fed a mixed diet of dog kibble, kangaroo pellets, vegetables, fruits and seeds. At Monarto, an open range zoo, wallabies were maintained in large enclosures with native vegetation, and fed ad lib meadow hay, with supplemental kangaroo pellets, lucerne chaff, and browse. Meta-data of experiment is presented at Supplementary 9.
Animals were anaesthetised with gaseous isoflurane in oxygen via face mask. Two subgingival plaque samples were collected from different quadrants of the mouth using sterile cuvettes, or whole teeth collected in the case of extraction. Animals with clinical disease were only included if they were sampled prior to commencement of antimicrobial treatment. Samples were placed immediately in transport media (10% glycerol in Wilkins Chalgren broth) and stored at either − 80 °C (AZ) or − 20 °C (MZ). Samples were transported on dry ice to the University of Adelaide, School of Animal and Veterinary Sciences, and stored at − 80 °C until analysis.
Medical records (including dental charts and diagnostic imaging where available) were used to classify oral health status at the time of sampling. Animals were classified similarly to Antiabong et al.24 as healthy, gingivitis and periodontitis-osteomyelitis cases, based predominantly on dental examination, with gingivitis and periodontitis-osteomyelitis cases respresenting early and advanced cases of MPPD, respectively. Individuals with gingivitis had gross swelling, redness of the gums, bleeding on swabbing, minor gingival recession and/or early periodontal pocket formation. Periodontitis-osteomyelitis was defined here as periodontitis with or without more progressive disease associated with soft tissue and/or bone involvement. Individuals were classified with periodontitis-osteomyelitis if, in addition to gingivitis, they had severe gingival recession, deep periodontal pockets, tooth mobility, bone necrosis, and/or other more severe lesions. Healthy animals were classified by the absence of the above gross lesions. For MPPD cases, the sample analysed was that collected from the affected tooth. Additional findings such as plaque in healthy animals (in the absence of gingivitis), other disease conditions present, episodes of previous dental disease and pouch status were also recorded.
DNA extraction
The periodontal plaque samples were thawed at 37 °C in an anaerobic chamber (Coy, Grasslakes, Michigan, USA) and then vortexed for 20 s. DNA extraction was performed using the QIAGEN DNeasy Blood and Tissue Kit (Qiagen Hilden, Germany) spin-column protocol, modified to improve DNA yield by using 400 µL of sample, 40 µL of Proteinase K, 400 µL buffer AL and 400 µL ethanol. The elution step was performed twice, using 70µL and then 30 µL of elution buffer. Sample DNA concentrations and quality were tested using Nanodrop 1000 spectrophotometry (Thermo Fisher, Waltham, Massachusetts, USA). Concentrations were also measured with a Qubit Fluorimeter, following the assay preparation instructions from Qubit dsDNA HS Assay Kits (Thermo Fisher).
Illumina Miseq 16S rRNA gene sequencing
Published primers were used for the amplification of the 550 bp V3-V4 region of the bacterial 16S rRNA gene. PCR products were visualised following electrophoresis in agarose and staining with Gel Red™ to confirm positive yield for each sample. Samples were submitted to the South Australian Health & Medical Research Institute DNA Sequencing Facility for 16S Microbiota library preparation and sequencing. Library preparation followed the Illumina library preparation protocol, with the following primers: forward CCTACGGGNGGCWGCAG, reverse GACTACHVGGGTATCTAATCC. Sequencing was carried out by Illumina Miseq V3 SBS Chemistry targeting machine. Amplicons were sequenced as paired reads with the length of 300 bp (2 × 300 bp).
Microbiota profiling
Adapter trimming, fixed length trimming, merging paired reads and filtering based on the number of reads (to remove the samples with low coverage) were performed to obtain high quality sequence reads with enough depth for microbiota profiling and comparison as previously described37. CLC Microbial Genomics Module (QIAGEN) Version 11 was used to assign taxonomy to the reads from different samples. To this end, reads were clustered using representative sequences of pseudo-species called OTUs (operational taxonomic units). The OTU clustering tool clusters the reads and reduces the read collection in each sample to representative sequences (cluster centroids) that are 97% similar to any member of the cluster they represent. The SILVA database was used as the reference of 16S rRNAs38. The number of reads assigned to each OTU and the relative abundance of each OTU was calculated39.
Statistical analysis
Linear discriminant analysis (LDA) effect size (LEfSe)40 algorithm, a test for high-dimensional biomarker discovery in metagenomic data, was used to find differentially abundant bacteria between periodontitis-osteomyelitis versus healthy as well as gingivitis versus healthy samples. The following criteria were used for selection of bacteria with statistically different relative abundance: alpha value = 0.05 and threshold of absolute logarithmic LDA score > 2. Analysis was performed in the Galaxy platform (https://huttenhower.sph.harvard.edu/galaxy/).
LEfSe algorithm benefits from a range of tests: (1) Non-parametric factorial Kruskal–Wallis (KW) sum-rank test to find bacteria with statistical significant differences between groups; (2) a set of pairwise tests among subclasses using the (unpaired) Wilcoxon rank-sum test; and (3) LDA for estimating the effect size of each differentially abundant bacteria as well as feature selection (dimension reduction)40.
Alpha diversity estimates the diversity within samples. Alpha diversity was performed based on Shannon Index. Kruskal–Wallis H test was used for measuring the statistical significance of alpha diversity. Kruskal–Wallis H assesses whether the values originate from the same distribution or whether their distribution is different depending on the group they belong to. This test is a nonparametric alternative to ANOVA. A significant p-value for the Kruskal–Wallis test means that at least one group has a different distribution. However, Kruskal–Wallis does not report which pairs have different distributions. Mann–Whitney U test was used to performs a pair-wise test to specifically find which pairs of groups follow different distributions.
Beta diversity measures the change in diversity between groups. Beta diversity was calculated in using CLC Microbial Genomics Module in two steps: (1) estimating the distance between each pair of samples; and (2) performing Principal Coordinate Analysis (PCoA) on the distance matrices. Bray–Curtis measurement was used to calculate the distance matrices.
PERMANOVA test41, also known as non-parameteric MANOVA, measures the effect of size and significance on beta diversity in comparisons of gingivitis group versus healthy and periodontitis-osteomyelitis versus healthy samples. PERMANOVA obtains its significance from a permutation test. The number of permutations was set to 99,999. Analysis was performed by CLC Microbial Genomics Module.
References
Smith, J. A. Macropod nutrition. Vet. Clin. N. Am. Exotic Anim. Pract. 12, 197–208 (2009).
Vogelnest, L. & Woods, R. Medicine of Australian Mammals (CSIRO Publishing, 2008).
Antiabong, J. F., Boardman, W., Moore, R. B., Brown, M. H. & Ball, A. S. The oral microbial community of gingivitis and lumpy jaw in captive macropods. Res. Vet. Sci. 95, 996–1005 (2013).
Graves, D. T., Kayal, R. A., Oates, T. & Garlet, G. P. Osteoimmunology 2nd edn, 325–344 (Elsevier, 2015).
Hughes, F. J. Stem Cell Biology and Tissue Engineering in Dental Sciences 433–444 (Elsevier, 2015).
Sampaio-Maia, B., Caldas, I., Pereira, M., Pérez-Mongiovi, D. & Araujo, R. Advances in applied microbiology Vol. 97, 171–210 (Elsevier, 2016).
Bodley, K. & McCracken, H. E. Dental disease in macropod species at Melbourne Zoo. Internal recommendations (2002).
Burton, J. D. Studies into Aspects of Lumpy Jaw in Macropods (University of Melbourne, 1981).
Butcher, R. et al. Periodontal disease in free-ranging koalas (Phascolarctos cinereus) from the Mount Lofty Ranges, South Australia, and its association with koala retrovirus infection. Aust. Vet. J. 98, 200–206 (2020).
Kolenbrander, P. E., Palmer, R. J., Periasamy, S. & Jakubovics, N. S. Oral multispecies biofilm development and the key role of cell–cell distance. Nat. Rev. Microbiol. 8, 471–480 (2010).
Welch, J. L. M., Rossetti, B. J., Rieken, C. W., Dewhirst, F. E. & Borisy, G. G. Biogeography of a human oral microbiome at the micron scale. Proc. Natl. Acad. Sci. 113, E791–E800 (2016).
Hienz, S. A. & Ivanovski, S. Functional Occlusion in Restorative Dentistry and Prosthodontics 23–32 (Elsevier, 2016).
Hajishengallis, G. & Lamont, R. J. Beyond the red complex and into more complexity: The polymicrobial synergy and dysbiosis (PSD) model of periodontal disease etiology. Mol. Oral Microbiol. 27, 409–419 (2012).
Rosier, B. T., De Jager, M., Zaura, E. & Krom, B. P. Historical and contemporary hypotheses on the development of oral diseases: Are we there yet?. Front. Cell. Infect. Microbiol. 4, 92 (2014).
Antiabong, J., Boardman, W. & Ball, A. What can we learn from the microbial ecological interactions associated with polymicrobial diseases?. Vet. Immunol. Immunopathol. 158, 30–36 (2014).
Bird, P. S. et al. Porphyromonas loveana sp. nov., isolated from the oral cavity of Australian marsupials. Int. J. Syst. Evol. Microbiol. 66, 3771–3778 (2016).
Oliphant, J., Parsons, R. & Smith, G. Aetiological agents of necrobacillosis in captive wallabies. Res. Vet. Sci. 36, 382–384 (1984).
Samuel, J. Jaw disease in macropod marsupials: Bacterial flora isolated from lesions and from the mouths of affected animals. Vet. Microbiol. 8, 373–387 (1983).
Brookins, M. D., Rajeev, S., Thornhill, T. D., Kreinheder, K. & Miller, D. L. Mandibular and maxillary osteomyelitis and myositis in a captive herd of red kangaroos (Macropus rufus). J. Vet. Diagn. Investig. 20, 846–849 (2008).
Keane, C. et al. Bacteroides ruminicola as a possible cause of “lumpy-jaw” in Bennett’s wallabies. Vet. Microbiol. 2, 179–183 (1977).
Asperger, M. Zur Ätiologie und Bekämpfung der Lumpy Jaw Disease bei Kängurus (2003).
Beighton, D. & Miller, W. A. A microbiological study of normal flora of macropod dental plaque. J. Dent. Res. 56, 995–1000 (1977).
Samuel, J. The normal flora of the mouths of macropods (Marsupialia: macropodidae). Arch. Oral Biol. 27, 141–146 (1982).
Antiabong, J. F. et al. A molecular survey of a captive wallaby population for periodontopathogens and the co-incidence of Fusobacterium necrophorum subspecies necrophorum with periodontal diseases. Vet. Microbiol. 163, 335–343 (2013).
Chhour, K.-L., Hinds, L. A., Jacques, N. A. & Deane, E. M. An observational study of the microbiome of the maternal pouch and saliva of the tammar wallaby, Macropus eugenii, and of the gastrointestinal tract of the pouch young. Microbiology 156, 798–808 (2010).
Gulino, L.-M. et al. Shedding light on the microbial community of the macropod foregut using 454-amplicon pyrosequencing. PLoS ONE 8, e61463 (2013).
Cheng, Y. et al. The Tasmanian devil microbiome—implications for conservation and management. Microbiome 3, 76 (2015).
Goupil, M. T., Banki, M. & Ferneini, E. M. Osteomyelitis and osteonecrosis of the jaws. in Head, Neck, and Orofacial Infections (ed Hupp, J. R.) 222–231 (Elsevier, St. Louis, 2016).
Li, Y. et al. Coinfection with Fusobacterium nucleatum can enhance the attachment and invasion of Porphyromonas gingivalis or Aggregatibacter actinomycetemcomitans to human gingival epithelial cells. Arch. Oral Biol. 60, 1387–1393 (2015).
Saito, A. et al. Porphyromonas gingivalis entry into gingival epithelial cells modulated by Fusobacterium nucleatum is dependent on lipid rafts. Microb. Pathog. 53, 234–242 (2012).
Ebbers, M. et al. Interplay between P. gingivalis, F. nucleatum and A. actinomycetemcomitans in murine alveolar bone loss, arthritis onset and progression. Sci. Rep. 8, 1–10 (2018).
Langendijk, P., Kulik, E., Sandmeier, H., Meyer, J. & Van Der Hoeven, J. Isolation of Desulfomicrobium orale sp. nov. and Desulfovibrio strain NY682, oral sulfate-reducing bacteria involved in human periodontal disease. Int. J. Syst. Evol. Microbiol. 51, 1035–1044 (2001).
Vartoukian, S., Palmer, R. & Wade, W. Diversity and morphology of members of the phylum “synergistetes” in periodontal health and disease. Appl. Environ. Microbiol. 75, 3777–3786 (2009).
Alfano, N. et al. Variation in koala microbiomes within and between individuals: Effect of body region and captivity status. Sci. Rep. 5, 10189 (2015).
Chen, G. et al. Digestion under saliva, simulated gastric and small intestinal conditions and fermentation in vitro by human intestinal microbiota of polysaccharides from Fuzhuan brick tea. Food Chem. 244, 331–339 (2018).
Jorth, P. et al. Metatranscriptomics of the human oral microbiome during health and disease. MBio 5, e01012-01014 (2014).
Speight, K. et al. Oxalate-degrading bacteria, including Oxalobacter formigenes, colonise the gastrointestinal tract of healthy koalas (Phascolarctos cinereus) and those with oxalate nephrosis. Aust. Vet. J. 97, 166–170 (2019).
Quast, C. et al. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2012).
Szafranski, S. P. et al. High-resolution taxonomic profiling of the subgingival microbiome for biomarker discovery and periodontitis diagnosis. Appl. Environ. Microbiol. 81, 1047–1058 (2015).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, 1–18 (2011).
Anderson, M. J. A new method for non-parametric multivariate analysis of variance. Austral. Ecol. 26, 32–46 (2001).
Acknowledgements
We would like to thank the University of Adelaide and Zoos South Australia for supporting this study.
Author information
Authors and Affiliations
Contributions
S. Y., M.MD. and E.E. wrote the main manuscript text. S.Y. performed experiments, and M.M.D. and E.E. performed the analyses. P.S.B. reviewed and revised the manuscript. D.J.T., D.J.M., and W.S.J.B., conceived the study, arranged funding and contributed to writing and editing of the manuscript. L.S.W. contributed expertise to the analysis of data and manuscript revision and S.S. helped in performing experiments.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yip, S., Dehcheshmeh, M.M., McLelland, D.J. et al. Porphyromonas spp., Fusobacterium spp., and Bacteroides spp. dominate microbiota in the course of macropod progressive periodontal disease. Sci Rep 11, 17775 (2021). https://doi.org/10.1038/s41598-021-97057-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-97057-1
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.