Abstract
The main types of thyroid neoplasms, follicular adenoma (FA), follicular thyroid carcinoma (FTC), classical and follicular variants of papillary carcinoma (clPTC and fvPTC), and anaplastic thyroid carcinoma (ATC), differ in prognosis, progression rate and metastatic behaviour. Specific patterns of lncRNAs involved in the development of clinical and morphological features can be presumed. LncRNA landscapes within distinct benign and malignant histological variants of thyroid neoplasms were not investigated. The aim of the study was to discover long noncoding RNA landscapes common and specific to major benign and malignant histological subtypes of thyroid neoplasms. LncRNA expression in FA, FTC, fvPTC, clPTC and ATC was analysed with comprehensive microarray and RNA-Seq datasets. Putative biological functions were evaluated via enrichment analysis of coexpressed coding genes. In the results, lncRNAs common and specific to FTC, clPTC, fvPTC, and ATC were identified. The discovered lncRNAs are putatively involved in L1CAM interactions, namely, pre-mRNA processing (lncRNAs specific to FTC); PCP/CE and WNT pathways (lncRNAs specific to fvPTC); extracellular matrix organization (lncRNAs specific to clPTC); and the cell cycle (lncRNAs specific to ATC). Known oncogenic and suppressor lncRNAs (RMST, CRNDE, SLC26A4-AS1, NR2F1-AS1, and LINC00511) were aberrantly expressed in thyroid carcinomas. These findings enhance the understanding of lncRNAs in the development of subtype-specific features in thyroid cancer.
Similar content being viewed by others
Introduction
The major types of thyroid cancer are papillary carcinoma (PTC) and follicular carcinoma (FTC), accounting for approximately 70–80% and 10–15% of all thyroid cancers, respectively. Both PTC and FTC are derived from follicular cells and are well-differentiated thyroid cancers (WDTCs), with distinct mutational landscapes, clinical behaviours, typical sites of metastasis and prognostic clinical markers1. Within PTC, several variants can be distinguished, and classical (clPTC) and follicular variants (fvPTC) are the most frequently identified. fvPTC has intermediate behaviour; it is composed of neoplastic follicles rather than papillae, but with follicular cells showing nuclear features characteristic of PTC. The benign counterpart of FTC is follicular adenoma (FA), and it is often challenging to differentiate them through cytology. FA, FTC and fvPTC compose follicular-pattern thyroid tumours, sharing common mutational prevalence and clinical features1,2. Anaplastic thyroid carcinoma (ATC) is the most advanced and aggressive thyroid cancer and the least likely to respond to treatment3,4. Based on the differences in mutational landscapes, morphology and clinical behaviour of histological subtypes, specific molecular patterns, including patterns of long noncoding RNAs (lncRNAs), are expected to be associated with these features.
Evidence of the important roles of lncRNAs in tumour suppression, cancer progression, invasion and metastatic potential and their prognostic and therapeutic value is increasing5. LncRNAs are RNA molecules of more than 200 nucleotides that typically do not have a functional open reading frame (however, bifunctional RNAs have been discovered that function as both protein-coding and noncoding RNAs). Many lncRNA genes have two or more exons and display 5′-capping, polyadenylation and alternative splicing. The functions of lncRNAs are realized in different ways: recruiting transcription factors, chromatin organizers, or chromatin modifiers, forming DNA–RNA triplex anchoring effector proteins to the gene promoter, acting as decoys for miRNAs and proteins, or interfering with protein posttranslational modification5,6,7,8. Relative to the coding genes, lncRNAs can be classified into intergenic (lincRNA); antisense (on the opposite strand of a protein-coding locus); sense intronic or overlapping (on the same strand, with transcript in introns of a coding gene, or containing a coding gene in its intron); retained intron (an alternatively spliced transcript containing an intronic sequence); bidirectional (originates from the promoter region of a protein-coding gene with transcription proceeding in the opposite direction on the other strand); and 3-prime overlapping (overlap the 3′UTR of a protein-coding locus on the same strand). Today, the number of annotated lncRNA genes has reached 14 720 according to Ensembl version 939.
In thyroid cancer, several lncRNAs have been shown to have pathogenic and predictive roles, including BANCR, FALEC, CNALPTC1, PVT1, NAMA, PTCSC1, PTCSC2, PTCSC3, and TNRC6C-AS110,11,12,13,14,15,16,17,18,19,20,21. However, all of the studies to date have considered only PTC, and mostly none of the previous works takes into account the difference between clPTC and fvPTC. There are no published studies describing the landscapes of lncRNAs in ATC, FTC and FA. Nevertheless, lncRNAs differentially expressed in ATC may reflect anaplastic features and be strong prognostic factors. As the morphology and behaviour of FTC differ from those of PTC, it is proposed that the landscape of lncRNAs in FTC may be different from that of PTC. Investigation of lncRNAs common and specific to FA and FTC is important in understanding their relations and revealing differential diagnostic markers.
This study aimed to identify lncRNAs specific and common to the main types of thyroid neoplasms (FA, FTC, fvPTC, clPTC and ATC). The expression data from microarray technology (8 datasets) and RNA-Seq technology (the PRJEB11591 dataset and TCGA transcriptome data) were analysed.
Results
LncRNAs differentially expressed in thyroid neoplasms
LncRNA expression was evaluated in the main histological subtypes of thyroid neoplasms, FA, FTC, fvPTC, clPTC, and ATC, compared to those in NT. The expression of 3910 lncRNA genes in the microarray dataset, 2587 in the RNA-Seq PRJEB11591 dataset and 3009 in the RNA-Seq TCGA dataset was analysed. The number of genes analysed corresponded to the total number of lncRNAs covered by uniquely mapped probes in the Affymetrix Human Genome U133 Plus 2.0 Array and the number of lncRNAs yielded after filtration by low number of counts for RNA-Seq datasets.
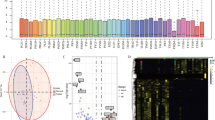
The numbers of lncRNAs found to be differentially expressed in each subtype compared to NT are presented in Table 1. The complete lists of the identified differentially expressed lncRNAs are shown in Supplementary files 1–3. Volcano plots representing the distribution of fold change and adjusted p values in the studied histological subtypes are shown in Supplementary file 4.
Hierarchical clustering of differentially expressed lncRNAs in the microarray and PRJEB11591 datasets is presented in Fig. 1. There was strong clustering of ATC, clustering of clPTC and weak clustering of fvPTC lncRNAs. No clustering of lncRNAs within the FTC or FA groups was observed (Fig. 1).
Via cross-dataset confirmation, 116 genes in clPTC were validated (45 genes found in all analysed datasets, 71 genes without probes in the microarray were found in both RNA-Seq datasets; Fig. 2A), and 62 genes in fvPTC were validated (Fig. 2B). These genes can be considered to have robustly differentially expressed lncRNAs. There are no datasets available for performing cross-dataset validation of FA, FTC or ATC genes.
In silico validation of differentially expressed lncRNAs in clPTC (A) and fvPTC (B). In clPTC, 116 genes were considered to be validated (differentially expressed in all datasets or differentially expressed in both RNA-Seq datasets but absent in microarray probes). In fvPTC, 62 genes were considered to be validated.
LncRNAs common and specific to each histological subtype were detected via intersection of the genes expressed differentially in each subtype compared to NT, and subsequent selection of lncRNAs validated in clPTC and fvPTC, and significantly differentially expressed in comparison between subtypes of neoplasms (Figs. 3, 4).
LncRNAs common to FA and WDTC
Of the 35 lncRNAs found to be differentially expressed in FA and WDTC (FTC, clPTC, and fvPTC) compared to NT, 13 genes were cross validated in clPTC and fvPTC (Figs. 3, 4, Table 2). The expression of LINC02555 and LINC02471 increased during the progression from adenoma to carcinomas and was significantly higher in fvPTC and clPTC than in FA. The expression of ENSG00000256542 and ENSG00000258117 decreased during the transition from FA to carcinomas and was significantly lower in fvPTC and clPTC than in FA or FTC.
LncRNAs common to WDTC
There were 32 lncRNAs differentially expressed in all the studied histological subtypes of WDTC (FTC, clPTC, and fvPTC) but not in FA (Fig. 3). Of these lncRNAs, 6 lncRNAs were validated to be significantly differentially expressed in clPTC and fvPTC compared to FA (Fig. 4, Table 3). None of the 32 lncRNAs were differentially expressed in FTC compared to FA.
LncRNAs common to papillary carcinomas
There were 22 genes differentially expressed in both clPTC and fvPTC, but not in FA or FTC (Fig. 3), validated and significantly differentially expressed compared to FA and FTC (Fig. 4, Table 4)—lnRNAs are putatively associated with papillary features in thyroid carcinomas.
LncRNA specific to histological subtypes of differentiated carcinomas
Nineteen lncRNAs were aberrantly expressed in FTC but not in the other studied neoplasms and were significantly differentially expressed compared to those in clPTC and fvPTC (Figs. 3, 4, Table 5). However, none of these lncRNAs was differentially expressed compared to those in FA.
Of the 29 genes differentially expressed in fvPTC but not in other differentiated carcinomas or FA (Figs. 3, 4), only the ENSG00000257647 gene was specific to fvPTC-validated and significantly differentially expressed in fvPTC compared to FA, FTC and clPTC.
The 32 genes were found to be differentially expressed in clPTC but not in other differentiated carcinomas or FA, validated, and significantly differentially expressed compared to those in fvPTC, FTC and FA-lncRNAs specific to clPTC (Figs. 3, 4, Table 6).
LncRNA specific to ATC
ATC samples were available only in the microarray dataset, which also included two variants of PTC. Of the 376 lncRNAs differentially expressed in ATC compared to NT, 252 were not differentially expressed in the other investigated histological subtypes, and 185 were significantly differentially expressed compared to those in clPTC and fvPTC-lncRNAs specific to ATC. The 30 most differentially expressed genes are presented in Table 7, and the full list is shown in Supplementary file 5.
Potential biological functions of aberrantly expressed lncRNAs
The coexpressed genes for each lncRNA from the top 5 most differentially expressed list for discussed groups were identified. The number of coexpressed genes was 138.5 (46.25 − 256.5) −median (Q1 − Q3).
Analysis of the enrichment of GO biological processes and GO molecular functions, KEGG, and Reactome terms with coexpressed coding genes allowed us to identify putative pathways involving common and specific lncRNAs (Fig. 5). The main functions of the lncRNAs common to FA and WDTC-cancers (Colorectal, Non-small cell lung, Thyroid) and p53 signalling; functions of the lncRNAs common to WDTC-Pathways in cancer and L1CAM interactions; functions of the lncRNAs common to papillary carcinomas-aldehyde dehydrogenase (NAD) activity; functions of the lncRNAs specific to FTC-processing of capped intron-containing pre-mRNA; functions of the lncRNAs specific to fvPTC-PCP/CE pathway and Beta-catenin independent WNT signalling; functions of the lncRNAs specific to clPTC-extracellular matrix organization and endoderm formation; and functions of the lncRNAs specific to ATC-cell cycle and mitotic processes.
Putative biological process involving aberrantly expressed lncRNAs in thyroid neoplasms. The enrichment analysis of GO Biological Process, GO Molecular Function, KEGG, and Reactome terms was performed for the 5 most differentially expressed lncRNAs, and terms with adjusted p values ≤ 0.05 were considered significantly enriched. For fvPTC and ATC, the 10 most significantly enriched terms are presented.
Discussion
Histological subtypes of follicular cell-derived thyroid carcinomas (FTC, PTC, and ATC) significantly differ in their mutational landscapes and clinical characteristics. Although FTC and clPTC are both WDTCs, FTC is characterized by a follicular growth pattern and tends more often to spread as metastases to distant organs, while clPTC typically has papillary architecture and spreads more often to lymph nodes in the neck. In FTC, K/H/NRAS and PAX8/PPARG mutations are prevalent, whereas BRAF mutations and tyrosine kinase fusions prevail in clPTC1. The clinical characteristics of fvPTC are intermediate; fvPTC is composed of neoplastic follicles not papillae, but with follicular cells showing nuclear features typical of PTC22. The mutational profile of fvPTC is most similar to that of FTC: both exhibit a prevalence of K/H/NRAS and PAX8/PPARG mutations. In a previous TCGA study, fvPTC was characterized as a Ras-like tumour, and its classification as a papillary carcinoma was questioned23. Recently, reclassification of encapsulated fvPTC as a “noninvasive follicular thyroid neoplasm with papillary-like nuclear features” (NIFTP) was proposed2. ATC is an advanced stage thyroid neoplasm and is the most aggressive thyroid cancer. It is expected that there are specific molecular features, including lncRNA patterns, associated with the clinical and histological features of WDTC and the aggressive behaviour of ATC. FA is thought to be a benign counterpart of FTC, and understanding the common and different molecular features of these neoplasms is important for the development of diagnostic and therapeutic strategies.
In this study, the expression of lncRNAs was evaluated in the main histological subtypes of thyroid neoplasms: FA, FTC, fvPTC, clPTC and ATC. Datasets analysed in the study (a microarray dataset of 8 independent experiments; RNA-Seq PRJEB11591; and RNA-Seq TCGA) allowed us to perform robust cross-dataset validation of the results for clPTC and fvPTC and to include representative sets of FA, FTC and ATC samples. LncRNA landscapes in FA, FTC and ATC were analysed for the first time. The highest number of genes aberrantly expressed compared to normal thyroid tissue were found in ATC (330 lncRNAs), followed by clPTC, FTC and fvPTC, which reflects the more advanced stage of ATC. Since the data for ATC, FA and FTC wer limited with one dataset only, the results for these subtypes are preliminary.
Intersection of the differentially expressed lncRNAs and subsequent comparison of the expression between subtypes of neoplasms led to the discovery of lncRNAs common to FA and WDTC (13 genes), common to WDTC (6 genes), common to classical and follicular variants of PTC (22 genes), and specific to FTC (19 genes), fvPTC (1 gene), clPTC (32 genes), and ATC (185 genes). The discovered lncRNAs were proposed to be involved in the development of clinical and morphological features of the studied subtypes. Putative biological processes involving common and specific lncRNAs were identified.
LncRNAs common to all studied thyroid neoplasms (including FA) and common to WDTC appear to be involved in carcinogenesis in different locations. L1CAM interactions found in this study to involve lncRNAs common to WDTC have been previously associated with well-described roles in tumour progression, metastases and the epithelial-to-mesenchymal transition24,25.
LncRNAs common to follicular and classical variants of papillary carcinoma (associated with papillary histology) are involved in aldehyde dehydrogenase (NAD) activity. Aldehyde dehydrogenase is known to maintain cancer stem cell properties in various cancers, including the thyroid26.
Biological processes involving lncRNAs specific to FTC include processes that are associated with splicing (Processing of Capped Intron-Containing Pre-mRNA, mRNA Splicing, and RNA processing). Accumulating evidence suggests that aberrant RNA splicing is a common and driving event in cancer development and progression. For instance, oncogenic Ras signalling via the ERK and PI3-K/Akt pathways regulates the phosphorylation of splicing factors such as SRSF1, SRSF7, and SPF45 and drives the switching of active and inactive states of tumour promoters and suppressors (MST1R, FAS, CD44, LBR, Casp-9, KLF6, and others) via alternative splicing27,28. None of the lncRNAs specific to FTC were differentially expressed compared to those in FA. The absence of lncRNAs differentially expressed in FTC and FA corresponds to the commonality of these subtypes and frequent difficulty in cytology-based differential diagnostics.
Only lncRNA ENSG00000257647 is specific to fvPTC, which might be explained by its intermediate morphology with features of both papillary and follicular carcinomas, leading to its debatable classification. LncRNA ENSG00000257647 appeared to be involved in WNT signalling, predominantly through the Beta-catenin-independent WNT pathway (especially, planar cell polarity that modulates cytoskeleton rearrangements through the activation of the small GTPases RhoA and Rac and their downstream effectors Rock and JNK). WNT signalling is known to play a crucial role in thyroid carcinogenesis, and several mechanisms of its deregulation have been described, including inhibition of the β-catenin degradation complex via its phosphorylation by RET/PTC, inhibition of E-cadherin expression through the MAPK/ERK pathway activated by BRAF mutations, and activation of both canonical and non-canonical Wnt pathways by RAS mutations29,30.
LncRNAs specific to clPTC are involved in extracellular matrix organization and endoderm and collagen formation. Extracellular matrix (ECM) disorganization is known to play a pivotal role in cancer initiation and progression. The major driving mutation in clPTC is BRAF p.V600E, and there is emerging evidence of ECM remodelling induced by BRAF p.V600E in PTCs31. Notably, it has been previously shown that the extracellular matrix of PTCs driven by BRAF p. V600E (but not mutant HRAS) is enriched with stromal-derived fibrillar collagen and facilitates cancer progression32.
lncRNAs specific to ATC are probably associated with its anaplastic features and aggressive behaviour. For these lncRNAs, there is a strong enrichment of cell cycle and mitotic pathways which possibly reflects the involvement of these lncRNAs in the loss of differentiation and high proliferation rate characteristic of ATC.
Of lncRNAs previously described in thyroid cancer, we found that PTCSC3 was downregulated in all investigated neoplasms, including FA; TNRC6C-AS1 was upregulated in papillary carcinomas; and PVT1 was specifically upregulated in ATC11,17,18. Other lncRNAs previously described in thyroid malignancy (BANCR, NAMA, CNALPTC1, FALEC, and PTCSC2) were not identified in our study10,12,13,16,20,21. A possible explanation is the strong association of these lncRNAs with specific mutations and the heterogeneity of driving mutations within the same subtype; for example, the aberrant expression of BANCR is driven by BRAF mutation.
The aberrant expression of lncRNAs with Ensembl annotation found by Liyanarachchi et al. (2016) in PTC was confirmed in our study14. Most of these lncRNAs were common to thyroid neoplasms (including FA) or common to classical and follicular variants of PTC. No lncRNAs found in our study to be subtype specific were discovered by Liyanarachchi.
In the present study, we identified some lncRNAs with known roles in tumorigenesis but not previously described in thyroid cancer33,34,35,36,37. The identified upregulated promoters of cancer progression included NR2F1-AS1 and LINC00511 in clPTC and CRNDE in ATC; downregulated tumour suppressors SLC26A4-AS1 in clPTC and RMST in ATC.
Conclusion
LncRNAs common to FA and WDTC, common to WDTC, common to carcinomas with papillary features, and specific to clPTC, fvPTC, FTC and ATC were discovered in the analysis performed with the most comprehensive datasets (combination of a microarray dataset and two RNA-Seq datasets). The similarity of the lncRNA landscapes in FTC and FA was revealed. The results showed that LncRNAs common to FA and WDTC and common to WDTC are involved in pathways in cancer at various sites, p53 signalling and L1CAM interactions; lncRNAs common to papillary carcinomas are involved in aldehyde dehydrogenase (NAD) activity; lncRNAs specific to FTC are involved in mRNA processing; a lncRNA specific to fvPTC is involved in planar cell polarity and WNT signalling; lncRNAs specific to clPTC are involved in extracellular matrix organization and endoderm formation; and lncRNAs specific to ATC are involved in the cell cycle and mitotic processes; and LncRNAs found to be specific to ATC, including CRNDE and RMST, are likely associated with cancer aggressiveness and cancer progression.
Materials and methods
Microarray datasets
The microarray datasets obtained from Affymetrix Human Genome U133 Plus 2.0 Array (Platform GPL570) were originally selected from the GEO database. The following datasets were included: GSE3467, GSE60542, GSE35570, GSE76039, GSE53157, GSE33630, GSE65144, and GSE29265. A total of 107 samples of normal tissue (NT) and 32 fvPTC, 48 clPTC, and 49 ATC samples were analysed. CEL files were downloaded, and normalization was performed using the gcrma R package. Microarray probes were annotated with Ensembl version 93 using the biomaRt package38.
RNA-Seq datasets
The RNA-Seq dataset PRJEB11591 of Yoo et al.39 was selected from the EBI European Nucleotide Archive database (https://www.ebi.ac.uk/ena/data/view/PRJEB11591). PRJEB11591 is the most comprehensive available RNA-Seq dataset containing benign and malignant thyroid neoplasms (FA, FTC, fvPTC, and clPTC). The PRJEB11591 samples included 81 NT, 26 FA, 30 FTC, 48 fvPTC and 77 clPTC samples. FASTQ files were downloaded, and alignment was performed by HISAT240. Counts were calculated using featureCounts (Rsubread package) with annotation by Ensembl version 93 and Ensemble gene ID for grouping attributes41. Genes with low counts (less than 2 counts in number of samples exceeding the size of the smallest sample group) were eliminated, and TMM normalization (edgeR package) and the voom method using the limma R package were applied.
In the TCGA transcriptome data, 58 NT, 356 clPTC and 101 fvPTC were selected. Samples of metastases and other minor histological subtypes were excluded. Raw counts (HTSeq-Counts Workflow Type, briefly, STAR 2-pass alignment followed by gene expression count assessment with HTSeq) were downloaded from Genomic Data Commons Data Portal (GDC, https://portal.gdc.cancer.gov/). Genes with low counts (less than 1 count in number of samples exceeding the size of smallest sample group) were eliminated, followed by TMM normalization (edgeR package) and voom analysis with limma42.
Selection of lncRNA genes
Protein-coding genes and genes attributed to Havana biotypes not related to lncRNAs were eliminated. Genes of the following Havana biotypes were included in the analysis: lincRNA, antisense, 3-prime overlapping ncRNA, bidirectional promoter lncRNA, misc RNA, processed transcript, sense intronic, and sense overlapping.
Statistical analysis
To identify differentially expressed lncRNAs, linear modelling using the limma package was performed43. Genes with FDR adjusted p value ≤ 0.01 and fold change (FC) ≥ 2.0 were considered to be differentially expressed. A heat map analysis of differentially expressed genes was performed using coolmap limma.
Validation
For clPTC and fvPTC, the sets of genes found to be significantly differentially expressed in a previous step in the microarray, RNA-Seq PRJEB11591, and RNA-Seq TCGA datasets were processed with intersection. Genes found in all three datasets and genes found in both RNA-Seq datasets but not in microarray probes were considered validated.
Selection of lncRNAs common and specific to histological subtypes
LncRNAs common and specific to FA and WDTC were selected via the intersection analysis of genes found to be significantly differentially expressed in each subtype compared to NT in the RNA-Seq dataset PRJEB11591 and subsequent application following criteria:
-
Common to FA and WDTC-confirmed through validation for clPTC and fvPTC;
-
Common to WDTC-confirmed through validation for clPTC and fvPTC, and significantly differentially expressed compared to FA;
-
Common to papillary carcinomas-confirmed through validation for clPTC and fvPTC, and significantly differentially expressed compared to FA and FTC;
-
Specific to clPTC, fvPTC, FTC-confirmed through validation for clPTC and fvPTC (not applied to FTC), and significantly differentially expressed compared to each studied subtype;
LncRNAs specific to ATC were selected from intersection with genes found in FA and WDTC with subsequent filtration of genes significantly differentially expressed compared to those in clPTC and fvPTC.
Evaluation of potential biological functions
To identify genes positively and negatively coexpressed with 5 most differentially expressed lncRNAs, pairwise Pearson correlation between the lncRNAs and all the genes was calculated using the RNA-Seq PRJEB11591 dataset (for FA, FTC, fvPTC and clPTC) and the microarray dataset (for ATC). Genes with an absolute r ≥ 0.7 and a significant correlation (p value < 0.05) were considered to be coexpressed. For coexpressed genes, enrichment of Gene Ontology (GO) Biological Process (2018), GO Molecular Function (2018), Kyoto Encyclopedia of Genes and Genomes (KEGG, 2019) and Reactome (2016) terms was estimated using Enrichr44,45. Terms with adjusted p values in Fisher's exact test ≤ 0.05 were considered significantly enriched42,43,44,45,46.
References
Acquaviva, G. et al. Molecular pathology of thyroid tumours of follicular cells: A review of genetic alterations and their clinicopathological relevance. Histopathology 72(1), 6–31 (2018).
Nikiforov, Y. E. et al. Nomenclature revision for encapsulated follicular variant of papillary thyroid carcinoma. JAMA Oncol. 2(8), 1023 (2016).
Cabanillas, M. E., Zafereo, M., Gunn, G. B. & Ferrarotto, R. Anaplastic thyroid carcinoma: Treatment in the age of molecular targeted therapy. J. Oncol. Pract. 12, 511–518 (2016).
Landa, I. et al. Genomic and transcriptomic hallmarks of poorly differentiated and anaplastic thyroid cancers. J. Clin. Invest. 126, 1052–1066 (2016).
Bure, I. V., Kuznetsova, E. B. & Zaletaev, D. V. Long noncoding RNAs and their role in oncogenesis. Mol. Biol. 52(6), 787–798 (2018).
Derrien, T. et al. The GENCODE v7 catalog of human long noncoding RNAs: Analysis of their gene structure, evolution, and expression. Genome Res. 22(9), 1775–1789 (2012).
Fang, Y. & Fullwood, M. J. Roles, functions, and mechanisms of long non-coding RNAs in cancer. Genom. Proteom. Bioinform. 14(1), 42–54 (2016).
Li, R., Zhu, H. & Luo, Y. Understanding the functions of long non-coding RNAs through their higher-order structures. Int. J. Mol. Sci. 17(5), E702 (2016).
Zerbino, D. R. et al. Ensembl 2018. Nucleic Acids Res. 46(D1), D754–D761 (2018).
Yoon, H. et al. Identification of a novel noncoding RNA gene, NAMA, that is downregulated in papillary thyroid carcinoma with BRAF mutation and associated with growth arrest. Int. J. Cancer. 121, 767–775 (2007).
He, H. et al. A susceptibility locus for papillary thyroid carcinoma on chromosome 8q24. Cancer Res. 69, 625–631 (2009).
Jeong, S. et al. Relationship of focally amplified long noncoding on chromosome 1 (FAL1) lncRNA with E2F transcription factors in thyroid cancer. Medicine 95, e2592 (2016).
Liao, T. et al. BRAF-activated LncRNA functions as a tumor suppressor in papillary thyroid cancer. Oncotarget https://doi.org/10.18632/oncotarget.10825 (2016).
Liyanarachchi, S. et al. Genome-wide expression screening discloses long noncoding RNAs involved in thyroid carcinogenesis. J. Clin. Endocrinol. Metab. https://doi.org/10.1210/jc.2016-1991 (2016).
Wang, Q. et al. Identification of specific long non-coding RNA expression: Profile and analysis of association with clinicopathologic characteristics and BRAF mutation in papillary thyroid cancer. Thyroid 26, 1719–1732 (2016).
Zheng, H. et al. BRAF-activated long noncoding RNA modulates papillary thyroid carcinoma cell proliferation through regulating thyroid stimulating hormone receptor. Cancer Res. Treat. 48, 698–707 (2016).
Zhou, Q., Chen, J., Feng, J. & Wang, J. Long noncoding RNA PVT1 modulates thyroid cancer cell proliferation by recruiting EZH2 and regulating thyroid-stimulating hormone receptor (TSHR). Tumor Biol. 37, 3105–3113 (2016).
Muhanhali, D. et al. Long non-coding antisense RNA TNRC6C-AS1 is activated in papillary thyroid cancer and promotes cancer progression by suppressing TNRC6C expression. Front. Endocrinol. (Lausanne) 9, 360 (2018).
Hou, S., Lin, Q., Guan, F. & Lin, C. LncRNA TNRC6C-AS1 regulates UNC5B in thyroid cancer to influence cell proliferation, migration, and invasion as a competing endogenous RNA of miR-129-5p. J. Cell. Biochem. https://doi.org/10.1002/jcb.26868 (2018).
Chen, C. et al. Long noncoding RNA CNALPTC1 promotes cell proliferation and migration of papillary thyroid cancer via sponging miR-30 family. Am. J. Cancer Res. 8, 192–206 (2018).
Huang, H. et al. LncRNA NR2F1-AS1 regulates hepatocellular carcinoma oxaliplatin resistance by targeting ABCC1 via miR-363. J. Cell. Mol. Med. 22, 3238–3245 (2018).
Daniels, G. H. Follicular variant of papillary thyroid carcinoma: Hybrid or mixture?. Thyroid 26(7), 872–874 (2016).
Agrawal, N. et al. Integrated genomic characterization of papillary thyroid carcinoma. Cell 159, 676–690 (2014).
Altevogt, P., Doberstein, K. & Fogel, M. L1CAM in human cancer. Int. J. Cancer 138(7), 1565–1576 (2016).
Maten, M. V., Reijnen, C., Pijnenborg, J. M. A. & Zegers, M. M. L1 cell adhesion molecule in cancer, a systematic review on domain-specific functions. Int. J. Mol. Sci. 20(17), 4180 (2019).
Lee, S., Bae, J. S., Jung, C. K. & Chung, W. Y. Extensive lymphatic spread of papillary thyroid microcarcinoma is associated with an increase in expression of genes involved in epithelial-mesenchymal transition and cancer stem cell-like properties. Cancer Med. 8(15), 6528–6537 (2019).
Gonçalves, V., Pereira, J. F. S. & Jordan, P. Signaling pathways driving aberrant splicing in cancer cells. Genes (Basel) 9(1), 9 (2017).
Yea, S. et al. Ras promotes growth by alternative splicing-mediated inactivation of the KLF6 tumor suppressor in hepatocellular carcinoma. Gastroenterology 134, 1521–1531 (2008).
Sastre-Perona, A., Riesco-Eizaguirre, G., Zaballos, M. A. & Santisteban, P. β-Catenin signaling is required for RAS-driven thyroid cancer through PI3K activation. Oncotarget 7, 49435–49449 (2016).
Ely, K. A., Bischoff, L. A. & Weiss, V. L. Wnt signaling in thyroid homeostasis and carcinogenesis. Genes (Basel) 9(4), 204 (2018).
Nucera, C., Lawler, J. & Parangi, S. BRAF(V600E) and microenvironment in thyroid cancer: A functional link to drive cancer progression. Cancer Res. 71, 2417–2422 (2011).
Jolly, L. A. Fibroblast-mediated collagen remodeling within the tumor microenvironment facilitates progression of thyroid cancers driven by BrafV600E and Pten loss. Cancer Res. 76(7), 1804–1813 (2016).
Sun, C.-C. et al. Long intergenic noncoding RNA 00511 acts as an oncogene in non–small-cell lung cancer by binding to EZH2 and suppressing p57. Mol. Ther. Nucleic Acids. 5, e385 (2016).
Xu, S., Kong, D., Chen, Q., Ping, Y. & Pang, D. Oncogenic long noncoding RNA landscape in breast cancer. Mol. Cancer 16(1), 129 (2017).
Han, P. et al. The lncRNA CRNDE promotes colorectal cancer cell proliferation and chemoresistance via miR-181a-5p-mediated regulation of Wnt/β-catenin signaling. Mol. Cancer 16(1), 9 (2017).
Xie, H. et al. Long non-coding RNA CRNDE in cancer prognosis: Review and meta-analysis. Clin. Chim. Acta 485, 262–271 (2018).
Wang, L. et al. Long non-coding RNA (LncRNA) RMST in triple-negative breast cancer (TNBC): Expression analysis and biological roles research. J. Cell. Physiol. 233, 6603–6612 (2018).
Durinck, S., Spellman, P. T., Birney, E. & Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 4, 1184–1191 (2009).
Yoo, S. K. et al. Comprehensive analysis of the transcriptional and mutational landscape of follicular and papillary thyroid cancers. PLoS Genet. 12(8), e1006239. https://doi.org/10.1371/journal.pgen.1006239 (2016).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods. 12, 357–360 (2015).
Liao, Y., Smyth, G. K. & Shi, W. The Subread aligner: Fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 41, e108–e108 (2013).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297 (2012).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Chen, E. Y. et al. Enrichr: Interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform. 14, 128 (2013).
Kuleshov, M. V. et al. Enrichr: A comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
He, H.-T. et al. Biomarker and competing endogenous RNA potential of tumor-specific long noncoding RNA in chromophobe renal cell carcinoma. Oncol. Targets Ther. 9(6399–6406), 6399–6406 (2016).
Funding
The research was supported by a Russian Science Foundation (project No. 19-75-00053).
Author information
Authors and Affiliations
Contributions
V.D.Y.—analysis, or interpretation of data for the work, conception or design of the work; V.V.S.—revising the work; A.S.T.—revising the work; A.V.L.—supervision, conception or design of the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yakushina, V.D., Strelnikov, V.V., Tanas, A.S. et al. Long noncoding RNA landscapes specific to benign and malignant thyroid neoplasms of distinct histological subtypes. Sci Rep 11, 16728 (2021). https://doi.org/10.1038/s41598-021-96149-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-96149-2
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.