Abstract
(+)-7-iso-Jasmonoyl-l-isoleucine (JA-Ile) is a lipid-derived phytohormone implicated in plant development, reproduction, and defense in response to pathogens and herbivorous insects. All these effects are instigated by the perception of JA-Ile by the COI1-JAZ co-receptor in the plant body, which in Arabidopsis thaliana is profoundly influenced by the short JAZ degron sequence (V/L)P(Q/I)AR(R/K) of the JAZ protein. Here, we report that SlJAZ-SlCOI1, the COI1-JAZ co-receptor found in the tomato plant, relies on the extended JAZ degron sequence (V/L)P(Q/I)AR(R/K)XSLX instead of the canonical JAZ degron. This finding illuminates our understanding of the mechanism of ligand perception by JA-Ile in this plant, and will inform both efforts to improve it by genetic modification of the SlCOI1-SlJAZ co-receptor, and the development of the synthetic agonists/antagonists.
Similar content being viewed by others
Introduction
Lipid-derived (+)-7-iso-jasmonoyl-l-isoleucine (JA-Ile) is a major class of phytohormones implicated in plant development, reproduction, and the defense response of plants against pathogens and herbivorous insects1,2. Exposure of plants to external stress causes jasmonate signaling, which is triggered by the production of JA-Ile from jasmonic acid (JA)3,4. Perception of JA-Ile by the COI1-JAZ co-receptor causes upregulation of JA-responsive genes, leading to the expression of JAZ (JASMONATE ZIM-DOMAIN) repressor protein –a hub of jasmonate signaling5,6.
The JAZ protein contains two major functional domains: Jas and TIFY. In the absence of JA-Ile, the Jas domain causes JAZ to interact with various transcription factors (TFs), including the master regulator MYC27. The TIFY domain of JAZ interacts with domain C of the adaptor protein Novel Interactor of JAZ (NINJA), the ethylene-responsive element binding factor-associated amphiphilic repression (EAR) domain of which recruits co-repressor TOPLESS (TPL)8. These interactions constitute a transcriptional repression machinery consisting of the MYC2-JAZ-NINJA-TPL complex, which represses the expression of JA-responsive gene. In the presence of JA-Ile, JAZ also plays an important role in releasing the repression of TFs. The Jas motif is also important for protein–protein interactions with COI1, a subunit of SCFCOI1E3 ubiquitin ligase, which forms the COI1-JA-Ile-JAZ co-receptor complex with the JA-Ile ligand9. JA-Ile causes COI1-JAZ co-receptor formation and subsequent ubiquitination and degradation of JAZ to derepress the expression of JA-responsive genes10,11,12.
Recently, it has become clear that the JAZ gene functions redundantly; however, each JAZ subfamily gene is also responsible for its own unique function13,14,15,16,17, and differences in the sequence of the Jas motif profoundly influence the unique function of each JAZ. This is partly because the function of JAZ depends on how strongly and with which of the many TFs it interacts13. The Jas motif also affects the lifespan of each JAZ in plant cells (through the affinity between JAZ and COI1), which in turn determines the duration of effect of each JAZ’s unique function13.
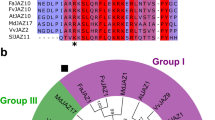
The crystal structure of the COI1-JA-Ile-JAZ1 degron peptide, which also includes inositol pentakisphosphate (InsP5), revealed that the short JAZ degron sequence in the Jas motif is responsible for COI1-JA-Ile-JAZ1 complex formation, based on the discovery of a hydrogen bond network between the JAZ1 degron sequence LPIARR and COI1-JA-Ile complex18. JAZ belongs to the TIFY protein family, but only the JAZ subfamily incorporates the Jas motif, including the short canonical degron sequence LPIAR(R/K) necessary for binding with JA-Ile (Fig. 1a,b). The JAZ degron sequence is highly conserved among plant species, except for the two non-canonical degron sequences IPMQRK of FaJAZ1 found in the strawberry plant (Fragaria × ananassa), and MPIARK of EcJAZ1 found in finger millet Eleusine coracana19,20,21.
Slight differences in the Arabidopsis JAZ degron sequences (identity 37.0% in Fig. 1a) are thought to account for the difference in affinity (KD) between JAZ and COI1-JA-Ile22. However, the 12 JAZs of Solanum lycopersicum have a remarkably well-conserved JAZ degron sequence (V/L)P(Q/I)AR(R/K) (identity 46.6% in Fig. 1a), suggesting their affinities (KD) for the SlCOI1-JA-Ile complex are very similar (Fig. 1a). Here, we report the first comprehensive study of the affinity of the SlJAZ-SlCOI1 co-receptor for JA-Ile, and demonstrate that the perception of JA-Ile depends on the extended JAZ degron sequence (V/L)P(Q/I)AR(R/K)XSLX. This use of an extended degron sequence (instead of the canonical short JAZ degron sequence) accounts for the different affinities of SlJAZ and SlCOI1-JA-Ile.
Results
The affinities of SlJAZ1-11/13 for SlCOI1-JA-Ile are different
All the TIFY sequences of the JAZ proteins of Solanum lycopersicum have been previously reported: 19 TIFY genes including 12 canonical JAZ (SlJAZ1-11/13) and non-canonical SlJAZ12 genes are encoded in the Solanum lycopersicum genome23,24,25. The Jas motifs of the 12 canonical SlJAZs are remarkably similar to the Arabidopsis consensus Jas motif SLX2FX2KRX2RX5PY, and the JAZ degron sequence (V/L)P(Q/I)AR(R/K), wherein the hydrophobic L/V is conjugated with P(Q/I)AR and followed by basic R/K is conserved in eight out of 12 SlJAZs (SlJAZ1-6/8/13) (Figs. 1a,b). Accordingly, SlCOI1-SlJAZ1-6/8/13 are expected to perceive JA-Ile with equal affinity, whereas SlCOI1-SlJAZ9-11 (which incorporate a non-canonical JAZ degron) are not expected to perceive it at all. Accordingly, we examined the affinities of 12 SlJAZ proteins for the SlCOI1-JA-Ile complex by pull-down assay.
SlJAZ genes were cloned from the tomato cultivar Micro-Tom and their FLAG-tag-fused proteins FLAG-SlJAZ1-11/13 expressed using the wheat germ-derived cell-free protein expression system (Supplementary Fig. S1). GST-fused SlCOI1 (GST-SlCOI1) was also expressed in Sf9 cultured insect cells (Supplementary Fig. S2). Arabidopsis ASK protein was co-expressed to improve the stability of SlCOI126. As expected, SlJAZ1-3/5–8 but not FLAG-SlJAZ9-11 pulled-down GST-SlCOI1 in the presence of 100 nM JA-Ile (Fig. 2a and Supplementary Fig. S3). However, SlJAZ4/13 (which incorporates the same canonical JAZ degron LPIARR as SlJAZ1-3) could not pull-down the GST-SlCOI1 under the same conditions (Figs. 1b and 2a). Identical results were obtained using coronatine (COR), a naturally occurring phytotoxin and known structural mimic of JA-Ile, suggesting that COR is perceived by the co-receptor in place of JA-Ile in a similar manner to JA-Ile (Supplementary Fig. S4). These results indicate that sequences other than the highly conserved SlJAZ degron in full-length JAZ affect the perception of JA-Ile by the SlCOI1-SlJAZ co-receptor.
(a) Pull down assay of GST-SlCOI1 with FLAG-SlJAZ (full-length proteins) in the presence of JA-Ile (100 nM). GST-SlCOI1 bound to FLAG-SlJAZ proteins was pulled down with anti-FLAG antibody and Protein G magnetic beads, and analyzed by immunoblotting (top: anti-GST-HRP conjugate for detection of GST-SlCOI1, bottom: anti-FLAG antibody and anti mouse-IgG HRP conjugate for detection of FLAG-SlJAZs). *or **show the bands derived from heavy chain or light chain of the anti-FLAG antibody. (b) Pull down assay of GST-SlCOI1 with Fl-SlJAZPs in the presence of JA-Ile (100 nM). GST-SlCOI1 bound to Fl-SlJAZPs was pulled down with anti-fluorescein antibody and Protein G magnetic beads, and analyzed by immunoblotting (anti-GST-HRP conjugate for detection of GST-SlCOI1).
To quantitatively examine the effect of the exo-degron sequence, we designed and synthesized fluorescein-tagged SlJAZ1-11/13 degron short peptides (SlJAZP1-11/13), each 27 amino acids in length (Fig. 3a and Supplementary Figs. S5–S6), based on previous work on Arabidopsis COI1-JAZ22. The pull-down assay using the GST-SlCOI1 and Fl-SlJAZ degron peptides in the presence of increasing concentrations of JA-Ile yielded very similar results to those obtained using full-length SlJAZs (Fig. 2b and Supplementary Fig. S7). Therefore, the affinity of full-length JAZ was confirmed to depend on the sequence in these short peptides. The observed affinities were quantitatively assessed in AlphaScreen luminescence proximity assays using SlJAZPs and GST-SlCOI1 in the presence of 0–30 µM JA-Ile (Fig. 3b and Supplementary Fig. S8)7,27, and found to be in good accordance with the results obtained by pull-down assay: 20 > Kd for SlJAZ1/5-8 of strong affinity, 150 > Kd for SlJAZ2/3 of weak affinity, Kd > 400 for SlJAZ4/9-11/13 of no/little affinity (Table 1). Similar results were obtained using COR (Supplementary Figs. S9 and S10).
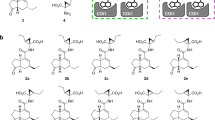
(a) Chemical structures of fluorescein-tagged SlJAZ1-11/13 degron short peptides (SlJAZP1-11/13). Canonical JAZ degron sequences are shown in blue and down-stream-of-degron (DOD) sequence are in orange. (b) AlphaScreen assays using Fl-SlJAZPs and GST-SlCOI1 with JA-Ile (0–30 µM). Experiments were performed in triplicate to obtain mean and S.D. (shown as error bars). The figure was created by KaleidaGraph 4.1.1 (Synergy, Software, US).
Hormone perception relies on extended JAZ degron sequences in tomato SlJAZs
Here, we focused on the relationship between the degron sequences of SlJAZPs and their Kd values. The short JAZ degron sequence (L/V)P(Q/I)AR(R/K) was highly conserved in SlJAZP1-6/8/13 (Fig. 3a). SlJAZP5/6/8, which incorporate the JAZ degron sequence VPQARK, all have strong affinity for SlCOI1-JA-Ile. In contrast, remarkable differences in affinity were observed for SlJAZP1-4/13, whose degron sequence is LPIARR: only SlJAZP1 showed a moderate affinity for SlCOI1-JA-Ile; the others had weak/no affinity. Among SlJAZ1-4/13, the difference can be found in the downstream-of-degron (DOD) sequence XSLX (Fig. 3a). This strongly suggests that sequences more extended than the canonical JAZ degron influence the affinity of SlJAZs for SlCOI1-JA-Ile.
To confirm the effect of the exo-JAZ-degron sequence within SlJAZPs on their affinity, we prepared chimeric SlJAZPs incorporating swapped sequences and submitted them to the AlphaScreen assay. First, we swapped the two JAZ-degron sequences VPQARK of SlJAZP5/6/8 and LPIARR of SlJAZP1-4/13 to examine the effect of the JAZ degron sequence on affinity. The N-terminal region of the high-affinity peptide SlJAZP5 including JAZ-degron VPQARK was swapped with that of the moderate/low-affinity SlJAZP1/4 including JAZ-degron LPIARR, to give the swapped peptide SlJAZP1/2/4-5 (Fig. 4a, Supplementary Figs. S11 and S12). Then, we examined whether the difference in the JAZ degron sequence of SlJAZs affected the affinity with SlCOI1-JA-Ile (Fig. 4b and Supplementary Fig. S13a). As shown in Fig. 4b, the high affinity of SlJAZP5 was moderately decreased by swapping with SlJAZP1/4 (Kd = 4.2 nM for SlJAZP5 to Kd = 16.5 nM for SlJAZP1/2/4-5). The effect of swapping was moderate, and the complete swapping of their affinities did not occur. This result suggested that the differences in JAZ degron alone cannot fully account for the difference in their affinities, which must therefore be influenced by exo-JAZ-degron sequences in addition to the canonical JAZ degron.
(a) Design of the swapped SlJAZ peptides (SlJAZ5, 1/2/4-5, 1-3DOD, 1-4DOD 5-2DOD, 2-5DOD, 2-5, 5-2). Canonical JAZ degron sequences are shown in blue and down-stream-of-degron (DOD) sequence are in orange. (b) AlphaScreen assay using GST-SlCOI1 and Fl-SlJAZP1/2/4-5 (black circle) or Fl-SlJAZ5 (red square) with JA-Ile (0–300 nM). (c–h) Alphascreen assays of swapped Fl-SlJAZPs (c: SlJAZ1-3DOD, d: 1-4DOD, e: 5-2DOD, f: 2-5DOD, g: 5–2, h: 2–5) to study the extended degron sequence. Signal intensity changes of AlphaScreen of swapped Fl-SlJAZPs and GST-SlCOI1 upon addition of JA-Ile (0–10 µM). Black circles show the results of swapped SlJAZPs and red squares those of the corresponding natural SlJAZPs, respectively. Experiments were performed in triplicate to obtain mean and S.D. (shown as error bars). Significant differences were evaluated by Students’ T-test (*p < 0.05, **p < 0.01). These figures were created by KaleidaGraph 4.1.1 (Synergy, Software, US).
To examine the effect of the DOD sequence, we studied SlJAZP1-4/13 which incorporate the same JAZ degron sequence LPIARR and alternative DOD sequence XSLX. We focused on three SlJAZPs: SlJAZP1, which is of strong affinity (Kd = 9.2 nM); SlJAZP3, of moderate affinity (Kd = 136 nM); and SlJAZP4, of no affinity (Kd = 1776 nM). We prepared SlJAZP1-3DOD and SlJAZP1-4DOD, wherein the DOD sequence of SlJAZP1 was swapped with those of SlJAZP3 and SlJAZP4, respectively (Fig. 4a, and Supplementary Figs. S11–12). As shown in Fig. 4c,d (and Supplementary Fig. S13b, c), their affinities for SlCOI1-JA-Ile were moderately decreased (Kd = 28.4 nM for SlJAZP1-3DOD and Kd = 46.9 nM for SlJAZP1-4DOD). This result confirmed that the DOD sequence, in addition to the JAZ degron, affected the affinity between SlJAZ and SlCOI1-JA-Ile. Next, we examined whether the DOD sequence also affected the affinity of SlJAZP5/6/8 with another JAZ degron sequence VPQARK. As SlJAZP5/6/8 has the same DOD sequence ASLA, we replaced the DOD of SlJAZP5 of strong affinity (Kd = 4.2 nM) with that of SlJAZ2 of weak affinity (Kd = 134 nM). We prepared DOD swapped peptides SlJAZP5-2DOD (ASLA of SlJAZ5 to NSLT of SlJAZ2) and SlJAZP2-5DOD (NSLT of SlJAZ2 to ASLA of SlJAZ5) (Fig. 4a, and Supplementary Figs. S11–12). Their affinities for SlCOI1-JA-Ile were affected by this swapping (Kd = 11.7 nM for SlJAZP5-2DOD and Kd = 42.7 nM for SlJAZP2-5DOD, Fig. 4e,f and Supplementary Fig. S13d, e). This result suggested that the DOD sequence in SlJAZ5/6/8 also affects affinity for SlCOI1-JA-Ile.
We also swapped the extended JAZ degron sequence including the JAZ-degron and DOD to prepare swapped peptides of SlJAZP5-2 (DLPIARRNSLT of SlJAZ2 into AVPQARKASLA of SlJAZ5) and SlJAZP2-5 (AVPQARKASLA of SlJAZ5 into DLPIARRNSLT of SlJAZ2) (Fig. 4a, and Supplementary Figs. S11–12). Their affinities with SlCOI1-JA-Ile were completely replaced by this sequence swapping (Kd = 88.8 nM for SlJAZP5-2 and Kd = 5.0 nM for SlJAZP2-5, respectively, Fig. 4g,h and Supplementary Fig. S13f, g). This result confirmed that the extended JAZ degron sequence including JAZ-degron and DOD affects the affinity of in SlJAZ with SlCOI1-JA-Ile.
From all of these results, we concluded that extended-JAZ-degron sequences, VPQARKASLA or LPIARRXSLX, determine the affinity of SlJAZs with SlCOI1-JA-Ile.
In silico simulation demonstrated the role of extended JAZ degron sequences in SlCOI1-JA-Ile-SlJAZ complex formation
In silico docking simulation studies using the crystal structure of Arabidopsis COI1-JA-Ile-JAZ1 have previously enabled the study of the JA-Ile binding mode of the COI1-JAZ co-receptor of other plant species, such as Phaseolus lunatus, Eleusine coracana (L.) Gaertn, Fragaria vesca, and Fragaria × ananassa20,21,28,29. The contribution of the extended JAZ degron sequence to SlCOI1-JA-Ile-SlJAZ complex formation was examined by comparing the crystal structure of Arabidopsis COI1-JA-Ile-JAZ1 with the in silico interaction models of SlCOI1-JAIle-SlJAZ1 and SlCOI1-JA-Ile-SlJAZ5. A homology model of SlCOI1 was obtained with the software Molecular Operating Environment (MOE) using the crystal structure of AtCOI1 (PDB ID: 3OGL) as a template (the sequence identity of SlCOI1 and AtCOI1 is 68.1%). AtJAZ1 in the homology model of SlCOI1-JA-Ile-AtJAZ1 was replaced with SlJAZ1 or SlJAZ5. The obtained models of these complexes (SlCOI1-JA-Ile-SlJAZ1 and SlCOI1-JA-Ile-SlJAZ5) were then used for subsequent molecular dynamics (MD) simulations. Root means square deviation (RMSD) values indicated that the structures reached equilibrium after 50 ns or more (Fig. 5a,b). There were no significant differences in the canonical JAZ degron sequence (LPIARR) interactions in AtJAZ1/SlCOI1 with AtCOI1-JA-Ile/SlJAZ1-JA-Ile, nor in the hydrogen-bonding network formed around JA-Ile (Figs. 5c–e, 6a–f, and Supplementary Fig. S14) in any of the three complexes. In addition, no direct interaction between the DOD sequences of SlJAZs and JA-Ile was observed in SlCOI1-JA-Ile-SlJAZ1/5. These results suggest that the DOD sequence of SlJAZs does not interact with JA-Ile. Therefore the DOD sequence may enhance the SlCOI1-SlJAZs interaction. Next, we compared the interaction between the DOD sequence and SlCOI1 in SlCOI1-JA-Ile-SlJAZ1/5 with the interaction between the corresponding sequence in AtJAZ1 (ASLH) and AtCOI1. In AtCOI1-JA-Ile-AtJAZ1, a weak interaction through one hydrogen bond was found between the DOD sequence and AtCOI1 (Fig. 6g), whereas in SlCOI1-JA-Ile-SlJAZ1 and SlCOI1-JA-Ile-SlJAZ5, a strong interaction through several hydrogen bonds or hydrophobic interactions was found (Fig. 6b,c,h,i, and Supplementary Fig. S15). This indicates that the DOD sequence in SlJAZ significantly contributes to the formation of the SlCOI1-JA-Ile-SlJAZ complex, consistent with the results of the previous experiments. However, the results in Table 1 in which SlCOI1-SlJAZ1 has a lower affinity for JA-Ile/COR than SlCOI1-SlJAZ5 cannot be explained by the in silico docking studies because of the slight differences in their affinities.
(a,b) The RMSDs of backbone atoms of the complex of SlCOI1-JA-Ile-SlJAZ1 (a) and SlCOI1-JA-Ile-SlJAZ5 (b) as a function of time. Each plot depicted in blue, red, orange, purple, or green was obtained from the five independent MD simulation. (c) The reported structure of the ligand binding pocket of AtCOI1-JA-Ile-AtJAZ1 (PDB ID: 3OGL). (d,e) The representative structure of the ligand binding pocket of SlCOI1-JA-Ile-SlJAZ1 (d) and that of SlCOI1-JA-Ile-SlJAZ5 (e), which was obtained by homology modeling and MD simulation. Hydrogen bonds are shown as yellow dotted lines. The RMSD plot was created by KaleidaGraph 4.1.1 (Synergy, Software, US), and the images of the 3D structures were created by MOE 2020.09.
In silico analyses of AtCOI1-JA-Ile-AtJAZ1 (PDB ID: 3OGL, a,d,g), SlCOI1-JA-Ile-SlJAZ1 (b,e,h), and SlCOI1-JA-Ile-SlJAZ5 (c,f,i) to show the binding mode of the extended degron. (a–c) The amino acid residues forming hydrogen bonds (H-bond) or hydrophobic interaction (Hydrophobic) between COI1 and JAZ observed in the crystal structure of AtCOI1-JA-Ile-AtJAZ1 (3OGL, a), MD simulation of SlCOI1-JA-Ile-SlJAZ1 (b) and MD simulation of SlCOI1-JA-Ile-SlJAZ5 (c). (d) The reported structure around the degron sequence of AtJAZ1 in the complex of AtCOI1-JA-Ile-AtJAZ1. (e,f) The obtained MD structure around the degron sequence of SlJAZ in the complex of SlCOI1-JA-Ile-SlJAZ1 (e) or SlCOI1-JA-Ile-SlJAZ5 (f). (g) The reported structure around the DOD sequence of AtJAZ1 in the complex of AtCOI1-JA-Ile-AtJAZ1. (h,i) The obtained MD structure around the DOD sequence of SlJAZ in the complex of SlCOI1-JA-Ile-SlJAZ1 (h) or SlCOI1-JA-Ile-SlJAZ5 (i). Hydrogen bonds are shown as yellow dotted lines. The images of the 3D structures were created by MOE 2020.09.
Discussion
JAZ functions as a repressor of numerous TFs in plant cells, and the differences in function between JAZ family proteins are profoundly affected by how strongly and with which of the many TFs each JAZ interacts. Since JAZs interact with many TFs through the Jas motif, differences in this motif can account for differences in the function of JAZs. In addition, JAZ interacts with the COI1-JA-Ile complex using the degron sequence within the Jas motif, and ubiquitinated and degraded as a substrate for E3 ubiquitin ligase. A slight difference in the degron sequence of each JAZ can profoundly impact the duration of JAZ function because it affects the lifetime of each JAZ in the plant cell. Therefore, differences in the strength of the interaction between each JAZ and the COI1-JA-Ile complex affect the turn-over of each JAZ30.
This study is the first comprehensive investigation into the affinity of the SlJAZ-SlCOI1 co-receptor for JA-Ile. SlJAZ9-11, having a non-canonical JAZ degron sequence QPQTVK, were found to have no affinity for SlCOI1-JA-Ile. However, in transcript expression of SlJAZs on JA-treated Micro-Tom, SlJAZ9-11 were also JA-responsive: SlJAZ9/10 were weakly induced in both roots and leaves, and SlJAZ11 was strongly induced in roots but weakly in leaves23. In Arabidopsis, non-canonical AtJAZ7/8/13 lacking a conserved degron sequence do not have affinity for AtCOI1-JA-Ile and play a unique role in transcriptional repression in plant cells31,32. AtJAZ10.4, an alternative splice variant of JAZ10, is involved in the negative feedback regulation of JA signaling and is responsible for the delayed repression of activated JA signals33. A similar function is inferred for JA-responsive SlJAZ9-11, despite its lack of affinity for SlCOI1-JA-Ile: SlJAZ9-11 may act as repressors, similar to AtJAZ7/8/13, or the SlCOI1-SlJAZ9-11 pairs could perceive another ligand. The same QPQTVK sequence is also observed in a JAZ member of Solanum tuberosum19.
The crystal structure of the Arabidopsis AtCOI1-JA-Ile-AtJAZ1 complex revealed important details regarding the interaction between AtJAZ and AtCOI1-JA-Ile complex. The short canonical degron sequence LPIARR of AtJAZ1 overlies the top of JA-Ile binding pocket of AtCOI1, covering the JA-Ile bound to AtCOI1, and interacting with both AtCOI1 and JA-Ile. Detailed analyses revealed that each amino acid in the LPIARR sequence interacts with JA-Ile by hydrogen-bond formation (LPIARR) or hydrophobic interaction (LPIARR). Thus, any change in the conserved LPIAR(R/K) sequence will affect the affinity between AtJAZ and AtCOI1-JA-Ile. Comparison of the amino acid sequences of the JAZ degron among the Arabidopsis functional JAZs showed slight differences from LPIARR of AtJAZ1, which will be responsible for the difference in affinity for AtCOI1-JA-Ile among the AtJAZs (Kd 7–34 for JA-Ile on fluorescence anisotropy assay)34. Small differences in the Arabidopsis JAZ degron sequences are considered responsible for the difference in their affinity.
In contrast, the degron sequence differences among functional SlJAZs are smaller than those of Arabidopsis JAZs, but each SlJAZ nevertheless binds with different affinities to SlCOI1-JA-Ile (Figs. 2, 3 and Table 1). In particular, significant discrepancies in the affinities of SlJAZ1-4/13 for SlCOI1-JA-Ile were observed, even though their JAZ degron sequences are identical.
We prepared the swapped SlJAZPs and found that their affinity with SlCOI1-JA-Ile strongly depends on the extended JAZ degron sequence of VPQARKASLA or LPIARRXSLX (Fig. 4). The short degron sequence (V/L)P(Q/I)AR(R/K) has been hypothesized to play a critical role in hormone reception due to its high degree of conservation across numerous plant species. The reported exceptions are the cases of strawberry JAZ1 (FaJAZ1) of the non-canonical JAZ degron sequence IPMQRK and finger millet EcJAZ1 of the non-canonical JAZ degron sequence MPIARK. Our result confirmed that tomato SlJAZs employ the extended JAZ degron sequence of (V/L)P(Q/I)AR(R/K)XSLX for the perception of JA-Ile. The contribution of this extended JAZ-degron sequence in SlCOI1-JA-Ile-SlJAZs complex formation was further studied using the in silico interaction models generated from the reported crystal structure of Arabidopsis COI1-JA-Ile-JAZ1 (Fig. 5). The DOD sequence in SlJAZ formed a strong hydrogen bond network with SlCOI1, stabilizing the complex (Fig. 6). In interaction models of SlCOI1-JA-Ile-SlJAZ1/5, the canonical JAZ degron of SlJAZs retained the same interaction as that of AtJAZ in AtCOI1-JA-Ile-AtJAZ118. Then, why do SlJAZ1/2/3/4/13, which have LPIARR as a common canonical JAZ degron, have different affinities for SlCOI1-JA-Ile? Swapping experiments have shown that this is due to differences in the DOD sequence; when the DOD sequence of SlJAZ1 was replaced with that of SlJAZ3/4, the affinity was markedly decreased (SlJAZ1-3/4DOD in Fig. 4a,c,d), indicating that the DOD sequence of SlJAZ3/4 negatively affects the interaction with SlCOI1-JA-Ile. Specifically, two point-mutations in the DOD sequence of SlJAZ1, A166S and T168H/Y, are presumed to negatively affect the interaction between the LPIARR sequence and SlCOI1-JA-Ile. This is the first report to demonstrate that sequences longer than the canonical JAZ degron are employed for hormone perception of COI1-JAZ co-receptor.
SlJAZ7 is unique among SlJAZs having three amino acid residues in the extended JAZ degron (175A, 176M, and 181T) which are not found in other SlJAZs (Fig. 3a). SlJAZ7 of a moderate affinity (Kd = 16.2 nM) (Table 1) has the extended JAZ degron sequence of LAMARRATLA in which the P(Q/I) in the canonical degron (V/L)P(Q/I)AR(R/K) is replaced by 175A176M and the highly conserved S in the DOD sequence XSLX is replaced by 181T. In the Arabidopsis COI1-JA-Ile-JAZ1 crystal structure, the PI sequence in the canonical JAZ degron of AtJAZ1 plays an important role in the interaction with AtCOI118. This suggests that SlJAZ7 may interact with SlCOI1-JA-Ile in a manner different from other SlJAZs. Remarkably, the extended JAZ degron sequence LAMARRATLA in SlJAZ7 (belonging to the strong affinity group) is also present in the other two Solanaceae species, Solanum tuberosum and Petunia axilaris19. Moreover, the LAMARRATLA sequence is only present in dicots, particularly in Solanaceae19, suggesting that the LAMARR degron is a new functional degron in addition to the most common JAZ degrons LPIARR and VPQARK.
Here, we report that the affinity of SlJAZ for SlCOI-JA-Ile depends on the extended degron sequence. Recently, a similar result was reported in the auxin co-receptor TIR1-AUX/IAA, in which additional, non-hormone interacting sequences of AUX/IAA re-enforcing auxin perception35. The role and flexibility of intrinsically disordered domains in hormone perception attract attentions of plant biologists.
Conclusion
The affinity of SlJAZ for SlCOI-JA-Ile depends on the extended degron sequence and not on the canonical degron sequence, based on the result of a comprehensive study of ligand perception of SlJAZ-SlCOI1. SlJAZ9-11 have no affinity for SlCOI-JA-Ile but are also JA-inducible23, which suggests that they have unique functions in plant cells, such as suppression of JA signaling. This finding illuminates our understanding of the mechanism of hormone perception in the edible tomato plant. Genetic modification of the SlCOI1-SlJAZ co-receptor may improve hormone perception, and lead to the development of the synthetic agonists/antagonists.
Materials and methods
All chemical reagents and solvents were obtained from commercial suppliers (Wako Pure Chemical Industries Co. Ltd., Nacalai Tesque Co., Ltd., Watanabe Chemical Industries Co. Ltd., Thermo Fisher Scientific K.K., GE Healthcare) and used without further purification. Coronatine (COR) and (+)-7-iso-JA-L-Ile (JA-Ile) were prepared as previously described1,36,37. DNA purification was accomplished using a GENE PERP STAR PI-80X (KURABO, Osaka, Japan). Ultraviolet (UV)-visible spectra were recorded on a UV-2600 spectrophotometer (Shimadzu, Kyoto, Japan). The AlphaScreen assay was carried out on an EnVision multimode plate reader (PerkinElmer, Inc., CA, US). SDS-PAGE and Western blotting were performed using Mini-Protean III (Bio-Rad Laboratories, Inc., US), Tras-Blot Turbo (Bio-Rad Laboratories, Inc., US), and iBind Flex (Thermo Fisher Scientific K.K., CA, US) electrophoresis apparatus. Chemiluminescent images were detected using the Amersham Imager 680 (GE Healthcare, CA, US). Reverse-phase high-performance liquid chromatography (HPLC) was performed on a PU-4180 plus with UV-4075 and MD-4010 detectors (JASCO, Tokyo, Japan). Absorbance at 220 nm and 488 nm was monitored by an MD-4010 photodiode array detector (PDA). MALDI-TOF MS analysis was performed on an Autoflex Max (Bruker Daltonics Inc., MA, US). 3D structures were constructed using MOE 2020.09 software (Chemical Computing Groups, Montreal, Canada).
Preparation of the SlCOI1 and SlJAZs proteins
Standard methods for cloning were used, and PCR-amplified DNA fragments were sequenced after cloning into the vectors. The plasmids of GST-fused AtCOI1 or AtASK1 (pFB-GTE-COI1 and pFB-HTB-ASK1) were obtained from Addgene (https://www.addgene.org/), and the plasmid for wheat-derived cell-free protein expression system (pEU-FLAG-GW-STOP) was a kind gift from Drs. Koji Miyamoto (Teikyo University), Kazunori Okada (The University of Tokyo), and Tatsuya Sawasaki (Ehime University). The full-length CDS of SlCOI1 was obtained from Osaka Prefecture University (kindly supported by Prof. Koh Aoki) and cloned into pFB-GTE-COI1, to prepare the plasmid of GST-fused SlCOI1. These SlCOI1 and AtASK1 proteins were co-expressed in insect cells and purified by Glutathione Sepharose 4B (GE Healthcare) according to the previous reports18,34,38. The full-length CDS of SlJAZ2/3/5/6/7 were obtained from Osaka Prefecture University (kindly supported by Prof. Koh Aoki), and PCR-amplified and cloned into the pDONR221 vector (Invitrogen, CA, US) using BP reaction (Gateway). The coding sequences of SlJAZ1/4 were isolated from Solanum cDNA using the primers. PCR-amplified SlJAZ1/4 DNA was cloned into pENTR/D-TOPO (Thermo Fisher Scientific, USA). The coding sequences of SlJAZ8/9/10/11/13 were synthesized by the manufacturers (Eurofins Genomics K.K., Japan), and were cloned into the pDONR221 vector using the BP reaction. The CDS was then inserted into pEU-FLAG-GW-STOP vector using the LR reaction (Gateway) to prepare the plasmid for FLAG tag-fused SlJAZ (pEU-FLAG-GW-SlJAZs). These SlJAZs proteins were expressed in wheat germ-derived cell-free protein expression system according to previous reports39, and used without purification. Cell-free translation reaction was performed according to the instruction protocol (Cell Free Sciences, Co., Ltd., Ehime, Japan) with minor modification. Briefly, the transcription reactions with pEU-FLAG-GW-SlJAZs (each 1 µg) were performed at 37 °C for 5 h. The obtained mixture of mRNA was added to a solution of creatine kinase and WEPRO7240, to prepare the translation mixture, carefully transferred to the bottom of a well containing translation buffer to form the bilayer, and then incubated at 15 °C for 20 h in a 96-well plate. The obtained protein mixture was centrifuged (20,000g for 15 min at 4 °C) and the supernatant was used for the pulldown experiments without any purification.
Synthesis of Fl-SlJAZPs
All SlJAZ peptides were prepared using the fully automated microwave peptide synthesizer Initiator + Alstra (Biotage Ltd, North Carolina, US) starting from Fmoc-Tyr-Wang resin (90 µm) as previously reported, with minor modifications34. A representative protocol is as follows. The resin was swollen in DMF at 70 °C for 20 min. The Fmoc protecting group was removed by treating with 20% piperidine in DMF twice. Amino acid coupling was accomplished by mixing the resin with Fmoc protected amino acids (3 eq), O-(1H-Benzotriazol-1-yl)-N,N,N′,N'-tetramethyluronium hexafluorophosphate (HBTU, 3 eq), 1-Hydroxy-1H-benzotriazole hydrate (HOBt·H2O) (3 eq), and DIPEA (6 eq) in DMF, and subjecting it to microwave irradiation at 50 °C for either 30 min (Fmoc-Arg-OH) or 10 min (all others). After the peptide had been fully elongated, the resin was mixed with 5-carboxy-fluorescein diacetate (3 eq), HBTU (5 eq) and DIPEA (5 eq) in DMF and incubated at r.t. for 2 h. After the reaction, the peptide was deprotected by stirring in TFA solution at r.t. for 1.5 h (deprotection of SlJAZ6 and 7 was performed using TFA solution containing thioanisole, anisole, and 1,2-ethanedithiol, for the avoidance of methionine oxidation). The reaction mixture was purified by HPLC using a Develosil ODS-HG-5 column (Φ 4.6 × 250 mm) eluting with a linear gradient (CH3CN (0.05% TFA):H2O (0.05% TFA) = 20:80 (5 min) to 50:50 (35 min)) to afford fluorescein-conjugated SlJAZ peptide. After lyophilization, conjugated SlJAZ peptide was dissolved in sterilized water to prepare the stock solution. The concentrations of the stock solution were determined by their absorbance at 494 nm in 0.1 N NaOH aqueous solution using a molar extinction coefficient of 75,000 M−1 cm−1. The purity of each peptide was confirmed by HPLC analyses, and these were characterized by MALDI-TOF MS as follows;
-
Fl-SlJAZ1: m/z [M + H]+ calcd for 3571,92, found 3571.92
-
Fl-SlJAZ2: m/z [M + H]+ calcd for 3573.86, found 3573.86
-
Fl-SlJAZ3: m/z [M + H]+ calcd for 3623.92, found 3623.90
-
Fl-SlJAZ4: m/z [M + H]+ calcd for 3651.85, found 3651.85
-
Fl-SlJAZ5: m/z [M + H]+ calcd for 3429.81, found 3429.80
-
Fl-SlJAZ6: m/z [M + H]+ calcd for 3500.83, found 3500.83
-
Fl-SlJAZ7: m/z [M + H]+ calcd for 3594.97, found 3594.99
-
Fl-SlJAZ8: m/z [M + H]+ calcd for 3443.80, found 3443.76
-
Fl-SlJAZ9: m/z [M + H]+ calcd for 3515.88, found 3515.84
-
Fl-SlJAZ10: m/z [M + H]+ calcd for 3656.97, found 3656.99
-
Fl-SlJAZ11: m/z [M + H]+ calcd for 3605.90, found 3605.89
-
Fl-SlJAZ13: m/z [M + H]+ calcd for 3729.97, found 3729.98
-
Fl-SlJAZ1/2/4-5: m/z [M + H]+ calcd for 3500.87, found 3500.84
-
Fl-SlJAZ1-3DOD: m/z [M + H]+ calcd for 3623.92, found 3623.93
-
Fl-SlJAZ1-4DOD: m/z [M + H]+ calcd for 3649.93, found 3649.92
-
Fl-SlJAZ2-5DOD: m/z [M + H]+ calcd for 3500.84, found 3500.84
-
Fl-SlJAZ5-2DOD: m/z [M + H]+ calcd for 3502.82, found 3502.82
-
Fl-SlJAZ2-5: m/z [M + H]+ calcd for 3429.81, found 3429.80
-
Fl-SlJAZ5-2: m/z [M + H]+ calcd for 3573.86, found 3573.87
Pulldown assay
All chemicals (JA-Ile or COR) were dissolved in ethanol to generate 10 mM stock solutions and diluted with 20% ethanol aq. for preparation of 100 μM stock solutions. For the pull-down experiments using full-length SlJAZ proteins, purified GST-COI1 (5 nM), FLAG-tagged SlJAZ (each 20 µL of the translation mixture), and the ligands (COR or JA-Ile) in 350 µL of incubation buffer (50 mM Tris–HCl buffer, pH 7.8, 100 mM NaCl, 20 mM 2-mercaptoethanol, 10% glycerol, 0.1% Tween20, 100 nM inositol-1,2,4,5,6-pentakisphosphate (IP5)) were combined with anti-FLAG antibody (0.2 µL, Sigma Aldrich, F1804, clone M2), and incubated for 10–15 h at 4 °C with rotation. After incubation, the samples were combined with SureBeads Protein G (10 µL in 50% incubation buffer slurry, Bio-Rad). After 3 h incubation at 4 °C with rotation, the samples were washed three times with 350 µL of fresh incubation buffer. The washed beads were resuspended in 35 µL of SDS-PAGE loading buffer containing dithiothreitol (DTT, 100 mM). After heating for 10 min at 60 °C, the samples were subjected to SDS-PAGE and analyzed by western blotting. The bound GST-COI1 protein was detected using anti-GST HRP conjugate (RPN1236, GE Healthcare, 5000-fold dilution in blocking buffer (Nakalai tesque, Inc., Japan). FLAG-JAZ proteins were detected using anti-FLAG antibody (1000-fold dilution in blocking buffer) and anti-mouse IgG-HRP antibody (Southern Biotech. Inc., Birmingham, US, 1031-05, 20,000-fold dilution in blocking buffer). Three independent replicates using proteins purified at different times were done with similar results (Supplementary Fig. S3).
For the pull-down experiments using fluorescein-tagged SlJAZ peptides (Fl-SlJAZps), purified GST-COI1 (5 nM), Fl-SlJAZp (10 nM), and JA-Ile (1 µM) in 350 µL of incubation buffer were combined with anti-fluorescein antibody (0.2 µL, GeneTex, CA, US), and incubated for 10–15 h at 4 °C with rotation. After incubation, the samples were combined with SureBeads Protein G (10 µL in 50% incubation buffer slurry, Bio-Rad). The washing and eluting protocols were same as the pulldown experiments using full-length JAZ proteins. Three independent replicates using proteins and peptides purified at different times were done with similar results (Supplementary Fig. S7).
AlphaScreen assay
AlphaScreen experiments were performed at 25 °C in the incubation buffer. 15 μL of the reaction mixture containing the incubation buffer, 5 nM SlCOI1, 10 nM Fl-SlJAZPs and various concentrations of COR or JA-Ile was added to a 1/2 Area AlphaPlate-96 (PerkinElmer), and then incubated for 1 h at 4 °C. Then, 10 μL of a detection mixture containing incubation buffer, 0.1 μL of FITC-coated donor beads, 0.1 μL of GST-coated acceptor beads was added to each well. Finally, the mixture was incubated for 12 h and the luminescence signals were detected using the Envision 2105 Multimode Plate Reader (PerkinElmer). The experiment was repeated three times, and the data are presented as average values with standard deviation. To clarify the difference in the affinity of the ligands with each COI1/JAZ co-receptor, the results of the AlphaScreen assay are shown with the normalized signal intensity change (Figs. 3, 4, and Supplementary Fig. S9) along with the raw signal intensity (Supplementary Figs. S8, S10, and S13)40.
In silico analyses
The homology modeling of SlCOI1 was obtained based on the crystal structure of AtCOI1–JA-Ile–AtJAZ1 (PDB ID: 3OGL). The structure preparation program MOE 2020.09 was used to deduce the structures of the absent residues (residues 550–562) of AtCOI1. The model structures of SlJAZ1, and 5 were constructed by mutating residues of the AtJAZ1 peptide of the complex.
MD simulations of SlCOI1-JA-Ile-SlJAZ1 and SlCOI1-JA-Ile-SlJAZ5 were performed under conditions of constant temperature and pressure (T = 300 K, P = 1 atm). A Parrinello–Rahman type thermostat41 and Nosé-Hoover barostat42 were used to control the system temperature and pressure, respectively. The Amber14SB43 and generalized amber force field (gaff)44 were assigned for the protein/peptide and the ligand molecule, respectively. The TIP3P model45 was used for water solvent. The cutoff length for van der Waals (vdW) and coulomb interactions in real space was 12 Å. The particle mesh Ewald (PME) method46 was used for the estimation of the coulomb interactions. The time step for integration of equations of motions was 2 fs. All MD calculations were done by GROMACS2018 program47.
Snapshot structures of SlCOI1-JA-Ile-SlJAZ1 and SlCOI1-JA-Ile-SlJAZ5 were sampled every 10 ps. First, we performed 10 ns MD simulations for energy minimization/equilibration of the systems, and subsequently conducted five independent 100 ns MD simulations (total 500 ns) for each system. The system equilibrations of the protein/peptide and the ligand were monitored by the root means square displacement (RMSD) values as the simulation time step. Ligand binding modes were confirmed to be little changed in all MD simulations. The last snapshot structures were used as the representative structures to investigate the binding forms of the complex.
References
Fonseca, S. et al. (+)-7-iso-Jasmonoyl-l-isoleucine is the endogenous bioactive jasmonate. Nat. Chem. Biol. 5, 344–350. https://doi.org/10.1038/nchembio.161 (2009).
Wasternack, C. & Song, S. Jasmonates: Biosynthesis, metabolism, and signaling by proteins activating and repressing transcription. J. Exp. Bot. 68, 1303–1321. https://doi.org/10.1093/jxb/erw443 (2017).
Wasternack, C. Jasmonates: An update on biosynthesis, signal transduction and action in plant stress response, growth and development. Ann. Bot. 100, 681–697. https://doi.org/10.1093/aob/mcm079 (2007).
Wasternack, C. & Hause, B. Jasmonates: Biosynthesis, perception, signal transduction and action in plant stress response, growth and development. An update to the 2007 review in Annals of Botany. Ann. Bot. 111, 1021–1058. https://doi.org/10.1093/aob/mct067 (2013).
Pauwels, L. & Goossens, A. The JAZ proteins: A crucial interface in the jasmonate signaling cascade. Plant Cell 23, 3089–3100. https://doi.org/10.1105/tpc.111.089300 (2011).
Kazan, K. & Manners, J. M. JAZ repressors and the orchestration of phytohormone crosstalk. Trends Plant Sci. 17, 22–31. https://doi.org/10.1016/j.tplants.2011.10.006 (2012).
Zhang, F. et al. Structural basis of JAZ repression of MYC transcription factors in jasmonate signalling. Nature 525, 269–273. https://doi.org/10.1038/nature14661 (2015).
Pauwels, L. et al. NINJA connects the co-repressor TOPLESS to jasmonate signalling. Nature 464, 788–791. https://doi.org/10.1038/nature08854 (2010).
Bai, Y., Meng, Y., Huang, D., Qi, Y. & Chen, M. Origin and evolutionary analysis of the plant-specific TIFY transcription factor family. Genomics 98, 128–136. https://doi.org/10.1016/j.ygeno.2011.05.002 (2011).
Chini, A. et al. The JAZ family of repressors is the missing link in jasmonate signalling. Nature 448, 666–671. https://doi.org/10.1038/nature06006 (2007).
Thines, B. et al. JAZ repressor proteins are targets of the SCF(COI1) complex during jasmonate signalling. Nature 448, 661–665. https://doi.org/10.1038/nature05960 (2007).
Yan, Y. et al. A downstream mediator in the growth repression limb of the jasmonate pathway. Plant Cell 19, 2470–2483. https://doi.org/10.1105/tpc.107.050708 (2007).
Chini, A., Gimenez-Ibanez, S., Goossens, A. & Solano, R. Redundancy and specificity in jasmonate signalling. Curr. Opin. Plant Biol. 33, 147–156. https://doi.org/10.1016/j.pbi.2016.07.005 (2016).
Gimenez-Ibanez, S. et al. JAZ2 controls stomata dynamics during bacterial invasion. New Phytol. 213, 1378–1392. https://doi.org/10.1111/nph.14354 (2017).
Major, I. T. et al. Regulation of growth-defense balance by the JASMONATE ZIM-DOMAIN (JAZ)-MYC transcriptional module. New Phytol. https://doi.org/10.1111/nph.14638 (2017).
Guo, Q. et al. JAZ repressors of metabolic defense promote growth and reproductive fitness in Arabidopsis. Proc. Natl. Acad. Sci. U. S. A. 115, E10768–E10777. https://doi.org/10.1073/pnas.1811828115 (2018).
Demianski, A. J., Chung, K. M. & Kunkel, B. N. Analysis of Arabidopsis JAZ gene expression during Pseudomonas syringae pathogenesis. Mol. Plant Pathol. 13, 46–57. https://doi.org/10.1111/j.1364-3703.2011.00727.x (2012).
Sheard, L. B. et al. Jasmonate perception by inositol-phosphate-potentiated COI1-JAZ co-receptor. Nature 468, 400–405. https://doi.org/10.1038/nature09430 (2010).
Garrido-Bigotes, A., Valenzuela-Riffo, F. & Figueroa, C. R. Evolutionary analysis of JAZ proteins in plants: An approach in search of the ancestral sequence. Int. J. Mol. Sci. 20, 5060 (2019).
Garrido-Bigotes, A. et al. A new functional JAZ degron sequence in strawberry JAZ1 revealed by structural and interaction studies on the COI1-JA-Ile/COR-JAZs complexes. Sci. Rep. 10, 11310. https://doi.org/10.1038/s41598-020-68213-w (2020).
Sen, S., Kundu, S. & Dutta, S. K. Proteomic analysis of JAZ interacting proteins under methyl jasmonate treatment in finger millet. Plant Physiol. Biochem. 108, 79–89. https://doi.org/10.1016/j.plaphy.2016.05.033 (2016).
Takaoka, Y. et al. A comprehensive in vitro fluorescence anisotropy assay system for screening ligands of the jasmonate COI1-JAZ co-receptor in plants. J. Biol. Chem. 294, 5074–5081. https://doi.org/10.1074/jbc.RA118.006639 (2019).
Chini, A., Ben-Romdhane, W., Hassairi, A. & Aboul-Soud, M. A. M. Identification of TIFY/JAZ family genes in Solanum lycopersicum and their regulation in response to abiotic stresses. PLoS One 12, e0177381. https://doi.org/10.1371/journal.pone.0177381 (2017).
Ishiga, Y., Ishiga, T., Uppalapati, S. R. & Mysore, K. S. Jasmonate ZIM-domain (JAZ) protein regulates host and nonhost pathogen-induced cell death in tomato and Nicotiana benthamiana. PLoS One 8, e75728. https://doi.org/10.1371/journal.pone.0075728 (2013).
Sun, J. Q., Jiang, H. L. & Li, C. Y. Systemin/Jasmonate-mediated systemic defense signaling in tomato. Mol. Plant 4, 607–615. https://doi.org/10.1093/mp/ssr008 (2011).
Li, H. et al. Efficient ASK-assisted system for expression and purification of plant F-box proteins. Plant J. 92, 736–743. https://doi.org/10.1111/tpj.13708 (2017).
Melcher, K. et al. A gate-latch-lock mechanism for hormone signalling by abscisic acid receptors. Nature 462, 602–608. https://doi.org/10.1038/nature08613 (2009).
Nakamura, Y. et al. Synthesis of 6-substituted 1-oxoindanoyl isoleucine conjugates and modeling studies with the COI1-JAZ co-receptor complex of lima bean. J. Chem. Ecol. 40, 687–699. https://doi.org/10.1007/s10886-014-0469-2 (2014).
Valenzuela-Riffo, F., Garrido-Bigotes, A., Figueroa, P. M., Morales-Quintana, L. & Figueroa, C. R. Structural analysis of the woodland strawberry COI1-JAZ1 co-receptor for the plant hormone jasmonoyl-isoleucine. J. Mol. Graph. Model. 85, 250–261. https://doi.org/10.1016/j.jmgm.2018.09.004 (2018).
Tian, J. et al. The OsJAZ1 degron modulates jasmonate signaling sensitivity during rice development. Development 146, dev173419. https://doi.org/10.1242/dev.173419 (2019).
Shyu, C. et al. JAZ8 lacks a canonical degron and has an EAR motif that mediates transcriptional repression of jasmonate responses in Arabidopsis. Plant Cell 24, 536–550. https://doi.org/10.1105/tpc.111.093005 (2012).
Thireault, C. et al. Repression of jasmonate signaling by a non-TIFY JAZ protein in Arabidopsis. Plant J. 82, 669–679. https://doi.org/10.1111/tpj.12841 (2015).
Moreno, J. E. et al. Negative feedback control of jasmonate signaling by an alternative splice variant of JAZ10. Plant Physiol. 162, 1006–1017. https://doi.org/10.1104/pp.113.218164 (2013).
Takaoka, Y., Hayashi, K., Suzuki, K., Azizah, I. N. & Ueda, M. A fluorescence anisotropy-based comprehensive method for the In vitro screening of COI1-JAZs agonists and antagonists. In Jasmonate in Plant Biology: Methods and Protocols (eds. Antony, C. & Laurent, L.) 145–160. https://doi.org/10.1007/978-1-0716-0142-6_11 (Springer US, 2020).
Niemeyer, M. et al. Flexibility of intrinsically disordered degrons in AUX/IAA proteins reinforces auxin co-receptor assemblies. Nat. Commun. 11, 2277. https://doi.org/10.1038/s41467-020-16147-2 (2020).
Kato, N. et al. A scalable synthesis of (+)-coronafacic acid. Chirality 32, 423–430. https://doi.org/10.1002/chir.23172 (2020).
Okada, M. et al. Total syntheses of coronatines by exo-selective Diels–Alder reaction and their biological activities on stomatal opening. Org. Biomol. Chem. 7, 3065–3073. https://doi.org/10.1039/b905159g (2009).
Takaoka, Y. et al. A rationally designed JAZ subtype-selective agonist of jasmonate perception. Nat. Commun. 9, 3654. https://doi.org/10.1038/s41467-018-06135-y (2018).
Sawasaki, T. et al. A bilayer cell-free protein synthesis system for high-throughput screening of gene products. FEBS Lett. 514, 102–105. https://doi.org/10.1016/S0014-5793(02)02329-3 (2002).
Ullman, E. F. et al. Luminescent oxygen channeling immunoassay: Measurement of particle binding kinetics by chemiluminescence. Proc. Natl. Acad. Sci. 91, 5426–5430. https://doi.org/10.1073/pnas.91.12.5426 (1994).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: A new molecular dynamics method. J. Appl. Phys. 52, 7182–7190. https://doi.org/10.1063/1.328693 (1981).
Hoover, W. G. Canonical dynamics: Equilibrium phase-space distributions. Phys. Rev. A 31, 1695–1697. https://doi.org/10.1103/PhysRevA.31.1695 (1985).
Maier, J. A. et al. ff14SB: Improving the accuracy of protein side chain and backbone parameters from ff99SB. J. Chem. Theory Comput. 11, 3696–3713. https://doi.org/10.1021/acs.jctc.5b00255 (2015).
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general amber force field. J. Comput. Chem. 25, 1157–1174. https://doi.org/10.1002/jcc.20035 (2004).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935. https://doi.org/10.1063/1.445869 (1983).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: An N⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092. https://doi.org/10.1063/1.464397 (1993).
Abraham, M. J. et al. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25. https://doi.org/10.1016/j.softx.2015.06.001 (2015).
Acknowledgements
This work was financially supported by a Grant-in-Aid for Scientific Research from JSPS, Japan (nos. 17H06407, 18KK0162, and 20H00402 for MU, nos. 18H02101 and 19H05283 for YT, and nos. 19K05378 and 20H04791 for HS), JSPS A3 Foresight Program (MU), and JSPS Core-to-Core Program Asian Chemical Biology Initiative (MU). And we also thank to Mr. Teppei Kamimura for his technical help. The computations in this study were performed using the Advanced Center for Computing and Communication of RIKEN, Research Center for Computational Science of Institute for Molecular Science (IMS), Research Center for Advanced Computing Infrastructure of Japan Advanced Institute of Science and Technology (JAIST).
Author information
Authors and Affiliations
Contributions
M.U. conceived, designed, and coordinated the research project. M.U., Y.T., R.S., K.H., and H.S. designed the experiments and examined data. R.S., K.H., M.N., H.N., S.Y., and T.M. performed the wet experiments. H.S., Y.T., and K.H. performed in silico studies. M.U., Y.T., K.H., and R.S. wrote the main manuscript text and all figures. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Saito, R., Hayashi, K., Nomoto, H. et al. Extended JAZ degron sequence for plant hormone binding in jasmonate co-receptor of tomato SlCOI1-SlJAZ. Sci Rep 11, 13612 (2021). https://doi.org/10.1038/s41598-021-93067-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-93067-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.