Abstract
We aimed to isolate Acinetobacter baumannii (A. baumannii) from wound infections, determine their resistance and virulence profile, and assess the impact of Silver nanoparticles (AgNPs) on the bacterial growth, virulence and biofilm-related gene expression. AgNPs were synthesized and characterized using TEM, XRD and FTIR spectroscopy. A. baumannii (n = 200) were isolated and identified. Resistance pattern was determined and virulence genes (afa/draBC, cnf1, cnf2, csgA, cvaC, fimH, fyuA, ibeA, iutA, kpsMT II, PAI, papC, PapG II, III, sfa/focDE and traT) were screened using PCR. Biofilm formation was evaluated using Microtiter plate method. Then, the antimicrobial activity of AgNPs was evaluated by the well-diffusion method, growth kinetics and MIC determination. Inhibition of biofilm formation and the ability to disperse biofilms in exposure to AgNPs were evaluated. The effect of AgNPs on the expression of virulence and biofilm-related genes (bap, OmpA, abaI, csuA/B, A1S_2091, A1S_1510, A1S_0690, A1S_0114) were estimated using QRT-PCR. In vitro infection model for analyzing the antibacterial activity of AgNPs was done using a co-culture infection model of A. baumannii with human fibroblast skin cell line HFF-1 or Vero cell lines. A. baumannii had high level of resistance to antibiotics. Most of the isolates harbored the fimH, afa/draBC, cnf1, csgA and cnf2, and the majority of A. baumannii produced strong biofilms. AgNPs inhibited the growth of A. baumannii efficiently with MIC ranging from 4 to 25 µg/ml. A. baumannii showed a reduced growth rate in the presence of AgNPs. The inhibitory activity and the anti-biofilm activity of AgNPs were more pronounced against the weak biofilm producers. Moreover, AgNPs decreased the expression of kpsMII , afa/draBC,bap, OmpA, and csuA/B genes. The in vitro infection model revealed a significant antibacterial activity of AgNPs against extracellular and intracellular A. baumannii. AgNPs highly interrupted bacterial multiplication and biofilm formation. AgNPs downregulated the transcription level of important virulence and biofilm-related genes. Our findings provide an additional step towards understanding the mechanisms by which sliver nanoparticles interfere with the microbial spread and persistence.
Similar content being viewed by others
Introduction
Acinetobacter species are commonly found in the environment, however, some species within this genus are generally associated with various habitats such as soil, water, sewage, human, foods and animals. Acinetobacter baumannii (A. baumannii) has an inevitable ability to sustain harsh conditions and spread as a serious pathogen1,2,3,4. A. baumannii infects moist tissues such as mucous membranes or areas of the skin that are exposed to this pathogen, either through wound or injury5,6.
The adhesion and colonization or biofilm formation include primary stage in bacterial infections. Major adhesion virulence factors in this step include type I fimbriae (FimH) and pilli structures for attachment to the host cells7,8. Furthermore, numerous bacteria secrete toxins and extracellular enzymes which play a crucial role in the apoptosis or necrosis of epithelial cells or immunocytes. Various virulence factors of A. baumannii such as adhesins genes like kpsMII (group 2 capsule synthesis) and fimH, tratT (serum resistance associated), fyuA (yersiniabactin receptor) and iutA (aerobactin receptor) have been investigated previously9,10. An important polysaccharide for biofilm formation is encoded by pgaABCD locus11. Biofilm production is a strategy to escape from harsh conditions and immune responses, hence play as reservoirs for drug-resistant systemic infections. Biofilm-producing A. baumannii has been isolated from several infectious origins such as pneumonia and devise-associated infections. Bacterial within biofilm can resist significantly more against antibiotics compared to planktonic mode of growth12. Hence, biofilm-mediated infections are in relapse more frequently13.
Therefore, there is an urgent need to enhance the effects of antimicrobials against pathogenic bacteria. In recent years, interest has enhanced towards application of nanoparticles as therapeutic regimens14,15,16,17,18,19,20,21. Silver nanoparticles (AgNPs), which have low toxicity in ecosystems and have high rate of surface capacity, can inhibit accumulation of biofilm materials responsible for evasion and protection22,23,24.
The aim of this study was to isolate A. baumannii from wound infections, determine their resistance and virulence profile, and assess the impact of AgNPs on the bacterial growth, virulence and biofilm-related gene expressions in the isolated strains.
Methods
Ethical statements for human/animal experiments
The study was approved by the Institutional Review Board and ethics committee of Assiut University, Egypt and Mustansiriyah University, Baghdad, Iraq. A written informed consent was obtained from participants. All experiments were performed in accordance with relevant guidelines and regulations.
Synthesis and characterization of silver nanoparticles (AgNPs)
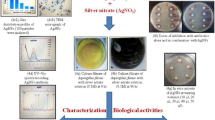
AgNPs (1000 µg/ml) were prepared via chemical reduction of silver nitrate in presence of polyvinylpyrrolidone (PVP) as a stabilizer according to25. The obtained samples were stirred, forming AgNPs. AgNPs formation was proved by Ultraviolet–visible spectral analysis. The absorbance spectra were determined using UV spectroscopy at a wavelength of 300–700 nm. The obtained NPs were characterized using transmission electron microscopy (TEM), X-ray diffraction methods (XRD) and Fourier transform infrared (FTIR) spectroscopy26,27. In TEM, a drop of NPs dispersion was placed on a carbon-coated copper grid and dried at room temperature and the micrograph of the preparation was taken.
In case of XRD, crystalline structure of AgNPs powder was identified using the X-ray diffractometer with CuKα radiation (λ = 1.5405 Å) in the 2θ range of 10°–80°. For FTIR, samples were prepared by KBr-discs technique and the spectra were recorded in the range of 400–4000 cm−1.
Bacterial isolates and identification of A. baumannii
Two hundred isolates of A. baumannii were identified from wound infections. Phenotypic identification included culture onto the MacConkey agar (OXOID, Basingstoke, England), Leed Acinetobacter agar (Hardy Diagnostics, Santa Maria, CA, USA), and CHROMagar Acinetobacter/MDR medium, and incubated under aerobic conditions at 37 C for 48 h. Gram staining, motility, oxidase, peroxidase, and oxidative-fermentation reactions were performed according to standard techniques. API 20NE bacterial identification system (Biomerieux, Marcy-l’Étoile, France) was used for species identification. Molecular identification of isolates was also conducted using PCR for detection of recA specific gene and bla-oxa51 gene as previously described28. The sequence of the primers are listed in supplemental Table 1.
Antimicrobial susceptibility patterns
Following Clinical and Laboratory Standards Institute guidelines29, bacterial cultures were plated onto the surface of Muller Hinton agar plates (HiMedia Laboratories, Mumbai, India) and susceptibility was measured by the disc-diffusion methods. The following antibiotics were tested: cephalothin (30 μg), cotrimoxazole (23.75/1.25 μg), nitrofurantoin (300 μg), trimethoprim (5 µg), ceftazidime (30 µg), tobramycin (10 µg), amikacin (30 µg), tetracycline (30 μg), gentamicin (10 µg), streptomycin (10 µg), erythromycin (15 µg), levofloxacin (5 µg), rifampicin (5 μg), chloramphenicol (30 μg), azithromycin (15 µg), imipenem (10 μg) and ciprofloxacin (5 μg). Staphylococcus aureus ATCC25923, A. baumannii ATCC 19,606 and Escherichia coli ATCC25922 were used as quality control organisms in the test29.
Screening for virulence-related genes
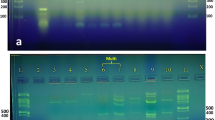
Virulence-related genes including afa/draBC, cnf1, cnf2, csgA, cvaC, fimH, fyuA, ibeA, iutA, kpsMT II, PAI, papC, PapG II, III, sfa/focDE and traT were screened using PCR. The primer sequence and product size are listed in Supplemental Table 1.
Screening for the antibacterial activity of AgNPs
The antimicrobial activity of the AgNPs was evaluated by the well-diffusion method. A. baumannii cultures (containing about 105 CFU/ml) were plated on the surface of Mueller Hinton agar plates (HiMedia Laboratories, India). Then, wells (5 mm is diameter) were made using a borer and 50 μg/ml of the AgNPs dispersions were added into the wells. Plates were then incubated at 37 °C for 24 h and the inhibition zone diameters were measured30.
Determination of the minimum inhibitory concentration (MIC) of AgNPs
MIC of the formulated AgNPs against clinically isolated A. baumannii was determined using the broth micro-dilution method according to the guidelines of the Clinical and Laboratory Standards Institute31. Briefly, AgNPs formulation was serially diluted in 100 μl Mueller–Hinton broth in wells of 96-well plate. Bacterial suspensions were added and plates were incubated for 24 h at 37 °C. MIC was determined visually by determining the wells with no visible bacterial growth.
Testing the biofilm strength using the microtiter plate method
The biofilm-forming abilities of the tested strains were carried out using the microtiter plate method as previously described32. Briefly, isolates were cultivated in Muller Hinton Broth (HiMedia Laboratories, India) containing 2% w/v sucrose (Sigma-Aldrich, USA) and incubated for 24 h at 37 °C in 96 well microtiter plates (1 × 105 CFU/ml, 200 μl /well). After incubation, wells were gently washed twice with PBS solution. Biofilm formed by adherent organisms was stained with crystal violet (0.2% w/v) and the excess dye was washed with distilled water. After drying, 95% ethanol was added to the wells and the optical density (OD) was measured at 570 nm using a microplate reader. Average OD values were determined for all tested isolates and the negative controls. Cut-off value was defined as the value of mean OD of the negative control plus 3X standard deviation of the negative control reads. Isolates were considered weak biofilm producers if they have a mean OD values ≤ two times the cut-off value, moderate biofilm producer if OD ≤ four times the cut-off value and strong biofilm producer if OD > four times the cut-off value.
In some experiments, the effect of AgNPs on biofilm formation was tested. To do that, the isolates were treated with 100 µl of 25 µg/ml AgNPs in BHI medium and analyzed as mentioned before. The ability of AgNPs to reduce biofilm formation was assessed by comparing the OD of the AgNPs treated wells versus control untreated wells. To test the ability of AgNPs to disperse the pre-formed biofilm layers, bacterial suspensions were allowed to grow overnight in the 96-well plates, then different aliquots of AgNPs were added to the overnight bacterial cultures. Plates were incubated for additional 24 h, and finally, the strength of the biofilms exposed to the AgNPs was re-evaluated. The experiment was performed in triplicate for each isolate.
Effect of AgNPs on microbial growth kinetics
Using the 96-well plate, we compared the effect of AgNPs on the growth of 3 representative A. baumannii strong, moderate and weak biofilm producers. Bacterial suspensions at OD595 in Muller Hinton broth were incubated at 37 °C in the presence of 5 μg/ml AgNPs (sub-MIC concentration) and OD595 was measured at 6, 24, 48 and 72 h using a microplate reader (Epoch, biotek, USA). Then, growth curves were plotted using the mean of three replicates for each strain ± SD33.
Quantitative detection of the effect of AgNPs virulence and biofilm-related genes using RT-PCR
The effect of AgNPs (25 µg/ml) on the expression of the virulence-related genes (kpsMII and afa/draBC genes) and biofilm-related genes (bap, OmpA, abaI, csuA/B, A1S_2091, A1S_1510, A1S_0690, A1S_0114) was evaluated using RT-PCR. RNA was extracted from bacteria treated with AgNPs using TRIzol reagent (Sigma-Aldrich, Switzerland) and untreated bacteria were used as controls. cDNA was synthesized subsequently by Superscript III kit (Invitrogen Inc., USA) according to the manufacturer protocol. PCR reactions were carried out using ePower SYBR Green PCR Master Mix (Applied Biosystems, USA) in a 7500 Sequence Detection System (CFX 96 biorad, USA). Samples were normalized to the expression of the 16S rRNA gene and the fold change in the expression of the target genes was calculated using 2−ΔΔCT method34. Expression level of each gene in AgNps treated cells was compared to the expression of untreated bacteria as a control.
In vitro infection model for analyzing the antibacterial activity of AgNPs
The ability of AgNPs to kill extracellular and intracellular A. baumannii was determined using a co-culture infection model of A. baumannii and HFF or Vero cells, as described previously35,36. Briefly, 1 × 105 HFF or Vero cells were seeded in a 24-well plate and incubated overnight at 37C with 5% CO2. Log phase bacterial suspension was prepared by incubating fresh colonies of A. baumannii overnight in Mueller–Hinton broth at 37C. Then, the bacterial suspension was diluted with broth and incubated for another 2 h at 37 C to reach the log phase of growth.
First, the extracellular killing activity was evaluated as previously described35. For this purpose, HFF or Vero cells were challenged with 2 × 105 CFU/mL of A. baumannii then AgNPs (25 µg/ml) were added in a total volume of 0.5 mL. Control samples treated with PBS were included. Kinetic studies were conducted at different time points to evaluate the extracellular killing activity at different contact times (5, 15, 30 and 60 min). Post-treatment samples were serially diluted in sterile PBS, plated on Mueller–Hinton agar plates, incubated for 24 h at 37C and CFUs were counted. The extracellular killing activity of AgNPs was evaluated by determining the percentage killing at the different time points, which was calculated by the following formula: (CFU of the control untreated samples − CFU of the AgNPs treated samples/CFU of the untreated control samples) × 100.
Second, the ability of AgNPs to kill the intracellular A. baumannii was carried out using the same co-culture infection model of A. baumannii and human fibroblast skin cell line HFF-1or Vero cells35. Briefly, cells were challenged with 1 × 108 CFU/mL log-phase A. baumannii and incubated for 2 h. Extracellular bacteria were removed by 2 h treatment with gentamycin then AgNPs (25 µg/ml ) or sterile PBS (plain control) were added and incubated for different time pointes (0, 2, 4, 8 and 16 h). Following incubation, cells were lysed by Triton X-100 (0.1%) and samples were diluted in PBS, plated on Mueller–Hinton agar plates, incubated for 24 h at 37C and CFUs were counted. The Intracellular killing activity of AgNPs was evaluated by determining the percentage killing at different time points relative to the control untreated cells as described previously. All experiments were performed in triplicate and the mean ± standard deviation was presented.
Analysis of in vitro cytotoxicity of AgNPs
The cytocompatibility of AgNPs was evaluated using two cell lines; the human fibroblast skin cell line HFF-1 and Vero cell line (kindly provided by VACSERA tissue culture laboratory, Dokki, Giza, Egypt) as previously described33. About 1 × 104 cells/200 ul were cultured in 96-well-plate at 37 C and 5% CO2 in DMEM (Gibco, Thermo Fisher Scientific, USA) supplemented with 10% Fetal Bovine Serum. After incubation of the cells overnight, AgNPs were added at different concentrations and incubated for 24 h. The viability of the fibroblast cells was measured by the MTT assay protocol (CellTiter 96 Non-Radioactive Cell Proliferation kit, Promega, USA) according to manufacturer instructions. Experiments were performed in triplicates and absorbance was measured at 570 nm using Epoch microplate reader (Bitek, USA).
Statistical analysis
All the obtained results were statistically analysed with the GraphPad Prism (version 6.0, GraphPad, San Diego, CA, USA) and the statistical software package for the Social Sciences software (SPSS, version 16). A p value < 0.05 was considered statistically significant.
Ethics approval and consent to participate
The study was approved by the Institutional Review Board and ethics committee of Assiut University, Egypt and Mustansiriyah University, Baghdad, Iraq. A written informed consent was obtained from participants. All experiments were performed in accordance with relevant guidelines and regulations.
Results
Characterization of synthesized AgNPs
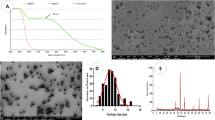
Shape and size of AgNPs, which were prepared via chemical reduction of Ag salts, were characterized using TEM. AgNPs had a spherical shape with particle size in the range of 10–50 nm (Fig. 1A). XRD measurement for a dried film from the concentrated suspension confirmed the crystalline structure of AgNPs (Fig. 1B). Figure 1C shows FTIR spectra for PVP and PVP/AgNPs. Spectrum of AgNPs stabilized with PVP was identical to that of pure PVP, showing PVP characteristic peaks at 1290 cm−1, 1670 cm−1 and 2900 cm−1, which correspond to CN bond, carbonyl group and C–H stretching, respectively37,38. Slight shifts in the spectrum of AgNPs indicate the formation of coordination bonds between oxygen or nitrogen of PVP with Ag atoms.
Characterization of synthesized silver nanoparticles (AgNPs). (A) The TEM image of AgNPs showed a spherical particles with size in nano range (10–50 nm, (B) X-ray diffraction analysis revealed the crystalline structure of AgNPs, (C) The FTIR analysis was done for AgNPs in the range of 400–4000 cm−1.
Antibacterial susceptibility testing
Antibacterial susceptibility of the isolated A. baumannii isolates were tested against 17 antibiotics. These isolates were 100% resistant to 12 antibacterial agents (cephalothin, cotrimoxazole, trimethoprim, ceftazidime, tobramycin, amikacin, tetracycline, streptomycin, erythromycin, levofloxacin, chloramphenicol and ciprofloxacin). Only azithromycin, nitrofurantoin, rifampicin, imipenem and gentamycin displayed limited degrees of resistance. The resistance profile of the 200 A. baumannii isolates and distribution of the virulence-related genes are shown in supplemental file 2.
Evaluation of bacterial virulence profile
Most of the A. baumannii isolates (n = 200) carried the fimH (n = 160, 80%), afa/draBC (n = 146, 73%), cnf1 (n = 112, 56%), csgA (n = 98, 49%) and cnf2 (n = 86, 43%), followed by ibeA (n = 82, 41%), cvaC (n = 80, 40%), iutA (n = 80, 40%), papC (n = 78, 39%), traT (n = 77, 38.5%), PAI (n = 62, 31%), fyuA (n = 48, 24%), kpsMII (n = 47, 23.5%), PapGII (n = 34, 17%) and papGIII (n = 8, 4%). The majority of A. baumannii (n = 165, 82.5%) produced strong biofilms, 22 (11%) isolates produced moderate and 13 (6.5%) isolates produced weak biofilms.
Evaluation of the antibacterial and anti-biofilm activities of AgNPs
Agar-well diffusion was used to test the inhibitory activity of AgNPs on bacterial growth. Exposure to 50 µg/mL AgNPs produced marked inhibition zones in all tested bacterial strains (mean = 16 mm and range = 6–27 mm). However, the biofilm formation seems to interfere with the formulation’s inhibitory activity. The inhibitory activity was more pronounced on weak biofilm producers, where AgNPs induced a mean inhibition zone diameter of 22 ± 5 mm, compared to 17 ± 4 mm and 10 ± 4 mm in moderate and strong A. baumannii biofilm producers (Fig. 2A).
Antibacterial activity of AgNPs on the isolated A. baumannii. (A) Inhibition zone diameters induced by the silver nanoparticles. (B) Growth kinetics of 3 representative A. baumannii from each group in the presence of AgNPs. The microbial growth was estimated by the optical density (OD595). (C) MIC values according to strength of biofilm formation. (D) Effect of silver nanoparticles on biofilm inhibition and biofilm dispersion. Untreated bacteria were used as control. Columns show the mean ± SD. *p < 0.05 indicates statistical significance as compared to control by Student’s t-test.
The growth curve of A. baumannii strains was determined in the presence of silver nanoparticles or its absence as a control. The growth rate was estimated by measuring the OD595 at different time points (6, 24, 48 and 72 h). Normal cells showed a marked increase in OD595 after 6 h and reached a stationary phase over the duration of 24–72 h. Generally, A. baumannii showed a reduced growth rate in the presence of AgNPs (at 24, 48 and 72 h), compared to untreated cells. Consistent with the resistance phenotype of the strong biofilm producers, the effect was more obvious with moderate and weak biofilm producing bacteria (Fig. 2B).
A similar effect was observed when MIC values of the formulated AgNPs against the isolated A. baumannii were determined. MIC values of the AgNPs against A. baumannii were in the range of 4–25 µg/mL. Lowest MIC values were observed for the weak biofilm producers, followed by the moderate and strong biofilm forming isolates (8 ± 4 µg/mL, 15 ± 4 µg/mL and 19 ± 6 µg/mL, respectively). Significant difference was observed between strong and moderate biofilm producers vs the weak group as shown in Fig. 2C.
Furthermore, exposure to AgNPs inhibited the strong, moderate and weak biofilm production significantly and biofilms reached 63 ± 15%, 58 ± 16% and 42 ± 9% of controls. AgNPs were also able to disperse the already formed strong, moderate and weak biofilm. Percentage of biofilms treated with AgNPs were 85 ± 12%, 77 ± 15% and 51 ± 11%, respectively (Fig. 2D).
Modulation of virulence and biofilm-related gene expression
The expression of kpsMII and afa/draBC genes were assessed with and without exposure to AgNPs. In presence of AgNPs, the expression of kpsMII and afa/draBC adhesin genes decreased by 4.6-folds (p = 0.001) and 3.4-folds (p = 0.001), respectively. RT-PCR analysis of total RNA isolated from strong biofilm producers treated with AgNPs showed that the transcription of bap, OmpA, and csuA/B was significantly decreased by 4.5-, 3.1- and 3.1-folds, respectively, when compared with the untrerated bacteria (Table 1). In contrast, the transcriptional expression of the A1S_2091, A1S_1510, A1S_0690 and A1S_0114 was not affected.
In vitro infection model for analyzing the antibacterial activity of AgNPs
An in vitro infection model was conducted to evaluate the antibacterial activity of the synthesized AgNPs against extracellular and intracellular A. baumannii. The bacterial killing efficacy increased with increasing the contact time of the treatment. The 25 μg/ml AgNPs displayed strong anti-Acinetobacter activity against extracellular bacteria, showing 10 ± 3% killing efficacy after 5 min and reaching complete killing after 60 min for Vero cells with a similar trend in case of HFF cell line (Fig. 3A). In addition, the killing efficacy against intracellular A. baumannii was investigated. There was an increase in killing efficacy with time; the percentage of killed intracellular bacteria was 29 ± 13% after 1 h, which increased to complete killing after 16 h in case of Vero cells and a similar trend was also observed in case of HFF cells (Fig. 3B).
Kinetic profile of the antibacterial activity of AgNPs against extracellular (A) and intracellular (B) A. baumannii. HFF or Vero cells were co-cultured with A. baumannii and AgNPs (25 μg/ml) were added. Extracellular killing was evaluated by plating the mixture on Mueller–Hinton agar plates and determination of the CFU, while intracellular killing was evaluated by lysing the cells with Triton X-100 following by culture. The percentage of killing caused by the AgNPs was calculated at the indicated time points relative to the control untreated cells. Experiment was performed in triplicate and the mean ± standard deviation is shown.
Evaluation of cytocompatibility of AgNPs on human foreskin fibroblast (HFF-1) and Vero cell lines
The biocompatibility of AgNPs was evaluated using the human foreskin fibroblast (HFF) and Vero cell lines. Cell viability test showed that exposure of the human fibroblasts to different AgNPs concentrations, starting from 0.25 to 25 µg/mL (the highest MIC value), did not affect the viability of the cells compared to untreated control cells, indicating that the nanoparticles are not harmful and biocompatible (Fig. 4).
Discussion
Severe infections associated with the multidrug-resistant bacteria, have increased significantly in the last decades39,40,41,42,43,44. The ability of bacteria to form biofilms is one of the main virulence factors that interferes with antibiotic activity and immune defense response mechanisms45,46,47,48. A. baumannii has developed many virulence factors and is responsible for severe life threatening infections49,50,51,52,53. The bacterium affects different sites, including wound infections, pneumonia, UTI. Biofilm formation abilities and various adhesins participate in the pathogenesis of their infections and resistance to antimicrobial drugs54. Herein, we observed that the majority of A. baumannii were resistant to most of antibiotics and could produce strong biofilms (82.5%). Interestingly, all the isolates which contained fimH and afa/draBC could produce strong biofilms. A. baumannii from wound infections mostly harbored the fimH, afa/draBC, cnf1, csgA and cnf255,56. It is worth considering that all of these strains were also MDR. The most effective antibiotic against MDR isolates included nitrofurantoin (12%) and imipenem (23%). Our results are in line with other studies which reported a high rate of biofilm formation among MDR-A. baumannii57.
In the last years new approaches have been developed to enhance the antimicrobial treatment such as the use antibiotic combinations, bacteriophage therapy, usage of antimicrobial NP-based formulations of old antimicrobial agent58,59,60. Recently, several reports have described the improved antimicrobial activity of AgNPs to be used for topical administration against multidrug-resistant bacteria61.
The anti-Acinetobater activity of the formulated AgNPs was shown by producing marked zones of inhibition and delay in microbial growth curves. Similar results were reported by previous studies using 14–27 µg/mL AgNPs62. The inhibitory activity of AgNPs on Gram-negative bacteria is more efficient than their activity on the Gram-positive bacteria63,64. AgNPs exerts potent antibacterial activity through different mechanisms. They have been reported to adsorb in the form of ionic silver (Ag+) onto the cytoplasmic membranes of Gram-negative bacteria which leads to destruction of the cell membrane, leakage of bacterial contents and death65. In addition, AgNPs also inhibit synthesis of bacterial cell wall66. They cause bacterial death through interaction with sulfur groups in essential proteins, generation of reactive oxygen species leading to bacterial death67. The high anti-Acinetobacter activity of the AgNPs could be attributed to the small size of the particles and the larger surface area, which allows more contact with the bacterial surface68.
Our results showed a potent ability of silver in inhibiting the strong biofilms produced by A. baumannii. Also, overnight incubation of sliver to well-formed biofilms lead to their partial removal. This allows the potential use of AgNPs as a biofilm-disrupting agent69. Consistent with our results, previous studies have shown efficient effect of AgNPs in inhibiting biofilms produced by different microorganisms70,71. Different investigators have evaluated the ability of silver nanoparticles to inhibit biofilm formation by A. baumannii, Pseudomonas aeruginosa (synergy with tobramycin), Staphylococcus aureus (such as synergy with vancomycin), Escherichia coli and Klebsiella pneumonia at similar concentrations23,72,73,74,75,76,77,78.
Importantly, this is the first report that shows that ability of AgNPs to inhibit the expression of different virulence-related genes (kpsMII and afa/draBC) and biofilm-related genes (bap, OmpA, and csuA/B), which adds to the known mechanism by which silver interferes with microbial growth. We observed that 25 µg/mL of AgNPs significantly decreased the expression of virulence and biofilm-related genes.
Studies have shown that the presence of Csu gene enables A. baumannii strains to attach and form strong biofilms through formation of pili79. When Csu is inactivated, the ability of pili production and subsequently the ability of bacteria to format biofilms is inhibited80. Finally, we concluded that decreasing the expression of bap, OmpA, and csuA/B by AgNPs is a main approach for decreasing biofilm activity of A. baumannii that will help in reducing antibiotic resistance mechanisms81. The limitations of our study included lack of an in vivo study, which we plan to study in the future work.
Conclusion
This study has enhanced our understanding of the characteristics of clinical A. baumannii isolates. The analysis of the isolates revealed that most isolates showed high level of resistance to antibiotics, carry bundle of virulence-related genes and different abilities to produce biofilms. Furthermore, treating the bacteria with AgNPs significantly interrupted bacterial growth and multiplication. The activity of AgNPs on A. baumannii growth kinetics, biofilm inhibition and dispersion was affected markedly by the strength of the produced biofilms. AgNPs downregulated the transcription level of important virulence and biofilm-related genes. Our findings provide an additional step towards understanding the mechanisms by which AgNPs interfere with the microbial spread and persistence.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Lee, C.-R. et al. Biology of Acinetobacter baumannii: pathogenesis, antibiotic resistance mechanisms, and prospective treatment options. Front. Cell. Infect. Microbiol. 7, 55 (2017).
Al-Kadmy, I. M. et al. Prevalence of genes involved in colistin resistance in Acinetobacter baumannii: first report from Iraq. Microb. Drug Resist. 26, 616–622 (2020).
Kareem, S. M., Al-Kadmy, I. M., Al-Kaabi, M. H., Aziz, S. N. & Ahmad, M. Acinetobacter baumannii virulence is enhanced by the combined presence of virulence factors genes phospholipase C (plcN) and elastase (lasB). Microb. Pathog. 110, 568–572 (2017).
El-Kazzaz, W. et al. Antibiogram, prevalence of OXA carbapenemase encoding genes, and RAPD-genotyping of multidrug-resistant Acinetobacter baumannii incriminated in hidden community-acquired infections. Antibiotics 9, 603 (2020).
Adewoyin, M. A. & Okoh, A. I. The natural environment as a reservoir of pathogenic and non-pathogenic Acinetobacter species. Rev. Environ. Health 33, 265–272 (2018).
Khazaal, S. S., Al-Kadmy, I. M. & Aziz, S. N. Mechanism of pathogenesis in multidrug resistant Acinetobacter baumannii isolated from intensive care unit. Gene Rep. 18, 100557 (2020).
Giles, S. K., Stroeher, U. H., Eijkelkamp, B. A. & Brown, M. H. Identification of genes essential for pellicle formation in Acinetobacter baumannii. BMC Microbiol. 15, 116 (2015).
Lee, J. C. et al. Adherence of Acinetobacter baumannii strains to human bronchial epithelial cells. Res. Microbiol. 157, 360–366 (2006).
Harding, C. M., Hennon, S. W. & Feldman, M. F. Uncovering the mechanisms of Acinetobacter baumannii virulence. Nat. Rev. Microbiol. 16, 91 (2018).
Kanaan, M. H. G., Al-Shadeedi, S. M., Al-Massody, A. J. & Ghasemian, A. Drug resistance and virulence traits of Acinetobacter baumannii from Turkey and chicken raw meat. Comp. Immunol. Microbiol. Infect. Dis. 70, 101451 (2020).
Fleming, I. D. et al. Modeling Acinetobacter baumannii wound infections: the critical role of iron. J. Trauma Acute Care Surg. 82, 557–565 (2017).
Qi, L. et al. Relationship between antibiotic resistance, biofilm formation, and biofilm-specific resistance in Acinetobacter baumannii. Front. Microbiol. 7, 483 (2016).
Pakharukova, N. et al. Structural basis for Acinetobacter baumannii biofilm formation. Proc. Natl. Acad. Sci. 115, 5558–5563 (2018).
Eid, A. M. et al. Endophytic Streptomyces laurentii mediated green synthesis of Ag-NPs with antibacterial and anticancer properties for developing functional textile fabric properties. Antibiotics 9, 641 (2020).
Abd Ellah, N. H., Gad, S. F., Muhammad, K., Batiha, E. G. & Hetta, H. F. Nanomedicine as a promising approach for diagnosis, treatment and prophylaxis against COVID-19. Nanomedicine 15, 2085–2102 (2020).
Wasef, L. et al. The potential ameliorative impacts of cerium oxide nanoparticles against fipronil-induced hepatic steatosis. Sci. Rep. 11, 1310. https://doi.org/10.1038/s41598-020-79479-5 (2021).
Hetta, H. F. et al. Modulation of rifampicin-induced hepatotoxicity using poly (lactic-co-glycolic acid) nanoparticles: a study on rat and cell culture models. Nanomedicine 15, 1375–1390 (2020).
Chaturvedi, V. K. et al. Pleurotus sajor-caju-mediated synthesis of silver and gold nanoparticles active against colon cancer cell lines: a new era of herbonanoceutics. Molecules 25, 3091 (2020).
Saleh, H. et al. Chemo-protective potential of cerium oxide nanoparticles against fipronil-induced oxidative stress, apoptosis, inflammation and reproductive dysfunction in male white albino rats. Molecules 25, 3479 (2020).
Abd Ellah, N. H., Tawfeek, H. M., John, J. & Hetta, H. F. Nanomedicine as a future therapeutic approach for Hepatitis C virus. Nanomedicine 14, 1471–1491 (2019).
Abd Ellah, N. H. et al. Metoclopramide nanoparticles modulate immune response in a diabetic rat model: association with regulatory T cells and proinflammatory cytokines. Int. J. Nanomed. 14, 2383 (2019).
Neethu, S., Midhun, S. J., Radhakrishnan, E. & Jyothis, M. Green synthesized silver nanoparticles by marine endophytic fungus Penicillium polonicum and its antibacterial efficacy against biofilm forming, multidrug-resistant Acinetobacter baumanii. Microb. Pathog. 116, 263–272 (2018).
Singh, R. et al. Antibacterial activities of bacteriagenic silver nanoparticles against nosocomial Acinetobacter baumannii. J. Nanosci. Nanotechnol. 18, 3806–3815 (2018).
Abo-Shama, U. H. et al. Synergistic and antagonistic effects of metal nanoparticles in combination with antibiotics against some reference strains of pathogenic microorganisms. Infect. Drug Resist. 13, 351 (2020).
Zielińska, A., Skwarek, E., Zaleska, A., Gazda, M. & Hupka, J. Preparation of silver nanoparticles with controlled particle size. Procedia Chem. 1, 1560–1566 (2009).
Jyoti, K., Baunthiyal, M. & Singh, A. Characterization of silver nanoparticles synthesized using Urtica dioica Linn. leaves and their synergistic effects with antibiotics. J. Radiat. Res. Appl. Sci. 9, 217–227 (2016).
Al-Dhabi, N. A., Ghilan, A.-K.M., Arasu, M. V. & Duraipandiyan, V. Green biosynthesis of silver nanoparticles produced from marine Streptomyces sp Al-Dhabi-89 and their potential applications against wound infection and drug resistant clinical pathogens. J. Photochem. Photobiol. B Biol. 189, 176–184 (2018).
Zander, E., Chmielarczyk, A., Heczko, P., Seifert, H. & Higgins, P. G. Conversion of OXA-66 into OXA-82 in clinical Acinetobacter baumannii isolates and association with altered carbapenem susceptibility. J. Antimicrob. Chemother. 68, 308–311 (2013).
CLSI. Performance Standards for Antimicrobial Susceptibility Testing. 30th ed. CLSI supplement M100. (Clinical and Laboratory Standards Institute, Wayne, PA, 2020).
Sancineto, L. et al. Diphenyl diselenide derivatives inhibit microbial biofilm formation involved in wound infection. BMC Microbiol 16, 220. https://doi.org/10.1186/s12866-016-0837-x (2016).
CLSI. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically; approved standard. 10th ed. M07-A11. (Clinical and Laboratory Standards Institute, Wayne, PA, 2018).
Elkhawaga, A. A., Hetta, H. F., Osman, N. S., Hosni, A. & El-Mokhtar, M. A. Emergence of Cronobacter sakazakii in cases of neonatal sepsis in upper Egypt: first report in North Africa. Front. Microbiol. 11, 215. https://doi.org/10.3389/fmicb.2020.00215 (2020).
Scutera, S. & Argenziano, M. Enhanced antimicrobial and antibiofilm effect of new colistin-loaded human albumin nanoparticles. Antibiotics 10, 2. https://doi.org/10.3390/antibiotics10010057 (2021).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Guzman, M., Dille, J. & Godet, S. Synthesis and antibacterial activity of silver nanoparticles against gram-positive and gram-negative bacteria. Nanomedicine 8, 37–45. https://doi.org/10.1016/j.nano.2011.05.007 (2012).
Noore, J., Noore, A. & Li, B. Cationic antimicrobial peptide LL-37 is effective against both extra- and intracellular Staphylococcus aureus. Antimicrob Agents Chemother 57, 1283–1290. https://doi.org/10.1128/aac.01650-12 (2013).
Bryaskova, R., Pencheva, D., Nikolov, S. & Kantardjiev, T. Synthesis and comparative study on the antimicrobial activity of hybrid materials based on silver nanoparticles (AgNps) stabilized by polyvinylpyrrolidone (PVP). J. Chem. Biol. 4, 185 (2011).
Malina, D., Sobczak-Kupiec, A., Wzorek, Z. & Kowalski, Z. Silver nanoparticles synthesis with different concentrations of polyvinylpyrrolidone. Dig. J. Nanomater. Biostruct. (DJNB) 7, 1527–1534 (2012).
Tacconelli, E. et al. ESCMID guidelines for the management of the infection control measures to reduce transmission of multidrug-resistant Gram-negative bacteria in hospitalized patients. Clin. Microbiol. Infect. 20(Suppl 1), 1–55. https://doi.org/10.1111/1469-0691.12427 (2014).
Makharita, R. R. et al. Antibiogram and genetic characterization of carbapenem-resistant gram-negative pathogens incriminated in healthcare-associated infections. Infect. Drug Resist. 13, 3991 (2020).
Algammal, A. M. et al. Methicillin-Resistant Staphylococcus aureus (MRSA): one health perspective approach to the bacterium epidemiology, virulence factors, antibiotic-resistance, and zoonotic impact. Infect. Drug Resist. 13, 3255 (2020).
Farhan, S. M., Ibrahim, R. A., Mahran, K. M., Hetta, H. F. & Abd El-Baky, R. M. Antimicrobial resistance pattern and molecular genetic distribution of metallo-β-lactamases producing Pseudomonas aeruginosa isolated from hospitals in Minia, Egypt. Infect. Drug Resist. 12, 2125 (2019).
El-Mokhtar, M. A. & Hetta, H. F. Ambulance vehicles as a source of multidrug-resistant infections: a multicenter study in Assiut City, Egypt. Infect. Drug Resist. 11, 587 (2018).
Ahmed, S. et al. Nosocomial vancomycin and methicillin resistant staphylococcal infections in intensive care units in Assiut University Hospitals. Egypt J. Med. Microbiol. 20 (2), 127–140 (2011).
El-Sayed Ahmed, M. A. E. et al. Colistin and its role in the Era of antibiotic resistance: an extended review (2000–2019). Emerg. Microbes Infect. 9, 868–885. https://doi.org/10.1080/22221751.2020.1754133 (2020).
Abd El-Baky, R. M., Sandle, T., John, J., Abuo-Rahma, G.E.-D.A. & Hetta, H. F. A novel mechanism of action of ketoconazole: inhibition of the NorA efflux pump system and biofilm formation in multidrug-resistant Staphylococcus aureus. Infect. Drug Resist. 12, 1703 (2019).
Algammal, A. M. et al. Virulence-determinants and antibiotic-resistance genes of MDR-E. coli isolated from secondary infections following FMD-outbreak in cattle. Sci. Rep. 10, 1–13 (2020).
Algammal, A. M. et al. Emerging MDR-Pseudomonas aeruginosa in fish commonly harbor opr L and tox A virulence genes and bla TEM, bla CTX-M, and tet A antibiotic-resistance genes. Sci. Rep. 10, 1–12 (2020).
Al Atrouni, A., Joly-Guillou, M.-L., Hamze, M. & Kempf, M. Reservoirs of non-baumannii Acinetobacter species. Front. Microbiol. 7, 49 (2016).
Andriamanantena, T. S. et al. Dissemination of multidrug resistant Acinetobacter baumannii in various hospitals of Antananarivo Madagascar. Ann. Clin. Microbiol. Antimicrob. 9, 17 (2010).
Antunes, L., Visca, P. & Towner, K. J. Acinetobacter baumannii: evolution of a global pathogen. Pathog. Dis. 71, 292–301 (2014).
Aydemir, H. et al. Risk factors and clinical responses of pneumonia patients with colistin-resistant Acinetobacter baumannii-calcoaceticus. World J. Clin. Cases 7, 1111 (2019).
Abd El-Baky, R. M., Farhan, S. M., Ibrahim, R. A., Mahran, K. M. & Hetta, H. F. Antimicrobial resistance pattern and molecular epidemiology of ESBL and MBL producing Acinetobacter baumannii isolated from hospitals in Minia, Egypt. Alex. J. Med. 56, 4–13 (2020).
Ghasemian, A., Mobarez, A. M., Peerayeh, S. N. & Abadi, A. B. The association of surface adhesin genes and the biofilm formation among Klebsiella oxytoca clinical isolates. New Microbes New Infect. 27, 36–39 (2019).
Tavakol, M., Momtaz, H., Mohajeri, P., Shokoohizadeh, L. & Tajbakhsh, E. Genotyping and distribution of putative virulence factors and antibiotic resistance genes of Acinetobacter baumannii strains isolated from raw meat. Antimicrob. Resist. Infect. Control 7, 120. https://doi.org/10.1186/s13756-018-0405-2 (2018).
Abdullah, R. M. & Ahmed, R. Z. T. Genotype detection of fimH gene of Acinetobacter baumannii isolated from different clinical cases. J. Infect. 3, 4 (2019).
Babapour, E., Haddadi, A., Mirnejad, R., Angaji, S.-A. & Amirmozafari, N. Biofilm formation in clinical isolates of nosocomial Acinetobacter baumannii and its relationship with multidrug resistance. Asian Pac. J. Trop. Biomed. 6, 528–533 (2016).
Mulani, M. S., Kamble, E. E., Kumkar, S. N., Tawre, M. S. & Pardesi, K. R. Emerging strategies to Combat ESKAPE pathogens in the era of antimicrobial resistance: a review. Front. Microbiol. 10, 539. https://doi.org/10.3389/fmicb.2019.00539 (2019).
Yeh, Y. C., Huang, T. H., Yang, S. C., Chen, C. C. & Fang, J. Y. Nano-based drug delivery or targeting to eradicate bacteria for infection mitigation: a review of recent advances. Front. Chem. 8, 286. https://doi.org/10.3389/fchem.2020.00286 (2020).
Abdelkader, A. et al. Ultrahigh antibacterial efficacy of meropenem-loaded chitosan nanoparticles in a septic animal model. Carbohydr. Polym. 174, 1041–1050. https://doi.org/10.1016/j.carbpol.2017.07.030 (2017).
Mekkawy, A. I. et al. In vitro and in vivo evaluation of biologically synthesized silver nanoparticles for topical applications: effect of surface coating and loading into hydrogels. Int. J. Nanomed. 12, 759–777. https://doi.org/10.2147/IJN.S124294 (2017).
Ahmad, N. et al. Biosynthesis of silver nanoparticles from Desmodium triflorum: a novel approach towards weed utilization. Biotechnol. Res. Int. 2011, 454090. https://doi.org/10.4061/2011/454090 (2011).
Ghosh, S. et al. Synthesis of silver nanoparticles using Dioscorea bulbifera tuber extract and evaluation of its synergistic potential in combination with antimicrobial agents. Int. J. Nanomed. 7, 483–496. https://doi.org/10.2147/IJN.S24793 (2012).
Shahverdi, A. R., Fakhimi, A., Shahverdi, H. R. & Minaian, S. Synthesis and effect of silver nanoparticles on the antibacterial activity of different antibiotics against Staphylococcus aureus and Escherichia coli. Nanomedicine 3, 168–171. https://doi.org/10.1016/j.nano.2007.02.001 (2007).
Deshpande, L. M. & Chopade, B. A. Plasmid mediated silver resistance in Acinetobacter baumannii. Biometals 7, 49–56. https://doi.org/10.1007/BF00205194 (1994).
Prabhu, S. & Poulose, E. K. Silver nanoparticles: mechanism of antimicrobial action, synthesis, medical applications, and toxicity effects. Int. Nano Lett. 2, 1–10 (2012).
Raffi, M. et al. Antibacterial characterization of silver nanoparticles against E. coli ATCC-15224. J. Mater. Sci. Technol. 24, 192–196 (2008).
Martínez-Castañon, G.-A., Nino-Martinez, N., Martinez-Gutierrez, F., Martinez-Mendoza, J. & Ruiz, F. Synthesis and antibacterial activity of silver nanoparticles with different sizes. J. Nanopart. Res. 10, 1343–1348 (2008).
Peulen, T. O. & Wilkinson, K. J. Diffusion of nanoparticles in a biofilm. Environ. Sci. Technol. 45, 3367–3373. https://doi.org/10.1021/es103450g (2011).
Hussain, Z., Thu, H. E., Sohail, M. & Khan, S. Hybridization and functionalization with biological macromolecules synergistically improve biomedical efficacy of silver nanoparticles: reconceptualization of in-vitro, in-vivo, and human clinical studies. J. Drug Deliv. Sci. Technol. 54, 101169 (2019).
Davis, S. C. et al. Preclinical evaluation of a novel silver gelling fiber dressing on Pseudomonas aeruginosa in a porcine wound infection model. Wound Repair Regen. 27, 360–365 (2019).
Habash, M. B. et al. Potentiation of tobramycin by silver nanoparticles against Pseudomonas aeruginosa biofilms. Antimicrob. Agents Chemother. 61, e00415-00417 (2017).
Thuptimdang, P., Limpiyakorn, T. & Khan, E. Dependence of toxicity of silver nanoparticles on Pseudomonas putida biofilm structure. Chemosphere 188, 199–207 (2017).
Singh, N., Rajwade, J. & Paknikar, K. Transcriptome analysis of silver nanoparticles treated Staphylococcus aureus reveals potential targets for biofilm inhibition. Colloids Surf. B 175, 487–497 (2019).
Hair, B. B., Conley, M. E., Wienclaw, T. M., Conley, M. J. & Berges, B. K. Synergistic activity of silver nanoparticles and vancomycin against a spectrum of Staphylococcus aureus biofilm types. J. Bacteriol. Mycol. 5(9), 1089 (2018).
Shafreen, R. B., Seema, S., Ahamed, A. P., Thajuddin, N. & Alharbi, S. A. Inhibitory effect of biosynthesized silver nanoparticles from extract of Nitzschia palea against curli-mediated biofilm of Escherichia coli. Appl. Biochem. Biotechnol. 183, 1351–1361 (2017).
Bala Subramaniyan, S., Senthilnathan, R., Arunachalam, J. & Anbazhagan, V. Revealing the significance of glycan binding property of butea monosperma seed lectin for enhancing the antibiofilm activity of silver nanoparticles against uropathogenic Escherichia coli. Bioconjug. Chem. 31, 139–148 (2020).
Farooq, U. et al. Rifampicin conjugated silver nanoparticles: a new arena for development of antibiofilm potential against methicillin resistant Staphylococcus aureus and Klebsiella pneumoniae. Int. J. Nanomed. 14, 3983 (2019).
Tomaras, A. P., Dorsey, C. W., Edelmann, R. E. & Actis, L. A. Attachment to and biofilm formation on abiotic surfaces by Acinetobacter baumannii: involvement of a novel chaperone-usher pili assembly system. Microbiology (Reading) 149, 3473–3484. https://doi.org/10.1099/mic.0.26541-0 (2003).
Gaddy, J. A. & Actis, L. A. Regulation of Acinetobacter baumannii biofilm formation. Future Microbiol. 4, 273–278. https://doi.org/10.2217/fmb.09.5 (2009).
Hosseini, A., Nejadsattari, T. & Zargar, M. In vitro anti-biofilm activity of curcumin nanoparticles in Acinetobacter baumannii: a culture-based and molecular approach. J. Arch. Clin. Infect. Dis. 14, e83263 (2019).
Acknowledgements
The authors would like to thank Mustansiriyah University (https://uomustansiriyah.edu.iq/)/Baghdad, Iraq for its support to complete this work.
Author information
Authors and Affiliations
Contributions
Conceptualization and design: H.F.H. and I.M.S.A. Methodology: H.F.H., I.M.S.A., S.S.K., S.A.,A.S., M.A.E., N.H.A., E.A.A., R.B.A., E.A.E., A.A.E., N.A.M. and A.M.A. Data analysis: H.F.H., I.M.S.A., S.S.K., S.A., A.S., M.A.E., N.H.A., E.A.A., R.B.A., E.A.E., G.E.B. and A.M.A. Manuscript writing and revision: H.F.H., I.M.S.A., S.S.K., S.A., A.S., M.A.E., N.H.A., E.A.A., R.B.A., E.A.E, G.E.B., A.A.E, N.A.M. and A.M.A. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hetta, H.F., Al-Kadmy, I.M.S., Khazaal, S.S. et al. Antibiofilm and antivirulence potential of silver nanoparticles against multidrug-resistant Acinetobacter baumannii. Sci Rep 11, 10751 (2021). https://doi.org/10.1038/s41598-021-90208-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-90208-4
This article is cited by
-
Ciprofloxacin- and levofloxacin-loaded nanoparticles efficiently suppressed fluoroquinolone resistance and biofilm formation in Acinetobacter baumannii
Scientific Reports (2024)
-
Utilization of Xanthan Gum-Silver Nitroprusside Nanoparticles for Prospective Advancements in Bacteriostasis and Wound Healing
Journal of Inorganic and Organometallic Polymers and Materials (2024)
-
Sequencing analysis and efficient biodiesel production by lipase from Pseudomonas aeruginosa
Molecular Biology Reports (2024)
-
Determination of the Cytotoxicity and Antibiofilm Potential Effect of Equisetum arvense Silver Nanoparticles
Applied Biochemistry and Biotechnology (2024)
-
Biogenic Ag2O nanoparticles with “Hoja Santa” (Piper auritum) extract: characterization and biological capabilities
BioMetals (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.