Abstract
Despite the advent of whole genome metagenomics, targeted approaches (such as 16S rRNA gene amplicon sequencing) continue to be valuable for determining the microbial composition of samples. Amplicon microbiome sequencing can be performed on clinical samples from a normally sterile site to determine the aetiology of an infection (usually single pathogen identification) or samples from more complex niches such as human mucosa or environmental samples where multiple microorganisms need to be identified. The methodologies are frequently applied to determine both presence of micro-organisms and their quantity or relative abundance. There are a number of technical steps required to perform microbial community profiling, many of which may have appreciable precision and bias that impacts final results. In order for these methods to be applied with the greatest accuracy, comparative studies across different laboratories are warranted. In this study we explored the impact of the bioinformatic approaches taken in different laboratories on microbiome assessment using 16S rRNA gene amplicon sequencing results. Data were generated from two mock microbial community samples which were amplified using primer sets spanning five different variable regions of 16S rRNA genes. The PCR-sequencing analysis included three technical repeats of the process to determine the repeatability of their methods. Thirteen laboratories participated in the study, and each analysed the same FASTQ files using their choice of pipeline. This study captured the methods used and the resulting sequence annotation and relative abundance output from bioinformatic analyses. Results were compared to digital PCR assessment of the absolute abundance of each target representing each organism in the mock microbial community samples and also to analyses of shotgun metagenome sequence data. This ring trial demonstrates that the choice of bioinformatic analysis pipeline alone can result in different estimations of the composition of the microbiome when using 16S rRNA gene amplicon sequencing data. The study observed differences in terms of both presence and abundance of organisms and provides a resource for ensuring reproducible pipeline development and application. The observed differences were especially prevalent when using custom databases and applying high stringency operational taxonomic unit (OTU) cut-off limits. In order to apply sequencing approaches with greater accuracy, the impact of different analytical steps needs to be clearly delineated and solutions devised to harmonise microbiome analysis results.
Similar content being viewed by others
Introduction
The analysis of the microbial composition of a sample can be performed using high-throughput sequencing methods by amplifying and sequencing selected regions of a gene (a metagenetic approach) or whole metagenomic DNA. Amplicon sequencing of regions such as the 16S ribosomal RNA gene (16S rRNA) continue to provide a simplified approach that is widely applied for bacterial identification and microbial community profiling1,2,3 even though there are newer approaches which involve sequencing of whole metagenomes. This is in part due to being an order of magnitude cheaper compared to a shotgun metagenomic approach while also being able to cope with the presence of a high background of contaminating (e.g. human) genomic DNA. In addition, targeted approaches require less computing power, and are well established, so have more complete, actively maintained and extensive databases and highly developed workflows. In this approach, highly conserved regions of the 16S rRNA gene are most often chosen as PCR primer binding sites to span variable region(s) that provide sequence clustering at the level of Operational Taxonomic Units (OTU). While this strategy is widely used, conserved regions of the 16S rRNA gene are not universally conserved across all microbial taxa4, and this sequence variability at primer-binding sites causes bias in microbial profiling experiments5,6. These biases can be further driven by the variable amplification efficiencies of different primer sets due to template-primer mismatches which will further distort the abundances of certain taxa when observing microbial community structure7. Conversely, shotgun metagenomic sequencing, which does not require sequence-dependent primer annealing, is thought to introduce less bias especially if it is prepared without PCR8.
There are many technical steps required in performing 16S rRNA gene amplicon sequencing experiments that can influence the results9. These include sampling (sampling site, method, sample transport and storage), extraction of nucleic acid material, choice of 16S rRNA primer, amplification, library preparation, sequencing and bioinformatic analysis pipeline. Previously, we used control materials (i.e., defined mock communities of mixed organism nucleic acids) to investigate how different steps in the process impact on the observed results10,11. Other studies have investigated how DNA extraction methods12,13,14, sample storage14, and variable 16S rRNA gene copy number can impact observed microbial community structures15. Hiergeist et al. used an inter-laboratory study to evaluate 16S rRNA gene amplicon sequencing of stool samples16. They concluded that investigators need to perform proficiency testing as all steps of the workflow can significantly affect the output of the procedure. However, the study did not evaluate the impact of the bioinformatic approach on error in isolation. Other studies have used simulated data sets to evaluate the bioinformatics approach17.
The use of control materials can enable the measurement of technical error. We previously investigated the application of a control material10 to investigate factors including choice of 16S rRNA gene amplicon strategy and impact of sequencing depth on whole genome sequencing data11. In that study, the control materials were characterised by absolute quantification of each organism contained in the mixture using a method orthogonal to sequencing called digital PCR (dPCR). dPCR is a highly accurate method for absolute quantification of DNA targets in samples18,19, and can be applied as a primary reference measurement procedure20. Previous studies have demonstrated that the choice of bioinformatic approach for analysis of microbiome sequence data can strongly impact the inferred microbial taxa composition21 and observed taxon relative abundance22. There are multiple decision points in bioinformatics pipelines, including quality filtering method, chimera removal method, 16S rRNA database choice, alignment method and assignment of taxonomy methodology. All of these steps can be performed by pipeline tools such as QIIME (Quantitative Insights Into Microbial Ecology)23, MG-RAST (Metagenomics Rapid Annotations using Subsystems Technology)24,25, MGnify26 and mothur27 or these tools can be combined with customised pipelines for the analysis of data. In addition to the broad choice of bioinformatic approaches, each tool has alterable parameters which affect the outcome. All of these options increase analysis variability, so it is important that a comprehensive description of data processing steps is included in the methodology. More stringent requirements in data reporting via a guideline system such as the Minimum Information about any (X) Sequence (MIxS) requirements for publishing will help28.
Inter-laboratory studies are a way to determine the reproducibility of methods and this is especially important in the field of multi-step advanced sequence analysis. In order to apply these methods with confidence, pipelines need to be validated through verification of inter-laboratory agreement using mock communities, as demonstrated in our previous study11. Few studies to date have investigated the reproducibility of results. Prior multi-centre studies have focused on next generation sequence based oncology tests29, detection of complex variants30 and whole genome sequence (WGS) based bacterial genotyping31. Other studies have looked into the benchmarking of 16S rRNA gene amplicon sequencing data32, investigating the tools for clustering of data in microbiome studies33 and making resources for benchmarking of data publicly available34. In addition to these studies, there are initiatives from the Global Microbial Identifier (GMI) initiative which has performed proficiency testing schemes for bacterial isolates35,36, and the European Molecular Genetics Quality Network (EMQN) which runs external quality assessment (EQA) schemes for germline and somatic mutation testing.
In this study, we used an inter-laboratory comparison to investigate the impact of the bioinformatic processing step on the prediction of the composition of control materials (i.e., genomic DNA from mock community samples, MCM2α and MCM2β). Raw sequence data, generated from PCR-next-generation amplicon sequencing of different 16S rRNA gene variable regions, which included technical repeats, were shared with multiple laboratories for bioinformatic pipeline comparison.
Methods
Preparation of Metagenomic Control Materials (MCM) MCM2α and MCM2β
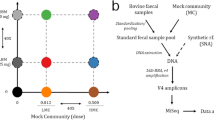
The metagenomic control material (MCM) 2α and β contained 15 different bacterial and one viral species, representing common human pathogens (Table 1). The two materials varied only by one species of Enterococcus but there were also subtle differences in the quantity of each organism in the mixture. This design was implemented in order to interrogate the ability of the sequencing approaches to identify these subtle differences.
The materials were prepared using genomic DNA (gDNA) sourced from ATCC (LGC Standards) and gDNA from E. coli O157, strain EDL 933 from IRMM (Institute for Reference Materials and Measurements). The concentration (ng/µL) of each gDNA preparation was determined by observing the mean value using triplicate measurements using a Qubit dsDNA BR Assay Kit (ThermoFisher) on the Qubit Fluorometer. The concentration of each gDNA preparation was also determined using specific assays for each of the organisms (Table 2) in the materials using dPCR using methods previously developed11. dPCR analyses were performed on a Bio-Rad QX200 droplet digital PCR system. This value was used when preparing the materials. The materials were diluted in TE pH 7.0 buffer and incubated at 4 °C for 4 h on a tube rotator. Aliquots of the materials were prepared in a final volume of 25 µL and stored at − 80 °C. Stability of the materials was determined as previously described11 using assays ctrA amplifying N. meningitidis and hin from MCM2α, mor and lytA from MCM2β. The material composition was determined using microfluidic dPCR as previously described11 using the assays in Table 2. The dMIQE (Minimum Information for publication of Quantitative Digital PCR Experiments) checklist is included in Additional File 1.
Amplicon sequencing
Amplicon sequencing was performed using two different primer sets; set one targeting variable regions 1 and 2 (V1–2) and set two targeting variable regions 4, 5 and 6 (V4–6) (Table 3). These priming strategies had been previously evaluated as strategy β, employing a combination of forward primers and one single primer in order to increase the specificity of the primer set for V1–2, and strategy γ which used degenerate bases to amplify the V4–6 regions11.
Amplicons were prepared of both materials, MCM2α and MCM2β, in triplicate using KAPA HiFi mastermix (Kapa Biosystems) to generate twelve samples for sequencing. Each reaction consisted of 1 × KAPA HiFi Hotstart ReadyMix, 0.3 µM of each primer and each template in a background of 50 ng human gDNA (Promega) in a final volume of 25 µL Nuclease Free Water (Ambion). The reactions were performed on a DNA Engine Tetrad 2 with the following cycling conditions: enzyme activation at 95 °C for 3 min, 30 cycles of denaturation at 98 °C for 15 s, annealing at 72 °C for 15 s and extension at 72 °C for 15 s, a final extension at 72 °C for 5 min and hold at 4 °C. Amplicons were visualised to determine product sizing using the Agilent DNA 1000 kit (Agilent Technologies) version 2.3 on the Agilent Bioanalyzer 2100 Instrument (Agilent Technologies).
Sequencing of the amplicons was performed. In total 12 libraries were prepared using Illumina TruSeq DNA PCR-Free Library Preparation kit processing according to the protocol for 350 bp input size for V1–2 and separately processing the V4–6 amplicons according to the 550 bp insert size (Revision D June 2015). Libraries were indexed using the Nextera indices (Illumina) and were pooled and quantified using KAPA SYBR FAST qPCR Master Mix (2X) Kit (Kapa Biosystems) according to manufacturer’s instructions. Libraries were visualised using a High Sensitivity DNA kit (Agilent Technologies) version 1.03 on the Bioanalyzer 2100. The libraries were diluted to 2 nM and after denaturing diluted to 10 pM. PhiX was spiked in at 5% using 20 pM library which should equate to around 7–10% of total cluster density. DNA sequencing was performed in a single run with an Illumina MiSeq platform using MiSeq V3 reagents (600 cycle chemistry), employing paired-end 300 base reads.
Shotgun metagenomic sequencing
Shotgun metagenomic sequencing was performed by LGC Genomics GmbH (Berlin, Germany). Libraries were prepared using 25 ng of DNA from three aliquots of MCM2α and MCM2β pools employing an Ovation Rapid DNA Library Preparation Kit (NuGEN). Genomic DNA was sheared to an average size of 400 bp using ultrasonication (Covaris S2 model). Libraries were sequenced on a NextSeq 500 sequencer (Illumina, San Diego, CA, USA) employing paired end 2 × 150 base sequencing. In addition to sequencing of the mixed samples, gDNA from each organism was sequenced as well using the same library preparation and sequencing protocols.
Data analysis
Paired end sequence reads generated from the shotgun metagenomic DNA libraries were trimmed to remove adapter sequences and bases with quality lower than Phred score 20. Sequences were then assembled using MEGAHIT37. Paired end reads were mapped back to the metagenomics assemblies and sequence bins were generated for each sample using metaBAT238. Taxonomy was assigned to individual sequence bins using kraken39 for each sample. The relative abundance of each organism was then calculated as the total metagenome length of each unique taxon in base pairs as a percentage of the total number of base pairs sequenced per sample.
Inter-laboratory study
In total 12 FASTQ data sets were generated from amplicon sequencing consisting of triplicate analysis of each material from sample to sequencing result. Laboratories were invited (Additional File 2) to participate in the analysis of these files using their standard bioinformatics pipelines for 16S rRNA gene amplicon sequence data. If they were in agreement, a link to the data and full description of the study hosted on the following URL: http://pathogenseq.lshtm.ac.uk/mcm.html was provided. They were asked to complete a submission form which collected pertinent information, including: (1) who processed the data, (2) the name of the laboratory, (3) results as biological observation matrices (BIOMs) based on annotation of OTU97 clusters, (4) a description of the analysis to include the command list for the whole bioinformatic process used to produce the results (Additional File 3).
Thirteen participating laboratories returned results. The results were collated in MS Excel 2010, and further analysis carried out using GraphPad Prism 6. The results from the 16S rRNA gene amplicon sequencing from each laboratory had to be normalised to be compared to the dPCR data to take into account that this operon can have different copy numbers depending on the genome. Data from each laboratory was compared to the dPCR analysis of the materials by calculating fold change. To assess agreement between different analytical methods, a cut-off of three-fold difference in relative abundance was applied, as described previously9.
The FASTQ files have been deposited in the European Nucleotide Archive (PRJEB34919).
Results
Amplicon sequencing
Mock community gDNA from two samples (MCM 2α and β) was PCR-amplified with 16S rRNA gene primers and sequenced using two different strategies. The mean number of reads per sample after sequencing were 845,651 (s.d. 246,478). The performance of these approaches have been previously determined11 where they were referred to as strategy β which used multiple forward primers to the same priming site amplifying a region spanning variable regions 1 and 2 and strategy γ which used degenerate bases in the primers to amplify a region spanning variable regions 4, 5 and 6.
The inter-laboratory study
Thirteen laboratories participated and submitted their results. The analysis steps applied are summarised in Table 4. The full commands used to run pipelines are available in Additional File 4. Most laboratories followed similar approaches except for one laboratory which used the online pipeline BIOiPLUG which implements the EzBioCloud database40. It is closed source and so could not be compared to the other pipelines in the same detail. Of the remaining twelve laboratories, eleven performed some form of filtering on the read data before clustering into OTUs and all laboratories merged overlaps between the read pairs. Sequence clustering, OTU assignment and taxonomic assignment differed between laboratories generally by those that used QIIME and mothur. Notable exceptions were the laboratories that used BLAST and USEARCH. There was an almost even split by those that assigned OTUs de novo and those using a closed reference method. Half of the participating laboratories applied further filtering steps to their sequence reads after OTU assignment. The different filtering steps are outlined in Table 4.
Results were reported as relative abundance with 97% OTU identity by the participating laboratories. As the taxonomic depth reported by the laboratories varied, it was decided that in order to compare the data the approach which would use the largest proportion of the data for cross-comparison was chosen; comparing the results using family level was therefore the optimal approach (Fig. 1A–D).
16S rRNA gene amplicon sequencing compared to dPCR results
The relative abundance of each taxonomic group was determined using 16S rRNA gene amplicon sequence data, and these relative abundance (RA) values were compared to the ‘true’ relative abundance, as determined through dPCR analysis of each taxon independently.
The dPCR approach in this study characterised the material according to single copy species specific genes whereas the 16S rRNA gene amplicon sequencing results have to be normalised based on the number of copies of this operon per genome. Differences are reported as fold-differences in RA between 16S amplicon results and dPCR results. In general, the results of the 16S rRNA amplicon sequencing approach differed by less than three-fold when compared to the dPCR value (Additional File 5). Differences greater than three-fold were described by 11/13 laboratories when reporting the abundance of Mycobacteriaceae (range 5.70–94.72 fold) and 10/13 laboratories when reporting the abundance of Pseudomonadaceae (range 3.26–466.34) in MCM2α v12. These families of organisms were the least abundant organisms in the sample; Pseudomonadaceae at 0.0015% and Mycobacteriaceae at 0.002% according to dPCR (Fig. 1). To determine if this observation was as a result of these families being the least abundant in the material, and therefore might represent a threshold which could be applied when analysing data for composition, we compared the results observed for the same variable regions but with the second mock DNA sample (MCM2β). This time, a different pattern was observed. The reported abundance of Neisseriaceae (the least abundant family of organisms, present at 0.005%) was very similar to the dPCR result, differing only by onefold on average except for laboratories 5 and 8 which differed by 61- and 11-fold respectively. Laboratory 5 used a custom database for taxonomic assignment, whereas the other laboratories used large public databases (Table 4). Laboratory 8 was the only participant to normalise their data using CSS, rather than taking a fraction from the total number of sequences within a sample. For most other laboratories, differences were less than three-fold. The differences comparing V1–2 amplicon sequencing to dPCR of MCM2β were greatest for the Mycobacteriaceae with 9/13 laboratories reporting fold differences greater than three.
When analysing the sequence data from V4–6 of MCM2α, all differences were less than three-fold compared to dPCR. For MCM2β differences in general were also less than three-fold except for large differences observed by laboratory 8 in terms of abundance of Mycobacteriaceae and Neisseriaceae (6.41 and 17.43 fold), Mycobacteriaceae reported by laboratory 6 (6.14 fold) in MCM2α (5, 33 and 49 fold respectively) and Neisseriaceae reported by laboratory 5 (73.29 fold).
In general, it was observed that the V4–6 region gave the most similar relative abundance compared to the dPCR results. It was observed for this primer set that many more OTUs were generated (as an example laboratory 13 reported an average 34 OTUs for the V1–2 region compared to 3240 OTUs for the V4–6 region). This could be due to the fact the V4–6 generates a much larger amplicon (~ 564 bp amplicon size for V4–6 v ~ 337 bp amplicon size for V1–2) and because of this could have a higher expected high error rate within those sequences which may lead to an over estimation of OTU richness.
To determine the influence that the different steps in the bioinformatic pipeline had on the results, results were clustered using NDMS (non-metric multidimensional scaling) plots to see if results group according to OTU assignment tool, OTU assignment database and taxa assignment database (Fig. 2A–C). Using ANOSIM we found statistically significant clustering of results due to the variable region amplified (p = 0.027) but not by laboratory, OTU assignment tool and taxa assignment database. The choice of OTU database or de novo methods also showed statistically significant clustering (p = 0.021). The choice of the Greengenes database for this step had a significant influence on the results, as observed by the data reported from laboratory 12 (Fig. 2A). This database choice however did not influence the results to this degree when laboratory 12 analysed V4–6 of both materials. In addition, results from laboratory 8 were more different compared to the other laboratories in the results reported for MCM2α and MCM2β perhaps due to normalisation strategy. It was observed that the results reported by laboratory 5 for MCM2β for V1–2 and V4–6 differed compared to the other laboratories due to using a custom database for taxa assignment rather than the choice of OTU database or assignment tool (Fig. 2C).
(A–C) NDMS (non-metric multidimensional scaling) plots to see if results from the 13 laboratories cluster according to OTU assignment tool (A), OTU assignment database (B) and taxa assignment database (C). OTU assignment database was included for laboratories that used closed reference OTU picking. They generally compared their sequences against a reference database of sequences that clustered the reads into OTUs based on sequence similarity. Later, many laboratories assigned taxonomic identifiers to each of these OTUs using a separate database which had sequence data (sometimes not clustered into OTUs) and which taxa that sequence originated from.
Some of the laboratories did not report the presence of some of families of organisms (Additional File 5). Enterococcaceae appeared to be the family most commonly unreported; by laboratory 4 in V1–2 of both materials, laboratory 5 in V4–6 of both materials, laboratory 13 in MCM2α V1–2 and MCM2β V4–6 and laboratory 2 in MCM2β V4–6. In addition, Mycobacteriaceae were not reported by laboratory 12 in V1–2 of both materials and Psuedomonadaceae were not reported by laboratory 5 in V4–6 of both materials.
Shotgun metagenomic sequencing compared to dPCR results
Shotgun metagenomic sequencing was performed on each of the gDNAs in addition to triplicate sequencing runs on the MCM2α and MCM2β (Additional File 6). The mean number of reads per sample after sequencing were 27,678,330 (s.d. 5,540,768). After metagenomic assembly of three MCM2α and three MCM2β replicates the average percentage of reads that mapped to the assembled contigs were 87%, 88% and 86% for MCM2α and 72%, 89% and 90% for MCM2β. The depth of sequencing of the assembled organisms in each sample differed with their relative abundance and ranged from a mean of 16 reads in low abundance organisms up to a mean of 900 reads in the high abundance samples. When compared to the dPCR data it was observed that the results were less than 1.8-fold different in terms of abundance of each organism between the two methods except for P. aeruginosa which was over-represented in both materials according to shotgun metagenomics sequencing (7.34 higher in MCM2α and 5.59 in MCM2β) (Additional File 7). Previous work analysing a different material had the same observation, but the underlying reason is not fully understood11. The source of P. aeruginosa was the same between the two studies but different batches were used. The shotgun sequencing method did not identify organisms present at the lowest abundance (S. pneumoniae and A. baumannii in MCM2α and S. enterica, A. baumannii and N. meningitidis in MCM2β). These organisms were of low abundance in the materials (≤ 0.003%) and so this could be explained by the sequencing coverage not being adequate to identify organisms at this level of abundance.
Additional taxa reported in the shotgun metagenomics data included Bradyrhizobiaceae and Bradyrhizobium, each present at 0.08%. These taxa were reported in all three units analysed. When each individual genomic DNA preparation was analysed by shotgun sequencing, this organism was present in the genomic preparation of hCMV (14% of total dataset), K. pneumoniae (12%) and S. aureus (0.03%). The Bradyrhizobiaceae family was also observed in the 16S rRNA analysis of V4–6 of MCM2α and MCM2β by laboratory 5 at 0.01% and 0.001% respectively. Laboratories 7, 9, 10 and 12 also reported this family but at a much lower abundance (< 0.00001%) and this was observed from both V4–6 and V1–2. For laboratories reporting Bradyrhizobiaceae presence it was not observed in all sequencing replicates unlike what was reported for the shotgun sequencing data. This family of bacteria has been observed as a contaminant in previous next generation sequencing studies41,42.
Discussion
The 16S rRNA gene amplicon sequencing procedure requires a multi-step process in order to determine the microbial composition of a sample. Bioinformatic analysis is crucial and involves multiple tools and steps as well as varying parameters. The bioinformatic tools used in this study represent relatively straightforward and simple to integrate for 16S rRNA gene amplicon analysis and are relatively less computationally intensive than when studying entire metagenomes. With the increasing diversity of bioinformatics tools and of the parameters which determine how each of these tools are used, it can be difficult to determine which tool will give the most accurate representation of the sample composition. In the face of this growing challenge, the use of control materials aids researchers in pipeline validation and choice of pipeline. These materials can also be used to evaluate current and future software versions. However, the choice of the most appropriate control material is itself an unresolved challenge and the ‘true’ composition of the reference community can be difficult to verify. Here we applied dPCR to determine the absolute and relative abundance of each organism in our control materials using species-specific assays that allowed for an accurate measurement of the composition of each of the materials and provides an absolute method for determining the composition of microbial standards.
In this study it was observed that in general there was good agreement when comparing the material composition according to the different 16S rRNA gene amplicon sequencing data results from the different laboratories to dPCR, with differences of less than three fold. Most large discrepancies were encountered with organisms present at low abundance. However, this result varied by analysis of variable regions under investigation. For example, analysis of 16S rRNA gene amplicon sequence data generated with the V4–6 primer set was more concordant with the dPCR analysis of the composition of the source materials than were amplicons covering the V1–2 region. However the method using this primer set as was observed for laboratory 5 was not concordant with the dPCR result. This laboratory used a custom database for taxonomic assignment, whereas the other laboratories used large public databases. It could be that the custom database was missing some key sequences that would lead to under-representation and over-representation of certain taxa.
Some of the laboratories did not identify families of organisms known to be present in the materials. This must have been because of the different methods applied. Laboratory 4, for example, applied a very stringent 0.1% relative abundance OTU cut-off which would remove many of the OTUs present, including the Enterococcaceae. It was observed that it was always the lowest abundance taxa that were found to be missing from the results when stringent filtering of the data was applied. The shotgun metagenomic sequencing approach also under-estimated the presence of this family. This could be due to the fact that this family is present at 0.6% in the MCM2α. Both 16S rRNA gene amplicon and metagenomic sequencing find it hard to differentiate between low abundance organisms and low-level contamination of bacterial DNA in each sample.
The inclusion of technical replicates allowed the reproducibility of the individual laboratory methods to be investigated. Overall they demonstrated good precision with coefficient of variation (CV) of < 10% in general (Table 5), apart from the results from laboratory 12 which, although used QIIME 1 like many others, performed no filtering of the OTU assigned reads which have previously been shown to be of poorer quality.
In this setting it was observed that the bioinformatic analysis of the V4–6 of 16S rRNA gene more closely resembled the determined composition using dPCR with V4–6 mean Bray–Curtis similarity to dPCR of 0.62 (95% CI 0.56–0.69) compared to 0.58 (95% CI 0.51–0.65) for V1–2. After omission of the results from laboratory 8 for MCM2α, all of the other pipelines were on average 0.84 fold different compared to dPCR (range 0.02—2.4). This is a very impressive result in terms of performance of the various 16S rRNA pipelines, all of which are composed of multiple steps, in analysing these materials.
Shotgun metagenomic sequencing results compared to dPCR results demonstrated for the most part good agreement, except for reporting of the abundance of P. aeruginosa where there were the largest differences, and also in the reporting of the lower abundance organisms in the materials. The precision of this approach was also determined and was demonstrated to be very good except for the lower abundance organisms (present at ≤ 0.01% abundance). It should be noted here that the comparison to the shotgun metagenomic sequencing approach was not from an inter-laboratory study, as was the case for the 16S rRNA data, but was from a single analysis workflow. So a further study is warranted to compare analysis tools for shotgun metagenomic data.
Conclusions
Determining the microbial composition of a sample can be undertaken by various high-throughput sequencing. Frequently this involves sequencing of variable regions of the 16S rRNA genes. We evaluated the multi-step process in assigning OTUs to a complex sample in terms of repeatability and reproducibility using control materials containing complex communities of microbes. The methods in general demonstrated high precision; however caution needs to be applied when drawing conclusions for microbiome data as variation between methods could significantly alter results.
In this study the reproducibility of the bioinformatics component was optimal when analysing the V4–6 regions which gave the most concordance with the dPCR analysis and the sequencing approach. While there was good agreement in general when comparing the different bioinformatics approaches, caution is required when using custom databases and applying high-stringency cut-offs that could misrepresent the relative abundance of organisms present. These findings are independent of software versions used and should be considered for current and future formats. This study provides compelling evidence of the importance of interrogating methods through the use of carefully designed control materials which could underpin future selection of the most appropriate methods to be applied to samples of interest.
Data availability
The datasets generated and/or analysed during the current study are available in the European Nucleotide Archive repository, https://www.ebi.ac.uk/ena/browser/view/PRJEB34919.
Abbreviations
- OTU:
-
Operational taxonomic unit
- dPCR:
-
Digital PCR
- QIIME:
-
Quantitative insights into microbial ecology
- MG-RAST:
-
Metagenomics rapid annotations using subsystems technology
- MIxS:
-
Minimum information about any (X) sequence
- WGS:
-
Whole genome sequencing
- GMI:
-
Global microbial identifier
- EMQN:
-
European molecular genetics quality network
- EQA:
-
External quality assessment
- MCM:
-
Metagenomic control material
- gDNA:
-
Genomic DNA
- NDMS:
-
Non-metric multidimensional scaling
References
Kitsios, G. D. et al. Respiratory microbiome profiling for etiologic diagnosis of pneumonia in mechanically ventilated patients. Front. Microbiol. 9(1413), 1 (2018).
Amelia, T. S. M., Amirul, A.-A.A., Saidin, J. & Bhubalan, K. Identification of cultivable bacteria from tropical marine sponges and their biotechnological potentials. Trop. Life Sci. Res. 29(2), 187–199 (2018).
Tringe, S. G. & Hugenholtz, P. A renaissance for the pioneering 16S rRNA gene. Curr. Opin. Microbiol. 11(5), 442–446 (2008).
Martinez-Porchas, M., Villalpando-Canchola, E., Ortiz Suarez, L. E. & Vargas-Albores, F. How conserved are the conserved 16S-rRNA regions?. Peer J. 5, e3036 (2017).
Klindworth, A. et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 41(1), e1 (2012).
Pinto, A. J. & Raskin, L. PCR biases distort bacterial and archaeal community structure in pyrosequencing datasets. PLoS ONE 7(8), e43093 (2012).
Frank, J. A. et al. Critical evaluation of two primers commonly used for amplification of bacterial 16S rRNA genes. Appl. Environ. Microbiol. 74(8), 2461–2470 (2008).
Ranjan, R., Rani, A., Metwally, A., McGee, H. S. & Perkins, D. L. Analysis of the microbiome: Advantages of whole genome shotgun versus 16S amplicon sequencing. Biochem. Biophys. Res. Commun. 469(4), 967–977 (2016).
Kim, D. et al. Optimizing methods and dodging pitfalls in microbiome research. Microbiome. 5(1), 52 (2017).
Huggett, J. F. et al. Considerations for the development and application of control materials to improve metagenomic microbial community profiling. Accred. Qual. Assur. 18(2), 77–83 (2013).
O’Sullivan, D. M. et al. Assessing the accuracy of quantitative molecular microbial profiling. Int. J. Mol. Sci. 15(11), 21476–21491 (2014).
Martin-Laurent, F. et al. DNA extraction from soils: Old bias for new microbial diversity analysis methods. Appl. Environ. Microbiol. 67(5), 2354–2359 (2001).
Velásquez-Mejía, E. P., de la Cuesta-Zuluaga, J. & Escobar, J. S. Impact of DNA extraction, sample dilution, and reagent contamination on 16S rRNA gene sequencing of human feces. Appl. Microbiol. Biotechnol. 102(1), 403–411 (2018).
Walker, A. W. et al. 16S rRNA gene-based profiling of the human infant gut microbiota is strongly influenced by sample processing and PCR primer choice. Microbiome 3, 26 (2015).
Louca, S., Doebeli, M. & Parfrey, L. W. Correcting for 16S rRNA gene copy numbers in microbiome surveys remains an unsolved problem. Microbiome. 6(1), 41 (2018).
Hiergeist, A., Reischl, U. & Gessner, A. Multicenter quality assessment of 16S ribosomal DNA-sequencing for microbiome analyses reveals high inter-center variability. Int. J. Med. Microbiol. 306(5), 334–342 (2016).
Almeida, A., Mitchell, A. L., Tarkowska, A. & Finn, R. D. Benchmarking taxonomic assignments based on 16S rRNA gene profiling of the microbiota from commonly sampled environments. GigaScience 7(5), 1 (2018).
Bhat, S., Herrmann, J., Armishaw, P., Corbisier, P. & Emslie, K. R. Single molecule detection in nanofluidic digital array enables accurate measurement of DNA copy number. Anal. Bioanal. Chem. 394(2), 457–467 (2009).
Sanders, R. et al. Evaluation of digital PCR for absolute DNA quantification. Anal. Chem. 83(17), 6474–6484 (2011).
Whale, A. S. et al. Assessment of digital PCR as a primary reference measurement procedure to support advances in precision medicine. Clin. Chem. 64(9), 1296–1307 (2018).
De Filippis, F., Parente, E., Zotta, T. & Ercolini, D. A comparison of bioinformatic approaches for 16S rRNA gene profiling of food bacterial microbiota. Int. J. Food Microbiol. 265, 9–17 (2018).
Allali, I. et al. A comparison of sequencing platforms and bioinformatics pipelines for compositional analysis of the gut microbiome. BMC Microbiol. 17(1), 194 (2017).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335 (2010).
Meyer, F. et al. The metagenomics RAST server: A public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinf. 9(1), 386 (2008).
Mitchell, A. L. et al. EBI Metagenomics in 2017: Enriching the analysis of microbial communities, from sequence reads to assemblies. Nucleic Acids Res. 46(D1), D726–D735 (2017).
Mitchell, A. L. et al. MGnify: The microbiome analysis resource in 2020. Nucleic Acids Res. 1, 1 (2019).
Schloss, P. D. et al. Introducing mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75(23), 7537–7541 (2009).
Yilmaz, P. et al. Minimum information about a marker gene sequence (MIMARKS) and minimum information about any (x) sequence (MIxS) specifications. Nat. Biotechnol. 29, 415 (2011).
Merker, J. D. et al. Proficiency testing of standardized samples shows very high interlaboratory agreement for clinical next-generation sequencing-based oncology assays. Arch. Pathol. Lab. Med. 143(4), 463–471 (2019).
Lincoln, S. E., Zook, J. M., Chowdhury, S., Mahamdallie, S., Fellowes, A., Klee, E. W., et al. An interlaboratory study of complex variant detection. BioRxiv. 218529 (2017).
Mellmann, A. et al. High interlaboratory reproducibility and accuracy of next-generation-sequencing-based bacterial genotyping in a ring trial. J. Clin. Microbiol. 55(3), 908–913 (2017).
Thorsen, J. et al. Large-scale benchmarking reveals false discoveries and count transformation sensitivity in 16S rRNA gene amplicon data analysis methods used in microbiome studies. Microbiome. 4(1), 62 (2016).
Zou, Q., Lin, G., Jiang, X., Liu, X. & Zeng, X. Sequence clustering in bioinformatics: An empirical study. Brief. Bioinform. 21(1), 1–10 (2018).
Bokulich, N. A. et al. mockrobiota: A public resource for microbiome bioinformatics benchmarking. mSystems 1(5), 1 (2016).
Wielinga, P. R. et al. Global Microbial Identifier. In Applied genomics of foodborrne pathogens (eds Deng, X. et al.) 13–31 (Springer International Publishing, 2017).
Hendriksen, R. S., Lukjancenko, O., Pedersen, S. K., Cisneros, J. L. B., Dynowski, L. D., Lund, O., et al. Report on the 2nd proficiency test trial for the Global Microbial Identifier (GMI) Initiative, Year 2016 (2017).
Li, D., Liu, C.-M., Luo, R., Sadakane, K. & Lam, T.-W. MEGAHIT: An ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31(10), 1674–1676 (2015).
Kang, D. D., Froula, J., Egan, R. & Wang, Z. MetaBAT, an efficient tool for accurately reconstructing single genomes from complex microbial communities. Peer J. 3, e1165 (2015).
Wood, D. E. & Salzberg, S. L. Kraken: Ultrafast metagenomic sequence classification using exact alignments. Genome Biol. 15(3), R46 (2014).
Yoon, S.-H. et al. Introducing EzBioCloud: A taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int. J. Syst. Evol. Microbiol. 67(5), 1613–1617 (2017).
Laurence, M., Hatzis, C. & Brash, D. E. Common contaminants in next-generation sequencing that hinder discovery of low-abundance microbes. PLoS ONE 9(5), e97876 (2014).
Salter, S. J. et al. Reagent and laboratory contamination can critically impact sequence-based microbiome analyses. BMC Biol. 12(1), 87 (2014).
Naserpour Farivar, T. et al. Development and evaluation of a Quadruplex Taq Man real-time PCR assay for simultaneous detection of clinical isolates of Enterococcus faecalis, Enterococcus faecium and their vanA and vanB genotypes. Iran J Microbiol. 6(5), 335–340 (2014).
Garcha, D. S. et al. Changes in prevalence and load of airway bacteria using quantitative PCR in stable and exacerbated COPD. Thorax. 67(12), 1075–1080 (2012).
Hartman, L. J. et al. Rapid Real-Time PCR Assays for Detection of Klebsiella pneumoniae with the rmpA or magA Genes Associated with the Hypermucoviscosity Phenotype: Screening of Nonhuman Primates. The Journal of Molecular Diagnostics. 11(5), 464–471 (2009).
Greiner, O., Day, P. J. R., Altwegg, M., Nadal, D. Quantitative Detection of Moraxella catarrhalis in Nasopharyngeal Secretions by Real-Time PCR. Journal of Clinical Microbiology. 41(4), 1386–1390 (2003).
Devonshire, A. S. et al. Highly Reproducible Absolute Quantification of Mycobacterium tuberculosis Complex by Digital PCR. Analytical chemistry. 87(7), 3706–3713 (2015).
Lee, D.-Y., Shannon, K., Beaudette, L. A. Detection of bacterial pathogens in municipal wastewater using an oligonucleotide microarray and real-time quantitative PCR. Journal of Microbiological Methods. 65(3), 453–467 (2006).
Francois, P. et al. Rapid Detection of Methicillin-Resistant Staphylococcus aureus Directly from Sterile or Nonsterile Clinical Samples by a New Molecular Assay. Journal of Clinical Microbiology. 41(1), 254–260 (2003).
Harris, K. A., Turner, P., Green, E. A., Hartley, J. C. Duplex Real-Time PCR Assay for Detection of Streptococcus pneumoniae in Clinical Samples and Determination of Penicillin Susceptibility. Journal of Clinical Microbiology. 46(8), 2751–2758 (2008).
Acknowledgements
DJS acknowledges the use of the University of Exeter’s Isca high-performance computing facility.
Funding
This work was supported by the UK National Measurement System and the European Metrology Programme for Innovation and Research (EMPIR) joint research project [HLT07] “AntiMicroResist” which has received funding from the EMPIR programme co-financed by the Participating States and the European Union’s Horizon 2020 research and innovation programme. TGC receives funding from the MRC UK (Grant no. MR/K000551/1, MR/M01360X/1, MR/N010469/1, MR/R020973/1) and BBSRC UK (BB/R013063/1). GLK and JOG received funding from the Quadram Institute Bioscience BBSRC Strategic Programme: Microbes in the Food Chain (project number BB/R012504/1) and its constituent projects BBS/E/F/000PR10348 and BBS/E/F/000PR10349 (JOG and GLK). The UK Antimicrobial Resistance Cross Council Initiative (no. MR/N013956/1, J.O.G. and G.L.K.). Part of the bioinformatics analysis was run on CLIMB-computing servers, an infrastructure supported by a grant from the UK Medical Research Council (no. MR/L015080/1). HD and RDF were supported by EMBL core funds. JRM at The Division of Digestive Disease at Imperial College London receives financial support from the National Institute of Health Research (NIHR) Imperial Biomedical Research Centre (BRC) based at Imperial College Healthcare NHS Trust and Imperial College London. This article is independent research funded by the NIHR BRC and NIHR Policy Research Programme (NIBSC Regulatory Science Research Unit), and the views expressed in this publication are those of the authors and not necessarily those of the NHS, NIHR, the Department of Health, ‘arms’ length bodies or other government departments.
Author information
Authors and Affiliations
Contributions
D.M.O.S., R.M.D., K.A.H. and J.F.H. conceived and designed the study. S.T. and A.S.W. prepared the materials. N.R. performed the sequencing. D.M.O.S., R.M.D., J.F.H. and J.M.G. drafted the manuscript. J.M.G. was involved in critical review of the manuscript. R.M.D., G.L., J.H., N.F., G.C.A.A., M.D.P., J.R.M., J.W., J.P., Y.M., H.D., R.D.F., G.L.K., J.O.G., E.R.J., H.W., E.L., D.J.S., E.D.B., J.E.P., T.G.C. and J.M.G. performed the analysis. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
O’Sullivan, D.M., Doyle, R.M., Temisak, S. et al. An inter-laboratory study to investigate the impact of the bioinformatics component on microbiome analysis using mock communities. Sci Rep 11, 10590 (2021). https://doi.org/10.1038/s41598-021-89881-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-89881-2
This article is cited by
-
Variability and bias in microbiome metagenomic sequencing: an interlaboratory study comparing experimental protocols
Scientific Reports (2024)
-
Multicenter evaluation of gut microbiome profiling by next-generation sequencing reveals major biases in partial-length metabarcoding approach
Scientific Reports (2023)
-
Metrological framework to support accurate, reliable, and reproducible nucleic acid measurements
Analytical and Bioanalytical Chemistry (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.