Abstract
The human AlkB homolog family (ALKBH) of proteins play a critical role in some types of cancer. However, the expression and function of the lysine demethylase ALKBH4 in cancer are poorly understood. Here, we examined the expression and function of ALKBH4 in non-small-cell lung cancer (NSCLC) and found that ALKBH4 was highly expressed in NSCLC, as compared to that in adjacent normal lung tissues. ALKBH4 knockdown significantly induced the downregulation of NSCLC cell proliferation via cell cycle arrest at the G1 phase of in vivo tumour growth. ALKBH4 knockdown downregulated E2F transcription factor 1 (E2F1) and its target gene expression in NSCLC cells. ALKBH4 and E2F1 expression was significantly correlated in NSCLC clinical specimens. Moreover, patients with high ALKBH4 expression showed a poor prognosis, suggesting that ALKBH4 plays a pivotal tumour-promoting role in NSCLC.
Similar content being viewed by others
Introduction
Escherichia coli AlkB is a 2-oxoglutarate and Fe (II)-dependent dioxygenase that repairs alkylated DNA/RNA nucleotides by catalysing oxidative demethylation1,2,3. In humans, the existence of nine AlkB homolog (ALKBH) family members (ALKBH1–ALKBH9) has been reported4,5. Some of the ALKBH molecules recognise various kinds of substrates, including methylated single- or double-stranded DNA or RNA bases6. Recently, the high expression of ALKBH molecules, such as ALKBH27, ALKBH38,9,10,11,12 and ALKBH813,14, has been reported to have tumour promoting activity in several cancers.
ALKBH4, one of the ALKBH family proteins, has been reported to be a lysine demethylase of actin in HEK293T cells15. Moreover, ALKBH4 has been reported to regulate gene expression or the chromatin state through association with transcriptional factors and chromatin regulation factors in HEK293 cells16. Recently, ALKBH4 has been reported to function as a suppressor of metastasis as it binds competitively to WD repeat-containing protein 5 (WDR5) in colon cancer17. However, the expression, substrate and biological function of ALKBH4 in other cancers are currently unknown.
Non-small-cell lung cancer (NSCLC) is one of the leading causes of cancer death worldwide. NSCLC accounts for approximately 85% of all lung cancers18. NSCLC is divided into adenocarcinoma, squamous cell carcinoma and large cell carcinoma. The most widely recognised genomic alterations in NSCLC include epidermal growth factor receptor (EGFR) mutations and echinoderm microtubule-associated protein-like 4-anaplastic lymphoma kinase (EML-ALK) fusion gene mutations. Although, molecular-targeted drugs for the proteins with such genetic mutations, improved the prognosis of NSCLC patients and their continuous usage often induced acquired resistance19. Moreover, some NSCLC cases do not present such mutations. Therefore, a novel therapeutic target candidate or approach is eagerly required.
The E2 promoter-binding factor (E2F) family of transcriptional factors consists of E2F1, E2F2, and E2F3. The activation and overexpression of members of the E2F family can contribute to oncogenic transformation of rodent embryonic fibroblasts and to tumourigenesis20. E2F1 plays an important role in cell cycle progression, senescence and apoptosis21,22,23. Abnormalities in E2F1 gene expression are found in several human cancers. Retrospective studies of several cancers, including NSCLC, indicate that an upregulated E2F1 expression is frequently associated with high-grade tumours and a poor patient survival prognosis24,25,26.
In this study, we show that the high expression of ALKBH4 in NSCLC specimens is associated with a poor prognosis. Additionally, we demonstrate the tumour-promoting role of ALKBH4 via its enzymatic activity in NSCLC cells.
Results
ALKBH4 is highly expressed in NSCLC tissues
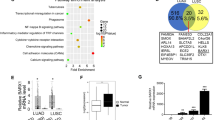
To obtain the ALKBH4 expression profile, we evaluated ALKBH4 expression in 89-matched pair NSCLC clinical specimens using qPCR. Compared to adjacent normal lung tissues, ALKBH4 expression was significantly higher in tumour tissues (Fig. 1A). Although there was no significant difference between either of the tumour stages, node stages (Fig. 1B,C), or pathological stages (Fig. 1D), ALKBH4 expression was higher in tumour tissues than in normal tissues, regardless of histologic subtypes (Fig. 1E,F). Interestingly, a higher ALKBH4 expression was detected in tumour tissues than in matched-pair normal tissues, regardless of the presence or absence of epidermal growth factor receptor (EGFR) gene mutations (Fig. 1G,H).
ALKBH4 is highly expressed in NSCLC. ALKBH4 expression levels were measured using qPCR and compared between normal and tumour tissues. (A) T classification (B), N classification (C), stages (D), histological types (E,F), and EGFR mutation (WT wild type, mt mutation) status (G,H) in NSCLC clinical specimens. Relative ALKBH4 expression normalised to ACTB is shown. Data are represented as mean ± S.D. The number of specimens examined is shown in parentheses. ***p < 0.001; ****p < 0.0001 for paired t-test (A) and Student’s t-test (B–H).
ALKBH4 knockdown reduced cell proliferation by inducing G1 phase arrest in NSCLC cells
To investigate the function of ALKBH4 in NSCLC cells, we first evaluated the expression of ALKBH4 in 22 NSCLC cell lines (adenocarcinoma: 15 cell lines; squamous cell carcinoma: 7 cell lines). Compared to the adjacent normal lung tissues obtained from postoperative tissues of NSCLC patients, all 22 NSCLC cell lines expressed high levels of ALKBH4 (Supplementary Fig. S1). A549 and II-18 cells with a high ALKBH4 expression were used for the subsequent knockdown and overexpression experiments. ALKBH4 knockdown using siRNAs significantly suppressed the proliferation of A549 (Fig. 2A) and II-18 cells (Fig. 2B). Conversely, ALKBH4 overexpression using pEB-ALKBH4 vector significantly promoted the proliferation of A549 cells (Fig. 2C). Also, in the case of NCI-H23 cells, the overexpression of ALKBH4 led to increased proliferation. However, overexpression of a catalytically inactive mutant ALKBH4 (H169A/D171A) had no significant effect on cell proliferation (Fig. 2D,E). Moreover, ALKBH4 knockdown inhibited the effect of overexpression of wild-type ALKBH4 on cell proliferation in NCI-H23 cells (Fig. 2D,E), suggesting that ALKBH4 promotes cell growth via its enzymatic activity. To clarify the tumour-promoting potential of ALKBH4 expression in vivo, A549 cells stably expressing control or ALKBH4 shRNA were constructed. We confirmed the decreased proliferation of A549 cells transfected with ALKBH4 shRNA, compared with those transfected with control shRNA, in vitro (Fig. 2F). These cells were used as an in vivo xenograft model. Suppression of tumour volume, as well as of tumour weight, was observed in mice xenografted with ALKBH4-knockdown A549 cells (shALKBH4), compared with the control cells (shControl) in Fig. 2G,H, respectively. These results suggest that ALKBH4 might function as a critical tumour promoter in NSCLC cells.
ALKBH4 knockdown suppressed NSCLC cells proliferation. Lysates of A549 (A) and II-18 (B) cells transfected with two ALKBH4 siRNAs (#1 and #2) or control siRNA, A549 cells transfected with pEB Multi-Neo-ALKBH4 or mock vector (C) were subjected to Western blot analysis with anti-ALKBH4 and anti-β-actin antibodies. Uncropped Western blot data are shown in Supplementary Fig. S9. Representative pictures of three independent experiments are shown (upper pictures). The cells which were transfected for 48 h were reseeded on a xCELLigence E-plate and their proliferation was detected using a xCELLigence DP system (lower panels). Degrees of cell proliferation expressed as cell index by the system are the means ± S.D. of three independent experiments (A–C). (D) Lysates of NCI-H23 cells co-transfected with pEB Multi-Neo-ALKBH4 wild-type (WT), pEB Multi-Neo-ALKBH4 mutant (mt), or mock vector and ALKBH4 siRNAs were subjected to western blot analysis with anti-ALKBH4 and anti-β-actin antibodies. Uncropped Western blot data are shown in Supplementary Fig. S9. Representative pictures of three independent experiments are shown (upper pictures). NCI-H23 cells were transfected for 48 h and reseeded in 96-well plates, and their proliferation was measured using the WST-8 assay (lower panels). The relative cell growth data on day 4 are presented in (E). The values are presented as the mean ± S.D. for each group. *p < 0.05; **p < 0.01 vs. wild-type ALKBH4 (one-way ANOVA with Bonferroni post-hoc tests). (F) A549 cells stably expressing ALKBH4 shRNA or control shRNA were subjected to Western blot analysis with anti-ALKBH4 and anti-β-actin antibodies. Uncropped Western blot data are shown in Supplementary Fig. S9. Representative pictures of three independent experiments are shown (upper pictures). Cell proliferation was detected using a xCELLigence DP system (lower panels). Degrees of cell proliferation expressed as cell index by the system are the means ± S.D. of three independent experiments (lower panels). (G) A549 cells stably expressing ALKBH4 shRNA (shALKBH4) and control shRNA (shControl), which are shown in (F), were subcutaneously injected into nude mice. The upper picture shows a xenograft tumour from mice inoculated with shControl- or shALKBH4-expressing A549 cells (shControl: n = 8; shALKBH4: n = 7). White scale bar, 1 cm. Tumour volume was calculated by measuring the tumour size every four days (lower panel). *p < 0.05 vs. control (Student’s t-test). (H) Tumour weight in shControl- and shALKBH4 xenografted mice. The values are presented as mean ± S.D. for each group. *p < 0.05 vs. shControl tumour (Student’s t-test).
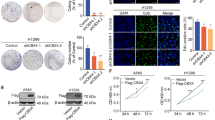
To clarify the function of ALKBH4 on cell proliferation, we conducted gene array analysis using the total RNA of A549 cells transfected with either ALKBH4 siRNA #1 or control siRNA. ALKBH4 knockdown affected the expression of 3572 genes (fold-change ≥|1.5|, 1561 upregulated genes, and 2011 downregulated genes) in A549 cells (Supplementary Table S2). Gene ontology enrichment analysis using several databases (Bioplanet, KEGG, and Reactome) revealed that these 2011 genes were mostly associated with cell cycle processes (Supplementary Table S3). Upregulated 1561 genes were associated with p53 signaling pathway and ECM-receptor interaction (Supplementary Table S4). To examine whether the suppression of cell proliferation induced by ALKBH4 knockdown was due to cell cycle arrest, we performed cell cycle analysis. ALKBH4 knockdown elevated the ratio of G1 phase cells and decreased the ratio of S phase cells in A549 (Fig. 3A) and II-18 cells (Fig. 3B). Moreover, cyclin-dependent kinase 2 (CDK2), cyclin-dependent kinase 4 (CDK4), and cyclin D3, the key proteins for the progression of the G1 phase, were markedly downregulated after ALKBH4 knockdown in A549 cells (Fig. 3C). Conversely, overexpression of wild-type ALKBH4 increased the ratio of S phase cells in NCI-H23 cells (Fig. 3D). Apoptosis analysis showed that ALKBH4 knockdown significantly induced apoptosis in A549 cells compared to control siRNA transfection (Fig. 3E). To further clarify the function of ALKBH4 in the cell cycle, we conducted G0-marker assays through β-galactosidase staining. ALKBH4 knockdown had no significant effect on senescence-associated β-galactosidase expression in A549 cells (Supplementary Fig. S2). These results suggest that highly expressed ALKBH4 promotes cell proliferation via regulation of the progression of the G1 phase in NSCLC cells.
ALKBH4 knockdown induced G1 phase arrest in NSCLC cells. A549 cells (A) or II-18 cells (B) were transfected with ALKBH4 siRNAs or control siRNA for 48 h and the cell cycle was analysed using a flow cytometer. Values are represented as means ± S.D. of three independent experiments. *p < 0.05; **p < 0.01 vs. control siRNA (One-way ANOVA with Bonferroni post-hoc tests). (C) Lysates of A549 cells transfected with ALKBH4 siRNAs or control siRNA were subjected to Western blot analysis with anti-CDK2, anti-CDK4, anti-Cyclin D3, and anti-β-actin antibodies. Uncropped Western blot data are shown in Supplementary Fig. S9. Representative pictures of three independent experiments are shown. (D) NCI-H23 cells were transfected with wild-type ALKBH4 or mutant ALKBH4 for 48 h and the cell cycle was analysed using a flow cytometer. Values are represented as means ± S.D. of three independent experiments. ***p < 0.001; ****p < 0.0001 (One-way ANOVA with Bonferroni post-hoc tests). (E) A549 cells were transfected with ALKBH4 siRNAs or control siRNA for 48 h and the apoptosis was analysed using a flow cytometer. Values are represented as means ± S.D. of three independent experiments. **p < 0.01 vs. control siRNA (One-way ANOVA with Bonferroni post-hoc tests).
Using Gene Expression Profiling Interactive Analysis (GEPIA)27, an interactive web server for analysing RNA sequencing expression data of tumours and normal samples from The Cancer Genome Atlas (TCGA) and The Genotype-Tissue Expression (GTEx) database (https://gtexportal.org/home/), we found that a broad range of cancer tissues, including lung adenocarcinoma and lung squamous cell carcinoma, express higher levels of ALKBH4 mRNA than each corresponding normal tissue (Supplementary Fig. S3A). To examine whether ALKBH4 specifically promotes cell proliferation in NSCLC cells or not, first, we examined the expression of ALKBH4 in some cancer cell lines (prostate, liver, lung, breast, renal, and pancreatic cancer cell lines). As shown in Supplementary Fig. S3B, ALKBH4 was highly expressed in many of the cancer cell lines examined. Then, we used MCF7 breast cancer cells to confirm the effect of ALKBH4 siRNAs on cell proliferation. ALKBH4 knockdown significantly suppressed the proliferation of MCF7 cells (Supplementary Fig. S3C,D). Moreover, ALKBH4 knockdown elevated the ratio of G1 phase cells and decreased the ratio of S phase cells in MCF7 cells (Supplementary Fig. S3E), suggesting that ALKBH4 plays an important role in the regulation of the proliferation of, not only NSCLC cells, but also other cancer cells.
ALKBH4 knockdown downregulated E2F1 signalling in NSCLC cells
Since ALKBH4 knockdown induced G1 phase arrest in NSCLC cells, we focused on E2F1 being known as a critical regulator of G1/S phase transition. ALKBH4 knockdown significantly reduced E2F1 expression in A549 and II-18 cells (Fig. 4A,B, respectively). Enrichment analysis using ChEA database28 showed that 215 E2F1-target genes were also downregulated via ALKBH4 knockdown (Supplementary Table S5). Since phospho-Ser/Thr phosphatase cdc25A (CDC25A), cyclin E1 (CCNE1), and Myb-like protein 2 (MYBL2) have been reported as tumour promoters29,30,31 and are known as cell cycle regulators in NSCLC, we focused on these genes. To verify the results of gene array analysis, the expression of E2F1-target genes was determined using qPCR. ALKBH4 knockdown significantly downregulated the target genes of E2F1 both in A549 and in II-18 cells (Fig. 4C–H).
ALKBH4 knockdown downregulated E2F1 and E2F1-target genes in A549 and II-18 cells. A549 cells (A,C–E) or II-18 cells (B,F–H) transfected with ALKBH4 siRNAs or control siRNA for 48 h were subjected to qPCR analysis of E2F1, CCNE1, MYBL2, and CDC25A expression. The relative expression values, which were normalised according to ACTB expression, are represented as mean ± S.D. of three independent experiments. *p < 0.05; ***p < 0.001; ****p < 0.0001 vs. control siRNA. (One-way ANOVA with Bonferroni post-hoc tests for (A,C–E), and Student’s t-test for (B,F–H)).
The relationship between ALKBH4 and E2F1 expression or those of E2F1-target genes was further confirmed in NSCLC specimens. Compared to the normal adjacent lung tissues, the expression of E2F1 (Fig. 5A) and of its target genes (Fig. 5B–D) was significantly high in tumour tissues. Importantly, there was a positive correlation (r = 0.46, p = 0.001) between ALKBH4 and E2F1 expression in NSCLC specimens (Fig. 5E). A significant positive correlation was also observed between ALKBH4 and each of the E2F1-target genes (Fig. 5F–H). The positive correlation between the expression of ALKBH4 and E2F1, as well as between that of ALKBH4 and of each of the E2F1-target genes was observed, regardless of the presence or absence of EGFR gene mutation (Supplementary Fig. S4A–H). In addition, only stage I, but not late stage (≥ stage II) NSCLC, showed a positive correlation between ALKBH4 and E2F1, or each of the E2F1-target genes (Supplementary Fig. S5A–H). Moreover, we recognised the positive correlation between the expression of ALKBH4 and E2F1, as well as between that of ALKBH4 and of each of the E2F1-target genes, regardless of the histologic subtypes, using TCGA database (Supplementary Fig. S6A,B) in NSCLC. These results suggested that ALKBH4 upregulates the expression of E2F1, followed by that of its target genes, in NSCLC.
The expression of ALKBH4 and E2F1of E2F1-target genes has a positive correlation in NSCLC clinical specimens. E2F1 (A) and E2F1-target gene ((B) CCNE1, (C) MYBL2, and (D) CDC25A) expression levels were measured using qPCR and were compared between normal and tumour tissues in NSCLC clinical specimens. Relative expression normalised to ACTB is shown. Data are represented as means ± S.D. ****p < 0.0001 for paired t-test. Correlation analysis was performed with the relative ALKBH4 expression and the relative E2F1 expression (E), CCNE1 expression (F), MYBL2 expression (G), and CDC25A expression (H) using 42-matched pairs of NSCLC samples. Pearson correlation analysis was conducted.
High ALKBH4 expression correlates with overall- and recurrence-free survival in NSCLC
Finally, to clarify the relationship between ALKBH4 expression and the prognosis of NSCLC patients, we performed immunohistochemical staining using anti-ALKBH4 antibody on NSCLC specimens. The immunohistochemical analysis showed that ALKBH4 was positive in 35 tumours and negative in 45 tumours (Fig. 6A). There was no significant intergroup heterogeneity regarding tumour size, presence or absence of pleural invasion, intrapulmonary metastasis, and nodal metastasis. During the follow up (median: 68.5 months; range: 2 to 110 months), 35 recurrences and 31 deaths occurred in a total of 80 patients. Kaplan–Meier survival curves are shown in Fig. 6B. The recurrence-free survival rate was significantly lower in the ALKBH4 positive group than in the ALKBH4 negative group (log rank test, p = 0.01). Likewise, the overall survival rate was significantly lower in the ALKBH4 positive group than in the ALKBH4 negative group (log rank test, p = 0.03). Additionally, we performed survival analysis using TCGA database. However, neither overall survival nor disease-free survival was significantly related to ALKBH4 mRNA expression levels in NSCLC (Supplementary Fig. S7).
Association of ALKBH4 expression levels with recurrence-free survival and overall survival. (A) Expression of ALKBH4 in NSCLC specimens was examined using immunohistochemical staining. Representative results are shown. Black scale bars, 200 μm. Association of ALKBH4 expression levels with recurrence-free survival (B) and overall survival (C). NSCLC tissue specimens stained with anti-ALKBH4 were divided into two groups, according to ALKBH4 expression: negative (45 samples) and positive (35 samples). Data was statistically analysed using Log-rank test.
Discussion
In the present study, we demonstrated that ALKBH4 is highly expressed in cell lines as well as in tumour tissues of NSCLC patients, and that it functions as a tumour promoter via its enzymatic activity. Knockdown of ALKBH4 downregulated the expression of E2F1, a critical regulator of the G1/S phase transition in NSCLC cells. E2F1 expression was positively correlated with the expression of ALKBH4 in NSCLC clinical samples. Moreover, the expression of E2F1-target genes (CDC25A, CCNE1 and MYBL2) was positively correlated with the expression of ALKBH4 in NSCLC clinical samples. ALKBH4 knockdown downregulated the expression of CDK2 and cyclin D3 encoding gene in A549 cells. Since CDK2 and CCND3 (cyclin D3) have also been reported as E2F1-target genes32,33,34,35, the downregulation of CDK2 and CCND3 might be due to the regulation of E2F1 expression by ALKBH4. It is well known that E2F1 is overexpressed in several cancers, including NSCLC36,37,38 and that an aberrant E2F1 expression is correlated with a lower overall survival of NSCLC patients26. Taken together, the upregulated ALKBH4 expression plays a pivotal NSCLC-promoting role, leading to a poor prognosis in NSCLC patients.
Although we have shown that ALKBH4 overexpression promotes cell growth via its enzymatic activity (Fig. 2D,E), the effect of ALKBH4 enzymatic activity on downstream effector molecules such as E2F1 and E2F1-target genes remains unclear and requires further investigation.
Recently, it was reported that ALKBH4 functions as a demethylase for N6-adenosine modification in DNA39, and the C. elegans ALKBH4 ortholog (denoted NMAD-1) also has such a demethylase function40. Abrogation of ALKBH4 (or NMAD-1) function appears to have strong effects on various phenomena related to cell cycle progression, such as cytokinesis, DNA replication, and meiosis15,41,42. As shown in Supplementary Table S3, ALKBH4 knockdown also affected DNA replication and meiosis-related genes in A549 cells. Moreover, the enzymatic domain of ALKBH4 was critical for the upregulation of cell proliferation in NSCLC cells, suggesting that ALKBH4 may function as a tumour promoter by targeting N6-adenosine modification in NSCLC cells.
Although few publications suggest that gene expression is mostly regulated at the mRNA level43,44, many studies have reported a discrepancy between mRNA and protein levels45,46. Protein levels more closely reflect the cancer phenotype because these proteins execute major intracellular molecular functions. Therefore, we believe that survival analysis using IHC data reflects the true nature of ALKBH4 in NSCLC.
A high expression of ALKBH4 is correlated with an overall- and recurrence-free survival in NSCLC. On the contrary, Supplementary Fig. S4 shows a positive correlation between the expression of ALKBH4 and E2F1, as well as between that of ALKBH4 and E2F1-target genes, which were only observed in early stage NSCLC tumour tissues, but not in those of late stage NSCLC. Among the Polo-like kinase (PLK) family, PLK1 and PLK4 are highly expressed in NSCLC47,48 and promote metastasis via epithelial–mesenchymal transition (EMT) induction49,50. Since ALKBH4 knockdown downregulated the expression of PLK1 and PLK4 (Supplementary Table S2), ALKBH4 may function as a tumour promoter by accelerating cancer cell proliferation via upregulation of E2F1 signalling in early stage, and by promoting metastasis via upregulation of PLK signalling in late-stage NSCLC.
ALKBH4 knockdown had no significant effect on the population of G0 phase cells in A549 cells. Although our gene array data showed that upstream regulators of E2F1 (Rb1 and cyclin D1) expression remained unchanged upon ALKBH4 knockdown, ubiquitin C-terminal hydrolase L5 (UCHL5, also known as UCH37, ubiquitin C-terminal hydrolase 37), which has been reported as a deubiquitinating enzyme of E2F151, was downregulated upon ALKBH4 knockdown (Supplementary Table S2, Fold-change = − 2.1). UCHL5 activates the E2F1 transcriptional activity by decreasing Lys-63-linked ubiquitination of E2F1. E2F1 can activate gene transcription of the E2F family (E2F1, E2F2, and E2F3), which in turn induces positive feedback on E2F1 gene transcription52,53,54,55. Moreover, ALKBH4 knockdown downregulated the expression of E2F2 (Supplementary Table S2). In addition, UCHL5 knockdown induces apoptosis in A549 cells56. Therefore, we believe that ALKBH4 knockdown-induced E2F1 reduction is due to the downregulation of UCHL5 expression in NSCLC cells.
We found that ALKBH4 functions as a tumour promoter in NSCLC. However, Shen et al. recently reported that ALKBH4 expression is decreased in colon cancer tissues, compared to adjacent normal tissues, and functions as a tumour suppressor by decreasing H3K4me3 levels by competitively binding to methyltransferase WDR517. WDR5 is overexpressed in colon cancer and the depletion of WDR5 reduces cell viability of colon cancer cell lines57. On the contrary, an overexpression of WDR5 was reported to induce G1 arrest in A549 cells, independently of the H3K4me3 status, and WDR5 was suggested to possibly have different functions in different cancers58. Therefore, although the underlying mechanism should be analysed, the tumour promoting role of ALKBH4 in NSCLC might partly rely on WDR5.
Cancer cells adapt their metabolism to promote tumour growth. One important metabolic feature of cancer cells is the Warburg effect, which leads to high rates of glucose utilisation and lactate production59. It has been reported that E2F1 promotes this effect by enhancing glycolysis and repressing glucose oxidation in the mitochondria60,61. Moreover, E2F1-mediated repression of oxidative metabolism results in the self-renewal of cancer stem cells62, suggesting that ALKBH4 may confer the Warburg effect through an increased expression of E2F1, leading to efficient recurrence in NSCLC patients.
The specific types of mutations that confer drug sensitivity to EGFR-targeted drugs are present in the tyrosine kinase domain of the EGFR gene, corresponding to exons 19 and 21 with 5–20% of overall incidence63. In contrast, upregulated ALKBH4 expression was independent of the mutation status of the EGFR gene in NSCLC specimens (Fig. 1G,H). Moreover, an increased ALKBH4 expression was shown in broad types of cancer (Supplementary Fig. S3A). We showed that ALKBH4 knockdown induced the downregulation of cell proliferation and G1 arrest in breast cancer cell line MCF-7 cells, as well as in NSCLC cells (Supplementary Fig. S3D,E, respectively). Therefore, we propose that ALKBH4 may be an important molecular target for, not only NSCLC, but also for a wide range of cancers.
Materials and methods
Clinical specimens
Specimens of NSCLC tissues and adjacent non-cancerous tissues were obtained from the patients who had undergone primary curative resection of a lung tumour at Kagoshima University Hospital (Japan) as described before64. All enrolled patients were diagnosed via pathological examination staged by specialised oncologists via the 7th edition of the International Association for the Study of Lung Cancer TNM Classification. Written informed consent was obtained from each of the patients and the study was approved by the Ethical Committee of Kagoshima University Hospital (registration No. 351). Experiments using clinical specimens were approved by the institutional review boards of the Graduate School of Pharmaceutical Sciences, Osaka University. All methods were carried out in accordance with relevant guidelines and regulations. Clinical and histopathological data related to the clinical specimens are presented in Supplementary Table S1.
Antibodies
Monoclonal anti-ALKBH4 antibody (ab195379) was purchased from Abcam (Cambridge, UK) and used for Western blotting. Polyclonal anti-ALKBH4 antibody (NBP2-14737) was purchased from Novus Biologicals (Littleton, Colorado, USA) and used for IHC and ICC experiments. Monoclonal anti-CDK2 antibody was purchased from BD Transduction Laboratories (San Jose, CA, USA). Monoclonal anti-β-actin antibody was purchased from Sigma (St. Louis, Missouri, United States). Monoclonal anti-CDK4 and anti-cyclin D3 antibodies and anti-rabbit IgG, HRP-linked antibody (#7074) and anti-mouse IgG, HRP-linked antibody (#7076) were purchased from Cell signalling technology (Danvers, MA 01923, USA).
Cell culture
Twenty-three human lung cancer cell lines (Calu-3, NCI-H2228, NCI-H1792, NCI-H1437, NCI-H1650, NCI-H1755, NCI-H1838, RERF-LC-KJ, NCI-H23, NCI-H1975, PC-9, LC-2/ad, HLC-1, A549, II-18, RERF-LC-AI, NCI-H226, Sq-1, NCI-H520, LK-2, EBC-1, and LC-1F cells) obtained from the American Type Culture Collection (ATCC), were cultured in RPMI1640 medium (Wako) supplemented with 10% foetal bovine serum, 100 U/mL penicillin G, and 0.1 μg/mL streptomycin.
Western blotting
Western blotting was conducted as described before65. Protein samples were separated on a 7.5–15% sodium dodecyl sulphate (SDS)-polyacrylamide gel electrophoresis (PAGE) gel, and then transferred to a polyvinylidene difluoride (PVDF) membrane by using the Bio-Rad semidry transfer system (1 h, 12 V) (Hercules, CA, USA). Immunoreactive proteins were visualised by treatment with a detection reagent (ECL Prime western blotting detection reagent, GE Healthcare), using the antibodies described above, in an ImageQuant LAS 4000 mini system (GE Healthcare). Densitometric analysis was performed using the National Institute of Health (NIH) Image J software.
Construction of the ALKBH4 expression vector
PCR-amplified wild-type ALKBH4 was cloned into pEB Multi-Neo vector (Wako). The primer sequences used for the gene amplification were as follows: forward primer, 5′-GAATTCGCGGAAATGGCTGGGAGGGG-3′ and reverse primer, 5′-GGGACATAGTAGATTACAGGTGG-3′. Inserted ALKBH4 sequence was confirmed via sequencing. Catalytic domain-mutated ALKBH4 (H169A/D171A) was purchased from GenScript (Piscataway, NJ, USA).
Cell cycle analysis
A549 or II-18 cells were transfected with the ALKBH4 siRNA or control siRNA. NCI-H23 cells were transfected with the pEB Multi-Neo-ALKBH4 vector (wild-type or mutant). After incubation for 48 h, cells were fixed with 70% ethanol and were washed with phosphate buffered saline (PBS). Cells were suspended in a hypotonic solution containing 5 mg/mL propidium iodide (PI). PI-stained samples (1 × 105 cells) were analysed for fluorescence using an FACS Calibur (Becton–Dickinson, Franklin Lakes, New Jersey, USA).
Apoptosis analysis
A549 cells transfected with ALKBH4 or control siRNAs were cultured for 48 h and stained with ethidium homodimer III and fluorescein isothiocyanate-conjugated Annexin V (Apoptotic & Necrotic Cell Detection Kits, TaKaRa) according to the manufacturer's protocol. Flow cytometric analysis was conducted using MACSQuant X (MiltenyiBiotec, BergischGladbach, Germany).
G0-marker analysis
A549 cells transfected with ALKBH4 or control siRNAs were cultured for 72 h, and G0-marker assay was conducted using senescence-associated β-galactosidase (Senescence Detection Kit, BioVision, Milpitas, CA, USA) according to the manufacturer’s protocol. β-galactosidase-positive cells were counted in randomly selected 25 fields.
Cell proliferation assay
Cell proliferation was examined using the xCELLigenceReal Time Cell Analyzer Dual Purpose (RTCA DP) system (Roche, Basel, Switzerland). A549 or II-18 cells were transfected with the ALKBH4 siRNA or control siRNA. After 24 h of incubation, cells were reseeded on an E-plate 16 (A549 cells: 1 × 103 cells/well; II-18 cells: 3 × 103 cells/well) and incubated for the indicated times. Water-soluble tetrazolium salt-8 (WST-8) reagent (Dojindo) was used for the cell proliferation assay (Fig. 2D), which was conducted as described previously66. NCI-H23 cells transfected with the pEB Multi-Neo-ALKBH4 vector (wild-type or mutant) and ALKBH4 siRNAs were reseeded in a 96-well plate and incubated for the indicated times. After incubation for 2 h with WST-8 reagent at 37 °C and 5% CO2, the optical density was determined at 450/630 nm (Ex/Em).
siRNA and DNA transfection
siRNA duplexes were used to downregulate ALKBH4 mRNA (ALKBH4 stealth siRNA #1: ACAUACCGUUUCAUUUACUGCUCCG; ALKBH4 stealth siRNA #2: ACAGAGGAGUCUGACUUUGAGGGCU) and stealth RNAi siRNA negative control low GC were purchased from Life technologies (Carlsbad, CA, USA). For all siRNA transfection studies, A549 cells (4 × 104 cells/well) or II-18 (5 × 104 cells/well) were transfected on a 12-well plate using Lipofectamine RNAiMAX (Life Technologies). For DNA transfection, A549 cells (4 × 104 cells/well) seeded on a 24-well plate were transfected with pEB Multi-Neo vector or pEB Multi-Neo-ALKBH4 vector using Lipofectamine 3000 and P3000 reagent (Life Technologies).
Co-transfection of siRNA and expression vector
NCI-H23 cells (15 × 104 cells/well) seeded on a 12-well plate were transfected with pEB Multi-Neo-ALKBH4 vector (wild-type or mutant) and ALKBH4 siRNAs using Lipofectamine 2000 (Life Technologies).
Quantitative PCR (qPCR)
Total RNA was isolated by using RNeasy Plus Mini Kit (QIAGEN, Hilden, Germany). Prime Script RT reagent Kit (Takara, Mountain View, USA) was used to prepare cDNA from 500 ng of total RNA. The Light cycler 96 plate (Roche) was used for qPCR analysis. The thermal cycling conditions used were as follows: an initial step at 95 °C for 10 s and 40 cycles of 95 °C for 5 s, and 60 °C for 20 s for ALKBH4; an initial step at 95 °C for 10 s and 40 cycles of 95 °C for 5 s, and 60 °C for 20 s for β-actin gene (ACTB); an initial step at 95 °C for 30 s and 50 cycles of 95 °C for 5 s, 60 °C for 10 s, and 68 °C for 15 s for E2F1 and CDC25A; and an initial step at 95 °C for 30 s and 50 cycles of 95 °C for 5 s, 58 °C for 10 s and 68 °C for 15 s for CCNE1 and MYBL2. The primer sequences used for gene amplification were as follows: ALKBH4 forward, 5′-TGATGCTGATCGAGGACTTTGTG-3′, and reverse, 5′-AAGCCCTCGGTCTTTAGCTTCTG-3′; ACTB forward, 5′-TGGCACCCAGCACAATGAA-3′, and reverse, 5′-CTAAGTCATAGTCCGCCTAGAAGCA-3′; E2F1 forward, 5′-CAAGAAGTCCAAGAACCACATCC-3′, and reverse, 5′-AGATATTCATCAGGTGGTCCAGC-3′; CDC25A forward, 5′-TTGTTGTGTTTCACTGCGAGTTTT-3′, and reverse, 5′-AGGGTAGTGGAGTTTGGGGTATTC-3′; CCNE1 forward, 5′-GCCAGCCTTGGGACAATAATG-3′, and reverse, 5′-CTTGCACGTTGAGTTTGGGT-3′; MYBL2 forward, 5′-CATTGTGGATGAGGATGTGAAGC-3′, and reverse, 5′-TGGTTGAGCAAGCTGTTGTCTTC-3′.
Gene array analysis
Total RNA was obtained from the ALKBH4 knockdown cells and from cells with an overexpression of ALKBH4 by using miRNeasy Mini Kit (QIAGEN). Total RNA (100 ng) was converted to cDNA and biotinylated using Ambion WT Expression Kit (Thermo Fisher Scientific, Waltham, Massachusetts, United States). Subsequently, biotinylated cDNA was hybridised to the GeneChip Human 2.0 ST Array (Affymetrix, Santa Clara, California, United States) and scanned using GeneChip Scanner 3000 (Affymetrix). The obtained gene array data were analysed using Partek Genomic Suite 6.6 software. Enrichment analysis was conducted by using Enricher (https://amp.pharm.mssm.edu/Enrichr/)67.
Establishment of ALKBH4 shRNA stable cells
A549 cells were seeded on the day before lentivirus infection and cultured in DMEM (Wako) supplemented with 10% foetal bovine serum and 100 mg/L kanamycin at 37 °C under a 5% CO2 atmosphere. Lentiviral particles, which were purchased from Sigma Aldrich (control shRNA: SHC002V and ALKBH4 shRNA: SHCLNV-NM_17621), were added to the culture medium to 4 multiplicity of infection (MOI), and polybrene (Thermo Fisher Scientific) was added at a final concentration of 8 ng/µL. After selection by 5 µg/mL puromycin (Sigma Aldrich), the expression levels of ALKBH4 were confirmed using qPCR, and cell lines with a high ALKBH4 knockdown efficiency were used for the next experiments.
Establishment of ALKBH4 shRNA stable cell-xenografted mice
Female BALB/c nude mice were obtained from Oriental Yeast Corporation (Tokyo, JAPAN). Five-week-old mice were used for ALKBH4 shRNA stable cell-xenograft experiments. Animals were kept under 12 h light–dark cycles at 22–24 °C. A549 cells, which had been stably transfected with ALKBH4 shRNA (A549-shALKBH4) and control shRNA (A549-shControl), were both adjusted to a concentration of 0.6 × 107 cells in 100 μL of serum-free DMEM. The cell suspensions, together with 100 μL of Matrigel Matrix High Concentration (Corning, New York, USA) were then injected subcutaneously into the right flanks of BALB/c nude mice (A549-shALKBH4, n = 8; A549-shControl, n = 8). One xenografted mouse, which was inoculated with A549-shALKBH4, died during the experiment. The tumour volume was calculated as follows: (tumour length × tumour width2)/2. All procedures were performed under a protocol approved by the Animal Experimentation Committee at Osaka University. All methods were carried out in accordance with relevant ARRIVE guidelines and regulations. We confirmed that all methods were carried out in accordance with relevant guidelines and regulations. Developed tumours were resected 52 days after xenografts.
ALKBH4 immunohistochemistry in clinical cases
Eighty patients who underwent radical operation for primary lung adenocarcinoma at Kagoshima University Hospital from January 2001 to December 2007 were subjected to immunohistochemistry for ALKBH4. Immunohistochemistry was conducted as described before64. Paraffin-embedded section (3 μm of thickness) were deparaffinised and dehydrated. The endogenous peroxidase activity of specimens was blocked using a 0.3% hydrogen peroxide solution in methanol. The sections were blocked with 1% bovine serum albumin and were incubated with the rabbit polyclonal antibody against human ALKBH4 (1:200; Novus Biologicals, NBP2-14737) overnight at 4 °C, followed by staining with a streptavidin–biotin peroxidase kit (Vector Laboratories, CA, USA). The immune complex was visualised by incubating the sections with diaminobenzidine tetrahydrochloride. The sections were counterstained with haematoxylin and mounted. Non-cancerous colon samples were used as positive controls for ALKBH4. ALKBH4 expression was determined by counting the number of cancer cells in which the cytoplasm was stained with the anti-ALKBH4 antibody. Two investigators evaluated ALKBH4 expression via immunohistochemistry within each tumour by assessing a total of 1000 cancer cells in 10 selected fields (100 cells/field) using high-power (× 200) microscopy, in an independent manner. The average labelling index of ALKBH4 was assessed according to the proportion of positive cells present in each field. Tumours with an average labelling index of 20% or more were defined as ALKBH4-positive. The specificity of the anti-ALKBH4 antibody was verified through immunofluorescence staining using A549 cells transfected with shControl and shALKBH4 (Supplementary Fig. S8).
Statistics
The results were expressed as the mean ± standard deviation (S.D.). Differences between the values were statistically analysed using the Student’s t-test, paired t-test or one-way analysis of variance (ANOVA) with Bonferroni post-hoc tests (GraphPad Prism 6.0, GraphPad software). Pearson correlation analysis was used for the correlation analysis. A p-value < 0.05 was considered statistically significant.
Ethical approval
The animal experiments were approved by the Animal Experimentation Committee at Osaka University. All animal experiments were performed in accordance with relevant guideline and regulations. For human research, written informed consent was obtained from each of the patients and the study was approved by the Ethical Committee of Kagoshima University Hospital (registration No. 351). Experiments using clinical specimens were approved by the institutional review boards of the Graduate School of Pharmaceutical Sciences, Osaka University.
References
Falnes, P. Ø., Johansen, R. F. & Seeberg, E. AlkB-mediated oxidative demethylation reverses DNA damage in Escherichia coli. Nature 419, 178–182 (2002).
Trewick, S. C., Henshaw, T. F., Hausinger, R. P., Lindahl, T. & Sedgwick, B. Oxidative demethylation by Escherichia coli AlkB directly reverts DNA base damage. Nature 419, 174–178 (2002).
Aas, P. A. et al. Human and bacterial oxidative demethylases repair alkylation damage in both RNA and DNA. Nature 421, 859–863 (2003).
Kurowski, M. A., Bhagwat, A. S., Papaj, G. & Bujnicki, J. M. Phylogenomic identification of five new human homologs of the DNA repair enzyme AlkB. BMC Genomics 1, 48 (2003).
Gerken, T. et al. The obesity-associated FTO gene encodes a 2-oxoglutarate-dependent nucleic acid demethylase. Science 318, 1469–1472 (2007).
Ougland, R., Rognes, T., Klungland, A. & Larsen, E. Non-homologous functions of the AlkB homologs. J. Mol. Cell Biol. 7, 494–504 (2015).
Fujii, T., Shimada, K., Anai, S., Fujimoto, K. & Konishi, N. ALKBH2, a novel AlkB homologue, contributes to human bladder cancer progression by regulating MUC1 expression. Cancer Sci. 104, 321–327 (2013).
Hotta, K. et al. Clinical significance and therapeutic potential of prostate cancer antigen-1/ALKBH3 in human renal cell carcinoma. Oncol. Rep. 34, 648–654 (2015).
Tasaki, M., Shimada, K., Kimura, H., Tsujikawa, K. & Konishi, N. ALKBH3, a human AlkB homologue, contributes to cell survival in human non-small-cell lung cancer. Br. J. Cancer 104, 700–706 (2011).
Shimada, K. et al. ALKBH3 contributes to survival and angiogenesis of human urothelial carcinoma cells through NADPH oxidase and tweak/Fn14/VEGF signals. Clin. Cancer Res. 18, 5247–5255 (2012).
Yamato, I. et al. PCA-1/ALKBH3 contributes to pancreatic cancer by supporting apoptotic resistance and angiogenesis. Cancer Res. 72, 4829–4839 (2012).
Konishi, N. et al. High expression of a new marker PCA-1 in human prostate carcinoma. Clin. Cancer Res. 11, 5090–5097 (2005).
Ohshio, I. et al. ALKBH8 promotes bladder cancer growth and progression through regulating the expression of survivin. Biochem. Biophys. Res. Commun. 477, 413–418 (2016).
Shimada, K. et al. A novel human AlkB homologue, ALKBH8, contributes to human bladder cancer progression. Cancer Res. 69, 3157–3164 (2009).
Li, M. et al. ALKBH4-dependent demethylation of actin regulates actomyosin dynamics. Nat. Commun. 4, 1832 (2013).
Bjørnstad, L. G. et al. Human ALKBH4 interacts with proteins associated with transcription. PLoS ONE 7, e49045 (2012).
Shen, C. et al. ALKBH4 functions as a suppressor of colorectal cancer metastasis via competitively binding to WDR5. Front. Cell Dev. Biol. 8, 293 (2020).
Keating, G. M. Nivolumab: A review in advanced squamous non-small cell lung cancer. Drugs 75, 1925–1934 (2015).
Morgillo, F., Corte, C. M., Fasano, M. & Ciardiello, F. Mechanisms of resistance to EGFR-targeted drugs: Lung cancer. ESMO Open 1, e000060 (2016).
Yamasaki, L. et al. Loss of E2F-1 reduces tumorigenesis and extends the lifespan of Rb1(+/−) mice. Nat. Genet. 18, 360–364 (1998).
Wu, L. et al. The E2F1-3 transcription factors are essential for cellular proliferation. Nature 414, 457–462 (2001).
Park, C., Lee, I. & Kang, W. K. E2F–1 is a critical modulator of cellular senescence in human cancer. Int. J. Mol. Med. 17, 715–720 (2006).
Engelmann, D. & Pützer, B. M. The dark side of E2F1: in transit beyond apoptosis. Cancer Res. 72, 571–575 (2012).
Lee, J. et al. Expression signature of E2F1 and its associated genes predict superficial to invasive progression of bladder tumors. J. Clin. Oncol. 28, 2660–2667 (2010).
Haller, F. et al. Prognostic role of E2F1 and members of the CDKN2A network in gastrointestinal stromal tumors. Clin. Cancer Res. 11, 6589–6597 (2005).
Huang, C. et al. E2F1 overexpression correlates with thymidylate synthase and survivin gene expressions and tumor proliferation in non-smallcell lung cancer. Clin. Cancer Res. 13, 6938–6946 (2007).
Tang, Z. et al. GEPIA: A web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 45, W98–W102 (2017).
Lachmann, A. et al. ChEA: Transcription factor regulation inferred from integrating genome-wide ChIP-X experiments. Bioinformatics 26, 2438–2444 (2010).
Younis, R. et al. CDC25A(Q110del): A novel cell division cycle 25A isoform aberrantly expressed in non-small cell lung cancer. PLoS ONE 7, e46464 (2012).
Huang, L. N. et al. Meta-analysis for cyclin E in lung cancer survival. Clin. Chim. Acta 413, 663–668 (2012).
Xiong, Y. C., Wang, J., Cheng, Y., Zhang, X. Y. & Ye, X. Q. Overexpression of MYBL2 promotes proliferation and migration of non-small-cell lung cancer via upregulating NCAPH. Mol. Cell. Biochem. 468, 185–193 (2020).
Ishida, S. et al. Role for E2F in control of both DNA replication and mitotic functions as revealed from DNA microarray analysis. Mol. Cell Biol. 14, 4684–4699 (2001).
Ren, B. et al. E2F integrates cell cycle progression with DNA repair, replication, and G2/M checkpoints. Genes Dev. 2, 245–256 (2002).
Stanelle, J., Stiewe, T., Theseling, C. C., Peter, M. & Pützer, B. M. Gene expression changes in response to E2F1 activation. Nucleic Acids Res. 8, 1859–1867 (2002).
Müller, H. et al. E2Fs regulate the expression of genes involved in differentiation, development, proliferation, and apoptosis. Genes Dev. 3, 267–285 (2001).
Kasahara, M. et al. Thymidylate synthase expression correlates closely with E2F1 expression in colon cancer. Clin. Cancer Res. 6, 2707–2711 (2000).
Sowers, R. et al. mRNA expression levels of E2F transcription factors correlate with dihydrofolate reductase, reduced folate carrier, and thymidylate synthase mRNA expression in osteosarcoma. Mol. Cancer Ther. 2, 535–541 (2003).
Suzuki, T. et al. Expression of the E2F family in human gastrointestinal carcinomas. Int. J. Cancer 81, 535–538 (1999).
Kweon, S. M. et al. An adversarial DNA N 6-methyladenine-sensor network preserves polycombsilencing. Mol. Cell 74, 1138–1147 (2019).
Greer, E. et al. DNA methylation on N6-adenine in C. elegans. Cell 161, 868–878 (2015).
Nilsen, A., Fusser, M., Greggains, G., Fedorcsak, P. & Klungland, A. ALKBH4 depletion in mice leads to spermatogenic defects. PLoS ONE 9, e105113 (2014).
Wang, S. et al. The demethylase NMAD-1 regulates DNA replication and repair in the Caenorhabditis elegans germline. PLoS Genet. 15, e1008252 (2019).
Jovanovic, M. et al. Immunogenetics. Dynamic profiling of the protein life cycle in response to pathogens. Science 347, 1259038 (2015).
Li, J., Bickel, P. & Biggin, M. System wide analyses have underestimated protein abundances and the importance of transcription in mammals. PeerJ 2, e270 (2014).
VogelC, M. E. Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nat. Rev. Genet. 13, 227–232 (2012).
Schwanhäusser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011).
Wolf, G. et al. Prognostic significance of Polo-like kinase (PLK) expression in non-small cell lung cancer. Oncogene 14, 543–549 (1997).
Zhou, Q., Fan, G. & Dong, Y. Polo-like kinase 4 correlates with greater tumor size, lymph node metastasis and confers poor survival in non-small cell lung cancer. Lab. Anal. 34, e23152 (2020).
Shin, S., Jang, H., Xu, R., Won, J. & Yim, H. Active PLK1-driven metastasis is amplified by TGF-β signaling that forms a positive feedback loop in non-small cell lung cancer. Oncogene 39, 767–785 (2020).
Tian, X. et al. Polo-like kinase 4 mediates epithelial-mesenchymal transition in neuroblastoma via PI3K/Akt signaling pathway. Cell Death Dis. 9, 54 (2018).
Mahanic, C., Budhavarapu, V., Graves, J., Li, G. & Lin, W. C. Regulation of E2 promoter binding factor 1 (E2F1) transcriptional activity through a deubiquitinating enzyme, UCH37. J. Biol. Chem. 290, 26508–26522 (2015).
Liu, K., Luo, Y., Lin, F. & Lin, W. TopBP1 recruits Brg1/Brm to repress E2F1-induced apoptosis, a novel pRb-independent and E2F1-specific control for cell survival. Genes Dev. 18, 673–686 (2004).
Araki, K., Nakajima, Y., Eto, K. & Ikeda, M. Distinct recruitment of E2F family members to specific E2F-binding sites mediates activation and repression of the E2F1 promoter. Oncogene 22, 7632–7641 (2003).
Martinez, L. A. et al. E2F3 is a mediator of DNA damage-induced apoptosis. Mol. Cell Biol. 30, 524–536 (2010).
Iaquinta, P. J. & Lees, J. A. Life and death decisions by the E2F transcription factors. Curr. Opin. Cell Biol. 19, 649–657 (2007).
Chen, Z. et al. Effect of ubiquitin carboxy-terminal hydrolase 37 on apoptotic in A549 cells. Cell Biochem. Funct. 29, 142–148 (2011).
Neilsen, B. et al. WDR5 supports colon cancer cells by promoting methylation of H3K4 and suppressing DNA damage. BMC Cancer 18, 673 (2018).
Xie, Q., Li, Z. & Chen, J. WDR5 positively regulates p53 stability by inhibiting p53 ubiquitination. Biochem. Biophys. Res. Commun. 487, 333–338 (2017).
Liberti, M. V. & Locasale, J. W. The warburg effect: How does it benefit cancer cells?. Trends Biochem. Sci. 3, 211–218 (2016).
Tarangelo, A. et al. Recruitment of Pontin/Reptin by E2f1 amplifies E2f transcriptional response during cancer progression. Nat. Commun. 6, 10028 (2015).
Wu, M., Seto, E. & Zhang, J. E2F1 enhances glycolysis through suppressing Sirt6 transcription in cancer cells. Oncotarget 6, 11252–11263. https://doi.org/10.18632/oncotarget.3594 (2015).
Chen, C. et al. NANOG metabolically reprograms tumor-initiating stem-like cells through tumorigenic changes in oxidative phosphorylation and fatty acid metabolism. Cell Metab. 23, 206–219 (2016).
Gazdar, A. F. Activating and resistance mutations of EGFR in non-small-cell lung cancer: role in clinical response to EGFR tyrosine kinase inhibitors. Oncogene 28, S24–S31 (2009).
Aoki, M. et al. Expression of bone morphogenetic protein-7 significantly correlates with non-small cell lung cancer progression and prognosis: A retrospective cohort study. Clin. Med. Insights Oncol. 13, 1179554919852087 (2019).
Hirono, T. et al. MicroRNA-130b functions as an oncomiRNA in non-small cell lung cancer by targeting tissue inhibitor of metalloproteinase-2. Sci. Rep. 9, 6956 (2019).
Jingushi, K. et al. Leukocyte-associated immunoglobulin-like receptor 1 promotes tumorigenesis in RCC. Oncol. Rep. 41, 1293–1303 (2019).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinform. 14, 128 (2013).
Acknowledgements
This research was partially supported by the Platform Project for Supporting Drug Discovery and Life Science Research (Basis for Supporting Innovative Drug Discovery and Life Science Research (BINDS)) from AMED, under Grant Number JP20am0101084 and JP20am0101123.
Author information
Authors and Affiliations
Contributions
Conception and design: K.J., M.Ao. and K.T. Acquisition of data: K.J., M.Ao., K.U., M.T., M.A., T.M., K.M., Y.U., K.K., H.H., T.N., A.T., M.Y., K.Ka., T.F., M.S., K.T., Y.M., T.K. and K.Ta. Analysis and interpretation of data: K.J., M.Ao., K.U., M.T., M.A., T.M., K.M., T.N., A.T., M.Y., K.Ka., T.F., M.S., K.T., Y.M., T.K. and K.Ta. Writing, review, and/or revision of the manuscript: K.J., M.Ao., K.U., M.T., M.A., T.M., K.M., T.N., A.T., M.Y., K.Ka., T.F., M.S. and K.T.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jingushi, K., Aoki, M., Ueda, K. et al. ALKBH4 promotes tumourigenesis with a poor prognosis in non-small-cell lung cancer. Sci Rep 11, 8677 (2021). https://doi.org/10.1038/s41598-021-87763-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-87763-1
This article is cited by
-
ALKBH4 impedes 5-FU Sensitivity through suppressing GSDME induced pyroptosis in gastric cancer
Cell Death & Disease (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.