Abstract
The inhibitory effects of programmed cell death 1/programmed cell death ligand 1 (PD-1/PD-L1) modulates T-cell depletion. T-cell depletion is one of the key mechanisms of hepatitis B virus (HBV) persistence, in particular liver disease progression and the development of hepatocellular carcinoma (HCC). This case–control study aimed to understand the significance of PD-1 polymorphisms (PD-1.5 and PD-1.9) association with HBV infection risk and HBV-induced liver disease progression. Genotyping of PD-1.5 and PD-1.9 variants was performed by direct Sanger sequencing in 682 HBV-infected patients including chronic hepatitis (CHB, n = 193), liver cirrhosis (LC, n = 183), hepatocellular carcinoma (HCC, n = 306) and 283 healthy controls (HC). To analyze the association of PD-1 variants with liver disease progression, a binary logistic regression, adjusted for age and gender, was performed using different genetic models. The PD-1.9 T allele and PD-1.9 TT genotype are significantly associated with increased risk of LC, HCC, and LC + HCC. The frequencies of PD-1.5 TT genotype and PD-1.5 T allele are significantly higher in HCC compared to LC patients. The haplotype CT (PD-1.5 C and PD-1.9 T) was significantly associated with increased risk of LC, HCC, and LC + HCC. In addition, the TC (PD-1.5 T and PD-1.9 C) haplotype was associated with the risk of HCC compared to non-HCC. The PD-1.5 CC, PD-1.9 TT, genotype, and the CC (PD-1.5 C and PD-1.9) haplotype are associated with unfavorable laboratory parameters in chronic hepatitis B patients. PD-1.5 and PD1.9 are useful prognostic predictors for HBV infection risk and liver disease progression.
Similar content being viewed by others
Introduction
Chronic hepatitis B virus (CHB) infection causes long-term clinical complications, including acute liver failure, liver cirrhosis (LC) and hepatocellular carcinoma (HCC). WHO estimates 257 million cases of CHB and 780,000 HBV-related deaths annually1. Although an effective vaccination program has reduced the burden of hepatitis B virus (HBV), the persistence of CHB in adult populations remains a significant health burden, particularly in low and middle-income countries, including Vietnam1,2. The incidence of HBV is > 10% in the Vietnamese population and is one of the major contributors of mortality1.
During the clinical course of HBV, both innate and adaptive immune responses largely modulate the liver disease progression and subsequent liver damage. Several compelling evidences reveal that human cellular immune response is crucial for HBV pathogenesis and such responses are mediated by CD8+ and CD4+T cells, especially by HBV-specific CD8+T cells3,4,5,6,7. HBV clearance is arbitrated by HBV-specific CD8+T cells, by cytopathic and noncytolytic activity7,8,9, largely during the acute phase. A relatively weak HBV-specific CD8+T cell response was equally observed during the chronic phase4,10,11,12,13. The HBV-specific CD8+T cell depletion occurs primarily by the activation of the inhibitory checkpoint PD-1/PD-L1 (Programmed cell death 1/Programmed cell death ligand 1) signaling pathway14,15,16,17,18,19,20.
PD-1 is a transmembrane protein and an immunoinhibitory receptor, that are largely expressed on the surface of T cells, B cells, natural killer T cells and activated monocytes21,22,23. The PD-1 immunomodulatory ligands (PD-L1 and PD-L2) maintain immune homeostasis, but can also attenuate T-cell proliferation and regulate cytokine responses24. The phenomenon of persistent overexpression of PD-1 and its ligands is a common feature in chronic infections and cancer25,26,27, and blocking inhibitory signal transduction has led to increased T-cell function, and thus improved clinical outcomes. Blocking the PD-1/PD-L1 signaling with anti PD-1 or anti-PD-L1 molecules in combination with antiviral therapies has led to better immunological and clinical responses in HBV infections28,29. Clinical studies have also used expression of PD-1 and CTLA-4 as prognostic indicators, in evaluating antiviral HBV therapy responses30.
Several genetic association studies had investigated genetic variants of programmed cell death 1 (PD-1) and programmed death ligand 1 (PD-L1) with cancer, autoimmune and infectious diseases in different world populations31,32,33,34,35. In particular, five single nucleotide polymorphisms of PD1: PD-1.1 (-538G/A), PD-1.3 (+ 7146G/A), PD-1.5 (+ 7785T/C), PD-1.6 (+ 8669G/A) and PD-1.9 (+ 7625C/T) were associated with human malignancies and is highly expressed in several cancers32,36,37,38,39. Genetic variations in the promoter and at the transcription binding site can alter gene function. The PD-1.1 (-538G/A) promoter variant, located on the translation start codon, while PD-1.3 (+ 7146A/G) is located on intron 4, has been described to alter the gene transcription, since there are four tandem repeats consisting of several putative binding sequences of transcription factors40. PD-1.5 (+ 7785T/C), a synonymous mutation in exon 5, which is in linkage equilibrium with other PD-1 variants31. Relatively few studies have investigated on the association of PD-1 variants in CHB infection and liver disease progression, but the results are debatable41,42,43,44,45.
The investigated PD-1 variants are located in the 5′ upstream regions of the PD-1 locus (Genbank ID: 5133). The variant PD-1.5 (rs2227981) is associated with cancer susceptibility46; while the variant PD-1.9 (rs2227982) is associated with risk of HBV infection and clinical outcome47. In this context, we conducted a case–control study with the aim to determine the association of PD1 (PD-1.5 and PD-1.9) variants with HBV infection risk, liver disease progression and clinical outcome in the Vietnamese population.
Materials and methods
Ethics statement
This study was performed in accordance with the relevant national guidelines and regulations. Informed written consent was obtained from all participants after explanation of the study at the time of sampling. Informed consent was obtained from a parent and/or legal guardian for minors. The study was approved by the institutional review board of the 108 Institute of Clinical Medical and Pharmaceutical Sciences, Hanoi, Vietnam.
Patients and liver specimens
A total of 682 unrelated Vietnamese HBV-infected patients were randomly recruited in a case–control design at 108 Institute of Clinical Medical and Pharmaceutical Sciences, Hanoi, Vietnam, between 2012 and 2015. Patients were assigned to subgroups of disease based on clinical manifestations, liver function tests, imaging modalities and histological confirmation for all HCC cases. Subgroups were chronic hepatitis (CHB, n = 193), liver cirrhosis (LC, n = 183), hepatocellular carcinoma (HCC, n = 306). The diagnostic criteria for the CHB patients and the HBV-related LC were previously described48. The HBV-related HCC group was characterized as patients infected with HBV and was diagnosed based on the AASLD practice guideline49. None of these HBV-infected patients had evidence of chronic comorbidities/conditions such as: autoimmune diseases, alcoholic liver disease, type 2 diabetes, cancers, addiction to smoking and alcohol. All patients were confirmed negative for anti-HCV and anti-HIV by ELISA assays. Laboratory parameters including HBV-DNA loads and liver function tests including alanine transaminase (ALT), aspartate transaminase (AST), total bilirubin and direct bilirubin, albumin, prothrombin were obtained from the patients’ medical records. In addition, 283 healthy individuals (HC) were collected and assigned as the control group. These individuals were healthy individuals who presented themselves to the hospital for a routine medical check-up. Hematological and biochemical parameters were normal and serological tests were negative for hepatitis C virus and human immune deficiency virus. Finally, plasma and blood cells were separated and frozen at − 80 °C until use.
PD-1 genotyping
Genomic DNA was isolated from 200 µl of whole blood using a DNA isolation kit (Qiagen, Hilden, Germany), following manufacturer’s instructions. The process of PD-1 genotyping was followed the study protocol as described50. The amplicon containing the variants PD-1.5 (rs2227981) and PD-1.9 (rs2227982) was amplified by PCR (PCR1) using the primer pairs PD-1.5/9_F: 5′-GCA AGA ATG CCA GGG ACA TTT CAG AG-3′ and PD-1.5/9_R: 5′-TGC CTG GTG CAG GTG CAG-3′. PCR amplification was performed out in 25 μl reaction volumes containing: 1 × PCR buffer, 0.2 mM dNTPs, 1 mM MgCl2, 0.15 mM of each primer, 1 unit of Taq polymerase and 50 ng of genomic DNA. Cycling conditions: denaturation at 95 °C for 5 min, followed by 35 cycles of three-step cycling with denaturation at 94 °C for 40 s, annealing at 66 °C (PCR1) and 60 °C (PCR2) for 40 s, and extension at 72 °C for 45 s and a final extension at 72 °C for 7 min. PCR products were purified using the Exo-SAP-IT PCR product cleanup reagent (Affymetrix Santa Clara, USA) 5 µl of purified PCR products were used as templates. Sequencing was performed using the BigDye terminator v.1.1 cycle sequencing kit (Applied Biosystems, Foster City, CA, USA) on an ABI 3130XL DNA sequencer according to the manufacturer’s instructions.
Statistical analysis
All statistical analysis was performed using R version 3.1.2 (http://www.r-project.org). Statistical analysis plan was tailored as described previously50. In brief, genotype and allelic frequencies were determined by simple gene counting and the haplotype frequency was estimated using the expectation-maximum algorithm method implemented in the Arlequin v.3.5.2.2. The deviations from Hardy–Weinberg equilibrium were calculated for each group. We used a binary logistic regression adjusted for age and gender to analyze association of PD-1 variants with HBV-related liver diseases applying for different genetic models and adjusted odds ratios (aOR) with 95% confidence intervals (CI) were calculated. Chi-square tests were used to test for significant differences of categorical variables and Mann–Whitney-Wilcoxon and Kruskal–Wallis tests were applied to compare quantitative variables between groups. Significance was set at a value of p < 0.05.
Results
Baseline characteristics of HBV patients
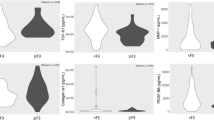
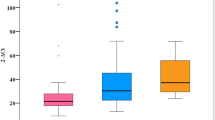
The baseline profiles of 682 HBV-infected patients and 283 healthy controls (HC) are described in Table 1 and Fig. 1. Most of the HBV and HC patients are male (88% and 63%, respectively) and the HCs were younger than HBV patients. The ALT, AST liver enzymes were significantly higher in CHB patients than other subgroups (p < 0.0001). Albumin and prothrombin levels, red blood cells and platelets were lower in LC patients than in other patient groups (p < 0.0001). HCC patients had higher AFP levels than CHB and LC patients (p < 0.001).
Evaluation of clinical parameters in HBV patient subgroups. CHB chronic hepatitis B, LC liver cirrhosis, HCC hepatocellular carcinoma, PLT platelets, AST and ALT aspartate and alanine amino transferase, WBC white blood cells, RBC red blood cells, PLT platelet, IU international unit. Box-plots illustrate medians with 25 and 75 percentiles with whiskers to 10 and 90 percentiles. p-values were calculated by Kruskal–Wallis test.
PD-1 variants and HBV related liver disease progression
The genotype and allele frequencies of two PD-1 variants (PD-1.5 and PD-1.9) in clinically classified 682 HBV patients and 283 HCs are summarized in the Tables 2 and 3. The variants were in Hardy–Weinberg equilibrium (p > 0.05) for both the cases and the controls. No significant association with HBV infection and clinical course was observed for PD-1.5 variant. For PD-1.9 variant, the heterozygous genotype PD-1.9 CT showed significant association with HBV infection (HBV patients vs. HC: OR = 1.4; 95% CI = 1.01–1.98; padj = 0.04).
A significant difference in PD-1.5 allelic and genotype frequencies was observed between HCC and LC patients [PD-1.5 Allele T OR = 1.6 (1.13–2.1), padj = 0.006; PD-1.5 genotype TT co-dominant model: OR = 4.4 (1.2–15.6), padj = 0.016; dominant model: OR = 1.6 (1.1–2.35), padj = 0.013; recessive model: OR = 3.7 (1.1–13.1), padj = 0.02]. For the PD-1.9 variant significant differences in distribution of allele and genotypes were observed in LC, HCC and LC + HCC patients compared to CHB patients. The minor allele PD-1.9 T increased CHB patients’ risk towards disease progression either as LC, HCC or LC + HCC [LC vs. CHB: OR = 1.7(1.2–2.4), padj = 0.00014; HCC vs. CHB: OR = 1.3 (1.1–1.8), padj = 0.029; LC + HCC vs. CHB: OR = 1.5 (1.1–2.0); padj = 0.0043]. The PD-1.9 homozygous genotype TT was observed frequently among patients with LC, HCC and LC + HCC groups compared to CHB patients in the co-dominant [LC vs. CHB: OR = 3.8(1.8–8.0), padj = 0.0007; HCC vs. CHB: OR = 2.1(1.01–4.3), padj = 0.047; LC + HCC vs. CHB: OR = 2.9(1.5–5.6), padj = 0.004] and recessive [LC vs. CHB: OR = 3.4(1.7–6.7), padj = 0.00019; HCC vs. CHB: OR = 1.8(1.09–3.6), padj = 0.042; LC + HCC vs. CHB: OR = 2.5(1.4–4.7), padj = 0.0015] models respectively.
PD-1 haplotypes and HBV-related liver diseases
A total of four haplotypes were reconstructed from both PD-1.5 and PD-1.9 variants and the frequencies were presented in the Table 4. No significant association in the distribution of haplotypes in HC and HBV and patient subgroups was observed. The analysis within HBV patient subgroups showed that the haplotype CT (PD-1.5 C and PD-1.9 T) was frequently observed in patients with advanced liver disease rather than those with CHB [LC vs. CHB: OR = 1.6(1.1–2.4), padj = 0.009; HCC vs. CHB: OR = 1.5 (1.1–2.2), padj = 0.016; LC + HCC vs. CHB: OR = 1.6(1.2–2.2), padj = 0.0028]. In addition, haplotype TC (PD-1.5 T and PD-1.9 C) was a prognostic indicator for HCC [HCC vs. non-HCC: OR = 1.4 (1.1–1.9); padj = 0.019]. Both CT and TC haplotypes contribute to liver disease progression and clinical outcome in HBV infected patients.
PD-1 genetic variant and laboratory parameters
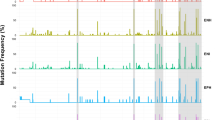
The PD-1.5 and PD-1.9 genetic variants were associated with distinct biochemical and laboratory parameters (Figs. 2 and 3). The total and direct bilirubin levels were higher in patients with genotype PD-1.5 CC compared to those with genotypes CT/TT (p = 0.016 and p = 0.05, respectively). In addition, prothrombin levels were significantly decreased among those with PD-1.5 CC genotypes (p = 0.00036). Equally other blood cell counts such as platelets, white and red blood cells were also significantly decreased among those carrying PD-1.5 CC genotypes (p = 0.042, 0.018 0.056 respectively). In the PD-1.9 variant, patients with the genotype TT had lower platelet, red blood cell, prothrombin and albumin levels compared to the CC/CT genotype (p = 0.009, 0.0019, 0.00025 and 0.047, respectively). In contrast, total bilirubin levels were significantly higher in the genotype TT than in the genotype CC/CT. There was no significant correlation between PD-1.9 and other laboratory parameters such as liver enzymes, HBV DNA and AFP.
We also analyzed the association between constructed haplotypes and laboratory parameters (Fig. 4). We observed that patients with the TC haplotype had higher prothrombin and platelet levels than other CC and CT haplotypes (p = 0.0041, 0.0398, respectively). The TC haplotype was significantly associated with lower levels of total bilirubin compared to CC haplotypes (p = 0.026).
Discussion
PD-1 is a member of the immunoglobulin superfamily and its cytoplasmic domain contain immunoreceptor tyrosine inhibitory motif associated with inhibitory signaling. The interaction between PD-1 and PD-L1 is known to contribute primarily to the depletion of T cells and thus involved in carcinogenesis. Several variants in the PD-1 gene have been identified to associate with various infectious and autoimmune diseases, and cancers. In this case–control study, we investigated on the PD-1.5 (+ 7785T/C) and PD-1.9 (+ 7625C/T) variants in Vietnamese HBV-infected patients and controls. Two variants PD-1.5 and PD-1.9 were observed to influence the clinical outcome of HBV, in particular progression of liver disease.
PD-1.5 is a synonymous substitution, whereas PD-1.9 is a non-synonymous substitution (valine to alanine) that modulates structural and functional properties of PD-1. Several studies have investigated on these two variants with different human malignancies31,32, autoimmune disorders51,52,53 and infectious diseases, including HBV infection41,47,54. PD-1.5 and PD-1.9 variants susceptibility to HBV infection were earlier studied47,54, and in particular, the PD-1.9 T allele is considered as a predisposing factor for HBV susceptibility47. In addition, the frequency of TT and CT genotypes in HBV patients was significantly higher than healthy controls. Based on the reported results, Huang et al. pointed out that the CC genotype of rs2227982 and the C allele are protective factors in HBV infection47. In our study, we also reported that when analyzed with the age- and sex-adjusted logistic regression model, the frequency of CT genotype was significantly higher in HBV patients than in controls [co-dominant (CT:CC) model: OR = 1.4 (1.01–1.98), P = 0.042]. However, another study found no association between this variant and HBV infection55. For the PD-1.5 polymorphism, we found no significant association with HBV infection. Our conclusions were in contrast to a previous study, which found an association between this polymorphism and susceptibility to chronic HBV infection in the Chinese population54. In addition, this polymorphism was also reported to be associated with HCV infection56.
Few studies suggest that PD-1 as a convincing genetic indicator for different cancers in different world populations, however results were contradictory and ambiguous31,32,33,34,35,46. To date, comprehensive meta-analyses have been conducted to investigate the association of PD-1 polymorphisms, notably PD-1.5 and PD-1.9, with risk of cancer32,46,57,58. Concerning the PD-1.5 polymorphism, Dong et al. showed that the allele T (compared to the C allele) is associated with a reduced risk of cancer in the general population32. In addition, two other meta-analyses have shown that the PD-1.5 TT genotype (TT compared to CT/CC) significantly reduces the overall risk of cancer {OR = 0.82, 95% CI = 0.68–0. 99, P = 0.0457]; [OR = 0.75, 95% CI: 0.64–0.86, P < 0.000146} and {OR = 0.65, 95% CI: 0.47–0.90, P < 0.05,58}. In regards to PD-1.9 polymorphism, no significant association between this variant and overall cancer susceptibility was established in these meta-analyses32,46,57,58. However, Hashemi et al.57 performed stratified analyses and indicated that the PD-1.9 polymorphism was associated with increased risk of general cancer in hospital-based study (OR = 1.2, 95% CI = 1.05–1.37, P = 0.008, CT/TT vs. CC). Additionally, stratified analyses found the association between PD-1.9 and increased risk of gastrointestinal cancer (OR = 1.16, 95% CI = 1.03–1.30, P = 0.017, CT/TT vs. CC) but decreased risk of breast cancer (OR = 0.73, 95% CI = 0.60–0.89, P = 0.002, CT/TT vs. CC).
It should be noted that these above meta-analyses included several case–control studies, but no studies have examined the association between PD-1.5 and PD-1.9 polymorphisms and HCC risk. To our knowledge, this is the first study in which we have examined the link between PD-1.5 and PD-1.9 polymorphisms and the progression of liver disease including HCC. We have shown that the PD-1.5 and PD-1.9 polymorphisms are associated with the progression of HBV-related liver diseases. Particularly, the variant PD-1.9 shows a recessively homozygous advantage in disease progression and might be a risk factor for the advanced disease progression including LC and HCC. The results from our study indicate that PD-1.9 polymorphism may influence liver disease progression and HCC development in chronic HBV infection. There is growing evidence that PD-1 overexpression is associated with T-cell dysfunction and exhaustion in chronic HBV infection and HCC development59,60,61.
As a single SNP site could not well represent the influence of PD-1 genetic variants in HBV infection, we analyzed the association of haplotypes constructed from variants PD-1.5 and PD-1.9 with the liver disease progression and outcomes in HBV-infected patients. The study results could show a way by integrating different sites to completely elucidate the potent role of PD-1 genetic variants in the risk of HBV infection and HBV-related clinical presentation. Still more works are needed to be done to examine the clinical significance of PD-1 genetic variants as the host genetic perspective in HBV infection.
This study has two limitations. First, a longitudinal monitoring of HCC patients would be an added advantage in understanding the overall survival rates among patients with HBV-related HCC. Secondly, quantification of PD-1 gene expression in liver tissues of HCC patients may additionally warrant on the relevance these variants in clinical routine. Taken together, this study demonstrates on the role of PD-1.5 and PD-1.9 genetic variants in predicting HBV infection risk and subsequent liver disease progression.
Change history
01 February 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41598-022-06068-z
References
WHO. Global Hepatitis Report, 2017 (2017).
Liang, X. et al. Epidemiological serosurvey of hepatitis B in China—Declining HBV prevalence due to hepatitis B vaccination. Vaccine. 27, 6550–6557. https://www.ncbi.nlm.nih.gov/pubmed/19729084 (2009).
Chisari, F. V. & Ferrari, C. Hepatitis B virus immunopathogenesis. Annu. Rev. Immunol. 13, 29–60. http://www.ncbi.nlm.nih.gov/pubmed/7612225 (1995).
Chang, J. J. & Lewin, S. R. Immunopathogenesis of hepatitis B virus infection. Immunol. Cell Biol. 85, 16–23. http://www.ncbi.nlm.nih.gov/pubmed/17130898 (2007).
Rehermann, B. & Nascimbeni, M. Immunology of hepatitis B virus and hepatitis C virus infection. Nat. Rev. Immunol. 5, 215–229. http://www.ncbi.nlm.nih.gov/pubmed/15738952 (2005).
Shuai, Z. et al. Adaptive immunity in the liver. Cell Mol. Immunol. 13, 354–368. http://www.ncbi.nlm.nih.gov/pubmed/26996069 (2016).
Guidotti, L. G. et al. Viral clearance without destruction of infected cells during acute HBV infection. Science. 284, 825–829. http://www.ncbi.nlm.nih.gov/pubmed/10221919 (1999).
Guidotti, L. G. & Chisari, F. V. Immunobiology and pathogenesis of viral hepatitis. Annu. Rev. Pathol. 1, 23–61. http://www.ncbi.nlm.nih.gov/pubmed/18039107 (2006).
Schuch, A., Hoh, A. & Thimme, R. The role of natural killer cells and CD8(+) T cells in hepatitis B virus infection. Front. Immunol. 5, 258. http://www.ncbi.nlm.nih.gov/pubmed/24917866 (2014).
Phillips, S. et al. CD8(+) T cell control of hepatitis B virus replication: Direct comparison between cytolytic and noncytolytic functions. J. Immunol. 184, 287–295. http://www.ncbi.nlm.nih.gov/pubmed/19949099 (2010).
Yang, P. L. et al. Immune effectors required for hepatitis B virus clearance. Proc. Natl. Acad. Sci. U.S.A. 107, 798–802. http://www.ncbi.nlm.nih.gov/pubmed/20080755 (2010).
Bertoletti, A. & Naoumov, N. V. Translation of immunological knowledge into better treatments of chronic hepatitis B. J. Hepatol. 39, 115–124. http://www.ncbi.nlm.nih.gov/pubmed/12821053 (2003).
Webster, G. J. et al. Incubation phase of acute hepatitis B in man: Dynamic of cellular immune mechanisms. Hepatology. 32, 1117–1124. http://www.ncbi.nlm.nih.gov/pubmed/11050064 (2000).
Boni, C. et al. Characterization of hepatitis B virus (HBV)-specific T-cell dysfunction in chronic HBV infection. J. Virol. 81, 4215–4225. https://www.ncbi.nlm.nih.gov/pubmed/17287266 (2007).
Maier, H., Isogawa, M., Freeman, G. J. & Chisari, F. V. PD-1:PD-L1 interactions contribute to the functional suppression of virus-specific CD8+ T lymphocytes in the liver. J. Immunol. 178, 2714–2720. https://www.ncbi.nlm.nih.gov/pubmed/17312113 (2007).
Peng, G. et al. PD-1 upregulation is associated with HBV-specific T cell dysfunction in chronic hepatitis B patients. Mol. Immunol. 45, 963–970. https://www.ncbi.nlm.nih.gov/pubmed/17868872 (2008).
Fisicaro, P. et al. Antiviral intrahepatic T-cell responses can be restored by blocking programmed death-1 pathway in chronic hepatitis B. Gastroenterology. 138, 682–693, 693 e681–684. https://www.ncbi.nlm.nih.gov/pubmed/19800335 (2010).
Zhang, Z. et al. Dynamic programmed death 1 expression by virus-specific CD8 T cells correlates with the outcome of acute hepatitis B. Gastroenterology. 134, 1938–1949, 1949 e1931–1933. https://www.ncbi.nlm.nih.gov/pubmed/18455515 (2008).
Ye, B. et al. T-cell exhaustion in chronic hepatitis B infection: current knowledge and clinical significance. Cell Death Dis. 6, e1694. https://www.ncbi.nlm.nih.gov/pubmed/25789969 (2015).
Watanabe, T., Bertoletti, A. & Tanoto, T. A. PD-1/PD-L1 pathway and T-cell exhaustion in chronic hepatitis virus infection. J. Viral Hepatol. 17, 453–458. https://www.ncbi.nlm.nih.gov/pubmed/20487259 (2010).
Bardhan, K., Anagnostou, T. & Boussiotis, V. A. The PD1:PD-L1/2 pathway from discovery to clinical implementation. Front. Immunol. 7, 550. https://www.ncbi.nlm.nih.gov/pubmed/28018338 (2016).
Agata, Y. et al. Expression of the PD-1 antigen on the surface of stimulated mouse T and B lymphocytes. Int. Immunol. 8, 765–772. https://www.ncbi.nlm.nih.gov/pubmed/8671665 (1996).
Greenwald, R. J., Freeman, G. J. & Sharpe, A. H. The B7 family revisited. Annu. Rev. Immunol. 23, 515–548. https://www.ncbi.nlm.nih.gov/pubmed/15771580 (2005).
Carter, L. et al. PD-1:PD-L inhibitory pathway affects both CD4(+) and CD8(+) T cells and is overcome by IL-2. Eur. J. Immunol. 32, 634–643. https://www.ncbi.nlm.nih.gov/pubmed/11857337 (2002).
Pauken, K. E. & Wherry, E. J. Overcoming T cell exhaustion in infection and cancer. Trends Immunol. 36, 265–276. https://www.ncbi.nlm.nih.gov/pubmed/25797516 (2015).
Barber, D. L. et al. Restoring function in exhausted CD8 T cells during chronic viral infection. Nature 439, 682–687. https://www.ncbi.nlm.nih.gov/pubmed/16382236 (2006).
Saito, H. et al. Highly activated PD-1/PD-L1 pathway in gastric cancer with PD-L1 expression. Anticancer Res. 38, 107–112. https://www.ncbi.nlm.nih.gov/pubmed/29277762 (2018).
Liu, J. et al. Enhancing virus-specific immunity in vivo by combining therapeutic vaccination and PD-L1 blockade in chronic hepadnaviral infection. PLoS Pathog. 10, e1003856. https://www.ncbi.nlm.nih.gov/pubmed/24391505 (2014).
E. Gane et al. A phase1 study evaluating anti-PD-1 treatment with or without GS-4774 in HBeAg negative chronic hepatitis B patients. J. Hepatol. 66, S26–S27 (2017).
Janssen, H. L. In HBV Cure Meeting—Boston (2016).
Salmaninejad, A. et al. PD-1 and cancer: molecular mechanisms and polymorphisms. Immunogenetics 70, 73–86. https://www.ncbi.nlm.nih.gov/pubmed/28642997 (2018).
Dong, W. et al. Programmed cell death-1 polymorphisms decrease the cancer risk: A meta-analysis involving twelve case-control studies. PLoS One. 11, e0152448. https://www.ncbi.nlm.nih.gov/pubmed/27031235 (2016).
Lee, S. Y. et al. Functional polymorphisms in PD-L1 gene are associated with the prognosis of patients with early stage non-small cell lung cancer. Gene 599, 28–35. https://www.ncbi.nlm.nih.gov/pubmed/27838455 (2017).
Yang, Q., Liu, Y., Liu, D., Zhang, Y. & Mu, K. Association of polymorphisms in the programmed cell death 1 (PD-1) and PD-1 ligand genes with ankylosing spondylitis in a Chinese population. Clin. Exp. Rheumatol. 29, 13–18. https://www.ncbi.nlm.nih.gov/pubmed/21269571 (2011).
Yeo, M. K. et al. Association of PD-L1 expression and PD-L1 gene polymorphism with poor prognosis in lung adenocarcinoma and squamous cell carcinoma. Hum. Pathol. 68, 103–111. https://www.ncbi.nlm.nih.gov/pubmed/28851662 (2017).
Yin, L., Guo, H., Zhao, L. & Wang, J. The programmed death-1 gene polymorphism (PD-1.5 C/T) is associated with non-small cell lung cancer risk in a Chinese Han population. Int. J. Clin. Exp. Med. 7, 5832–5836. https://www.ncbi.nlm.nih.gov/pubmed/25664115 (2014).
Haghshenas, M. R., Dabbaghmanesh, M. H., Miri, A., Ghaderi, A. & Erfani, N. Association of PDCD1 gene markers with susceptibility to thyroid cancer. J. Endocrinol. Invest. 40, 481–486. https://www.ncbi.nlm.nih.gov/pubmed/27943063 (2017).
Li, X. F., Jiang, X. Q., Zhang, J. W. & Jia, Y. J. Association of the programmed cell death-1 PD1.5 C>T polymorphism with cervical cancer risk in a Chinese population. Genet. Mol. Res. 15. https://www.ncbi.nlm.nih.gov/pubmed/27050970 (2016).
Ren, H. T. et al. PD-1 rs2227982 polymorphism is associated with the decreased risk of breast cancer in northwest Chinese women: A hospital-based observational study. Medicine (Baltimore). 95, e3760. https://www.ncbi.nlm.nih.gov/pubmed/27227944 (2016).
Mahmoudi, M. et al. PDCD1 single nucleotide genes polymorphisms confer susceptibility to juvenile-onset systemic lupus erythematosus. Autoimmunity 48, 488–493. https://www.ncbi.nlm.nih.gov/pubmed/26108738 (2015).
Zhang, G. et al. Association of polymorphisms of programmed cell death-1 gene with chronic hepatitis B virus infection. Hum. Immunol. 71, 1209–1213. https://www.ncbi.nlm.nih.gov/pubmed/20837075 (2010).
E. Galmozzi, F. Facchetti, G. G., A. Loglio, M. C. & Lampertico, P. In International Liver Congress (2018).
Li, Z. et al. Immune checkpoint proteins PD-1 and TIM-3 are both highly expressed in liver tissues and correlate with their gene polymorphisms in patients with HBV-related hepatocellular carcinoma. Medicine (Baltimore). 95, e5749. https://www.ncbi.nlm.nih.gov/pubmed/28033288 (2016).
Li, Z. et al. Genetic polymorphisms of immune checkpoint proteins PD-1 and TIM-3 are associated with survival of patients with hepatitis B virus-related hepatocellular carcinoma. Oncotarget 7, 26168–26180. https://www.ncbi.nlm.nih.gov/pubmed/27034168 (2016).
Bayram, S. et al. Lack of an association of programmed cell death-1 PD1.3 polymorphism with risk of hepatocellular carcinoma susceptibility in Turkish population: a case-control study. Gene 511, 308–313. https://www.ncbi.nlm.nih.gov/pubmed/23041554 (2012).
Zhang, J., Zhao, T., Xu, C., Huang, J. & Yu, H. The association between polymorphisms in the PDCD1 gene and the risk of cancer: A PRISMA-compliant meta-analysis. Medicine (Baltimore). 95, e4423. https://www.ncbi.nlm.nih.gov/pubmed/27749524 (2016).
Huang, C. et al. Association of rs10204525 genotype GG and rs2227982 CC combination in programmed cell death 1 with hepatitis B virus infection risk. Medicine (Baltimore). 98, e16972. https://www.ncbi.nlm.nih.gov/pubmed/31464942 (2019).
Song, l. H. et al. Mannose-binding lectin gene polymorphisms and hepatitis B virus infection in Vietnamese patients. Mutat. Res. 522, 119–125. http://www.ncbi.nlm.nih.gov/pubmed/12517417 (2003).
Bruix, J. & Sherman, M. Management of hepatocellular carcinoma: An update. Hepatology 53, 1020–1022. http://www.ncbi.nlm.nih.gov/pubmed/21374666 (2011).
Hoan, N. X. et al. Interferon-stimulated gene 15 in hepatitis B-related liver diseases. Oncotarget 7, 67777–67787. https://www.ncbi.nlm.nih.gov/pubmed/27626177 (2016).
Gao, J. et al. Meta-analysis of programmed cell death 1 polymorphisms with systemic lupus erythematosus risk. Oncotarget. 8, 36885–36897. https://www.ncbi.nlm.nih.gov/pubmed/28415570 (2017).
Fathi, F. et al. Effects of the programmed cell death 1 (PDCD1) polymorphisms in susceptibility to systemic lupus erythematosus. Int. J. Immunogenet. 47, 57–64. https://www.ncbi.nlm.nih.gov/pubmed/31565862 (2020).
Tahoori, M. T. et al. Association of programmed cell death-1 (PDCD-1) gene polymorphisms with rheumatoid arthritis in Iranian patients. Clin. Exp. Rheumatol. 29, 763–767. https://www.ncbi.nlm.nih.gov/pubmed/21961966 (2011).
Hou, Z., Zhou, Q., Lu, M., Tan, D. & Xu, X. A programmed cell death-1 haplotype is associated with clearance of hepatitis B virus. Ann. Clin. Lab. Sci. 47, 334–343. https://www.ncbi.nlm.nih.gov/pubmed/28667037 (2017).
Li, F. et al. Genetic association and interaction of PD1 and TIM3 polymorphisms in susceptibility of chronic hepatitis B virus infection and hepatocarcinogenesis. Discov. Med. 27, 79–92. https://www.ncbi.nlm.nih.gov/pubmed/30939292 (2019).
Sarvari, J., Dowran, R., Hosseini, S. Y., Fattahi, M. R. & Erfani, N. Association of PD-1 gene with outcome of hepatitis C virus infection. Excli J. 17, 935–944. https://www.ncbi.nlm.nih.gov/pubmed/30564072 (2018).
Hashemi, M. et al. Association between PD-1 and PD-L1 polymorphisms and the risk of cancer: A meta-analysis of case-control studies. Cancers (Basel). 11. https://www.ncbi.nlm.nih.gov/pubmed/31405171 (2019).
Tang, W. et al. Programmed death-1 (PD-1) rs2227981 C > T polymorphism is associated with cancer susceptibility: A meta-analysis. Int. J. Clin. Exp. Med. 8, 22278–22285. https://www.ncbi.nlm.nih.gov/pubmed/26885204 (2015).
Isogawa, M., Chung, J., Murata, Y., Kakimi, K. & Chisari, F. V. CD40 activation rescues antiviral CD8(+) T cells from PD-1-mediated exhaustion. PLoS Pathog. 9, e1003490. https://www.ncbi.nlm.nih.gov/pubmed/23853599 (2013).
Shi, F. et al. PD-1 and PD-L1 upregulation promotes CD8(+) T-cell apoptosis and postoperative recurrence in hepatocellular carcinoma patients. Int. J. Cancer. 128, 887–896. https://www.ncbi.nlm.nih.gov/pubmed/20473887 (2011).
Cheng, H. Y. et al. Circulating programmed death-1 as a marker for sustained high hepatitis B viral load and risk of hepatocellular carcinoma. PLoS One. 9, e95870. https://www.ncbi.nlm.nih.gov/pubmed/25427199 (2014).
Acknowledgements
We thank all study subjects for their participation. This study was funded by the Vietnam National Foundation for Science and Technology Development (NAFOSTED), Grant number: 108.02-2018.315. The author TPV acknowledges the German Federal Ministry of Education and Research for exchange of Vietnamese scientists (BMBF01DP19006A, BMBF01DP17047). The funder has no role in the study design, data collection and analysis, decision to publish or preparation of the manuscript. We expand our acknowledgements to Miss. Ngo Thi Uyen, Hoang Anh Ha and Mr. Bui Dinh Tung for experimental helps.
Author information
Authors and Affiliations
Contributions
N.X.H., L.H.S., T.P.V. designed the study and N.X.H. wrote the first draft. N.X.H., P.T.M.H., B.T.L., D.P.G., S.R.P. performed the experiments. N.X.H. performed the statistical analyses. NXH and TPV interpreted data. N.X.H., L.H.S., M.T.B., P.T.M.H., D.P.G, N.T.T. recruited patients. D.T.N.D. contributed to data management. T.P.V. revised the first draft. M.H.B., L.H.S., P.G.K. and T.P.V. contributed to materials and reagents.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: In the original version of this Article, Peter G. Kremsner was incorrectly affiliated with ‘Vietnamese-German Center for Medical Research (VG-CARE), Hanoi, Vietnam’. Additionally, an affiliation was omitted. The correct affiliations are ‘Institute for Tropical Medicine, University Hospital Tübingen, Tübingen, Germany’ and ‘Centre de Recherches Médicales de Lambaréné (CERMEL), Lambaréné, Gabon’.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hoan, N.X., Huyen, P.T.M., Binh, M.T. et al. Genetic variants of programmed cell death 1 are associated with HBV infection and liver disease progression. Sci Rep 11, 7772 (2021). https://doi.org/10.1038/s41598-021-87537-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-87537-9
This article is cited by
-
Analysis of angiogenesis-related subtypes of hepatocellular carcinoma and tumor microenvironment infiltration feature in hepatocellular carcinoma
Clinical and Translational Oncology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.