Abstract
Bacterial wilt caused by the Ralstonia solanacearum species complex (RSSC) is among the most important plant diseases worldwide, severely affecting a high number of crops and ornamental plants in tropical regions. Only a limited number of phages infecting R. solanacearum have been isolated over the years, despite the importance of this bacterium and the associated plant disease. The antibacterial effect or morphological traits of these R. solanacearum viruses have been well studied, but not their genomic features, which need deeper consideration. This study reports the full genome of 23 new phages infecting RSSC isolated from agricultural samples collected in Mauritius and Reunion islands, particularly affected by this plant bacterial pathogen and considered biodiversity hotspots in the Southwest Indian Ocean. The complete genomic information and phylogenetic classification is provided, revealing high genetic diversity between them and weak similarities with previous related phages. The results support our proposal of 13 new species and seven new genera of R. solanacearum phages. Our findings highlight the wide prevalence of phages of RSSC in infected agricultural settings and the underlying genetic diversity. Discoveries of this kind lead more insight into the diversity of phages in general and to optimizing their use as biocontrol agents of bacterial diseases of plants in agriculture.
Similar content being viewed by others
Introduction
Bacteria from the Ralstonia solanacearum species complex (RSSC) are major plant pathogens, causing bacterial wilt disease in a wide range of important agricultural and wild plant hosts1. Its distribution is worldwide, with special incidence in tropical areas, and detailed epidemiological information is available from many regions2,3. It is considered a bacterial species complex that comprises three species (R. pseudosolanacearum, R. solanacearum and R. syzygii), four genetic groups or phylotypes (I, II, III and IV) and more than 50 monophyletic groups, named sequevars4. Ralstonia solanacearum is included in the A2 (high risk) list of quarantine organisms in Europe and Canada, and some sequevars are listed as bioterrorist agents in the US, because of their potential danger to agriculture. Climate change and international commercial exchanges are increasing the risk of bacterial wilt outbreaks in presently unaffected areas5,6.
Bacteriophages or phages, the viruses of bacteria, are present in all environments and are an important ecological and evolutionary force that shape microbial communities7. It has been demonstrated that R. solanacearum phages can influence plant health or disease. Virulent phages can be used as biocontrol agents to limit the incidence of the disease, as shown recently8. Then, a particular group of phages, the non-lethal and filamentous Inoviridae, can either increase or decrease bacterial virulence, by providing with new genetic repertoire or modifying behaviours such as motility or biofilm formation in bacteria9,10. Several R. solanacearum phages have been studied, but the understanding of their prevalence in nature, or their complete characterisation is still poor.
The complete genome of only 35 R. solanacearum phages are fully classified according to the International Committee on Taxonomy of Viruses (ICTV) guides. This number is low compared, for example, to the 224 phages of Pseudomonas (comprising all bacterial species), or phages infecting human associated bacteria such as the 270 genomes of Escherichia phages. The amount of R. solanacearum phages described is however comparable to phages of the other five main plant pathogenic bacteria with worldwide distribution, with 45 Erwinia, 35 Pectobacterium, 21 Rhizobium, 18 Xanthomonas or 5 Xylella complete phage genomes published so far. The main geographical origin of the published complete R. solanacearum phage genomes is Asia, with 64% of all phages coming from Japan (e.g.11). There are no R. solanacearum phages from European sources among those fully-classified R. solanacearum phage genomes, even if recent work has isolated and tested phages as biocontrol agents of RSSC12. Only one phage genome is available from Africa13, despite that the agriculture of this continent suffers from bacterial wilt disease, and RSSC are present in most tropical areas. Despite the difficulties of analysis on non-conserved highly recombinant phage genomes, to obtain well-annotated genomes of virulent phages is essential for phage applied purposes, ensuring that they do not encode genes associated with toxicity in bacteria, or allowing to assess better the outcome of the treatment14. More genomic and metagenomic data of phages is helping to acquire detailed knowledge on phage ecology and evolution in varied ecosystems15.
New methods of phage genome analysis give the opportunity to undertake deep genetic characterisation of all phages and to do comparative genomics16,17. The ICTV has recently re-organized the taxonomy of phages, providing with guide-lines to classify phages18. The GenBank database includes the updated taxonomy of the 35 R. solanacearum phages. Thirteen are classified as Autographiviridae, eleven as Inoviridae, seven as Myoviridae, two as Ackermanviridae and only one as Siphoviridae and one as Podoviridae. Regarding the non Inoviridae, the Ackermanviridae, Siphoviridae and Podoviridae have not been assigned to genera. Of the remaining 20 Autographiviridae and Myoviridae, they are all comprised in 14 genera. The current study examined R. solanacearum phages from two African islands in the Southwest Indian Ocean, considered biodiversity hotspots: Mauritius and Reunion. The aim was to isolate and characterize R. solanacearum phages from a non-explored geographical region and thus to contribute knowledge on viral diversity. We report the discovery of 23 new R. solanacearum phages, and propose 13 new species and 7 new genera, based on genomic and morphological analyses. We also complete the taxonomic classification of three R. solanacearum phages already described but not fully characterised. We highlight the genetic diversity of these newly isolated phages that contributes considerably to expanding our understanding of R. solanacearum phages.
Methods
Sampling, isolation and purification of phages
Sampling of R. solanacearum phages in this study was carried out at four agricultural sites in Southwestern Reunion Island in February 2018 and at six sites in Mauritius Island in March 2018 (see exact locations and maps in Supplementary Fig. 1). The criteria for site selection was the recent incidence of bacterial wilt disease in solanaceous plants, irrespective of location, host plant species or cultivation method (greenhouse, field, off-ground, organic, etc.). Different types of samples were collected: diseased plant roots, soil and irrigation water. The locations and characteristics of the samples from each of the sites with positive samples are presented in Supplementary Table 1.
An enrichment technique with multiple bacterial strains was used to widen the diversity of phage isolation19. We used the most frequent R. pseudosolanacearum strains founded in each island as well as other strains that represent the genetic diversity of RSSC (Supplementary Table 2). The strains were not mixed in the same tube due to the possible production of bacteriocins by RSSC20. Each agricultural sample was divided and enriched with every strain in separate tubes. Using the overlay assay, single plaques were identified and transferred three times. A RSSC strain from sequevar I-31, the main genetic variant found in Reunion, was used as the common host for subsequent amplification of Reunion Island phage clones3. Each phage clone of Mauritius island was amplified with its corresponding specific sequevar host (I-15, I-31, I-33, see Supplementary Table 1 for details). High titers were produced by infecting 25 mL of the host strains with 0.1 mL of each phage isolate. Following a standard chloroform protocol phages lysates were produced and kept at 4 °C until needed21.
Bacterial strains and culture media
All Ralstonia solanacearum species complex strains used here belong to the reference collection of the Pôle de Protection des Plantes in Reunion Island (France). Details about their phylogenetic classification and origin are provided in Supplementary Table 2. All inoculations and bacterial cultures were carried out in a semi-selective Kelman culture medium containing 1% m/m triphenyltetrazolium chloride. All bacterial cultures were incubated at 28 °C. Liquid cultures were agitated at 80 rpm when they included phages or 120 rpm for bacteria only.
Phage genome extraction and sequencing
Genomic DNA extraction of 81 phages from a high titer phage lysate was done with the Wizard DNA clean-up kit from Promega. Restriction analysis of phage genomic DNA was performed with the enzymes NcoI, PstI and HindIII (Promega), after failed DNA digestion with EcoRI, XbaI et KnpI, and thus phages classified by restriction profile group. One phage per group was selected for whole genome sequencing. These results and the genomic DNA quality guided the selection of 39 phages for whole genome sequencing. DNA phage libraries were prepared with the TruSeq Nano kit (Illumina) for 39 phage genomes and sequenced to 2 × 250 pb reads length using Illumina HiSeq at the sequencing platform GeT-PlaGe of GenoToul (Toulouse, France).
Phage genome assembly and annotation
Reads were trimmed with Sickle 1.33.2 and assembled with SPAdes 3.12.022,23. Coding sequences (CDSs) were predicted and annotated using Prokka 1.14.5, and manual refinement database searches were performed using BLASTp, HHpred and VIRFam to predict head-neck-tail module genes24,25,26,27.
Phage life-style prediction
Temperate phages were identified as in other works28. Briefly all phage genomes were analysed using PHACTS, PHASTER29,30. When the results were not conclusive we did a blast of the aminoacid sequences for the presence of recombinases using PFAM v26 and HMMER3 with default options31,32. These predictions were manually curated.
Phage phylogenetic analysis
Phylogenetic tree based on nucleotides were constructed using VICTOR17. The branch lengths were scaled in terms of the Genome BLAST Distance Phylogeny (GBDP) intergenomic distance formula d0 for nucleotide based VICTOR phylogenetic trees17. A proteomic tree was constructed using the ViPTree server based on genome-wide sequence similarities computed by tBLASTx33.
TEM micrographs
Electron micrographs of R. solanacearum phages were generated as in other studies34. Briefly, the specimen was picked up with 400-mesh copper grids (Delta microscopies, Mauressac, French), negative stained with 1% (W/V) ammonium molybdate, and analysed using a HT7800 Hitachi transmission electron microscope (Hitachi, Tokyo, Japan) at an acceleration voltage of 80 kV. Electron micrographs were taken with an AMT XR401, sCMOS-camera (AMT imaging, Woburn, MA-US).
Results and discussion
Sampling and geographical distribution of phages
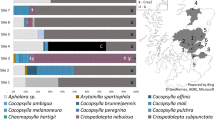
R. solanacearum phages were broadly present in agricultural material associated with bacterial wilt from Reunion (76%, N = 26), and Mauritius (56%, N = 32) (Fig. 1a, Supplementary Fig. 1, Supplementary Table 2). There were no significant differences in the proportion of positive samples between the two Southwest Indian Ocean islands (quasi-Poisson GLM: F = 1.887, d.f. = 1, p-value = 0.303, Fig. 1a). Samples came from greenhouses or fields, with no differences between these agricultural settings in terms of phage abundance (quasi-Poisson GLM: F = 0.06, d.f. = 1, p-value = 0.830, Fig. 1a). R. solanacearum phages were found in around 50% or more of all types of samples, from plants, soil from uprooted fields and water (Fig. 1b). The highest proportion of positive samples were observed in aubergine (Solanum melongena), pepper (Capsicum annum), and irrigation water samples (Fig. 1b).
Not all bacterial strains were susceptible to the phages, as revealed by the subset of RSSC strains that were attacked by the phages in the enrichment step of the phage isolation (Supplementary Table 1). Only strains from phylotype I resulted in the isolation of phages, a result likely derived from the epidemiology of this bacterium in the islands, where this is the main RSSC genetic type present, and the specificity and local adaptation of phages3. After purification and differentiation of individual lysis plaques, a total of 35 and 46 individual phages were isolated from Reunion and Mauritius, but only 17 and 21 phages were sequenced, respectively, after discarding identical DNA digestion profiles through analysis with different restriction enzymes. Sequencing allowed the identification of ten different phage genomes from Reunion and 13 from Mauritius, and the rest were identical clones (Table 1). We named Mauritius phages after historical cyclones that hit the island, and Reunion ones after legendary anti-colonialist slaves. We included a gender balance criteria in the choice of phage names. The most abundant phage clones were Ralstonia phage Bakoly (Ralstonia virus Bakoly) found five times in Mauritius, and Ralstonia phage Cimandef (Ralstonia virus Cimandef) found three times in Reunion, in different host plants (Supplementary Table 1).
As it will be detailed in the next section, the 23 phages of the study represent 13 new species, within seven new genera which belong to four different phage families. Three genera out of the seven described were found in both islands, one was so far endemic of Reunion and one of Mauritius, and two contained Mauritian and foreign phages. Five different phage species were distinguished in Reunion, whereas eight different ones were found in Mauritius. Most phage species were geographically restricted to one of the islands, except Ralstonia virus Raharianne, that was found in both Mauritius and Reunion. Even for this shared phage, Ralstonia virus Raharianne, clones were unique from each island, with two representatives in Reunion (Ralstonia phage Raharianne and Ralstonia phage Albius) and one in Mauritius (Ralstonia phage Hyacinthe).
The sampling design of this study was not exhaustive enough to present general conclusions on the geographical distribution of R. solanacearum phages in the islands, or the association with specific host plants and agricultural practices, but the results support the presence of R. solanacearum phages in all agricultural environments where bacterial wilt was recorded. We provide an important collection of R. solanacearum phages that considerably widens its known genetic diversity. Mauritius phages are more diverse, with more genera and species, compared to Reunion ones in this collection, a non surprising fact, in light of the more complex and spatially structured RSSC epidemiology in Mauritius3. All clones are only located in one of the islands, suggesting an evolutionary process of local adaptation, although some genera are shared, indicating a phylogenetic relationship of R. solanacearum phages within the Southwest Indian Ocean, likely linked to the related bacterial epidemiology3.
Taxonomic analysis and support for the description of seven new R. solanacearum phage genera
The 23 phages of our study represent 13 new species, within seven new genera, according to the scored phylogenomic distances calculated using VIRIDIC (Virus Intergenomic Distance Calculator), which integrates the ICTV criteria (Table 2). Four genera (Anchaingvirus, Dinavirus, Firingavirus and Rahariannevirus) include only one phage species. The genera Bakolyvirus and Gervaisevirus include two new R. solanacearum phage species each, and five new phage species belong to the Cimandefvirus genus, being the most diverse one of the seven (Table 2). The BLASTn analysis of the 23 phages isolated in this study did not reveal any significant genetic similarities to other phages in databases (last accessed in July 2020) at the nucleotide level (< 75%, ICTV criterion), except for 3 R. solanacearum phages sampled in China and Japan that were relatively similar to ours: Ralstonia phage RPZH6, Ralstonia phage GP4 and Ralstonia phage RSK1, (Supplementary Table 3)35. These three phages have not been completely taxonomically classified and, according to the nucleotide similarity with the presently studied phages, we suggest to include them in the new genera Gervaisevirus (RPZH6 and GP4) and Firingavirus (RSK1).
Phylogenetic trees using the full genome of the new 23 phages and including nine closest relatives provide additional support to the genera classification by VIRIDIC, with highly consistent branches (higher or equal to 96 bootstrap replicates) (Fig. 2a). The closest viruses, included in this tree, are four R. solanacearum phages, four Burkholderia phages and one Halomonas phage isolated from China, Japan and the USA35,36,37,38. Taxonomically, Burkholderia and Ralstonia bacteria are from different genus (Ralstonia and Burkholderia), but the same family (Burkholderiaceae), and thus the distant relationship of phages that attack them is not unexpected. However, Halomonas bacteria are classified as Gammaproteobacteria, a different bacterial class than the one assessed to Ralstonia (Betaproteobacteria). Yet, the genetic homology between the related Halomonas phage and our R. solanacearum phages is only 4% (Supplementary Table 3). Just two complete genomes of Halomonas phages have been described so far, according to GenBank records. The proteomic tree generated with ViPTree confirmed the taxonomic relationships among the R. solanacearum phages of the present study, as well as their genomic uniqueness compared to previously described R. solanacearum, Burkholderia and Halomonas phages (Fig. 2b). This may be explained by the geographical distance of the Southwest Indian Ocean islands from previous phage samples, the genetic diversity of the RSSC and overall biodiversity present. The high diversity and quantity of new phage genera and species described in our study is all the same surprising.
Phylogenetic trees based on the full genome of the new 23 R. solanacearum phages (in red those from Reunion island and in blue the ones from Mauritius). (a) Tree based in nucleotides including also the nine closest phage relatives (in black), calculated with VICTOR methods. The seven proposed genera are indicated by coloured dots on the left side of the tree. A bootstrap value of 100 was used in the analysis. (b) Circular proteomic tree generated by ViPTree including also all R. solanacearum (non-Inoviridae and non-jumbo), Burkholderia and Halomonas phages, coloured by virus families and host taxonomic groups. The new 23 R. solanacearum phages are indicated by red stars.
The electron micrographs confirm the Virfam genetic-based predictions of phage families in this phage collection, that is, the seven genera are classified as four Podoviridae, one Siphoviridae and two Myoviridae25 (Fig. 3). The presence of RNA and DNA polymerases in the genome of Ralstonia virus Anchaing revealed that this Podoviridae is in fact an Autographiviridae39. Surprisingly, only one Podoviridae and one Siphoviridae had been described before amongst the previous R. solanacearum phage species. In the present R. solanacearum phage collection we assess nine species as new Podoviridae, one new Siphoviridae and one new Autographiviridae R. solanacearum phage (Table 1, Fig. 3).
Transmission electron micrographs of the representative R. solanacearum phage species of five new genera with unique viruses (non described before): (a) Ralstonia phage Cimandef (Cimandefvirus, Podoviridae); (b) Ralstonia phage Anchaing (Anchaingvirus, Autographiviridae); (c) Ralstonia phage Raharianne (Rahariannevirus, Myoviridae); (d) Ralstonia phage Bakoly (Bakolyvirus, Myoviridae); (e) Ralstonia phage Dina (Dinavirus, Siphoviridae).
Genomic comparison of the seven new genera
The average genome size of the 23 Southwest Indian Ocean phages was 50,108 bp, the average GC content 63%, and the average number of encoded proteins was 65. Compared to the genome size of 22 other non-Inoviridae and non-jumbo (giant) R. solanacearum phages already described (41,536 bp), the present phage collection average genome size is bigger. The biggest genome size in the present collection was the 61,164 bp of Ralstonia phage Gervaise, and the smallest genome was that of Ralstonia phage Dina, with 39,161 bp. This genus, Dinavirus had also the highest GC content (65.27%). This is consistent with previous analysis that correlate GC % with low genome size in phages40. It was not the case of the two phage species of the genus Firingavirus, with small genome size (39,610 bp) and the lowest GC % (59.56) of the entire collection. As seen for other phages, the average GC% of all 23 phages is lower than that of the bacterial host, which has a median GC% of 66.840. As for the number of proteins encoded, there was variability within the new phage genera, and overall it ranged from the 51 of Ralstonia phage Firinga to the 78 of Ralstonia phage Gerry.
The ability of phages to perform lysogenic life-cycles (temperate phages) is not always elucidated or discussed in genomic or phenotypic studies. From the seven R. solanacearum phage genera studied here, three of them are probably composed by temperate phages: Dinavirus, Anchaingvirus and Gervaisevirus, as suggested by the presence of a gene encoding a recombinase in their genome. The lack of experimental proof of the integrase activity in these or the homologous genes in current databases, precludes more solid conclusions. Phages from the other four genera are with high probability virulent viruses, thus only capable of performing lytic cycles. This implies a significant (26%) proportion of R. solanacearum phages potentially able to undergo a lysogenic cycle, but a higher proportion of mostly virulent phages and therefore adapted for biocontrol purposes. Interestingly, from the four temperate phage species, three were sampled in Mauritius and only one in Reunion island (Ralstonia virus Anchaing). One potential cause of the abundance of temperate phages in Mauritius is the tight molecular interactions and genetic exchanges with their hosts, related to the more complex RSSC epidemiology in this island3. In Reunion, a majority of virulent phages may have faster effects for the more homogeneous RSSC bacterial populations. Further surveys and experiments would be needed to test these ideas.
Finally, substantial genomic homology can be observed between the reference phages of the genera Cimandefvirus and Gervaisevirus, really weak between Gervaisevirus, Rahariannevirus and Firingavirus, and completely absent for Dinavirus, Anchaingvirus and Bakolyvirus (Fig. 4). The absence of synteny provides further evidence of the non-conserved genomic architecture between this particular group of R. solanacearum phages (Fig. 4).
Conclusions
There is a substantial imbalance in virtually all host–parasite interactions, with multiple parasites attacking the same host species. This is the case for bacteria and phages, but the knowledge and amount of isolated or sequenced bacteria surpass that of their natural enemies, bacteriophages41. This study reports a wide diversity of newly isolated R. solanacearum phages in two islands of the Southwest Indian Ocean, and demonstrates the prevalence of the parasites of this bacterium in agricultural environments affected by bacterial wilt disease. Only 35 phages of R. solanacearum have been previously fully analysed at the genomic level. Earlier phages were mostly sampled in eastern Asiatic countries, and other geographic origins have been largely disregarded. The 20 non-Inoviridae R. solanacearum phages previously known are classified into fourteen genera, revealing a high taxonomic variety. Genetic differences between our phages and other fully-sequenced R. solanacearum phages reported in current databases are substantial, in most cases characterized by the complete absence of genetic homology. We identify nonetheless, three phages belonging to the newly defined genera. Following the ICTV guidelines, we propose seven new genera of R. solanacearum phages, which include twenty-three new phage clones within thirteen new species. A small proportion of the isolated phages are probably temperate, and specially abundant in Mauritius. In this island, phage genetic diversity is significantly higher than in Reunion. Taken together, these results suggest different phage-bacteria dynamics and evolutionary history of R. solanacearum phages in each island, which deserve additional investigations.
This work is singular for many reasons: the quantity and diversity of new R. solanacearum phages presented, their unprecedented geographical origin, the complete disclosure of the sampling results (including negative results, nature of the samples and exact location), and the deep genomic analysis characterisation. Our work expands substantially the existing R. solanacearum phage records and points in the direction that there is much more phage genetic diversity waiting to be discovered in agricultural ecosystems. Furthermore, broad comprehension of the genetic features of phages and their association with their bacterial hosts’ local epidemiology is essential to fine-tune biocontrol strategies based on phages against bacterial pathogens.
Data availability
Assembled and annotated genomes of the 23 R. solanacearum phages were uploaded to GenBank under accession numbers MT740725 to MT740747. All data generated or analysed during this study are included in this published article (and its “Supplementary Information files”).
References
Mansfield, J. et al. Top ten plant pathogenic bacteria in molecular plant pathology. Mol. Plant Pathol. 13, 614–629 (2012).
Bhunchoth, A. et al. Isolation of Ralstonia solanacearum-infecting bacteriophages from tomato fields in Chiang Mai, Thailand, and their experimental use as biocontrol agents. J. Appl. Microbiol. 118, 1023–1033 (2015).
Yahiaoui, N. et al. Genetic diversity of the Ralstonia solanacearum species complex in the Southwest Indian Ocean Islands. Front. Plant Sci. 8, 2139 (2017).
Prior, P. et al. Genomic and proteomic evidence supporting the division of the plant pathogen Ralstonia solanacearum into three species. BMC Genom. 17, 90 (2016).
Bergsma-Vlami, M. et al. Phylogenetic Assignment of Ralstonia pseudosolanacearum (Ralstonia solanacearum Phylotype I) Isolated from Rosa spp. Plant Dis. 102, 2258–2267 (2018).
Norman, D. J., Yuen, J. M. F. & Bocsanczy, A. M. Threat of brown rot of potato and existing resistance. Am. J. Potato Res. 97, 272–277 (2020).
Koskella, B. & Taylor, T. B. Multifaceted impacts of bacteriophages in the plant microbiome. Annu. Rev. Phytopathol. 56, 361–380 (2018).
Wang, X. et al. Phage combination therapies for bacterial wilt disease in tomato. Nat. Biotechnol. https://doi.org/10.1038/s41587-019-0328-3 (2019).
Addy, H. S., Askora, A., Kawasaki, T., Fujie, M. & Yamada, T. Utilization of filamentous phage φRSM3 to control bacterial wilt caused by Ralstonia solanacearum. Plant Dis. 96, 1204–1209 (2012).
Addy, H. S., Askora, A., Kawasaki, T., Fujie, M. & Yamada, T. The filamentous phage φrSS1 enhances virulence of phytopathogenic Ralstonia solanacearum on tomato. Phytopathology 102, 244–251 (2012).
Fujiwara, A., Kawasaki, T., Usami, S., Fujie, M. & Yamada, T. Genomic characterization of Ralstonia solanacearum phage phiRSA1 and its related prophage (phiRSX) in strain GMI1000. J. Bacteriol. 190, 143–156 (2008).
Álvarez, B., López, M. M. & Biosca, E. G. Biocontrol of the major plant pathogen Ralstonia solanacearum in Irrigation water and host plants by novel waterborne lytic bacteriophages. Front. Microbiol. https://doi.org/10.3389/fmicb.2019.02813 (2019).
Ahmad, A. A. et al. Sequencing, genome analysis and host range of a novel Ralstonia phage, RsoP1EGY, isolated in Egypt. Arch. Virol. 163, 2271–2274 (2018).
Philipson, C. et al. Characterizing phage genomes for therapeutic applications. Viruses 10, 188 (2018).
Dion, M. B., Oechslin, F. & Moineau, S. Phage diversity, genomics and phylogeny. Nat. Rev. Microbiol. 18, 125–138 (2020).
Moraru, C., Varsani, A. & Kropinski, A. M. VIRIDIC—A novel tool to calculate the intergenomic 1 similarities of prokaryote-infecting viruses 2. bioRxiv. https://doi.org/10.1101/2020.07.05.188268 (2020).
Meier-Kolthoff, J. P. & Göker, M. VICTOR: Genome-based phylogeny and classification of prokaryotic viruses. Bioinformatics 33, 3396–3404 (2017).
Adriaenssens, E. M. & Rodney Brister, J. How to name and classify your phage: An informal guide. Viruses 9, 70 (2017).
Ross, A., Ward, S. & Hyman, P. More is better: Selecting for broad host range bacteriophages. Front. Microbiol. 7, 1352 (2016).
Huerta, A. I., Milling, A. & Allen, C. Tropical strains of Ralstonia solanacearum outcompete race 3 biovar 2 strains at lowland tropical temperatures. Appl. Environ. Microbiol. 81, 3542–3551 (2015).
Betts, A., Vasse, M., Kaltz, O. & Hochberg, M. E. Back to the future: Evolving bacteriophages to increase their effectiveness against the pathogen Pseudomonas aeruginosa PAO1. Evol. Appl. 6, 1054–1063 (2013).
Joshi, N. & Fass, J. Sickle: A sliding-window, adaptive, quality-based trimming tool for FastQ files (Version 1.33) [Software]. https://github.com/najoshi/sickle (2011).
Bankevich, A. et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. https://doi.org/10.1089/cmb.2012.0021 (2012).
Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Lopes, A., Amarir-Bouhram, J., Faure, G., Petit, M. A. & Guerois, R. Detection of novel recombinases in bacteriophage genomes unveils Rad52, Rad51 and Gp2.5 remote homologs. Nucleic Acids Res. 38, 3952–3962 (2010).
Zimmermann, L. et al. A completely reimplemented MPI bioinformatics toolkit with a new HHpred server at its core. J. Mol. Biol. https://doi.org/10.1016/j.jmb.2017.12.007 (2018).
Bateman, A. UniProt: A worldwide hub of protein knowledge. Nucleic Acids Res. https://doi.org/10.1093/nar/gky1049 (2019).
Touchon, M., Bernheim, A. & Rocha, E. P. Genetic and life-history traits associated with the distribution of prophages in bacteria. ISME J. 10, 2744–2754 (2016).
McNair, K., Bailey, B. A. & Edwards, R. A. PHACTS, a computational approach to classifying the lifestyle of phages. Bioinformatics 28, 614–618 (2012).
Arndt, D. et al. PHASTER: A better, faster version of the PHAST phage search tool. Nucleic Acids Res. 44, W16–W21 (2016).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Finn, R. D. et al. The Pfam protein families database. Nucleic Acids Res. 36, D281–D288 (2008).
Nishimura, Y. et al. ViPTree: The viral proteomic tree server. Bioinformatics 33, 2379–2380 (2017).
Ackermann, H. W. Basic phage electron microscopy. Methods Mol. Biol. 501, 113–126 (2009).
Wang, R. et al. Characterization and complete genome sequence analysis of phage GP4, a novel lytic Bcep22-like podovirus. Arch. Virol. 164, 2339–2343 (2019).
Fu, C. et al. Complete genome sequence of Halomonas ventosae virulent halovirus QHHSV-1. Arch. Virol. 162, 3215–3219 (2017).
Gill, J. J. et al. Genomes and characterization of phages Bcep22 and BcepIL02, founders of a novel phage type in Burkholderia cenocepacia. J. Bacteriol. 193, 5300–5313 (2011).
Summer, E. J. et al. Divergence and mosaicism among virulent soil phages of the Burkholderia cepacia complex. J. Bacteriol. 188, 255–268 (2006).
Lavigne, R., Seto, D., Mahadevan, P., Ackermann, H. W. & Kropinski, A. M. Unifying classical and molecular taxonomic classification: Analysis of the Podoviridae using BLASTP-based tools. Res. Microbiol. 159, 406–414 (2008).
Almpanis, A., Swain, M., Gatherer, D. & McEwan, N. Correlation between bacterial G+C content, genome size and the G+C content of associated plasmids and bacteriophages. Microb. Genom. 4, e000168 (2018).
Haerter, J. O., Mitarai, N. & Sneppen, K. Phage and bacteria support mutual diversity in a narrowing staircase of coexistence. ISME J. 8, 2317–2326 (2014).
Acknowledgements
We thank farmers of Reunion and Mauritius islands for their help with the sampling and their enthusiastic interest in the project. We are thankful to Evelien Adriaenssens, Andrew Kropinsky and members of the Phages.fr network, for helpful discussions on phage genomic analysis and taxonomy. This work was supported by the microscopy facilities of the Platform 3A, funded by the European Regional Development Fund, the French Ministry of Research, Higher Education and Innovation, the Provence-Alpes-Côte d'Azur region, the Departmental Council of Vaucluse and the Urban Community of Avignon. We are grateful to the INRAE MIGALE bioinformatics facility (Migale bioinformatics Facility, https://doi.org/10.15454/1.5572390655343293E12) for providing computing and storage resources. This work was co-funded by the European Union: European Regional Development Fund (ERDF), by the Conseil Régional de La Réunion and by the Centre de Coopération Internationale en Recherche Agronomique pour le Développement (CIRAD). The authors acknowledge the Plant Protection Platform (3P, IBISA).
Author information
Authors and Affiliations
Contributions
C.T.B. conceived the study; M.J.B.P. guided sampling in Mauritius; C.T.B. and C.B. isolated phages and extracted their DNA; A.T., C.S. and C.T.B. performed bioinformatic analyses; I.B. performed TEM analysis; C.T.B. wrote the manuscript. All authors revised and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Trotereau, A., Boyer, C., Bornard, I. et al. High genomic diversity of novel phages infecting the plant pathogen Ralstonia solanacearum, isolated in Mauritius and Reunion islands. Sci Rep 11, 5382 (2021). https://doi.org/10.1038/s41598-021-84305-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-84305-7
This article is cited by
-
Isolation and Characterization of Novel Lytic Bacteriophage vB_RsoP_BMB50 infecting Ralstonia solanacearum
Current Microbiology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.