Abstract
Fibroblast growth factor receptor 2 (FGFR2) has two isoforms: IIIb type and IIIc type. Clinicopathologic significance of these two FGFR2 subtypes in gastric cancer remains to be known. This study aimed to clarify the clinicopathologic difference of FGFR2IIIb and/or FGFR2IIIc overexpression. A total of 562 patients who underwent gastrectomy was enrolled. The expressions of FGFR2IIIb and FGFR2IIIc were retrospectively examined by immunohistochemistry or fluorescence in situ hybridization (FISH) using the 562 gastric tumors. We evaluated the correlation between clinicopathologic features and FGFR2IIIb overexpression and/or FGFR2IIIc overexpression in gastric cancer. FGFR2IIIb overexpression was observed in 28 cases (4.9%), and FGFR2IIIc overexpression was observed in four cases (0.7%). All four FGFR2IIIc cases were also positive for FGFR2IIIb, but not in the same cancer cells. FGFR2IIIb and/or FGFR2IIIc overexpression was significantly correlated with lymph node metastasis and clinical stage. Both FGFR2IIIb and FGFR2IIIc were significantly associated with poor overall survival. A multivariate analysis showed that FGFR2IIIc expression was significantly correlated with overall survival. FISH analysis indicated that FGFR2 amplification was correlated with FGFR2IIIb and/or FGFR2IIIc overexpression. These findings suggested that gastric tumor overexpressed FGFR2IIIc and/or FGFR2IIIb at the frequency of 4.9%. FGFR2IIIc overexpression might be independent prognostic factor for patients with gastric cancer.
Similar content being viewed by others
Introduction
Fibroblast growth factor receptor 2 (FGFR2) is one of four FGFRs that encode a transmembrane receptor tyrosine kinase1. FGFR2 signaling is associated with the proliferation, migration, and angiogenesis of carcinomas2,3. FGFR2-overexpressed cancer might be a therapeutic target4,5,6. Several FGFR2 inhibitors have been developed for the treatment of cancer patients with enhanced expression of FGFR2 signaling7,8,9.
Gastric cancer is one of the most common malignancies in the world, and the prognosis of advanced gastric cancer remains to be poor10. The aberrant regulation of FGFR2 pathway has been implicated in anti-apoptosis, drug resistance, and epithelial-to-mesenchymal transition (EMT)8. FGFR2 amplification has been reported in several types of solid carcinoma11,12,13 including gastric cancer14,15,16. We previously reported that the conditioned medium derived from gastric fibroblasts stimulates the growth of gastric cancer cells, which are mediated by FGF7/FGFR2 signaling17,18. Gastric cancers with FGFR2 amplification, which was observed in 3–10% of all gastric cancers19,20,21,22,23,24, has been found to be associated with malignant progression25,26,27. Since FGFRs have an important role in the progression of gastric cancer, their use as a therapeutic candidate for the development of targeted anticancer agents should attract substantial attention6,8,28, while no clinical FGFR inhibitor has been approved. One of the reasons for the no clinical approvement of FGFR inhibitors for gastric cancer treatment might be a lack of tool that allows optimal patient selection. The establishment of beneficial markers is necessary to select for FGFR-targeted therapy2,29.
To date, the molecular heterogeneity has been gradually elucidated30,31. Following the understanding of gastric cancer biology, targeted therapies have been evaluated in experimental studies and transferred promptly to clinical trials. FGFR2 has two isoforms, i.e., the IIIb type and the IIIc type based on the alternative splicing within the C-terminal half of the third Ig loop (D3) in the extracellular FGF binding domain, which are alternatively spliced by exon 8 and by exon 9, respectively1,7,32. The spliced isoforms differ in binding ligands: FGFR2IIIb is a high-affinity receptor for FGF1, -3, -7, -10, and -22, whereas FGFR2IIIc binds FGF1, -2, -4, -6, -8, -9 -17 and -1833,34. The FGFR2IIIb isoform is expressed mainly in epithelial cells, and it preferentially binds secreted FGF ligands from adjacent mesenchymal cells33. In contrast, the FGFR2IIIc isoform is preferentially expressed in mesenchymal cells and usually binds ligands secreted from the adjacent epithelial cells35. These findings suggested that the clinical significance of FGFR2 signaling in cancer might differ between FGFR2IIIb and FGFR2IIIc.
There are a few reports of FGFR2IIIb and FGFR2IIIc expression in some other types of solid cancers including gastric carcinomas36,37,38. In this study, we conducted to examine the clinicopathologic significance of the expression of FGFR2IIIb and that of FGFR2IIIc using a large sample of gastric cancer.
Results
FGFR2IIIb-positive and FGFR2IIIc-positive patients
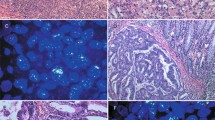
FGFR2IIIb and FGFR2IIIc were mainly expressed at the cell membrane of the cancer cells (Fig. 1A). Among the 562 gastric cancers, FGFR2IIIb and FGFR2IIIc were positive in 28 cases (4.9%) and four cases (0.7%), respectively. All four FGFR2IIIc-positive tumors were also positive for FGFR2IIIb in the same tumor, but both were not positive at the same cancer cells. Case 1 and Case 2 showed heterogeneous expression of both FGFR2IIIb isoform and FGFR2IIIc isoform in a primary tumor (Fig. 1B). In contrast, most of FGFR2-positive tumors were positive for FGFR2IIIb but not FGFR2IIIc (Case 3). The relationships between the clinicopathologic features and the FGFR2IIIb expression or FGFR2IIIc expression were summarized in Table 1. There was a significant correlation between FGFR2IIIb expression and Borrmann's type 4 (p < 0.001), undifferentiated type, depth of invasion (p < 0.001), lymph node metastasis (p < 0.001), lymphatic invasion (p < 0.001), venous invasion (p = 0.015), and clinical stage (p < 0.001). FGFR2IIIc expression was significantly correlated with lymph node metastasis (p = 0.039) and clinical stage (p = 0.014).
Representative pictures of FGFR2IIIb and FGFR2IIIc expression in gastric cancer. (A) IHC score of FGFR2IIIb and FGFR2IIIC. FGFR2IIIb and FGFR2IIIc were mainly expressed at cell membrane (arrows and inset). The immunoreactivity of FGFR2IIIb and FGFR2IIIc were evaluated according to the intensity of membranous staining score 0–3, as follows: 0, negative; 1 +, weak at cytosol; 2 +, moderate at cell membrane; 3 +, strong at cell membrane. FGFR2IIIb and FGFR2IIIc expression were considered positive when scores were ≥ 2. Bar = 100 μm. (B) Representative cases stained with H&E, FGFR2IIIb, FGFR2IIIc, pan-FGFR2, and CEA of gastric cancer. Heterogeneous expressions of FGFR2IIIb isoform and FGFR2IIIc isoform were found in a tumor (Case 1 and Case 2). The pan-FGFR2 detected both FGFR2IIIb isoforms and FGFR2IIIc isoforms. FGFR2IIIb expression (arrows) and FGFR2IIIc expression (asterisks) were positive in the same tumor, but both were not positive in the same tumor cells. FGFR2IIIc dominant expression in a metastatic lymph node disease was found in Case 1. CEA determined cancer cells in the primary tumor and the lymph node. In contrast, Case 3 shows FGFR2IIIb expression, but not FGFR2IIIc expression. × 100. Bar = 200 μm.
FGFR2IIIb and FGFR2IIIc expression on lymph nodes
Among the 28 gastric cancers with FGFR2IIIb and/or FGFR2IIIc expression, 25 cases had lymph node metastasis. FGFR2IIIb expression on lymph nodes was observed in 10 (40%) of the 25 cases with lymph node metastasis. FGFR2IIIc expression on lymph nodes was observed in 4 (16%) of the 25 cases, and the 4 cases were the same cases with FGFR2IIIc expression at the primary tumor. All four cases with FGFR2IIIc expression were also positive for FGFR2IIIb expression in lymph nodes, but both were not positive at the same cancer cells, which was as well as the primary tumors. In Case 1, FGFR2IIIc dominant expression was found in a metastatic lymph node (Fig. 1B).
FGFR2 gene amplification analyzed by FISH
A total of 33 gastric cancer including 28 FGFR2-positive tumors and five FGFR2-negative tumors were examined FGFR2 amplification by FISH. Figure 2 Eleven (39.3%) of the 28 FGFR2-IIIb positive cases had FGFR2 gene amplification: FGFR2 amplification was observed in nine of the 11 cases with an immunohistochemistry score of 3 +, two of the 17 cases with a score of 2 +, and none of the five cases with a score of 1 + or 0. FGFR2/CEN10 ratio of cancer cells with FGFR2IIIb and/or FGFR2IIIc overexpression was significantly (p < 0.01) correlated to that of IHC score by immunohistochemical staining (Table 2). All four cases with FGFR2IIIc expression had FGFR2 amplification. The average FGFR2/CEN10 ratio was 9.1 ± 4.8 in the 11 cases of immunohistochemistry score 3 + , 1.8 ± 2.3 in the 17 cases of score 2 +, and 1.0 ± 0.081 in the five cases with score 1 + and/or 0.
Correlation between FGFR2 expression/amplification and the patients' survival
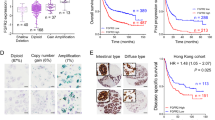
The overall survival and disease-specific survival of the gastric cancer patients with FGFR2IIIb and/or FGFR2IIIc expression (n = 28) was significantly poorer than that of the patients with FGFR2-negative expression (n = 534) (p = 0.008 and 0.002, respectively) (Fig. 3A,B). Among the 28 patients with an FGFR2-positive tumor, the survival rate of the patients who were positive for FGFR3IIIc (n = 4) was significantly poorer than that of FGFR2IIIc-negative patients (n = 24) (p = 0.003, log-rank; Fig. 3C). FGFR2 amplification cases (n = 11) tended to be associated with a poorer outcome (log-rank test, p = 0.095) compared to the non-amplified cases (n = 22) (Fig. 3D).
Survival curve. (A) Overall survival of patients with FGFR2IIIb-positive tumor was significantly poorer (p = 0.008) than that of patients with FGFR2IIIb-negative tumor. (B) Disease specific survival of patients with FGFR2IIIb-positive tumor was significantly poorer (p = 0.002) than that of patients with FGFR2IIIb-negative tumor. (C) In patients with FGFR2-positive tumor, overall survival of patients with FGFR2IIIc-positive tumor was significantly (p = 0.003) poorer than that with FGFR2IIIc-negative. (D) FGFR2 amplification tended to be associated with a poorer outcome, compared with non-amplified cases (p = 0.095).
The univariate analysis revealed that the overall survival was significantly correlated with FGFR2IIIb expression, FGFRIIIc expression, invasion depth, macroscopic type, histological type, lymph node metastasis, peritoneal dissemination, lymphatic invasion, and cytology. Among these clinicopathologic factors, the multivariate analysis revealed that FGFR2IIIc expression (p = 0.045), macroscopic type (p < 0.001), invasion depth (p = 0.003), lymph node metastasis (p < 0.001), hepatic metastasis (p = 0.005), and peritoneal dissemination (p < 0.001) was significantly correlated with overall survival (Table 3).
Discussion
FGFR2 has two isoforms, IIIb type and the IIIc type, which are alternatively spliced by exon 8 and by exon 9, respectively39. In this study, we used two new mouse-rat chimeric monoclonal antibodies, mFR2-10b and mFR2-28c, which recognize human cells with FGFR2IIIb overexpression and FGFR2IIIc overexpression, respectively. Also, we compared the IHC results for both FGFR2IIIb and FGFR2IIIc isoforms using an available pan-FGFR2 antibody, which supported specificity of protein isoform expression. Both antibodies stained the cell membrane of the cancer cells with FGFR2 amplification. These findings suggested that the two new recombinant chimeric antibodies, mFR2-10b against FGFR2IIIb and mFR2-28c against FGFR2IIIc, might be useful for the determination of carcinomas with FGFR2IIIb isoform and FGFR2IIIc isoform. The scoring system of FGFR2IIIb and FGFR2IIIc immunoreactivity were evaluated according to the intensity of membranous staining in the deepest level of the tumor cells. Therefore, scoring system will allow reporting possible heterogeneity of FGFR IIIb and IIIc in a case, but not in the individual tumor cells.
FGFR2, one of the known driver genes of solid tumors, has been a therapeutic target for gastric cancer9,40,41. We also have reported that an FGFR2 phosphorylation inhibitor is useful for the treatment of gastric cancer with FGFR2 amplification18,41,42. Although a reliable biomarker is desired to identify the gastric cancer patients who are candidates for FGFR2 targeting therapy, however suitable FGFR2 antibodies that are specific for the FGFR2IIIb and FGFR2IIIc types are not yet available, so far. We herein obtained that FGFR2IIIb and FGFR2IIIc antibodies specifically bind to the cell membrane of cancer cells. In human gastric carcinomas, FISH analysis results also indicated that FGFR2 amplification was correlated with FGFR2IIIb overexpression and/or FGFR2IIIc overexpression. In this study, our two antibodies, FGFR2IIIb and FGFR2IIIc, might be useful tool for the determination of carcinomas with FGFR2 overexpression and treatment of FGFR2 inhibitors. Analyses of FGFR2 expression using these antibodies may be reliable for the identification of patients who are appropriate for FGFR2 targeting therapy.
Our present analyses demonstrated that the overexpression of FGFR2IIIb and/or FGFR2IIIc was 4.9% in 562 gastric cancers. In human gastric carcinomas, FISH analysis results also indicated that FGFR2 amplification was correlated with FGFR2IIIb overexpression and/or FGFR2IIIc overexpression. Several immunohistochemical studies previously reported that FGFR2 overexpression ranged from 10 to 51% of gastric cancer cases43,44,45,46. FGFR2 amplification has been reported to be observed in 3%–10% of gastric cancers20,21,22,23,24. Currently, several studies have defined the genomic landscape of GC using the next-generation sequencing (NGS) technology47,48,49,50,51, and indicated that the genetic alternations of FGFR2 is around 3–10%50. These findings might suggest that FGFR2 overexpression might be present in around 5% of all gastric cancers, which is lower than previous reports.
Since FGFR2 has two isoforms, IIIb type and IIIc type, it is important to examine the frequency of FGFR2-IIIc expression among FGFR2 overexpressing gastric cancers. However few reports of FGFR2 subclass of IIIb and IIIc expression are available in gastric cancer36. In this study, the overexpression of FGFR2IIIb was 4.9% and that of FGFR2IIIc was 0.7% in 562 gastric cancers. FGFR2IIIb-positive expression was frequently observed in the cases with Borrmann's type 4, undifferentiated type, high invasion depth, lymph node metastasis, lymphatic invasion, venous invasion, and clinical stage. The prognosis of the FGFR2IIb-positive patients was significantly poorer compared to that of the patients with FGFR2IIIb-negative cancer. These findings may indicate that FGFR2IIIb signaling is associated with the malignant progress of gastric cancer. In contrast, no report of FGFR2IIIc expression in gastric cancer is available, whereas It has been reported that FGFR2IIIc immunoreactivity was expressed in 49% and 27% in endometrial endometrioid carcinoma52 and colorectal carcinoma12,37. The expression rate of FGFR2IIIc in 562 gastric cancer cases was only 0.7%, suggesting that gastric cancer expressed FGFR2IIIc extremely few in compared to the expression rate of FGFR2IIIb. These findings suggested that FGFR2IIIb signaling inhibition might be more wide indication for GC patients bearing FGFR2.
Several FGFR2 tyrosine kinase inhibitors (TKIs), including AZD4547 (NCT01457846, SHINE trial), dovitinib (NCT01719549), E7090 (NCT02275910), and TAS-120 (NCT02052778), have been evaluated for patients of gastric cancer with FGFR2 amplification16,40,41,53. Recently, FGFR2 specific antibodies, FPA144 (NCT02318329) have been developed to treat patients bearing high FGFR2b-overexpressing gastric cancers. FPA144, Bemarituzumab, is glycol-engineered for enhanced antibody-dependent cell-mediated cytotoxicity (ADCC), is currently being evaluated in a phase III trial as front-line therapy for patients with FGFR2-IIIb overexpressing advanced-stage54,55. FGFR2 signaling is associated with the proliferation, migration, and angiogenesis of carcinomas11,17, therefore, these FGFR2 signaling inhibitors have been proposed to be a key drug for the treatment of solid tumors with FGFR2 overexpression. However several studies of FGFR2 inhibitors did not improve the prognosis of GC patients. FGFR2 gene amplification was detected in 39.3% of our FGFR2IIIb-positive cases and all of the FGFR2IIIc-positive cases. All FGFR2IIIc-positive tumors were also positive for FGFR2IIIb in the same tumor, but both were not positive in the same cancer cells. One of the reasons for the negative results might be this tumor heterogeneity. Kim et al.56 reported that a difference in sensitivity to FGFR2 inhibitor of AZD4547 may be related to baseline FGFR2 IIIC expression level caused by tumor heterogeneity using patient-derived tumor cell models. The molecular heterogeneity of FGFR2 IIIb type and FGFR2 IIIc might be one of the mechanisms of resistance of FGFR2 signaling therapy.
Among our 28 patients with FGFR2IIIb-positive gastric cancer, the prognoses of the four FGFR2IIIc-positive patients were significantly poorer than those of the 24 FGFR2IIIc-negative patients, and FGFR2IIIc was an independent prognostic factor in this study. FGFR2IIIc expression was reported to be correlated with the EMT32,57. In our present cohort, FGFR2IIIc expression was frequently observed in the cases with lymph node disease compared to the expressions in the primary tumors. These findings indicated that FGFR2IIIc signaling might be associated with metastatic ability of gastric cancer cells. EMT is a critical process in cancer progression that provides cancer cells with the ability to escape from the primary foci, invade stromal tissues, and metastasize to secondary regions due to decreased cell–cell adhesion58,59.
All FGFR2IIIc-positive tumors were also positive for FGFR2IIIb in the same tumor, but both were not positive in the same cancer cells. The IIIb type and the IIIc type are alternatively spliced by exon 8 and by exon 9, respectively39. We previously reported that splicing variant of FGFR2 IIIb type and FGFR2 IIIc type might be regulated the oxidative level of tumor microenvironment25. Although both FGFR2IIIc expression and FGFR2IIIb expression were observed in the same tumors, they were not positive in the same cells. In normal human tissues, FGFR2IIIb is expressed mainly on epithelial cells whereas FGFR2IIIc is expressed mostly in mesenchymal cells35. FGFR2IIIc signaling might be associated with the EMT, which is one of the malignant phenotypes of cancer progression. FGFR2IIIc expression was frequently observed in lymph node disease in the present study, and it was significantly correlated with poor survival. These findings might indicate that the switch from FGFR2IIIb to FGFR2IIIc is one of the mechanisms underlying the malignant progression of gastric cancer cells.
In conclusion, FGFR2IIIc and/or FGFR2IIIb overexpression was observed in 4.9% of gastric cancer. FGFR2IIIc might be useful independent prognostic marker for patients with gastric cancer.
Methods
Clinical materials
A total of 562 patients who were histologically confirmed to have primary gastric cancer was enrolled in this study. All patients underwent a resection of gastric tumor and regional lymph nodes at Osaka City University. None of patients had undergone preoperative radiation and/or chemotherapy. The pathologic diagnoses and classifications were made according to the UICC TNM classification of malignant tumors. This study was approved by Osaka City University ethics committee (Reference number 924). Informed consent was obtained from all patients. All methods with respect to humans were carried out in accordance with relevant guidelines and regulations.
Antibodies
Mouse-rat chimeric monoclonal antibodies which recognize human FGFR2-IIIb (mFR2-10b) and human FGFR2-IIIc (mFR2-28c) were produced and provided from Daiichi Sankyo Co., Ltd. as follows. Female WKY/Izm rats (Japan SLC, Inc.) were immunized with recombinant human FGFR2b (IIIb)/Fc Chimera or FGFR2c (IIIc)/Fc Chimera (R&D Systems, Inc.), and the lymphoid cells or splenocytes collected from immunized rats were fused with SP2/0-Ag14 mouse myeloma cells (ATCC: CRL-1581) by conventional hybridoma techniques. Hybridoma culture supernatants obtained from the established hybridomas were screened for FGFR2 binding by the flowcytometry analysis using FGFR2-expressing cells. The selected established hybridomas were further screened for their ability to apply immunohistochemistry using FGFR2-expressing cells embedded in FFPE. Rat FR2-10b (FGFR2IIIb specific) and rat FR2-28c (FGFR2IIIc specific) were selected (Supplement Fig. S1), and their variable regions were cloned and fused into expression vector containing constant region of mouse IgG1. The recombinant chimeric antibodies of mouse FR2-10b (mFR2-10b) and mouse FR2-28c (mFR2-28c) were produced in culture supernatant from FREESTYLE 293F cells (Thermo Fisher Scientific Inc.), and purified by MABSELECTSURE column chromatography (GE Healthcare Bioscience). All methods with respect to animals were carried out in accordance with relevant guidelines and regulations. An Anti-CEA antibody (sc-48364; 1:200, Santa Cruz, Dallas, TX, USA) was used to determine histologically cancer cells in the tumor tissue. A pan-FGFR2 antibody (#23328, 1:200; Cell Signaling, Danvers, MA, USA) was used to detect cancer cells with both FGFR2IIIb and/or FGFR2IIIc expression. All methods with respect to animals were carried out in accordance with relevant ARRIVE guidelines.
Immunostaining of the FGFR2-IIIb and FGFR2-IIIc
Immunohistochemical study was performed as follows. In brief, Paraffin-embedded sections from 562 patients were heated for 40 min at 98 °C by heater in Target Retrieval Solution High pH (DAKO, Carpinteria, CA). Then sections were incubated with 3% hydrogen peroxide to block endogenous peroxidase activity and the sections were immersed in protein block (DAKO protein block, serum free) to block nonspecific binding. The specimens were incubated with anti-FGFR2-IIIb (dilution, 1:333) for 30 min at room temperature, and anti-FGFR2-IIIc (dilution, 1:333) for overnight at 4 °C. These sections were incubated with mouse linker (ENVISION FLEX Mouse+, DAKO) for 20 min, and peroxidase-labeled polymer (ENVISION FLEX/HRP, DAKO) for 20 min. The sections were counterstained with hematoxylin.
Immunohistochemical determination
The immunoreactivity of FGFR2IIIb and FGFR2IIIc were evaluated according to the intensity of membranous staining in the deepest level of the tumor. FGFR2IIIb and FGFR2IIIc were mainly expressed at the cell membrane and weakly expressed in the cytosol of cancer cells. Immunostaining intensity score was rated 0–3 as follows: 0, negative; 1 +, weak at cytosol; 2 +, moderate at cell membrane; 3 +, strong at cell membrane (Fig. 1A). FGFR2IIIb or FGFR2IIIc expression were considered positive when scores were ≥ 2.
Fluorescence in situ hybridization (FISH) of FGFR2
All of 28 FGFR2-positive tumors and 5 of FGFR2-negative tumors were examined FGFR2 gene amplification by FISH analysis. The area most strongly immunostained of tumor paraffin-embedded sections were subjected to FISH. Tumor sections were cut to 4 μm thickness, followed by deparaffinization with the pretreatment reagent (Abbott Molecular Inc., 2J06-30) at 80 ± 1 °C for 30 min. Protease digestion procedures were performed using the protease reagent (Abbott Molecular Inc., 2J08-32) at 37 ± 1 °C for 60 min. FGFR2/CEN10p Dual Color FISH Probe conjugated with Texas Red/FITC (GSPLab., Inc, GC018) were hybridized at 75 °C for 5 min, and 37 °C for 48 h (Supplement Fig. S2). After hybridization, the slides were washed in 2 × saline-sodium citrate/0.3% NP-40 at 72 °C for 5 min, and counterstained with PROLONG Gold antifade reagent with DAPI (Thermo Fischer Scientific, P36935). The specimens were examined with ARIOL SL-200 (Leica Biosystems). FGFR2/CEN10 signals of 40 tumor cell nuclei were counted and FGFR2/CEN10 ratio was calculated according to the evaluation method of The PATHVYSION HER-2 DNA Probe Kit (Abbott). An FGFR2/CEN10 ratio 2.0 or higher were defined as gene amplification positive.
Statistical analysis
We used the χ2 test or Fisher’s exact test to determine the significance of the difference between the covariates. The survival durations were calculated using the Kaplan–Meier method and analyzed by the log-rank test to compare the cumulative survival durations in the patient groups. In addition, the Cox proportional hazards model was used to compute multivariate hazards ratios for the study parameters. In all of the tests, a p value < 0.05 was defined as statistically significant. Statistical analysis was conducted in R (R Project for Statistical Computing, V.3.5.3).
References
Kelleher, F. C., O’Sullivan, H., Smyth, E., McDermott, R. & Viterbo, A. Fibroblast growth factor receptors, developmental corruption and malignant disease. Carcinogenesis 34, 2198–2205. https://doi.org/10.1093/carcin/bgt254 (2013).
Kuroda, K. et al. Circulating tumor cells with FGFR2 expression might be useful to identify patients with existing FGFR2-overexpressing tumor. Cancer Sci. 111, 4500–4509. https://doi.org/10.1111/cas.14654 (2020).
Krook, M. A. et al. Fibroblast growth factor receptors in cancer: Genetic alterations, diagnostics, therapeutic targets and mechanisms of resistance. Br. J. Cancer https://doi.org/10.1038/s41416-020-01157-0 (2020).
Weaver, A. & Bossaer, J. B. Fibroblast growth factor receptor (FGFR) inhibitors: A review of a novel therapeutic class. J. Oncol. Pharm. Pract. https://doi.org/10.1177/1078155220983425 (2020).
Okuno, T. et al. Establishment of a new scirrhous gastric cancer cell line with FGFR2 overexpression, OCUM-14. Ann. Surg. Oncol. 26, 1093–1102. https://doi.org/10.1245/s10434-018-07145-2 (2019).
Yashiro, M., Matsuoka, T. & Ohira, M. The significance of scirrhous gastric cancer cell lines: The molecular characterization using cell lines and mouse models. Hum. Cell 31, 271–281. https://doi.org/10.1007/s13577-018-0211-4 (2018).
Zhao, W. M. et al. Monoclonal antibodies to fibroblast growth factor receptor 2 effectively inhibit growth of gastric tumor xenografts. Clin. Cancer Res. 16, 5750–5758. https://doi.org/10.1158/1078-0432.CCR-10-0531 (2010).
Yashiro, M. & Matsuoka, T. Fibroblast growth factor receptor signaling as therapeutic targets in gastric cancer. World J. Gastroenterol. 22, 2415–2423. https://doi.org/10.3748/wjg.v22.i8.2415 (2016).
Qiu, H., Yashiro, M., Zhang, X., Miwa, A. & Hirakawa, K. A FGFR2 inhibitor, Ki23057, enhances the chemosensitivity of drug-resistant gastric cancer cells. Cancer Lett. 307, 47–52 (2011).
Matsuoka, T. & Yashiro, M. Precision medicine for gastrointestinal cancer: Recent progress and future perspective. World J. Gastrointest. Oncol. 12, 1–20. https://doi.org/10.4251/wjgo.v12.i1.1 (2020).
Turner, N. & Grose, R. Fibroblast growth factor signalling: From development to cancer. Nat. Rev. Cancer 10, 116–129. https://doi.org/10.1038/nrc2780 (2010).
Ishiwata, T. et al. Enhanced expression of fibroblast growth factor receptor 2 IIIc promotes human pancreatic cancer cell proliferation. Am. J. Pathol. 180, 1928–1941. https://doi.org/10.1016/j.ajpath.2012.01.020 (2012).
Knights, V. & Cook, S. J. De-regulated FGF receptors as therapeutic targets in cancer. Pharmacol. Ther. 125, 105–117. https://doi.org/10.1016/j.pharmthera.2009.10.001 (2010).
Hattori, Y. et al. K-sam, an amplified gene in stomach cancer, is a member of the heparin-binding growth factor receptor genes. Proc. Natl. Acad. Sci. USA 87, 5983–5987 (1990).
Dienstmann, R. et al. Genomic aberrations in the FGFR pathway: Opportunities for targeted therapies in solid tumors. Ann. Oncol. 25, 552–563. https://doi.org/10.1093/annonc/mdt419 (2014).
Kunii, K. et al. FGFR2-amplified gastric cancer cell lines require FGFR2 and Erbb3 signaling for growth and survival. Cancer Res. 68, 2340–2348 (2008).
Nakazawa, K., Yashiro, M. & Hirakawa, K. Keratinocyte growth factor produced by gastric fibroblasts specifically stimulates proliferation of cancer cells from scirrhous gastric carcinoma. Cancer Res. 63, 8848–8852 (2003).
Yashiro, M. et al. Synergistic antitumor effects of FGFR2 inhibitor with 5-fluorouracil on scirrhous gastric carcinoma. Int. J. Cancer 126, 1004–1016 (2010).
Kim, P. T. et al. Aberrant right hepatic artery in pancreaticoduodenectomy for adenocarcinoma: Impact on resectability and postoperative outcomes. HPB (Oxford) 16, 204–211. https://doi.org/10.1111/hpb.12120 (2014).
Matsumoto, K. et al. FGFR2 gene amplification and clinicopathological features in gastric cancer. Br. J. Cancer 106, 727–732. https://doi.org/10.1038/bjc.2011.603 (2012).
Jung, E. J., Jung, E. J., Min, S. Y., Kim, M. A. & Kim, W. H. Fibroblast growth factor receptor 2 gene amplification status and its clinicopathologic significance in gastric carcinoma. Hum. Pathol. 43, 1559–1566 (2012).
Su, X. et al. FGFR2 amplification has prognostic significance in gastric cancer: Results from a large international multicentre study. Br. J. Cancer 110, 967–975 (2014).
Mor, O. et al. DNA amplification in human gastric carcinomas. Cancer Genet. Cytogenet. 65, 111–114 (1993).
Hara, T. et al. Amplification of c-myc, K-sam, and c-met in gastric cancers: Detection by fluorescence in situ hybridization. Lab. Investig. 78, 1143–1153 (1998).
Kinoshita, H. et al. Diffuse-type gastric cancer cells switch their driver pathways from FGFR2 signaling to SDF1/CXCR4 axis in hypoxic tumor microenvironments. Carcinogenesis 36, 1511–1520. https://doi.org/10.1093/carcin/bgv134 (2015).
Toyokawa, T., Yashiro, M. & Hirakawa, K. Co-expression of keratinocyte growth factor and K-sam is an independent prognostic factor in gastric carcinoma. Oncol. Rep. 21, 875–880 (2009).
Liu, Y. J. et al. HER2, MET and FGFR2 oncogenic driver alterations define distinct molecular segments for targeted therapies in gastric carcinoma. Br. J. Cancer 110, 1169–1178 (2014).
Zhang, J. et al. Targeting the oncogenic FGF-FGFR axis in gastric carcinogenesis. Cells 8, 637. https://doi.org/10.3390/cells8060637 (2019).
Matsuoka, T. & Yashiro, M. Biomarkers of gastric cancer: Current topics and future perspective. World J. Gastroenterol. 24, 2818–2832. https://doi.org/10.3748/wjg.v24.i26.2818 (2018).
Holbrook, J. D. et al. Deep sequencing of gastric carcinoma reveals somatic mutations relevant to personalized medicine. J. Transl. Med. 9, 119 (2011).
Nadauld, L. D. et al. Metastatic tumor evolution and organoid modeling implicate TGFBR2 as a cancer driver in diffuse gastric cancer. Genome Biol. 15, 428 (2014).
Ishiwata, T. Role of fibroblast growth factor receptor-2 splicing in normal and cancer cells. Front. Biosci. (Landmark Ed) 23, 626–639. https://doi.org/10.2741/4609 (2018).
Gong, S. G. Isoforms of receptors of fibroblast growth factors. J. Cell Physiol. 229, 1887–1895. https://doi.org/10.1002/jcp.24649 (2014).
Holzmann, K. et al. Alternative splicing of fibroblast growth factor receptor IgIII loops in cancer. J. Nucleic Acids 2012, 950508. https://doi.org/10.1155/2012/950508 (2012).
Orr-Urtreger, A. et al. Developmental localization of the splicing alternatives of fibroblast growth factor receptor-2 (FGFR2). Dev. Biol. 158, 475–486. https://doi.org/10.1006/dbio.1993.1205 (1993).
Teles, S. P. et al. Integrated analysis of structural variation and RNA expression of FGFR2 and its splicing modulator ESRP1 highlight the ESRP1(amp)-FGFR2(norm)-FGFR2-IIIc(high) axis in diffuse gastric cancer. Cancers 12, 70. https://doi.org/10.3390/cancers12010070 (2019).
Matsuda, Y., Hagio, M., Seya, T. & Ishiwata, T. Fibroblast growth factor receptor 2 IIIc as a therapeutic target for colorectal cancer cells. Mol. Cancer Ther. 11, 2010–2020. https://doi.org/10.1158/1535-7163.MCT-12-0243 (2012).
Chaffer, C. L. et al. Mesenchymal-to-epithelial transition facilitates bladder cancer metastasis: Role of fibroblast growth factor receptor-2. Cancer Res. 66, 11271–11278. https://doi.org/10.1158/0008-5472.CAN-06-2044 (2006).
Carstens, R. P., Eaton, J. V., Krigman, H. R., Walther, P. J. & Garcia-Blanco, M. A. Oncogene 15, 3059–3065 (1997).
Deng, N. et al. A comprehensive survey of genomic alterations in gastric cancer reveals systematic patterns of molecular exclusivity and co-occurrence among distinct therapeutic targets. Gut 61, 673–684 (2012).
Nakamura, K. et al. A novel molecular targeting compound as K-samII/FGF-R2 phosphorylation inhibitor, Ki23057, for Scirrhous gastric cancer. Gastroenterology 131, 1530–1541 (2006).
Yashiro, M. Co-expression of keratinocyte growth factor and K-sam is an independent prognostic factor in gastric carcinoma. Oncol. Rep. 21(875–880), 2009. https://doi.org/10.3892/or_00000297 (2009).
Hattori, Y. et al. Immunohistochemical detection of K-sam protein in stomach cancer. Clin. Cancer Res. 2, 1373–1381 (1996).
Matsunobu, T. et al. Expression of keratinocyte growth factor receptor correlates with expansive growth and early stage of gastric cancer. Int. J. Oncol. 28, 307–314 (2006).
Tani, H., Saito, N., Kobayashi, M. & Kameoka, S. Clinical significance of keratinocyte growth factor and K-sam gene expression in gastric cancer. Mol. Med. Rep. 7, 1381–1386. https://doi.org/10.3892/mmr.2013.1397 (2013).
Murase, H. et al. Prognostic significance of the co-overexpression of fibroblast growth factor receptors 1, 2 and 4 in gastric cancer. Mol. Clin. Oncol. 2, 509–517 (2014).
Cai, H. et al. Mutational landscape of gastric cancer and clinical application of genomic profiling based on target next-generation sequencing. J. Transl. Med. 17, 12. https://doi.org/10.1186/s12967-019-1941-0 (2019).
Cristescu, R. et al. Molecular analysis of gastric cancer identifies subtypes associated with distinct clinical outcomes. Nat. Med. 21, 449. https://doi.org/10.1038/nm.3850 (2015).
Bass, A. J. et al. Comprehensive molecular characterization of gastric adenocarcinoma. Nature 513, 202–209. https://doi.org/10.1038/nature13480 (2014).
Ichikawa, H. et al. Actionable gene-based classification toward precision medicine in gastric cancer. Genome Med. 9, 93. https://doi.org/10.1186/s13073-017-0484-3 (2017).
Katona, B. W. & Rustgi, A. K. Gastric cancer genomics: Advances and future directions. Cell. Mol. Gastroenterol. Hepatol. 3, 211–217. https://doi.org/10.1016/j.jcmgh.2017.01.003 (2017).
Peng, W. X., Kudo, M., Fujii, T., Teduka, K. & Naito, Z. Altered expression of fibroblast growth factor receptor 2 isoform IIIc: Relevance to endometrioid adenocarcinoma carcinogenesis and histological differentiation. Int. J. Clin. Exp. Pathol. 7, 1069–1076 (2014).
Xie, L. et al. FGFR2 gene amplification in gastric cancer predicts sensitivity to the selective FGFR inhibitor AZD4547. Clin. Cancer Res. 19, 2572–2583 (2013).
Catenacci, D. V. et al. Bemarituzumab with modified FOLFOX6 for advanced FGFR2-positive gastroesophageal cancer: FIGHT Phase III study design. Future Oncol. 15, 2073–2082. https://doi.org/10.2217/fon-2019-0141 (2019).
Catenacci, D. V. T. et al. Phase I escalation and expansion study of bemarituzumab (FPA144) in patients with advanced solid tumors and FGFR2b-selected gastroesophageal adenocarcinoma. J. Clin. Oncol. 2, 5. https://doi.org/10.1200/JCO.19.01834 (2020).
Kim, S. T. et al. 3-Dimensional micropillar drug screening identifies FGFR2-IIIC overexpression as a potential target in metastatic giant cell tumor. Oncotarget 8, 36484–36491. https://doi.org/10.18632/oncotarget.16883 (2017).
Ueda, J. et al. Epithelial splicing regulatory protein 1 is a favorable prognostic factor in pancreatic cancer that attenuates pancreatic metastases. Oncogene 33, 4485–4495. https://doi.org/10.1038/onc.2013.392 (2014).
Katoh, M. & Nakagama, H. FGF receptors: cancer biology and therapeutics. Med. Res. Rev. 34, 280–300 (2014).
Zhu, D. Y. et al. Twist1 correlates with poor differentiation and progression in gastric adenocarcinoma via elevation of FGFR2 expression. World J. Gastroenterol. 20, 18306–18315 (2014).
Acknowledgements
We thank Kokichi Honda (Daiichi Sankyo Co., Ltd.), Reimi Kawaida (Daiichi Sankyo Co., Ltd.), and Toshiaki Ohtsuka (Daiichi Sankyo Co., Ltd.) for providing human FGFR2-IIIb antibody (mFR2-10b) and human FGFR2-IIIc antibody (mFR2-28c) with helpful advice. We thank Maiko Nagai (Osaka City University Graduate School of Medicine), for technical assistance.
Funding
This study is partially founded by KAKENHI (Grant-in-Aid for Scientific Research) 18H02883.
Author information
Authors and Affiliations
Contributions
M.Y. and K.K. designed and performed the experiments and co-wrote the manuscript; M.Y. and K.K. contributed equally; G.M., T.O., Y.M., Y.Y., T.S., A.S., S.K., S.N., S.T., collected tumor specimens and contributed to the in vitro experiments; M.O. suggested and co-designed the study. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
Masakazu Yashiro received research funding from Daiichi Sankyo Co., Ltd. The other authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yashiro, M., Kuroda, K., Masuda, G. et al. Clinical difference between fibroblast growth factor receptor 2 subclass, type IIIb and type IIIc, in gastric cancer. Sci Rep 11, 4698 (2021). https://doi.org/10.1038/s41598-021-84107-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-84107-x
This article is cited by
-
FGFR-targeted therapeutics: clinical activity, mechanisms of resistance and new directions
Nature Reviews Clinical Oncology (2024)
-
TTN-AS1 delivered by gastric cancer cell–derived exosome induces gastric cancer progression through in vivo and in vitro studies
Cell Biology and Toxicology (2023)
-
FGFR2-amplified tumor clones are markedly heterogeneously distributed in carcinomas of the upper gastrointestinal tract
Journal of Cancer Research and Clinical Oncology (2023)
-
Therapeutisch relevante prädiktive Biomarker beim Adenokarzinom des Ösophagus
Die Onkologie (2023)
-
Significance of tumor heterogeneity of p-Smad2 and c-Met in HER2-positive gastric carcinoma with lymph node metastasis
BMC Cancer (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.