Abstract
Patients requiring diagnostic testing for coronavirus disease 2019 (COVID-19) are routinely assessed by reverse-transcription quantitative polymerase chain reaction (RT-qPCR) amplification of Sars-CoV-2 virus RNA extracted from oro/nasopharyngeal swabs. Despite the good specificity of the assays certified for SARS-CoV-2 molecular detection, and a theoretical sensitivity of few viral gene copies per reaction, a relatively high rate of false negatives continues to be reported. This is an important challenge in the management of patients on hospital admission and for correct monitoring of the infectivity after the acute phase. In the present report, we show that the use of digital PCR, a high sensitivity method to detect low amplicon numbers, allowed us to correctly detecting infection in swab material in a significant number of false negatives. We show that the implementation of digital PCR methods in the diagnostic assessment of COVID-19 could resolve, at least in part, this timely issue.
Similar content being viewed by others
Introduction
As outlined in several reports1,2,3, the problem of false-negative detection of Sars-CoV-2 in oro/nasopharyngeal swabs material continues to have a major impact on the management of the patients. Despite as of today exist more than 100 different certified tests for viral nucleic acid amplification, mainly based on reverse transcriptase quantitative polymerase chain reaction (RT-qPCR)4,5, and further nucleic acid6,7 or even protein detection systems—e.g. MALDI-mass spectrometry8—are coming into the scenario, this problem has not been solved yet. In the case of negative results of the molecular diagnostic method, other parameters are taken into account to diagnose COVID-19, such as the typical radiologic appearance of the lungs as detected by a Computer Tomography scan (CT-scan)9. The use of nucleic acid amplification methods such as the droplet digital PCR (ddPCR) has been proposed to resolve this problem, enabling detection of SARS-CoV-2 virus in superior airways with a theoretical sensitivity of 1 copy/reaction10,11. The present work was designed to retrospectively assess with a chip-based digital PCR platform (herewith-named dPCR) the presence of low viral titers in the RNA extracted from swab samples of patients with radiologic features of COVID-19 pneumonia testing negative to conventional diagnostic RT-qPCR.
Materials and methods
Ethical information
The present study was conducted after notification to the competent ethical committee, and conforming to the current laws on the management of patient personal data, and to the principles outlined in the Declaration of Helsinki of 1964. The approval of an informed consent was waived according to a specific FAQ (“Data processing in clinical trials and medical research in the context of the COVID-19 health emergency”—article 3), published by the Italian Data Protection Authority to rule the use of patients material for experimental studies on COVID-19 (See https://www.garanteprivacy.it/temi/coronavirus/faq#English) for more information.
Patient characteristics
This is a retrospective, observational cohort study. We included in the study 64 consecutive patients admitted to our Hospital between February 27 and April 29, 2020, with clinical symptoms of pneumonia and COVID-19-typical chest computed tomography (CT) features12. Patients without COVID-19 imaging features or with contraindication to CT were excluded.
Chest computed tomography
CT-scans were performed using a 256-slices CT scanner (Revolution CT; GE Healthcare, Milwaukee, WI). No contrast media were administered to the patients. CTs were recorded as positive in the presence of viral pneumonia imaging features12. In particular, we assessed ground-glass opacities (GGO), distribution, consolidations, multilobar involvement, crazy paving, air bronchogram, and the amount of infected lung using multiplane reconstructions.
Diagnostic RT-qPCR assay
Our microbiological diagnostic laboratory adopts the GeneFinder COVID-19 Plus RealAmp Kit, a One-Step Reverse Transcription Real-Time PCR (RT-PCR) Kit designed to detect the presence of SARS-CoV-2 by quantitative amplification of the RdRp, E, and N genes (Elitech). The GeneFinder™ COVID-19 PLUS RealAmp Kit is used with RNA extracted in a robotized 12-channel RNA extraction/ RT-PCR amplification platform (ELITe InGenius SP200) using nasopharyngeal swabs material (NPS) collected in UTM medium (3 mL) within 72 h after collection under appropriate storage conditions (+ 2 to + 8 °C).
The diagnostic runs are performed as follows: after thawing working solutions (COVID-19 PLUS Reaction Mixture and COVID-19 PLUS Probe Mixture), an RT-PCR master mix is prepared and stored at 4 °C in a thermally controlled rack in the platform. In parallel, an operator manually dispenses 0.2 mL of the NPS material in dedicated sample tubes installed in the sample line in the platform. The Ingenius instrument is set to perform RNA extraction using a bead-based protocol and produces a 100 μL total volume of eluted RNA, which is automatically transferred to recovery tubes and immediately used for RT-PCR. The eluted RNAs produced by the platform were immediately frozen at the end of each diagnostic run.
The protocol for diagnostic SARS-CoV-2 genes detection is performed by automatic dispensing 5 μL of the RNA eluate together with 15 μL of the RT-PCR Master Mixture into preloaded PCR tubes, followed by a 45 cycles amplification program. The expected detection targets consist of viral RdRp, N, and E RNAs and RNAseP cellular mRNA (as an internal control, IC) with these respective fluorophores: FAM, VIC, Texas Red, and Cy5. A sample is considered positive when one or more of the genes are detected below 40 threshold cycles (Ct) or below 35 Ct for IC. The specificity of the procedure is declared to be 100%, with an analytical sensitivity of 10 copies/test for each gene by the Manufacturer (https://www.elitechgroup.com/documents?search=SARS).
ACE2 detection by RT-PCR
Detection of the ACE2 mRNA was performed using 4 μL of the eluted RNAs produced by the automated platform. The RT-qPCR analysis was performed with an ABI Prism 7900 HT (Applied Biosystems) with the FAM chemistry (Thermo Fisher). As a PCR reaction mix, we used the TaqMan fast one-step Master Mix (Thermo Fischer), the ACE2 primers (Hs01085333_m; Thermo Fischer), the RPLP0 primers (Hs99999902_m1; Thermo Fischer), or the RPL32 primers (Hs00851655_g1; Thermo Fischer). Reverse transcription (RT) was performed at 50 °C for 5 min, followed by RT inactivation/initial denaturation at 95 °C for 20 s. PCR cycling consisted of a denaturation step at 95 °C for 5 s followed by annealing/extension at 60 °C for 30 s (45 cycles). For ACE2 as well as two reference genes (RPLP0 and RPL32), the cycle threshold (Ct) value was determined and the dCt value was calculated (target Ct – mean reference gene Ct).
Digital PCR assay
To proceed with a direct comparison of the efficiency of digital vs. conventional RT-qPCR assay, dPCR was performed using 5 μL of the eluted RNAs collected from the automated platform performing the RT-qPCR diagnostic assay, and appropriately stored at − 80 °C. The dPCR analysis was performed with a QuantStudio 3D Digital PCR System platform consisting of a QuantStudio 3D Instrument, a Dual Flat Block GeneAmp PCR System 9700, and a QuantStudio 3D Digital PCR Chip Loader (all from Thermo Fisher). As a PCR reaction mix, we used the TaqMan fast Virus one-step Master Mix (Thermo Fischer). We used primers and probes authorized from the Center for Disease Control and Prevention (CDC) for the SARS-CoV-2 N gene (N1 and N2) (referred in7). As a positive control, we included primers specific for the RNAseP (a non-viral transcript); water was added as a negative control. Reverse transcription (RT) was performed at 50 °C for 10 min, followed by RT inactivation/initial denaturation at 96 °C for 5 min. PCR cycling consisted of a denaturation step at 98 °C for 30 s followed by annealing/extension at 56 °C for 1 min (40 cycles), and a final extension at 60 °C for 5 min. Data analysis was performed with the online version of the QuantStudio 3D AnalysisSuite (Thermo Fisher Cloud). The expected detection targets consist of the viral N (nucleocapsid protein gene) and cellular RNAseP (as an internal control, IC) mRNAs with FAM fluorophore. The lower limit of detection (LOD) of the dPCR method was set by adding three standard deviations to the mean of the background signal achieved in three PCR amplifications with water (0.149 ± 0.0132 and 0.163 ± 0.008 copies/µL, for N1 and N2 primers, respectively). This limit corresponded to ~ 2.2 copies/mL per each primers set.
Statistical analysis
Clinical and CT characteristics are expressed as counts and proportions in the case of categorical variables. Most clinical continuous variables did not pass the D'Agostino-Pearson omnibus normality test and, thus, are reported as medians and interquartile ranges. No imputation was made for missing data points. Categorical variables were compared by the χ2 test or Fisher’s exact test, as needed. Group comparisons for continuous data were performed with the Kruskal–Wallis test, followed by Dwass–Steel–Critchlow–Fligner pairwise comparisons if the previous test was significant. We directly compared the sensitivity of dPCR with diagnostic RT-qPCR assay by computing the Pearson’s correlation coefficient (rP) between log10 copies/mL of N1 or N2 amplicons detected by the dPCR test and the Ct values of the N gene detected by the diagnostic test. The level of statistical significance was set at P < 0.05. Statistical analysis was performed using the jamovi software version 1.2.17 (https://www.jamovi.org).
Results
Patient population characteristics, as well as CT findings, are listed in Table 1. The overall median age was 67.0 (55.3–77.3) years with 39 males and 25 females. In the overall population, the prevalent symptoms on admission were fatigue (44 out of 64 patients, 68.8%), dyspnea (41 out of 64, 64.1%), and fever (34 out of 64, 53.1%). On admission, lymphocytopenia was present in 60.9% of the patients (39 out of 64). Most of the patients had elevated levels of C-reactive protein (PCR) and B-type natriuretic peptide (BNP). Only 7 patients presented with moderate acute respiratory distress syndrome (PaO2/FiO2 > 100 mmHg and ≤ 200 mmHg). Most of the patients had typical COVID-19 CT features12, such as GGO and consolidations with peripheral distribution and multi-lobar involvement with vascular enlargement (Fig. 1a–d). In this cohort, we retrospectively identified a group of 18 subjects who tested negative for SARS-CoV-2 with the conventional diagnostic RT-qPCR assay performed on oro/nasopharyngeal swab material. These patients were screened once (n = 5) or twice (n = 12) at variable time intervals (1–25 days), and never exhibited conversion to positivity. One patient (#15 in Table S1) was assessed four times in 12 days, always testing negative.
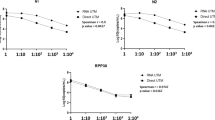
(a–d) Chest CT axial and coronal projections at the level of tracheal bifurcation in a SARS-CoV-2 positive patient (a,c) and patient #15 (b,d), admitted to our Center. (e) Correlation between the copies of N1/N2 amplicons (expressed as log10 copies/mL) detected by dPCR and the Ct values of the N gene by the single primer present in the diagnostic test. It is evident a better correlation of the data below ~ 36 Ct (evidenced by the dotted line in the graph). (f) Individual results of digital PCR analysis of the 18 patients scoring negative in the conventional diagnostic test. In patients 1–7, the digital PCR test was unable to detect the virus in the diagnostic eluted RNA. Patients 8–18 exhibited a low copy number of N1, N2 amplicons, or both. The line labeled with LOD indicates the lower detection limit of ~ 2.2 copies/mL, calculated as described in materials and methods section. (g) Patient #15 was tested in four consecutive swabbing procedures and was invariantly negative wih the diagnostic assay. Re-testing by digital PCR of the eluted RNA showed a clear SARS-CoV-2 positivity by N2 sequence amplification. (h) The ACE2 mRNA was detected by conventional RT-PCR using the eluate RNA extracted from the swab material. ACE2 expression levels showed a decreased trend in negative vs. positive patients (P = 0.0515 by t test). (i) no differences were observed in the levels of ACE2 expression in the true-negative (TN) and false-negative (FN) subjects. In panels (h) and (i) data are reported as average ± standard deviation.
To reconcile the negative results of the diagnostics RT-qPCR with a diagnosis of COVID-19 pneumonia based on clinical evaluation and CT-scan, we re-tested the eluted RNAs produced by the diagnostic pipeline using dPCR and primers for the SARS-CoV-2 N gene sequence approved by the U.S. CDC (referred in7) (N1 and N2; Figure S1). Method calibration was performed using eluted RNAs from positive patients: we found a good correlation between viral copy numbers detected with digital PCR and Ct values scored by diagnostic RT-qPCR, at least up to ~ 36 Ct, corresponding to ~ 1000 copies/mL (Fig. 1e), the lower copy number for reliable viral detection as demonstrated elsewhere13. The analysis revealed that 11 (61%) of the 18 RT-qPCR negative patients had a detectable number of SARS-CoV-2 copies, with either one or both primers’ sets (Fig. 1f) when tested with dPCR. Interestingly, patient #15 (invariantly negative with the RT-qPCR test) showed conversion from negativity to positivity and vice-versa with dPCR, with a peak in copy number in the second swab test (Fig. 1g).
We then analyzed whether there were differences between groups in the clinical, laboratory, and CT imaging characteristics when patients were stratified according to dPCR positivity for SARS-CoV-2. We distinguished patients into: (i) RT-qPCRneg/dPCRneg (herewith referred as to ‘true negatives’; n = 7), when both the diagnostic test and the digital PCR test were negative. (ii) RT-qPCRneg/dPCRpos (herewith referred as to ‘false negatives’; n = 11), when the diagnostic test was negative but the digital PCR test was positive. (iii) RT-qPCRpos (herewith referred as to ‘positives’; n = 46), when samples were positive with the conventional diagnostic test. As shown in Table 1, we did not observe differences in the demographic characteristics and symptoms on admission between the three groups. Likewise, there were no differences in comorbidities, except for a higher prevalence of previous lung diseases (chronic obstructive pulmonary disease or asthma) in the false-negative group than in the positive group (27.3% vs. 0%, respectively; P = 0.0056). Blood count, coagulation tests, hepatic and renal functions, and blood glucose were similar among the groups. Conversely, BNP median plasma concentration was higher in true negatives than in positives (1546 [740–2271] vs. 130 [25.9–424], respectively; P = 0.0122) and CRP levels were more elevated in the false-negative than in the positive group (80 [20.9–151] vs. 17.2 [6.1–49.5]; P = 0.0346). Interestingly, there was a decreasing trend in white blood cell count, which dropped from a median of 11.0 × 103/µL (8.9–16.2) in true negatives to 7.9 × 103/µL (6.5–10.3) in false negatives, to 6.8 × 103/µL (4.9–9.5) in positives, with a significant difference between false negatives and positives at post-hoc analysis (P = 0.037). Finally, there were no differences in chest CT features and outcomes among the three groups, except for a higher prevalence of peripherally distributed GGOs or consolidations in positive than in true negative patients (80.4% vs. 28.6%, respectively; P = 0.0104).
Recently, a single-cell RNA profiling of SARS-CoV-2/coronavirus-associated receptors and factors (SCARFs) expression has been performed in several human tissues including the nasal epithelium14. Since a relatively high variability in the expression of these receptors (e.g. ACE2) was reported, with possible implications for the severity of the infection in superior airways, we measured ACE2 in the RNA contained in the swab eluates and correlated this level to the positive/negative status of our patients. As shown in Fig. 1h, the ACE2 level exhibited a lower trend in negative subjects, with no discrimination between true and false negatives (Fig. 1i).
Discussion
Our results show that in the cohort of subjects with COVID-19 pneumonia who scored negative with the diagnostic SARS-CoV-2 RT-qPCR (18/64), 11/64 (~ 17%) turned out to be false negatives with dPCR amplification, thus increasing the overall sensitivity of the virus molecular detection from ~ 72 to ~ 89%. This finding consolidates the utility of high-resolution amplification methods, as ‘second-level’ assays to detect SARS-CoV-2 infection in subjects with clear COVID-19 pneumonia and low viral replication in the superior airways3,11,15. Limitations in the use of dPCR still exist, considering that a number of subjects in our cohort (the true negatives; 7/64; ~ 11%) did not exhibit positive amplification of the viral N gene sequences even with this high sensitivity technique. Other factors such as the quality of the swabbing procedure and the reported absence of detectable viral replication in superior airways2 could account for the failure to detect SARS-CoV-2 in the RNA material of these subjects, even with the highest performance detection methods. A preventive strategy to minimize this problem is to perform more thorough and comprehensive material collection, e.g. by concentrating on multiple respiratory sites16, repeat tests at different times during the course of the illness, or test broncho-alveolar aspirate in addition to the superior airways material17.
Although we cannot provide an explanation for the different replication of the virus in superior airways and its potential relationships with alterations of laboratory markers and severity of the pathology18, we noticed a lower trend in the expression ACE2 (one of the major SARS-CoV-2 receptors19) in patients with negative RT-qPCR diagnosis (Fig. 1h). If these trends were confirmed in studies with adequate sample size, it would provide additional criteria to diagnose patients with higher accuracy.
In summary, confirming previous reports, we show the superior performance of dPCR amplification for the diagnosis of SARS-CoV-2 infection in subjects with low viral titers in conventional swab tests. Furthermore, given the existence of multiple receptors and intracellular pathways involved in virus entry into cells14, this method seems useful to investigate the biological basis of low SARS-CoV-2 replication in the superior airways in a significant proportion of symptomatic COVID-19 patients, with benefits for the overall outbreak management.
References
Xiao, A. T., Tong, Y. X. & Zhang, S. False negative of RT-PCR and prolonged nucleic acid conversion in COVID-19: Rather than recurrence. J. Med. Virol. https://doi.org/10.1002/jmv.25855 (2020).
West, C. P., Montori, V. M. & Sampathkumar, P. COVID-19 testing: The threat of false-negative results. Mayo Clin. Proc. 95, 1127–1129. https://doi.org/10.1016/j.mayocp.2020.04.004 (2020).
Wiersinga, W. J., Rhodes, A., Cheng, A. C., Peacock, S. J. & Prescott, H. C. Pathophysiology, transmission, diagnosis, and treatment of coronavirus disease 2019 (COVID-19): A review. JAMA https://doi.org/10.1001/jama.2020.12839 (2020).
Sharfstein, J. M., Becker, S. J. & Mello, M. M. Diagnostic testing for the novel coronavirus. JAMA J. Am. Med. Assoc. 323, 1437–1438. https://doi.org/10.1001/jama.2020.3864 (2020).
Pokhrel, P., Hu, C. & Mao, H. Detecting the coronavirus (COVID-19). ACS Sens. https://doi.org/10.1021/acssensors.0c01153 (2020).
Esbin, M. N. et al. Overcoming the bottleneck to widespread testing: A rapid review of nucleic acid testing approaches for COVID-19 detection. RNA 26, 771–783. https://doi.org/10.1261/rna.076232.120 (2020).
Udugama, B. et al. Diagnosing COVID-19: The disease and tools for detection. ACS Nano 14, 3822–3835. https://doi.org/10.1021/acsnano.0c02624 (2020).
Nachtigall, F. M., Pereira, A., Trofymchuk, O. S. & Santos, L. S. Detection of SARS-CoV-2 in nasal swabs using MALDI-MS. Nat. Biotechnol. https://doi.org/10.1038/s41587-020-0644-7 (2020).
Long, C. et al. Diagnosis of the coronavirus disease (COVID-19): rRT-PCR or CT?. Eur. J. Radiol. 126, 108961. https://doi.org/10.1016/j.ejrad.2020.108961 (2020).
Falzone, L. et al. Sensitivity assessment of droplet digital PCR for SARS-CoV-2 detection. Int. J. Mol. Med. 46, 957–964. https://doi.org/10.3892/ijmm.2020.4673 (2020).
Suo, T. et al. ddPCR: A more sensitive and accurate tool for SARS-CoV-2 detection in low viral load specimens. medRxiv https://doi.org/10.1101/2020.02.29.20029439 (2020).
Ye, Z., Zhang, Y., Wang, Y., Huang, Z. & Song, B. Chest CT manifestations of new coronavirus disease 2019 (COVID-19): A pictorial review. Eur. Radiol. 30, 4381–4389. https://doi.org/10.1007/s00330-020-06801-0 (2020).
Callahan, C. et al. Nasal-swab testing misses patients with low SARS-CoV-2 viral loads. medRxiv. https://doi.org/10.1101/2020.06.12.20128736 (2020).
Singh, M., Bansal, V. & Feschotte, C. A single-cell RNA expression map of human coronavirus entry factors. Cell Rep. 32, 108175. https://doi.org/10.1016/j.celrep.2020.108175 (2020).
Liu, Y. et al. Aerodynamic analysis of SARS-CoV-2 in two Wuhan hospitals. Nature 582, 557–560. https://doi.org/10.1038/s41586-020-2271-3 (2020).
Mohammadi, A., Esmaeilzadeh, E., Li, Y., Bosch, R. J. & Li, J. Z. SARS-CoV-2 detection in different respiratory sites: A systematic review and meta-analysis. EBioMedicine https://doi.org/10.1016/j.ebiom.2020.102903 (2020).
Loeffelholz, M. J. & Tang, Y. W. Laboratory diagnosis of emerging human coronavirus infections—The state of the art. Emerg. Microbes Infect. 9, 747–756. https://doi.org/10.1080/22221751.2020.1745095 (2020).
Velavan, T. P. & Meyer, C. G. Mild versus severe COVID-19: Laboratory markers. Int. J. Infect. Dis. 95, 304–307. https://doi.org/10.1016/j.ijid.2020.04.061 (2020).
Wang, Q. et al. Structural and functional basis of SARS-CoV-2 entry by using human ACE2. Cell https://doi.org/10.1016/j.cell.2020.03.045 (2020).
Funding
The present research was in part funded from Institutional resources (Ricerca 5 per mille, Italian Ministery of Health) and in part from a grant issued to MP by Regione Lombardia (POR FESR 2014–2020-LINEA 2A COVID-grant no. 1850333).
Author information
Authors and Affiliations
Contributions
P.P., P.S., C.V., V.R., and G.G. performed experiments. V.A.M., M.E.M., A.F., D.A., E.M.A., and P.A. performed clinical/radiological observations. P.P., C.B., S.S.B., L.P., A.R., A.S., E.S., M.C.V., M.C., G.I.C., and M.P. collected material. P.P., P.S., D.C., M.L.B., M.C., D.A., P.A., G.I.C., and M.P. analyzed the data. G.I.C., P.P., and G.I.C. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Poggio, P., Songia, P., Vavassori, C. et al. Digital PCR for high sensitivity viral detection in false-negative SARS-CoV-2 patients. Sci Rep 11, 4310 (2021). https://doi.org/10.1038/s41598-021-83723-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-83723-x
This article is cited by
-
Multi-omics in thoracic aortic aneurysm: the complex road to the simplification
Cell & Bioscience (2023)
-
Flt1 produced by lung endothelial cells impairs ATII cell transdifferentiation and repair in pulmonary fibrosis
Cell Death & Disease (2023)
-
Multiplex real-time RT-PCR method for the diagnosis of SARS-CoV-2 by targeting viral N, RdRP and human RP genes
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.