Abstract
In the United States, farm-raised shrimp accounts for ~ 80% of the market share. Farmed shrimp are cultivated as monoculture and are susceptible to infections. The aquaculture industry is dependent on the application of antibiotics for disease prevention, resulting in the selection of antibiotic-resistant bacteria. We aimed to characterize the prevalence of antibiotic-resistant bacteria and gut microbiome communities in commercially available shrimp. Thirty-one raw and cooked shrimp samples were purchased from supermarkets in Florida and Georgia (U.S.) between March–September 2019. The samples were processed for the isolation of antibiotic-resistant bacteria, and isolates were characterized using an array of molecular and antibiotic susceptibility tests. Aerobic plate counts of the cooked samples (n = 13) varied from < 25 to 6.2 log CFU/g. Isolates obtained (n = 110) were spread across 18 genera, comprised of coliforms and opportunistic pathogens. Interestingly, isolates from cooked shrimp showed higher resistance towards chloramphenicol (18.6%) and tetracycline (20%), while those from raw shrimp exhibited low levels of resistance towards nalidixic acid (10%) and tetracycline (8.2%). Compared to wild-caught shrimp, the imported farm-raised shrimp harbored distinct gut microbiota communities and a higher prevalence of antibiotic-resistance genes in their gut. The presence of antibiotic-resistant strains in cooked shrimps calls for change in processing for their mitigation.
Similar content being viewed by others
Introduction
Antibiotic resistance is one of the severe threats faced by the world. In the U.S., more than 2.8 million people are infected with antibiotic-resistant strains each year, resulting in at least 35,000 deaths1. A review on global antibiotic resistance reported that by the year 2050, 10 million lives per year are at risk due to the rise of infection by antimicrobial-resistant strains2. According to the United States Department of Agriculture (USDA), ~ 97% of fish and shellfish consumed in the U.S. are imported from China, Ecuador, India, Indonesia, Thailand, and Vietnam3,4. In the large aquaculture operations, where farmed shrimps are cultivated as monoculture at high density, the prevalence of pathogens is one of the biggest challenges faced by the aquaculture industry5. Infection caused by pathogens in aquaculture operations can lead to massive production loss6 that can completely wipe out the shrimp farms7. The post-larvae infection caused by bacterial pathogens in shrimps in 2013 resulted in 1 billion dollars in production loss6. To prevent the production loss associated with bacterial infections, the aquaculture industry and shrimp hatcheries are heavily dependent on the application of antibiotics as prophylactic and therapeutic agents8,9,10,11,12. The use of multiple classes of antibiotics, e.g., tetracyclines (chlortetracycline, oxytetracycline, and tetracyclines), fluoroquinolone (enrofloxacin and ciprofloxacin), quinolones (oxolinic acid and norfloxacin), sulfonamides, chloramphenicol, and nitrofuran are practiced in shrimp farming5,13,14. Antibiotics are administered in aquaculture by mixing them in their diet or by adding them in rearing water. Continuous application of antibiotics by the shrimp hatcheries and farms facilitates the development of antibiotic-resistant bacterial (ARB) strains15. Non-biodegradable antibiotics with a long activity period apply selective pressures in an environment16. For instance, application of quinolones (oxolinic acid)17 in a high microbial load environment, forms a perfect combination for the selection of antibiotic-resistant strains in farm-raised shrimp. Once an antibiotic-resistance determinant has been selected, it can easily be transferred to other strains.
Popular foods like cooked shrimp or seafood mix are processed using mild heat treatment18 and can act as perfect vehicles for transferring ARB strains. Currently, FDA cooking recommendations are focused on mitigating Listeria monocytogenes in cooked shrimp19, which may not be effective to eliminate antibiotic-resistant bacteria spread over multiple genera. Further, antibiotic-resistant strains can be more tolerant to milder temperature treatments used for shrimp processing20,21. Cooked shrimp samples are mostly harvested and processed overseas. Each country and processing facility has its own processing specifications, due to which there is a considerable variation in the microbiological specifications of cooked shrimp available in the retail market. These samples that can be directly consumed after thawing or short heat treatment can be a perfect vehicle for ARB strains and opportunistic pathogens. Past studies have extensively studied the prevalence of antibiotic resistance in fresh produce22, beef23, poultry24, and manure from different livestock25,26. However, limited research has been conducted evaluating the prevalence of ARB strains in shrimp. Cross contact of raw shrimp with processed shrimp is another issue of concern27. The presence of ARB strains and the opportunistic pathogen can be detrimental to human health and limit treatment options.
Microbiome diversity has a significant impact on host health28. The competitive exclusion theory entails that the higher gut microbial diversity, the lower the possibility for pathogenic colonization29. The application of antibiotics at hatcheries and shrimp farms can cause dysbiosis. Researchers in the past have characterized and observed major gut microbiome composition differences among diseased and healthy shrimps30. Similarly, a comparison of the gut microbiome composition of farmed shrimp raised with antibiotics and wild-caught shrimp raised without antibiotics can be used for developing alternate sustainable methods for shrimp farming. Therefore, the aim of this study was to (a) isolate and characterize ARB strains among commercially available shrimp samples and (b) characterize the gut microbiome diversity of wild-caught shrimp from the U.S. compared to farm-raised shrimp imported from Ecuador.
Results
Distinct arrays of antibiotic resistance spectrum and the prevalence of antibiotic-resistant bacteria in raw versus cooked shrimp
A total of 31 shrimp samples (13 cooked and 18 raw shrimp) (Table 1) were tested for the presence of antibiotic-resistant bacteria. These shrimp samples originated from eight different countries i.e., Argentina (n = 2), Ecuador (n = 3), India (n = 6), Indonesia (n = 6), Panama (n = 1), Thailand (n = 3), U.S. (n = 7) and Vietnam (n = 3). Aerobic plate count (APC) and coliform count of cooked shrimp samples (n = 13) varied from < 25 CFU/g to 6.2 log CFU/g and < 25 CFU/g to 2.6 log CFU/g, respectively. All shrimp samples tested negative for Salmonella. Presumptive black colonies obtained on Xylose Lysine Deoxycholate agar with tergitol (XLDtn) and Hektoen Enteric (HE) agar plates tested negative using the Salmonella specific real-time PCR assay.
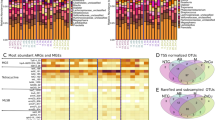
The shrimp samples (Table 1) were subsequently used for the isolation of extended-spectrum beta-lactam (ESBL) and carbapenem-resistant bacterial strains. After selective enrichment and plating on MacConkey agar media supplemented with specific antibiotics, a total of 120 bacterial isolates were obtained. Interestingly, no isolate was obtained from sample number 21 (cooked shrimp, farm-raised, Indonesia). Out of 120 isolates, 16S rRNA gene sequences were generated for 110 isolates following BLAST analysis. The 16S rRNA gene sequences generated were deposited to the NCBI nucleotide database and are available under NCBI GenBank accession numbers MT470929.1 to MT471033.1. The final 110 isolates represented 18 bacterial genera: Acinetobacter (n = 15), Aeromonas (n = 2), Alcaligenes (n = 1), Citrobacter (n = 18), Escherichia (n = 8), Enterobacter (n = 12), Enterococcus (n = 4), Hafnia (n = 1), Klebsiella (n = 2), Lelliottia (n = 5), Morganella (n = 14), Obesumbacterium (n = 2), Pantoea (n = 1), Proteus (n = 7), Providencia (n = 1), Serratia (n = 5), Stenotrophomonas (n = 3) and Vibrio (n = 9). The isolates obtained from cooked samples (n = 45) were dominated by Morganella (24%, n = 11), Proteus (11%, n = 5), Lelliottia (11.5%, n = 5), Enterobacter (9%, n = 4), Enterococcus (9%, n = 4) and Citrobacter (9%, n = 4) isolates. Whereas Acinetobacter (22%, n = 13), Citrobacter (23%, n = 13), Enterobacter (13%, n = 8), Vibrio (13%, n = 8), and Escherichia (10%, n = 6) were the top five genera isolated from raw shrimp samples.
Disc diffusion susceptibility assay was performed on 107 isolates. Three isolates [Citrobacter freundii (17B MM1), Morganella morganii (20A MM1), and Vibrio parahaemolyticus (7B MM)] were excluded from the study due to the lack of susceptibility standards. Data from the disc diffusion assay showed higher resistance towards tetracycline (9/45, 20%) and chloramphenicol (8/43, 19%) among isolates obtained from the cooked sample (Fig. 1a). Intermediate levels of resistance were observed among cooked sample isolates for ciprofloxacin, imipenem, and chloramphenicol at 17%, 12%, 7%, respectively. The most common resistance phenotype observed among cooked samples was tetracycline-chloramphenicol (C-TE). Interestingly, strains of Stenotrophomonas maltophilia were isolated from cooked, medium-sized shrimp samples imported from Vietnam. To our knowledge, this is the first report of the isolation of S. maltophilia from cooked shrimp samples.
Differences in the prevalence of antibiotic-resistant bacteria in raw versus cooked shrimp. (a) Percentage resistance observed in isolates from cooked and raw shrimp samples by disc diffusion assay. (b) Real-time multiplex PCR assay for the detection of ESBLs and carbapenem genes. Isolates from cooked sample (n = 45) and raw samples (n = 62). (c) Generalized linear model demonstrating the profiles of antibiotic resistance percentage and the prevalence of resistance genes between raw versus cooked shrimp. (d) Heatmap clustering analysis showing the clusters of antibiotic resistance profiles and the resistant genes in raw versus cooked shrimp. Aztreonam (ATM-30), Cefotaxime (CTX-30), Ceftazidime (CAZ-30), Imipenem (IPM-10), Ertapenem (ETP-10), Meropenem (MEM-10), Tetracycline (Te-30), Ciprofloxacin (CIP-5), Nalidixic Acid (NA-30), Chloramphenicol (C-30). *p < 0.05.
Among isolates obtained from raw shrimp samples, lower levels of antibiotic-resistance were observed. Resistance towards tetracycline, nalidixic acid, chloramphenicol, and imipenem was observed at 8.1% (5/62), 10% (4/40), 5% (2/40), and 1.6% (1/62), respectively (Fig. 1a). However, high levels of intermediate resistance were observed towards cefotaxime (24.2%, n = 15/62), nalidixic acid (22.5%, n = 9/40), ciprofloxacin (11%, n = 6/53), and chloramphenicol (10%, n = 4/40). Surprisingly, all Acinetobacter isolates showed intermediate resistance towards cefotaxime. Raw shrimp isolates were found to be highly susceptible to ceftazidime, ciprofloxacin, meropenem, aztreonam, and ertapenem antibiotics by disc diffusion assay.
The minimum inhibitory concentration (MIC) data using the GN2F Sensititer Gram-Negative plates for six strains confirmed the presence of multiple drug resistance (MDR) strains in raw as well as cooked shrimp samples (Supplementary Table 1). The isolates from cooked shrimp (i.e., M. morganii 31A MNC, Proteus mirabilis 20B-MC1 and Serratia marcescens 10A-MNC) showed resistance and intermediate resistance towards multiple antibiotics belonging to beta-lactams, cephalosporins, and nitrofurantoin class (Supplementary Table 1). However, the raw shrimp isolates E. hormaechei 2B-MC1, S. marcescens 28B-MC2, and Vibrio parahaemolyticus 24B-MC2 were resistant to 3 to 7 different antibiotics belonging to ampicillin, cephalosporin, fluoroquinolones, and nitrofurantoin. Further, whole-genome sequencing of isolate E. hormaechei 2B-MC1 showed the presence of sul1, sul2, qnrA1, oqxB, dfrA23, blaACT, floR, fosA, tet(A), aph(6)-Id, and aph(3″)-Ib resistance genes31. Based on the observed antibiotic profile, all isolates were classified as MDR isolates. The real-time PCR assay for the detection of ESBL resistance and carbapenem resistance genes showed a high prevalence of blaTEM (31.1%), blaOXA-48 (9.2%), and blaCMY (9.2%) genes among isolates (Fig. 1b). As shown in Fig. 1c, further analysis of the generalized linear model on the antibiotic susceptibility profiles and the prevalence of resistance genes in raw versus cooked shrimp also revealed a significant (p = 0.018) difference between the two groups. Figure 1d, in the form of hierarchical heatmap clustering analysis, also clearly demonstrates distinct signatures of antibiotics susceptibility array and the prevalence of resistance genes between the raw versus cooked shrimp.
Distinct signatures of gut microbiome communities and antibiotic-resistant bacteria in wild-caught versus farm-raised shrimp
To further investigate whether and how the overall gut microbiome communities associated with these shrimps differ by geographical origin as well as the rearing method (farm vs. wild-caught), the 16S rRNA metagenomics analysis of the gut microbiome associated with the two groups of shrimps, i.e., farm-raised (from Ecuador) versus wild-caught (from the U.S.) was performed. Shrimp samples collected from countries with a smaller number of samples were removed from the analysis. Interestingly, the analysis revealed remarkably distinct spectrums of microbiome signatures between the two groups of shrimps (Fig. 2). The analysis of β-diversity (in terms of the principal coordinate analysis of the Bray–Curtis index) clearly demonstrated significantly discrete spectrums of microbiome signatures between shrimp from the U.S. versus those from Ecuador (Fig. 2a). Further α-diversity analyses in terms of observed operational taxonomic units (OTU), Chao1 index as well as the Shannon diversity index revealed higher bacterial species richness and diversity in wild-caught shrimps from the U.S. compared with shrimps from Ecuador (Fig. 2b–d). Accordingly, further analysis at the level of major bacterial phyla and genera showed clearly distinct microbiome composition between the two groups of shrimps (Fig. 2e–f). Compared to farm-raised shrimps from Ecuador, the wild-caught shrimps from the U.S. harbored a higher abundance of Proteobacteria and Planctomyces and had a lower abundance of Tenericutes and Bacteroidetes (Fig. 2e). Interestingly, the analysis of organism-level phenotypes showed that the shrimps from the U.S. harbor a significantly higher overall ratio of Gram-negative to Gram-positive bacterial taxa as compared to shrimps from Ecuador (Fig. 2g). In addition, the proportion of aerobic taxa was higher while that of facultative anaerobic taxa was lower in shrimps from the U.S. versus those from Ecuador (Fig. 2h). Linear discriminatory analysis (LDA) effect size (LEfSe) analysis was performed to infer the relative abundance of unique taxonomic clades and Kyoto Encyclopedia of Genes and Genomes (KEGG) metagenome orthologs that were significantly (p < 0.01) over-or under-represented between shrimp from Ecuador and the U.S.. The LEfSe analysis identified several unique bacterial taxa that drive differences in the gut communities between the two groups of shrimps (Fig. 2i). Figure 2j further simplifies these data and presents the LDA scores of the taxa that were significantly (p < 0.01) different between the two groups of shrimps. The shrimps from the U.S. were uniquely distinguished by a higher proportion of Proteobacteria, Alteromonadales, Rhizobiales, Synechococcaceae, Myxococcales, and Planctomyces, whereas shrimps from Ecuador were characterized by a higher abundance of Tenericutes, Flavobacteriales, Desulfovibrio, Oscillatoriales, Mycobacterium, and Bacteroidales (Fig. 2i–j). In addition, data from the study showed several bacterial species that were significantly different between the two groups of shrimps (Supplementary Fig. 1a). These differentially abundant markers could be considered as potential metagenomic biomarkers for the identification of shrimp from Ecuador (farm-raised) and the U.S. (wild-caught). Further, the PICRUSt (Phylogenetic Investigation of Communities by Reconstruction of Unobserved States) analysis of the KEGG orthologs associated with these bacterial signatures also revealed distinct clusters among the two groups (Supplementary Fig. 1b). The shrimps from the U.S. were characterized mainly by a higher (p < 0.01) abundance of bacterial taxa associated with the metabolism of several amino acids, lipids, cofactors, and vitamins in addition to the xenobiotic degradation (Supplementary Fig. 1b). In contrast, the shrimps from Ecuador presented a higher proportion of taxa associated with the metabolism of amino acid-related enzymes and gluconeogenesis in addition to the taxa related to primary immunodeficiency (Supplementary Fig. 1b).
Distinct bacterial microbiome signatures in shrimp from U.S. (wild-caught) versus Ecuador (farm-raised) shrimps. (a) Beta-diversity (principal coordinate analysis; Bray–Curtis dissimilarity index; (b–d) alpha-diversity indices; (e,f) microbiome composition at phylum and genus level; (g,h) ratio of gram-positive to gram-negative bacteria and aerobic to anaerobic bacteria; and (i,j) linear discriminatory analysis effect size (LEfSe) cladogram representing the significantly unique bacterial taxa driving the difference between the wild-caught (U.S.) versus farm-raised (Ecuador) shrimp.
In addition to gut microbiome analysis, the bacterial genomic DNA from raw shrimp gut from the U.S. (wild-caught) and Ecuador (farm-raised) was subjected to real-time PCR assays for examining the prevalence of antibiotic-resistance genes. Interestingly, compared to raw shrimp sample of U.S. origin (wild-caught), the shrimp from Ecuador (farm-raised) showed a significantly higher prevalence of blasul-1, blasul-2, blaqnrS, blaaadA, blamec-A, blaCMY, blafloR, blacat, and blatet-A (p < 0.01) while demonstrating a lower prevalence of blaCTX-M and blavan-A (p < 0.01) (Fig. 3).
Discussion
According to the Food and Agricultural Organization (FAO), the microbiological limits for frozen cooked shrimp samples with aerobic plate count 107 CFU/g is the boundary between marginally acceptable counts and unacceptable counts32. In our study, the APC of cooked shrimp samples ranged from < 25 CFU/g to 6.2 logs CFU/g. Cooked shrimp samples from India (sample No. = 30) and Vietnam (sample No. = 31) had a high APC. Variations in hazard analysis critical control point and processing plans in different countries and temperature fluctuation during transportation may be possible reasons for the observed variation. Some samples showed as low as < 25 CFU/g APC, while other samples had high APC and even tested positive for the presence of antibiotic-resistant bacteria. We found that 22% (n = 24/107) of the bacterial isolates were resistant to one or more antibiotics. Interestingly, compared to isolates from raw shrimp, the resistance among isolates from cooked shrimp was significantly higher towards chloramphenicol (19%), aztreonam (7.7%), cefotaxime (7.3%), and meropenem (7.3%) (p < 0.01) (Fig. 1a). In a similar study, Nawaz et al.33 reported resistance to nalidixic acid, ampicillin, tetracycline, and chloramphenicol in 52% (55/105) of E. coli isolates from imported black Asian shrimp (n = 330).
Shrimp processing facilities rely on the application of a shorter and milder thermal treatment, which may not be sufficient for mitigation of ARB strains spread over multiple genera. Several studies have demonstrated the interplay between stress associated with food processing, antibiotic resistance, and vice-versa34. In fact, exposure to mild temperatures has been shown to enhance the selection of ARB strains. Ebinesh et al. observed a reduction in the inhibition zone for amikacin, imipenem, meropenem, norfloxacin, piperacillin, and tazobactam among six A. baumannii strains following heat treatment at 45 °C21. In another study, the ARB strains of E. coli O157:H7 were found to be less susceptible to 55 °C treatment compared to the wild strain20. Hwang et al. reported survival of antibiotic-resistant strains of E. coli with the tet(A) gene in the human gastrointestinal tract after consumption of retail ready-to-eat food contaminated ARB E. coli strains35. They further reported the transfer of the resistance gene from the ARB strain associated with food to antibiotic sensitive cells in the human gastrointestinal tract35. Commercially available cooked shrimp samples with high APC are not likely to cause infection in healthy individuals. However, the consumption of frozen shrimp harboring antibiotic-resistant strains of opportunistic pathogens can lead to serious illnesses, particularly in immunocompromised populations. The presence of ARB strains in cooked food may promote the transfer of resistance genes to bacterial strains in the human gut microbiome or pathogens.
Oxytetracycline is widely used by hatcheries across the world36,37. Along with the widespread use of oxytetracycline for aquaculture, resistance to this drug is commonly observed in bacterial isolates from shrimp38. In the present study, tetracycline resistance was associated with Enterobacter hormaechei, Enterococcus faecalis, Morganella morganii, Proteus mirabilis, and Vibrio parahaemolyticus isolates from the shrimp imported from India, Vietnam, and Indonesia. Chloramphenicol, a broad-spectrum antibiotic, is banned for use in food-producing animals by the U.S., European Union, and many other countries due to its toxic side-effects in humans39. However, the presence of chloramphenicol residue is a common reason for the rejection of imported shrimp by the FDA40. The presence of chloramphenicol residues reflects the indiscriminate use of banned antibiotics in shrimp-exporting counties. In the present study, chloramphenicol resistance was associated with Enterobacter hormaechei, Enterobacter lignolyticus, Proteus mirabilis, Morganella morganii, and Serratia marcescens isolates. In cooked samples, chloramphenicol resistance was associated with Enterococcus faecalis, M. morganii, P. mirabilis, and S. maltophilia, all of which are known to be opportunistic nosocomial pathogens and cause infection in the immunosuppressed population41. Not many reports are available on the isolation of carbapenem-resistant bacteria from food-producing animals, including shrimp and their environment42. A pilot study conducted by the National Antimicrobial Resistance Monitoring System (NARMS) reported carbapenem-resistant isolates with blaNDM-1 resistance genes in Acinetobacter baumannii and Aeromonas sobria from seafood samples collected from the U.S.43. In our study, carbapenem-resistance was observed in strains belonging to E. hormaechei, P. mirabilis, and S. maltophilia. Additionally, a small percentage of isolates from both raw and cooked shrimp samples tested positive for the presence of blaKPC, blaNDM, blaOXA48, and blaIMP carbapenem-resistance genes. These genes were associated with Acinetobacter pittii (blaOXA48), Citrobacter spp. (blaOXA48), C. freundii (blaKPC and blaOXA48), Enterobacter cloacae (blaKPC and blaNDM), E. hormaechei (blaNDM and blaOXA48), E. lignolyticus (blaOXA48), Klebsiella pneumoniae (blaOXA48), M. morganii (blaOXA48), Vibrio fluvialis (blaIMP), and V. parahaemolyticus (blaOXA48). The carbapenem-resistance genes are plasmid-borne, which can facilitate their spread in the food-producing environment and can promote the development of co-resistance42.
In the present study, low levels of resistance were observed towards ESBL antibiotics. All isolates were susceptible to ceftazidime, 7.7% of isolates showed resistance to cefotaxime and aztreonam, and 16.6% of isolates showed intermediate resistance to cefotaxime. Intermediate resistance to cefotaxime was primarily associated with strains belonging to the Acinetobacter pittii. A previous study also reported resistance to third-generation (cefotaxime, ceftriaxone, and ceftazidime) and fourth-generation (cefepime) cephalosporins in two E. cloacae isolate from fresh seafood44. A similar study reported the isolation of ESBL-producing Enterobacteriaceae from fresh seafood samples collected from the India market, wherein 80% (n = 53/66) of E. coli isolates were found to be ESBL producers45. MDR phenotype was reported among 169 isolates with resistance to third-generation cephalosporins, aztreonam, ertapenem, and meropenem45. Further, the isolates from the study were found to be positive for ESBL-encoding genes blaCTX, blaSHV, and blaTEM and the metallo-β-lactamase gene blaNDM-1. The extensive use of unapproved antibiotics for aquaculture operations in India can be a possible reason for this higher prevalence of antibiotic-resistant bacteria in shrimp harvested from India. In 2019, a total of 61 shrimp entry lines were refused by the FDA due to the presence of banned antibiotic residues from different countries. Among all refused products, shrimp products exported from India (n = 33/61) have consistently tested positive for banned antibiotics reflecting extensive use of banned antibiotics for aquaculture in India46,47,48.
The strains of S. maltophilia were isolated from farm-raised shrimp from Vietnam and following the disc diffusion assay antimicrobial susceptibility testing (AST) analysis of the strain confirmed its intrinsic resistance (i.e., towards chloramphenicol and ceftazidime)49. Vietnam allows the use of 32 antibiotics for aquaculture50. Due to the extensive use of a wide range of antibiotics in Vietnam, 2-hydroxy-3 phenylpyrazine (HPP), ampicillin, and β-lactam antibiotic degraded products were detected in 60 water samples collected from rivers (11/26) and household ponds (49/62)51. Yamasaki et al.52 reported ciprofloxacin (4/247), enrofloxacin (12/247), oxolinic acid (1/247), sulfamethazine (2/247), sulfamethoxazole (2/247), and trimethoprim (1/247) residues in shrimp samples collected from Vietnam. Indiscriminate use of a wide range of antibiotics for aquaculture operation and contamination of aquaculture ponds by untreated hospital waste can be one of the possible reasons for the presence of MDR strains of M. morganii and S. maltophilia isolated in our study.
Based on the observed antibiotic resistance profile by disc diffusion assay, we selected six isolates and further characterized their antibiotic-susceptibility array using Sensititer Gram-Negative GN2F plates. The data revealed resistance to beta-lactams, cephalosporins, quinolones, dihydrofolate reductase inhibitor/sulfonamide, and fluoroquinolones class of antibiotics and intermediate resistance to carbapenem antibiotic (i.e., imipenem) (Supplementary Table 1). Based on the updated definition of MDR viz. "the isolate is non-susceptible to at least 1 agent in ≥ 3 antimicrobial categories"53, all of the isolates tested herein using Sensititer Gram-Negative GN2F plates were characterized as MDR isolates. The widespread trade of shrimp can act as a vehicle for the dissemination of antibiotic-resistant genes. Currently, the FDA relies on only testing antibiotic residue54 in imported seafood. Hence, the presence of MDR strains in cooked shrimp samples points toward a gap in monitoring measures.
Despite emerging interest and research on marine and seafood-related microbiome, the microbial communities associated with shrimp gut remain relatively underexplored. In our study, the gut microbiota of shrimp samples collected from the U.S. had a higher abundance of Proteobacteria and Planctomyces and had a lower abundance of Tenericutes and Bacteroidetes. A study characterizing the gut microbiota of adult Litopenaeus vannamei raised in farms located in China reported that the adult shrimp gut from farm-A was dominated by Proteobacteria, Tenericutes, Firmicutes, Planctomyces, and Verrucomicrobia55. Parallel findings were reported wherein L. vannamei samples were collected from Mexico, showing a higher abundance of Proteobacteria, Cyanobacteria, Actinobacteria, Gemmatimonadetes, Bacteroidetes, and Firmicutes56.
Further, we observed higher bacterial species richness and diversity in wild-caught shrimps from the U.S. when compared to farm-raised shrimps from Ecuador. Rungrassamee et al. reported the presence of higher bacterial diversity in wild-caught Penaeus monodon (Tiger shrimp) compared to farmed-raised samples57. The major factor affecting the composition of aquatic organism gut microbiota are egg microbiota58, larval rearing water59, shrimp diet60,61,62, and environmental factors63. Fish on an herbivorous diet has been shown to have higher diversity compared to fish on a carnivorous diet60,61,62,64. Farm-raised shrimp are commonly fed an artificially formulated diet consisting of various animal proteins. A similar significant difference (p < 0.05) in the diversity indices were observed among the wild-caught and farm-raised L. vannamei samples collected from Mexico56, wherein Actinobacteria and Nitrospirae were found to be significantly abundant in wild-caught L. vannamei samples compared to Bacteroidetes, Gemmatimonadetes, Fusobacteria and Spirochaetes phyla, which were abundant in farm-raised samples56.
The two-dimensional PCoA plot for the beta-diversity for the shrimp samples from the U.S. and Ecuador showed clear clustering of representative samples into two distinct groups. Shrimp samples collected in our study were comprised of Pandalus borealis (Pink shrimp) and Litopenaeus setiferus (White shrimp). These two shrimp species clustered together in the PCoA plot, showing that the region of shrimp origin is a more crucial factor affecting gut microbiota than host genetics. Tzeng et al.65 characterized the gut microbiome composition of two closely related shrimp species, i.e., Macrobrachium nipponense (oriental river prawn) and M. asperulum, harvested from river and lake water located in Taiwan. Penaeus monodon samples were used as control. Data from the study showed that the host genetics and habitat both play important roles in determining the gut microbial composition of shrimp. Further, the PCoA plot from the study revealed that the host habitat has a much greater impact on the shrimp gut microbial composition. The effect of habitat on the gut microbial composition can be attributed to the similar diet and exposure to similar aquatic microbiota among different species of shrimp from a geographical region.
The PICRUSt data from a previous study showed an abundance of pathways dedicated to amino acid metabolism, lipid metabolism, and xenobiotics biodegradation in wild-caught L. vannamei56, which is similar to the PICRUSt data for wild-caught shrimp collected from the U.S. in our study. These distinct microbiome signatures observed among the U.S. wild-caught shrimp, when confirmed by further studies, could be used for designing region-specific probiotic therapies for improving shrimp health and can support the aquaculture of shrimp in the U.S. Pure culture strains of E. hormaechei were isolated in this study, which has been used in the past as a shrimp probiotic66.
In the present study, the prevalence of sulfonamide (blasul1 and blasul2), quinolone (blaqnrS), aminoglycoside (blaaadA), beta-lactam (blamecA and blaCMY), phenicol (blafloR and blacat), and tetracycline (blatetA) resistance genes was significantly higher in the gut of shrimp imported from Ecuador (Fig. 3). The higher abundance of these antibiotic-resistance genes in farm-raised shrimp can be attributed to the application of antibiotics for aquaculture operations facilitating the selection of strains with resistance genes. The presence of these genetic elements can facilitate the transfer of selected genes to genetically related and unrelated strains via the food chain67. A previous study has also shown a strong positive correlation of the presence of blasul1 and blaqnrD genes with the abundance of Proteobacteria; blacmlA and blafloR genes with the abundance of Bacteroidetes; blafloR and blasul2 with Firmicutes; and blacmlA, blatetG, blaqnrB, and blaqnrA with the abundance of Acidobacteria and Cyanobacteria55. Notably, these five phyla (i.e., Proteobacteria, Bacteroidetes, Firmicutes, Acidobacteria, and Cyanobacteria) were dominant phyla among farm-raised shrimp from Ecuador55.
One of the limitations of this study was our dependence on Sanger sequencing of 16S rDNA region for the identification of the obtained isolates. 16S rDNA sequencing is not very accurate for the differentiation of closely related species. To our knowledge, this is the first report demonstrating the isolation and prevalence of MDR strains of S. maltophilia from imported cooked shrimp. Selected cooked shrimp showed high APC as well as coliform counts. The data demonstrate that the presence of MDR bacterial strains and opportunistic pathogens in imported cooked shrimp products may pose a threat to human health and calls for the need for standardization of effective cooking time–temperature combination for potential mitigation of antibiotic-resistant strains in cooked shrimp samples. Further, data from our study can be used as baseline data for the U.S. wild-caught shrimp microbiome composition, and with larger validation studies, the microbiome markers identified in our study can be used for the development of novel PCR-based detection assays.
Methods
Shrimp samples
Raw (n = 18) and cooked (n = 13) shrimp samples were purchased from March to September 2019 from stores, supermarkets, and retailers located in the states of Florida and Georgia. The collected shrimp samples were either frozen pre-packaged or thawed shrimp placed on ice in bulk. The pre-packaged samples had a clear country of origin label. Similarly, thawed shrimp placed on ice in bulk collected from the supermarket had a country-of-origin label. Cooked shrimp samples were already peeled, deveined, and pre-cooked, and could be directly consumed after thawing. Sample details are listed in Table 1.
Aerobic plate count and coliform count
Twenty-five grams of cooked shrimp samples from each collected sample were weighed in Whirl–Pak filtered stomacher bags (Nasco, MI, U.S.), and samples were diluted using 225 mL of sterile 0.1% peptone water (w/v). Samples were stomached at 230 rpm for 2 min and were serially diluted using 9 mL of 0.1% (w/v) peptone water tubes. Diluted samples were plated on plate count agar (Hardy Diagnostics, CA, U.S.) and violet red bile agar (Hardy Diagnostics, CA, U.S.). Plates were incubated at 37 °C for 48 h, and colonies were enumerated after the incubation period, and obtained counts were expressed in log10 CFU/mL.
Isolation of Salmonella
Twenty-thirty grams of all shrimp samples were individually weighed in Whirl–Pak stomacher bags (Nasco, MI, U.S.). Samples were hand crushed and enriched with lactose broth (1:9) (Hardy Diagnostics, CA, U.S.) at 37 °C for 24 h. One mL and 0.1 mL of each enriched sample were transferred to freshly prepared 10 mL tetrathionate broth (T.T; Hardy Diagnostics, CA, U.S.) and Rappaport–Vassiliadis (R.V; Hardy Diagnostics, CA, U.S.) broths for selective enrichment, respectively. Inoculated samples in R.V. and T.T. tubes were incubated for 24 h at 42 ± 1 °C and 35 ± 1 °C, respectively. Samples after selective enrichment were streaked onto an XLD plate (Hardy Diagnostics, CA, U.S.) containing 4.6 mL/L tergitol (Niaproof; Sigma, MO, U.S.) and 15 mg/L of novobiocin (Alfa Aesar, MA, U.S.) and HE agar plates. After a 24 h incubation period, plates were observed for black colonies. DNA from black colonies was isolated using PrepMan Ultra Sample Preparation reagent (Applied Biosystems, CA, U.S.) and used for real-time PCR assay for specific detection of Salmonella68.
Isolation of ESBL and carbapenem-resistant bacterial strains
Collected shrimp samples were screened for the presence of ESBL resistant isolates as previously described. Briefly, 20–30 g of shrimp sample were transferred to Whirl–Pak stomacher bags (Nasco, MI, U.S.). As the exoskeleton of shrimp can rupture the stomacher bags, samples were hand crushed by applying pressure on the stomacher bags. Each shrimp sample was diluted with 225 mL of phosphate-buffered tryptic soy broth (TSB) with 2 µg/mL cefotaxime. After 18 h incubation, samples were streaked on following agars: (a) MacConkey agar with 4 µg/mL cefepime; (b) MacConkey agar with 4 µg/mL cefoxitin; (c) MacConkey agar with 8 µg/mL cefoxitin, 4 µg/mL cefepime, and 0.5 µg/mL meropenem. Two colonies with distinct colony morphology were selected from each plate and streaked on their respective plates for further purification. Isolated colonies were sub-cultured in TSB and preserved as a 20% (v/v) glycerol stock and stored in a − 20 °C freezer.
Molecular characterization of bacterial isolates
DNA was isolated from overnight broth cultures using PrepMan Ultra Sample Preparation reagent (Applied Biosystems, CA, U.S.). The quality and quantity of isolated DNA were measured using a NanoDrop One Spectrophotometer (Thermo Fisher Scientific, MA, U.S.), and samples were diluted to 20 ng/µL. The 16S rDNA sequences were amplified using 27F: AGAGTTTGATCMTGGCTCAG and 511R: GCGGCTGCTGGCACRKAGT primer pair69. Sanger sequencing of resulting amplicons were performed at the DNA Sequencing Facility of the Florida State University (Tallahassee, Florida). Sequence data were BLAST analyzed for the confirmation of genus and species identification of isolates.
Antibiotic susceptibility testing
The AST of all obtained isolates were performed using the disk diffusion method according to Clinical and Laboratory Standards Institute (CLSI) guidelines with Escherichia coli ATCC 25922 and Staphylococcus aureus ATCC 25923 as quality control strains. Enterobacteriaceae strains were tested for resistance to tetracycline (TE), beta-lactam antibiotics [aztreonam (ATM-30)], cephalosporins [cefotaxime (CTX-30), ceftazidime (CAZ-30)], carbapenem antibiotics [imipenem (IMP-10), ertapenem (ETP-10), meropenem (MEM-10)], fluoroquinolones [ciprofloxacin (CIP-5) and nalidixic acid (NA-30)] and phenicols [chloramphenicol (C-30)] (Hardy Diagnostics, CA, U.S.). Acinetobacter isolates were tested against IMP, CAZ, CTX, CIP, T.E., and MEM; Enterococcus isolates were tested against C, CIP, and T.E.; Stenotrophomonas isolates were tested against CAZ and C, and resistance of Vibrio isolates were tested using IMP, CAZ, CTX, T.E., and MEM antibiotic discs. Data obtained from the disc diffusion assay were used to categorize the strains as susceptible, intermediate, or resistant based on CLSI guidelines (document; M100-S23, M100-S24, and M100-S26)49,70,71.
The MIC of six selected strains was determined using GN2F Sensititer Gram-Negative plates (TREK Diagnostic Systems, OH, U.S.) as per manufacturer instructions. E. coli ATCC 25922 and S. aureus ATCC 25923 were used as quality control strains. The six isolates were selected based on MDR profiles towards different antibiotic classes observed by the disc diffusion assay, i.e., Enterobacter hormaechei 2B-MC1, Morganella morganii 31A-MNC, Proteus mirabilis 20B-MC1-1, Serratia marcescens 10A-MNC, Serratia marcescens 28B-MC2, and Vibrio parahaemolyticus 24B-MC2. The MIC data were analyzed based on CLSI guidelines (document M100-S29)72 and M45-02, 2010 edition for Vibrio parahaemolyticus73.
Detection of antibiotic-resistance genes using real-time PCR assays
Pure culture bacterial DNA isolated from broth culture was used for real-time PCR assays. DNA samples were tested for the presence of antibiotic resistance genes (blaKPC, blaOXA-48, blaNDM, blaVIM, blaCTX-M, blaIMP, blaCMY, blaACC, blaSHV, and blaTEM) by real-time PCR assay as previously described55,68,74,75.
Microbiome analysis
Shrimp gut microbiome was analyzed according to our previously described methods76,77,78. In brief, the Earth Microbiome Project (EMP) benchmarked protocol79 was used for the characterization of shrimp gut microbiome. Microbial community analysis, protocols, and standards80 were employed by following a barcoded high-throughput sequencing approach, as described by Caporaso et al.81. Bacterial genomic DNA from the gastrointestinal tract of raw shrimp samples (Table 2) from the U.S. and Ecuador was extracted using the PowerFecal DNA kit (Qiagen, CA, U.S.). The V4 hypervariable region of the 16S rDNA gene was PCR-amplified using the universal primer pair 515F/806R81; the resulting uniquely barcoded amplicons were purified using Agencourt AMPure XP magnetic purification beads (Beckman Coulter, CA, U.S.) and quantified with Qubit fluorimeter (Qubit3; Invitrogen, CA, U.S.) and the dsDNA H.S. assay kit (Life Technologies, CA, U.S.); and the amplicon library was generated according to methods of Caporaso et al.81. The purified PCR products were pooled in equal molar concentrations and sequenced on one 2 × 300-bp Illumina MiSeq run (Miseq reagent kit v3; Illumina Inc., CA, U.S.) for paired-end sequencing. The sequencing quality control was carried out with on-board Miseq Control Software and Miseq Reporter (Illumina Inc.). The resultant sequences were de-multiplexed, quality-filtered, clustered and taxonomically-assigned (at 97% similarity level against GreenGenes database, May 2013 version) with Ribosomal Database Project (RDP)-classifier workflow as described by Wang et al.82 using QIIME software suite83 as described previously76,78. Quality filtering was performed as described previously84. Subsequent data processing included filtering by clustering similar sequences with less than 3% dissimilarity by using the USEARCH algorithm version 5.2.3285 with an open-reference OTU clustering method using the Greengenes database and detecting and removing chimeras by using the UCHIME algorithm86. The most abundant sequence in each OTU was selected as the representative sequence and the resulting OTUs were assigned to taxa by using the Ribosomal Database Project classifier82 trained on the Greengenes reference database. A total of 2,183,494 sequences (mean ± SEM: 77,981.929 ± 7707.829) were obtained after quality-filtering. To avoid the bias of sequencing depth, the sequences were rarefied to the minimum sequence reads per sample for downstream analyses. To avoid the bias of sequencing errors or low-level contaminations, the OTUs with very low read count (less than 4) in very few samples (less than 10% prevalence) were filtered out from the subsequent analyses. The data of taxon abundance were subjected to the total sum scaling, and the taxa with less than 1% mean relative abundance were further excluded from the subsequent downstream analyses. To avoid the bias of nucleic acid extraction, PCR reaction conditions, and primers on community composition obtained by amplicon sequencing, all the samples were batch-processed identically and simultaneously. Bacterial community composition was measured at taxonomic levels of phylum, class, order, family, and genus. Alpha-diversity measures were calculated within QIIME. Beta-diversity was computed using principal coordinate analysis (PCoA) of the Bray–Curtis dissimilarity index, as described previously87. The functional activities of the bacterial communities were analyzed using the open-source bioinformatics tool PICRUSt (Phylogenetic Investigation of Communities by Reconstruction of Unobserved States; http://picrust.github.io/picrust/index.html) as described previously88, wherein the sequencing dataset was normalized and the metagenomes were predicted against the PICRUSt-formatted, characterized-protein, functional database of Kyoto Encyclopedia of Genes and Genomes (KEGG www.kegg.jp/kegg/kegg1.html)89.
Prevalence of antibiotic resistance genes in raw shrimp gut
Prevalence of antibiotic resistance genes in shrimp gut was determined using DNA isolated from shrimp gut and real-time PCR assays. Presence of antibiotic resistance gene (i.e., blasul-1, blasul-2, blaqnrS, blaqnrD, blaaadA, blaTEM, blaSHV, blaCTX-M, blaCMY, blaKPC, blaNDM, blamecA, blavanA, blavanB, blavanC, blacat, blacmlA, blafloR, and blatetA) were tested as using primer-pair and PCR conditions as previously described55,68,75,90.
Data analysis
Alpha-diversity indices and bacterial abundance data between the two groups of shrimps were compared by two-tailed unpaired t-test. The data of the prevalence of antibiotic resistance and the associated genes were compared with Fisher's exact test. LEfSe (Linear discriminatory analysis [LDA] Effect Size) was applied to identify the bacterial taxa driving the differences between the different groups of shrimps91, wherein the alpha parameter significance threshold for the Kruskal–Wallis as well as the Wilcoxon test implemented among classes was set to 0.01, and the logarithmic LDA score cut-off was set to 3.0. Bray–Curtis similarity scores inferred from the taxonomic data were reduced to a two-dimensional space using principal coordinate analysis (PCoA) for the computation of structural similarity (beta-diversity) of bacteriomes from different groups of shrimps. Differences in beta-diversity were computed by permutational multivariate analysis of variance (PERMANOVA) using the algorithm of web-based tool Microbiome Analyst92. Hierarchical clustering heatmaps were constructed within 'R' statistical software package (version 3.6.1; https://www.r-project.org/) using the 'heatmap.2’ and "ggplots" packages. Unless otherwise stated, the data are presented as means ± SEM. P < 0.05 was considered statistically significant unless specified.
Data availability
All data generated or analyzed during this study are included in this published article (and its Supplementary Information files). The raw sequences have been deposited at the Sequence Read Archive (SRA) under the SRA number (SUB7840299) and BioProject number PRJNA648917 for unrestricted, public access. Other information/ data related to the current study are available from the corresponding author on reasonable request.
Abbreviations
- APC:
-
Aerobic plate count
- ATM-30:
-
Aztreonam
- C-30:
-
Chloramphenicol
- CAZ-30:
-
Ceftazidime
- CFU:
-
Colony-forming unit
- CIP-5:
-
Ciprofloxacin
- CTX-30:
-
Cefotaxime
- EMP:
-
Earth microbiome project
- ESBL:
-
Extended-spectrum β-lactam
- ETP-10:
-
Ertapenem
- HE:
-
Hektoen enteric
- IMP-10:
-
Imipenem
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- LDA:
-
Linear discriminatory analysis
- LEfSe:
-
Linear discriminatory analysis effect size
- MEM-10:
-
Meropenem
- NA-30:
-
Nalidixic acid
- PCoA:
-
Principal coordinate analysis
- PICRUSt:
-
Phylogenetic Investigation of Communities by Reconstruction of Unobserved States
- R.V.:
-
Rappaport–Vassiliadis
- T.E.:
-
Tetracycline
- TSB:
-
Tryptic soy broth
- T.T.:
-
Tetrathionate broth
- XLDtn:
-
Xylose lysine deoxycholate agar with tergitol
References
Centers for Disease Control and Prevention (CDC). Biggest threats and data. https://www.cdc.gov/drugresistance/biggest-threats.html (2019).
O'Neill, J. Review on antimicrobial resistance: tackling a crisis for the health and wealth of nations. 2014. London: HM Government (2016).
United States Department of Agriculture (USDA). Import share of consumption. http://www.ers.usda.gov/topics/international-markets-trade/us-agricultural-trade/import-share-of-consumption.aspxExternal (2016).
National Oceanic and Atmospheric Administration (NOAA). Shrimp imports. https://www.st.nmfs.noaa.gov/apex/f?p=169:2 (2020).
Holmström, K. et al. Antibiotic use in shrimp farming and implications for environmental impacts and human health. Int. J. Food Sci. Technol. 38, 255–266 (2003).
Food and Agriculture Organization (FAO). Report of the FAO/MARD technical workshop on early mortality syndrome (EMS) or acute hepatopancreatic necrosis syndrome (AHPNS) of cultured shrimp (under TCP/VIE/3304). (2013).
Thornber, K. et al. Evaluating antimicrobial resistance in the global shrimp industry. Rev. Aquac. 12, 966–986 (2020).
Hinchliffe, S., Butcher, A. & Rahman, M. M. The AMR problem: Demanding economies, biological margins, and co-producing alternative strategies. Palgrave Commun. 4, 1–12 (2018).
Chi, T. T. K., Clausen, J. H., Van, P. T., Tersbøl, B. & Dalsgaard, A. Use practices of antimicrobials and other compounds by shrimp and fish farmers in Northern Vietnam. Aquac. Rep. 7, 40–47 (2017).
Ali, H., Rico, A., Murshed-e-Jahan, K. & Belton, B. An assessment of chemical and biological product use in aquaculture in Bangladesh. Aquaculture 454, 199–209 (2016).
Uddin, S. A. & Kader, M. A. The use of antibiotics in shrimp hatcheries in Bangladesh. J. Fish. Aquat. Sci 1, 64–67 (2006).
Elbashir, S. et al. Seafood pathogens and information on antimicrobial resistance: A review. Food Microbiol. 70, 85–93 (2018).
Roque, A., Molina-Aja, A., Bolán-Mejıa, C. & Gomez-Gil, B. In vitro susceptibility to 15 antibiotics of vibrios isolated from penaeid shrimps in Northwestern Mexico. Int. J. Antimicrob. Agents 17, 383–387 (2001).
Soto-Rodríguez, S., Armenta, M. & Gomez-Gil, B. Effects of enrofloxacin and florfenicol on survival and bacterial population in an experimental infection with luminescent Vibrio campbellii in shrimp larvae of Litopenaeus vannamei. Aquaculture 255, 48–54 (2006).
Zhang, Y. B., Li, Y. & Sun, X. L. Antibiotic resistance of bacteria isolated from shrimp hatcheries and cultural ponds on Donghai Island, China. Mar. Pollut. Bull. 62, 2299–2307 (2011).
Cabello, F. C. Heavy use of prophylactic antibiotics in aquaculture: A growing problem for human and animal health and for the environment. Environ. Microbiol. 8, 1137–1144 (2006).
Samuelsen, O. B., Lunestad, B. T., Husevåg, B., Hølleland, T. & Ervik, A. Residues of oxolinic acid in wild fauna following medication in fish farms. Dis. Aquat. Org. 12, 111–119 (1992).
Stryker, R. B., Dixon, E. M. & Rippen, T. E. Process to produce safe pasteurized shrimp and other shellfish of high sensory quality and extended refrigerated shelf-life. U.S. Patent Application No. 14/590,501 (Google patent, 2015).
Food and Drug Administration (FDA). Control of Listeria monocytogenes in ready-to-eat foods: guidance for industry. Draft Guidance, US Department of Health and Human Services Food and Drug Administration Center for Food Safety and Applied Nutrition (2017).
Duffy, G., Walsh, C., Blair, I. & McDowell, D. Survival of antibiotic resistant and antibiotic sensitive strains of E. coli O157 and E. coli O26 in food matrices. Int. J. Food Microbiol. 109, 179–186 (2006).
Ebinesh, A., Vijaykumar, G. & Kiran, T. Exposure to stress minimizes the zone of antimicrobial action: A phenotypic demonstration with six Acinetobacter baumannii strains. MicroMedicine 6, 16–35 (2018).
Hassan, M. et al. Monitoring the presence of chloramphenicol and nitrofuran metabolites in cultured prawn, shrimp and feed in the Southwest coastal region of Bangladesh. Egypt. J. Aquat. Res. 39, 51–58 (2013).
Vikram, A. et al. Similar levels of antimicrobial resistance in US food service ground beef products with and without a “Raised without Antibiotics” claim. J. Food Prot. 81, 2007–2018 (2018).
Ren, D. et al. Phenotypes and antimicrobial resistance genes in Salmonella isolated from retail chicken and pork in Changchun, China. J. Food Saf. 37, e12314 (2017).
Tien, Y.-C. et al. Impact of dairy manure pre-application treatment on manure composition, soil dynamics of antibiotic resistance genes, and abundance of antibiotic-resistance genes on vegetables at harvest. Sci. Total Environ. 581, 32–39 (2017).
Wang, F.-H., Qiao, M., Chen, Z., Su, J.-Q. & Zhu, Y.-G. Antibiotic resistance genes in manure-amended soil and vegetables at harvest. J. Hazard. Mater. 299, 215–221 (2015).
Global Aquaculture Alliance (GAA). GAA response to CBC marketplace about detection of antibiotic-resistant bacteria in retail samples of shrimp. https://www.aquaculturealliance.org/wp-content/uploads/2019/03/GAA_Statement_To_CBC_Marketplace_FINAL-1.pdf (2019).
Bass, D., Stentiford, G. D., Wang, H.-C., Koskella, B. & Tyler, C. R. The pathobiome in animal and plant diseases. Trends Ecol. Evol. 34, 996–1008 (2019).
Lawley, T. D. & Walker, A. W. Intestinal colonization resistance. Immunology 138, 1–11 (2013).
Holt, C. C., Bass, D., Stentiford, G. D. & van der Giezen, M. Understanding the role of the shrimp gut microbiome in health and disease. J. Invert. Pathol. 107387 (2020).
Indugu, N., Sharma, L., Jackson, C. R. & Singh, P. Whole-genome sequence analysis of multidrug-resistant Enterobacter hormaechei isolated from imported retail shrimp. Microbiol. Resour. Announc. 9, e01103-20 (2020).
Huss, H. H. Assurance of Seafood Quality. 169 (Food & Agriculture Org., 1994).
Nawaz, M. et al. Characterisation of novel mutations involved in quinolone resistance in Escherichia coli isolated from imported shrimp. Int. J. Antimicrob. Agents 45, 471–476 (2015).
Liao, X. et al. Interplay of antibiotic resistance and food-associated stress tolerance in foodborne pathogens. Trends Food Sci. Technol. 95, 97–106 (2020).
Hwang, D., Kim, S. M. & Kim, H. J. Modelling of tetracycline resistance gene transfer by commensal Escherichia coli food isolates that survived in gastric fluid conditions. Int. J. Antimicrob. Agents 49, 81–87 (2017).
Seyfried, E. E., Newton, R. J., Rubert, K. F., Pedersen, J. A. & McMahon, K. D. Occurrence of tetracycline resistance genes in aquaculture facilities with varying use of oxytetracycline. Microb. Ecol. 59, 799–807 (2010).
FDA. Approved aquaculture drugs. https://www.fda.gov/animal-veterinary/aquaculture/approved-aquaculture-drugs (2020).
Tendencia, E. A. & de la Peña, L. D. Antibiotic resistance of bacteria from shrimp ponds. Aquaculture 195, 193–204 (2001).
Shakila, R. J., Vyla, S. A. P., Kumar, R. S., Jeyasekaran, G. & Jasmine, G. I. Stability of chloramphenicol residues in shrimp subjected to heat processing treatments. Food Microbiol. 23, 47–51 (2006).
Food and Drug Administration (FDA). Approved aquaculture drugs. https://www.accessdata.fda.gov/cms_ia/importalert_29.html (2020).
Adegoke, A. A., Stenström, T. A. & Okoh, A. I. Stenotrophomonas maltophilia as an emerging ubiquitous pathogen: Looking beyond contemporary antibiotic therapy. Front. Microbiol. 8, 2276 (2017).
Hazards, E. P. O. B. Scientific Opinion on Carbapenem resistance in food animal ecosystems. EFSA J. 11, 3501 (2013).
McDermott, P. The NARMS seafood pilot project. https://www.hhs.gov/sites/default/files/7.3-mcdermott-narms-508.pdf (2020).
Almeida, M. V. A. d., Cangussú, Í. M., Carvalho, A. L. S. d., Brito, I. L. P. & Costa, R. A. Drug resistance, AmpC-β-lactamase and extended-spectrum β-lactamase-producing Enterobacteriaceae isolated from fish and shrimp. Revista do Instituto de Medicina Tropical de São Paulo. 59, e70 (2017).
Sanjit Singh, A., Lekshmi, M., Prakasan, S., Nayak, B. B. & Kumar, S. Multiple antibiotic-resistant, extended spectrum-β-Lactamase (ESBL)-producing enterobacteria in fresh seafood. Microorganisms 5, 53 (2017).
Food and Drug Administration (FDA). Import alert 16–127. https://www.accessdata.fda.gov/cms_ia/importalert_29.html (2020).
Food and Drug Administration (FDA). Import alert 16–129. https://www.accessdata.fda.gov/cms_ia/importalert_31.html (2020).
Southern Shrimp Alliance (SSA). Indian shrimp shipment again rejected for presence of banned antibiotics in February. https://www.shrimpalliance.com/indian-shrimp-shipment-again-rejected-for-presence-of-banned-antibiotics-in-february/ (2020).
Clinical and Laboratory Standards Institute (CLSI). Performance standards for antimicrobial susceptibility testing; twenty-fourth informational supplement (M100-S24). (Clinical and Laboratory Standards Institute, 2014).
Government Accountability Office (GAO). Seafood safety: FDA needs to improve oversight of imported seafood and better leverage limited resources. http://www.gao.gov/assets/320/317734.pdf (2011).
Sy, N. V. et al. Residues of 2-hydroxy-3-phenylpyrazine, a degradation product of some β-lactam antibiotics, in environmental water in Vietnam. Chemosphere 172, 355–362 (2017).
Yamasaki, S., Le, T. D., Vien, M. Q., Van Dang, C. & Yamamoto, Y. Prevalence of extended-spectrum β-lactamase-producing Escherichia coli and residual antimicrobials in the environment in Vietnam. Anim. Health Res. Rev. 18, 128–135 (2017).
Magiorakos, A. P. et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 18, 268–281 (2012).
Food and Drug Administration (FDA). Import Alert 16–131. https://www.accessdata.fda.gov/cms_ia/importalert_33.html (2020).
Su, H. et al. Persistence and spatial variation of antibiotic resistance genes and bacterial populations change in reared shrimp in South China. Environ. Int. 119, 327–333 (2018).
Cornejo-Granados, F. et al. Microbiome of Pacific Whiteleg shrimp reveals differential bacterial community composition between Wild, Aquacultured and AHPND/EMS outbreak conditions. Sci. Rep. 7, 1–15 (2017).
Rungrassamee, W. et al. Characterization of intestinal bacteria in wild and domesticated adult black tiger shrimp (Penaeus monodon). PLoS ONE 9, e91853 (2014).
Bell, G. R., Hoskins, G. E. & Hodgkiss, W. Aspects of the characterization, identification, and ecology of the bacterial flora associated with the surface of stream-incubating Pacific salmon (Oncorhynchus) eggs. J. Fish. Board Can. 28, 1511–1525 (1971).
Giatsis, C. et al. The impact of rearing environment on the development of gut microbiota in tilapia larvae. Sci. Rep. 5, 1–15 (2015).
Li, J. et al. Comparative study on gastrointestinal microbiota of eight fish species with different feeding habits. J. Appl. Microbiol. 117, 1750–1760 (2014).
Li, X., Zhu, Y., Yan, Q., Ringø, E. & Yang, D. Do the intestinal microbiotas differ between paddlefish (Polyodon spathala) and bighead carp (Aristichthys nobilis) reared in the same pond?. J. Appl. Microbiol. 117, 1245–1252 (2014).
Miyake, S., Ngugi, D. K. & Stingl, U. Diet strongly influences the gut microbiota of surgeonfishes. Mol. Ecol. 24, 656–672 (2015).
Huang, F., Pan, L., Song, M., Tian, C. & Gao, S. Microbiota assemblages of water, sediment, and intestine and their associations with environmental factors and shrimp physiological health. Appl. Microbiol. Biotechnol. 102, 8585–8598 (2018).
Cornejo-Granados, F., Gallardo-Becerra, L., Leonardo-Reza, M., Ochoa-Romo, J. P. & Ochoa-Leyva, A. A meta-analysis reveals the environmental and host factors shaping the structure and function of the shrimp microbiota. PeerJ 6, e5382 (2018).
Tzeng, T.-D. et al. Effects of host phylogeny and habitats on gut microbiomes of oriental river prawn (Macrobrachium nipponense). PLoS ONE 10, e0132860 (2015).
Knipe, H., Temperton, B., Lange, A., Bass, D. & Tyler, C. R. Probiotics and competitive exclusion of pathogens in shrimp aquaculture. Rev. Aquac. 13, 324–352 (2020).
Singh, B., Tyagi, A., Billekallu Thammegowda, N. K. & Ansal, M. D. Prevalence and antimicrobial resistance of vibrios of human health significance in inland saline aquaculture areas. Aquac. Res. 49, 2166–2174 (2018).
Singh, P., Pfeifer, Y. & Mustapha, A. Multiplex real-time PCR assay for the detection of extended-spectrum β-lactamase and carbapenemase genes using melting curve analysis. J. Microbiol. Methods 124, 72–78 (2016).
Liu, A.-C., Chou, C.-Y., Chen, L.-L. & Kuo, C.-H. Bacterial community dynamics in a swine wastewater anaerobic reactor revealed by 16S rDNA sequence analysis. J. Biotechnol. 194, 124–131 (2015).
Cockerill, F. et al. Performance Standards for Antimicrobial Susceptibility Testing: Twenty-Third Informational Supplement; M100–S23 (CLSI, Wayne, 2013).
Clinical and Laboratory Standards Institute (CLSI). Performance standards for antimicrobial susceptibility testing: Vol. M100. (Clinical and Laboratory Standards Institute, 2016).
Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing: Supplement M100 (ed 29) (Clinical & Laboratory Standards Institute, 2019).
Clinical and Laboratory Standards Institute (CLSI). Methods for antimicrobial dilution and disk susceptibility testing of infrequently isolated or fastidious bacteria; Approved guideline—second edition. (Clinical and Laboratory Standards Institute, 2010).
Singh, P. & Mustapha, A. Development of a real-time PCR melt curve assay for simultaneous detection of virulent and antibiotic resistant Salmonella. Food Microbiol. 44, 6–14 (2014).
Vikram, A. et al. Impact of “raised without antibiotics” beef cattle production practices on occurrences of antimicrobial resistance. Appl. Environ. Microbiol. 83, e01682-e11617 (2017).
Nagpal, R., Neth, B. J., Wang, S., Craft, S. & Yadav, H. Modified Mediterranean-ketogenic diet modulates gut microbiome and short-chain fatty acids in association with Alzheimer’s disease markers in subjects with mild cognitive impairment. EBioMedicine 47, 529–542 (2019).
Nagpal, R. et al. Gut microbiome composition in non-human primates consuming a Western or Mediterranean diet. Front. Nutr. 5, 28 (2018).
Ahmadi, S. et al. Prebiotics from acorn and sago prevent high-fat-diet-induced insulin resistance via microbiome–gut–brain axis modulation. J. Nutr. Biochem. 67, 1–13 (2019).
Thompson, L. R. et al. A communal catalogue reveals Earth’s multiscale microbial diversity. Nature 551, 457–463 (2017).
Earth microbiome project (EMP). Protocols and standards: Earth microbiome project. https://earthmicrobiome.org/protocols-and-standards/ (2020).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–1624 (2012).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73, 5261–5267 (2007).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335 (2010).
Bokulich, N. A. et al. Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat. Methods 10, 57–59 (2013).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200 (2011).
Chong, J., Liu, P., Zhou, G. & Xia, J. Using MicrobiomeAnalyst for comprehensive statistical, functional, and meta-analysis of microbiome data. Nat. Protoc. 15, 799–821 (2020).
Langille, M. G. et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 31, 814 (2013).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Stedtfeld, R. D. et al. Primer set 2.0 for highly parallel qPCR array targeting antibiotic resistance genes and mobile genetic elements. FEMS Microbiol. Ecol. 94, fiy130 (2018).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Dhariwal, A. et al. MicrobiomeAnalyst: A web-based tool for comprehensive statistical, visual and meta-analysis of microbiome data. Nucleic Acids Res. 45, W180–W188 (2017).
Acknowledgements
The authors would like to thank John William (Southern Shrimp Alliance) and Bob Jones (Florida Fisheries Foundation LLC) for their help in shrimp sample collection.
Funding
The study was funded by the Florida State University startup funds.
Author information
Authors and Affiliations
Contributions
P.S.: conceived, designed and supervised the study; L.S.: performed experiments; R.N.: analyzed microbiome data; P.S., C.J., R.N., L.S.: interpreted data; P.S.: collected sample; L.S., P.S., D.P.: bacterial isolation; P.S., L.S., R.N.: wrote the manuscript; R.N., C.J.: revised manuscript; P.S., L.S., R.N., C.J., D.P.: approved the final version of the manuscript. All the authors reviewed data, read the manuscript, and provided critical feedback to be included in the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sharma, L., Nagpal, R., Jackson, C.R. et al. Antibiotic-resistant bacteria and gut microbiome communities associated with wild-caught shrimp from the United States versus imported farm-raised retail shrimp. Sci Rep 11, 3356 (2021). https://doi.org/10.1038/s41598-021-82823-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-82823-y
This article is cited by
-
Response of gut microbiota, antioxidation, and disease resistance to Pacific shrimp fed distiller’s dried grains with solubles replaced soybean meal
Aquaculture International (2024)
-
The CRISPR/Cas system as an antimicrobial resistance strategy in aquatic ecosystems
Functional & Integrative Genomics (2024)
-
In vivo assessment of Lactobacillus plantarum strains in black tiger shrimp (Penaeus monodon): implications for growth performance, probiotic-pathogen interaction, and defense against AHPND infection
Aquaculture International (2024)
-
Supplementation of ex situ produced bioflocs improves immune response against AHPND in Pacific whiteleg shrimp (Litopenaeus vannamei) postlarvae
Applied Microbiology and Biotechnology (2022)
-
Selection of Beneficial Bacterial Strains With Potential as Oral Probiotic Candidates
Probiotics and Antimicrobial Proteins (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.