Abstract
This study was conducted to investigate the impact of genetic variants of immune checkpoint genes on the treatment outcome in small cell lung cancer (SCLC). In the present study, 261 platinum doublet-treated SCLC patients were enrolled. A total of 96 polymorphisms in 33 immune checkpoint-related genes were selected, and their association with chemotherapy response and survival outcomes were analyzed. Among the polymorphisms studied, CD155 rs1058402G > A (Ala67Thr, A67T) and CD226 rs763361C > T (Gly307Ser, G307S) were significantly associated with SCLC treatment outcome. The rs1058402G > A had a worse chemotherapy response and overall survival (under a dominant model, adjusted odds ratio [aOR] = 0.52, 95% confidence interval [CI] = 0.27–0.99, P = 0.05; adjusted hazard ratio [aHR] = 1.55, 95% CI = 1.12–2.14, P = 0.01, respectively). The rs763361C > T had better chemotherapy response and overall survival (under a dominant model, aOR = 2.03, 95% CI = 1.10–3.75, P = 0.02; aHR = 0.69, 95% CI = 0.51–0.94, P = 0.02, respectively). When the rs1058402GA/AA and rs763361CC genotypes were combined, the chemotherapy response and overall survival were significantly decreased as the number of bad genotypes increased (aOR = 0.52, 95% CI = 0.33–0.81, Ptrend = 0.004; aHR = 1.48, 95% CI = 1.19–1.84, Ptrend = 4 × 10−4, respectively). The 3-D structural model showed that CD155 A67T created a new hydrogen bond and structural change on CD155. These changes resulted in extending the distance and losing the hydrogen bonds between CD155 and CD226, thus weakening CD155/CD226 binding activity. In conclusion, CD155 rs1058402G > A and CD226 rs763361C > T may be useful for predicting the clinical outcomes of SCLC patients after chemotherapy.
Similar content being viewed by others
Introduction
Lung cancer is still the leading cause of cancer-related deaths1. Approximately 15% of lung cancer is categorized as small cell lung cancer (SCLC), which is characterized as having a more rapid doubling time, higher growth rate, and earlier metastasis than non-small cell lung cancer (NSCLC)2. Smoking is the most potent known cause of SCLC2. One-third of patients with SCLC are diagnosed with limited-stage disease if the cancer is confined to the ipsilateral hemithorax. The others are diagnosed with extensive-stage disease if it is beyond the ipsilateral hemithorax, including distant metastases. The development of SCLC therapy has been stagnant for decades, although there has been recent improvement in the treatment of NSCLC3. The combined modality treatment with chemotherapy and radiotherapy is the current treatment standard for limited-stage SCLC2. Chemotherapy with platinum doublet is the cornerstone of therapy for extensive-stage SCLC. SCLC is very sensitive to initial chemotherapy, but most SCLC patients relapse and eventually die. Therefore, efforts are needed to predict chemotherapy responses and find prognostic markers.
The immune system defends our bodies from infectious organisms and other invaders. The immune system is also important in preventing and eradicating cancers4. In general, immune cells can recognize tumor cells and destroy them. However, cancer cells have developed ways of escaping the immune system by suppressing the activation of immune cells5. Therefore, cancer cells can survive and spread beyond the patient’s immune system. Many researchers have tried to control immune-escaping tumors5,6,7. Immune checkpoint inhibitors are drugs that can block inhibitor signals from cancer cells, restore the immune system, and reactivate immune cells to kill cancer cells. Recently, immune checkpoint inhibitors have been actively used in the treatment of cancers, including lung cancer8,9,10,11.
The genetic variants in immune checkpoints may affect the body’s response to chemotherapy or lung cancer prognosis, considering the immune system’s ability to prevent and kill cancer cells. We previously reported that genetic variants in immune checkpoint genes were associated with the prognosis of surgically resected NSCLC12. Programmed cell death-ligand 1 (PD-L1) polymorphisms were associated with chemotherapy response and clinical outcomes of advanced NSCLC13. Nomizo et al. reported that the polymorphisms in immune checkpoint genes had an effect on the response rate and progression survival in NSCLC patients with nivolumab treatment14. However, only a few studies had been conducted regarding SCLC. Therefore, we investigated the effects of genetic variants in immune checkpoint genes on the chemotherapy response and prognosis in SCLC.
Materials and methods
Study population
This study is observational retrospective study. This study included 261 patients diagnosed with SCLC at Kyungpook National University Hospital (KNUH) in Daegu, Korea between March 2001 and November 2017. The flow of selection of the patients is shown in Fig. 1. All the patients were treated with at least two cycles of platinum doublet chemotherapy as first-line treatment. Patients with limited-stage SCLC who underwent chemotherapy with concurrent radiotherapy were excluded to avoid the confounding effect of radiation on the chemotherapy response. If radiotherapy was conducted sequentially after 2 cycles of chemotherapy, these patients were included. In this case, the response after 2 cycles of chemotherapy was considered as the best response to chemotherapy. There were two chemotherapy regimens. One consists of cisplatin 60 mg/m2 on day 1 and etoposide 100 mg/m2 on day 1, 2, and 3 every 3 weeks. The other consists of cisplatin 60 mg/m2 on day 1 and irinotecan 60 mg/m2 on day 1, 8, and 15 every 4 weeks. The treatment was discontinued in case of disease progression, major toxicities, or according to patient’s or physician’s decision. Response assessment was carried out every two cycles of chemotherapy using the Response Evaluation Criteria in Solid Tumors15. Chemotherapy response fell into two categories: responder and non-responder. Responder means the best response was a complete response or a partial response. A stable disease or progressive disease was defined as non-responder.
This study was approved by the Institutional Review Boards of KNUH. The blood samples for genotyping were provided by the National Biobank of KNUH, which is supported by the Ministry of Health, Welfare, and Family Affairs. All blood samples were obtained before the 1st chemotherapy session. All subjects are 18 years of age or older and informed consent was obtained prior to chemotherapy.
Polymorphism selection and genotyping
We selected 38 genes involved in the immune checkpoint pathway by searching public databases and related literatures16,17,18. To collect polymorphisms in immune checkpoint genes, we searched the public single-nucleotide polymorphisms (SNPs) database (http://www.ncbi.nlm.nih.gov/SNP) and RegulomeDB (http://www.regulomedb.org/). After excluding minor allele frequencies of ≤ 0.05 by the HapMap JPT data, 216 potentially functional SNPs were collected using the FuncPred utility for functional SNP prediction in the SNPinfo web server (https://snpinfo.niehs.nih.gov/). After excluding those in linkage disequilibrium (LD) (r2 ≥ 0.8) using the TagSNP utility for LD tag SNP selection, 123 SNPs were selected for genotyping. Among the 123 SNPs, 27 of them with call rates of < 95% or P value for Hardy–Weinberg equilibrium (HWE) of < 0.05 were excluded from further analysis. Finally, the remaining 96 SNPs in 33 immune checkpoint genes were analyzed for the association and response study (Supplementary Table S1). Genotyping was performed using Sequenom MassARRAY iPLEX Platform (Agena Bioscience, San Diego, USA). All genotyping was conducted blindly with respect to patient status. All methods were performed in accordance with the relevant guidelines and regulations.
Statistical analysis
The HWE was tested using a goodness-of-fit χ2 test with one degree of freedom. For comparison between clinical variables or genotypes and chemotherapy response, the odds ratio (OR) and 95% confidence interval (CI) were calculated using unconditional logistic regression analysis. Overall survival (OS) was counted from the 1st chemotherapy session date to death or last follow-up. Progression-free survival (PFS) was measured from the day of 1st chemotherapy until disease progression or death from any cause. Kaplan–Meier method was used to estimate the survival outcome. Log-rank test was used to compare OS across different groups or genotypes. Multivariate Cox proportional hazards models were used to estimate the hazard ratio (HR) and 95% CIs. Adjusting factors were age, gender, smoking status, stage, Eastern Cooperative Oncology Group (ECOG) performance status, weight loss, neuron specific enolase (NSE) level, 1st chemotherapy regimen, second-line chemotherapy, and radiation to primary tumor. A P value of less than 0.05 was considered statistically significant. Statistical analyses were performed using the Statistical Analysis System for Windows, version 9.4 (SAS Institute, Cary, NC, USA).
Structure modeling of CD155 A67T variant.
In order to evaluate the effect of the rs1058402G > A of CD155, which substitutes alanine with threonine at codon 67, The mutant structure of CD155(A67T) was modeled using MODELLER v9.12 with the crystal structures of wildtype CD155 (PDB : 6ISC). The least violated 10 structures were further refined by model/refine loops (loop modeling protocol DOPE) using UCSF Chimera v1.14 and a structure showed no violation and the lowest energy was selected as a model. All images of CD226/CD155 (wild type vs. A67T mutant) were made in PyMOL (https://pymol.org/2/).
Results
Patient characteristics and clinical predictors
Chemotherapy response and OS according to patients’ clinical characteristics are shown in Table 1. The overall response rate was 72.8% (95% CI = 67.4–78.2%). The median survival time (MST) was 10.4 months (95% CI = 9.1–11.1 months). The response rate was higher in the irinotecan/cisplatin regimen than the etoposide/cisplatin regimen (78.7% vs. 67.2%, P = 0.04), but OS was not different based on regimens (MST, 10.0 months vs. 10.7 months, P = 0.88, Table 1). OS was associated with age, stage, ECOG performance status, weight loss, NSE level, second-line chemotherapy, and radiation to tumor (Table 1).
Effect of polymorphisms on treatment outcome
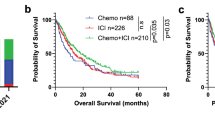
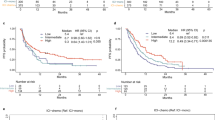
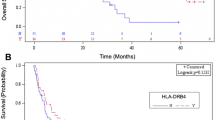
Among the observed 96 polymorphisms, CD155 rs1058402G > A (Ala67Thr, A67T) and CD226 rs763361C > T (Gly307Ser, G307S) were associated with both chemotherapy response and OS. The rs1058402G > A was significantly associated with worse chemotherapy response and OS (under a dominant model, adjusted OR [aOR] = 0.52, 95% CI = 0.27–0.99, P = 0.05 and adjusted HR [aHR] = 1.55, 95% CI = 1.12–2.14, P = 0.01; Table 2 and Fig. 2A). The effect of rs1058402 on PFS had the same trend as OS, although it was not statistically significant (under a dominant model, aHR = 1.30, 95% CI = 0.96–1.76, P = 0.09). The rs763361C > T showed significantly better chemotherapy response, OS, and PFS, respectively (under a dominant model, adjusted aOR = 2.03, 95% CI = 1.10–3.75, P = 0.02; aHR = 0.69, 95% CI = 0.51–0.94, P = 0.02; aHR = 0.73, 95% CI = 0.54–0.97, P = 0.03, respectively, Table 2 and Fig. 2B).
We also analyzed survival outcomes in the extensive-stage only. In the extensive-stage SCLC, both rs1058402 and rs763361 were associated with OS in univariated analysis (under a dominant model, Log Rank P = 0.03 and 0.006, respectively), but only rs1058402 was associated with OS in multivariated analysis (under a dominant model, aHR = 1.75, 95% CI = 1.21–2.53, P = 0.003) (Supplementary Table S2). PFS showed the same trend as OS, although it was not statistically significant (under a dominant model, aHR for rs1058402 = 1.34, 95% CI = 0.94–1.90, P = 0.11; aHR for rs763361 = 0.79, 95% CI = 0.56–1.10, P = 0.16, respectively, Supplementary Table S2).
The effects of the two variants on chemotherapy response and OS did not differ according to clinical variables, such as age, gender, etc., when data were categorized by these factors, except for chemotherapy regimen on chemotherapy response (P value for homogeneity test > 0.05, Supplementary Table S3).
Combined effects of rs1058402 and rs763361 on treatment outcomes
The rs1058402GA/AA and rs763361CC genotypes were associated with worse chemotherapy response and OS. When these genotypes were considered bad genotypes, chemotherapy response was decreased as the number of bad genotypes increased (responder 80.4% with 0 bad genotype, 71.0% with 1 bad genotype, and 61.3% with 2 bad genotypes; aOR = 0.52, 95% CI = 0.33–0.81, Ptrend = 0.004; Table 3). The MST was also significantly decreased as the number of bad genotypes increased (MST = 11.5 months with 0 bad genotype, 9.5 months with 1 bad genotype, and 7.2 months with 2 bad genotypes; aHR = 1.48, 95% CI = 1.19–1.84, Ptrend = 4 × 10−4; Table 3 and Fig. 2C). PFS also significantly decreased as the number of bad genotypes increased (Ptrend = 0.009). In an analysis of only patients with extensive-stage, the combined effect of rs1058402 and rs763361 was still significant in chemotherapy response, OS, and PFS, respectively (aOR = 0.55, 95% CI = 0.33–0.93, Ptrend = 0.03; aHR = 1.55, 95% CI = 1.21–1.98, Ptrend = 6 × 10–4: aHR = 1.28, 95% CI = 1.07–1.59, Ptrend = 0.04, respectively, Supplementary Table S4).
Functional prediction of CD155 rs1058402G > A (A67T)
The rs1058402G > A changes the amino acid of alanine to threonine at codon 67 of CD155. CD155 bound to CD226 of the immune cell and acts as co-stimulatory signal19. We evaluated whether this amino acid change affects the function of CD155 by creating a 3-D structural model using PyMOL (https://pymol.org/2/). As shown in Fig. 3, the alanine to threonine change at codon 67 provides distance shortening to S74 and G73 of CD155 (6.3→2.9 and 4.4→3.3 Å, respectively; Fig. 3B,C). The shortened distance between T67 and S74 created a new hydrogen bond (Fig. 3C). This change resulted in distance lengthening between S74 of CD155 and N116 of CD226 (2.7→6.0 Å) and losing the hydrogen bond (Fig. 3B,C). In addition, CD155 A67T also influenced other important interactions between G70 of CD155 and E185 of CD226 by distance extension (3.3→6.0 Å), resulting in the loss of the hydrogen bond (Fig. 3D). The increased distances and loss of hydrogen bonds will weaken the binding activity between CD155 and CD226.
Structural model of CD226/CD155 (wild-type or A67T mutant) complex. (A) 3-D structure of the mCD226-ecto (Gray) and hCD155-Domain1 (Orange) structures (PDB ID: 6ISC) aligned with the structural model of hCD155-T67 (Green). (B,C) Structural detail of the interaction between CD226-Domain1-N116 and CD155-G73, S74, A67/T67. (D) Structural detail of the interaction between CD226-Domain2-E185 and CD155-wild-type/mutant model G70. The related amino acid residues are shown as sticks (nitrogen atoms: blue, oxygen atoms: red). Inter-atomic distance (in Angstrom) is shown as dotted black lines. The putative polar contacts are shown as red lines. All images of CD226/CD155 (wild-type vs. A67T mutant) were made using PyMOL (https://pymol.org/2/).
Discussion
The immune system is evidently important not only in the development and progression of cancer but also in its treatment4,20. The interaction between tumors and their microenvironment (including immune cells such as tumor infiltrating lymphocytes) can affect chemotherapy response20,21,22. Genetic variants in immune genes can also influence the host’s immune activity, which affects the clinical cancer treatment outcome12,13,23,24. In this study, we investigated the association between variants in immune checkpoint genes and the clinical outcome of SCLC. This is the first study to investigate the effects of genetic variants in immune checkpoint genes on chemotherapy response and prognosis in SCLC. We found that two variants, namely CD155 rs1058402G > A (A67T) and CD226 rs763361C > T (G307S), were significantly associated with chemotherapy response and survival outcomes.
The development of drugs to block immune checkpoints has opened a new era in cancer treatment. Programmed death 1 (PD-1) and PD-L1 inhibitors are used actively in NSCLC treatment10,11. The PD-1/PD-L1 interaction provides inhibitory signals to suppress immune cell responses, so blocking the PD-1/PD-L1 pathway can reactivate immune cells to attack cancer. In NSCLC, the polymorphisms in PD-L1 were reported to affect chemotherapy/immunotherapy response and prognosis12,13,14. However, data on the association between polymorphisms in PD-1/PD-L1 and clinical outcomes in SCLC are limited. In this study, the polymorphisms in PD-1/PD-L1 were not associated with clinical outcomes in SCLC patients (Supplementary Table S1).
CD155, also called PVR or Necl-5, encodes a transmembrane glycoprotein belonging to the immunoglobulin superfamily. CD155 was initially known to play roles in cell adhesion and migration, but recently, its roles in immunology and oncology have been noted25,26. CD226 encodes a glycoprotein expressed on the surface of natural killer (NK) cells, T cells, a subset of B cells, and monocytes, and plays an important role in their activation and inhibition27,28. CD155 on antigen-presenting cells or tumor cells binds to CD226 on T cells and NK cells, which is similar to the interaction between PD-1 and PD-L119,29. The difference is that PD-1/PD-L1 interaction provides an inhibitory signal to suppress T cell response, but CD155/CD226 binding is a co-stimulatory interaction for T cell or NK cell activation19,30. In cancer immunology, CD155 overexpression was reported in several types of human malignancies, including lung adenocarcinoma, and was correlated with unfavorable prognosis31,32,33. CD226 has been reported to be involved in anti-tumor response by regulating NK cells34,35.
The rs1058402G > A is a missense mutation. The rs1058402G > A changes alanine amino acid to threonine at codon 67 of CD155. 3-D structure model showed that changing alanine to threonine at codon 67 shortened the distance to S74/G73 of CD155. The shortened distance made a new hydrogen bond between T67 and S74 of CD155, and this change resulted in distance extension and loss of hydrogen bond between the S74 of CD155 and N116 of CD226. In addition, the structure modification due to rs1058402G > A (A67T) increased the coupling distance between the G70 of CD 155 and E185 of CD226, and resulted in hydrogen bond loss. In the interaction between CD155 and CD226, the S74/G70 of CD155 and N116/E185 of CD226 are important binding points19. In this study, the 1058402G > A (A67T) was significantly associated with worse chemotherapy response and OS in SCLC patients. As shown in the 3-D structural model, the 1058402G > A (A67T) increased the binding point distance between CD155 and CD226. The increased distance and loss of hydrogen bonds lead to a weakening of the binding force of CD155/CD226, thereby reducing the co-stimulatory signal to immune cells. The decreased immune response would had a negative effect on SCLC clinical outcomes. However, further functional studies are needed to determine whether CD155 rs1058402G > A (A67T) affects the CD155/CD226 interaction. This is the first review to report that a CD155 variant was associated with cancer clinical outcomes. This variant may also affect the therapeutic effect of the drug to be developed in the future, considering that CD155 is being studied as a new therapeutic target in tumor immunology26,36.
In the present study, CD226 rs763361C > T was associated with better chemotherapy response and OS in SCLC patients. The rs763361C > T is located in exon 7 encoding the cytoplasmic tail of CD226, which harbors two phosphorylation sites and is a non-synonymous mutation37. The rs763361 C-to-T change results in the replacement of glycine to serine at codon 307. A Gly307Ser substitution believed to affect CD226 expression by changing the phosphorylation or altering RNA splicing by disrupting the exon-splicing silencer sequence37,38. CD226 rs763361C > T is associated with multiple autoimmune diseases, including multiple sclerosis and rheumatoid arthritis37,38,39,40. Regarding cancer, the T allele of rs763361 has been reported to increase the risk of NSCLC in the Chinese Han population41. However, the exact biological function of CD226 rs763361C > T is not well-known, and further researches are needed. The effect on clinical outcomes may be due to other causal variants in LD. The CD226 rs1790947G > T has a strong LD (r2 = 0.91) with CD226 rs763361C > T and is located in the 3′ untranslated region. Using RegulomDB (http://www.regulomedb.org/), the rs1790947G > T was likely expected to affect binding and linked to CD226 expression. RegulomeDB is a novel approach and database, which provides interpretation of regulatory variants in the human genome42. The association patterns between the rs1790947 genotypes and chemotherapy response and OS were similar to that of rs763361 (Supplementary Table S5). Hence, the selected polymorphism was rs763361C > T, but rs1790947G > T could actually affect CD226 expression and clinical outcomes.
In this study, polymorphisms in the CD155 and CD226 was associated with survival outcomes in SCLC patients who received chemotherapy. CD155 and CD226 are one of targets in immuno-oncology43,44. Therefore, variants in CD155 and CD226 may have a greater impact on clinical outcomes with immunotherapy alone or in combination with chemotherapy than with chemotherapy alone. Recently, immunotherapy has been shown to be effective in the treatment of SCLC45. The addition of atezolizumab to etoposide/carboplatin in the first-line treatment of extensive-stage SCLC showed significant longer OS and PFS than chemotherapy alone46. Because CD155 and CD226 are co-stimulatory signal for immune cells, investigating the role of variants in CD155 and CD226 in SCLC patients who received combination immunotherapy and chemotherapy may yield interesting results.
Although a theoretical model for the interactional changes caused by mutations of CD155 and CD226 has been presented, one of limitations is that it has not been confirmed experimentally. Unlike NSCLC, surgical treatment is not usually performed in patients with SCLC. The difficulty of obtaining sufficient tumor tissues for experiments is an obstacle to overcome in SCLC studies. Observational retrospective study at a single center can be another limitation of this study.
In summary, we investigated the association between the variants in immune checkpoint genes and SCLC prognosis. Two variants, namely CD155 rs1058402G > A (A67T) and CD226 rs763361C > T (G307S), were associated with chemotherapy response and OS outcomes. CD155 rs1058402G > A (A67T) seems to influence the interaction between CD155 and CD226. However, further studies are warranted to confirm this finding.
References
Siegel RL, Miller KD, Jemal A.. Cancer statistics, 2018. CA: A Cancer Journal for Clinicians 2018; 68(1):7–30.
Sher T, GK Dy, and AA Adjei. Small cell lung cancer. Mayo Clin Proc 2008; 83(3): p. 355–67.
Wang, S. et al. Survival changes in patients with small cell lung cancer and disparities between different sexes, socioeconomic statuses and ages. Sci. Rep. 7(1), 1339 (2017).
Candeias, S. M. & Gaipl, U. S. The immune system in cancer prevention, development and therapy. Anticancer Agents Med. Chem. 16(1), 101–107 (2016).
Beatty, G. L. & Gladney, W. L. Immune escape mechanisms as a guide for cancer immunotherapy. Clin. Cancer Res. 21(4), 687–692 (2015).
Kalos, M. & June, C. H. Adoptive T cell transfer for cancer immunotherapy in the era of synthetic biology. Immunity 39(1), 49–60 (2013).
Wei, S. C., Duffy, C. R. & Allison, J. P. Fundamental mechanisms of immune checkpoint blockade therapy. Cancer Discov. 8(9), 1069–1086 (2018).
Darvin, P. et al. Immune checkpoint inhibitors: recent progress and potential biomarkers. Exp. Mol. Med. 50(12), 165 (2018).
Ott, P. A., Hodi, F. S. & Robert, C. CTLA-4 and PD-1/PD-L1 blockade: new immunotherapeutic modalities with durable clinical benefit in melanoma patients. Clin. Cancer Res. 19(19), 5300–5309 (2013).
Reck, M. et al. Pembrolizumab versus chemotherapy for PD-L1-positive non-small-cell lung cancer. N. Engl. J. Med. 375(19), 1823–1833 (2016).
Borghaei, H. et al. Nivolumab versus docetaxel in advanced nonsquamous non-small-cell lung cancer. N. Engl. J. Med. 373(17), 1627–1639 (2015).
Lee, S. Y. et al. Functional polymorphisms in PD-L1 gene are associated with the prognosis of patients with early stage non-small cell lung cancer. Gene 599, 28–35 (2017).
Lee, S. Y. et al. PD-L1 polymorphism can predict clinical outcomes of non-small cell lung cancer patients treated with first-line paclitaxel-cisplatin chemotherapy. Sci. Rep. 6, 25952 (2016).
Nomizo, T. et al. Clinical impact of single nucleotide polymorphism in PD-L1 on response to nivolumab for advanced non-small-cell lung cancer patients. Sci. Rep. 7, 45124 (2017).
Eisenhauer EA, et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur. J. Cancer 2009; 45(2):228–47.
Pardoll, D. M. The blockade of immune checkpoints in cancer immunotherapy. Nat. Rev. Cancer 12(4), 252–264 (2012).
Nirschl, C. J. & Drake, C. G. Molecular pathways: coexpression of immune checkpoint molecules: signaling pathways and implications for cancer immunotherapy. Clin. Cancer Res. 19(18), 4917–4924 (2013).
Galluzzi, L., Kroemer, G. & Eggermont, A. Novel immune checkpoint blocker approved for the treatment of advanced melanoma. Oncoimmunology 3(11), e967147 (2014).
Wang, H. et al. Binding mode of the side-by-side two-IgV molecule CD226/DNAM-1 to its ligand CD155/Necl-5. Proc. Natl. Acad. Sci. USA 116(3), 988–996 (2019).
Galluzzi, L. et al. The secret ally: immunostimulation by anticancer drugs. Nat. Rev. Drug. Discov. 11(3), 215–233 (2012).
Blankenstein, T. The role of tumor stroma in the interaction between tumor and immune system. Curr. Opin. Immunol. 17(2), 180–186 (2005).
Zitvogel, L. et al. Immunological aspects of cancer chemotherapy. Nat. Rev. Immunol. 8(1), 59–73 (2008).
DeMichele, A. et al. Host genetic variants in the interleukin-6 promoter predict poor outcome in patients with estrogen receptor-positive, node-positive breast cancer. Cancer Res. 69(10), 4184–4191 (2009).
Schoof, N. et al. Favorable impact of the interleukin-4 receptor allelic variant I75 on the survival of diffuse large B-cell lymphoma patients demonstrated in a large prospective clinical trial. Ann. Oncol. 20(9), 1548–1554 (2009).
Takai, Y. et al. Nectins and nectin-like molecules: roles in cell adhesion, migration, and polarization. Cancer Sci. 94(8), 655–667 (2003).
Gao, J. et al. CD155, an onco-immunologic molecule in human tumors. Cancer Sci. 108(10), 1934–1938 (2017).
Lanier, L. L. NK cell recognition. Annu. Rev. Immunol. 23, 225–274 (2005).
Shibuya, K. et al. CD226 (DNAM-1) is involved in lymphocyte function-associated antigen 1 costimulatory signal for naive T cell differentiation and proliferation. J. Exp. Med. 198(12), 1829–1839 (2003).
Pende, D. et al. Expression of the DNAM-1 ligands, Nectin-2 (CD112) and poliovirus receptor (CD155), on dendritic cells: relevance for natural killer-dendritic cell interaction. Blood 107(5), 2030–2036 (2006).
Lozano, E. et al. The CD226/CD155 interaction regulates the proinflammatory (Th1/Th17)/anti-inflammatory (Th2) balance in humans. J. Immunol. 191(7), 3673–3680 (2013).
Nakai, R. et al. Overexpression of Necl-5 correlates with unfavorable prognosis in patients with lung adenocarcinoma. Cancer Sci. 101(5), 1326–1330 (2010).
Bevelacqua, V. et al. Nectin like-5 overexpression correlates with the malignant phenotype in cutaneous melanoma. Oncotarget 3(8), 882–892 (2012).
Nishiwada, S. et al. Clinical significance of CD155 expression in human pancreatic cancer. Anticancer Res. 35(4), 2287–2297 (2015).
Du, X. et al. CD226 regulates natural killer cell antitumor responses via phosphorylation-mediated inactivation of transcription factor FOXO1. Proc. Natl. Acad. Sci. USA 115(50), E11731-e11740 (2018).
Bottino, C. et al. Identification of PVR (CD155) and Nectin-2 (CD112) as cell surface ligands for the human DNAM-1 (CD226) activating molecule. J. Exp. Med. 198(4), 557–567 (2003).
Kucan, B. P. et al. Targeting PVR (CD155) and its receptors in anti-tumor therapy. Cell Mol. Immunol. 16(1), 40–52 (2019).
Mattana, T. C. et al. CD226 rs763361 is associated with the susceptibility to type 1 diabetes and greater frequency of GAD65 autoantibody in a Brazilian cohort. Mediators Inflamm 2014, 694948 (2014).
Todd, J. A. et al. Robust associations of four new chromosome regions from genome-wide analyses of type 1 diabetes. Nat. Genet. 39(7), 857–864 (2007).
Hafler, J. P. et al. CD226 Gly307Ser association with multiple autoimmune diseases. Genes Immun. 10(1), 5–10 (2009).
Qiu, Z. X. et al. CD226 Gly307Ser association with multiple autoimmune diseases: a meta-analysis. Hum. Immunol. 74(2), 249–255 (2013).
Qiu, Z. X., Peng, Y. & Li, W. M. CD226 gene polymorphisms are associated with non-small-cell lung cancer in the Chinese Han population. Ther. Clin. Risk Manag. 11, 1259–1264 (2015).
Boyle, A. P. et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 22(9), 1790–1797 (2012).
Lepletier, A. et al. Tumor CD155 expression is associated with resistance to anti-PD1 immunotherapy in metastatic melanoma. Clin. Cancer Res. 26(14), 3671–3681 (2020).
Jin, H.S. and M. Ko, CD226(hi)CD8(+) T Cells Are a Prerequisite for Anti-TIGIT Immunotherapy. 2020. 8(7): p. 912–5.
Pakkala, S. and T.K. Owonikoko, Immune checkpoint inhibitors in small cell lung cancer. Journal of Thoracic Disease, 2018: p. S460–7.
Horn, L. et al. First-line atezolizumab plus chemotherapy in extensive-stage small-cell lung cancer. N. Engl. J. Med. 379(23), 2220–2229 (2018).
Acknowledgements
This work has supported by the National Research Foundation of Korea (NRF) Grant funded by the Korea government (MSIT) (No. NRF-2018R1A2B2003038). This research was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (NRF-2017R1D1A3B03034445).
Author information
Authors and Affiliations
Contributions
Conception/Design (J.H.L., S.S.Y., J.Y.P.); Provision of study material or patients (S.S.Y., S.H.C., Y.H.L., H.W.S., J.H.L., S.Y.L., S.I.C., C.H.K., J.Y.P.); Collection of data (J.H.L., S.S.Y., S.Y.K., M.J.H., J.E.C, H.G.K., S.K.D., J.H.K., S.A.B., J.D.Y., S.H.C., Y.H.L., H.W.S.); Analysis and interpretation of Data (J.H.L., S.S.Y., S.Y.K., M.J.H., J.E.C., W.K.L., S.Y.L., S.I.C., C.H.K., J.Y.P.) Manuscript writing: (J.H.L., S.S.Y., J.Y.P.).
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, J.H., Yoo, S.S., Hong, M.J. et al. Impact of immune checkpoint gene CD155 Ala67Thr and CD226 Gly307Ser polymorphisms on small cell lung cancer clinical outcome. Sci Rep 11, 1794 (2021). https://doi.org/10.1038/s41598-021-81260-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-81260-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.