Abstract
Bacterial antibiotic resistance is an important global health threat and the interfaces of antibiotic resistance between humans, animals and the environment are complex. We aimed to determine the associations and overtime trends of antibiotic resistance between humans, animals and water sources from the same area and time and estimate attribution of the other sources to cases of human antibiotic resistance. A total of 125 children (aged 1–3 years old) had stool samples analysed for antibiotic-resistant bacteria at seven time points over two years, with simultaneous collection of samples of animal stools and water sources in a rural Indian community. Newey–West regression models were used to calculate temporal associations, the source with the most statistically significant relationships was household drinking water. This is supported by use of SourceR attribution modelling, that estimated the mean attribution of cases of antibiotic resistance in the children from animals, household drinking water and wastewater, at each time point and location, to be 12.6% (95% CI 4.4–20.9%), 12.1% (CI 3.4–20.7%) and 10.3% (CI 3.2–17.3%) respectively. This underlines the importance of the ‘one health’ concept and requires further research. Also, most of the significant trends over time were negative, suggesting a possible generalised improvement locally.
Similar content being viewed by others
Introduction
Antibiotic resistance of bacteria threatens the core of modern medicine1,2,3 and is an important global health threat. However, the increasing exposure to antibiotics, due to excessive use of antibiotic agents in agriculture and healthcare, contribute to rising bacterial antibiotic resistance1,4. The development of resistance occurs not only in pathogens, but also in commensal bacteria5 such as Escherichia coli (E. coli), which act as a reservoir for resistance genes6. The use of antibiotics not only applies selective pressure towards antibiotic resistance, but also stimulates transfer of resistance determinants between microorganisms4 and the resistance can spread to pathogens7,8.
The complex and interlinking drivers of antibiotic resistance have led international organisations and research communities to recommend a ‘one-health’ approach; a collaboration of health science and food and agriculture professions to attain optimal health for humans, animals and the environment1,9. Antibiotic misuse, poor antibiotic quality, lack of antibiotic stewardship, poor infection control procedures and a lack of surveillance and governance act as risk factors for antibiotic resistance7,10. Risk factors for onward transmission include a lack of sanitation, inadequate clean water access, low standards of infection control, poor access to diagnostics, migration, travel and reduced access to quality antibiotics4. Antibiotic resistance can be transmitted from human to human and from the environment or animals to humans and vice versa11. The pathways are complex, with the environment acting as an important reservoir of resistance12, and direct and indirect contact with animals can transmit resistance7,12,13. Antibiotic residues and resistance have also been found in various water sources, both natural and human-made, across the world14,15,16,17,18,19,20,21,22. In countries and communities where water is sourced from groundwater or untreated surface water there is increased risk of exposure to antibiotic resistance and harm to human health23. It is suggested that drinking water may contribute to development of resistance in humans23 and it is likely to be of more importance in countries with poor sewage and water treatment9.
There is considerable variation of antibiotic resistance globally, but it is a problem that concerns every country24. It is predicted that most deaths in the future from antibiotic resistance will be in Africa and Asia, with India, Nigeria, Indonesia and Russia particularly at risk24. The level of antibiotic resistance in India is increasing25, but there is insufficient surveillance of the resistance trends, with particular scarcity in rural settings25,26,27.
Mathematical modelling has already been used to help understand antibiotic resistance transmission networks between the three domains of the ‘one health’ concept28. However, there are difficulties in applying mathematical models to processes as complicated and inexact as antibiotic resistance. Models need to incorporate factors across the diverse biospheres28 underlining the importance in the ‘one-health’ concept, even when primarily considering human health. SourceR is an open-access statistical model that considers all aspects of ‘one health’ and has prior identified chickens as the most important source of a human campylobacteriosis outbreak29.
There has been substantial work in improving knowledge of drivers and transmission pathways of antibiotic resistance. However, the exact relationships between resistance in humans and animals7,30 and humans and their environment remain unclear7,23. Therefore, we aimed to determine the temporal associations and over time trends of antibiotic resistance of commensal E. coli isolated from children and their household drinking water, source drinking water, wastewater and from animals sharing the same community environment in rural India, over a 2-year period. This was supported by the application of SourceR to assess attribution of resistance.
Results
Overall level of antibiotic resistance
With the exception of colistin, E.coli isolated from all of the sources exhibited resistance to all antibiotics at one point in the study. There was no identified colistin resistance in animals, source drinking water and wastewater sources. The overall range of prevalence of resistance to individual antibiotics at each time point in humans and the measured sources varied greatly from 0.0 to 97.2%, see Supplementary Table 1. An overview of the pattern of resistance (see Supplementary Fig. 1), shows that overall resistance prevalence were higher in human samples (24.7%) than the other sources (10.0%, 11.7%, 14.6% and 15.5% in animals, household drinking water, source drinking water and wastewater respectively).
Temporal association of human antibiotic resistance to that of the other sources
For Newey–West regression model calculated associations between human proportions of antibiotic resistance and the other sources (n = 69 analyses done), 18 combinations of antibiotic and source showed a statistically significant association between the two groups (see Table 1). Colistin was excluded from analysis for the animal, source drinking water and wastewater groups as no resistance was seen in any of the three time-points it was measured. In this analysis, the regression coefficients relate to the statistical percentage increase/decrease in the dependent variable (human) as the percentage of independent variable increases/decreases for each time point. Of the 18 statistically significant associations between human and other source, three were with village animals (ampicillin, nalidixic acid and imipenem), eight with household drinking water sources (ampicillin, cefotaxime, cefepime, tetracycline, imipenem, sulfamethoxazole, extended spectrum beta-lactamase (ESBL) and multi-drug resistance (MDR)), two with source drinking water (ampicillin and imipenem) and five with wastewater (ampicillin, cefotaxime, tigecycline, imipenem and co-trimoxazole). Both ampicillin and imipenem had statistically significant associations between human resistance levels and all other sources measured.
Case attribution of human antibiotic resistance from other sources
The attribution modelling with SourceR estimated that more human isolates were more associated with those in animals, human drinking water and wastewater than source drinking water. The mean of human cases attributable to each environmental source at each time point and location was 38.3 (95% Confidence Interval (CI) 14.3–62.3), 47.7 (CI 10.8–84.6) and 36.7 (CI 9.0–64.4) cases for animals, household drinking water and wastewater respectively, but it was only 0.2 (CI − 0.2–0.67) for source drinking water (Fig. 1). The mean of the percentage of human cases attributable to each source at each time point and location was found to be 12.6% (CI 4.4–20.9%), 12.1% (CI 3.4–20.7%) and 10.3% (CI 3.2–17.3%) for animals, human drinking water and wastewater respectively; for source drinking water it was 0.1% (CI − 0.1–0.2%) (Fig. 2).
Violin Plot of the proportion of Human Cases Attributable to Each Environmental Source. The bold horizontal line for each source represents the mean for each source. The bold horizontal line for each source represents the mean for each source. HDW Household Drinking Water, SDW Source Drinking Water.
Temporal trends of antibiotic resistance in humans and other sources
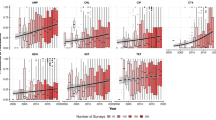
Of the temporal trends of percentage of antibiotic resistance (n = 87), 23 showed a statistically significant trend (see Table 2). Again, colistin was excluded from analysis for the animal, source drinking water and wastewater groups as no resistance was seen in any of the three time points it was measured. The regression coefficient relates to the trend of increase or decrease in resistance as a percentage per time unit (four months). Of these statistically significant trends, two were in human samples (amikacin and ESBL), five in animal samples (ampicillin, cefepime, gentamicin, amikacin and sulfamethoxazole), four in household water samples (ampicillin, gentamicin, tetracycline and ESBL), seven in source drinking water (cefotaxime, nalidixic acid, ciprofloxacin, tetracycline, meropenem, ESBL and MDR) and five in wastewater samples (nalidixic acid, ciprofloxacin, gentamicin, tigecycline and sulfamethoxazole). Of the 23 statistically significant results, five had an increasing and 18 a decreasing trend. In the antibiotic groups that had more than one statistically significant trend, the direction of the regression coefficient was the same in each source: ampicillin, see Fig. 3, (animal and household drinking water, increase), nalidixic acid (source drinking water and wastewater, decrease), ciprofloxacin (source drinking water and wastewater, decrease), gentamicin (animal, household drinking water and wastewater, decrease), amikacin (human and animal, increase), tetracycline (household drinking water and source drinking water, decrease), sulfamethoxazole (animal and wastewater, decrease) and ESBL, see Fig. 4, (human, household drinking water and source drinking water, decrease). The other statistically significant regression coefficients were cefotaxime (source drinking water, decrease), cefepime (animal, increase), tigecycline (wastewater, decrease) and meropenem (source drinking water, decrease).
Discussion
We conducted an examination of antibiotic resistance in human (young children) stools, village animals’ stools, household drinking water, source drinking water and wastewater in a rural Indian community. We have investigated, for the first time, temporal associations between antibiotic resistance in humans and the other sources. Previous studies, including a meta-analysis, have compared prevalence of resistance of simultaneously collected specimens from humans, animals and the environment. However, they have not explored the association between them, often due to methodological limitations30,31,32. Of the analyses of temporal association between sources (n = 69), 18 were statistically significant and the vast majority of these were positively associated. Of the significant associations, most were with household drinking water, suggesting a stronger relationship between human antibiotic resistance and that of their household drinking water compared to the other sources. This is in accordance with the review of antibiotic resistance in drinking water systems that concluded that the consumption of contaminated water may contribute to antibiotic resistance in humans23. However, with regards to the temporal association calculations, our results cannot be used to indicate direction of transmission or direct causality. Nevertheless, given that all sources showed some statistically significant associations to human antibiotic resistance in the Newey–West regression models, further research is needed to greater understand the level and importance of the associations between humans and the animals and environmental sources local to them. Our findings are supported by our previous analysis of the same data, where similar patterns of co-resistance and gene carriage were identified between each of the sources11. As this is the first study that quantitatively analyses temporal association, in this case using Newey–West regression modelling, between humans and environmental sources, further research is also required to investigate whether our results are repeatable and transfer to other locations, particularly where humans have different interactions with animals and their environment. A study from Europe found a mirroring of resistance patterns in wastewater samples and clinical samples in the same area, but no measure of association was made14. Furthermore, this study was from wastewater plants rather than open wastewater as we studied, which makes direct comparison difficult.
The findings of the calculations of temporal association were supported by the results when we applied the attribution model, SourceR. The case attribution results estimated that most human cases of antibiotic resistance were attributed to household drinking water per time point and location (mean = 47.7 (CI 10.8–84.6) cases of antimicrobial resistance in each village at each time point). This was closely followed by animal (38.3, CI 14.3–62.3) and wastewater (36.7, CI 9.0–64.4), however source drinking water attributed to only 0.2 (CI − 0.2–0.67) cases. A similar pattern of attribution was seen when considering the proportion of human cases attributed to each source per time point and location. However, the source with the highest percentage of cases attributed was animals (12.6% (CI 4.4–20.9%)), then household drinking water and wastewater (12.1% (CI 3.4–20.7%)) and 10.3% (CI 3.2–17.3%) respectively). Again, the lowest level of case attribution was seen in source drinking water (0.1% (CI − 0.1–0.2%)). It is unclear, however, why there are so few human cases of antibiotic resistance attributed to source drinking water, when compared to the other sources. In the measures of temporal association, the lowest amount of statistically significant associations was found between humans and source drinking water, but the difference between the same sources was not of the same scale as that seen in case attribution calculations. The overall results, however, do suggest that the relationship between human antibiotic resistance and their drinking water, in this setting, is more closely related to how humans interact with their drinking water than the source of the water itself. This is also the first study, to our knowledge, that has applied an attribution modelling to cases of human antibiotic resistance from environmental sources. Therefore, more research is required to further understand and clarify the contribution of each of these sources to human antibiotic resistance. However, this work does provide a promising start to use of mathematical modelling to understand and quantify the effects of environmental sources on human antibiotic resistance.
The level of temporal association, determined by Newey–West regression models, between the children and their environment, and the level of case attribution, estimated by SourceR, of each source to antibiotic resistance in children, support the concept of ‘one-health’. They highlight the overlapping nature of levels of antibiotic resistance in humans and that of the animal and environmental biospheres. The ‘one-health’ concept should therefore be used to inform interventions and policies to combat antibiotic resistance, as is already suggested by international organisations1,2,33,34,35. Several previous studies have shown evidence of transmission between these distinct disciplines23,36,37,38,39,40,41,42 and the theories are widely accepted and integrated into international plans for managing antibiotic resistance1,7,9,43. However, to date, this is the first study that has directly investigated the relationship between the three disciplines over time. Furthermore, we were able to analyse this relationship, with real-life observations, over real-time. To further understand these associations, more research that incorporates the observation over time is needed, as we showed a level of fluctuation between each time point. When observing the trend for ampicillin (see Fig. 3), there was an observable spike in resistance to all sources at time point 6 (Summer 2016) that we cannot explain. This may have occurred for many reasons, including, a change in antibiotic prescription, use or disposal in the area, or this may have been caused by an event or multiple events, such as the Simhastha Kumbh Mela; a religious pilgrimage and festival in the nearby city of Ujjain that occurred a few months prior to this collection point, but it is unclear what, if any, relationship that may have had.
When considering the temporal trends in antibiotic resistance, our results were mixed. We found, perhaps surprisingly, that where there was a statistically significant trend, they were more likely to be negative, suggesting a possible overall downward trend within the community. However, most of our results did not show a clear trend over time; 23 out of 87 regression coefficients were found to be statistically significant and relatively evenly spread out within the sources. It is also interesting that 18 of the 23 significant results came from just 7 antibiotics (including ESBL) and that within these individual antibiotics the direction of the trend was always only positive or negative. This suggests there has been a possible overall decrease in resistance in nalidixic acid, ciprofloxacin, tetracycline, sulfamethoxazole and ESBL, but an increase in resistance in ampicillin and amikacin. Our findings are similar of that found in another study from France, which found that most of the trends in antibiotic resistance in humans, pets and food-animals were either stationary or negative30. However, the consensus of the literature and guidance from international organisations is that antibiotic resistance is increasing globally1,3,4,24,44. More specifically, most publications suggest antibiotic resistance is increasing in India25,45. Our results, therefore, cannot be used in isolation to determine antibiotic resistance trends across the country, region or globally, but add to the existing evidence base. Fortunately, in the future, the quality of data, regarding over time trends of resistance, is likely to be improved by the implementation of the Global Antimicrobial Resistance Surveillance System from the World Health Organization enabling continuous monitoring26,46, although surveillance across all aspects of one-health and community data is required. Another point of consideration when comparing to other studies was that our study area was a rural community, with children as the study population. This may account for some of the differences in results, as most research is from tertiary centres and in adults47,48,49,50; for example, our participants were clinically well and assumed to be antibiotic naive, whereas adults and hospital patients are more likely to be or have been exposed to antibiotics. Although paediatric antibiotic resistance is itself of importance47, it is unclear if these results will remain similar for different age groups.
It is already established that antibiotic resistance increases with antibiotic use1,4, so an understanding of prescribing patterns is required to control the spread of antibiotic resistance6. The strength of selection for resistance differs depending on the balance of the cost and benefits of resistance28. It seems that antibiotic resistance confers a cost to the bacterium, which is expressed as diminished competitive ability when antibiotics are absent51. Therefore, where certain antibiotics have a reduced entry into the transmission pathways, a decline in resistance specific to that antibiotic may be seen28,52. Therefore, to understand the results of resistance patterns and trends better, they need to be studied alongside the local antibiotic prescription behaviours6. This is important in India, where antibiotic prescription practices vary across the country and require greater stewardship53. This is not within our remit, but briefly, the local rates of antibiotic prescription are high, across several hospital departments and while there is variation of type of antibiotic prescribed, they tend to be broad-spectrum and often not indicated54,55,56,57. Additionally, informal healthcare providers also prescribe high levels of antibiotics in the area58. Further, there is high antibiotic use in agriculture in India, although the exact figures for usage is unknown59. However, antibiotic consumption alone does not explain variations in antibiotic resistance, so other factors need considering from both a microbiological and from a one-health behavioural aspect, such as interaction of determinants of resistance, bystander selection26,60 or disposal of antibiotics. In keeping with ‘one health’, it is also important to consider agricultural practices, with regards to both antibiotic use, animal waste and subsequent wastewater flow61,62,63, with these factors highlighted with regards to antibiotic resistance in India59. These factors further underline the importance of multi-disciplinary and -sectorial work with close collaboration as part of a concerted effort59.
One of the strengths of this study is the collection of data over an extended period. It incorporates time into the analysis and recognises that the level of resistance at one point is dependent on previous findings. Temporal analysis, such as time series, has been used in previous studies into antibiotic resistance, but they have largely focused on comparing resistance to antibiotic use or the effects of antibiotic stewardship programmes64,65,66,67,68. However, none to date have analysed the association of resistance between different sources, incorporating temporality. The data was collected simultaneously and collated homogenously, which allowed the study of associations between the sources. Previous studies of antibiotic resistance that have included all three components of the ‘one health’ concept have been limited by not collecting data in this way, hence these studies only describe patterns and trends30,31. However, there are weaknesses within this study design as well. Whilst we believe this study offers a novel approach to attribution of human antibiotic resistance from environmental sources, this could have been strengthened further by incorporating genotypes into the model, possibly allowing a more precise measure of transmission, however this was not possible within this study. This is, in part, a reflection of the resources available, with the setting being a lower middle income country, and within this setting the utilisation of this tool may be more likely to be repeated with less expensive phenotypic analysis rather than genotypic. Further, this study phenotypically determined antibiotic resistance, as in clinical practice the greater interest is in the antibiotics and the groups of antibiotics to which there are resistance, rather than the mechanism of resistance. This is the first use of SourceR to calculate attribution for antibiotic resistance and using phenotypic measures of resistance rather than genotypic. It therefore may be prudent to regard the output as a suggestion of relative association between sources and humans of case attribution levels rather than an absolute calculation, this is highlighted by the variability within the results. Further, SourceR is designed for use with cross-sectional data, from which causality and directionality cannot be reliably determined. The number of samples collected at each time varied depending on the source and number of samples collected where E. coli was isolated. In total, 41 children were not available for all seven time points and samples were not received for a total of 63 times, but this represents only a small amount of the planned total samples for children (n = 875), with only 7.2% of samples not received. While power calculations for linear regressions and ANOVAs cannot be used in time series data69, based on simple statistics, the proportion of resistance is likely to be less accurate where smaller numbers of E. coli were isolated. However, Newey–West regression modelling of temporal data uses ordinary least squares70 which is robust and works well for small sample sizes71. A high number of statistical tests were performed; thus, we cannot rule out that some of the results are due to chance. However, the number of statistically significant results from the Newey–West calculations were higher than that expected by chance alone with a statistically significant p-value set at 0.05 (26.1% of calculations of association were significant and 26.4% of trends). No correction of multiple statistical tests has been done, as the analysis done here was multiple use of one simple test and it is the individual tests and their results are of importance72. The Kirby-Bauer disc diffusion method is robust for antibiotic susceptibility testing for all included antibiotics, other than colistin27. This was accounted for by simultaneous detection of the mcr-1 resistance gene on all isolates11,27. Colistin was also included late in the study, with collection at three time points only and with a smaller number of samples. However, the aim of this inclusion was to get a baseline for colistin resistance in this community and as such limited analysis of over time trends and associations have been done and it was excluded from the attribution modelling.
In conclusion, there was an observable temporal association between human antibiotic resistance prevalence and that of the animals and environmental sources in proximity. The closest of these relationships appears to be between human antibiotic resistance levels and household drinking water. This is supported by case attribution modelling, with the highest levels of attribution to human cases of antibiotic resistance seen in household drinking water and animals. Our results add valuable data to the overall picture of over time trends, but perhaps more importantly we provide a platform to study the relationships between the resistance levels in the three pillars of the ‘one-health’ concept. Where there was a statistically significant over time trend in resistance pattern, most of them showed a negative trend. The exceptions were in ampicillin and amikacin, which showed positive trends in more than one of the sources studied.
Methods
Study design
This prospective cohort study collected data from different sources: child stool, animal (cattle, hen, dog, goat and horse) stool, household drinking water, source drinking water and wastewater sources. Both the human and animal stool samples were representative of commensal, enteric E. coli, due to clinical absence of infection. Samples were collected at regular, four-month intervals, for two years, totalling 7 time points.
Study setting
The data was collected in the rural Palwa demographic surveillance site of Ruxmaniben Deepchand Gardi Medical College (RDGMC) Ujjain District, Madhya Pradesh, India. Madhya Pradesh, centrally located in India, is the second largest state and approximately 75% of the inhabitants live in rural areas27. As many as 80% of the healthcare workers, both formal and informal, are concentrated in one village within the surveillance site6, hereafter referred to as the central village.
Sampling selection
Further details of sampling are outlined in previous studies6,11,27. In brief, the six villages that met the inclusion criteria were purposively selected from 60 villages. The inclusion criteria were: to be within 5 km of the central village; total village population of 500 people; more than 15 children aged between 1 and 3 years old available in the village and a transport time of less than 45 min to the RDGMC Central Research Laboratory from each village. Further inclusion criteria for the human participants were: a child aged between 1 and 3 years old; living in the village for at least the past year; planning on living in the village for the three upcoming years and willing to participate in the study. Subsequently, simple random sampling was performed to select a total sample of 125 children.
Data collection
Details of the collection, transport and laboratory methods are outlined in previous studies6,11,27. The data was collected between August 2014 and September 2016. Sample collection was organised by trained research assistants. The data was collected in the three seasons experienced in the study setting; namely summer (March to June), monsoon (late June to September) and post-monsoon/winter (October to February). Each selected child provided a stool sample and a sample from their household’s drinking water storage vessel, which was collected by their parents or guardians. In each village, a sample of drinking water was collected from two source drinking water sites and a wastewater sample was collected from two discharge sites. The source drinking water sites are communal drinking water collection points in the village that are typically used to fill the household drinking water storage vessel. Stool samples from five animals (cattle, hen, dog, goat and horse) were collected in each village. The animals do not live within the households, but all commonly share the environment with the children.
Details of the preparation of the samples are also outlined in prior studies6,73. A total of six E. coli colonies were isolated from each sample. Antibiotic susceptibility testing was performed using the Kirby-Bauer disc diffusion method74, as per the current Clinical and Laboratory Standards Institute (CLSI) guidelines75. Antibiotics were chosen following the antibiotics used in the study area for gram-negative coliform infections and the CLSI guidelines75. The following antibiotics were chosen: ampicillin, cefotaxime, ceftazidime, cefepime, nalidixic acid, ciprofloxacin, nitrofurantoin, gentamicin, amikacin, tetracycline, tigecycline, imipenem, meropenem, co-trimoxazole, sulfamethoxazole and colistin. Measurements of colistin resistance were done by detecting mcr-1 gene and by disc diffusion. These measurements were taken for the three last time points as colistin is not regularly used in this setting but was later included in order to have a baseline measurement of community resistance in this important, last-line antibiotic. Measurements of mono-resistance (resistance to one antibiotic) and multi-drug resistance (MDR) (resistance involving three or more antibiotics of three different groups) were undertaken. The intermediate isolates were then grouped together with resistant isolates and classified as resistant.
If full collection of data was not possible, for example if the participant left the study area, the household was excluded from the study. However, if one data collection sample was not received from the caregiver on the day of collection, they were marked as ‘Sample Not Received’ for that collection round but continued in the study. If the sample was not received or it was not possible to isolate E. coli in one of the samples then they were excluded from the analysis of the results of that time point.
Statistical analysis
The first analysis of the trends and relationships was done using Stata Version 16.1 (StataCorp. College Station, TX, USA) and Excel (Microsoft Corp., Redmond, WA, USA). Firstly, the proportions and percentages of resistant isolates, per data collection point, per source, to each antibiotic were calculated. A measurement of the mean proportion of resistant isolates was made for each source, for each antibiotic. The data was grouped ordinally, by sample collection date, to create a data series of the percentage of resistance in each antibiotic at each of the seven times samples were collected. Newey–West regression modelling of temporal data70 was used to test the temporal trend of percentage of resistance for each antibiotic, for each source, against time. Newey–West can be used to estimate the covariance matrix of parameters of a regression where typical assumptions of a regression-model are not true. In this instance, it was used because the level of resistance at a time point is dependent on the previous time point. By using this method we incorporate time into the analysis. The command “tsset” was used in Stata to identify the collection point as an ordinal time variable for each Newey–West regression model. The dependent variable was the percentage of resistance at each time point for the antibiotic and source being calculated, with a separate model calculated for each antibiotic and source, and the independent variable was the collection point. A lag of one time point was used for each model. A total of 87 models were used to calculate the overtime trends of antibiotic resistance. The same method was used to assess the temporal association between proportion of resistance in human samples and the other sources individually. The collection point was used as the time variable in this model and the lag was one time point for each model. The dependent variable was the percentage of human antibiotic resistance to the antibiotic being calculated at each time point. The independent variable was of the percentage of resistance to the specified antibiotic at each time point for either the animal, household drinking water, source drinking water or wastewater sources. A separate model was created for each of these independent variables for each antibiotic. Thus, a total of 69 individual models were used to estimate the association of human antibiotic resistance to that of the other sources. Fitting of the attribution model was done via RStudio version. 4.0.2, using sourceR, which is available from the Comprehensive R Archive Network and released under a GPL-3 licence. The antibiotics were grouped as per their class to represent phenotypic resistance, this was defined as “Type” in the existing model and similarly “Cases” were defined as human antibiotic resistance29. Indicative priors were calculated with a Dirichlet process where the concentration parameter was one. A total of 1000 iterations were used for the MCMC chain, with a burn-in of 10,000 and thinning of 500, as used previously in SourceR modelling29. The measurements of human cases attributable to each environmental source and the proportion of cases attributable to each source were extracted from the model for each time point and location and expressed as medians. Means and 95% confidence intervals of these results were calculated as well as percentages of attribution, giving an overall figure of human cases attributable to each source. Colistin was excluded from the modelling, as due to incomplete collection it was not compatible with the rest of the model. Measures of multi-drug resistance were also excluded from the modelling, as it is not a measure of resistance to a specific antibiotic or group of antibiotics.
Ethical considerations.
Ethical review number: 2013/07/17-311 (Institutional Ethics committee RDGMC, Ujjain, India). Oral and written informed consent was obtained for each participant from a parent or legal guardian. There were no harmful or invasive procedures performed in this study. All children identified as needing medical care were referred to and treated by paediatric services in collaboration with the RDGMC.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
WHO, Global Action Plan on Antimicrobial Resistance, The World Health Organization https://apps.who.int/iris/bitstream/handle/10665/193736/9789241509763_eng.pdf?sequence=1 (2015).
The World Bank Operational Framework for Strengthening Human, Animal, and Environmental Public Health Systems at their Interface, The World Bank http://documents.worldbank.org/curated/en/961101524657708673/pdf/122980-REVISED-PUBLIC-World-Bank-One-Health-Framework-2018.pdf (2018).
Cecchini, M., Langer, J. & Slawomirski, L. Antimicrobial Resistance in G7 Countries and Beyond: Economic Issues, Policies and Options for Action, OECD http://www.oecd.org/els/health-systems/Antimicrobial-Resistance-in-G7-Countries-and-Beyond.pdf (2015).
Holmes, A. H. et al. Understanding the mechanisms and drivers of antimicrobial resistance. Lancet 387(10014), 176–187 (2016).
Macfarlane, S. Antibiotic treatments and microbes in the gut. Environ. Microbiol. 16(4), 919–924 (2014).
Stålsby Lundborg, C. et al. Protocol: A “One health” two year follow-up, mixed methods study on antibiotic resistance, focusing children under 5 and their environment in rural India. BMC Public Health 15, 1321 (2014).
Laxminarayan, R. et al. Antibiotic resistance-the need for global solutions. Lancet. Infect. Dis 13(12), 1057–1098 (2013).
Juhász, J. et al. Colonization dynamics of multidrug-resistant klebsiella pneumoniae are dictated by microbiota-cluster group behavior over individual antibiotic susceptibility: A metataxonomic analysis. Antibiotics 10, 268 (2021).
McEwen, S. A. & Collignon, P. J. Antimicrobial resistance: A one health perspective. Microbiol. Spectrum. 6(2), 6 (2018).
Aslam, A. et al. Evidence of the practice of self-medication with antibiotics among the lay public in low- and middle-income countries: A scoping review. Antibiotics (Basel) 9(9), 597 (2020).
Purohit, M. R. et al. Antibiotic resistance in an indian rural community: A “one-health” observational study on commensal coliform from humans, animals, and water. Int. J. Environ. Res. Public Health 14(4), 386 (2017).
Economou, V. & Gousia, P. Agriculture and food animals as a source of antimicrobial-resistant bacteria. Infect. Drug Resist. 2015(8), 49–61 (2015).
Winokur, P. L., Vonstein, D. L., Hoffman, L. J., Uhlenhopp, E. K. & Doern, G. V. Evidence for transfer of CMY-2 AmpC beta-lactamase plasmids between Escherichia coli and Salmonella isolates from food animals and humans. Antimicrob. Agents Chemother. AAC. 45(10), 2716–2722 (2001).
Pärnänen, K. M. M. et al. Antibiotic resistance in European wastewater treatment plants mirrors the pattern of clinical antibiotic resistance prevalence. Sci. Adv. 5(3), eaau9124 (2019).
Manaia, C. M. et al. Antibiotic resistance in wastewater treatment plants: Tackling the black box. Environ. Int. 115, 312–324 (2018).
Lorenzo, P. et al. Antibiotic resistance in urban and hospital wastewaters and their impact on a receiving freshwater ecosystem. Chemosphere 206, 70–82 (2018).
Chen, B. et al. Complex pollution of antibiotic resistance genes due to beta-lactam and aminoglycoside use in aquaculture farming. Water Res. 134, 200–208 (2018).
Yuan, K. et al. Occurrence of antibiotic resistance genes in extracellular and intracellular DNA from sediments collected from two types of aquaculture farms. Chemosphere 234, 520–527 (2019).
Diwan, V. et al. Antibiotics and antibiotic-resistant bacteria in waters associated with a hospital in Ujjain, India. BMC Public Health 10, 414 (2010).
Diwan, V. et al. Seasonal variations in water-quality, antibiotic residues, resistant bacteria and antibiotic resistance genes of escherichia coli isolates from water and sediments of the kshipra river in Central India. Int. J. Environ. Res. Public Health. 15(6), 1281 (2018).
Harada, K. Antibiotic residue in environmental water in Vietnam. Yakugaku Zasshi 138(3), 271–275 (2018).
Wilson, M. E. The afterlife of antibiotics. J. Travel Med. 27(1), taz102 (2020).
Sanganyado, E. & Gwenzi, W. Antibiotic resistance in drinking water systems: Occurrence, removal, and human health risks. Sci. Total Environ. 669, 785–797 (2019).
O'Neill, J. Antimicrobial Resistance: Tackling a crisis for the health and wealth of nations, Review on Antimicrobial Resistance https://amr-review.org/sites/default/files/AMR%20Review%20Paper%20-%20Tackling%20a%20crisis%20for%20the%20health%20and%20wealth%20of%20nations_1.pdf (2014).
Wattal, C. & Goel, N. Tackling antibiotic resistance in India. Expert Rev. Anti Infect. Ther. 12(12), 1427–1440 (2014).
Veeraraghavan, B. & Walia, K. Antimicrobial susceptibility profile & resistance mechanisms of Global Antimicrobial Resistance Surveillance System (GLASS) priority pathogens from India. Indian J. Med. Res. 149(2), 87–96 (2019).
Purohit, M. R., Lindahl, L. F., Diwan, V., Marrone, G. & Lundborg, C. S. High levels of drug resistance in commensal E. coli in a cohort of children from rural central India. Sci. Rep. 9(1), 6682 (2019).
Knight, G. M. et al. Mathematical modelling for antibiotic resistance control policy: Do we know enough?. BMC Infect. Dis. 19(1), 1011 (2019).
Miller, P., Marshall, J., French, N. & Jewell, C. sourceR: Classification and source attribution of infectious agents among heterogeneous populations. PLoS Comput. Biol. 13(5), e1005564 (2017).
Bourély, C. et al. Trends in antimicrobial resistance among Escherichia coli from defined infections in humans and animals. J. Antimicrob. Chemother. 75(6), 1525–1529 (2020).
Boonyasiri, A. et al. Prevalence of antibiotic resistant bacteria in healthy adults, foods, food animals, and the environment in selected areas in Thailand. Pathogens Glob. Health. 108(5), 235–245 (2014).
Pormohammad, A., Nasiri, M. J. & Azimi, T. Prevalence of antibiotic resistance in Escherichia coli strains simultaneously isolated from humans, animals, food, and the environment: A systematic review and meta-analysis. Infect. Drug Resist. 12, 1181–1197 (2019).
FAO et al. Contributing to One World, One Health: A Strategic Framework for Reducing Risks of Infectious Diseases at the Animal–Human–Ecosystems Interface, Food and Agricultural Organization of the United Nations , Office International des Epizooties, World Health Organization, United Nations System Influenza Coordination, United Nations Childrens Fund, The World Bank https://www.oie.int/doc/ged/D5720.PDF (2008).
FAO, OIE & WHO The FAO-OIE-WHO Collaboration: Sharing responsibilities and coordinating global activities to address health risks at the animal-humans-ecosystems interface: A Tripartite Concept Note, FAO, OIE, WHO https://www.who.int/influenza/resources/documents/tripartite_concept_note_hanoi_042011_en.pdf (2010).
FAO, OIE & WHO Memorandum of Understanding Between The United Nations Food and Agricultural Organization and The World Organization for Animal Health and The World Health Organization: Regarding cooperation to combat health risks at the animal-humans-ecosystems interface in the context of the 'one health' approach and including antimicrobial resistance, FAO, OIE, WHO https://www.who.int/zoonoses/MoU-Tripartite-May-2018.pdf (2018).
Grönthal, T. et al. Sharing more than friendship - transmission of NDM-5 ST167 and CTX-M-9 ST69 Escherichia coli between dogs and humans in a family, Finland, 2015. Euro Surv. Bull. européen sur les maladies transmissibles Eur. Commun. Dis. Bull. 23(27), 1700497 (2018).
Salinas, L. et al. Diverse commensal escherichia coli clones and plasmids disseminate antimicrobial resistance genes in domestic animals and children in a semirural community in ecuador. Sphere 4(3), e00316-e319 (2019).
Lazarus, B., Paterson, D. L., Mollinger, J. L. & Rogers, B. A. Do human extraintestinal Escherichia coli infections resistant to expanded-spectrum cephalosporins originate from food-producing animals? A systematic review. Clin. Infect. Dis. 60(3), 439–452 (2015).
Johnson, J. R., Clabots, C. & Kuskowski, M. A. Multiple-host sharing, long-term persistence, and virulence of Escherichia coli clones from human and animal household members. J. Clin. Microbiol. JCM. 46(12), 4078–4082 (2008).
Johnson, J. R., Miller, S., Johnston, B., Clabots, C. & Debroy, C. Sharing of Escherichia coli sequence type ST131 and other multidrug-resistant and Urovirulent E. coli strains among dogs and cats within a household. J. Clin. Microbiol. JCM. 47(11), 3721–3725 (2009).
Liu, Y., Han, C., Chen, Z., Guo, D. & Ye, X. Relationship between livestock exposure and methicillin-resistant Staphylococcus aureus carriage in humans: A systematic review and dose-response meta-analysis. Int. J. Antimicrob. Agents. 55(1), 105810 (2020).
Liu, Y.-Y. et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: A microbiological and molecular biological study. Lancet. Infect. Dis 16(2), 161–168 (2016).
Walsh, T. R. A one-health approach to antimicrobial resistance. Nat. Microbiol. 3(8), 854–855 (2018).
Friedman, N. D., Temkin, E. & Carmeli, Y. The negative impact of antibiotic resistance. Clin. Microbiol. Infect. 22(5), 416–422 (2016).
Gajamer, V. R. et al. The first report of phenotypic and molecular characterization of extended-spectrum beta-lactamase-producing uropathogens in Sikkim and Darjeeling hills of India. Microb. Drug Resist. 24(9), 1284–1288 (2018).
WHO, Global Antimicrobial Resistance Surveillance System: Manual for early implentation, WHO https://apps.who.int/iris/bitstream/handle/10665/188783/9789241549400_eng.pdf?sequence=1 (2015).
Dharmapalan, D., Shet, A., Yewale, V. & Sharland, M. High reported rates of antimicrobial resistance in indian neonatal and pediatric blood stream infections. J. Pediatric Infect. Dis. Soc. 6(3), e62–e68 (2017).
Jajoo, M. et al. Alarming rates of antimicrobial resistance and fungal sepsis in outborn neonates in North India. PLoS ONE 13(6), e0180705 (2018).
Makkar, A. et al. Epidemiological profile and antimicrobial resistance pattern of enteric fever in a tertiary care hospital of north india: A seven year ambispective study. Acta Medica (Hradec Kralove) 61(4), 125–130 (2018).
Sekar, R., Mythreyee, M., Srivani, S. & Amudhan, M. Prevalence of antimicrobial resistance in Escherichia coli and Klebsiella spp. in rural South India. J. Glob. Antimicrob. Resist. 5, 80–85 (2016).
Vogwill, T. & MacLean, R. C. The genetic basis of the fitness costs of antimicrobial resistance: A meta-analysis approach. Evol. Appl. 8(3), 284–295 (2015).
Andersson, D. I. & Hughes, D. Antibiotic resistance and its cost: Is it possible to reverse resistance?. Nat. Rev. 8(4), 260–271 (2010).
Singh, S. K. et al. Variations in antibiotic use across India: Multi-centre study through Global Point Prevalence survey. J. Hosp. Infect. 103(3), 280–283 (2019).
Landstedt, K., Sharma, A., Johansson, F., Stålsby Lundborg, C. & Sharma, M. Antibiotic prescriptions for inpatients having non-bacterial diagnosis at medicine departments of two private sector hospitals in Madhya Pradesh, India: A cross-sectional study. BMJ Open 7(4), e012974 (2017).
Hauge, C., Stålsby Lundborg, C., Mandaliya, J., Marrone, G. & Sharma, M. Up to 89% of neonates received antibiotics in cross-sectional Indian study including those with no infections and unclear diagnoses. Acta Pædiatrica. 106(10), 1674–1683 (2017).
Sharma, M., Eriksson, B., Marrone, G., Dhaneria, S. & Lundborg, C. S. Antibiotic prescribing in two private sector hospitals; one teaching and one non-teaching: A cross-sectional study in Ujjain, India. BMC Infect. Dis. 12, 155 (2012).
Sharma, M. et al. Antibiotic prescribing in women during and after delivery in a non-teaching, tertiary care hospital in Ujjain, India: A prospective cross-sectional study. J. Pharm. Policy Pract. 6, 9 (2013).
Khare, S. et al. Antibiotic prescribing by informal healthcare providers for common illnesses: A repeated cross-sectional study in Rural India. Antibiotics. 8(3), 139 (2019).
Taneja, N. & Sharma, M. Antimicrobial resistance in the environment: The Indian scenario. Indian J Med Res. 149(2), 119–128 (2019).
Gajdács, M., Paulik, E. & Szabó, A. Knowledge, attitude and practice of community pharmacists regarding antibiotic use and infectious diseases: A cross-sectional survey in hungary (KAPPhA-HU). Antibiotics (Basel) 9(2), 41 (2020).
Obimakinde, S., Fatoki, O., Opeolu, B. & Olatunji, O. Veterinary pharmaceuticals in aqueous systems and associated effects: An update. Environ. Sci. Pollut. Res. Int. 24(4), 3274–3297 (2017).
Zhi, S., Shen, S., Zhou, J., Ding, G. & Zhang, K. Systematic analysis of occurrence, density and ecological risks of 45 veterinary antibiotics: Focused on family livestock farms in Erhai Lake basin, Yunnan, China. Environ. Pollut. 267, 115539 (2020).
Samreen Ahmad, I., Malak, H. A. & Abulreesh, H. H. Environmental antimicrobial resistance and its drivers: A potential threat to public health. J. Glob. Antimicrob. Resist. 27, 101–111 (2021).
Davey, P. et al. Interventions to improve antibiotic prescribing practices for hospital inpatients. Cochrane Library. 2, CD003543 (2017).
Athanasiou, C. I. & Kopsini, A. Systematic review of the use of time series data in the study of antimicrobial consumption and Pseudomonas aeruginosa resistance. J. Glob. Antimicrob. Resist. 15, 69–73 (2018).
Zeng, S. et al. Time series analysis of antibacterial usage and bacterial resistance in China: Observations from a tertiary hospital from 2014 to 2018. Infect. Drug Resist. 12, 2683–2691 (2019).
Peragine, C. et al. Impact of a comprehensive antimicrobial stewardship program on institutional burden of antimicrobial resistance: A 14-year controlled interrupted time series study. Clin. Infect. Dis. 71(11), 2897–2904 (2020).
Zou, Y. M. et al. Trends and correlation of antibacterial usage and bacterial resistance: Time series analysis for antibacterial stewardship in a Chinese teaching hospital (2009–2013). Eur. J. Clin. Microbiol. Infect. Dis. 34(4), 795–803 (2015).
Beard, E. et al. Understanding and using time series analyses in addiction research. Addiction 114(10), 1866–1884 (2019).
Newey, W. K. & West, K. D. A simple, positive semi-definite, heteroskedasticity and autocorrelation consistent covariance matrix. Econometrica 55(3), 703–708 (1987).
Judkins, D. R. & Porter, K. E. Robustness of ordinary least squares in randomized clinical trials. Stat. Med. 35(11), 1763–1773 (2016).
Armstrong, R. A. When to use the Bonferroni correction. Ophthalmic Physiol. Opt. 34(5), 502–508 (2014).
Hope, M. C. & Neill, A. H. The use of the membrane filter technique for testing water supplies in the field. Public Health Rep. 1878 71(11), 1093–1096 (1956).
Bauer, A. W., Kirby, W. M., Sherris, J. C. & Turck, M. Antibiotic susceptibility testing by a standardized single disk method. Am. J. Clin. Pathol. 45(4), 493–496 (1966).
CLSI. Performance Standards for Antimicrobial Susceptibility Testing. Twenty-Third Informational Supplement (CLSI Document M100-S23). Clin. Lab. Stand. Inst. 33, 1–184 (2013).
Funding
Open access funding provided by Karolinska Institute.
Author information
Authors and Affiliations
Contributions
J.M. was responsible for the formulation of a part of the study for a master’s thesis, performed the statistical analysis, ran the SourceR model and prepared the manuscript. M.R.P. began the formulation of the study, was the responsible person for the microbiological work and analysis and helped prepare the manuscript. M.R.P., V.D. and C.S.L. were involved in conceptualising the study and study design. G.M. provided help with statistical analysis and helped prepare the manuscript. C.P.J. and J.M.R. provided assistance with use of SourceR and the output of the model. V.D. was responsible for data collection. C.S.L. initiated the project and is the principal investigator of the project to which this study belongs.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mitchell, J., Purohit, M., Jewell, C.P. et al. Trends, relationships and case attribution of antibiotic resistance between children and environmental sources in rural India. Sci Rep 11, 22599 (2021). https://doi.org/10.1038/s41598-021-01174-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-01174-w
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.